Human Endogenous Retrovirus Reactivation: Implications for Cancer Immunotherapy
Simple Summary
Abstract
1. Background
2. Human Endogenous Retroviruses
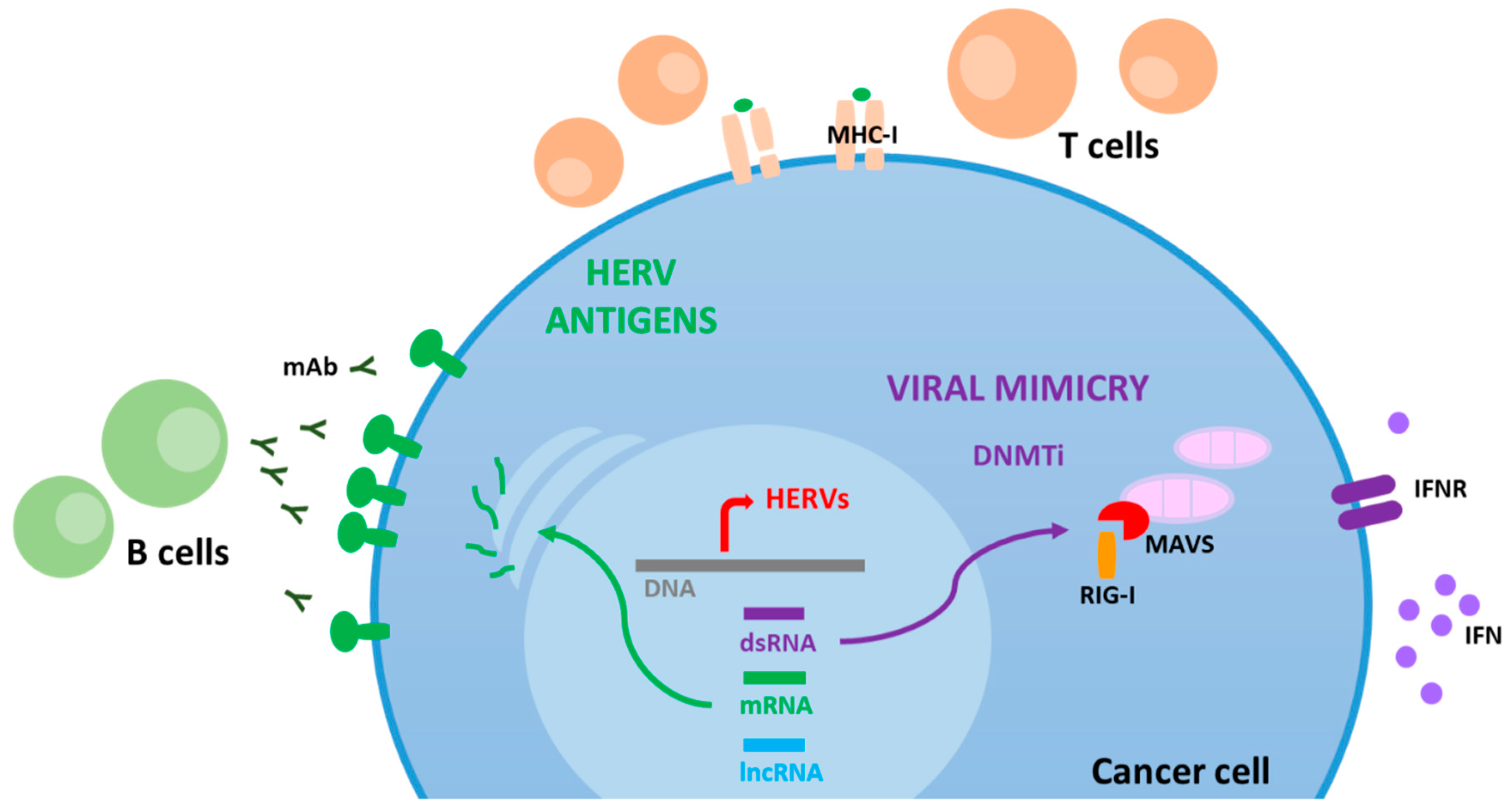
3. Viral Mimicry State
4. HERV Antigens
5. HERVs and Immune Checkpoint Blockade
6. HERVs and Liver Cancer
7. Concluding Remarks
Author Contributions
Funding
Data Availability Statement
Conflicts of Interest
References
- Andersen, K.G.; Rambaut, A.; Lipkin, W.I.; Holmes, E.C.; Garry, R.F. The proximal origin of SARS-CoV-2. Nat. Med. 2020, 26, 450–452. [Google Scholar] [CrossRef] [PubMed]
- Aswad, A.; Katzourakis, A. Paleovirology and virally derived immunity. Trends Ecol. Evol. 2012, 27, 627–636. [Google Scholar] [CrossRef] [PubMed]
- Kanda, R.K.; Coulson, T. The effect of life history on retroviral genome invasions. PLoS ONE 2015, 10, e0117442. [Google Scholar] [CrossRef] [PubMed]
- Nelson, P.N.; Carnegie, P.R.; Martin, J.; Davari, E.H.; Hooley, P.; Roden, D.; Rowland-Jones, S.; Warren, P.; Astley, J.; Murray, P.G. Demystified. Human endogenous retroviruses. Mol. Pathol. 2003, 56, 11–18. [Google Scholar] [CrossRef] [PubMed]
- Lavialle, C.; Cornelis, G.; Dupressoir, A.; Esnault, C.; Heidmann, O.; Vernochet, C.; Heidmann, T. Paleovirology of ‘syncytins’, retroviral env genes exapted for a role in placentation. Philos. Trans. R. Soc. Lond. B Biol. Sci. 2013, 368, 20120507. [Google Scholar] [CrossRef]
- Patel, M.R.; Emerman, M.; Malik, H.S. Paleovirology—Ghosts and gifts of viruses past. Curr. Opin. Virol. 2011, 1, 304–309. [Google Scholar] [CrossRef] [PubMed]
- Katzourakis, A. Paleovirology: Inferring viral evolution from host genome sequence data. Philos. Trans. R. Soc. Lond. B Biol. Sci. 2013, 368, 20120493. [Google Scholar] [CrossRef] [PubMed]
- Griffiths, D.J. Endogenous retroviruses in the human genome sequence. Genome Biol. 2001, 2, REVIEWS1017-2. [Google Scholar] [CrossRef] [PubMed]
- Gifford, R.J.; Blomberg, J.; Coffin, J.M.; Fan, H.; Heidmann, T.; Mayer, J.; Stoye, J.; Tristem, M.; Johnson, W.E. Nomenclature for endogenous retrovirus (ERV) loci. Retrovirology 2018, 15, 59–0442. [Google Scholar] [CrossRef]
- Blomberg, J.; Benachenhou, F.; Blikstad, V.; Sperber, G.; Mayer, J. Classification and nomenclature of endogenous retroviral sequences (ERVs): Problems and recommendations. Gene 2009, 448, 115–123. [Google Scholar] [CrossRef]
- Garcia-Montojo, M.; Doucet-O’Hare, T.; Henderson, L.; Nath, A. Human endogenous retrovirus-K (HML-2): A comprehensive review. Crit. Rev. Microbiol. 2018, 44, 715–738. [Google Scholar] [CrossRef] [PubMed]
- Gonzalez-Cao, M.; Iduma, P.; Karachaliou, N.; Santarpia, M.; Blanco, J.; Rosell, R. Human endogenous retroviruses and cancer. Cancer Biol. Med. 2016, 13, 483–488. [Google Scholar] [PubMed]
- Grandi, N.; Tramontano, E. HERV Envelope Proteins: Physiological Role and Pathogenic Potential in Cancer and Autoimmunity. Front. Microbiol. 2018, 9, 462. [Google Scholar] [CrossRef] [PubMed]
