MACC1-Induced Collective Migration Is Promoted by Proliferation Rather Than Single Cell Biomechanics
Simple Summary
Abstract
1. Introduction
2. Materials and Methods
2.1. Cell Culture
2.2. Quantitative Real-Time Reverse Transcription PCR (qRT-PCR)
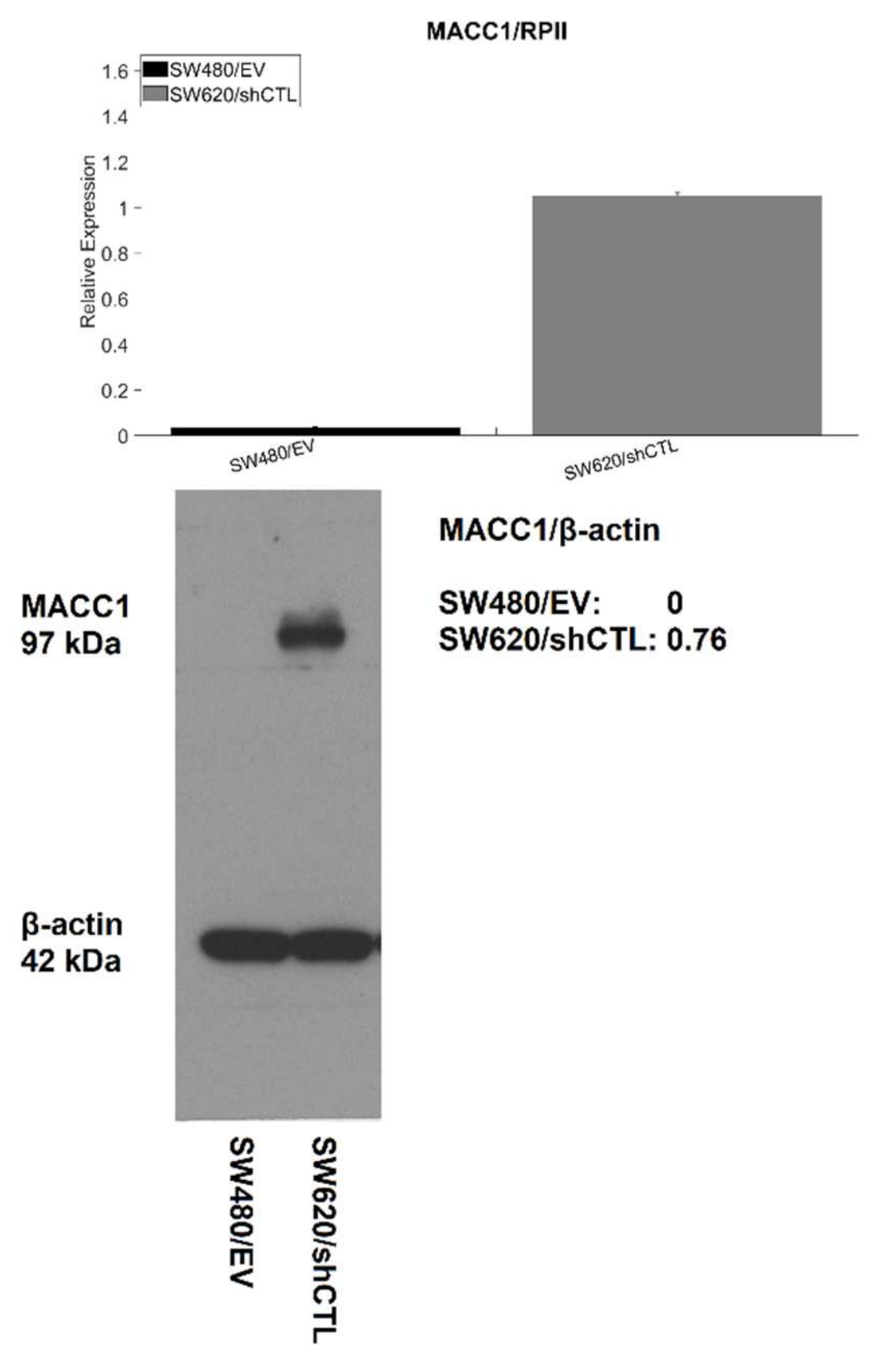
2.3. Western Blot Analysis
2.4. Atomic Force Microscopy
2.5. Single Cell Motility and Doubling Time
2.6. Measurement of the Properties of Collective Cell Migration
2.7. Measuring Colony Expansion
2.8. Simulation of a Persistent Random Walker with Directional Constraints
2.9. Statistics
3. Results
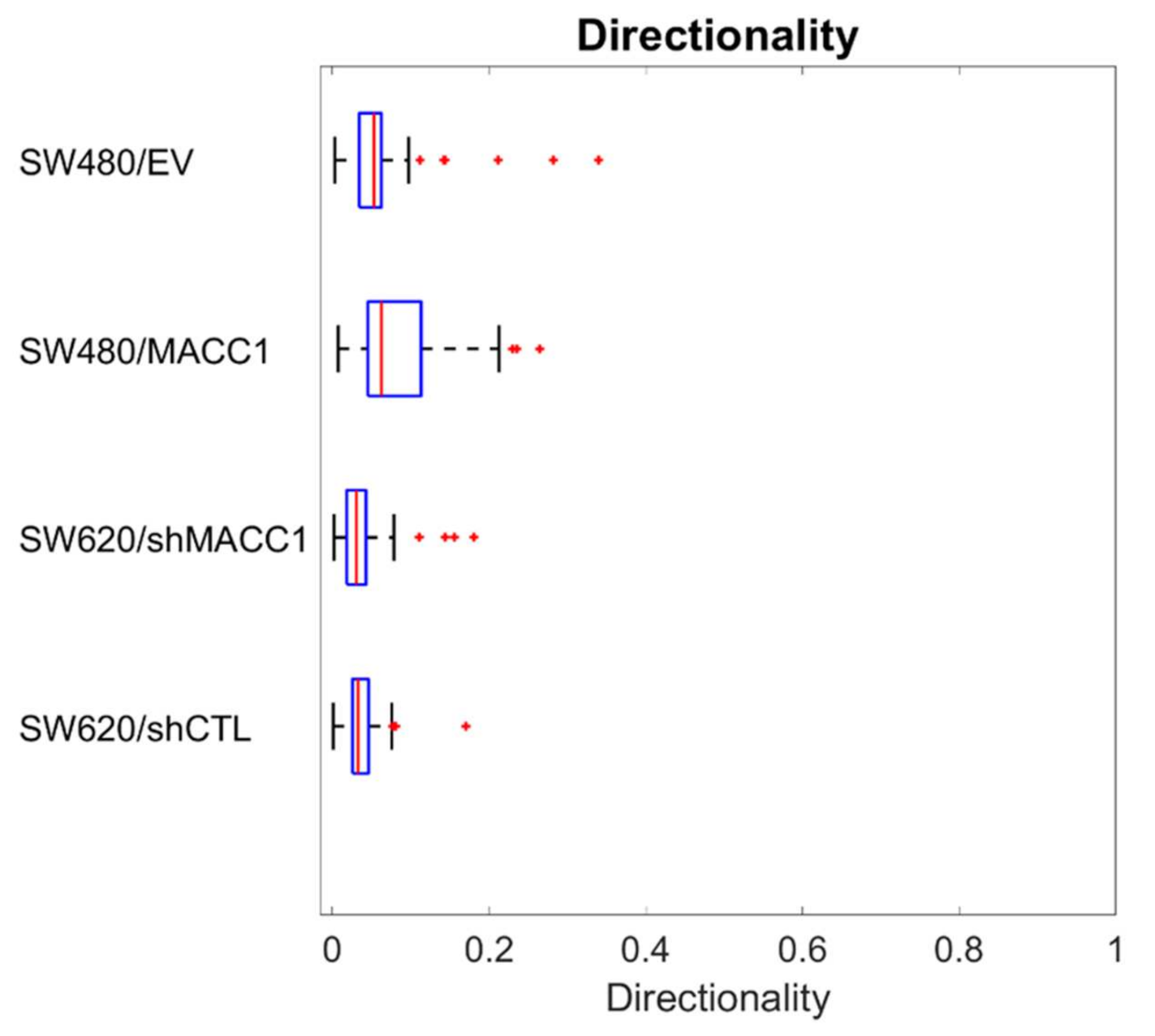
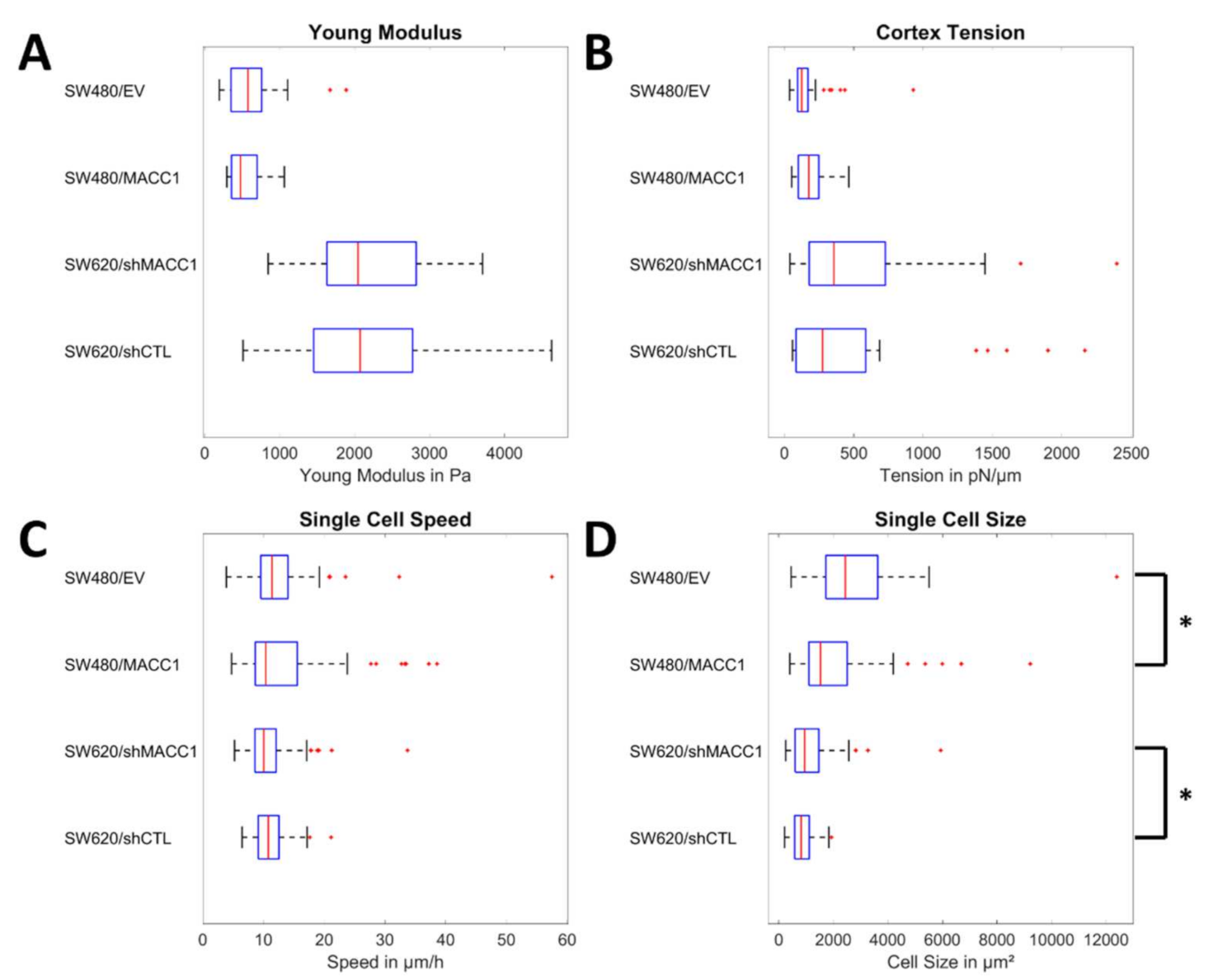
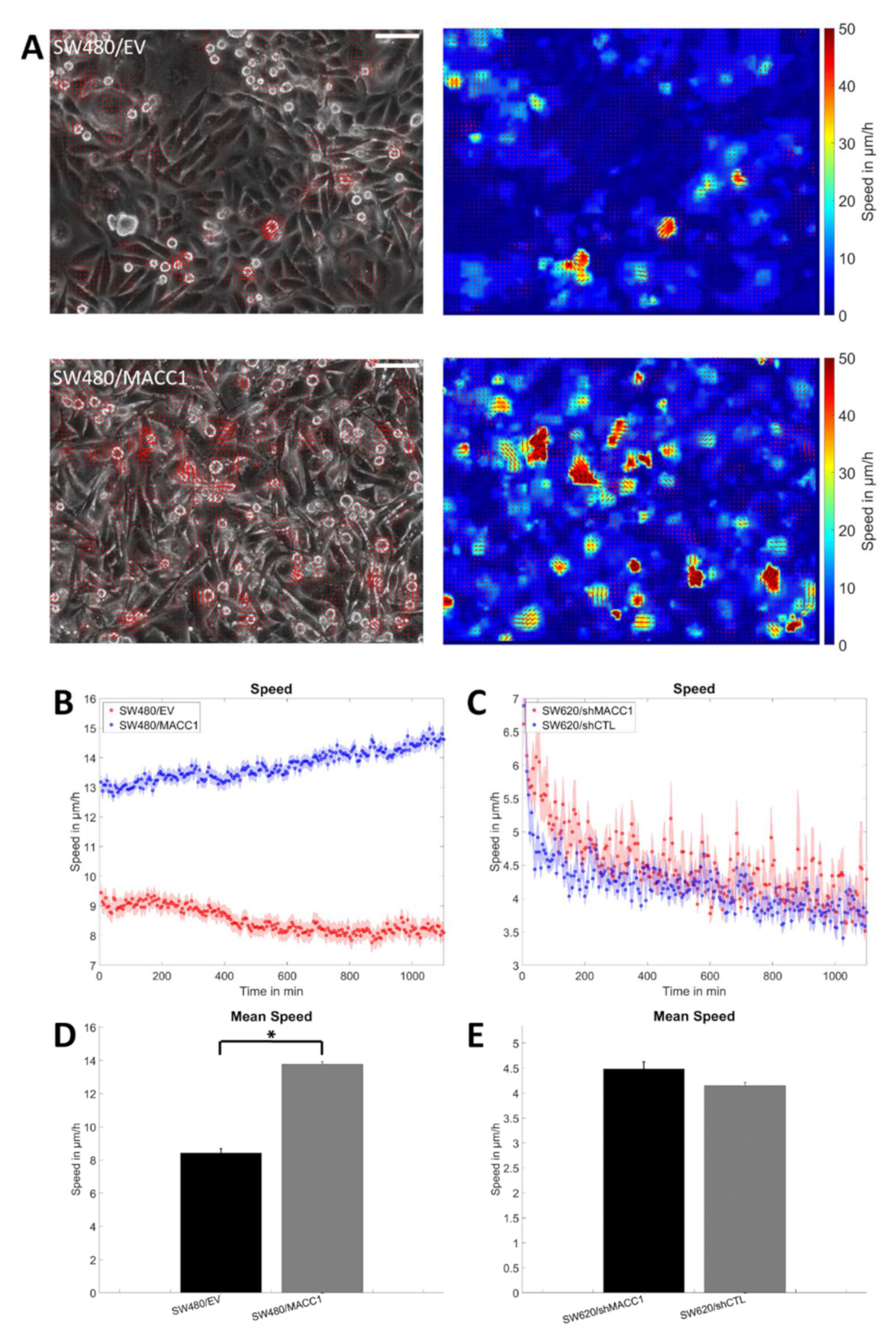
3.1. Effect of MACC1 Expression on Single Cell Properties
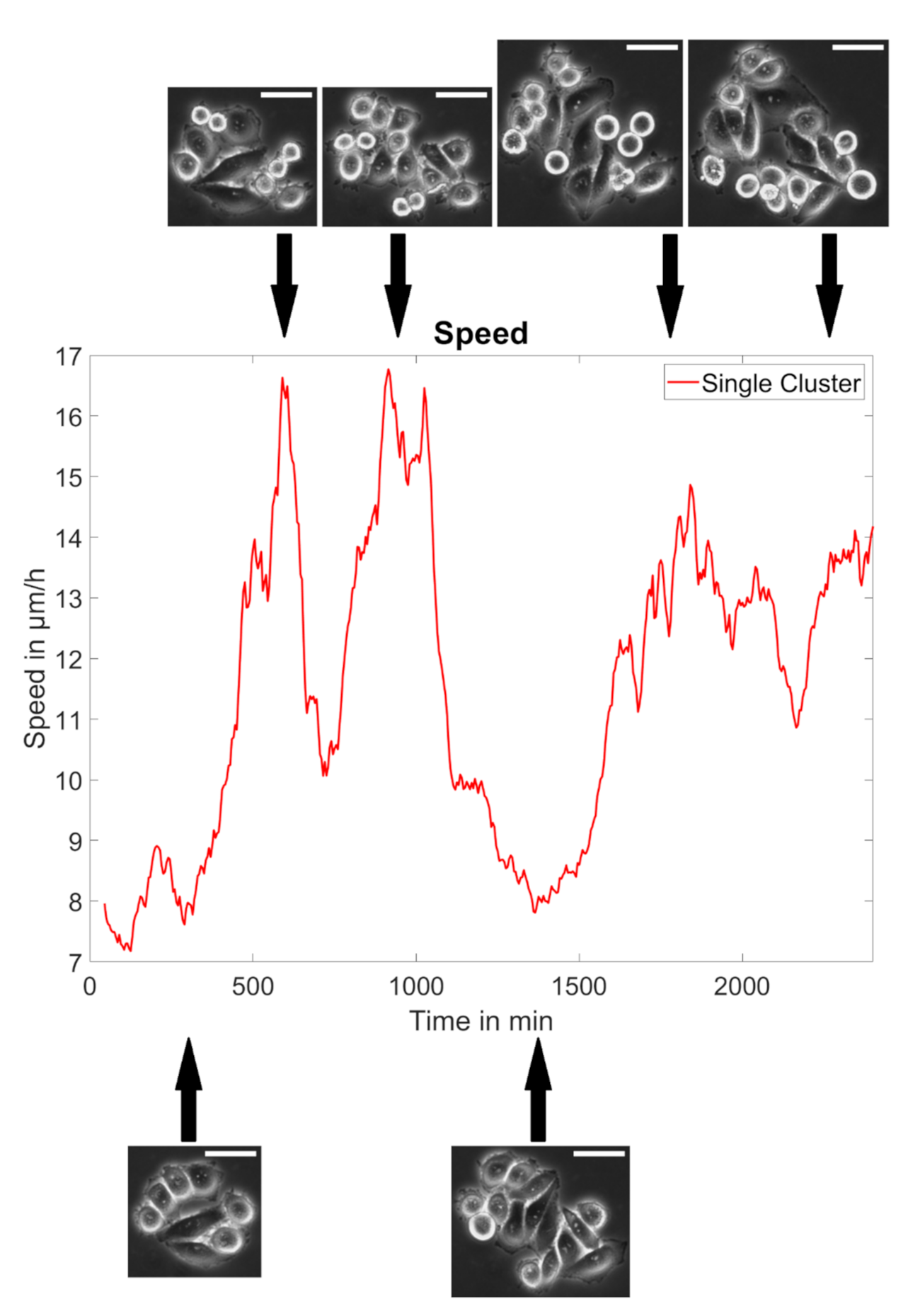
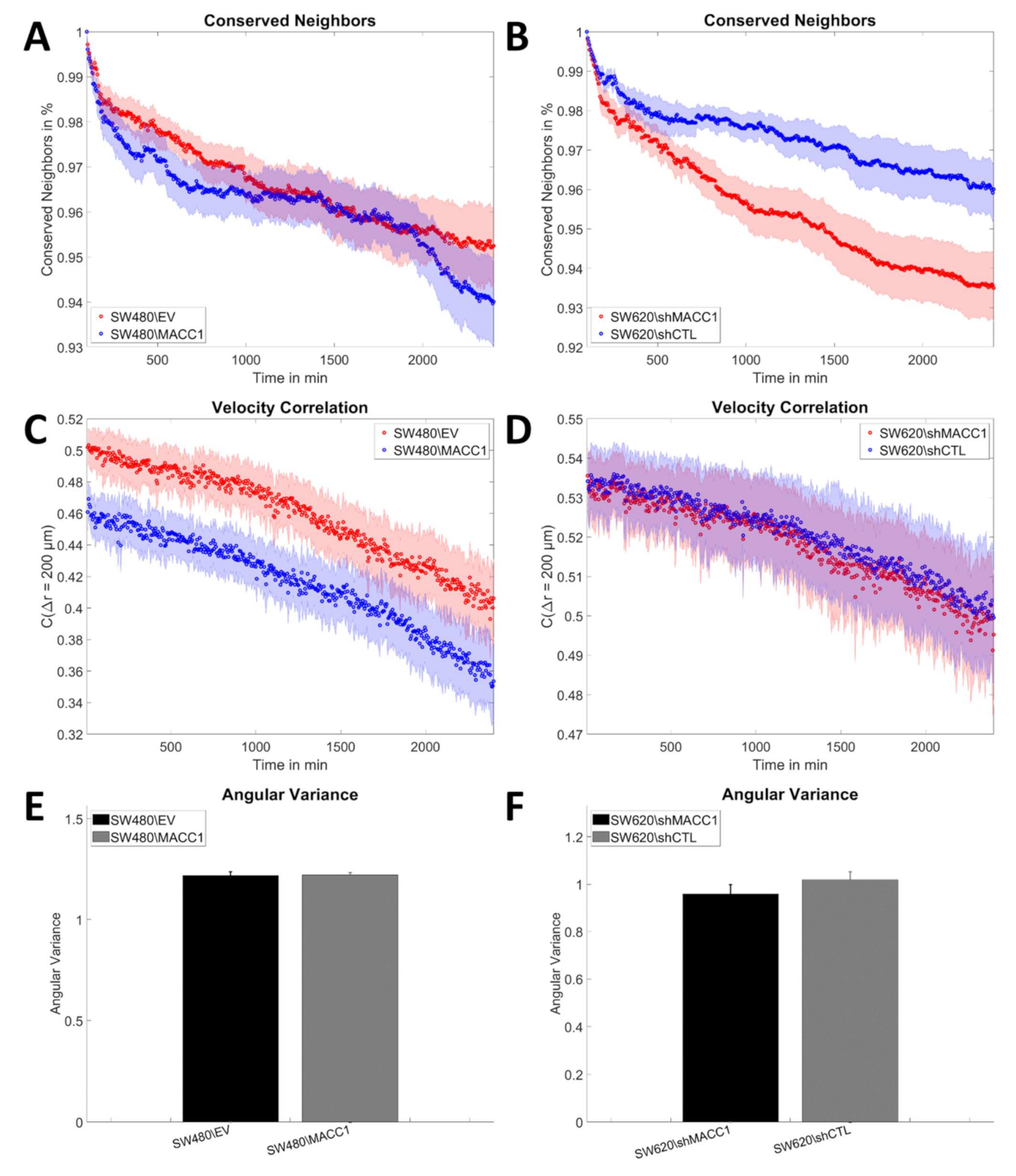
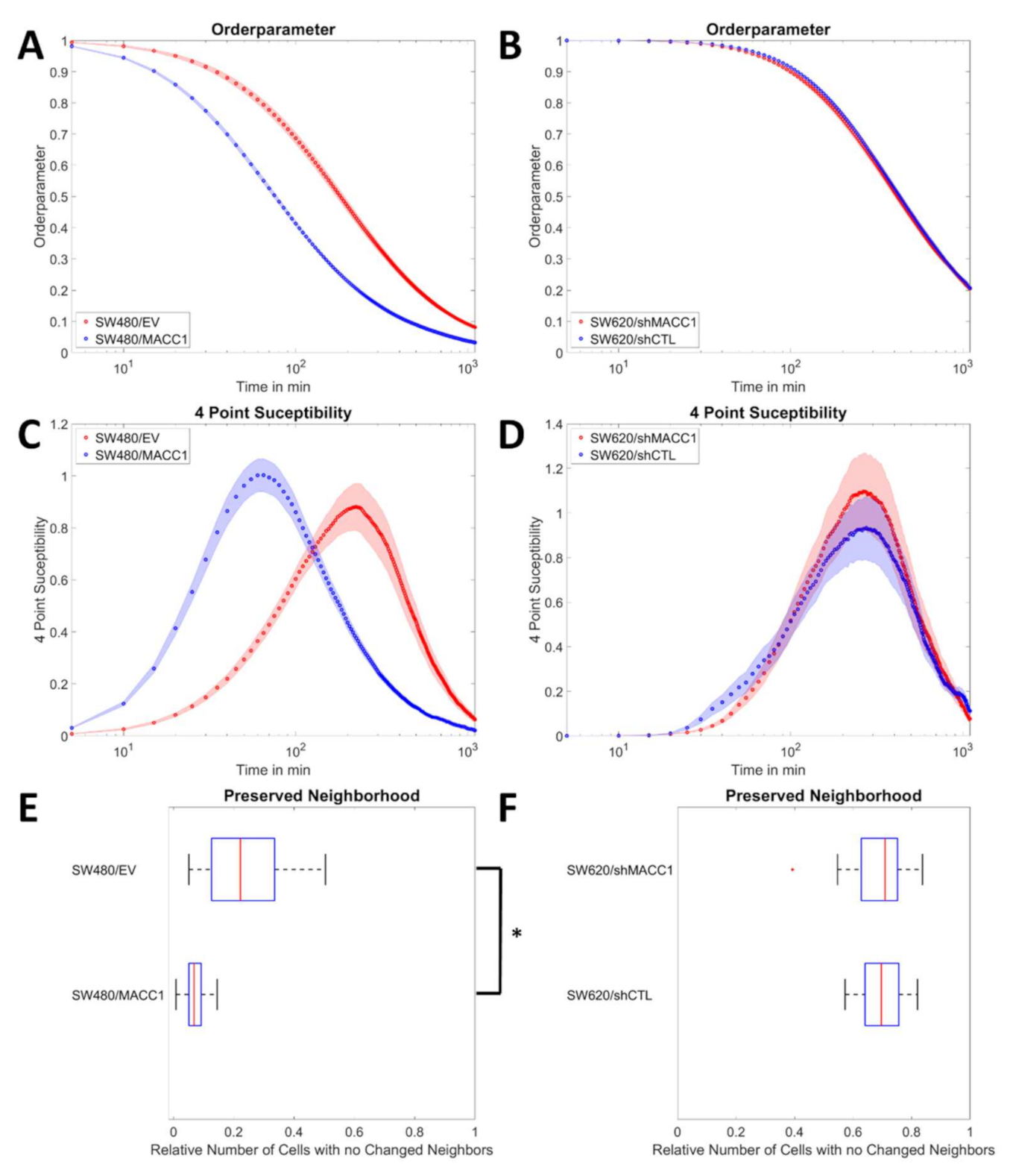
3.2. MACC1 Promotes Collective Migration in a Cell Line-Dependent Manner
3.3. MACC1 Promotes Colony Expansion and Migration Dependent on Proliferation
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
Appendix A
Persistent Random Walk Model
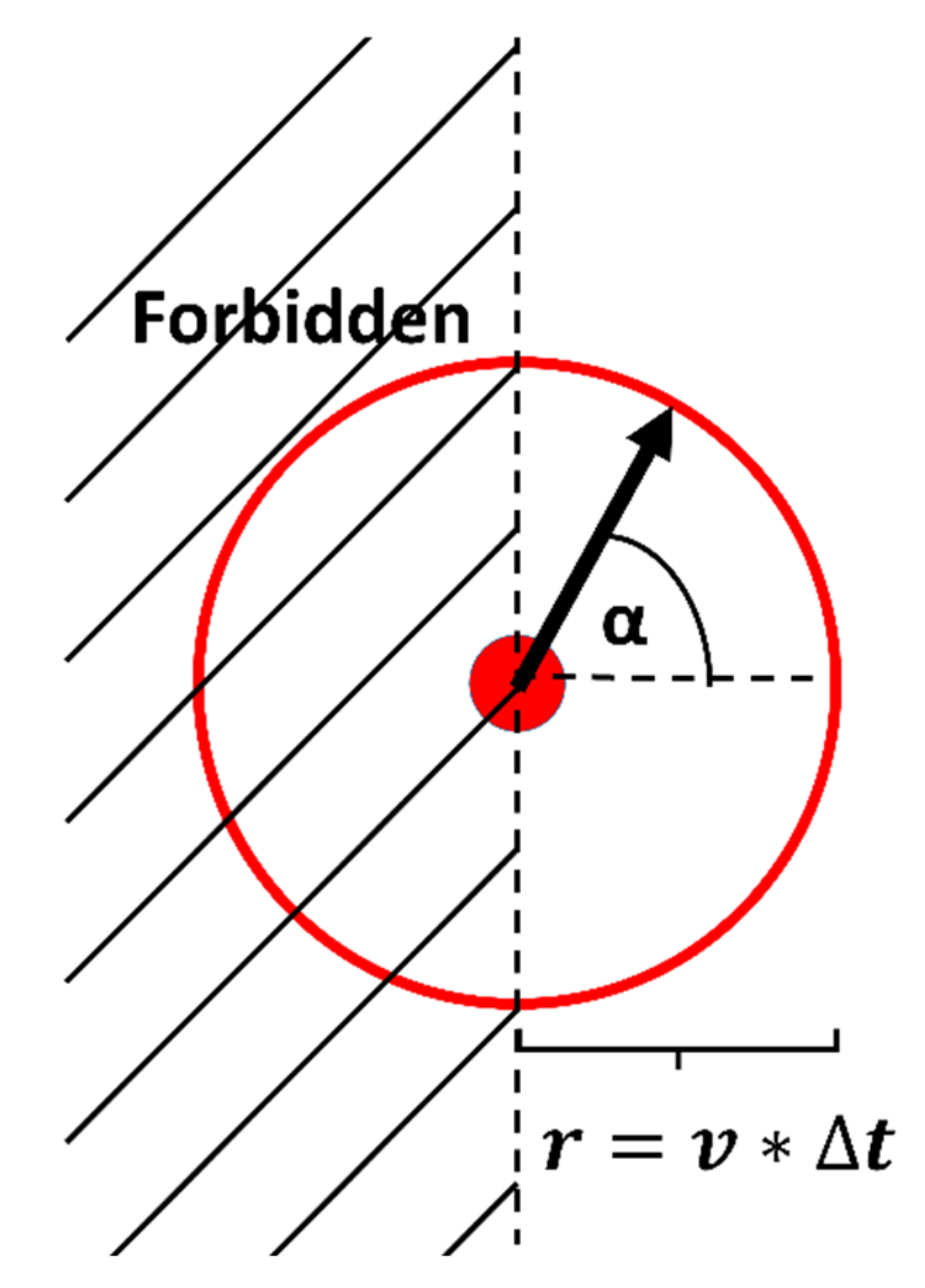
Appendix B
| Parameters | SW480\EV | SW480\MACC1 | SW620\shMACC1 | SW620\shCTL |
|---|---|---|---|---|
| Young Modulus | 35 | 33 | 40 | 40 |
| Cortical Tension | 33 | 31 | 25 | 26 |
| Single Cell Speed/Contact Area | 66 | 98 | 102 | 111 |
| Collective Migration | 75 | 75 | 32 | 32 |
| Colony Expansion | 14 | 14 | 18 | 17 |
| Colony Expansion + Mitomycin | 18 | 15 | --- | --- |
References
- Siegel, R.L.; Miller, K.D.; Fuchs, H.E.; Jemal, A. Cancer Statistics, 2022. CA Cancer J. Clin. 2022, 72, 7–33. [Google Scholar] [CrossRef] [PubMed]
- Siegel, R.L.; Miller, K.D.; Jemal, A. Cancer Statistics, 2020. CA Cancer J. Clin. 2020, 70, 7–30. [Google Scholar] [CrossRef] [PubMed]
- Siegel, R.L.; Miller, K.D.; Goding Sauer, A.; Fedewa, S.A.; Butterly, L.F.; Anderson, J.C.; Cercek, A.; Smith, R.A.; Jemal, A. Colorectal Cancer Statistics, 2020. CA Cancer J. Clin. 2020, 70, 145–164. [Google Scholar] [CrossRef]
