The Mycobiome: Cancer Pathogenesis, Diagnosis, and Therapy
Abstract
Simple Summary
Abstract
1. Introduction
2. Role of the Mycobiome in Cancer Pathogenesis
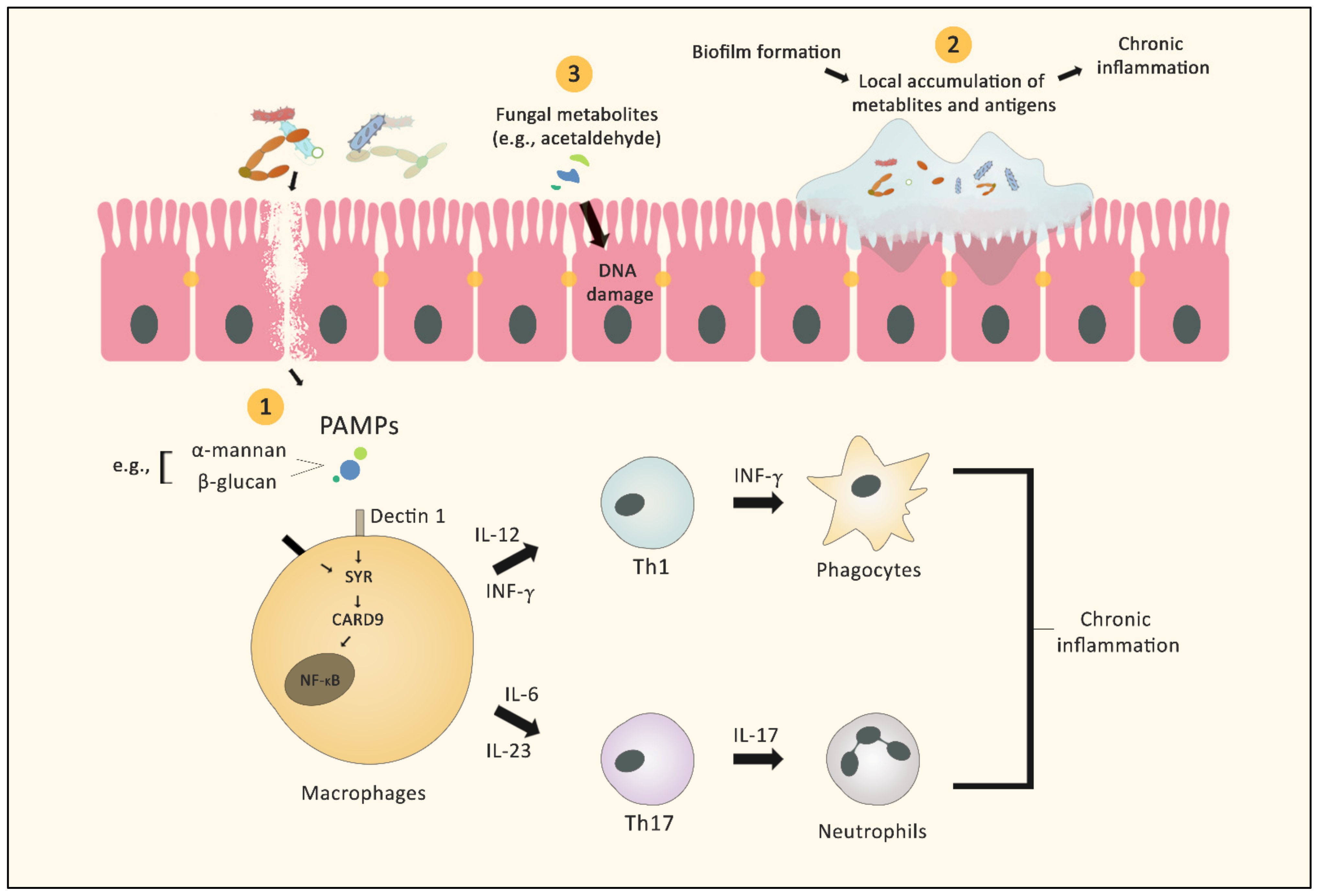
2.1. Microbial Inflammation
2.2. Biofilm Formation
2.3. Fungus-Derived Metabolites and Cancer
3. Diagnostic Implications of the Mycobiome in Cancer Patients
3.1. Cancers of the Gastrointestinal Tract
3.2. Cancers beyond the Gastrointestinal Tract
4. Mycobiome and Cancer Therapy
4.1. Effect of Mycobiome on Anticancer Therapy
4.1.1. Tumor Microenvironment Modification
4.1.2. Probiotics as Adjuvant Therapy
4.1.3. Effect on Radiotherapy
4.1.4. Fungal Metabolites
4.1.5. Other Promising Treatments
4.2. Effect of Anticancer Therapy on the Mycobiome
5. Future Perspectives
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Piñeros, M.; Mery, L.; Soerjomataram, I.; Bray, F.; Steliarova-Foucher, E. Scaling up the surveillance of childhood cancer: A global roadmap. J. Natl. Cancer Inst. 2021, 113, 9–15. [Google Scholar] [CrossRef] [PubMed]
- Garrett, W.S. Cancer and the microbiota. Science 2015, 348, 80–86. [Google Scholar] [CrossRef] [PubMed]
- Sears, C.L.; Garrett, W.S. Microbes, microbiota, and colon cancer. Cell Host Microbe 2014, 15, 317–328. [Google Scholar] [CrossRef] [PubMed]
- Vemuri, R.; Shankar, E.M.; Chieppa, M.; Eri, R.; Kavanagh, K. Beyond Just Bacteria: Functional Biomes in the Gut Ecosystem Including Virome, Mycobiome, Archaeome and Helminths. Microorganisms 2020, 8, 483. [Google Scholar] [CrossRef] [PubMed]
- Matijasic, M.; Mestrovic, T.; Paljetak, H.C.; Peric, M.; Baresic, A.; Verbanac, D. Gut Microbiota beyond Bacteria-Mycobiome, Virome, Archaeome, and Eukaryotic Parasites in IBD. Int. J. Mol. Sci. 2020, 21, 2668. [Google Scholar] [CrossRef] [PubMed]
- Pattaroni, C.; Macowan, M.; Chatzis, R.; Daunt, C.; Custovic, A.; Shields, M.D.; Power, U.F.; Grigg, J.; Roberts, G.; Ghazal, P.; et al. Early life inter-kingdom interactions shape the immunological environment of the airways. Microbiome 2022, 10, 34. [Google Scholar] [CrossRef]
- Arvanitis, M.; Mylonakis, E. Fungal-bacterial interactions and their relevance in health. Cell Microbiol. 2015, 17, 1442–1446. [Google Scholar] [CrossRef]
- Coker, O.O.; Nakatsu, G.; Dai, R.Z.; Wu, W.K.K.; Wong, S.H.; Ng, S.C.; Chan, F.K.L.; Sung, J.J.Y.; Yu, J. Enteric fungal microbiota dysbiosis and ecological alterations in colorectal cancer. Gut 2019, 68, 654–662. [Google Scholar] [CrossRef]
- Seed, P.C. The human mycobiome. Cold Spring Harb. Perspect. Med. 2015, 5, a019810. [Google Scholar] [CrossRef]
- Zhang, E.; Tanaka, T.; Tajima, M.; Tsuboi, R.; Nishikawa, A.; Sugita, T. Characterization of the skin fungal microbiota in patients with atopic dermatitis and in healthy subjects. Microbiol. Immunol. 2011, 55, 625–632. [Google Scholar] [CrossRef]
- Nguyen, L.D.; Viscogliosi, E.; Delhaes, L. The lung mycobiome: An emerging field of the human respiratory microbiome. Front. Microbiol. 2015, 6, 89. [Google Scholar] [CrossRef] [PubMed]
- Ghannoum, M.A.; Jurevic, R.J.; Mukherjee, P.K.; Cui, F.; Sikaroodi, M.; Naqvi, A.; Gillevet, P.M. Characterization of the oral fungal microbiome (mycobiome) in healthy individuals. PLoS Pathog. 2010, 6, e1000713. [Google Scholar] [CrossRef] [PubMed]
- Drell, T.; Lillsaar, T.; Tummeleht, L.; Simm, J.; Aaspollu, A.; Vain, E.; Saarma, I.; Salumets, A.; Donders, G.G.; Metsis, M. Characterization of the vaginal micro- and mycobiome in asymptomatic reproductive-age Estonian women. PLoS ONE 2013, 8, e54379. [Google Scholar] [CrossRef] [PubMed]
- Huseyin, C.E.; Rubio, R.C.; O’Sullivan, O.; Cotter, P.D.; Scanlan, P.D. The Fungal Frontier: A Comparative Analysis of Methods Used in the Study of the Human Gut Mycobiome. Front. Microbiol. 2017, 8, 1432. [Google Scholar] [CrossRef] [PubMed]
- Witherden, E.A.; Moyes, D.L. Mycobiome and gut inflammation: Implications in gut disease. In Immunity and Inflammation in Health and Disease; Chatterjee, S., Jungraithmayr, W., Bagchi, D., Eds.; Academic Press: Cambridge, MA, USA, 2018; Chapter 22; pp. 271–280. [Google Scholar]
- Hoffmann, C.; Dollive, S.; Grunberg, S.; Chen, J.; Li, H.; Wu, G.D.; Lewis, J.D.; Bushman, F.D. Archaea and fungi of the human gut microbiome: Correlations with diet and bacterial residents. PLoS ONE 2013, 8, e66019. [Google Scholar] [CrossRef]
- van Woerden, H.C.; Gregory, C.; Brown, R.; Marchesi, J.R.; Hoogendoorn, B.; Matthews, I.P. Differences in fungi present in induced sputum samples from asthma patients and non-atopic controls: A community based case control study. BMC Infect. Dis. 2013, 13, 69. [Google Scholar] [CrossRef]
- Paterson, M.J.; Oh, S.; Underhill, D.M. Host–microbe interactions: Commensal fungi in the gut. Curr. Opin. Microbiol. 2017, 40, 131–137. [Google Scholar] [CrossRef]
- Richard, M.L.; Sokol, H. The gut mycobiota: Insights into analysis, environmental interactions and role in gastrointestinal diseases. Nat. Rev. Gastroenterol. Hepatol. 2019, 16, 331–345. [Google Scholar] [CrossRef]
- Vallianou, N.; Kounatidis, D.; Christodoulatos, G.S.; Panagopoulos, F.; Karampela, I.; Dalamaga, M. Mycobiome and Cancer: What Is the Evidence? Cancers 2021, 13, 3149. [Google Scholar] [CrossRef]
- Luan, C.; Xie, L.; Yang, X.; Miao, H.; Lv, N.; Zhang, R.; Xiao, X.; Hu, Y.; Liu, Y.; Wu, N.; et al. Dysbiosis of fungal microbiota in the intestinal mucosa of patients with colorectal adenomas. Sci. Rep. 2015, 5, 7980. [Google Scholar] [CrossRef]
- Gao, R.; Kong, C.; Li, H.; Huang, L.; Qu, X.; Qin, N.; Qin, H. Dysbiosis signature of mycobiota in colon polyp and colorectal cancer. Eur. J. Clin. Microbiol. Infect. Dis. 2017, 36, 2457–2468. [Google Scholar] [CrossRef] [PubMed]
- Richard, M.L.; Liguori, G.; Lamas, B.; Brandi, G.; da Costa, G.; Hoffmann, T.W.; Pierluigi Di Simone, M.; Calabrese, C.; Poggioli, G.; Langella, P.; et al. Mucosa-associated microbiota dysbiosis in colitis associated cancer. Gut Microbes 2018, 9, 131–142. [Google Scholar] [CrossRef]
