Crosstalk of Oxidative Phosphorylation-Related Subtypes, Establishment of a Prognostic Signature and Immune Infiltration Characteristics in Colorectal Adenocarcinoma
Abstract
Simple Summary
Abstract
1. Introduction
2. Materials and Method
2.1. Data Acquisition
2.2. Identification of OXPHOS-Related Prognostic Signature
2.3. Functional and Gene Set Enrichment Analysis
2.4. Construction of Risk Score Signature
2.5. Cell Culture
2.6. Quantitative RT-PCR
2.7. Tumor Microenvironment (TME) Analysis
2.8. Mutational Analysis
2.9. Immunophenoscore (IPS) and Chemotherapy Analysis
2.10. Statistical Analysis
3. Results
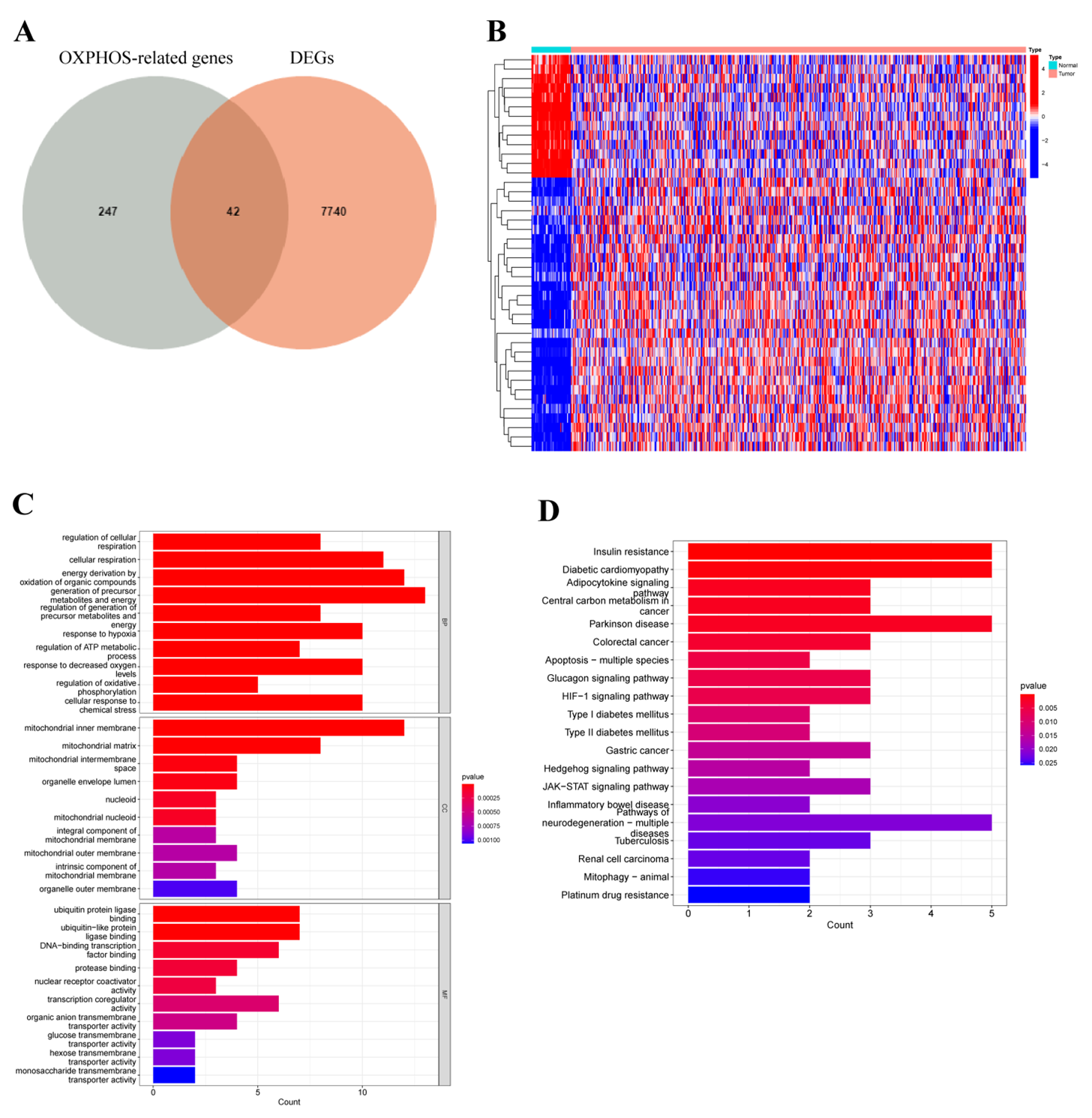
3.1. Differentially Expressed Oxidative Phosphorylation-Related Genes
3.2. Prediction Signature Construction
3.3. OXPHOS-Related Prognostic Signature Validation
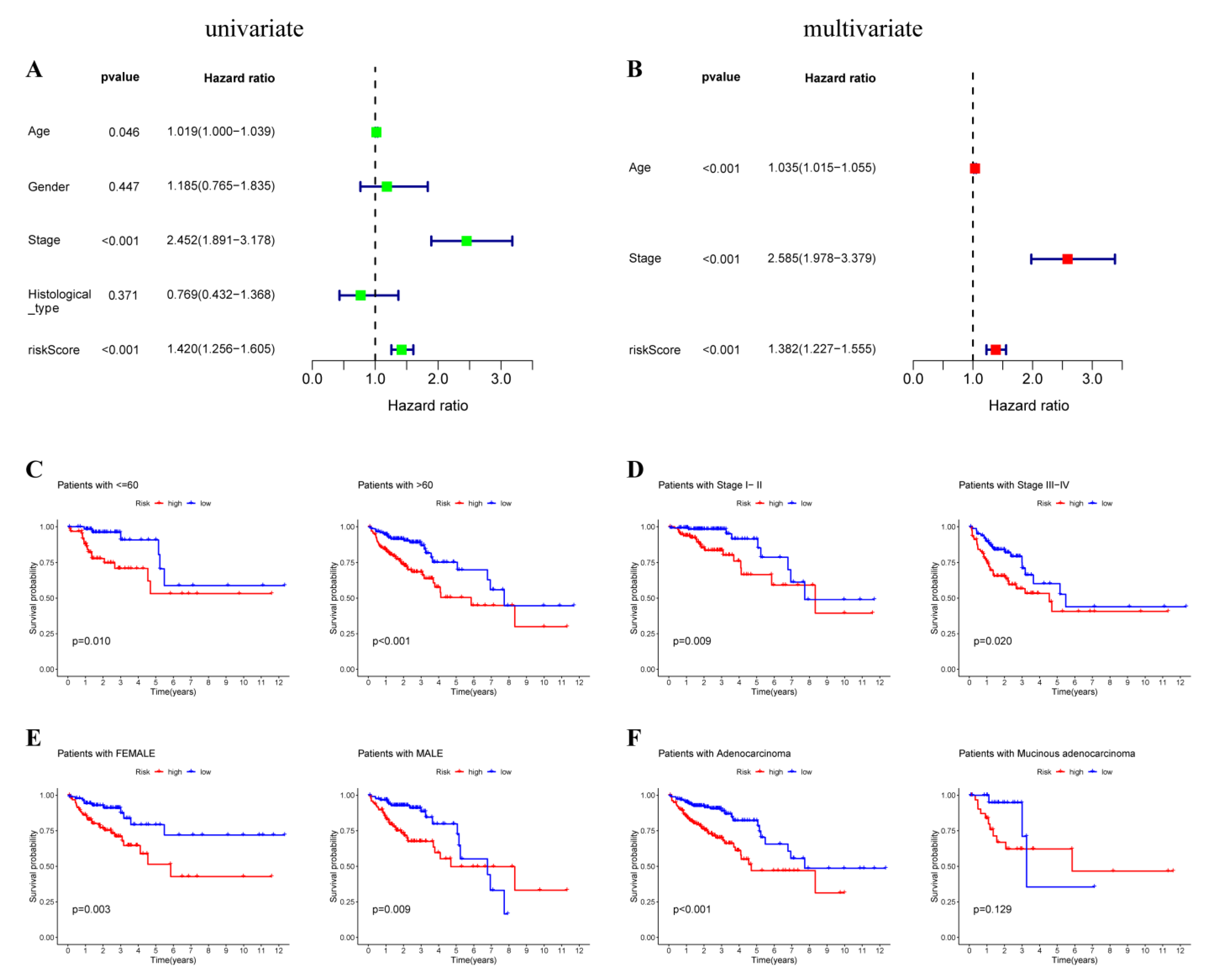
3.4. Prediction Value of OXPHOS-Related Prognostic Signature
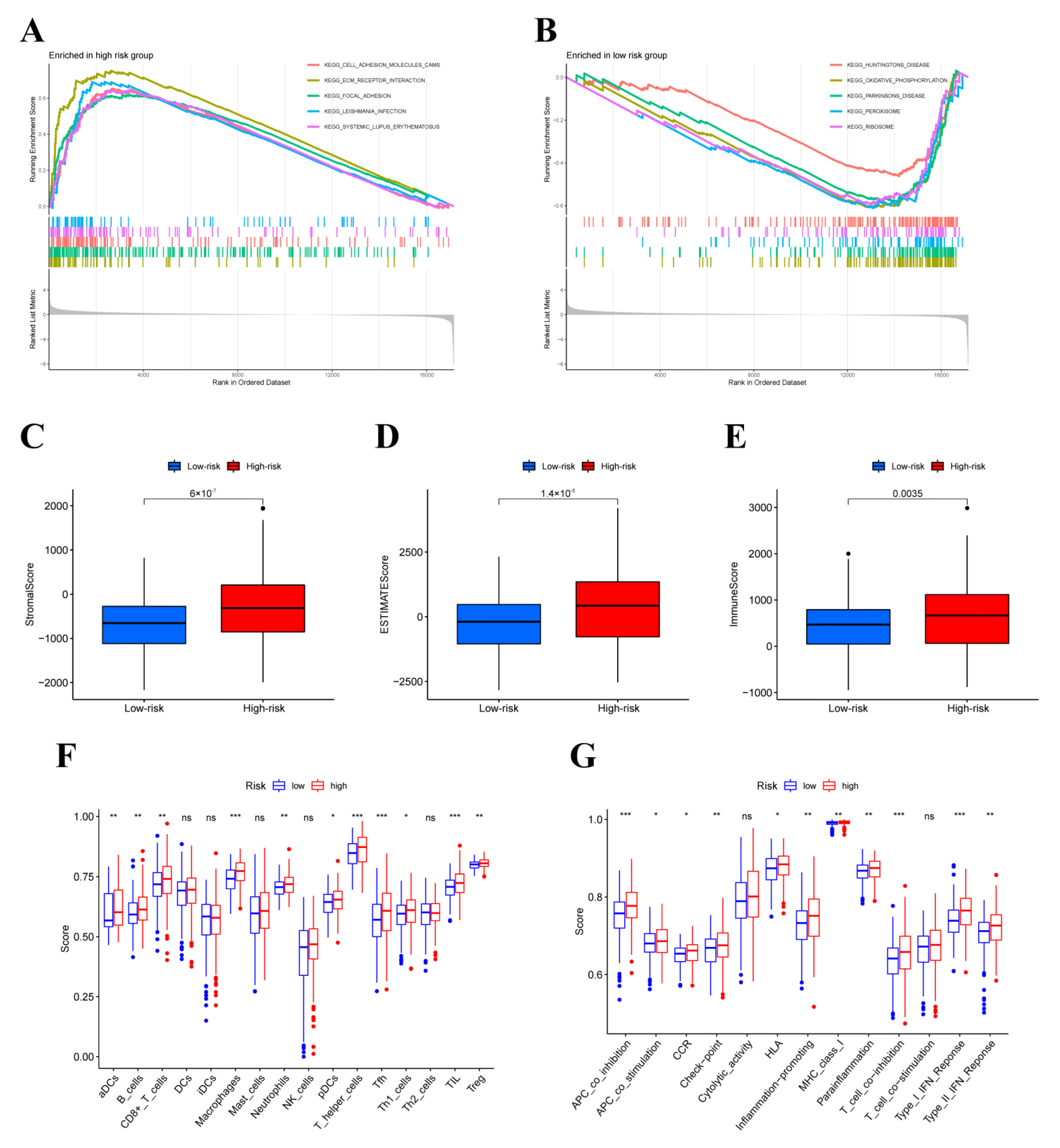
3.5. GESA, ESTIMATE and ssGSEA Analysis
3.6. Tumor-Infiltrating Immune Cells (TIICs) Analysis
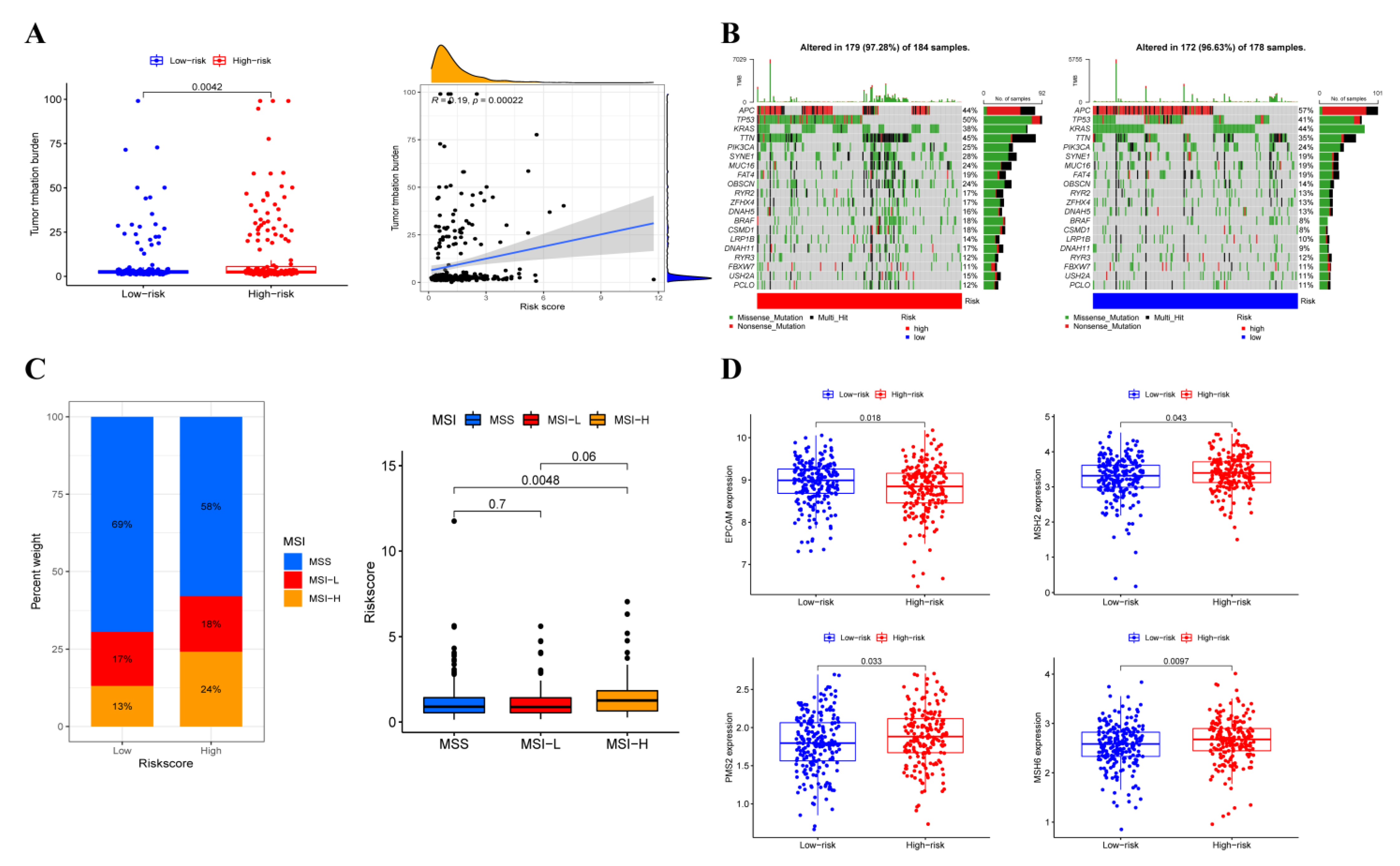
3.7. Somatic Variation Analysis
3.8. Immunotherapy Efficacy Analysis
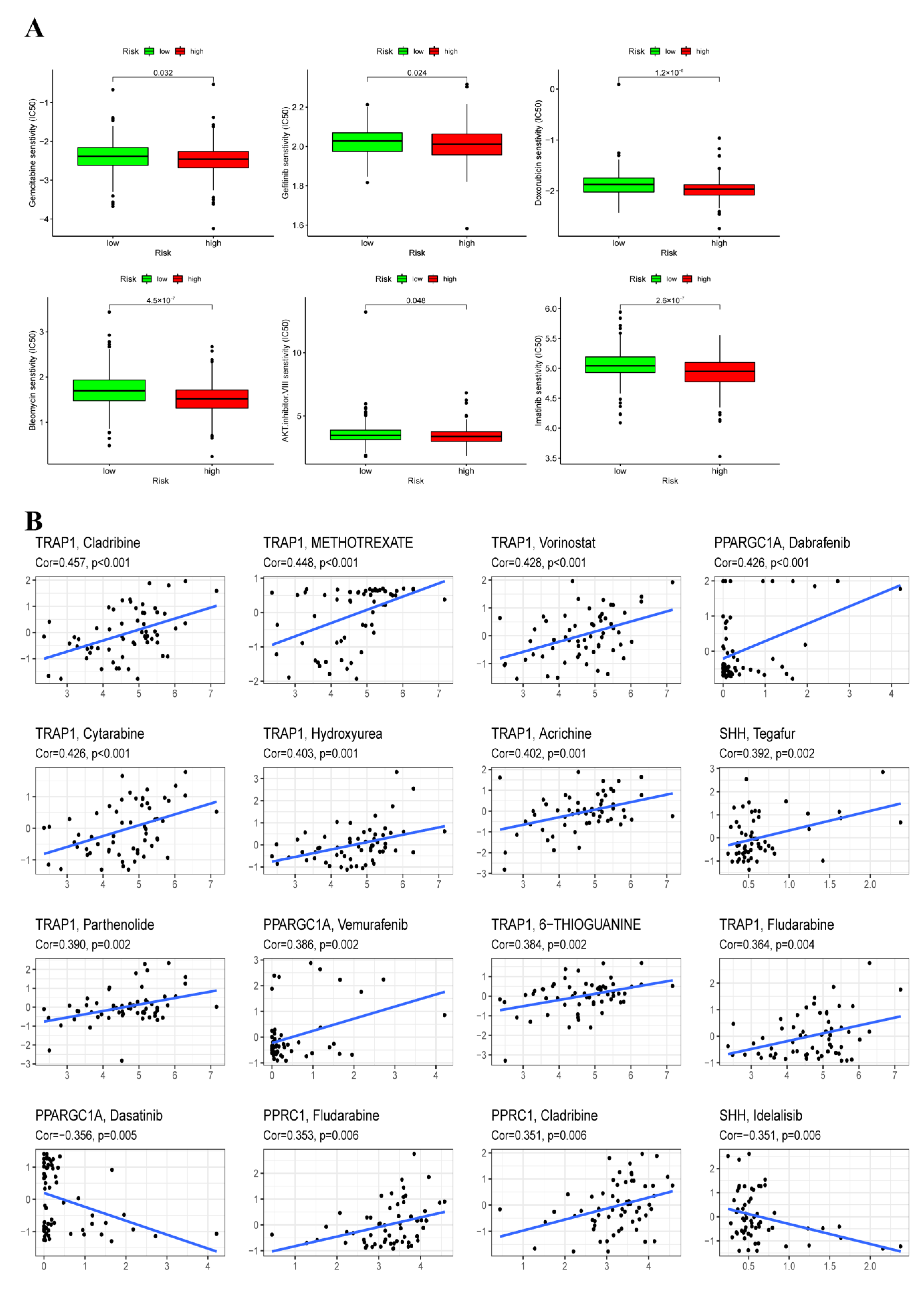
3.9. Chemotherapy Sensitivity Analysis
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Sung, H.; Ferlay, J.; Siegel, R.L.; Laversanne, M.; Soerjomataram, I.; Jemal, A.; Bray, F. Global Cancer Statistics 2020: GLOBOCAN Estimates of Incidence and Mortality Worldwide for 36 Cancers in 185 Countries. CA A Cancer J. Clin. 2021, 71, 209–249. [Google Scholar] [CrossRef] [PubMed]
- Pecci, F.; Cantini, L.; Bittoni, A.; Lenci, E.; Lupi, A.; Crocetti, S.; Giglio, E.; Giampieri, R.; Berardi, R. Beyond Microsatellite Instability: Evolving Strategies Integrating Immunotherapy for Microsatellite Stable Colorectal Cancer. Curr. Treat. Options Oncol. 2021, 22, 69. [Google Scholar] [CrossRef] [PubMed]
- Panza, A.; Castellana, S.; Biscaglia, G.; Piepoli, A.; Parca, L.; Gentile, A.; Latiano, A.; Mazza, T.; Perri, F.; Andriulli, A.; et al. Transcriptome and Gene Fusion Analysis of Synchronous Lesions Reveals lncMRPS31P5 as a Novel Transcript Involved in Colorectal Cancer. Int. J. Mol. Sci. 2020, 21, 7120. [Google Scholar] [CrossRef]
- Codrich, M.; Dalla, E.; Mio, C.; Antoniali, G.; Malfatti, M.C.; Marzinotto, S.; Pierobon, M.; Baldelli, E.; di Loreto, C.; Damante, G.; et al. Integrated multi-omics analyses on patient-derived CRC organoids highlight altered molecular pathways in colorectal cancer progression involving PTEN. J. Exp. Clin. Cancer Res. CR 2021, 40, 198. [Google Scholar] [CrossRef] [PubMed]
