Clinical Assay for the Early Detection of Colorectal Cancer Using Mass Spectrometric Wheat Germ Agglutinin Multiple Reaction Monitoring
Simple Summary
Abstract
1. Introduction
2. Materials and Methods
2.1. Materials
2.2. Patients and Specimens
2.3. Wheat Germ Agglutinin Chromatograhpy and Sample Preparation
2.4. Nano-LC-MS/MS Analyses
2.5. MRM Method
2.6. Statistical Anaylses
3. Results
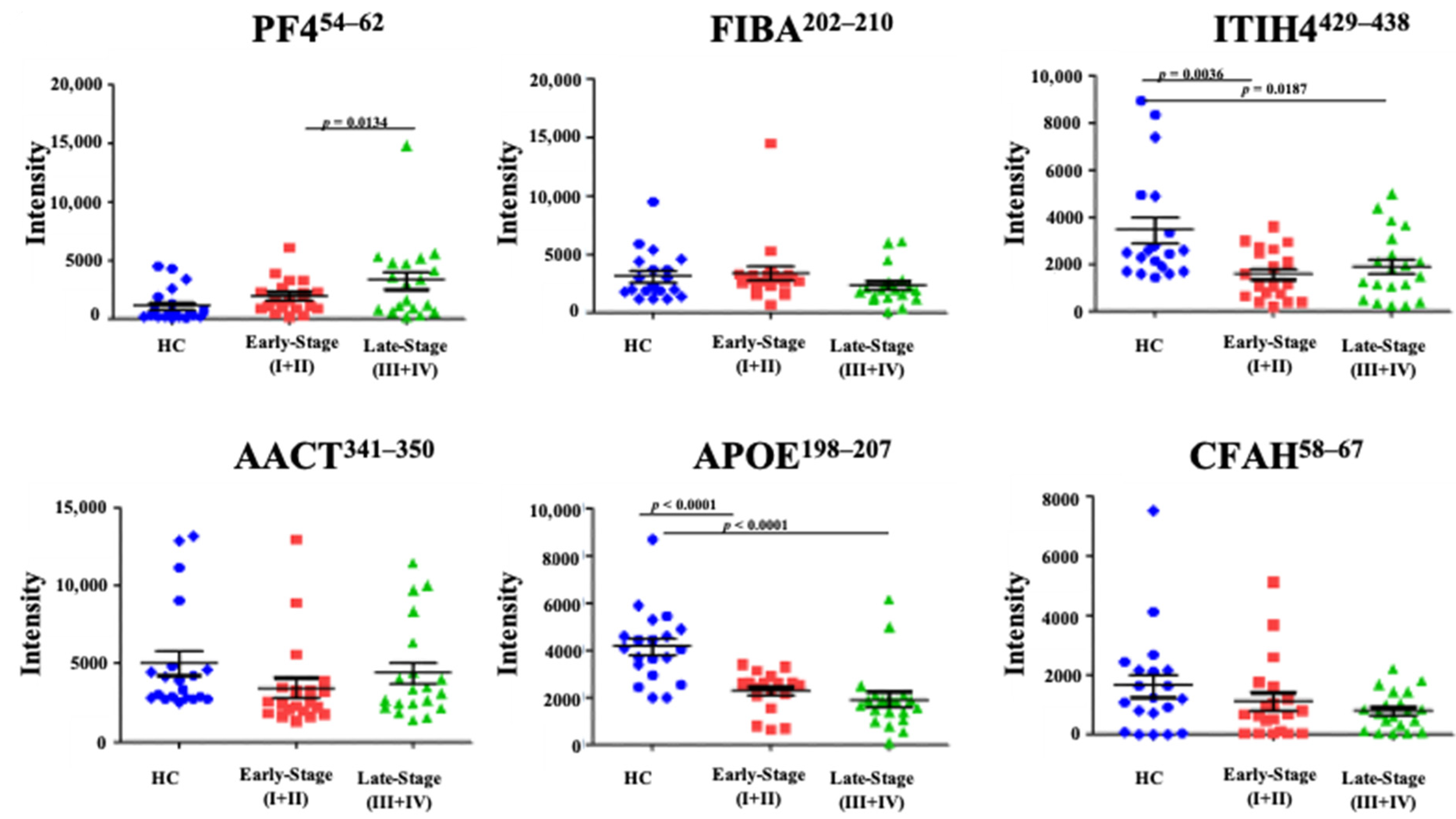
3.1. Discovery MS
3.2. Targeted LC-MS/MS
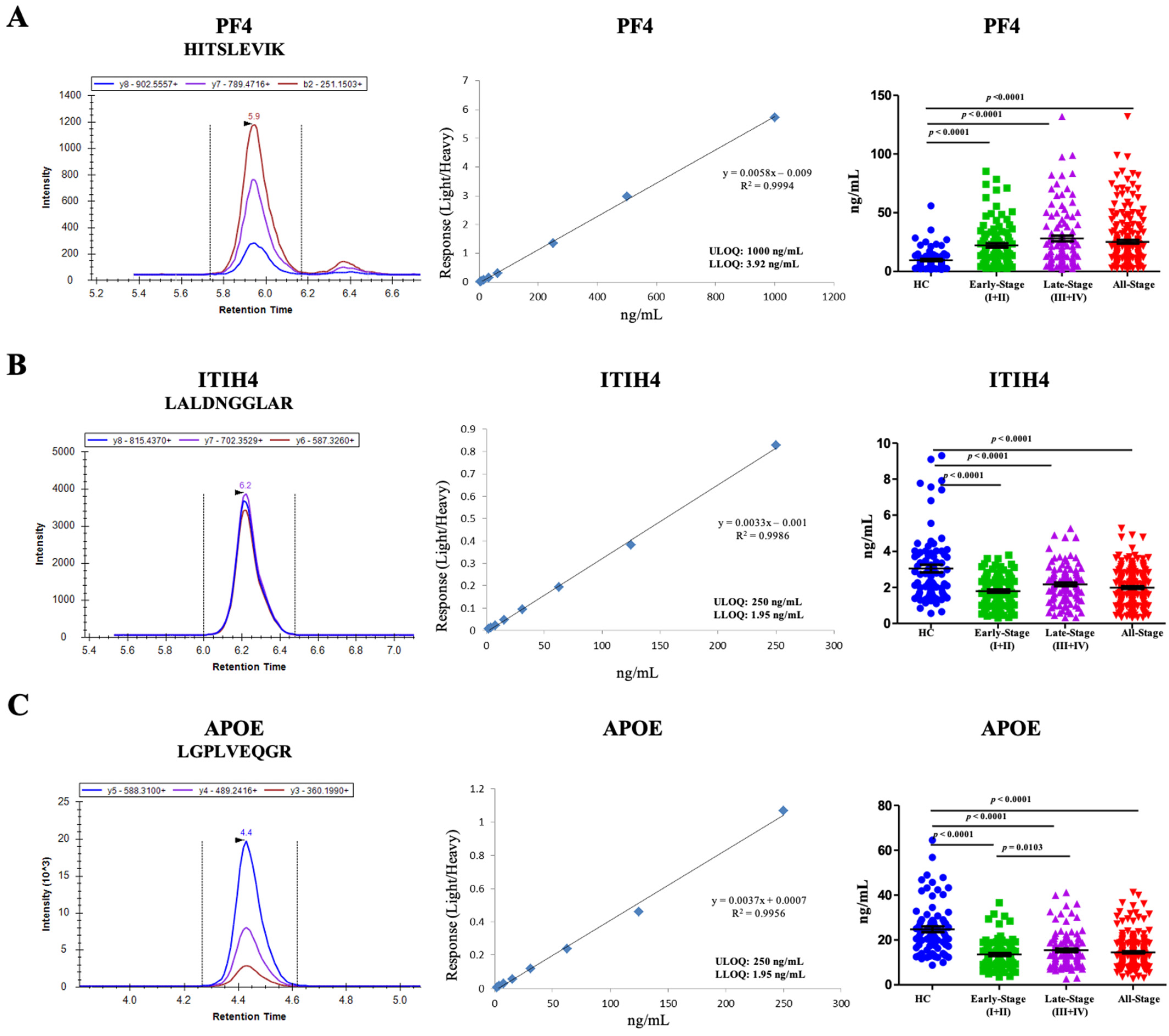
3.3. Analytical Method Development
3.4. Semi-Quantification of Peptides in Large Samples
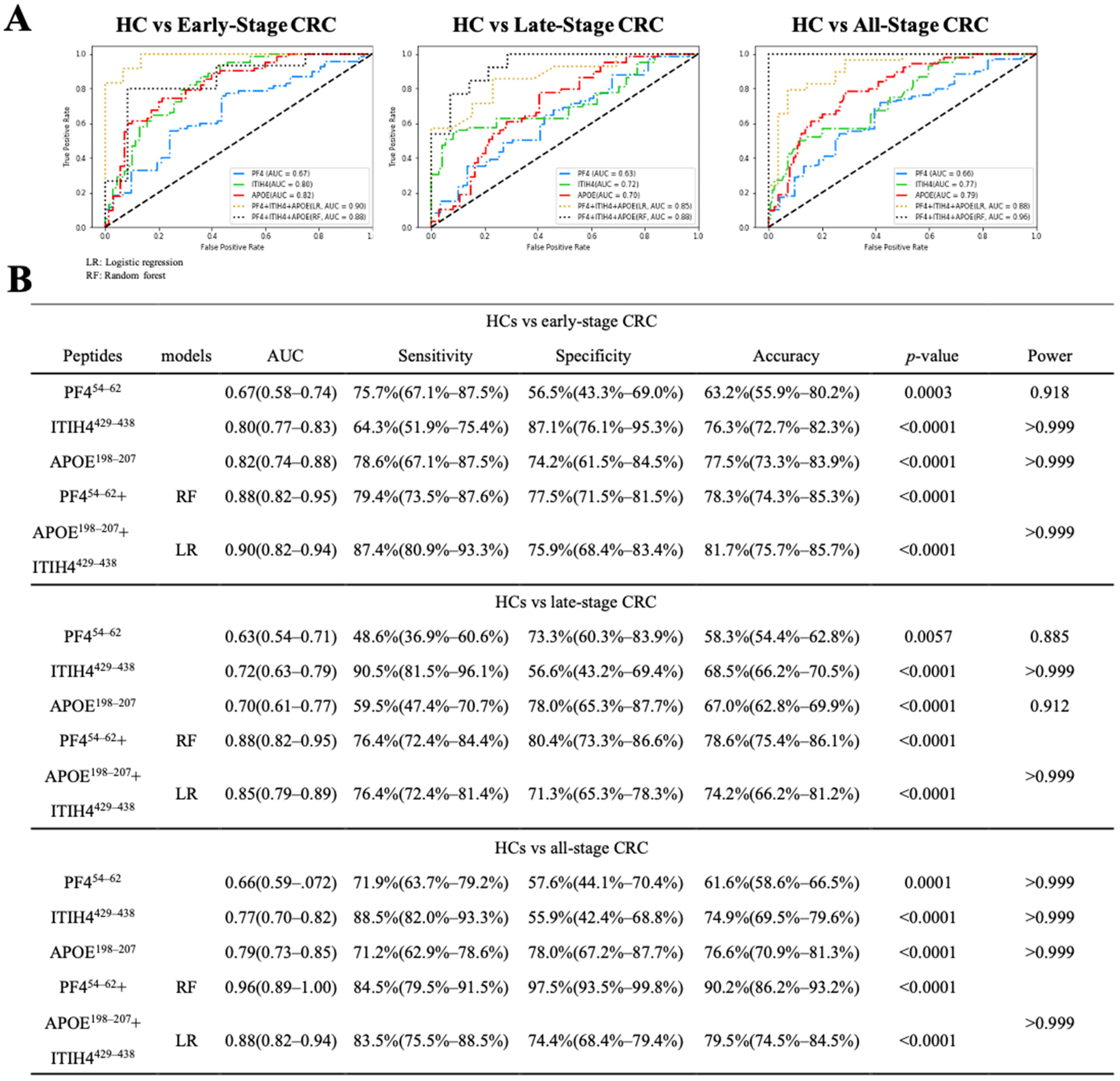
3.5. Diagnostic Performance
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Kuo, C.-N.; Liao, Y.-M.; Kuo, L.-N.; Tsai, H.-J.; Chang, W.-C.; Yen, Y. Cancers in Taiwan: Practical insight from epidemiology, treatments, biomarkers, and cost. J. Formos. Med. Assoc. 2020, 119, 1731–1741. [Google Scholar] [CrossRef]
- Bhandari, A.; Woodhouse, M.; Gupta, S. Colorectal cancer is a leading cause of cancer incidence and mortality among adults younger than 50 years in the USA: A SEER-based analysis with comparison to other young-onset cancers. J. Investig. Med. 2017, 65, 311–315. [Google Scholar] [CrossRef]
- Douaiher, J.; Ravipati, A.; Grams, B.; Chowdhury, S.; Alatise, O.; Are, C. Colorectal cancer-global burden, trends, and geographical variations. J. Surg. Oncol. 2017, 115, 619–630. [Google Scholar] [CrossRef] [PubMed]
- Ang, C.S.; Baker, M.S.; Nice, E.C. Mass Spectrometry-Based Analysis for the Discovery and Validation of Potential Colorectal Cancer Stool Biomarkers. Methods Enzymol. 2017, 586, 247–274. [Google Scholar] [CrossRef] [PubMed]
- Huo, Y.R.; Huang, Y.; Liauw, W.; Zhao, J.; Morris, D.L. Prognostic Value of Carcinoembryonic Antigen (CEA), AFP, CA19-9 and CA125 for Patients with Colorectal Cancer with Peritoneal Carcinomatosis Treated by Cytoreductive Surgery and Intraperitoneal Chemotherapy. Anticancer Res. 2016, 36, 1041–1049. [Google Scholar] [PubMed]
- Gao, Y.; Wang, J.; Zhou, Y.; Sheng, S.; Qian, S.Y.; Huo, X. Evaluation of Serum CEA, CA19-9, CA72-4, CA125 and Ferritin as Diagnostic Markers and Factors of Clinical Parameters for Colorectal Cancer. Sci. Rep. 2018, 8, 2732. [Google Scholar] [CrossRef]
- Han, X.; Aslanian, A.; Yates, J.R., 3rd. Mass spectrometry for proteomics. Curr. Opin. Chem. Biol. 2008, 12, 483–490. [Google Scholar] [CrossRef]
- Saleem, S.; Tariq, S.; Aleem, I.; Sadr-Ul, S.; Tahseen, M.; Atiq, A.; Hassan, S.; Abu Bakar, M.; Khattak, S.; Syed, A.A.; et al. Proteomics analysis of colon cancer progression. Clin. Proteom. 2019, 16, 44. [Google Scholar] [CrossRef]