- Chin, S.-F.; Megat Mohd Azlan, P.I.H.; Mazlan, L.; Neoh, H.-m. Identification of Schizosaccharomyces pombe in the guts of healthy individuals and patients with colorectal cancer: Preliminary evidence from a gut microbiome secretome study. Gut Pathog. 2018, 10, 29. [Google Scholar] [CrossRef] [PubMed]
- Wang, T.; Fan, C.; Yao, A.; Xu, X.; Zheng, G.; You, Y.; Jiang, C.; Zhao, X.; Hou, Y.; Hung, M.-C. The adaptor protein CARD9 protects against colon cancer by restricting mycobiota-mediated expansion of myeloid-derived suppressor cells. Immunity 2018, 49, 504–514.e504. [Google Scholar] [CrossRef] [PubMed]
- Aykut, B.; Pushalkar, S.; Chen, R.; Li, Q.; Abengozar, R.; Kim, J.I.; Shadaloey, S.A.; Wu, D.; Preiss, P.; Verma, N.; et al. The fungal mycobiome promotes pancreatic oncogenesis via activation of MBL. Nature 2019, 574, 264–267. [Google Scholar] [CrossRef]
- Zhong, M.; Xiong, Y.; Zhao, J.; Gao, Z.; Ma, J.; Wu, Z.; Song, Y.; Hong, X. Candida albicans disorder is associated with gastric carcinogenesis. Theranostics 2021, 11, 4945–4956. [Google Scholar] [CrossRef]
- Perera, M.; Al-Hebshi, N.N.; Perera, I.; Ipe, D.; Ulett, G.C.; Speicher, D.J.; Chen, T.; Johnson, N.W. A dysbiotic mycobiome dominated by Candida albicans is identified within oral squamous-cell carcinomas. J. Oral. Microbiol. 2017, 9, 1385369. [Google Scholar] [CrossRef]
- Vesty, A.; Gear, K.; Biswas, K.; Radcliff, F.J.; Taylor, M.W.; Douglas, R.G. Microbial and inflammatory-based salivary biomarkers of head and neck squamous cell carcinoma. Clin. Exp. Dent. Res. 2018, 4, 255–262. [Google Scholar] [CrossRef]
- Shay, E.; Sangwan, N.; Padmanabhan, R.; Lundy, S.; Burkey, B.; Eng, C. Bacteriome and mycobiome and bacteriome-mycobiome interactions in head and neck squamous cell carcinoma. Oncotarget 2020, 11, 2375–2386. [Google Scholar] [CrossRef]
- Mäkinen, A.; Nawaz, A.; Mäkitie, A.A.; Meurman, J.H. Role of Non-Albicans Candida and Candida Albicans in Oral Squamous Cell Cancer Patients. J. Oral Maxillofac. Surg. 2018, 76, 2564–2571. [Google Scholar] [CrossRef]
- Dongari-Bagtzoglou, A.; Kashleva, H. Candida albicans triggers interleukin-8 secretion by oral epithelial cells. Microb. Pathog. 2003, 34, 169–177. [Google Scholar] [CrossRef]
- Arzmi, M.H.; Dashper, S.; McCullough, M. Polymicrobial interactions of Candida albicans and its role in oral carcinogenesis. J. Oral Pathol. Med. 2019, 48, 546–551. [Google Scholar] [CrossRef] [PubMed]
- Greten, F.R.; Grivennikov, S.I. Inflammation and Cancer: Triggers, Mechanisms, and Consequences. Immunity 2019, 51, 27–41. [Google Scholar] [CrossRef]
- Underhill, D.M.; Iliev, I.D. The mycobiota: Interactions between commensal fungi and the host immune system. Nat. Rev. Immunol. 2014, 14, 405–416. [Google Scholar] [CrossRef]
- Qin, X.; Gu, Y.; Liu, T.; Wang, C.; Zhong, W.; Wang, B.; Cao, H. Gut mycobiome: A promising target for colorectal cancer. Biochim. Biophys. Acta (BBA) Rev. Cancer 2020, 1875, 188489. [Google Scholar] [CrossRef]
- Li, S.S.; Ogbomo, H.; Mansour, M.K.; Xiang, R.F.; Szabo, L.; Munro, F.; Mukherjee, P.; Mariuzza, R.A.; Amrein, M.; Vyas, J.M.; et al. Identification of the fungal ligand triggering cytotoxic PRR-mediated NK cell killing of Cryptococcus and Candida. Nat. Commun. 2018, 9, 751. [Google Scholar] [CrossRef] [PubMed]
- Drummond, R.A.; Franco, L.M.; Lionakis, M.S. Human CARD9: A Critical Molecule of Fungal Immune Surveillance. Front. Immunol. 2018, 9, 1836. [Google Scholar] [CrossRef]
- Wang, Y.; Zhang, D.; Hou, Y.; Shen, S.; Wang, T. The adaptor protein CARD9, from fungal immunity to tumorigenesis. Am. J. Cancer Res. 2020, 10, 2203–2225. [Google Scholar]
- Salcedo, R.; Worschech, A.; Cardone, M.; Jones, Y.; Gyulai, Z.; Dai, R.M.; Wang, E.; Ma, W.; Haines, D.; O’HUigin, C.; et al. MyD88-mediated signaling prevents development of adenocarcinomas of the colon: Role of interleukin 18. J. Exp. Med. 2010, 207, 1625–1636. [Google Scholar] [CrossRef]
- Khor, B.; Gardet, A.; Xavier, R.J. Genetics and pathogenesis of inflammatory bowel disease. Nature 2011, 474, 307–317. [Google Scholar] [CrossRef]
- Velcich, A.; Yang, W.; Heyer, J.; Fragale, A.; Nicholas, C.; Viani, S.; Kucherlapati, R.; Lipkin, M.; Yang, K.; Augenlicht, L. Colorectal cancer in mice genetically deficient in the mucin Muc2. Science 2002, 295, 1726–1729. [Google Scholar] [CrossRef] [PubMed]
- Moyes, D.L.; Wilson, D.; Richardson, J.P.; Mogavero, S.; Tang, S.X.; Wernecke, J.; Höfs, S.; Gratacap, R.L.; Robbins, J.; Runglall, M.; et al. Candidalysin is a fungal peptide toxin critical for mucosal infection. Nature 2016, 532, 64–68. [Google Scholar] [CrossRef] [PubMed]
- Ho, J.; Yang, X.; Nikou, S.A.; Kichik, N.; Donkin, A.; Ponde, N.O.; Richardson, J.P.; Gratacap, R.L.; Archambault, L.S.; Zwirner, C.P.; et al. Candidalysin activates innate epithelial immune responses via epidermal growth factor receptor. Nat. Commun. 2019, 10, 2297. [Google Scholar] [CrossRef] [PubMed]
- Vandenbroucke, R.E.; Libert, C. Is there new hope for therapeutic matrix metalloproteinase inhibition? Nat Rev Drug Discov 2014, 13, 904–927. [Google Scholar] [CrossRef]
- Sigismund, S.; Avanzato, D.; Lanzetti, L. Emerging functions of the EGFR in cancer. Mol. Oncol. 2018, 12, 3–20. [Google Scholar] [CrossRef]
- Román, E.; Correia, I.; Prieto, D.; Alonso, R.; Pla, J. The HOG MAPK pathway in Candida albicans: More than an osmosensing pathway. Int. Microbiol. 2020, 23, 23–29. [Google Scholar] [CrossRef]
- Frank, C.F.; Hostetter, M.K. Cleavage of E-cadherin: A mechanism for disruption of the intestinal epithelial barrier by Candida albicans. Transl. Res. 2007, 149, 211–222. [Google Scholar] [CrossRef]
- Ashman, R.B.; Papadimitriou, J.M. Endothelial cell proliferation associated with lesions of murine systemic candidiasis. Infect. Immun. 1994, 62, 5151–5153. [Google Scholar] [CrossRef]
- Barker, K.S.; Park, H.; Phan, Q.T.; Xu, L.; Homayouni, R.; Rogers, P.D.; Filler, S.G. Transcriptome profile of the vascular endothelial cell response to Candida albicans. J. Infect. Dis. 2008, 198, 193–202. [Google Scholar] [CrossRef]