- Stein, U.; Walther, W.; Arlt, F.; Schwabe, H.; Smith, J.; Fichtner, I.; Birchmeier, W.; Schlag, P.M. MACC1, a Newly Identified Key Regulator of HGF-MET Signaling, Predicts Colon Cancer Metastasis. Nat. Med. 2009, 15, 59–67. [Google Scholar] [CrossRef] [PubMed]
- Radhakrishnan, H.; Walther, W.; Zincke, F.; Kobelt, D.; Imbastari, F.; Erdem, M.; Kortüm, B.; Dahlmann, M.; Stein, U. MACC1—the First Decade of a Key Metastasis Molecule from Gene Discovery to Clinical Translation. Cancer Metastasis Rev. 2019, 37, 805–820. [Google Scholar] [CrossRef]
- Pichorner, A.; Sack, U.; Kobelt, D.; Kelch, I.; Arlt, F.; Smith, J.; Walther, W.; Schlag, P.M.; Stein, U. In Vivo Imaging of Colorectal Cancer Growth and Metastasis by Targeting MACC1 with ShRNA in Xenografted Mice. Clin. Exp. Metastasis 2012, 29, 573–583. [Google Scholar] [CrossRef]
- Wang, G.; Fu, Z.; Li, D. MACC1 Overexpression and Survival in Solid Tumors: A Meta-Analysis. Tumor Biol. 2015, 36, 1055–1065. [Google Scholar] [CrossRef]
- Wu, Z.; Zhou, R.; Su, Y.; Sun, L.; Liao, Y.; Liao, W. Prognostic Value of MACC1 in Digestive System Neoplasms: A Systematic Review and Meta-Analysis. Biomed. Res. Int. 2015, 2015, 252043. [Google Scholar] [CrossRef]
- Sun, D.W.; Zhang, Y.Y.; Qi, Y.; Liu, G.Q.; Chen, Y.G.; Ma, J.; Lv, G.Y. Prognostic and Clinicopathological Significance of MACC1 Expression in Hepatocellular Carcinoma Patients: A Meta-Analysis. Int. J. Clin. Exp. Med. 2015, 8, 4769–4777. [Google Scholar]
- Wang, L.; Fan, L.; Xu, H.; Jiang, H. Prognostic Significance of the Expression of Metastasis-Associated in Colon Cancer-1 in Gynecologic Cancers and Breast Cancer: A Protocol for Systematic Review and Meta-Analysis. Medicine 2021, 100, e24255. [Google Scholar] [CrossRef]
- Zhao, Y.; Dai, C.; Wang, M.; Kang, H.; Lin, S.; Yang, P.; Liu, X.; Liu, K.; Xu, P.; Zheng, Y.; et al. Clinicopathological and Prognostic Significance of Metastasisassociated in Colon Cancer-1 (MACC1) Overexpression in Colorectal Cancer: A Meta-Analysis. Oncotarget 2016, 7, 62966–62975. [Google Scholar] [CrossRef] [PubMed]
