Prevalence and Prognostic Role of IDH Mutations in Acute Myeloid Leukemia: Results of the GIMEMA AML1516 Protocol
Abstract
Simple Summary
Abstract
1. Introduction
2. Materials and Methods
2.1. GIMEMA AML1516 Study Design
2.2. Statistical Analysis
3. Results
3.1. Study Population
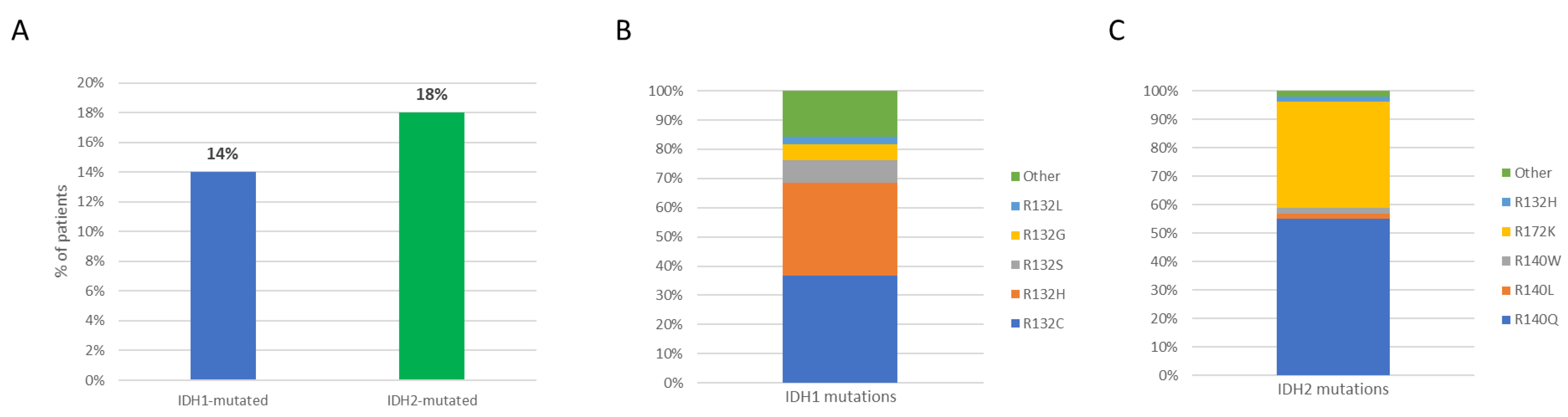
3.2. Incidence, Type of IDH1/2 Mutations, and Patients’ Clinico-Biological Features
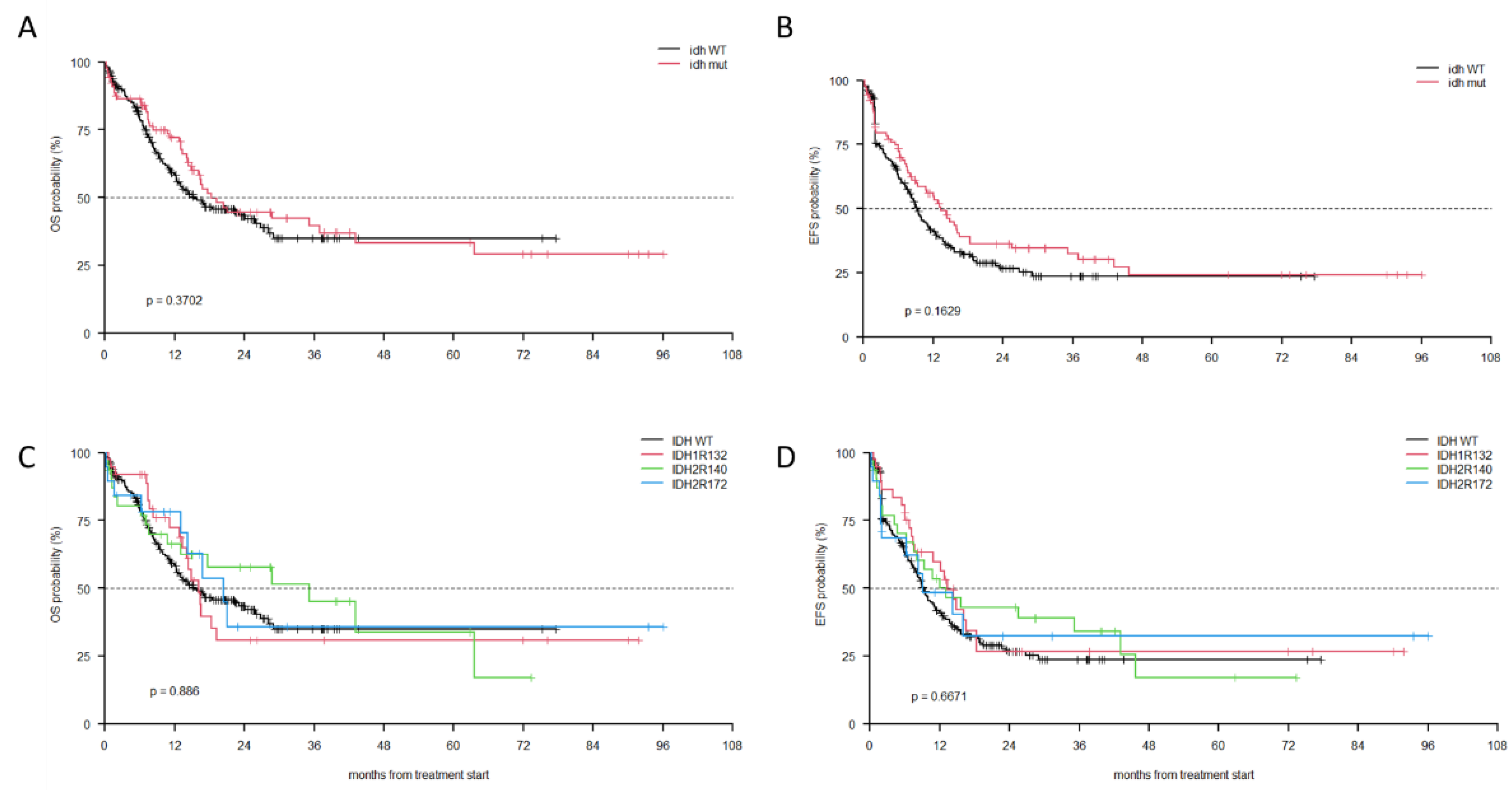
3.3. Treatment Response and Survival According to IDH1/2 Mutations
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Ley, T.J.; Miller, C.; Ding, L.; Raphael, B.J.; Mungall, A.J.; Robertson, A.; Hoadley, K.; Triche, T.J., Jr.; Laird, P.W. Cancer Genome Atlas Research Network, Genomic and epigenomic landscapes of adult de novo acute myeloid leukemia. N. Engl. J. Med. 2013, 368, 2059–2074. [Google Scholar] [CrossRef] [PubMed]
- Tyner, J.W.; Tognon, C.E.; Bottomly, D.; Wilmot, B.; Kurtz, S.E.; Savage, S.L.; Long, N.; Schultz, A.R.; Traer, E.; Abel, M.; et al. Functional genomic landscape of acute myeloid leukaemia. Nature 2018, 562, 526–531. [Google Scholar] [CrossRef]
- Mardis, E.R.; Ding, L.; Dooling, D.J.; Larson, D.E.; McLellan, M.D.; Chen, K.; Koboldt, D.C.; Fulton, R.S.; Delehaunty, K.D.; McGrath, S.D.; et al. Recurring mutations found by sequencing an acute myeloid leukemia genome. N. Engl. J. Med. 2009, 361, 1058–1066. [Google Scholar] [CrossRef] [PubMed]
- Figueroa, M.E.; Abdel-Wahab, O.; Lu, C.; Ward, P.S.; Patel, J.; Shih, A.; Li, Y.; Bhagwat, N.; Vasanthakumar, A.; Fernandez, H.F.; et al. Leukemic IDH1 and IDH2 mutations result in a hypermethylation phenotype, disrupt TET2 function, and impair hematopoietic differentiation. Cancer Cell. 2010, 18, 553–567. [Google Scholar] [CrossRef] [PubMed]
- Paschka, P.; Schlenk, R.F.; Gaidzik, V.I.; Habdank, M.; Krönke, J.; Bullinger, L.; Späth, D.; Kayser, S.; Zucknick, M.; Götze, K.; et al. IDH1 and IDH2 mutations are frequent genetic alterations in acute myeloid leukemia and confer adverse prognosis in cytogenetically normal acute myeloid leukemia with NPM1 mutation without FLT3 internal tandem duplication. J. Clin. Oncol. 2010, 28, 3636–3643. [Google Scholar] [CrossRef] [PubMed]