- Simpson, K.L.; Whetton, A.D.; Dive, C. Quantitative mass spectrometry-based techniques for clinical use: Biomarker identification and quantification. J. Chromatogr. B Anal. Technol. Biomed. Life Sci. 2009, 877, 1240–1249. [Google Scholar] [CrossRef] [PubMed]
- Shiromizu, T.; Kume, H.; Ishida, M.; Adachi, J.; Kano, M.; Matsubara, H.; Tomonaga, T. Quantitation of putative colorectal cancer biomarker candidates in serum extracellular vesicles by targeted proteomics. Sci. Rep. 2017, 7, 12782. [Google Scholar] [CrossRef]
- Beretov, J.; Wasinger, V.C.; Millar, E.K.A.; Schwartz, P.; Graham, P.H.; Li, Y. Proteomic Analysis of Urine to Identify Breast Cancer Biomarker Candidates Using a Label-Free LC-MS/MS Approach. PLoS ONE 2015, 10, e0141876. [Google Scholar] [CrossRef]
- Kumar, V.; Ray, S.; Ghantasala, S.; Srivastava, S. An Integrated Quantitative Proteomics Workflow for Cancer Biomarker Discovery and Validation in Plasma. Front. Oncol. 2020, 10. [Google Scholar] [CrossRef]
- Dardé, V.M.; Barderas, M.G.; Vivanco, F. Multiple reaction monitoring (MRM) of plasma proteins in cardiovascular proteomics. Methods Mol. Biol 2013, 1000, 191–199. [Google Scholar] [CrossRef]
- Arora, A.; Patil, V.; Kundu, P.; Kondaiah, P.; Hegde, A.S.; Arivazhagan, A.; Santosh, V.; Pal, D.; Somasundaram, K. Serum biomarkers identification by iTRAQ and verification by MRM: S100A8/S100A9 levels predict tumor-stroma involvement and prognosis in Glioblastoma. Sci. Rep. 2019, 9, 2749. [Google Scholar] [CrossRef]
- Kobata, A.; Amano, J. Altered glycosylation of proteins produced by malignant cells, and application for the diagnosis and immunotherapy of tumours. Immunol. Cell Biol. 2005, 83, 429–439. [Google Scholar] [CrossRef]
- Taniguchi, N.; Kizuka, Y. Glycans and cancer: Role of N-glycans in cancer biomarker, progression and metastasis, and therapeutics. Adv. Cancer Res. 2015, 126, 11–51. [Google Scholar] [CrossRef]
- Durand, G.; Seta, N. Protein glycosylation and diseases: Blood and urinary oligosaccharides as markers for diagnosis and therapeutic monitoring. Clin. Chem. 2000, 46, 795–805. [Google Scholar] [CrossRef] [PubMed]
- Zhao, J.; Qiu, W.; Simeone, D.M.; Lubman, D.M. N-linked glycosylation profiling of pancreatic cancer serum using capillary liquid phase separation coupled with mass spectrometric analysis. J. Proteome Res. 2007, 6, 1126–1138. [Google Scholar] [CrossRef] [PubMed]
- Wei, H.; Cheng, Z.; Ouyang, C.; Zhang, Y.; Hu, Y.; Chen, S.; Wang, C.; Lu, F.; Zhang, J.; Wang, Y.; et al. Glycoprotein screening in colorectal cancer based on differentially expressed Tn antigen. Oncol. Rep. 2016, 36, 1313–1324. [Google Scholar] [CrossRef] [PubMed]
