The TCGA Molecular Classification of Endometrial Cancer and Its Possible Impact on Adjuvant Treatment Decisions
Abstract
Simple Summary
Abstract
1. Introduction
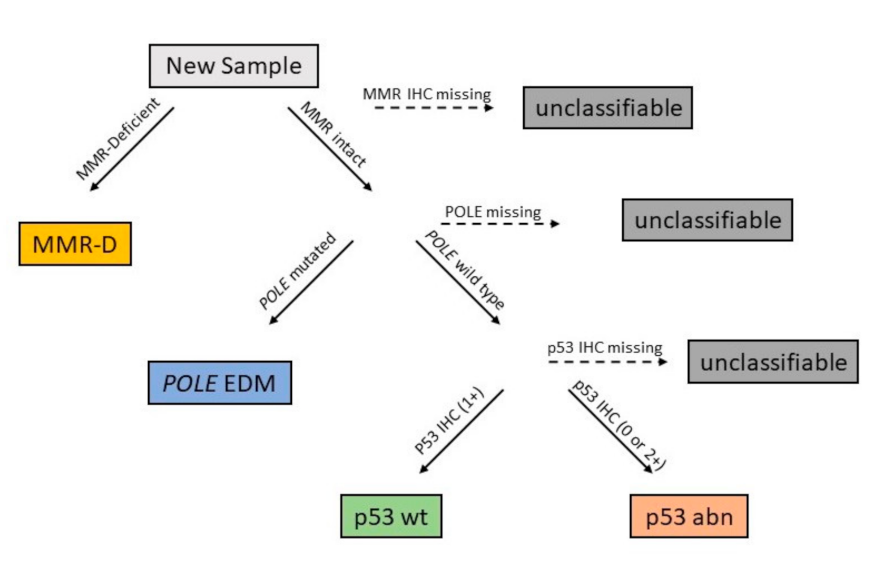
2. Development of a New Molecular Classification
3. Main Features of the Molecular Subtypes
3.1. Polymerase Epsilon (POLE)
3.2. Mismatch Repair Deficiency (MMRd)
3.3. p53 Abn
3.4. p53 wt
4. Published Trials
5. Ongoing Trial and Conclusions
Funding
Conflicts of Interest
References
- Bray, F.; Ferlay, J.; Soerjomataram, I.; Siegel, R.L.; Torre, L.A.; Jemal, A. Global Cancer Statistics 2018: GLOBOCAN Estimates of Incidence and Mortality Worldwide for 36 Cancers in 185 Countries. CA A Cancer J. Clin. 2018, 68, 394–424. [Google Scholar] [CrossRef] [PubMed]
- NIH National Cancer Institute Surveillance Epidemiology, and End Results Program. Cancer Stat Facts: Endometrial Cancer. 2020. Available online: https://seer.cancer.gov/statfacts/html/corp.html (accessed on 4 July 2020).
- Colombo, N.; Preti, E.; Landoni, F.; Carinelli, S.; Colombo, A.; Marini, C.; Sessa, C. Endometrial Cancer: ESMO Clinical Practice Guidelines for Diagnosis, Treatment and Follow-Up. Ann. Oncol. 2013, 24, vi33–vi38. [Google Scholar] [CrossRef] [PubMed]
- Bokhman, J.V. Two Pathogenetic Types of Endometrial Carcinoma. Gynecol. Oncol. 1983, 15, 10–17. [Google Scholar] [CrossRef]
- Setiawan, V.W.; Yang, H.P.; Pike, M.C.; McCann, S.E.; Yu, H.; Xiang, Y.-B.; Wolk, A.; Wentzensen, N.; Weiss, N.S.; Webb, P.M.; et al. Type I and II Endometrial Cancers: Have They Different Risk Factors? J. Clin. Oncol. 2013, 31, 2607–2618. [Google Scholar] [CrossRef]
- Murali, R.; Soslow, R.A.; Weigelt, B. Classification of Endometrial Carcinoma: More than Two Types. Lancet Oncol. 2014, 15, e268–e278. [Google Scholar] [CrossRef]
- World Health Organization. Female Genital Tumours; World Health Organization: Geneva, Switzerland, 2020; ISBN 978-92-832-4504-9. [Google Scholar]
- Gilks, C.B.; Oliva, E.; Soslow, R.A. Poor Interobserver Reproducibility in the Diagnosis of High-Grade Endometrial Carcinoma. Am. J. Surg. Pathol. 2013, 37, 874–881. [Google Scholar] [CrossRef]
- Han, G.; Sidhu, D.; Duggan, M.A.; Arseneau, J.; Cesari, M.; Clement, P.B.; Ewanowich, C.A.; Kalloger, S.E.; Köbel, M. Reproducibility of Histological Cell Type in High-Grade Endometrial Carcinoma. Mod. Pathol. 2013, 26, 1594–1604. [Google Scholar] [CrossRef]
- Colombo, N.; Creutzberg, C.; Amant, F.; Bosse, T.; González-Martín, A.; Ledermann, J.; Marth, C.; Nout, R.; Querleu, D.; Mirza, M.R.; et al. ESMO-ESGO-ESTRO Consensus Conference on Endometrial Cancer. Int. J. Gynecol. Cancer 2016, 26, 2–30. [Google Scholar] [CrossRef]