- Bannert, N.; Hofmann, H.; Block, A.; Hohn, O. HERVs New Role in Cancer: From Accused Perpetrators to Cheerful Protectors. Front. Microbiol. 2018, 9, 178. [Google Scholar] [CrossRef]
- Lemaitre, C.; Tsang, J.; Bireau, C.; Heidmann, T.; Dewannieux, M. A human endogenous retrovirus-derived gene that can contribute to oncogenesis by activating the ERK pathway and inducing migration and invasion. PLoS Pathog. 2017, 13, e1006451. [Google Scholar] [CrossRef] [PubMed]
- Singh, M.; Cai, H.; Bunse, M.; Feschotte, C.; Izsvak, Z. Human Endogenous Retrovirus K Rec forms a Regulatory Loop with MITF that Opposes the Progression of Melanoma to an Invasive Stage. Viruses 2020, 12, 1303. [Google Scholar] [CrossRef] [PubMed]
- Hurst, T.P.; Magiorkinis, G. Epigenetic Control of Human Endogenous Retrovirus Expression: Focus on Regulation of Long-Terminal Repeats (LTRs). Viruses 2017, 9, 130. [Google Scholar] [CrossRef] [PubMed]
- Benesova, M.; Trejbalova, K.; Kovarova, D.; Vernerova, Z.; Hron, T.; Kucerova, D.; Hejnar, J. DNA hypomethylation and aberrant expression of the human endogenous retrovirus ERVWE1/syncytin-1 in seminomas. Retrovirology 2017, 14, 20–0342. [Google Scholar] [CrossRef] [PubMed]
- Ohtani, H.; Liu, M.; Zhou, W.; Liang, G.; Jones, P.A. Switching roles for DNA and histone methylation depend on evolutionary ages of human endogenous retroviruses. Genome Res. 2018, 28, 1147–1157. [Google Scholar] [CrossRef]
- Liu, N.; Lee, C.H.; Swigut, T.; Grow, E.; Gu, B.; Bassik, M.C.; Wysocka, J. Selective silencing of euchromatic L1s revealed by genome-wide screens for L1 regulators. Nature 2018, 553, 228–232. [Google Scholar] [CrossRef]
- Robbez-Masson, L.; Tie, C.H.C.; Conde, L.; Tunbak, H.; Husovsky, C.; Tchasovnikarova, I.A.; Timms, R.T.; Herrero, J.; Lehner, P.J.; Rowe, H.M. The HUSH complex cooperates with TRIM28 to repress young retrotransposons and new genes. Genome Res. 2018, 28, 836–845. [Google Scholar] [CrossRef]
- Liu, M.; Thomas, S.L.; DeWitt, A.K.; Zhou, W.; Madaj, Z.B.; Ohtani, H.; Baylin, S.B.; Liang, G.; Jones, P.A. Dual Inhibition of DNA and Histone Methyltransferases Increases Viral Mimicry in Ovarian Cancer Cells. Cancer Res. 2018, 78, 5754–5766. [Google Scholar] [CrossRef] [PubMed]
- Jones, P.A.; Ohtani, H.; Chakravarthy, A.; de Carvalho, D.D. Epigenetic therapy in immune-oncology. Nat. Rev. Cancer 2019, 19, 151–161. [Google Scholar] [CrossRef] [PubMed]
- Saito, Y.; Nakaoka, T.; Saito, H. A New Molecular Mechanism Underlying the Antitumor Effect of DNA Methylation Inhibitors via an Antiviral Immune Response. Adv. Protein Chem. Struct. Biol. 2017, 106, 227–242. [Google Scholar] [CrossRef] [PubMed]
- Attermann, A.S.; Bjerregaard, A.M.; Saini, S.K.; Gronbaek, K.; Hadrup, S.R. Human endogenous retroviruses and their implication for immunotherapeutics of cancer. Ann. Oncol. 2018, 29, 2183–2191. [Google Scholar] [CrossRef] [PubMed]