- Ju, L.; Wang, Y.; Xie, Q.; Xu, X.; Li, Y.; Chen, Z.; Li, Y. Elevated level of serum glycoprotein bifucosylation and prognostic value in Chinese breast cancer. Glycobiology 2016, 26, 460–471. [Google Scholar] [CrossRef]
- Duskova, K.; Vesely, S. Prostate specific antigen. Current clinical application and future prospects. Biomed. Pap. Med. Fac. Univ. Palacky Olomouc Czech Repub. 2015, 159, 18–26. [Google Scholar] [CrossRef] [PubMed]
- Kailemia, M.J.; Park, D.; Lebrilla, C.B. Glycans and glycoproteins as specific biomarkers for cancer. Anal. Bioanal. Chem. 2017, 409, 395–410. [Google Scholar] [CrossRef] [PubMed]
- Poruk, K.E.; Gay, D.Z.; Brown, K.; Mulvihill, J.D.; Boucher, K.M.; Scaife, C.L.; Firpo, M.A.; Mulvihill, S.J. The clinical utility of CA 19-9 in pancreatic adenocarcinoma: Diagnostic and prognostic updates. Curr. Mol. Med. 2013, 13, 340–351. [Google Scholar] [CrossRef]
- Ideo, H.; Kondo, J.; Nomura, T.; Nonomura, N.; Inoue, M.; Amano, J. Study of glycosylation of prostate-specific antigen secreted by cancer tissue-originated spheroids reveals new candidates for prostate cancer detection. Sci. Rep. 2020, 10, 2708. [Google Scholar] [CrossRef] [PubMed]
- Aimjongjun, S.; Reamtong, O.; Janvilisri, T. Lectin affinity chromatography and quantitative proteomic analysis reveal that galectin-3 is associated with metastasis in nasopharyngeal carcinoma. Sci. Rep. 2020, 10, 16462. [Google Scholar] [CrossRef] [PubMed]
- Totten, S.M.; Adusumilli, R.; Kullolli, M.; Tanimoto, C.; Brooks, J.D.; Mallick, P.; Pitteri, S.J. Multi-lectin Affinity Chromatography and Quantitative Proteomic Analysis Reveal Differential Glycoform Levels between Prostate Cancer and Benign Prostatic Hyperplasia Sera. Sci. Rep. 2018, 8, 6509. [Google Scholar] [CrossRef]
- Zhao, J.; Patwa, T.H.; Lubman, D.M.; Simeone, D.M. Protein biomarkers in cancer: Natural glycoprotein microarray approaches. Curr. Opin. Mol. 2008, 10, 602–610. [Google Scholar]
- Lee, A.; Nakano, M.; Hincapie, M.; Kolarich, D.; Baker, M.S.; Hancock, W.S.; Packer, N.H. The lectin riddle: Glycoproteins fractionated from complex mixtures have similar glycomic profiles. Omics 2010, 14, 487–499. [Google Scholar] [CrossRef]
- Chou, T.Y.; Hart, G.W. O-linked N-acetylglucosamine and cancer: Messages from the glycosylation of c-Myc. Adv. Exp. Med. Biol. 2001, 491, 413–418. [Google Scholar] [CrossRef]
- Song, E.; Zhu, R.; Hammoud, Z.T.; Mechref, Y. LC–MS/MS Quantitation of Esophagus Disease Blood Serum Glycoproteins by Enrichment with Hydrazide Chemistry and Lectin Affinity Chromatography. J. Proteome Res. 2014, 13, 4808–4820. [Google Scholar] [CrossRef]