- Biller, L.H.; Schrag, D. Diagnosis and Treatment of Metastatic Colorectal Cancer: A Review. JAMA 2021, 325, 669–685. [Google Scholar] [CrossRef]
- Poturnajova, M.; Furielova, T.; Balintova, S.; Schmidtova, S.; Kucerova, L.; Matuskova, M. Molecular features and gene expression signature of metastatic colorectal cancer (Review). Oncol. Rep. 2021, 45, 10. [Google Scholar] [CrossRef]
- Andrei, P.; Battuello, P.; Grasso, G.; Rovera, E.; Tesio, N.; Bardelli, A. Integrated approaches for precision oncology in colorectal cancer: The more you know, the better. Semin. Cancer Biol. 2022, 84, 199–213. [Google Scholar] [CrossRef]
- Van Cutsem, E.; Köhne, C.H.; Hitre, E.; Zaluski, J.; Chang Chien, C.R.; Makhson, A.; D’Haens, G.; Pintér, T.; Lim, R.; Bodoky, G.; et al. Cetuximab and chemotherapy as initial treatment for metastatic colorectal cancer. N. Engl. J. Med. 2009, 360, 1408–1417. [Google Scholar] [CrossRef]
- Brody, H. Colorectal cancer. Nature 2015, 521, S1. [Google Scholar] [CrossRef]
- Sepulveda, A.R.; Hamilton, S.R.; Allegra, C.J.; Grody, W.; Cushman-Vokoun, A.M.; Funkhouser, W.K.; Kopetz, S.E.; Lieu, C.; Lindor, N.M.; Minsky, B.D.; et al. Molecular Biomarkers for the Evaluation of Colorectal Cancer: Guideline from the American Society for Clinical Pathology, College of American Pathologists, Association for Molecular Pathology, and the American Society of Clinical Oncology. J. Clin. Oncol. Off. J. Am. Soc. Clin. Oncol. 2017, 35, 1453–1486. [Google Scholar] [CrossRef]
- Puga Bermúdez, R.; Ramia, J.M.; de la Plaza Llamas, R.; Sabater Maroto, C.; García Fernández, E. Adenosquamous and squamous carcinomas of the colon and rectum. Gastroenterol. Y Hepatol. 2015, 38, 283–285. [Google Scholar] [CrossRef] [PubMed]
- Ganapathy-Kanniappan, S.; Geschwind, J.F. Tumor glycolysis as a target for cancer therapy: Progress and prospects. Mol. Cancer 2013, 12, 152. [Google Scholar] [CrossRef] [PubMed]
- Yu, L.; Chen, X.; Sun, X.; Wang, L.; Chen, S. The Glycolytic Switch in Tumors: How Many Players Are Involved? J. Cancer 2017, 8, 3430–3440. [Google Scholar] [CrossRef] [PubMed]
- Vander Heiden, M.G.; Cantley, L.C.; Thompson, C.B. Understanding the Warburg effect: The metabolic requirements of cell proliferation. Science 2009, 324, 1029–1033. [Google Scholar] [CrossRef] [PubMed]
- Frederick, M.; Skinner, H.D.; Kazi, S.A.; Sikora, A.G.; Sandulache, V.C. High expression of oxidative phosphorylation genes predicts improved survival in squamous cell carcinomas of the head and neck and lung. Sci. Rep. 2020, 10, 6380. [Google Scholar] [CrossRef]