- Concin, N.; Matias-Guiu, X.; Vergote, I.; Cibula, D.; Mirza, M.R.; Marnitz, S.; Ledermann, J.; Bosse, T.; Chargari, C.; Fagotti, A.; et al. ESGO/ESTRO/ESP Guidelines for the Management of Patients with Endometrial Carcinoma. Int. J. Gynecol. Cancer 2021, 31, 12–39. [Google Scholar] [CrossRef] [PubMed]
- Kommoss, F.; Kommoss, F.; Grevenkamp, F.; Bunz, A.-K.; Taran, F.-A.; Fend, F.; Brucker, S.Y.; Wallwiener, D.; Schönfisch, B.; Greif, K.; et al. L1CAM: Amending the “Low-Risk” Category in Endometrial Carcinoma. J. Cancer Res. Clin. Oncol. 2017, 143, 255–262. [Google Scholar] [CrossRef] [PubMed]
- Fogel, M.; Gutwein, P.; Mechtersheimer, S.; Riedle, S.; Stoeck, A.; Smirnov, A.; Edler, L.; Ben-Arie, A.; Huszar, M.; Altevogt, P. L1 Expression as a Predictor of Progression and Survival in Patients with Uterine and Ovarian Carcinomas. Lancet 2003, 362, 869–875. [Google Scholar] [CrossRef]
- Zeimet, A.G.; Reimer, D.; Huszar, M.; Winterhoff, B.; Puistola, U.; Abdel Azim, S.; Müller-Holzner, E.; Ben-Arie, A.; van Kempen, L.C.; Petru, E.; et al. L1CAM in Early-Stage Type I Endometrial Cancer: Results of a Large Multicenter Evaluation. J. Natl. Cancer Inst. 2013, 105, 1142–1150. [Google Scholar] [CrossRef] [PubMed]
- Bosse, T.; Nout, R.A.; Stelloo, E.; Dreef, E.; Nijman, H.W.; Jürgenliemk-Schulz, I.M.; Jobsen, J.J.; Creutzberg, C.L.; Smit, V.T.H.B.M. L1 Cell Adhesion Molecule Is a Strong Predictor for Distant Recurrence and Overall Survival in Early Stage Endometrial Cancer: Pooled PORTEC Trial Results. Eur. J. Cancer 2014, 50, 2602–2610. [Google Scholar] [CrossRef]
- Briët, J.M.; Hollema, H.; Reesink, N.; Aalders, J.G.; Mourits, M.J.E.; ten Hoor, K.A.; Pras, E.; Boezen, H.M.; van der Zee, A.G.J.; Nijman, H.W. Lymphvascular Space Involvement: An Independent Prognostic Factor in Endometrial Cancer. Gynecol. Oncol. 2005, 96, 799–804. [Google Scholar] [CrossRef] [PubMed]
- Gemer, O.; Ben Arie, A.; Levy, T.; Gdalevich, M.; Lorian, M.; Barak, F.; Anteby, E.; Lavie, O. Lymphvascular Space Involvement Compromises the Survival of Patients with Stage I Endometrial Cancer: Results of a Multicenter Study. Eur. J. Surg. Oncol. 2007, 33, 644–647. [Google Scholar] [CrossRef] [PubMed]
- Bosse, T.; Peters, E.E.M.; Creutzberg, C.L.; Jürgenliemk-Schulz, I.M.; Jobsen, J.J.; Mens, J.W.M.; Lutgens, L.C.H.W.; van der Steen-Banasik, E.M.; Smit, V.T.H.B.M.; Nout, R.A. Substantial Lymph-Vascular Space Invasion (LVSI) Is a Significant Risk Factor for Recurrence in Endometrial Cancer—A Pooled Analysis of PORTEC 1 and 2 Trials. Eur. J. Cancer 2015, 51, 1742–1750. [Google Scholar] [CrossRef]
- Kim, S.; Jeong, S. Mutation Hotspots in the β-Catenin Gene: Lessons from the Human Cancer Genome Databases. Mol. Cells 2019, 42, 8–16. [Google Scholar] [CrossRef]