- Drake, P.M.; Schilling, B.; Niles, R.K.; Prakobphol, A.; Li, B.; Jung, K.; Cho, W.; Braten, M.; Inerowicz, H.D.; Williams, K.; et al. Lectin Chromatography/Mass Spectrometry Discovery Workflow Identifies Putative Biomarkers of Aggressive Breast Cancers. J. Proteome Res. 2012, 11, 2508–2520. [Google Scholar] [CrossRef]
- Ahn, Y.H.; Shin, P.M.; Oh, N.R.; Park, G.W.; Kim, H.; Yoo, J.S. A lectin-coupled, targeted proteomic mass spectrometry (MRM MS) platform for identification of multiple liver cancer biomarkers in human plasma. J. Proteom. 2012, 75, 5507–5515. [Google Scholar] [CrossRef] [PubMed]
- Deutsch, E.W.; Csordas, A.; Sun, Z.; Jarnuczak, A.; Perez-Riverol, Y.; Ternent, T.; Campbell, D.S.; Bernal-Llinares, M.; Okuda, S.; Kawano, S.; et al. The ProteomeXchange consortium in 2017: Supporting the cultural change in proteomics public data deposition. Nucleic Acids Res. 2017, 45, D1100–D1106. [Google Scholar] [CrossRef]
- Rauniyar, N. Parallel Reaction Monitoring: A Targeted Experiment Performed Using High Resolution and High Mass Accuracy Mass Spectrometry. Int. J. Mol. Sci. 2015, 16, 28566–28581. [Google Scholar] [CrossRef] [PubMed]
- Yuan, L.; Zhu, M. Quantitative Bioanalysis of Proteins by Mass Spectrometry. Mater. Methods 2015, 5. [Google Scholar] [CrossRef]
- Tsai, K.-L.; Chang, C.-C.; Chang, Y.-S.; Lu, Y.-Y.; Tsai, I.J.; Chen, J.-H.; Lin, S.-H.; Tai, C.-C.; Lin, Y.-F.; Chang, H.-W.; et al. Isotypes of autoantibodies against novel differential 4-hydroxy-2-nonenal-modified peptide adducts in serum is associated with rheumatoid arthritis in Taiwanese women. BMC Med. Inform. Decis. Mak. 2021, 21, 49. [Google Scholar] [CrossRef]
- Diz, A.P.; Truebano, M.; Skibinski, D.O.F. The consequences of sample pooling in proteomics: An empirical study. Electrophoresis 2009, 30, 2967–2975. [Google Scholar] [CrossRef] [PubMed]
- Rodríguez-Paredes, M.; Lyko, F. The importance of non-histone protein methylation in cancer therapy. Nat. Rev. Mol. Cell Biol. 2019, 20, 569–570. [Google Scholar] [CrossRef]
- Stowell, S.R.; Ju, T.; Cummings, R.D. Protein glycosylation in cancer. Annu. Rev. Pathol. 2015, 10, 473–510. [Google Scholar] [CrossRef] [PubMed]
- Illiano, A.; Pinto, G.; Melchiorre, C.; Carpentieri, A.; Faraco, V.; Amoresano, A. Protein Glycosylation Investigated by Mass Spectrometry: An Overview. Cells 2020, 9, 1986. [Google Scholar] [CrossRef] [PubMed]
- Phueaouan, T.; Chaiyawat, P.; Netsirisawan, P.; Chokchaichamnankit, D.; Punyarit, P.; Srisomsap, C.; Svasti, J.; Champattanachai, V. Aberrant O-GlcNAc-modified proteins expressed in primary colorectal cancer. Oncol. Rep. 2013, 30, 2929–2936. [Google Scholar] [CrossRef] [PubMed]