- Vellanki, S.; Huh, E.Y.; Saville, S.P.; Lee, S.C. Candida albicans Morphology-Dependent Host FGF-2 Response as a Potential Therapeutic Target. J. Fungi 2019, 5, 22. [Google Scholar] [CrossRef]
- Masters, E.A.; Trombetta, R.P.; de Mesy Bentley, K.L.; Boyce, B.F.; Gill, A.L.; Gill, S.R.; Nishitani, K.; Ishikawa, M.; Morita, Y.; Ito, H.; et al. Evolving concepts in bone infection: Redefining “biofilm”, “acute vs. chronic osteomyelitis”, “the immune proteome” and “local antibiotic therapy”. Bone Res. 2019, 7, 20. [Google Scholar] [CrossRef] [PubMed]
- Bjarnsholt, T. The role of bacterial biofilms in chronic infections. APMIS 2013, 121, 1–58. [Google Scholar] [CrossRef] [PubMed]
- Khatoon, Z.; McTiernan, C.D.; Suuronen, E.J.; Mah, T.-F.; Alarcon, E.I. Bacterial biofilm formation on implantable devices and approaches to its treatment and prevention. Heliyon 2018, 4, e01067. [Google Scholar] [CrossRef] [PubMed]
- Hager, C.L.; Ghannoum, M.A. The mycobiome: Role in health and disease, and as a potential probiotic target in gastrointestinal disease. Dig. Liver Dis. 2017, 49, 1171–1176. [Google Scholar] [CrossRef]
- Tomkovich, S.; Dejea, C.M.; Winglee, K.; Drewes, J.L.; Chung, L.; Housseau, F.; Pope, J.L.; Gauthier, J.; Sun, X.; Muhlbauer, M.; et al. Human colon mucosal biofilms from healthy or colon cancer hosts are carcinogenic. J. Clin. Invest. 2019, 129, 1699–1712. [Google Scholar] [CrossRef]
- Sovran, B.; Planchais, J.; Jegou, S.; Straube, M.; Lamas, B.; Natividad, J.M.; Agus, A.; Dupraz, L.; Glodt, J.; Da Costa, G.; et al. Enterobacteriaceae are essential for the modulation of colitis severity by fungi. Microbiome 2018, 6, 152. [Google Scholar] [CrossRef]
- Hager, C.L.; Isham, N.; Schrom, K.P.; Chandra, J.; McCormick, T.; Miyagi, M.; Ghannoum, M.A. Effects of a Novel Probiotic Combination on Pathogenic Bacterial-Fungal Polymicrobial Biofilms. mBio 2019, 10, e00338-19. [Google Scholar] [CrossRef]
- Sun, C.H.; Li, B.B.; Wang, B.; Zhao, J.; Zhang, X.Y.; Li, T.T.; Li, W.B.; Tang, D.; Qiu, M.J.; Wang, X.C.; et al. The role of Fusobacterium nucleatum in colorectal cancer: From carcinogenesis to clinical management. Chronic Dis. Transl. Med. 2019, 5, 178–187. [Google Scholar] [CrossRef]
- Wu, J.; Li, Q.; Fu, X. Fusobacterium nucleatum Contributes to the Carcinogenesis of Colorectal Cancer by Inducing Inflammation and Suppressing Host Immunity. Transl. Oncol. 2019, 12, 846–851. [Google Scholar] [CrossRef]
- Peleg, A.Y.; Hogan, D.A.; Mylonakis, E. Medically important bacterial-fungal interactions. Nat. Rev. Microbiol. 2010, 8, 340–349. [Google Scholar] [CrossRef]
- Baena-Monroy, T.; Moreno-Maldonado, V.; Franco-Martinez, F.; Aldape-Barrios, B.; Quindos, G.; Sanchez-Vargas, L.O. Candida albicans, Staphylococcus aureus and Streptococcus mutans colonization in patients wearing dental prosthesis. Med. Oral Patol. Oral Cir. Bucal 2005, 10 (Suppl. 1), E27–E39. [Google Scholar] [PubMed]
- Gupta, N.; Haque, A.; Mukhopadhyay, G.; Narayan, R.P.; Prasad, R. Interactions between bacteria and Candida in the burn wound. Burns 2005, 31, 375–378. [Google Scholar] [CrossRef] [PubMed]
- Allison, D.L.; Willems, H.M.E.; Jayatilake, J.; Bruno, V.M.; Peters, B.M.; Shirtliff, M.E. Candida-Bacteria Interactions: Their Impact on Human Disease. Microbiol. Spectr. 2016, 4. [Google Scholar] [CrossRef] [PubMed]
- Hogan, D.A.; Vik, A.; Kolter, R. A Pseudomonas aeruginosa quorum-sensing molecule influences Candida albicans morphology. Mol. Microbiol. 2004, 54, 1212–1223. [Google Scholar] [CrossRef]
- Ikeda, T.; Suegara, N.; Abe, S.; Yamaguchi, H. Efficacy of antibacterial drugs in mice with complex infection by Candida albicans and Escherichia coli. J. Antibiot. 1999, 52, 552–558. [Google Scholar] [CrossRef][Green Version]
- Yang, W.; Zhou, Y.; Wu, C.; Tang, J. Enterohemorrhagic Escherichia coli promotes the invasion and tissue damage of enterocytes infected with Candida albicans in vitro. Sci. Rep. 2016, 6, 37485. [Google Scholar] [CrossRef]
- Liu, M.; Luo, F.; Ding, C.; Albeituni, S.; Hu, X.; Ma, Y.; Cai, Y.; McNally, L.; Sanders, M.A.; Jain, D.; et al. Dectin-1 Activation by a Natural Product beta-Glucan Converts Immunosuppressive Macrophages into an M1-like Phenotype. J. Immunol. 2015, 195, 5055–5065. [Google Scholar] [CrossRef]
- Lambooij, J.M.; Hoogenkamp, M.A.; Brandt, B.W.; Janus, M.M.; Krom, B.P. Fungal mitochondrial oxygen consumption induces the growth of strict anaerobic bacteria. Fungal Genet. Biol. 2017, 109, 1–6. [Google Scholar] [CrossRef]
- Sánchez-Alonzo, K.; Parra-Sepúlveda, C.; Vega, S.; Bernasconi, H.; Campos, V.L.; Smith, C.T.; Sáez, K.; García-Cancino, A. In Vitro Incorporation of Helicobacter pylori into Candida albicans Caused by Acidic pH Stress. Pathogens 2020, 9, 489. [Google Scholar] [CrossRef]
- Santus, W.; Devlin, J.R.; Behnsen, J. Crossing Kingdoms: How the Mycobiota and Fungal-Bacterial Interactions Impact Host Health and Disease. Infect. Immun. 2021, 89, e00648-20. [Google Scholar] [CrossRef]
- van Leeuwen, P.T.; van der Peet, J.M.; Bikker, F.J.; Hoogenkamp, M.A.; Oliveira Paiva, A.M.; Kostidis, S.; Mayboroda, O.A.; Smits, W.K.; Krom, B.P. Interspecies Interactions between Clostridium difficile and Candida albicans. mSphere 2016, 1, e00187-16. [Google Scholar] [CrossRef]
- Nieminen, M.T.; Uittamo, J.; Salaspuro, M.; Rautemaa, R. Acetaldehyde production from ethanol and glucose by non-Candida albicans yeasts in vitro. Oral Oncol 2009, 45, e245–e248. [Google Scholar] [CrossRef] [PubMed]
- Uittamo, J.; Siikala, E.; Kaihovaara, P.; Salaspuro, M.; Rautemaa, R. Chronic candidosis and oral cancer in APECED-patients: Production of carcinogenic acetaldehyde from glucose and ethanol by Candida albicans. Int. J. Cancer 2009, 124, 754–756. [Google Scholar] [CrossRef]
- Tillonen, J.; Homann, N.; Rautio, M.; Jousimies-Somer, H.; Salaspuro, M. Role of Yeasts in the Salivary Acetaldehyde Production from Ethanol Among Risk Groups for Ethanol-Associated Oral Cavity Cancer. Alcohol. Clin. Exp. Res. 1999, 23, 1409–1411. [Google Scholar] [CrossRef] [PubMed]