- Boissel, N.; Nibourel, O.; Renneville, A.; Gardin, C.; Reman, O.; Contentin, N.; Bordessoule, D.; Pautas, C.; de Revel, T.; Quesnel, B.; et al. Prognostic impact of isocitrate dehydrogenase enzyme isoforms 1 and 2 mutations in acute myeloid leukemia: A study by the Acute Leukemia French Association group. J. Clin. Oncol. 2010, 28, 3717–3723. [Google Scholar] [CrossRef]
- Damm, F.; Thol, F.; Hollink, I.; Zimmermann, M.; Reinhardt, K.; van den Heuvel-Eibrink, M.M.; Zwaan, C.M.; de Haas, V.; Creutzig, U.; Klusmann, J.H.; et al. Prevalence and prognostic value of IDH1 and IDH2 mutations in childhood AML: A study of the AML-BFM and DCOG study groups. Leukemias 2011, 25, 1704–1710. [Google Scholar] [CrossRef][Green Version]
- Marcucci, G.; Maharry, K.; Wu, Y.Z.; Radmacher, M.D.; Mrózek, K.; Margeson, D.; Holland, K.B.; Whitman, S.P.; Becker, H.; Schwind, S.; et al. IDH1 and IDH2 gene mutations identify novel molecular subsets within de novo cytogenetically normal acute myeloid leukemia: A Cancer and Leukemia Group B study. J. Clin. Oncol. 2010, 28, 2348–2355. [Google Scholar] [CrossRef]
- Ye, D.; Xiong, Y.; Guan, K.L. The mechanisms of IDH mutations in tumorigenesis. Cell Res. 2012, 22, 1102–1104. [Google Scholar] [CrossRef][Green Version]
- Papaemmanuil, E.; Gerstung, M.; Bullinger, L.; Gaidzik, V.I.; Paschka, P.; Roberts, N.D.; Potter, N.E.; Heuser, M.; Thol, F.; Bolli, N.; et al. Genomic Classification and Prognosis in Acute Myeloid Leukemia. N. Engl. J. Med. 2016, 374, 2209–2221. [Google Scholar] [CrossRef]
- Simonetti, G.; Mengucci, C.; Padella, A.; Fonzi, E.; Picone, G.; Delpino, C.; Nanni, J.; De Tommaso, R.; Franchini, E.; Papayannidis, C.; et al. Integrated genomic-metabolic classification of acute myeloid leukemia defines a subgroup with NPM1 and cohesin/DNA damage mutations. Leukemias 2021, 35, 2813–2826. [Google Scholar] [CrossRef] [PubMed]
- Chou, W.C.; Lei, W.C.; Ko, B.S.; Hou, H.A.; Chen, C.Y.; Tang, J.L.; Yao, M.; Tsay, W.; Wu, S.J.; Huang, S.Y.; et al. The prognostic impact and stability of Isocitrate dehydrogenase 2 mutation in adult patients with acute myeloid leukemia. Leukemias 2011, 25, 246–253. [Google Scholar] [CrossRef] [PubMed]
- Falini, B.; Spinelli, O.; Meggendorfer, M.; Martelli, M.P.; Bigerna, B.; Ascani SStein, H.; Rambaldi, A.; Haferlach, T. IDH1-R132 changes vary according to NPM1 and other mutations status in AML. Leukemias 2019, 33, 1043–1047. [Google Scholar] [CrossRef]