- Krześlak, A.; Jóźwiak, P.; Lipińska, A. Down-regulation of β-N-acetyl-D-glucosaminidase increases Akt1 activity in thyroid anaplastic cancer cells. Oncol. Rep. 2011, 26, 743–749. [Google Scholar] [CrossRef]
- Zauber, H.; Schüler, V.; Schulze, W. Systematic evaluation of reference protein normalization in proteomic experiments. Front. Plant Sci. 2013, 4, 25. [Google Scholar] [CrossRef] [PubMed]
- Faria, M.; Halquist, M.S.; Yuan, M.; Mylott, W., Jr.; Jenkins, R.G.; Karnes, H.T. An extended stable isotope-labeled signature peptide internal standard for tracking immunocapture of human plasma osteopontin for LC-MS/MS quantification. Biomed. Chromatogr. BMC 2015, 29, 1780–1782. [Google Scholar] [CrossRef]
- Faria, M.; Halquist, M.S. Internal Standards for Absolute Quantification of Large Molecules (Proteins) from Biological Matrices by LC-MS/MS. Calibration Valid. Anal. Methods A Sampl. Curr. Approaches 2018, 61. [Google Scholar] [CrossRef]
- Chen, H.F.; Shiao, C.Y.; Wu, M.Y.; Lin, Y.C.; Chen, H.H.; Chang, W.C.; Wu, M.S.; Kao, C.C.; Tsai, I.L. Quantitative determination of human IgA subclasses and their Fc-glycosylation patterns in plasma by using a peptide analogue internal standard and ultra-high-performance liquid chromatography/triple quadrupole mass spectrometry. Rapid Commun. Mass Spectrom. 2020, 34, e8606. [Google Scholar] [CrossRef]
- Kim, H.; Sohn, A.; Yeo, I.; Yu, S.J.; Yoon, J.H.; Kim, Y. Clinical Assay for AFP-L3 by Using Multiple Reaction Monitoring-Mass Spectrometry for Diagnosing Hepatocellular Carcinoma. Clin. Chem. 2018, 64, 1230–1238. [Google Scholar] [CrossRef] [PubMed]
- Kato, K. Algorithm for in vitro diagnostic multivariate index assay. Breast Cancer 2009, 16, 248–251. [Google Scholar] [CrossRef]
- Zang, X.; Jones, C.M.; Long, T.Q.; Monge, M.E.; Zhou, M.; Walker, L.D.; Mezencev, R.; Gray, A.; McDonald, J.F.; Fernández, F.M. Feasibility of detecting prostate cancer by ultraperformance liquid chromatography-mass spectrometry serum metabolomics. J. Proteome Res. 2014, 13, 3444–3454. [Google Scholar] [CrossRef]
- Lee, H.J.; Kim, Y.T.; Park, P.J.; Shin, Y.S.; Kang, K.N.; Kim, Y.; Kim, C.W. A novel detection method of non-small cell lung cancer using multiplexed bead-based serum biomarker profiling. J. Thorac. Cardiovasc. Surg. 2012, 143, 421–427. [Google Scholar] [CrossRef] [PubMed]
- Marín-Vicente, C.; Mendes, M.; de los Ríos, V.; Fernández-Aceñero, M.J.; Casal, J.I. Identification and Validation of Stage-Associated Serum Biomarkers in Colorectal Cancer Using MS-Based Procedures. Proteom. Clin. Appl. 2020, 14, 1900052. [Google Scholar] [CrossRef]