- Jin, Y.; Zhou, K.; Zhao, W.; Han, R.; Huo, X.; Yang, F.; Chen, J. Clinicopathological and Prognostic Significance of Metastasis-Associated in Colon Cancer-1 in Gastric Cancer: A Meta-Analysis. Int. J. Biol. Markers 2019, 34, 27–32. [Google Scholar] [CrossRef] [PubMed]
- Lemos, C.; Hardt, M.S.; Juneja, M.; Voss, C.; Forster, S.; Jerchow, B.; Haider, W.; Blaker, H.; Stein, U. MACC1 Induces Tumor Progression in Transgenic Mice and Colorectal Cancer Patients via Increased Pluripotency Markers Nanog and Oct4. Clin. Cancer Res. 2016, 22, 2812–2824. [Google Scholar] [CrossRef] [PubMed]
- Hohmann, T.; Grabiec, U.; Ghadban, C.; Feese, K.; Dehghani, F. The Influence of Biomechanical Properties and Cannabinoids on Tumor Invasion. Cell Adh. Migr. 2017, 11, 54–67. [Google Scholar] [CrossRef]
- Hohmann, T.; Dehghani, F. Measuring Mechanical and Adhesive Properties of Single Cells Using an Atomic Force Microscope. In Metastasis: Methods and Protocols; Stein, U.S., Ed.; Springer: New York, NY, USA, 2021; pp. 81–92. ISBN 9781071613504. [Google Scholar]
- Cartagena-Rivera, A.X.; Logue, J.S.; Waterman, C.M.; Chadwick, R.S. Actomyosin Cortical Mechanical Properties in Nonadherent Cells Determined by Atomic Force Microscopy. Biophys. J. 2016, 110, 2528–2539. [Google Scholar] [CrossRef]
- Logue, J.S.; Cartagena-Rivera, A.X.; Baird, M.A.; Davidson, M.W.; Chadwick, R.S.; Waterman, C.M. Erk Regulation of Actin Capping and Bundling by Eps8 Promotes Cortex Tension and Leader Bleb- Based Migration. Elife 2015, 4, e08314. [Google Scholar] [CrossRef]
- Hohmann, T.; Feese, K.; Ghadban, C.; Dehghani, F.; Grabiec, U. On the Influence of Cannabinoids on Cell Morphology and Motility of Glioblastoma Cells. PLoS ONE 2019, 14, e0212037. [Google Scholar] [CrossRef]
- Campos, D.; Méndez, V.; Llopis, I. Persistent Random Motion: Uncovering Cell Migration Dynamics. J. Theor. Biol. 2010, 267, 526–534. [Google Scholar] [CrossRef]
- Dabov, K.; Foi, A.; Katkovnik, V.; Egiazarian, K. Image Denoising by Sparse 3-D Transform-Domain Collaborative Filtering. IEEE Trans. Image Process. 2007, 16, 2080–2095. [Google Scholar] [CrossRef]