- Solaini, G.; Sgarbi, G.; Baracca, A. Oxidative phosphorylation in cancer cells. Biochim. Biophys. Acta 2011, 1807, 534–542. [Google Scholar] [CrossRef]
- Evans, K.W.; Yuca, E.; Scott, S.S.; Zhao, M.; Paez Arango, N.; Cruz Pico, C.X.; Saridogan, T.; Shariati, M.; Class, C.A.; Bristow, C.A.; et al. Oxidative Phosphorylation Is a Metabolic Vulnerability in Chemotherapy-Resistant Triple-Negative Breast Cancer. Cancer Res. 2021, 81, 5572–5581. [Google Scholar] [CrossRef]
- Zacksenhaus, E.; Shrestha, M.; Liu, J.C.; Vorobieva, I.; Chung, P.E.D.; Ju, Y.; Nir, U.; Jiang, Z. Mitochondrial OXPHOS Induced by RB1 Deficiency in Breast Cancer: Implications for Anabolic Metabolism, Stemness, and Metastasis. Trends Cancer 2017, 3, 768–779. [Google Scholar] [CrossRef]
- Xu, Y.; Xue, D.; Bankhead, A., 3rd; Neamati, N. Why All the Fuss about Oxidative Phosphorylation (OXPHOS)? J. Med. Chem. 2020, 63, 14276–14307. [Google Scholar] [CrossRef]
- Sica, V.; Bravo-San Pedro, J.M.; Stoll, G.; Kroemer, G. Oxidative phosphorylation as a potential therapeutic target for cancer therapy. Int. J. Cancer 2020, 146, 10–17. [Google Scholar] [CrossRef]
- Ashton, T.M.; McKenna, W.G.; Kunz-Schughart, L.A.; Higgins, G.S. Oxidative Phosphorylation as an Emerging Target in Cancer Therapy. Clin. Cancer Res. Off. J. Am. Assoc. Cancer Res. 2018, 24, 2482–2490. [Google Scholar] [CrossRef] [PubMed]
- Ge, C.; Wang, Y.; Feng, Y.; Wang, S.; Zhang, K.; Xu, X.; Zhang, Z.; Zhao, Y.; Wang, Y.; Gao, L.; et al. Suppression of oxidative phosphorylation and IDH2 sensitizes colorectal cancer to a naphthalimide derivative and mitoxantrone. Cancer Lett. 2021, 519, 30–45. [Google Scholar] [CrossRef] [PubMed]
- Lascorz, J.; Bevier, M.; Schönfels, W.V.; Kalthoff, H.; Aselmann, H.; Beckmann, J.; Egberts, J.; Buch, S.; Becker, T.; Schreiber, S.; et al. Polymorphisms in the mitochondrial oxidative phosphorylation chain genes as prognostic markers for colorectal cancer. BMC Med. Genet. 2012, 13, 31. [Google Scholar] [CrossRef] [PubMed]
- Sun, X.; Zhan, L.; Chen, Y.; Wang, G.; He, L.; Wang, Q.; Zhou, F.; Yang, F.; Wu, J.; Wu, Y.; et al. Increased mtDNA copy number promotes cancer progression by enhancing mitochondrial oxidative phosphorylation in microsatellite-stable colorectal cancer. Signal Transduct. Target. Ther. 2018, 3, 8. [Google Scholar] [CrossRef] [PubMed]
- Vellinga, T.T.; Borovski, T.; de Boer, V.C.; Fatrai, S.; van Schelven, S.; Trumpi, K.; Verheem, A.; Snoeren, N.; Emmink, B.L.; Koster, J.; et al. SIRT1/PGC1α-Dependent Increase in Oxidative Phosphorylation Supports Chemotherapy Resistance of Colon Cancer. Clin. Cancer Res. Off. J. Am. Assoc. Cancer Res. 2015, 21, 2870–2879. [Google Scholar] [CrossRef]