- Xie, Y.; Chen, L.; Lv, X.; Hou, G.; Wang, Y.; Jiang, C.; Zhu, H.; Xu, N.; Wu, L.; Lou, X.; et al. The levels of serine proteases in colon tissue interstitial fluid and serum serve as an indicator of colorectal cancer progression. Oncotarget 2016, 7, 32592–32606. [Google Scholar] [CrossRef] [PubMed]
- Bhardwaj, M.; Gies, A.; Weigl, K.; Tikk, K.; Benner, A.; Schrotz-King, P.; Borchers, C.H.; Brenner, H. Evaluation and Validation of Plasma Proteins Using Two Different Protein Detection Methods for Early Detection of Colorectal Cancer. Cancers 2019, 11, 1426. [Google Scholar] [CrossRef] [PubMed]
- Bhardwaj, M.; Weigl, K.; Tikk, K.; Holland-Letz, T.; Schrotz-King, P.; Borchers, C.H.; Brenner, H. Multiplex quantitation of 270 plasma protein markers to identify a signature for early detection of colorectal cancer. Eur. J. Cancer 2020, 127, 30–40. [Google Scholar] [CrossRef] [PubMed]
- Lord, S.J.; Irwig, L.; Simes, R.J. When is measuring sensitivity and specificity sufficient to evaluate a diagnostic test, and when do we need randomized trials? Ann. Intern. Med. 2006, 144, 850–855. [Google Scholar] [CrossRef] [PubMed]
- Ramdzan, A.R.; Abd Rahim, M.A.; Mohamad Zaki, A.; Zaidun, Z.; Mohammed Nawi, A. Diagnostic Accuracy of FOBT and Colorectal Cancer Genetic Testing: A Systematic Review & Meta-Analysis. Ann. Glob. Health 2019, 85. [Google Scholar] [CrossRef]
- Ruytinx, P.; Proost, P.; Struyf, S. CXCL4 and CXCL4L1 in cancer. Cytokine 2018, 109, 65–71. [Google Scholar] [CrossRef]
- Lord, M.S.; Cheng, B.; Farrugia, B.L.; McCarthy, S.; Whitelock, J.M. Platelet Factor 4 Binds to Vascular Proteoglycans and Controls Both Growth Factor Activities and Platelet Activation. J. Biol. Chem. 2017, 292, 4054–4063. [Google Scholar] [CrossRef]
- Sikara, M.P.; Routsias, J.G.; Samiotaki, M.; Panayotou, G.; Moutsopoulos, H.M.; Vlachoyiannopoulos, P.G. β2 Glycoprotein I (β2GPI) binds platelet factor 4 (PF4): Implications for the pathogenesis of antiphospholipid syndrome. Blood 2010, 115, 713–723. [Google Scholar] [CrossRef]
- Muramatsu, T.; Muramatsu, H.; Kojima, T. Identification of proteoglycan-binding proteins. Methods Enzymol. 2006, 416, 263–278. [Google Scholar] [CrossRef] [PubMed]
- Pucci, F.; Rickelt, S.; Newton, A.P.; Garris, C.; Nunes, E.; Evavold, C.; Pfirschke, C.; Engblom, C.; Mino-Kenudson, M.; Hynes, R.O.; et al. PF4 Promotes Platelet Production and Lung Cancer Growth. Cell Rep. 2016, 17, 1764–1772. [Google Scholar] [CrossRef]