- Imboden, S.; Tapia, C.; Scheiwiller, N.; Kocbek, V.; Altermatt, H.J.; Janzen, J.; Mueller, M.D.; McKinnon, B. Early-Stage Endometrial Cancer, CTNNB1 Mutations, and the Relation between Lymphovascular Space Invasion and Recurrence. Acta Obstet. Et Gynecol. Scand. 2020, 99, 196–203. [Google Scholar] [CrossRef]
- Kurnit, K.C.; Kim, G.; Fellman, B.M.; Urbauer, D.; Mills, G.B.; Zhang, W.; Broaddus, R.R. CTNNB1 (Beta-Catenin) Mutation Identifies Low Grade, Early Stage Endometrial Cancer Patients at Increased Risk of Recurrence. Mod. Pathol. 2017, 30, 1032–1041. [Google Scholar] [CrossRef]
- Liu, Y.; Patel, L.; Mills, G.B.; Lu, K.H.; Sood, A.K.; Ding, L.; Kucherlapati, R.; Mardis, E.R.; Levine, D.A.; Shmulevich, I.; et al. Clinical Significance of CTNNB1 Mutation and Wnt Pathway Activation in Endometrioid Endometrial Carcinoma. J. Natl. Cancer Inst. 2014, 106. [Google Scholar] [CrossRef]
- Meissner, W.A.; Sommers, S.C.; Sherman, G. Endometrial Hyperplasia, Endometrial Carcinoma, and Endometriosis Produced Experimentally by Estrogen. Cancer 1957, 10, 500–509. [Google Scholar] [CrossRef]
- Fu, Y.S.; Gambone, J.C.; Berek, J.S. Pathophysiology and Management of Endometrial Hyperplasia and Carcinoma. West. J. Med. 1990, 153, 50–61. [Google Scholar]
- Stelloo, E.; Nout, R.A.; Osse, E.M.; Jürgenliemk-Schulz, I.J.; Jobsen, J.J.; Lutgens, L.C.; van der Steen-Banasik, E.M.; Nijman, H.W.; Putter, H.; Bosse, T.; et al. Improved Risk Assessment by Integrating Molecular and Clinicopathological Factors in Early-Stage Endometrial Cancer-Combined Analysis of the PORTEC Cohorts. Clin. Cancer Res. 2016, 22, 4215–4224. [Google Scholar] [CrossRef] [PubMed]
- Blanco-Aparicio, C.; Renner, O.; Leal, J.F.M.; Carnero, A. PTEN, More than the AKT Pathway. Carcinogenesis 2007, 28, 1379–1386. [Google Scholar] [CrossRef] [PubMed]
- Nero, C.; Ciccarone, F.; Pietragalla, A.; Scambia, G. PTEN and Gynecological Cancers. Cancers 2019, 11, 1458. [Google Scholar] [CrossRef] [PubMed]
- Raffone, A.; Travaglino, A.; Saccone, G.; Viggiani, M.; Giampaolino, P.; Insabato, L.; Mollo, A.; De Placido, G.; Zullo, F. PTEN Expression in Endometrial Hyperplasia and Risk of Cancer: A Systematic Review and Meta-Analysis. Arch. Gynecol. Obs. 2019, 299, 1511–1524. [Google Scholar] [CrossRef] [PubMed]
- The Cancer Genome Atlas Research Network. Integrated Genomic Characterization of Endometrial Carcinoma. Nature 2013, 497, 67–73. [Google Scholar] [CrossRef] [PubMed]
- Talhouk, A.; McConechy, M.K.; Leung, S.; Li-Chang, H.H.; Kwon, J.S.; Melnyk, N.; Yang, W.; Senz, J.; Boyd, N.; Karnezis, A.N.; et al. A Clinically Applicable Molecular-Based Classification for Endometrial Cancers. Br. J. Cancer 2015, 113, 299–310. [Google Scholar] [CrossRef]