- Xu, Z.; Wu, Z.; Zhang, J.; Zhou, R.; Ye, L.; Yang, P.; Yu, B. Development and validation of an oxidative phosphorylation-related gene signature in lung adenocarcinoma. Epigenomics 2020, 12, 1333–1348. [Google Scholar] [CrossRef]
- Liu, J.; Chen, T.; Yang, M.; Zhong, Z.; Ni, S.; Yang, S.; Shao, F.; Cai, L.; Bai, J.; Yu, H. Development of an Oxidative Phosphorylation-Related and Immune Microenvironment Prognostic Signature in Uterine Corpus Endometrial Carcinoma. Front. Cell Dev. Biol. 2021, 9, 753004. [Google Scholar] [CrossRef]
- Dekker, E.; Tanis, P.J.; Vleugels, J.L.A.; Kasi, P.M.; Wallace, M.B. Colorectal cancer. Lancet 2019, 394, 1467–1480. [Google Scholar] [CrossRef]
- Ritchie, M.E.; Phipson, B.; Wu, D.; Hu, Y.; Law, C.W.; Shi, W.; Smyth, G.K. Limma powers differential expression analyses for RNA-sequencing and microarray studies. Nucleic Acids Res. 2015, 43, e47. [Google Scholar] [CrossRef]
- Yu, G.; Wang, L.G.; Han, Y.; He, Q.Y. ClusterProfiler: An R package for comparing biological themes among gene clusters. Omics A J. Integr. Biol. 2012, 16, 284–287. [Google Scholar] [CrossRef]
- Lian, H.; Han, Y.P.; Zhang, Y.C.; Zhao, Y.; Yan, S.; Li, Q.F.; Wang, B.C.; Wang, J.J.; Meng, W.; Yang, J.; et al. Integrative analysis of gene expression and DNA methylation through one-class logistic regression machine learning identifies stemness features in medulloblastoma. Mol. Oncol. 2019, 13, 2227–2245. [Google Scholar] [CrossRef] [PubMed]
- Ranstam, J.; Cook, J.A. Kaplan-Meier curve. Br. J. Surg. 2017, 104, 442. [Google Scholar] [CrossRef] [PubMed]
- Yoon, P.S.; del Piccolo, N.; Shirure, V.S.; Peng, Y.; Kirane, A.; Canter, R.J.; Fields, R.C.; George, S.C.; Gholami, S. Advances in Modeling the Immune Microenvironment of Colorectal Cancer. Front. Immunol. 2020, 11, 614300. [Google Scholar] [CrossRef] [PubMed]
- Ke, Z.B.; Wu, Y.P.; Huang, P.; Hou, J.; Chen, Y.H.; Dong, R.N.; Lin, F.; Wei, Y.; Xue, X.Y.; Ng, C.F.; et al. Identification of novel genes in testicular cancer microenvironment based on ESTIMATE algorithm-derived immune scores. J. Cell. Physiol. 2021, 236, 706–713. [Google Scholar] [CrossRef]
- Subramanian, A.; Tamayo, P.; Mootha, V.K.; Mukherjee, S.; Ebert, B.L.; Gillette, M.A.; Paulovich, A.; Pomeroy, S.L.; Golub, T.R.; Lander, E.S.; et al. Gene set enrichment analysis: A knowledge-based approach for interpreting genome-wide expression profiles. Proc. Natl. Acad. Sci. USA 2005, 102, 15545–15550. [Google Scholar] [CrossRef]