- Smith, C.C.; Selitsky, S.R.; Chai, S.; Armistead, P.M.; Vincent, B.G.; Serody, J.S. Alternative tumour-specific antigens. Nat. Rev. Cancer 2019, 19, 465–478. [Google Scholar] [CrossRef] [PubMed]
- Hashimoto, K.; Suzuki, A.M.; Dos, S.A.; Desterke, C.; Collino, A.; Ghisletti, S.; Braun, E.; Bonetti, A.; Fort, A.; Qin, X.Y.; et al. CAGE profiling of ncRNAs in hepatocellular carcinoma reveals widespread activation of retroviral LTR promoters in virus-induced tumors. Genome Res. 2015, 25, 1812–1824. [Google Scholar] [CrossRef] [PubMed]
- Yu, P.; Lubben, W.; Slomka, H.; Gebler, J.; Konert, M.; Cai, C.; Neubrandt, L.; da, C.O.P.; Paul, S.; Dehnert, S.; et al. Nucleic acid-sensing Toll-like receptors are essential for the control of endogenous retrovirus viremia and ERV-induced tumors. Immunity 2012, 37, 867–879. [Google Scholar] [CrossRef] [PubMed]
- Roulois, D.; Loo, Y.H.; Singhania, R.; Wang, Y.; Danesh, A.; Shen, S.Y.; Han, H.; Liang, G.; Jones, P.A.; Pugh, T.J.; et al. DNA-Demethylating Agents Target Colorectal Cancer Cells by Inducing Viral Mimicry by Endogenous Transcripts. Cell 2015, 162, 961–973. [Google Scholar] [CrossRef] [PubMed]
- Buonaguro, L.; Petrizzo, A.; Tornesello, M.L.; Buonaguro, F.M. Innate immunity and hepatitis C virus infection: A microarray’s view. Infect. Agent. Cancer 2012, 7, 7. [Google Scholar] [CrossRef]
- Gonzalez-Cao, M.; Karachaliou, N.; Santarpia, M.; Viteri, S.; Meyerhans, A.; Rosell, R. Activation of viral defense signaling in cancer. Ther. Adv. Med. Oncol. 2018, 10. [Google Scholar] [CrossRef] [PubMed]
- Alcazer, V.; Bonaventura, P.; Depil, S. Human Endogenous Retroviruses (HERVs): Shaping the Innate Immune Response in Cancers. Cancers 2020, 12, 610. [Google Scholar] [CrossRef] [PubMed]
- Canadas, I.; Thummalapalli, R.; Kim, J.W.; Kitajima, S.; Jenkins, R.W.; Christensen, C.L.; Campisi, M.; Kuang, Y.; Zhang, Y.; Gjini, E.; et al. Tumor innate immunity primed by specific interferon-stimulated endogenous retroviruses. Nat. Med. 2018, 24, 1143–1150. [Google Scholar] [CrossRef] [PubMed]
- Wu, Y.; Zhao, Y.; Huan, L.; Zhao, J.; Zhou, Y.; Xu, L.; Hu, Z.; Liu, Y.; Chen, Z.; Wang, L.; et al. An LTR Retrotransposon-Derived Long Noncoding RNA lncMER52A Promotes Hepatocellular Carcinoma Progression by Binding p120-Catenin. Cancer Res. 2020, 80, 976–987. [Google Scholar] [CrossRef] [PubMed]
- Dembny, P.; Newman, A.G.; Singh, M.; Hinz, M.; Szczepek, M.; Kruger, C.; Adalbert, R.; Dzaye, O.; Trimbuch, T.; Wallach, T.; et al. Human endogenous retrovirus HERV-K(HML-2) RNA causes neurodegeneration through Toll-like receptors. JCI Insight 2020, 5, e131093. [Google Scholar] [CrossRef]
- Strick, R.; Strissel, P.L.; Baylin, S.B.; Chiappinelli, K.B. Unraveling the molecular pathways of DNA-methylation inhibitors: Human endogenous retroviruses induce the innate immune response in tumors. Oncoimmunology 2015, 5, e1122160. [Google Scholar] [CrossRef]