- Meggendorfer, M.; Cappelli, L.V.; Walter, W.; Haferlach, C.; Kern, W.; Falini, B.; Haferlach, T. IDH1R132, IDH2R140 and IDH2R172 in AML: Different genetic landscapes correlate with outcome and may influence targeted treatment strategies. Leukemias 2018, 32, 1249–1253. [Google Scholar] [CrossRef]
- Thol, F.; Damm, F.; Wagner, K.; Göhring, G.; Schlegelberger, B.; Hoelzer, D.; Lübbert, M.; Heit, W.; Kanz, L.; Schlimok, G.; et al. Prognostic impact of IDH2 mutations in cytogenetically normal acute myeloid leukemia. Blood 2010, 116, 614–616. [Google Scholar] [CrossRef] [PubMed]
- Abbas, S.; Lugthart, S.; Kavelaars, F.G.; Schelen, A.; Koenders, J.E.; Zeilemaker, A.; van Putten, W.J.; Rijneveld, A.W.; Löwenberg, B.; Valk, P.J. Acquired mutations in the genes 525 encoding IDH1 and IDH2 both are recurrent aberrations in acute myeloid 526 leukemia: Prevalence and prognostic value. Blood 2010, 116, 2122–2126. [Google Scholar] [CrossRef]
- Chotirat, S.; Thongnoppakhun, W.; Promsuwicha, O.; Boonthimat, C.; Auewarakul, C.U. Molecular alterations of isocitrate dehydrogenase 1 and 2 (IDH1 and IDH2) metabolic genes and additional genetic mutations in newly diagnosed acute myeloid leukemia patients. J. Hematol. Oncol. 2012, 5, 5. [Google Scholar] [CrossRef]
- DiNardo, C.D.; Ravandi, F.; Agresta, S.; Konopleva, M.; Takahashi, K.; Kadia, T.; Routbort, M.; Patel, K.P.; Brandt, M.; Pierce, S.; et al. Characteristics, clinical outcome, and 538 prognostic significance of IDH mutations in AML. Am. J. Hematol. 2015, 90, 732–736. [Google Scholar] [CrossRef]
- Stein, E.M.; DiNardo, C.D.; Pollyea, D.A.; Fathi, A.T.; Roboz, G.J.; Altman, J.K.; Stone, R.M.; DeAngelo, D.J.; Levine, R.L.; Flinn, I.W.; et al. Enasidenib in mutant IDH2 relapsed or refractory acute myeloid leukemia. Blood 2017, 130, 722–731. [Google Scholar] [CrossRef]
- DiNardo, C.D.; Stein, E.M.; de Botton, S.; Roboz, G.J.; Altman, J.K.; Mims, A.S.; Swords, R.; Collins, R.H.; Mannis, G.N.; Pollyea, D.A.; et al. Durable Remissions with Ivosidenib in IDH1-Mutated Relapsed or Refractory AML. N. Engl. J. Med. 2018, 378, 2386–2398. [Google Scholar] [CrossRef]