- IARC Working Group on the Evaluation of Carcinogenic Risks to Humans. Chemical Agents and Related Occupations; International Agency for Research on Cancer: Lyon, France, 2012; Volume 100, pp. 9–562. [Google Scholar]
- Vakevainen, S.; Tillonen, J.; Agarwal, D.P.; Srivastava, N.; Salaspuro, M. High salivary acetaldehyde after a moderate dose of alcohol in ALDH2-deficient subjects: Strong evidence for the local carcinogenic action of acetaldehyde. Alcohol. Clin. Exp. Res. 2000, 24, 873–877. [Google Scholar] [CrossRef] [PubMed]
- Salaspuro, M.P. Acetaldehyde, microbes, and cancer of the digestive tract. Crit. Rev. Clin. Lab. Sci. 2003, 40, 183–208. [Google Scholar] [CrossRef]
- Homann, N.; Tillonen, J.; Meurman, J.H.; Rintamaki, H.; Lindqvist, C.; Rautio, M.; Jousimies-Somer, H.; Salaspuro, M. Increased salivary acetaldehyde levels in heavy drinkers and smokers: A microbiological approach to oral cavity cancer. Carcinogenesis 2000, 21, 663–668. [Google Scholar] [CrossRef]
- Mukherjee, P.K.; Mohamed, S.; Chandra, J.; Kuhn, D.; Liu, S.; Antar, O.S.; Munyon, R.; Mitchell, A.P.; Andes, D.; Chance, M.R.; et al. Alcohol dehydrogenase restricts the ability of the pathogen Candida albicans to form a biofilm on catheter surfaces through an ethanol-based mechanism. Infect. Immun. 2006, 74, 3804–3816. [Google Scholar] [CrossRef]
- Sankari, S.L.; Gayathri, K.; Balachander, N.; Malathi, L. Candida in potentially malignant oral disorders. J. Pharm. Bioallied. Sci. 2015, 7, S162–S164. [Google Scholar] [CrossRef]
- Liew, W.P.; Mohd-Redzwan, S. Mycotoxin: Its Impact on Gut Health and Microbiota. Front. Cell Infect. Microbiol. 2018, 8, 60. [Google Scholar] [CrossRef]
- Tang, F.; Yuan, J.; Yuan, B.F.; Wang, Y. DNA-Protein Cross-Linking Sequencing for Genome-Wide Mapping of Thymidine Glycol. J. Am. Chem. Soc. 2022, 144, 454–462. [Google Scholar] [CrossRef] [PubMed]
- Tang, F.; Liu, S.; Li, Q.Y.; Yuan, J.; Li, L.; Wang, Y.; Yuan, B.F.; Feng, Y.Q. Location analysis of 8-oxo-7,8-dihydroguanine in DNA by polymerase-mediated differential coding. Chem. Sci. 2019, 10, 4272–4281. [Google Scholar] [CrossRef] [PubMed]
- Puel, O.; Galtier, P.; Oswald, I.P. Biosynthesis and toxicological effects of patulin. Toxins 2010, 2, 613–631. [Google Scholar] [CrossRef]
- Barhoumi, R.; Burghardt, R.C. Kinetic analysis of the chronology of patulin- and gossypol-induced cytotoxicity in vitro. Fundam. Appl. Toxicol. 1996, 30, 290–297. [Google Scholar] [CrossRef]
- Liu, B.H.; Wu, T.S.; Yu, F.Y.; Su, C.C. Induction of oxidative stress response by the mycotoxin patulin in mammalian cells. Toxicol. Sci. 2007, 95, 340–347. [Google Scholar] [CrossRef]
- de Jager, P.; Burgerhof, J.G.; Koopman, A.A.; Markhorst, D.G.; Kneyber, M.C. Physiologic responses to a staircase lung volume optimization maneuver in pediatric high-frequency oscillatory ventilation. Ann. Intensive Care 2020, 10, 153. [Google Scholar] [CrossRef] [PubMed]
- Saxena, N.; Ansari, K.M.; Kumar, R.; Chaudhari, B.P.; Dwivedi, P.D.; Das, M. Role of mitogen activated protein kinases in skin tumorigenicity of patulin. Toxicol. Appl. Pharm. 2011, 257, 264–271. [Google Scholar] [CrossRef]
- Chilaka, C.A.; De Boevre, M.; Atanda, O.O.; De Saeger, S. The Status of Fusarium Mycotoxins in Sub-Saharan Africa: A Review of Emerging Trends and Post-Harvest Mitigation Strategies towards Food Control. Toxins 2017, 9, 19. [Google Scholar] [CrossRef]
- Minervini, F.; Garbetta, A.; D’Antuono, I.; Cardinali, A.; Martino, N.A.; Debellis, L.; Visconti, A. Toxic mechanisms induced by fumonisin b1 mycotoxin on human intestinal cell line. Arch. Environ. Contam. Toxicol. 2014, 67, 115–123. [Google Scholar] [CrossRef]
- Angius, F.; Spolitu, S.; Uda, S.; Deligia, S.; Frau, A.; Banni, S.; Collu, M.; Accossu, S.; Madeddu, C.; Serpe, R.; et al. High-density lipoprotein contribute to G0-G1/S transition in Swiss NIH/3T3 fibroblasts. Sci. Rep. 2015, 5, 17812. [Google Scholar] [CrossRef]
- Das, A.; O’Herlihy, E.; Shanahan, F.; O’Toole, P.W.; Jeffery, I.B. The fecal mycobiome in patients with Irritable Bowel Syndrome. Sci. Rep. 2021, 11, 124. [Google Scholar] [CrossRef]
- Chen, Q.; Zhang, B.; Xu, H.; Ren, J.; Zhang, X. IDDF2019-ABS-0226 The potential intestinal fungal biomarkers in patients with colonic polyps. Gut 2019, 68, A105. [Google Scholar] [CrossRef]
- Banerjee, S.; Tian, T.; Wei, Z.; Peck, K.N.; Shih, N.; Chalian, A.A.; O’Malley, B.W.; Weinstein, G.S.; Feldman, M.D.; Alwine, J.; et al. Microbial Signatures Associated with Oropharyngeal and Oral Squamous Cell Carcinomas. Sci. Rep. 2017, 7, 4036. [Google Scholar] [CrossRef] [PubMed]
- Mukherjee, P.K.; Wang, H.; Retuerto, M.; Zhang, H.; Burkey, B.; Ghannoum, M.A.; Eng, C. Bacteriome and mycobiome associations in oral tongue cancer. Oncotarget 2017, 8, 97273–97289. [Google Scholar] [CrossRef] [PubMed]
- Banerjee, S.; Tian, T.; Wei, Z.; Shih, N.; Feldman, M.D.; Alwine, J.C.; Coukos, G.; Robertson, E.S. The ovarian cancer oncobiome. Oncotarget 2017, 8, 36225–36245. [Google Scholar] [CrossRef]
- Banerjee, S.; Wei, Z.; Tian, T.; Bose, D.; Shih, N.N.C.; Feldman, M.D.; Khoury, T.; De Michele, A.; Robertson, E.S. Prognostic correlations with the microbiome of breast cancer subtypes. Cell Death Dis. 2021, 12, 831. [Google Scholar] [CrossRef]
- Vitali, F.; Colucci, R.; Di Paola, M.; Pindo, M.; De Filippo, C.; Moretti, S.; Cavalieri, D. Early melanoma invasivity correlates with gut fungal and bacterial profiles. Br. J. Dermathol. 2022, 186, 106–116. [Google Scholar] [CrossRef]
- Ahmed, N.; Ghannoum, M.; Gallogly, M.; de Lima, M.; Malek, E. Influence of gut microbiome on multiple myeloma: Friend or foe? J. Immunother. Cancer. 2020, 8, e000576. [Google Scholar] [CrossRef]
- Chung, L.M.; Liang, J.A.; Lin, C.L.; Sun, L.M.; Kao, C.H. Cancer risk in patients with candidiasis: A nationwide population-based cohort study. Oncotarget 2017, 8, 63562–63573. [Google Scholar] [CrossRef]
- Chimonidou, M.; Strati, A.; Tzitzira, A.; Sotiropoulou, G.; Malamos, N.; Georgoulias, V.; Lianidou, E.S. DNA methylation of tumor suppressor and metastasis suppressor genes in circulating tumor cells. Clin. Chem. 2011, 57, 1169–1177. [Google Scholar] [CrossRef]