- Angelini, T.E.; Hannezo, E.; Trepatc, X.; Marquez, M.; Fredberg, J.J.; Weitz, D.A. Glass-like Dynamics of Collective Cell Migration. Proc. Natl. Acad. Sci. USA 2011, 108, 4714–4719. [Google Scholar] [CrossRef]
- Park, J.A.; Kim, J.H.; Bi, D.; Mitchel, J.A.; Qazvini, N.T.; Tantisira, K.; Park, C.Y.; McGill, M.; Kim, S.H.; Gweon, B.; et al. Unjamming and Cell Shape in the Asthmatic Airway Epithelium. Nat. Mater. 2015, 14, 1040–1048. [Google Scholar] [CrossRef] [PubMed]
- Garcia, S.; Hannezo, E.; Elgeti, J.; Joanny, J.F.; Silberzan, P.; Gov, N.S. Physics of Active Jamming during Collective Cellular Motion in a Monolayer. Proc. Natl. Acad. Sci. USA 2015, 112, 15314–15319. [Google Scholar] [CrossRef] [PubMed]
- Abate, A.R.; Durian, D.J. Topological Persistence and Dynamical Heterogeneities near Jamming. Phys. Rev. E-Stat. Nonlinear Soft Matter Phys. 2007, 76, 021306. [Google Scholar] [CrossRef] [PubMed]
- Tambe, D.T.; Corey Hardin, C.; Angelini, T.E.; Rajendran, K.; Park, C.Y.; Serra-Picamal, X.; Zhou, E.H.; Zaman, M.H.; Butler, J.P.; Weitz, D.A.; et al. Collective Cell Guidance by Cooperative Intercellular Forces. Nat. Mater. 2011, 10, 469–475. [Google Scholar] [CrossRef]
- Kobelt, D.; Perez-Hernandez, D.; Fleuter, C.; Dahlmann, M.; Zincke, F.; Smith, J.; Migotti, R.; Popp, O.; Burock, S.; Walther, W.; et al. The Newly Identified MEK1 Tyrosine Phosphorylation Target MACC1 Is Druggable by Approved MEK1 Inhibitors to Restrict Colorectal Cancer Metastasis. Oncogene 2021, 40, 5286–5301. [Google Scholar] [CrossRef]
- Gohlke, B.; Zincke, F.; Eckert, A.; Kobelt, D.; Preissner, S.; Liebeskind, J.M.; Gunkel, N.; Putzker, K.; Lewis, J.; Preissner, S.; et al. Real-world Evidence for Preventive Effects of Statins on Cancer Incidence: A Trans-Atlantic Analysis. Clin. Transl. Med. 2022, 12, e726. [Google Scholar] [CrossRef]
- Hagemann, C.; Fuchs, S.; Monoranu, C.M.; Herrmann, P.; Smith, J.; Hohmann, T.; Grabiec, U.; Kessler, A.F.; Dehghani, F.; Lo, M.; et al. Impact of MACC1 on Human Malignant Glioma Progression and Patients ’ Unfavorable Prognosis. J. Neurooncol. 2013, 15, 1696–1709. [Google Scholar] [CrossRef]