- DiNardo, C.D.; Stein, A.S.; Stein, E.M.; Fathi, A.T.; Frankfurt, O.; Schuh, A.C.; Döhner, H.; Martinelli, G.; Patel, P.A.; Raffoux, E.; et al. Mutant Isocitrate Dehydrogenase 1 Inhibitor Ivosidenib in Combination With Azacitidine for Newly Diagnosed Acute Myeloid Leukemia. J. Clin. Oncol. 2021, 39, 57–65. [Google Scholar] [CrossRef] [PubMed]
- DiNardo, C.D.; Schuh, A.C.; Stein, E.M.; Montesinos, P.; Wei, A.H.; de Botton, S.; Zeidan, A.M.; Fathi, A.T.; Kantarjian, H.M.; Bennett, J.M.; et al. Enasidenib plus azacitidine versus azacitidine alone in patients with newly diagnosed, mutant-IDH2 acute myeloid leukaemia (AG221-AML-005): A single-arm, phase 1b and randomised, phase 2 trial. Lancet Oncol. 2021, 22, 1597–1608. [Google Scholar] [CrossRef]
- Aguilera-Diaz, A.; Vazquez, I.; Ariceta, B.; Mañú, A.; Blasco-Iturri, Z.; Palomino-Echeverría, S.; Larrayoz, M.J.; García-Sanz, R.; Prieto-Conde, M.I.; Del Carmen Chillón, M.; et al. Assessment of the clinical utility of four NGS panels in myeloid malignancies. Suggestions for NGS panel choice or design. PLoS ONE 2020, 15, e0227986. [Google Scholar] [CrossRef] [PubMed]
- Harris, P.A.; Taylor, R.; Thielke, R.; Payne, J.; Gonzalez, N.; Conde, J.G. Research electronic data capture (REDCap)—A metadata-driven methodology and workflow process for providing translational research informatics support. J. Biomed. Inform. 2009, 42, 377–381. [Google Scholar] [CrossRef] [PubMed]
- Harris, P.A.; Taylor, R.; Minor, B.L.; Elliott, V.; Fernandez, M.; O’Neal, L.; McLeod, L.; Delacqua, G.; Delacqua, F.; Kirby, J.; et al. REDCap Consortium, The REDCap consortium: Building an international community of software partners. J. Biomed. Inform. 2019, 95, 103208. [Google Scholar] [CrossRef] [PubMed]
- Middeke, J.M.; Metzeler, K.H.; Röllig, C.; Kramer, M.; Eckardt, J.N.; Stasik, S.; Greif, P.A.; Spiekermann, K.; Rothenberg-Thurley, M.; Krug, U.; et al. Differential impact of IDH1/2 mutational subclasses on outcome in adult AML: Results from a large multicenter study. Blood Adv. 2022, 6, 1394–1405. [Google Scholar] [CrossRef]
- Chou, W.C.; Peng, K.Y.; Lei, W.C.; Ko, B.S.; Tsay, W.; Kuo, C.H.; Tien, H.F. Persistence of mutant isocitrate dehydrogenase in patients with acute myeloid leukemia in remission. Leukemias 2012, 26, 527–529. [Google Scholar] [CrossRef][Green Version]