- Shiao, S.L.; Kershaw, K.M.; Limon, J.J.; You, S.; Yoon, J.; Ko, E.Y.; Guarnerio, J.; Potdar, A.A.; McGovern, D.P.B.; Bose, S.; et al. Commensal bacteria and fungi differentially regulate tumor responses to radiation therapy. Cancer Cell 2021, 39, 1202–1213.e1206. [Google Scholar] [CrossRef] [PubMed]
- El Jurdi, N.; Filali-Mouhim, A.; Salem, I.; Retuerto, M.; Dambrosio, N.M.; Baer, L.; Lazarus, H.M.; Caimi, P.; Cooper, B.; Tomlinson, B.; et al. Gastrointestinal Microbiome and Mycobiome Changes during Autologous Transplantation for Multiple Myeloma: Results of a Prospective Pilot Study. Biol. Blood Marrow Transpl. 2019, 25, 1511–1519. [Google Scholar] [CrossRef] [PubMed]
- Robinson, S.; Peterson, C.B.; Sahasrabhojane, P.; Ajami, N.J.; Shelburne, S.A.; Kontoyiannis, D.P.; Galloway-Pena, J.R. Observational Cohort Study of Oral Mycobiome and Interkingdom Interactions over the Course of Induction Therapy for Leukemia. mSphere 2020, 5, e00048-20. [Google Scholar] [CrossRef] [PubMed]
- Hong, B.Y.; Sobue, T.; Choquette, L.; Dupuy, A.K.; Thompson, A.; Burleson, J.A.; Salner, A.L.; Schauer, P.K.; Joshi, P.; Fox, E.; et al. Chemotherapy-induced oral mucositis is associated with detrimental bacterial dysbiosis. Microbiome 2019, 7, 66. [Google Scholar] [CrossRef]
- Hong, F.; Yan, J.; Baran, J.T.; Allendorf, D.J.; Hansen, R.D.; Ostroff, G.R.; Xing, P.X.; Cheung, N.K.; Ross, G.D. Mechanism by which orally administered beta-1,3-glucans enhance the tumoricidal activity of antitumor monoclonal antibodies in murine tumor models. J. Immunol. 2004, 173, 797–806. [Google Scholar] [CrossRef]
- Wang, W.J.; Wu, Y.S.; Chen, S.; Liu, C.F.; Chen, S.N. Mushroom beta-Glucan May Immunomodulate the Tumor-Associated Macrophages in the Lewis Lung Carcinoma. Biomed. Res. Int. 2015, 2015, 604385. [Google Scholar] [CrossRef]
- Demir, G.; Klein, H.O.; Mandel-Molinas, N.; Tuzuner, N. Beta glucan induces proliferation and activation of monocytes in peripheral blood of patients with advanced breast cancer. Int. Immunopharmacol. 2007, 7, 113–116. [Google Scholar] [CrossRef]
- Yoon, T.J.; Kim, T.J.; Lee, H.; Shin, K.S.; Yun, Y.P.; Moon, W.K.; Kim, D.W.; Lee, K.H. Anti-tumor metastatic activity of beta-glucan purified from mutated Saccharomyces cerevisiae. Int. Immunopharmacol. 2008, 8, 36–42. [Google Scholar] [CrossRef]
- Kim, W.-J.; Yoon, T.J.; Kim, D.-W.; Moon, W.-K.; Lee, K.H. Immunostimulating Activity of Beta-Glucan Isolated from the Cell Wall of Mutant Saccharomyces cerevisiae, and Its Anti-Tumor Application in Combination with Cisplatin. Korean J. Food Nutr. 2010, 23, 141–146. [Google Scholar]
- Vetvicka, V.; Vetvickova, J. Glucan Supplementation Has Strong Anti-melanoma Effects: Role of NK Cells. Anticancer Res. 2015, 35, 5287–5292. [Google Scholar]
- Zhang, M.; Chun, L.; Sandoval, V.; Graor, H.; Myers, J.; Nthale, J.; Rauhe, P.; Senders, Z.; Choong, K.; Huang, A.Y.; et al. Systemic administration of beta-glucan of 200 kDa modulates melanoma microenvironment and suppresses metastatic cancer. Oncoimmunology 2018, 7, e1387347. [Google Scholar] [CrossRef] [PubMed]
- Chan, A.S.; Jonas, A.B.; Qiu, X.; Ottoson, N.R.; Walsh, R.M.; Gorden, K.B.; Harrison, B.; Maimonis, P.J.; Leonardo, S.M.; Ertelt, K.E.; et al. Imprime PGG-Mediated Anti-Cancer Immune Activation Requires Immune Complex Formation. PLoS ONE 2016, 11, e0165909. [Google Scholar] [CrossRef] [PubMed]
- Qiu, X.; Chan, A.S.H.; Jonas, A.B.; Kangas, T.; Ottoson, N.R.; Graff, J.R.; Bose, N. Imprime PGG, a yeast β-glucan PAMP elicits a coordinated immune response in combination with anti-PD1 antibody. J. Immunol. 2016, 196, 214–216. [Google Scholar]
- Chen, X.; Fruehauf, J.; Goldsmith, J.D.; Xu, H.; Katchar, K.K.; Koon, H.W.; Zhao, D.; Kokkotou, E.G.; Pothoulakis, C.; Kelly, C.P. Saccharomyces boulardii inhibits EGF receptor signaling and intestinal tumor growth in Apc(min) mice. Gastroenterology 2009, 137, 914–923. [Google Scholar] [CrossRef] [PubMed]
- Chen, S.M.; Chieng, W.W.; Huang, S.W.; Hsu, L.J.; Jan, M.S. The synergistic tumor growth-inhibitory effect of probiotic Lactobacillus on transgenic mouse model of pancreatic cancer treated with gemcitabine. Sci. Rep. 2020, 10, 20319. [Google Scholar] [CrossRef]
- Davar, D.; Dzutsev, A.K.; McCulloch, J.A.; Rodrigues, R.R.; Chauvin, J.M.; Morrison, R.M.; Deblasio, R.N.; Menna, C.; Ding, Q.; Pagliano, O.; et al. Fecal microbiota transplant overcomes resistance to anti-PD-1 therapy in melanoma patients. Science 2021, 371, 595–602. [Google Scholar] [CrossRef]
- Beelen, D.W.; Elmaagacli, A.; Müller, K.-D.; Hirche, H.; Schaefer, U.W. Influence of Intestinal Bacterial Decontamination Using Metronidazole and Ciprofloxacin or Ciprofloxacin Alone on the Development of Acute Graft-Versus-Host Disease After Marrow Transplantation in Patients with Hematologic Malignancies: Final Results and Long-Term Follow-Up of an Open-Label Prospective Randomized Trial. Blood 1999, 93, 3267–3275. [Google Scholar] [CrossRef]
- Miao, L.; Wang, Y.Y.; Zhu, L.; Wu, Z.J.; Zhou, R.M. Endophytic fungi from four plant species: Their isolation and antitumor activity. Microbiol. 2009, 36, 865–869. [Google Scholar]
- He, Q.; Zeng, Q.; Shao, Y.; Zhou, H.; Li, T.; Song, F.; Liu, W. Anti-cervical cancer activity of secondary metabolites of endophytic fungi from Ginkgo biloba. Cancer Biomark 2020, 28, 371–379. [Google Scholar] [CrossRef]
- Geller, L.T.; Barzily-Rokni, M.; Danino, T.; Jonas, O.H.; Shental, N.; Nejman, D.; Gavert, N.; Zwang, Y.; Cooper, Z.A.; Shee, K.; et al. Potential role of intratumor bacteria in mediating tumor resistance to the chemotherapeutic drug gemcitabine. Science 2017, 357, 1156–1160. [Google Scholar] [CrossRef]
- Ikeda, Y.; Adachi, Y.; Ishii, T.; Tamura, H.; Aketagawa, J.; Tanaka, S.; Ohno, N. Blocking effect of anti-Dectin-1 antibodies on the anti-tumor activity of 1,3-beta-glucan and the binding of Dectin-1 to 1,3-beta-glucan. Biol. Pharm. Bull. 2007, 30, 1384–1389. [Google Scholar] [CrossRef] [PubMed]