- Zhang, Y.; Gao, J.; Wang, X.; Deng, S.; Ye, H.; Guan, W.; Wu, M.; Zhu, S.; Yu, Y.; Han, W. CXCL4 mediates tumor regrowth after chemotherapy by suppression of antitumor immunity. Cancer Biol. 2015, 16, 1775–1783. [Google Scholar] [CrossRef]
- Peterson, J.E.; Zurakowski, D.; Italiano, J.E., Jr.; Michel, L.V.; Connors, S.; Oenick, M.; D’Amato, R.J.; Klement, G.L.; Folkman, J. VEGF, PF4 and PDGF are elevated in platelets of colorectal cancer patients. Angiogenesis 2012, 15, 265–273. [Google Scholar] [CrossRef] [PubMed]
- Bhanumathy, C.D.; Tang, Y.; Monga, S.P.; Katuri, V.; Cox, J.A.; Mishra, B.; Mishra, L. Itih-4, a serine protease inhibitor regulated in interleukin-6-dependent liver formation: Role in liver development and regeneration. Dev. Dyn. 2002, 223, 59–69. [Google Scholar] [CrossRef] [PubMed]
- Tang, Y.; Kitisin, K.; Jogunoori, W.; Li, C.; Deng, C.X.; Mueller, S.C.; Ressom, H.W.; Rashid, A.; He, A.R.; Mendelson, J.S.; et al. Progenitor/stem cells give rise to liver cancer due to aberrant TGF-beta and IL-6 signaling. Proc. Natl. Acad. Sci. USA 2008, 105, 2445–2450. [Google Scholar] [CrossRef] [PubMed]
- Huang, M.; Zhang, W.; Zhao, B.; Li, L. Relationship between inter-α-trypsin inhibitor heavy chain 4 and ovarian cancer. Chin. J. Cancer Res. 2019, 31, 955–964. [Google Scholar] [CrossRef] [PubMed]
- Lee, E.J.; Yang, S.H.; Kim, K.J.; Cha, H.; Lee, S.J.; Kim, J.H.; Song, J.; Chun, K.H.; Seong, J. Inter-alpha Inhibitor H4 as a Potential Biomarker Predicting the Treatment Outcomes in Patients with Hepatocellular Carcinoma. Cancer Res. Treat. 2018, 50, 646–657. [Google Scholar] [CrossRef] [PubMed]
- Hamm, A.; Veeck, J.; Bektas, N.; Wild, P.J.; Hartmann, A.; Heindrichs, U.; Kristiansen, G.; Werbowetski-Ogilvie, T.; Del Maestro, R.; Knuechel, R.; et al. Frequent expression loss of Inter-alpha-trypsin inhibitor heavy chain (ITIH) genes in multiple human solid tumors: A systematic expression analysis. BMC Cancer 2008, 8, 25. [Google Scholar] [CrossRef]
- Jiang, X.; Bai, X.Y.; Li, B.; Li, Y.; Xia, K.; Wang, M.; Li, S.; Wu, H. Plasma Inter-Alpha-Trypsin Inhibitor Heavy Chains H3 and H4 Serve as Novel Diagnostic Biomarkers in Human Colorectal Cancer. Dis. Markers 2019, 2019, 5069614. [Google Scholar] [CrossRef]
- Marais, A.D. Apolipoprotein E in lipoprotein metabolism, health and cardiovascular disease. Pathology 2019, 51, 165–176. [Google Scholar] [CrossRef]
- Long, J.; Zhang, C.-J.; Zhu, N.; Du, K.; Yin, Y.-F.; Tan, X.; Liao, D.-F.; Qin, L. Lipid metabolism and carcinogenesis, cancer development. Am. J. Cancer Res. 2018, 8, 778–791. [Google Scholar] [PubMed]
- Pencheva, N.; Tran, H.; Buss, C.; Huh, D.; Drobnjak, M.; Busam, K.; Tavazoie, S.F. Convergent multi-miRNA targeting of ApoE drives LRP1/LRP8-dependent melanoma metastasis and angiogenesis. Cell 2012, 151, 1068–1082. [Google Scholar] [CrossRef] [PubMed]