- Roboz, G.J.; DiNardo, C.D.; Stein, E.M.; de Botton, S.; Mims, A.S.; Prince, G.T.; Altman, J.K.; Arellano, M.L.; Donnellan, W.; Erba, H.P.; et al. Ivosidenib induces deep durable remissions in patients with newly diagnosed IDH1-mutant acute myeloid leukemia. Blood 2020, 135, 463–471. [Google Scholar] [CrossRef]
- Stein, E.M.; DiNardo, C.D.; Fathi, A.T.; Mims, A.S.; Pratz, K.W.; Savona, M.R.; Stein, A.S.; Stone, R.M.; Winer, E.S.; Seet, C.S.; et al. Ivosidenib or enasidenib combined with intensive chemotherapy in patients with newly diagnosed AML: A phase 1 study. Blood 2021, 137, 1792–1803. [Google Scholar] [CrossRef]
- Venugopal, S.; Takahashi, K.; Daver, N.; Maiti, A.; Borthakur, G.; Loghavi, S.; Short, N.J.; Ohanian, M.; Masarova, L.; Issa, G.; et al. Efficacy and safety of enasidenib and azacitidine combination in patients with IDH2 mutated acute myeloid leukemia and not eligible for intensive chemotherapy. Blood Cancer J. 2022, 12, 10. [Google Scholar] [CrossRef]
- DiNardo, C.D.; Tiong, I.S.; Quaglieri, A.; MacRaild, S.; Loghavi, S.; Brown, F.C.; Thijssen, R.; Pomilio, G.; Ivey, A.; Salmon, J.M.; et al. Molecular patterns of response and treatment failure after frontline venetoclax combinations in older patients with AML. Blood 2020, 135, 791–803. [Google Scholar] [CrossRef] [PubMed]
- Lachowiez, C.A.; Borthakur, G.; Loghavi, S.; Zeng, Z.; Kadia, T.M.; Masarova, L.; Takahashi, K.; Tippett, G.D.; Naqvi, K.; Bose, P.; et al. A phase Ib/II study of ivosidenib with venetoclax +/− azacitidine in IDH1-mutated myeloid malignancies. J. Clin. Oncol. 2020, 38, 7500. [Google Scholar] [CrossRef]
| IDH1-IDH2 Mutated vs. IDH1-IDH2 (Both)WT | |||||
|---|---|---|---|---|---|
| Characteristic | Overall, n = 284 | IDH1-IDH2 WT, n = 195 | IDH1-Mut, n = 38 | IDH2-Mut, n = 51 | p-Value 1 |
| Gender, n (%) | 0.15 | ||||
| M | 145 (51%) | 93 (48%) | 20 (53%) | 32 (63%) | |
| F | 139 (49%) | 102 (52%) | 18 (47%) | 19 (37%) | |
| Age starting treatment, median (range) | 65 (19, 86) | 65 (19, 85) | 66 (22, 86) | 65 (32, 85) | 0.86 |
| WBC (109/L), median (range) | 7 (0.5, 800) | 8 (0.5, 347) | 5 (1, 600) | 4 (0.4, 800) | 0.063 |
| HB (g/dL), median (range) | 9.00 (2.50, 14.9) | 9.00 (4.50, 14.2) | 8.80 (7.20, 13.20) | 9.30 (2.50, 14.90) | 0.28 |