- Sheng, X.J.; Li, Z.; Sun, M.; Wang, Z.H.; Zhou, D.M.; Li, J.Q.; Zhao, Q.; Sun, X.F.; Liu, Q.C. MACC1 Induces Metastasis in Ovarian Carcinoma by Upregulating Hepatocyte Growth Factor Receptor C-MET. Oncol. Lett. 2014, 8, 891–897. [Google Scholar] [CrossRef]
- Zhang, C.; Hai, L.; Zhu, M.; Yu, S.P.; Li, T.; Lin, Y.; Liu, B.; Zhou, X.C.; Chen, L.; Zhao, P.F.; et al. Actin Cytoskeleton Regulator Arp2/3 Complex Is Required for DLL1 Activating Notch1 Signaling to Maintain the Stem Cell Phenotype of Glioma Initiating Cells. Oncotarget 2017, 8, 33353–33364. [Google Scholar] [CrossRef]
- Wang, J.; Wang, W.; Cai, H.; Du, B.; Zhang, L.; Ma, W.; Hu, Y.; Feng, S.; Miao, G. MACC1 Facilitates Chemoresistance and Cancer Stem Cell-like Properties of Colon Cancer Cells through the PI3K/AKT Signaling Pathway. Mol. Med. Rep. 2017, 16, 8747–8754. [Google Scholar] [CrossRef]
- Zhou, W.; Liu, L.; Xue, Y.; Zheng, J.; Liu, X.; Ma, J.; Li, Z.; Liu, Y. Combination of Endothelial-Monocyte-Activating Polypeptide-II with Temozolomide Suppress Malignant Biological Behaviors of Human Glioblastoma Stem Cells via MiR-590-3p/MACC1 Inhibiting PI3K/AKT/MTOR Signal Pathway. Front. Mol. Neurosci. 2017, 10, 68. [Google Scholar] [CrossRef] [PubMed]
- Xiang, C.; Chen, J.; Fu, P. HGF/Met Signaling in Cancer Invasion: The Impact on Cytoskeleton Remodeling. Cancers 2017, 9, 44. [Google Scholar] [CrossRef] [PubMed]
- Schober, J.M.; Cain, J.M.; Komarova, Y.A.; Borisy, G.G. Migration and Actin Protrusion in Melanoma Cells Are Regulated by EB1 Protein. Cancer Lett. 2009, 284, 30–36. [Google Scholar] [CrossRef] [PubMed]
- Martini, M.; De Santis, M.C.; Braccini, L.; Gulluni, F.; Hirsch, E. PI3K/AKT Signaling Pathway and Cancer: An Updated Review. Ann. Med. 2014, 46, 372–383. [Google Scholar] [CrossRef] [PubMed]
- Hohmann, T.; Hohmann, U.; Kolbe, M.R.; Dahlmann, M.; Kobelt, D.; Stein, U. MACC1 Driven Alterations in Cellular Biomechanics Facilitate Cell Motility in Glioblastoma. Cell Commun. Signal. 2020, 18, 85. [Google Scholar] [CrossRef] [PubMed]
- Chai, H.; Yang, Y. Effects of MACC1 SiRNA on Biological Behaviors of HeLa. Arch. Gynecol. Obstet. 2014, 289, 1271–1280. [Google Scholar] [CrossRef] [PubMed]