- Buss, L.A.; Mandani, A.; Phillips, E.; Scott, N.J.A.; Currie, M.J.; Dachs, G.U. Characterisation of a Mouse Model of Breast Cancer with Metabolic Syndrome. In Vivo 2018, 32, 1071–1080. [Google Scholar] [CrossRef] [PubMed]
- El-Bahrawy, A.; Tarhuni, A.; Kim, H.; Subramaniam, V.; Benslimane, I.; Abd Elmageed, Z.Y.; Okpechi, S.; Ghonim, M.; Hemeida, R.; Abo-Yousef, A.; et al. ApoEdeficiency promotes colon inflammation and enhances inflammatory potential oxidized-LDL and TNF-α in colon epithelial cells. Biosci. Rep. 2016, 36. [Google Scholar] [CrossRef]
- An, H.J.; Koh, H.M.; Song, D.H. Apolipoprotein E is a predictive marker for assessing non-small cell lung cancer patients with lymph node metastasis. Pathol. Res. Pract. 2019, 215, 152607. [Google Scholar] [CrossRef]
| Varible | Discovery Set | Validation Set | ||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| group | HC | early-stage CRC | late-stage CRC | HC | early-stage CRC | late-stage CRC | ||||
| number of samples | n = 40 | n = 40 | n = 40 | n = 80 | n = 100 | n = 106 | ||||
| stage I | stage II | stage III | stage IV | stage I | stage II | stage III | stage IV | |||
| n =20 | n =20 | n =20 | n =20 | n = 47 | n =53 | n =50 | n =56 | |||
| male: female ratio | 24:16 | 24:16 | 24:16 | 46:34 | 71:29 | 66:40 | ||||
| mean age years ± SD | 57.65 ± 3.48 | 52.5 ± 6.87 | 51.25 ± 4.85 | 39.43 ± 11.14 | 70.84 ± 10.7 | 67.33 ± 10.06 | ||||
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Tsai, I.-J.; Su, E.C.-Y.; Tsai, I.-L.; Lin, C.-Y. Clinical Assay for the Early Detection of Colorectal Cancer Using Mass Spectrometric Wheat Germ Agglutinin Multiple Reaction Monitoring. Cancers 2021, 13, 2190. https://doi.org/10.3390/cancers13092190
Tsai I-J, Su EC-Y, Tsai I-L, Lin C-Y. Clinical Assay for the Early Detection of Colorectal Cancer Using Mass Spectrometric Wheat Germ Agglutinin Multiple Reaction Monitoring. Cancers. 2021; 13(9):2190. https://doi.org/10.3390/cancers13092190
Chicago/Turabian StyleTsai, I-Jung, Emily Chia-Yu Su, I-Lin Tsai, and Ching-Yu Lin. 2021. "Clinical Assay for the Early Detection of Colorectal Cancer Using Mass Spectrometric Wheat Germ Agglutinin Multiple Reaction Monitoring" Cancers 13, no. 9: 2190. https://doi.org/10.3390/cancers13092190
APA StyleTsai, I.-J., Su, E. C.-Y., Tsai, I.-L., & Lin, C.-Y. (2021). Clinical Assay for the Early Detection of Colorectal Cancer Using Mass Spectrometric Wheat Germ Agglutinin Multiple Reaction Monitoring. Cancers, 13(9), 2190. https://doi.org/10.3390/cancers13092190