| PLTS (109/L), median (range) | 56 (4, 789) | 53 (4, 664) | 110 (6, 742) | 56 (10, 789) | 0.036 |
| Blasts (% in BM), median (range) | 50 (3, 99) | 48 (4, 99) | 75 (3, 96) | 70 (4, 95) | 0.027 |
| WHO PS, n (%) | <0.001 | ||||
| 0 | 118 (44%) | 79 (43%) | 15 (39%) | 24 (50%) | |
| I | 111 (41%) | 89 (48%) | 12 (32%) | 10 (21%) | |
| II | 34 (13%) | 16 (8.6%) | 7 (18%) | 11 (23%) | |
| III | 8 (3.0%) | 1 (0.5%) | 4 (11%) | 3 (6.2%) | |
| AML type, n (%) | 0.63 | ||||
| de novo | 229 (81%) | 154 (80%) | 31 (82%) | 44 (86%) | |
| secondary | 37 (13%) | 25 (13%) | 6 (16%) | 6 (12%) | |
| therapy related | 16 (5.7%) | 14 (7.3%) | 1 (2.6%) | 1 (2.0%) | |
| AML secondary, n (%) | 0.71 | ||||
| MDS | 24 (65%) | 16 (64%) | 4 (67%) | 4 (67%) | |
| ET | 0 (0%) | 0 (0%) | 0 (0%) | 0 (0%) | |
| PV | 3 (8.1%) | 3 (12%) | 0 (0%) | 0 (0%) | |
| MF | 5 (14%) | 2 (8.0%) | 1 (17%) | 2 (33%) | |
| FLT3, n (%) | 0.177 | ||||
| ITD | 48 (18%) | 29 (15%) | 13 (38%) | 6 (14%) | |
| TKD | 10 (3.7%) | 9 (4.6%) | 1 (2.9%) | 0 (0%) | |
| ITD and TKD | 2 (0.7%) | 2 (1.0%) | 0 (0%) | 0 (0%) | |
| Mutated NPM1, n (%) | 71 (27%) | 44 (23%) | 17 (50%) | 10 (25%) | 0.007 |
| Mutated TP53, n (%) | 1 (6.2%) | 1 (8.3%) | 0 (NA%) | 0 (0%) | >0.99 |
| Mutated CEBPA, n (%) | 3 (9.7%) | 2 (15%) | 1 (10%) | 0 (0%) | 0.77 |
| Mutated IDH1, n (%) | 38 (13%) | 0 (0%) | 38 (100%) | 0 (0%) | <0.001 |
| Mutated IDH2, n (%) | 51 (18%) | 0 (0%) | 0 (0%) | 51 (100%) | <0.001 |
| Complex karyotype, n (%) | 29 (11%) | 26 (14%) | 0 (0%) | 3 (6.4%) | 0.021 |
| Treatment, n (%) | 0.071 | ||||
| Conventional CHT | 201 (71%) | 136 (70%) | 29 (76%) | 36 (71%) | |
| Hypomethylating | 76 (27%) | 57 (29%) | 8 (21%) | 11 (22%) | |
| AML | ||||||
| Univariate | Multivariate | |||||
| Characteristic | HR 1 | 95% CI 1 | p-Value | HR 1 | 95% CI 1 | p-Value |
| Age | 1.04 | 1.02, 1.05 | <0.001 | 1.03 | 1.01, 1.05 | 0.002 |
| WBC | 1.00 | 1.00, 1.00 | 0.004 | 1.00 | 1.00, 1.00 | 0.026 |
| WHO PS | ||||||
| 0 | – | — | ||||
| I | 1.96 | 1.34, 2.88 | <0.001 | 1.65 | 1.09, 2.49 | 0.018 |
| II | 2.70 | 1.63, 4.47 | <0.001 | 2.45 | 1.32, 4.53 | 0.005 |
| III | 0.70 | 0.17, 2.88 | 0.62 | 0.54 | 0.07, 3.99 | 0.55 |
| Complex karyotype vs. other | ||||||