- Ishak, C.A.; Classon, M.; de Carvalho, D.D. Deregulation of Retroelements as an Emerging Therapeutic Opportunity in Cancer. Trends Cancer 2018, 4, 583–597. [Google Scholar] [CrossRef]
- Diaz-Carballo, D.; Saka, S.; Acikelli, A.H.; Homp, E.; Erwes, J.; Demmig, R.; Klein, J.; Schroer, K.; Malak, S.; D’Souza, F.; et al. Enhanced antitumoral activity of TLR7 agonists via activation of human endogenous retroviruses by HDAC inhibitors. Commun. Biol. 2021, 4, 276–01800. [Google Scholar] [CrossRef]
- Chiappinelli, K.B.; Strissel, P.L.; Desrichard, A.; Li, H.; Henke, C.; Akman, B.; Hein, A.; Rote, N.S.; Cope, L.M.; Snyder, A.; et al. Inhibiting DNA Methylation Causes an Interferon Response in Cancer via dsRNA Including Endogenous Retroviruses. Cell 2015, 162, 974–986. [Google Scholar] [CrossRef]
- Smiraglia, D.J.; Szymanska, J.; Kraggerud, S.M.; Lothe, R.A.; Peltomaki, P.; Plass, C. Distinct epigenetic phenotypes in seminomatous and nonseminomatous testicular germ cell tumors. Oncogene 2002, 21, 3909–3916. [Google Scholar] [CrossRef]
- Peltomaki, P. DNA methylation changes in human testicular cancer. Biochim. Biophys. Acta 1991, 1096, 187–196. [Google Scholar] [CrossRef]
- Benesova, M.; Trejbalova, K.; Kucerova, D.; Vernerova, Z.; Hron, T.; Szabo, A.; Amouroux, R.; Klezl, P.; Hajkova, P.; Hejnar, J. Overexpression of TET dioxygenases in seminomas associates with low levels of DNA methylation and hydroxymethylation. Mol. Carcinog. 2017, 56, 1837–1850. [Google Scholar] [CrossRef] [PubMed]
- Haffner, M.C.; Taheri, D.; Luidy-Imada, E.; Palsgrove, D.N.; Eich, M.L.; Netto, G.J.; Matoso, A.; Nirschl, T.R.; Zheng, Q.; Hicks, J.L.; et al. Hypomethylation, endogenous retrovirus expression, and interferon signaling in testicular germ cell tumors. Proc. Natl. Acad. Sci. USA 2018, 115, E8580–E8582. [Google Scholar] [CrossRef] [PubMed]
- Tokuyama, M.; Kong, Y.; Song, E.; Jayewickreme, T.; Kang, I.; Iwasaki, A. ERVmap analysis reveals genome-wide transcription of human endogenous retroviruses. Proc. Natl. Acad. Sci. USA 2018, 115, 12565–12572. [Google Scholar] [CrossRef] [PubMed]
- Li, M.; Radvanyi, L.; Yin, B.; Rycaj, K.; Li, J.; Chivukula, R.; Lin, K.; Lu, Y.; Shen, J.; Chang, D.Z.; et al. Downregulation of Human Endogenous Retrovirus Type K (HERV-K) Viral env RNA in Pancreatic Cancer Cells Decreases Cell Proliferation and Tumor Growth. Clin. Cancer Res. 2017, 23, 5892–5911. [Google Scholar] [CrossRef] [PubMed]
- Montesion, M.; Bhardwaj, N.; Williams, Z.H.; Kuperwasser, C.; Coffin, J.M. Mechanisms of HERV-K (HML-2) Transcription during Human Mammary Epithelial Cell Transformation. J. Virol. 2017, 92, e01258-17. [Google Scholar] [CrossRef] [PubMed]