- Chen, B.; Khodadoust, M.S.; Liu, C.L.; Newman, A.M.; Alizadeh, A.A. Profiling Tumor Infiltrating Immune Cells with CIBERSORT. Methods Mol. Biol. 2018, 1711, 243–259. [Google Scholar]
- Merino, D.M.; McShane, L.M.; Fabrizio, D.; Funari, V.; Chen, S.J.; White, J.R.; Wenz, P.; Baden, J.; Barrett, J.C.; Chaudhary, R.; et al. Establishing guidelines to harmonize tumor mutational burden (TMB): In silico assessment of variation in TMB quantification across diagnostic platforms: Phase I of the Friends of Cancer Research TMB Harmonization Project. J. Immunother. Cancer 2020, 8, e000147. [Google Scholar] [CrossRef]
- Mayakonda, A.; Lin, D.C.; Assenov, Y.; Plass, C.; Koeffler, H.P. Maftools: Efficient and comprehensive analysis of somatic variants in cancer. Genome Res. 2018, 28, 1747–1756. [Google Scholar] [CrossRef]
- Yi, L.; Huang, P.; Zou, X.; Guo, L.; Gu, Y.; Wen, C.; Wu, G. Integrative stemness characteristics associated with prognosis and the immune microenvironment in esophageal cancer. Pharmacol. Res. 2020, 161, 105144. [Google Scholar] [CrossRef]
- Charoentong, P.; Finotello, F.; Angelova, M.; Mayer, C.; Efremova, M.; Rieder, D.; Hackl, H.; Trajanoski, Z. Pan-cancer Immunogenomic Analyses Reveal Genotype-Immunophenotype Relationships and Predictors of Response to Checkpoint Blockade. Cell Rep. 2017, 18, 248–262. [Google Scholar] [CrossRef]
- Yang, W.; Soares, J.; Greninger, P.; Edelman, E.J.; Lightfoot, H.; Forbes, S.; Bindal, N.; Beare, D.; Smith, J.A.; Thompson, I.R.; et al. Genomics of Drug Sensitivity in Cancer (GDSC): A resource for therapeutic biomarker discovery in cancer cells. Nucleic Acids Res. 2013, 41, D955–D961. [Google Scholar] [CrossRef]
- Shankavaram, U.T.; Varma, S.; Kane, D.; Sunshine, M.; Chary, K.K.; Reinhold, W.C.; Pommier, Y.; Weinstein, J.N. CellMiner: A relational database and query tool for the NCI-60 cancer cell lines. BMC Genom. 2009, 10, 277. [Google Scholar] [CrossRef]
- Lin, W.R.; Chiang, J.M.; Lim, S.N.; Su, M.Y.; Chen, T.H.; Huang, S.W.; Chen, C.W.; Wu, R.C.; Tsai, C.L.; Lin, Y.H.; et al. Dynamic bioenergetic alterations in colorectal adenomatous polyps and adenocarcinomas. EBioMedicine 2019, 44, 334–345. [Google Scholar] [CrossRef] [PubMed]
- Stintzing, S.; Tejpar, S.; Gibbs, P.; Thiebach, L.; Lenz, H.J. Understanding the role of primary tumour localisation in colorectal cancer treatment and outcomes. Eur. J. Cancer 2017, 84, 69–80. [Google Scholar] [CrossRef] [PubMed]
- Brenner, H.; Kloor, M.; Pox, C.P. Colorectal cancer. Lancet 2014, 383, 1490–1502. [Google Scholar] [CrossRef]