| other karyotype | - | - | - | - | ||
| complex karyotype | 2.83 | 1.79, 4.45 | <0.001 | 3.17 | 1.91, 5.26 | <0.001 |
| Treatment | ||||||
| Standard CHT | — | — | ||||
| Hypomethylating | 2.20 | 1.56, 3.10 | <0.001 | 1.07 | 0.66, 1.76 | 0.78 |
| IDH1/2-mutated AML | ||||||
| Univariate | Multivariate | |||||
| Characteristic | HR 1 | 95% CI 1 | p-Value | HR 1 | 95% CI 1 | p-Value 2 |
| Age | 1.03 | 1.00, 1.06 | 0.019 | |||
| WBC | 1.00 | 1.00, 1.00 | 0.007 | 1.00 | 1.00, 1.01 | 0.005 |
| Hb | 0.84 | 0.70, 1.00 | 0.049 | |||
| Treatment | ||||||
| Standard CHT | — | — | ||||
| Hypomethylating | 2.03 | 1.07, 3.83 | 0.03 | 1.09 | 0.48, 2.48 | 0.84 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Messina, M.; Piciocchi, A.; Ottone, T.; Paolini, S.; Papayannidis, C.; Lessi, F.; Fracchiolla, N.S.; Forghieri, F.; Candoni, A.; Mengarelli, A.; et al. Prevalence and Prognostic Role of IDH Mutations in Acute Myeloid Leukemia: Results of the GIMEMA AML1516 Protocol. Cancers 2022, 14, 3012. https://doi.org/10.3390/cancers14123012
Messina M, Piciocchi A, Ottone T, Paolini S, Papayannidis C, Lessi F, Fracchiolla NS, Forghieri F, Candoni A, Mengarelli A, et al. Prevalence and Prognostic Role of IDH Mutations in Acute Myeloid Leukemia: Results of the GIMEMA AML1516 Protocol. Cancers. 2022; 14(12):3012. https://doi.org/10.3390/cancers14123012
Chicago/Turabian StyleMessina, Monica, Alfonso Piciocchi, Tiziana Ottone, Stefania Paolini, Cristina Papayannidis, Federica Lessi, Nicola Stefano Fracchiolla, Fabio Forghieri, Anna Candoni, Andrea Mengarelli, and et al. 2022. "Prevalence and Prognostic Role of IDH Mutations in Acute Myeloid Leukemia: Results of the GIMEMA AML1516 Protocol" Cancers 14, no. 12: 3012. https://doi.org/10.3390/cancers14123012
APA StyleMessina, M., Piciocchi, A., Ottone, T., Paolini, S., Papayannidis, C., Lessi, F., Fracchiolla, N. S., Forghieri, F., Candoni, A., Mengarelli, A., Martelli, M. P., Venditti, A., Carella, A. M., Albano, F., Mancini, V., Massimo, B., Arena, V., Sargentini, V., Sciumè, M., ... Voso, M. T. (2022). Prevalence and Prognostic Role of IDH Mutations in Acute Myeloid Leukemia: Results of the GIMEMA AML1516 Protocol. Cancers, 14(12), 3012. https://doi.org/10.3390/cancers14123012