- Wang-Johanning, F.; Radvanyi, L.; Rycaj, K.; Plummer, J.B.; Yan, P.; Sastry, K.J.; Piyathilake, C.J.; Hunt, K.K.; Johanning, G.L. Human endogenous retrovirus K triggers an antigen-specific immune response in breast cancer patients. Cancer Res. 2008, 68, 5869–5877. [Google Scholar] [CrossRef] [PubMed]
- Rycaj, K.; Plummer, J.B.; Yin, B.; Li, M.; Garza, J.; Radvanyi, L.; Ramondetta, L.M.; Lin, K.; Johanning, G.L.; Tang, D.G.; et al. Cytotoxicity of human endogenous retrovirus K-specific T cells toward autologous ovarian cancer cells. Clin. Cancer Res. 2015, 21, 471–483. [Google Scholar] [CrossRef] [PubMed]
- Saini, S.K.; Orskov, A.D.; Bjerregaard, A.M.; Unnikrishnan, A.; Holmberg-Thyden, S.; Borch, A.; Jensen, K.V.; Anande, G.; Bentzen, A.K.; Marquard, A.M.; et al. Human endogenous retroviruses form a reservoir of T cell targets in hematological cancers. Nat. Commun. 2020, 11, 5660–19464. [Google Scholar] [CrossRef] [PubMed]
- Krishnamurthy, J.; Rabinovich, B.A.; Mi, T.; Switzer, K.C.; Olivares, S.; Maiti, S.N.; Plummer, J.B.; Singh, H.; Kumaresan, P.R.; Huls, H.M.; et al. Genetic Engineering of T Cells to Target HERV-K, an Ancient Retrovirus on Melanoma. Clin. Cancer Res. 2015, 21, 3241–3251. [Google Scholar] [CrossRef]
- Panda, A.; de Cubas, A.A.; Stein, M.; Riedlinger, G.; Kra, J.; Mayer, T.; Smith, C.C.; Vincent, B.G.; Serody, J.S.; Beckermann, K.E.; et al. Endogenous retrovirus expression is associated with response to immune checkpoint blockade in clear cell renal cell carcinoma. JCI Insight 2018, 3, e121522. [Google Scholar] [CrossRef] [PubMed]
- Chiappinelli, K.B.; Zahnow, C.A.; Ahuja, N.; Baylin, S.B. Combining Epigenetic and Immunotherapy to Combat Cancer. Cancer Res. 2016, 76, 1683–1689. [Google Scholar] [CrossRef] [PubMed]
- Sung, H.; Ferlay, J.; Siegel, R.L.; Laversanne, M.; Soerjomataram, I.; Jemal, A.; Bray, F. Global cancer statistics 2020: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA Cancer J. Clin. 2021, in press. [Google Scholar] [CrossRef] [PubMed]
- Grabski, D.F.; Ratan, A.; Gray, L.R.; Bekiranov, S.; Rekosh, D.; Hammarskjold, M.L.; Rasmussen, S.K. Upregulation of human endogenous retrovirus-K (HML-2) mRNAs in hepatoblastoma: Identification of potential new immunotherapeutic targets and biomarkers. J. Pediatr. Surg. 2021, 56, 286–292. [Google Scholar] [CrossRef] [PubMed]
- Ma, W.; Hong, Z.; Liu, H.; Chen, X.; Ding, L.; Liu, Z.; Zhou, F.; Yuan, Y. Human Endogenous Retroviruses-K (HML-2) Expression Is Correlated with Prognosis and Progress of Hepatocellular Carcinoma. Biomed. Res. Int. 2016, 2016, 8201642. [Google Scholar] [CrossRef] [PubMed]
| Class | Family | Subgroups |
|---|---|---|
| Class I | HERV-H, HERV-F, HERV-W, HERV-R, HERV-P, HERV-E, HERV-I, HERV-T, ERV-FTD, ERV-FRD | |
| Class II | HERV-K | HML 1-11 |
| Class III | HERV-L |
| Cancer | HERV | Study Setting | Study Main Findings | Reference |
|---|---|---|---|---|