- Vodenkova, S.; Buchler, T.; Cervena, K.; Veskrnova, V.; Vodicka, P.; Vymetalkova, V. 5-fluorouracil and other fluoropyrimidines in colorectal cancer: Past, present and future. Pharmacol. Ther. 2020, 206, 107447. [Google Scholar] [CrossRef]
- Marjaneh, R.M.; Khazaei, M.; Ferns, G.A.; Avan, A.; Aghaee-Bakhtiari, S.H. The role of microRNAs in 5-FU resistance of colorectal cancer: Possible mechanisms. J. Cell. Physiol. 2019, 234, 2306–2316. [Google Scholar] [CrossRef]
- Rasola, A.; Neckers, L.; Picard, D. Mitochondrial oxidative phosphorylation TRAP(1)ped in tumor cells. Trends Cell Biol. 2014, 24, 455–463. [Google Scholar] [CrossRef]
- Gao, J.Y.; Song, B.R.; Peng, J.J.; Lu, Y.M. Correlation between mitochondrial TRAP-1 expression and lymph node metastasis in colorectal cancer. World J. Gastroenterol. 2012, 18, 5965–5971. [Google Scholar] [CrossRef]
- Costantino, E.; Maddalena, F.; Calise, S.; Piscazzi, A.; Tirino, V.; Fersini, A.; Ambrosi, A.; Neri, V.; Esposito, F.; Landriscina, M. TRAP1, a novel mitochondrial chaperone responsible for multi-drug resistance and protection from apoptotis in human colorectal carcinoma cells. Cancer Lett. 2009, 279, 39–46. [Google Scholar] [CrossRef]
- Masuda, Y.; Shima, G.; Aiuchi, T.; Horie, M.; Hori, K.; Nakajo, S.; Kajimoto, S.; Shibayama-Imazu, T.; Nakaya, K. Involvement of tumor necrosis factor receptor-associated protein 1 (TRAP1) in apoptosis induced by beta-hydroxyisovalerylshikonin. J. Biol. Chem. 2004, 279, 42503–42515. [Google Scholar] [CrossRef] [PubMed]
- LeBleu, V.S.; O’Connell, J.T.; Gonzalez Herrera, K.N.; Wikman, H.; Pantel, K.; Haigis, M.C.; de Carvalho, F.M.; Damascena, A.; Domingos Chinen, L.T.; Rocha, R.M.; et al. PGC-1α mediates mitochondrial biogenesis and oxidative phosphorylation in cancer cells to promote metastasis. Nat. Cell Biol. 2014, 16, 992–1003. [Google Scholar] [CrossRef] [PubMed]
- Dickinson, A.; Yeung, K.Y.; Donoghue, J.; Baker, M.J.; Kelly, R.D.; McKenzie, M.; Johns, T.G.; St John, J.C. The regulation of mitochondrial DNA copy number in glioblastoma cells. Cell Death Differ. 2013, 20, 1644–1653. [Google Scholar] [CrossRef] [PubMed]
- Mori, M.P.; Souza-Pinto, N.C. PPRC1, but not PGC-1α, levels directly correlate with expression of mitochondrial proteins in human dermal fibroblasts. Genet. Mol. Biol. 2020, 43 (Suppl. 1), e20190083. [Google Scholar] [CrossRef] [PubMed]
- Tsai, H.F.; Chang, Y.C.; Li, C.H.; Chan, M.H.; Chen, C.L.; Tsai, W.C.; Hsiao, M. Type V collagen alpha 1 chain promotes the malignancy of glioblastoma through PPRC1-ESM1 axis activation and extracellular matrix remodeling. Cell Death Discov. 2021, 7, 313. [Google Scholar] [CrossRef]