- Chen, S.; Zong, Z.H.; Wu, D.D.; Sun, K.X.; Liu, B.L.; Zhao, Y. The Role of Metastasis-Associated in Colon Cancer 1 (MACC1) in Endometrial Carcinoma Tumorigenesis and Progression. Mol. Carcinog. 2017, 56, 1361–1371. [Google Scholar] [CrossRef]
- Sun, L.; Li, G.; Dai, B.; Tan, W.; Zhao, H.; Li, X.; Wang, A. Silence of MACC1 Expression by RNA Interference Inhibits Proliferation, Invasion and Metastasis, and Promotes Apoptosis in U251 Human Malignant Glioma Cells. Mol. Med. Rep. 2015, 12, 3423–3431. [Google Scholar] [CrossRef]
- Mei, J.; Zhu, C.Y.; Pan, L.L.; Li, M. MACC1 Regulates the AKT/STAT3 Signaling Pathway to Induce Migration, Invasion, Cancer Stemness, and Suppress Apoptosis in Cervical Cancer Cells. Bioengineered 2022, 13, 61–70. [Google Scholar] [CrossRef]
- Vazquez, K.; Saraswathibhatla, A.; Notbohm, J. Effect of Substrate Stiffness on Friction in Collective Cell Migration. Sci. Rep. 2022, 12, 2474. [Google Scholar] [CrossRef]
- Blauth, E.; Kubitschke, H.; Gottheil, P.; Grosser, S.; Käs, J.A. Jamming in Embryogenesis and Cancer Progression. Front. Phys. 2021, 9, 666709. [Google Scholar] [CrossRef]
- Wang, L.; Wu, Y.; Lin, L.; Liu, P.; Huang, H.; Liao, W.; Zheng, D.; Zuo, Q.; Sun, L.; Huang, N.; et al. Metastasis-Associated in Colon Cancer-1 Upregulation Predicts a Poor Prognosis of Gastric Cancer, and Promotes Tumor Cell Proliferation and Invasion. Int. J. Cancer 2013, 133, 1419–1430. [Google Scholar] [CrossRef] [PubMed]
- Cheng, H.; Zhou, L.; Long, Y.; Xiang, J.; Chen, L. MACC1 Is Associated With Epithelial–Mesenchymal Transition and Can Predict Poor Prognosis in Nasopharyngeal Carcinoma. Front. Oncol. 2021, 11, 644120. [Google Scholar] [CrossRef] [PubMed]
- Gupta, V.K.; Nam, S.; Yim, D.; Camuglia, J.; Martin, J.L.; Sanders, E.N.; O’Brien, L.E.; Martin, A.C.; Kim, T.; Chaudhuri, O. The Nature of Cell Division Forces in Epithelial Monolayers. J. Cell Biol. 2021, 220, e202011106. [Google Scholar] [CrossRef]
- Ranft, J.; Basan, M.; Elgeti, J.; Joanny, J.F.; Prost, J.; Jülicher, F. Fluidization of Tissues by Cell Division and Apoptosis. Proc. Natl. Acad. Sci. USA 2010, 107, 20863–20868. [Google Scholar] [CrossRef]