- Zhong, K.; Tong, L.; Liu, L.; Zhou, X.; Liu, X.; Zhang, Q.; Zhou, S. Immunoregulatory and antitumor activity of schizophyllan under ultrasonic treatment. Int. J. Biol. Macromol. 2015, 80, 302–308. [Google Scholar] [CrossRef] [PubMed]
- Lopez-Legarda, X.; Rostro-Alanis, M.; Parra-Saldivar, R.; Villa-Pulgarin, J.A.; Segura-Sanchez, F. Submerged cultivation, characterization and in vitro antitumor activity of polysaccharides from Schizophyllum radiatum. Int. J. Biol. Macromol. 2021, 186, 919–932. [Google Scholar] [CrossRef] [PubMed]
- van der Velden, W.J.; Plantinga, T.S.; Feuth, T.; Donnelly, J.P.; Netea, M.G.; Blijlevens, N.M. The incidence of acute graft-versus-host disease increases with Candida colonization depending the dectin-1 gene status. Clin. Immunol. 2010, 136, 302–306. [Google Scholar] [CrossRef]
- Yu, T.; Guo, F.; Yu, Y.; Sun, T.; Ma, D.; Han, J.; Qian, Y.; Kryczek, I.; Sun, D.; Nagarsheth, N.; et al. Fusobacterium nucleatum Promotes Chemoresistance to Colorectal Cancer by Modulating Autophagy. Cell 2017, 170, 548–563.e516. [Google Scholar] [CrossRef]
- Heshiki, Y.; Vazquez-Uribe, R.; Li, J.; Ni, Y.; Quainoo, S.; Imamovic, L.; Li, J.; Sørensen, M.; Chow, B.K.C.; Weiss, G.J.; et al. Predictable modulation of cancer treatment outcomes by the gut microbiota. Microbiome 2020, 8, 28. [Google Scholar] [CrossRef]
- Shi, W.; Shen, L.; Zou, W.; Wang, J.; Yang, J.; Wang, Y.; Liu, B.; Xie, L.; Zhu, J.; Zhang, Z. The Gut Microbiome Is Associated with Therapeutic Responses and Toxicities of Neoadjuvant Chemoradiotherapy in Rectal Cancer Patients—A Pilot Study. Front. Cell. Infect. Microbiol. 2020, 10, 562463. [Google Scholar] [CrossRef]
- Panebianco, C.; Adamberg, K.; Jaagura, M.; Copetti, M.; Fontana, A.; Adamberg, S.; Kolk, K.; Vilu, R.; Andriulli, A.; Pazienza, V. Influence of gemcitabine chemotherapy on the microbiota of pancreatic cancer xenografted mice. Cancer Chemother. Pharmacol. 2018, 81, 773–782. [Google Scholar] [CrossRef]
- Matson, V.; Fessler, J.; Bao, R.; Chongsuwat, T.; Zha, Y.; Alegre, M.-L.; Luke, J.J.; Gajewski, T.F. The commensal microbiome is associated with anti–PD-1 efficacy in metastatic melanoma patients. Science 2018, 359, 104–108. [Google Scholar] [CrossRef]
- Peters, B.A.; Wilson, M.; Moran, U.; Pavlick, A.; Izsak, A.; Wechter, T.; Weber, J.S.; Osman, I.; Ahn, J. Relating the gut metagenome and metatranscriptome to immunotherapy responses in melanoma patients. Genome Med. 2019, 11, 61. [Google Scholar] [CrossRef]
- Botticelli, A.; Putignani, L.; Zizzari, I.; Chierico, F.D.; Reddel, S.; Pietro, F.D.; Quagliarello, A.; Onesti, C.E.; Raffaele, G.; Mazzuca, F.; et al. Changes of microbiome profile during nivolumab treatment in NSCLC patients. J. Clin. Oncol. 2018, 36, e15020. [Google Scholar] [CrossRef]
- Fukuoka, S.; Daisuke, M.; Togashi, Y.; Sugiyama, E.; Udagawa, H.; Kirita, K.; Kamada, T.; Kawazoe, A.; Goto, K.; Doi, T.; et al. Association of gut microbiome with immune status and clinical response in solid tumor patients who received on anti-PD-1 therapies. J. Clin. Oncol. 2018, 36, 3011. [Google Scholar] [CrossRef]
- Gopalakrishnan, V.; Spencer, C.N.; Nezi, L.; Reuben, A.; Andrews, M.C.; Karpinets, T.V.; Prieto, P.A.; Vicente, D.; Hoffman, K.; Wei, S.C.; et al. Gut microbiome modulates response to anti–PD-1 immunotherapy in melanoma patients. Science 2018, 359, 97–103. [Google Scholar] [CrossRef] [PubMed]
- Baruch Erez, N.; Youngster, I.; Ben-Betzalel, G.; Ortenberg, R.; Lahat, A.; Katz, L.; Adler, K.; Dick-Necula, D.; Raskin, S.; Bloch, N.; et al. Fecal microbiota transplant promotes response in immunotherapy-refractory melanoma patients. Science 2021, 371, 602–609. [Google Scholar] [CrossRef]
- Zhang, M.; Kim, J.A.; Huang, A.Y. Optimizing Tumor Microenvironment for Cancer Immunotherapy: Beta-Glucan-Based Nanoparticles. Front. Immunol. 2018, 9, 341. [Google Scholar] [CrossRef]
- Park, D.R.; Thomsen, A.R.; Frevert, C.W.; Pham, U.; Skerrett, S.J.; Kiener, P.A.; Liles, W.C. Fas (CD95) Induces Proinflammatory Cytokine Responses by Human Monocytes and Monocyte-Derived Macrophages. J. Immunol. 2003, 170, 6209. [Google Scholar] [CrossRef]
- Koido, S.; Ohkusa, T.; Homma, S.; Namiki, Y.; Takakura, K.; Saito, K.; Ito, Z.; Kobayashi, H.; Kajihara, M.; Uchiyama, K.; et al. Immunotherapy for colorectal cancer. World J. Gastroenterol. 2013, 19, 8531–8542. [Google Scholar] [CrossRef]
- Edwards-Ingram, L.; Gitsham, P.; Burton, N.; Warhurst, G.; Clarke, I.; Hoyle, D.; Oliver, S.G.; Stateva, L. Genotypic and physiological characterization of Saccharomyces boulardii, the probiotic strain of Saccharomyces cerevisiae. Appl. Environ. Microbiol. 2007, 73, 2458–2467. [Google Scholar] [CrossRef]
- Chen, X.; Yang, G.; Song, J.H.; Xu, H.; Li, D.; Goldsmith, J.; Zeng, H.; Parsons-Wingerter, P.A.; Reinecker, H.C.; Kelly, C.P. Probiotic yeast inhibits VEGFR signaling and angiogenesis in intestinal inflammation. PLoS ONE 2013, 8, e64227. [Google Scholar] [CrossRef]
- Kazmierczak-Siedlecka, K.; Ruszkowski, J.; Fic, M.; Folwarski, M.; Makarewicz, W. Saccharomyces boulardii CNCM I-745: A Non-bacterial Microorganism Used as Probiotic Agent in Supporting Treatment of Selected Diseases. Curr. Microbiol. 2020, 77, 1987–1996. [Google Scholar] [CrossRef]
- Nakayama, H.; Kinouchi, T.; Kataoka, K.; Akimoto, S.; Matsuda, Y.; Ohnishi, Y. Intestinal anaerobic bacteria hydrolyse sorivudine, producing the high blood concentration of 5-(E)-(2-bromovinyl)uracil that increases the level and toxicity of 5-fluorouracil. Pharmacogenetics 1997, 7, 35–43. [Google Scholar] [CrossRef] [PubMed]
- Chu, D.T.; Nguyen, T.T.; Tien, N.L.B.; Tran, D.K.; Jeong, J.H.; Anh, P.G.; Thanh, V.V.; Truong, D.T.; Dinh, T.C. Recent Progress of Stem Cell Therapy in Cancer Treatment: Molecular Mechanisms and Potential Applications. Cells 2020, 9, 563. [Google Scholar] [CrossRef] [PubMed]
- Side Effects of a Bone Marrow Transplant (Stem Cell Transplant). Available online: https://www.cancer.net/navigating-cancer-care/how-cancer-treated/bone-marrowstem-cell-transplantation/side-effects-bone-marrow-transplant-stem-cell-transplant (accessed on 20 April 2022).