- Missiaglia, E.; Jacobs, B.; D’Ario, G.; di Narzo, A.F.; Soneson, C.; Budinska, E.; Popovici, V.; Vecchione, L.; Gerster, S.; Yan, P.; et al. Distal and proximal colon cancers differ in terms of molecular, pathological, and clinical features. Ann. Oncol. Off. J. Eur. Soc. Med. Oncol. 2014, 25, 1995–2001. [Google Scholar] [CrossRef]
- Lee, D.W.; Han, S.W.; Cha, Y.; Bae, J.M.; Kim, H.P.; Lyu, J.; Han, H.; Kim, H.; Jang, H.; Bang, D.; et al. Association between mutations of critical pathway genes and survival outcomes according to the tumor location in colorectal cancer. Cancer 2017, 123, 3513–3523. [Google Scholar] [CrossRef]
- Guinney, J.; Dienstmann, R.; Wang, X.; de Reyniès, A.; Schlicker, A.; Soneson, C.; Marisa, L.; Roepman, P.; Nyamundanda, G.; Angelino, P.; et al. The consensus molecular subtypes of colorectal cancer. Nat. Med. 2015, 21, 1350–1356. [Google Scholar] [CrossRef]
- Dienstmann, R.; Vermeulen, L.; Guinney, J.; Kopetz, S.; Tejpar, S.; Tabernero, J. Consensus molecular subtypes and the evolution of precision medicine in colorectal cancer. Nat. Rev. Cancer 2017, 17, 79–92. [Google Scholar] [CrossRef]
- Schwartz, B.; Yehuda-Shnaidman, E. Putative role of adipose tissue in growth and metabolism of colon cancer cells. Front. Oncol. 2014, 4, 164. [Google Scholar] [CrossRef]
- Artyomov, M.N.; Sergushichev, A.; Schilling, J.D. Integrating immunometabolism and macrophage diversity. Semin. Immunol. 2016, 28, 417–424. [Google Scholar] [CrossRef] [PubMed]
- Liu, Y.; Xu, R.; Gu, H.; Zhang, E.; Qu, J.; Cao, W.; Huang, X.; Yan, H.; He, J.; Cai, Z. Metabolic reprogramming in macrophage responses. Biomark. Res. 2021, 9, 1. [Google Scholar] [CrossRef] [PubMed]
- Habtetsion, T.; Ding, Z.C.; Pi, W.; Li, T.; Lu, C.; Chen, T.; Xi, C.; Spartz, H.; Liu, K.; Hao, Z.; et al. Alteration of Tumor Metabolism by CD4+ T Cells Leads to TNF-α-Dependent Intensification of Oxidative Stress and Tumor Cell Death. Cell Metab. 2018, 28, 228–242.e6. [Google Scholar] [CrossRef] [PubMed]
- Viswanathan, V.S.; Ryan, M.J.; Dhruv, H.D.; Gill, S.; Eichhoff, O.M.; Seashore-Ludlow, B.; Kaffenberger, S.D.; Eaton, J.K.; Shimada, K.; Aguirre, A.J.; et al. Dependency of a therapy-resistant state of cancer cells on a lipid peroxidase pathway. Nature 2017, 547, 453–457. [Google Scholar] [CrossRef]
- Beckermann, K.E.; Dudzinski, S.O.; Rathmell, J.C. Dysfunctional T cell metabolism in the tumor microenvironment. Cytokine Growth Factor Rev. 2017, 35, 7–14. [Google Scholar] [CrossRef]