- Devany, J.; Sussman, D.M.; Yamamoto, T.; Manning, M.L.; Gardel, M.L. Cell Cycle–Dependent Active Stress Drives Epithelia Remodeling. Proc. Natl. Acad. Sci. USA 2021, 118, e1917853118. [Google Scholar] [CrossRef]
- Rossen, N.S.; Tarp, J.M.; Mathiesen, J.; Jensen, M.H.; Oddershede, L.B. Long-Range Ordered Vorticity Patterns in Living Tissue Induced by Cell Division. Nat. Commun. 2014, 5, 5720. [Google Scholar] [CrossRef]
- Doostmohammadi, A.; Thampi, S.P.; Saw, T.B.; Lim, C.T.; Ladoux, B.; Yeomans, J.M. Celebrating Soft Matter’s 10th Anniversary: Cell Division: A Source of Active Stress in Cellular Monolayers. Soft Matter 2015, 11, 7328–7336. [Google Scholar] [CrossRef]
- Czajkowski, M.; Sussman, D.M.; Marchetti, M.C.; Manning, M.L. Glassy Dynamics in Models of Confluent Tissue with Mitosis and Apoptosis. Soft Matter 2019, 15, 9133–9149. [Google Scholar] [CrossRef]
- Matoz-Fernandez, D.A.; Martens, K.; Sknepnek, R.; Barrat, J.L.; Henkes, S. Cell Division and Death Inhibit Glassy Behaviour of Confluent Tissues. Soft Matter 2017, 13, 3205–3212. [Google Scholar] [CrossRef]
- Schnyder, S.K.; Molina, J.J.; Yamamoto, R. Control of Cell Colony Growth by Contact Inhibition. Sci. Rep. 2020, 10, 6173. [Google Scholar] [CrossRef] [PubMed]
- Puliafito, A.; Hufnagel, L.; Neveu, P.; Streichan, S.; Sigal, A.; Fygenson, D.K.; Shraiman, B.I. Collective and Single Cell Behavior in Epithelial Contact Inhibition. Proc. Natl. Acad. Sci. USA 2012, 109, 739–744. [Google Scholar] [CrossRef] [PubMed]
- Gauquelin, E.; Tlili, S.; Gay, C.; Peyret, G.; Mège, R.M.; Fardin, M.A.; Ladoux, B. Influence of Proliferation on the Motions of Epithelial Monolayers Invading Adherent Strips. Soft Matter 2019, 15, 2798–2810. [Google Scholar] [CrossRef] [PubMed]
- Bruun, J.; Kolberg, M.; Nesland, J.M.; Svindland, A.; Nesbakken, A.; Lothe, R.A. Prognostic Significance of β-Catenin, E-Cadherin, and SOX9 in Colorectal Cancer: Results from a Large Population-Representative Series. Front. Oncol. 2014, 4, 118. [Google Scholar] [CrossRef]