- Cooke, K.R.; Luznik, L.; Sarantopoulos, S.; Hakim, F.T.; Jagasia, M.; Fowler, D.H.; van den Brink, M.R.M.; Hansen, J.A.; Parkman, R.; Miklos, D.B.; et al. The Biology of Chronic Graft-versus-Host Disease: A Task Force Report from the National Institutes of Health Consensus Development Project on Criteria for Clinical Trials in Chronic Graft-versus-Host Disease. Biol. Blood Marrow Transpl. 2017, 23, 211–234. [Google Scholar] [CrossRef] [PubMed]
- Legoff, J.; Resche-Rigon, M.; Bouquet, J.; Robin, M.; Naccache, S.N.; Mercier-Delarue, S.; Federman, S.; Samayoa, E.; Rousseau, C.; Piron, P.; et al. The eukaryotic gut virome in hematopoietic stem cell transplantation: New clues in enteric graft-versus-host disease. Nat. Med. 2017, 23, 1080–1085. [Google Scholar] [CrossRef]
- Cooke, K.R.; Olkiewicz, K.; Erickson, N.; Ferrara, J.L.M. The role of endotoxin and the innate immune response in the pathophysiology of acute graft versus host disease. J. Endotoxin Res. 2002, 8, 441–448. [Google Scholar] [CrossRef]
- Rosshart, S.P.; Vassallo, B.G.; Angeletti, D.; Hutchinson, D.S.; Morgan, A.P.; Takeda, K.; Hickman, H.D.; McCulloch, J.A.; Badger, J.H.; Ajami, N.J.; et al. Wild Mouse Gut Microbiota Promotes Host Fitness and Improves Disease Resistance. Cell 2017, 171, 1015–1028.e1013. [Google Scholar] [CrossRef]
- Zackular, J.P.; Baxter, N.T.; Iverson, K.D.; Sadler, W.D.; Petrosino, J.F.; Chen, G.Y.; Schloss, P.D. The gut microbiome modulates colon tumorigenesis. mBio 2013, 4, e00692-13. [Google Scholar] [CrossRef]
- Zuo, T.; Wong, S.H.; Cheung, C.P.; Lam, K.; Lui, R.; Cheung, K.; Zhang, F.; Tang, W.; Ching, J.Y.L.; Wu, J.C.Y.; et al. Gut fungal dysbiosis correlates with reduced efficacy of fecal microbiota transplantation in Clostridium difficile infection. Nat. Commun. 2018, 9, 3663. [Google Scholar] [CrossRef]
- Fong, W.; Li, Q.; Yu, J. Gut microbiota modulation: A novel strategy for prevention and treatment of colorectal cancer. Oncogene 2020, 39, 4925–4943. [Google Scholar] [CrossRef]
- Pouncey, A.L.; Scott, A.J.; Alexander, J.L.; Marchesi, J.; Kinross, J. Gut microbiota, chemotherapy and the host: The influence of the gut microbiota on cancer treatment. Ecancermedicalscience 2018, 12, 868. [Google Scholar] [CrossRef]
- Ghannoum, M.A.; Long, L.; Bunick, C.G.; Del Rosso, J.Q.; Gamal, A.; Tyring, S.K.; McCormick, T.S.; Grada, A. Sarecycline Demonstrated Reduced Activity Compared to Minocycline against Microbial Species Representing Human Gastrointestinal Microbiota. Antibiotics 2022, 11, 324. [Google Scholar] [CrossRef] [PubMed]
- Routy, B.; Chatelier, E.L.; Derosa, L.; Duong, C.P.M.; Alou, M.T.; Daillère, R.; Fluckiger, A.; Messaoudene, M.; Rauber, C.; Roberti, M.P.; et al. Gut microbiome influences efficacy of PD-1–based immunotherapy against epithelial tumors. Science 2018, 359, 91–97. [Google Scholar] [CrossRef] [PubMed]
- Halley, A.; Leonetti, A.; Gregori, A.; Tiseo, M.; Deng, D.M.; Giovannetti, E.; Peters, G.J. The Role of the Microbiome in Cancer and Therapy Efficacy: Focus on Lung Cancer. Anticancer Res. 2020, 40, 4807–4818. [Google Scholar] [CrossRef] [PubMed]
| Type of Cancer | Evidence of Mycobiome Implication | Author/Year |
|---|---|---|
| Colorectal cancer | Increased Ascomycota, and Basidiomycota. | Luan et al., 2015 [21] Gao et al., 2017 [22] Richard et al., 2018 [23] |
| Decreased fungal diversity in polyps compared to adjacent tissue. | Luan et al., 2015 [21] Gao et al., 2017 [22] Coker et al., 2019 [8] | |
| Increased ratio of Ascomycota to Basidiomycota in CRC patients. | Coker et al., 2019 [8] Gao et al., 2017 [22] | |
| Increased opportunistic fungi; Trichosporon and Malassezia, which could be implicated in the progression to CRC. | Gao et al., 2017 [22] | |
| Distinctive sets of proteins secreted by Schizosaccharomyces pombe in patients’ stool samples. Increased Saccharomycetales in advanced adenoma versus non-advanced samples. | Chin et al., 2018 [24] | |
| CARD9 deficient mice exhibit fungal dysbiosis resulted in increased CRC tumor loads. | Luan et al., 2015 [21] | |
| Treatment with fluconazole suppressed tumor growth in mice. | Wang et al., 2018 [25] | |
| Pancreatic ductal adenocarcinoma (PDA) | In comparison to the normal pancreas, PDA tumors have a 3000-fold increase in fungi. In mice and humans, the fungal community infiltrating PDA was enriched in Malassezia. In both slowly progressive and invasive PDA models, fungal elimination with amphotericin B was tumor-protective, while re-population with Malassezia–but not Candida, Saccharomyces, or Aspergillus–promoted oncogenesis. Connection of MBL, that attaches fungal wall glycans to activate the complement pathway, was needed in the promotion of malignancy. Tumor growth was inhibited by MBL or C3 deletion in the extra-tumoral region or C3aR knockdown in tumor cells. Pathogenic fungi may promote PDA by inducing MBL, which activates the complement system. | Aykut et al., 2019 [26] |
| Gastric Cancer | Unique fungal profile was observed in gastric cancer biopsies. C. albicans, Arcopilus aureus, and Fusicolla spp. were enriched in GC compared to the control, whereas C. glabrata, Aspergillus montevidensis, Saitozyma and Penicillium were depleted. | Zhong et al., 2021 [27] |
| Head and neck SCC/Oral SCC | C. albicans, C. etchellsii, Hannaella, and Gibberella were prevalent in OSCC specimens, while Altenaria and Trametes were observed in larger abundance in polyps’ specimens | Perera et al., 2017 [28] Vesty et al., 2018 [29] Shay et al., 2020 [30] Makinen et al., 2018 [31] |
| Polyp specimens dominated by Malassezia restricta and Aspergillus tamarii. | Vesty et al., 2018 [29] | |
| Marked expression of IL-1, IL-6, and IL-8 by oral cancer cells which are usually associated with C. albicans. | Dongari-Bagtzoglou, and Kashleva, 2003 [32] Vesty et al., 2018 [29] Arzmi et al., 2019 [33] | |
| Compared with healthy controls, Schizophyllum commune was significantly lower in HNSCC patients. | Shay et al., 2020 [30] |
| Type of Cancer | Finding | Reference |
|---|---|---|
| I. Cancers of the Gastrointestinal tract | ||
| Colorectal cancer | Increase in the Ascomycota: Basidiomycota ratio and enrichment of Microascaceae and Sordariaceae_spp. in CRC patients compared to healthy subjects. Increase in Microbotryomycetes, Sordariomycetes, Microascaceae, Sordariales, Lasiosphaeriaceae, and Microascales, with a decrease in the abundance of Pleosporaceae and Alternaria was detected in late-stage CRC patients. | Gao et al., 2017 [22] |
| Basidiomycota was significantly enriched in adjacent biopsy samples. The phylum Chytridiomycota was significantly enriched in adenomas compared with adjacent biopsy samples. The order Saccharomycetales and phylum Basidiomycota organisms were significantly enriched in advanced versus non-advanced adenoma tissue biopsies. | Luan et al., 2015 [21] | |
| The abundance of Candida, Saccharomyces, and Ascomycete was elevated in patients with colonic polyps. | Chen et al., 2019 [94] | |
| Enrichment of Rhodotorula, Malassezia, and Acremonium with reduction of Saccharomycetes, especially S. cerevisiae, in fecal samples from CRC patients. | Coker et al., 2019 [8] | |
| Proteins secreted by Schizosaccharomyces pombe in the stool samples of CRC patients. | Chin et al., 2018 [24] | |
| Pancreatic ductal adenocarcinoma (PDA) | PDA patients were found to have a 3000-fold increase in fungal abundance compared to a cohort of subjects with a healthy pancreas. Anti-fungal medication in a mouse model of PDA protected against tumor progression, whereas repopulation with Malassezia resulted in rapid growth of pancreatic tumors. | Aykut et al., 2019 [26] |
| Head and neck SCC/Oral SCC | C. albicans was the most commonly isolated species among OSCC in most patients. | Mäkinen et al., 2018 [31] |
| Malassezia was reduced in abundance in patients with HNSCC. | Shay et al., 2020 [30] | |
| Rhodotorula, Geotrichum, and Pneumocystis were only detected in samples obtained from the cancer patient samples. Fonsecaea was found in both cancer samples and cancer-adjacent tissue from the same patients but not in oral tissue from healthy subjects. | Banerjee, Sagarika et al., 2017a [95] | |
| Glomeromycota was the only fungal phylum that was significantly decreased in the tumor group compared to their matched non-tumor tissues. High T-stage tumor samples exhibited an increased richness of both bacteria and fungi at the phylum level compared to low T-stage samples (p ≤ 0.047). | Mukherjee et al., 2017 [96] | |