| Colorectal cancer | HERV | Preclinical | DNA-demethylating agents act by inducing endogenous dsRNAs that activate an interferon response pathway. This anti-viral response reduces proliferation of colorectal cancer-initiating cells. | [26] |
| Ovarian cancer | HERV families | Preclinical/Clinical | DNA methyltransferase inhibitors upregulate endogenous retroviruses in tumor cells to induce a growth-inhibiting immune response. | [27] |
| Ovarian cancer | HERV families | Preclinical | Dual inhibition of DNA and histone methyltransferases in ovarian cancer cell lines induces synergistic anti-tumor effects by upregulation of endogenous retroviruses, and activation of the viral defense response. | [15] |
| Breast cancer | HERV-K | Clinincal | HERV-K env protein products are able of acting as tumor associated antigens, activating both T cell and B cell responses in breast cancer patients. | [28] |
| Ovarian cancer | HERV-K | Clinincal | Ovarian cancer cells in primary tumors express HERV transcripts, including HERV-K env protein. Ovarian cancer patient sera contain HERV-K immunoreactive antibodies. | [29] |
| Melanoma | HERV-K | Preclinical/Clinical | HERV-K env protein is expressed on melanoma but not in normal tissues. | [30] |
| Renal cell carcinoma | HERV families | Clinincal | Abnormal expression of ERVs is associated with ccRCC, and ERV3-2 expression is associated with response to ICB in ccRCC. | [31] |
| Hepatoblastoma | HERV-K (HML-2) | Clinical | HERV-K is expressed from multiple loci in hepatoblastoma. Expression is increased for several proviruses compared to normal liver controls. | [32] |
| Hepatocellular carcinoma | HERV-K (HML-2) | Clinical | Upregulation of HERV-K (HML-2) in HCC patients is significantly correlated to cancer progression and poor outcome. | [33] |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Petrizzo, A.; Ragone, C.; Cavalluzzo, B.; Mauriello, A.; Manolio, C.; Tagliamonte, M.; Buonaguro, L. Human Endogenous Retrovirus Reactivation: Implications for Cancer Immunotherapy. Cancers 2021, 13, 1999. https://doi.org/10.3390/cancers13091999
Petrizzo A, Ragone C, Cavalluzzo B, Mauriello A, Manolio C, Tagliamonte M, Buonaguro L. Human Endogenous Retrovirus Reactivation: Implications for Cancer Immunotherapy. Cancers. 2021; 13(9):1999. https://doi.org/10.3390/cancers13091999
Chicago/Turabian StylePetrizzo, Annacarmen, Concetta Ragone, Beatrice Cavalluzzo, Angela Mauriello, Carmen Manolio, Maria Tagliamonte, and Luigi Buonaguro. 2021. "Human Endogenous Retrovirus Reactivation: Implications for Cancer Immunotherapy" Cancers 13, no. 9: 1999. https://doi.org/10.3390/cancers13091999
APA StylePetrizzo, A., Ragone, C., Cavalluzzo, B., Mauriello, A., Manolio, C., Tagliamonte, M., & Buonaguro, L. (2021). Human Endogenous Retrovirus Reactivation: Implications for Cancer Immunotherapy. Cancers, 13(9), 1999. https://doi.org/10.3390/cancers13091999