- Arlt, F.; Stein, U. Colon Cancer Metastasis: MACC1 and Met as Metastatic Pacemakers. Int. J. Biochem. Cell Biol. 2009, 41, 2356–2359. [Google Scholar] [CrossRef]
- Zhang, K.; Tian, F.; Zhang, Y.; Zhu, Q.; Xue, N.; Zhu, H.; Wang, H.; Guo, X. MACC1 Is Involved in the Regulation of Proliferation, Colony Formation, Invasion Ability, Cell Cycle Distribution, Apoptosis and Tumorigenicity by Altering Akt Signaling Pathway in Human Osteosarcoma. Tumor Biol. 2014, 35, 2537–2548. [Google Scholar] [CrossRef]
- Escaff, D.; Toral, R.; Van Den Broeck, C.; Lindenberg, K. A Continuous-Time Persistent Random Walk Model for Flocking. Chaos 2018, 28, 075507. [Google Scholar] [CrossRef]
- Luzhansky, I.D.; Schwartz, A.D.; Cohen, J.D.; Macmunn, J.P.; Barney, L.E.; Jansen, L.E.; Peyton, S.R. Anomalously Diffusing and Persistently Migrating Cells in 2D and 3D Culture Environments. APL Bioeng. 2018, 2, 026112. [Google Scholar] [CrossRef]
- Ecker, N.; Kruse, K. Excitable Actin Dynamics and Amoeboid Cell Migration. PLoS ONE 2021, 16, e0246311. [Google Scholar] [CrossRef]

Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Hohmann, T.; Hohmann, U.; Dahlmann, M.; Kobelt, D.; Stein, U.; Dehghani, F. MACC1-Induced Collective Migration Is Promoted by Proliferation Rather Than Single Cell Biomechanics. Cancers 2022, 14, 2857. https://doi.org/10.3390/cancers14122857
Hohmann T, Hohmann U, Dahlmann M, Kobelt D, Stein U, Dehghani F. MACC1-Induced Collective Migration Is Promoted by Proliferation Rather Than Single Cell Biomechanics. Cancers. 2022; 14(12):2857. https://doi.org/10.3390/cancers14122857
Chicago/Turabian StyleHohmann, Tim, Urszula Hohmann, Mathias Dahlmann, Dennis Kobelt, Ulrike Stein, and Faramarz Dehghani. 2022. "MACC1-Induced Collective Migration Is Promoted by Proliferation Rather Than Single Cell Biomechanics" Cancers 14, no. 12: 2857. https://doi.org/10.3390/cancers14122857
APA StyleHohmann, T., Hohmann, U., Dahlmann, M., Kobelt, D., Stein, U., & Dehghani, F. (2022). MACC1-Induced Collective Migration Is Promoted by Proliferation Rather Than Single Cell Biomechanics. Cancers, 14(12), 2857. https://doi.org/10.3390/cancers14122857