- Liberti, M.V.; Dai, Z.; Wardell, S.E.; Baccile, J.A.; Liu, X.; Gao, X.; Baldi, R.; Mehrmohamadi, M.; Johnson, M.O.; Madhukar, N.S.; et al. A Predictive Model for Selective Targeting of the Warburg Effect through GAPDH Inhibition with a Natural Product. Cell Metab. 2017, 26, 648–659.e8. [Google Scholar] [CrossRef]
- Luengo, A.; Gui, D.Y.; Vander Heiden, M.G. Targeting Metabolism for Cancer Therapy. Cell Chem. Biol. 2017, 24, 1161–1180. [Google Scholar] [CrossRef]
- Ganesh, K.; Stadler, Z.K.; Cercek, A.; Mendelsohn, R.B.; Shia, J.; Segal, N.H.; Diaz, L.A., Jr. Immunotherapy in colorectal cancer: Rationale, challenges and potential. Nat. Rev. Gastroenterol. Hepatol. 2019, 16, 361–375. [Google Scholar] [CrossRef]
- Nair, S.S.; Weil, R.; Dovey, Z.; Davis, A.; Tewari, A.K. The Tumor Microenvironment and Immunotherapy in Prostate and Bladder Cancer. Urol. Clin. N. Am. 2020, 47, e17–e54. [Google Scholar] [CrossRef]




| Genes | HR | Low 95%CI | High 95%CI | p-Value |
|---|---|---|---|---|
| MRPS23 | 0.499 | 0.255 | 0.976 | 0.042 |
| PPARGC1A | 0.546 | 0.307 | 0.972 | 0.040 |
| SHH | 0.595 | 0.365 | 0.972 | 0.038 |
| FSCN1 | 1.346 | 1.052 | 1.721 | 0.018 |
| TRAP1 | 0.437 | 0.205 | 0.931 | 0.032 |
| MPC1 | 0.433 | 0.214 | 0.878 | 0.020 |
| PPRC1 | 2.288 | 1.098 | 4.766 | 0.027 |
| PPA1 | 0.516 | 0.266 | 1.000 | 0.050 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Wang, C.; Cui, G.; Wang, D.; Wang, M.; Chen, Q.; Wang, Y.; Lu, M.; Tang, X.; Yang, B. Crosstalk of Oxidative Phosphorylation-Related Subtypes, Establishment of a Prognostic Signature and Immune Infiltration Characteristics in Colorectal Adenocarcinoma. Cancers 2022, 14, 4503. https://doi.org/10.3390/cancers14184503
Wang C, Cui G, Wang D, Wang M, Chen Q, Wang Y, Lu M, Tang X, Yang B. Crosstalk of Oxidative Phosphorylation-Related Subtypes, Establishment of a Prognostic Signature and Immune Infiltration Characteristics in Colorectal Adenocarcinoma. Cancers. 2022; 14(18):4503. https://doi.org/10.3390/cancers14184503
Chicago/Turabian StyleWang, Can, Guoliang Cui, Dan Wang, Min Wang, Qi Chen, Yunshan Wang, Mengjie Lu, Xinyi Tang, and Bolin Yang. 2022. "Crosstalk of Oxidative Phosphorylation-Related Subtypes, Establishment of a Prognostic Signature and Immune Infiltration Characteristics in Colorectal Adenocarcinoma" Cancers 14, no. 18: 4503. https://doi.org/10.3390/cancers14184503
APA StyleWang, C., Cui, G., Wang, D., Wang, M., Chen, Q., Wang, Y., Lu, M., Tang, X., & Yang, B. (2022). Crosstalk of Oxidative Phosphorylation-Related Subtypes, Establishment of a Prognostic Signature and Immune Infiltration Characteristics in Colorectal Adenocarcinoma. Cancers, 14(18), 4503. https://doi.org/10.3390/cancers14184503