| Overgrowth of C. albicans in oral squamous cell malignant tissue. | Perera et al., 2017 [28] | |
| Increased concentration of C. albicans in the saliva of HNSCC patients. | Vesty et al., 2018 [29] | |
| II. Cancers beyond the Gastrointestinal Tract | ||
| Ovarian cancer | Eighteen fungal signatures were detected only in ovarian cancer samples including: Cladosporium, with the highest hybridization signal, Acremonium, and Candida. | Banerjee, S. et al., 2017b [97] |
| Breast cancer | The highest fungal diversity was detected in ER+ samples, while HR+ samples had the least fungal diversity. Arthroderma accounted for the highest average hybridization signal. Candida, Cryptococcus, Mucor, Penicillium, Trichophyton, and Trichosporon were observed in all cancer samples. | Banerjee, S. et al., 2021 [98] |
| Melanoma | α-diversity of the fungal community revealed significantly higher richness in melanoma patients compared to controls. Reduction in the fungal diversity as the disease progressed. Saccharomycetales was enriched in melanoma patients. One hundred and eighty zOTUs (including 63 fungal zOTUs) were enriched in melanoma patients compared to controls. Out of these zOTUs, 162 were mainly enriched in patients with in situ melanoma. Analysis of the gut microbiota revealed a significant difference between metastatic and nonmetastatic melanoma patients. | Vitali et al., 2022 [99] |
| Type of Study | Investigated Agent/Modality | Finding | Reference |
|---|---|---|---|
| In vitro and In vivo—Murine tumor model | β-glucan | Oral administration can boost phagocytes’ tumoricidal activity against iC3b-opsonized cancer cells. | (Hong et al., 2004) [107] |
| In vitro and In vivo—Murine tumor model | Oral administration decreased the tumor burden by inducing the transition of immunosuppressive M2 macrophages into inflammatory M1 macrophages via dectin-1 receptor. | (Liu et al., 2015; Wang et al., 2015) [68,108] | |
| Clinical Trial | Oral administration of two 10-mg capsules of soluble β-glucan derived from S. cerevisiae enhanced the proliferation and activation of peripheral blood monocytes with no clinical adverse effects (Mean monocyte count increased from 326 ± 124/mm3 to 496 ± 194/mm3 on day 15 post-administration p = 0.015). | (Demir et al., 2007) [109] | |
| In vitro and In vivo—Murine tumor model | Intravenous administration with β-glucan derived from mutated S. cerevisiae strain suppressed cancer cell proliferation in a dose-dependent manner. Pretreatment of mice with the same type of β-glucan, 2 days before inoculation with the tumor cells, enhanced mice survival time. Enhanced pro-inflammatory cytokine production and anticancer activity of peritoneal macrophages, and increased natural killer (NK) cell cytotoxicity. | (Yoon et al., 2008). [110] | |
| In vivo—Murine tumor model | Prophylactic intravenous administration of β-glucan, derived from mutated S. cerevisiae, in combination with cisplatin, had better efficacy compared to the chemotherapy agent alone. | Kim et al., 2010 [111] | |
| In vitro and In vivo—Murine tumor model | Oral administration of β-glucan extracted from S. cerevisiae suppressed the development of melanoma in a dose-dependent manner. | Vetvicka and Vetvickova, 2015 [112] | |
| In vitro and In vivo—Murine tumor model | Systemic administration (intraperitoneal or intranasal) of oat-derived β-(1-3)—(1-4)-glucan resulted in activation of M1-type macrophage, production of pro-inflammatory cytokines (such as IFN-γ, TNF-α, CXCL9, and CXCL10), IRF1, and PD-L1 expression, resulting in enhanced anticancer immune response compared to untreated controls. | Zhang et al., 2018 [113] | |
| In vitro | Monocyte treatment with Imprime (soluble type of β-glucan) resulted in enhanced production of M2 macrophages and dendritic cells with higher expression of PD-L1 and CD86, both of which can potentiate the activity of anti PD-1 antibodies. For Imprime to act efficiently, prior presence of anti-β-glucan antibodies is required. | Chan et al., 2016 [114] | |
| In vivo—Murine tumor model | In a synergistic tumor model, Imprime and anti-PD-1 antibodies combination group had a lower median tumor volume compared to the anti-PD-1 antibodies alone- treated group (172 mm3 vs. 936 mm3, respectively). | Qiu et al., 2016 [115] | |
| In vivo—Murine tumor model | Probiotics | Saccharomyces boulardii inhibited the EGFR-MEK-ERK signaling network and pro-apoptotic actions in tumor cells by suppressing Akt, thereby regulating the inflammatory responses and suppressing the gut cancer expansion. S. boulardii blocks vascular endothelial growth factor receptor (VEGFR) signaling and inhibits angiogenesis both in vitro and in vivo | Chen, X. et al., 2009 [116] |
| In vivo—Murine tumor model | Administration of Lactobacillus paracasei and Lactobacillus reuteri in combination with the anticancer agent gemcitabine in murine model was associated with a more reduction in the growth rate of pancreatic intraepithelial neoplasia compared to gemcitabine only treated group and untreated controls. | Chen, S.M. et al., 2020 [117]. | |
| Human subject and In vivo—Murine tumor model | Bacteriome and mycobiome modulation. | Responsiveness to radiation therapy was enhanced following antibiotic-mediated depletion or gnotobiotic exclusion of fungi. Depletion of bacteria reduced responsiveness. A negative association was noted between increased intra-tumoral expression of Dectin-1 and survival in patients with breast cancer, and was also required for the effects of commensal fungi in mouse models of radiation therapy | Shiao et al., 2021 [103] |
| Clinical Trial | FMT improved the resistance to anti-PD-1 therapy in melanoma patients. | Davar et al., 2021 [118] | |
| Human subject and In vivo—Murine tumor model | Mycobiome modulation. | Mycobiome ablation:
| Aykut et al., 2019 [26] |
| Clinical trial | Bacteriome Modulation | Antimicrobial chemotherapy targeted toward intestinal anaerobic bacteria in SCT patients showed a significant reduction in the severity of acute GVHD following bone marrow transplantation. | Beelen et al., 1999 [119] |
| In vitro | Fungal metabolites | Crude extracts of several endophytic fungal strains present in Ginkgo biloba exhibited anticancer activity at a test concentration of 200 μg/mL. | L. Miao, 2009 [120] |
| In vitro and In vivo—Murine tumor model | Three strains of endophytic fungi from the leaves of Ginkgo biloba were found to produce podophyllotoxin. Their metabolites demonstrated:
| He et al., 2020 [121] | |
| Human subject and In vivo—Murine tumor model | Overexpression of the enzyme cytidine deaminase is associated with reducing gemcitabine into its inactive metabolite. | Geller et al., 2017 [122] | |
| In vitro and In vivo—Murine tumor model | Some Schizophyllum species metabolites possess anticancer activity. Administration of dectin-1 antibodies resulted in reduced SPG activity. Schizophyllan anti-cancer effect is dependent on its interaction with Dectin-1. | Ikeda et al., 2007; Zhong et al., 2015; Lopez-Legarda et al., 2021 [123,124,125] | |
| Human subjects | Others | Colonization with Candida is accompanied by a significant increase in the incidence of acute GVHD compared to non-colonized patients. | Van Der Velden et al., 2010 [126] |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Gamal, A.; Elshaer, M.; Alabdely, M.; Kadry, A.; McCormick, T.S.; Ghannoum, M. The Mycobiome: Cancer Pathogenesis, Diagnosis, and Therapy. Cancers 2022, 14, 2875. https://doi.org/10.3390/cancers14122875
Gamal A, Elshaer M, Alabdely M, Kadry A, McCormick TS, Ghannoum M. The Mycobiome: Cancer Pathogenesis, Diagnosis, and Therapy. Cancers. 2022; 14(12):2875. https://doi.org/10.3390/cancers14122875
Chicago/Turabian StyleGamal, Ahmed, Mohammed Elshaer, Mayyadah Alabdely, Ahmed Kadry, Thomas S. McCormick, and Mahmoud Ghannoum. 2022. "The Mycobiome: Cancer Pathogenesis, Diagnosis, and Therapy" Cancers 14, no. 12: 2875. https://doi.org/10.3390/cancers14122875
APA StyleGamal, A., Elshaer, M., Alabdely, M., Kadry, A., McCormick, T. S., & Ghannoum, M. (2022). The Mycobiome: Cancer Pathogenesis, Diagnosis, and Therapy. Cancers, 14(12), 2875. https://doi.org/10.3390/cancers14122875