- Talhouk, A.; McConechy, M.K.; Leung, S.; Yang, W.; Lum, A.; Senz, J.; Boyd, N.; Pike, J.; Anglesio, M.; Kwon, J.S.; et al. Confirmation of ProMisE: A Simple, Genomics-Based Clinical Classifier for Endometrial Cancer. Cancer 2017, 123, 802–813. [Google Scholar] [CrossRef] [PubMed]
- Karnezis, A.N.; Leung, S.; Magrill, J.; McConechy, M.K.; Yang, W.; Chow, C.; Kobel, M.; Lee, C.-H.; Huntsman, D.G.; Talhouk, A.; et al. Evaluation of Endometrial Carcinoma Prognostic Immunohistochemistry Markers in the Context of Molecular Classification. J. Pathol. Clin. Res. 2017, 3, 279–293. [Google Scholar] [CrossRef] [PubMed]
- Cosgrove, C.M.; Tritchler, D.L.; Cohn, D.E.; Mutch, D.G.; Rush, C.M.; Lankes, H.A.; Creasman, W.T.; Miller, D.S.; Ramirez, N.C.; Geller, M.A.; et al. An NRG Oncology/GOG Study of Molecular Classification for Risk Prediction in Endometrioid Endometrial Cancer. Gynecol. Oncol. 2018, 148, 174–180. [Google Scholar] [CrossRef]
- Bosse, T.; Nout, R.A.; McAlpine, J.N.; McConechy, M.K.; Britton, H.; Hussein, Y.; Gonzalez, C.; Ganesan, R.; Steele, J.C.; Harrison, B.T.; et al. Molecular Classification of Grade 3 Endometrioid Endometrial Cancers Identifies Distinct Prognostic Subgroups. Am. J. Surg. Pathol. 2018, 42, 561–568. [Google Scholar] [CrossRef]
- McAlpine, J.; Leon-Castillo, A.; Bosse, T. The Rise of a Novel Classification System for Endometrial Carcinoma; Integration of Molecular Subclasses. J. Pathol. 2018, 244, 538–549. [Google Scholar] [CrossRef] [PubMed]
- Rayner, E.; van Gool, I.C.; Palles, C.; Kearsey, S.E.; Bosse, T.; Tomlinson, I.; Church, D.N. A Panoply of Errors: Polymerase Proofreading Domain Mutations in Cancer. Nat. Rev. Cancer 2016, 16, 71–81. [Google Scholar] [CrossRef] [PubMed]
- Mayer, A.; Schmidt, M.; Seeger, A.; Serras, F.A.; Vaupel, P.; Schmidberger, H. Microregional Mismatch of Markers Related to Hypoxia and the Warburg Effect in Squamous Cell Carcinomas of the Vulva. Strahlenther. Onkol. 2014, 190, 82–83. [Google Scholar]
- León-Castillo, A.; Britton, H.; McConechy, M.K.; McAlpine, J.N.; Nout, R.; Kommoss, S.; Brucker, S.Y.; Carlson, J.W.; Epstein, E.; Rau, T.T.; et al. Interpretation of Somatic POLE Mutations in Endometrial Carcinoma. J. Pathol. 2020, 250, 323–335. [Google Scholar] [CrossRef]
- Hause, R.J.; Pritchard, C.C.; Shendure, J.; Salipante, S.J. Classification and Characterization of Microsatellite Instability across 18 Cancer Types. Nat. Med. 2016, 22, 1342–1350. [Google Scholar] [CrossRef]
- Lynch, H.T.; Snyder, C.L.; Shaw, T.G.; Heinen, C.D.; Hitchins, M.P. Milestones of Lynch Syndrome: 1895–2015. Nat. Rev. Cancer 2015, 15, 181–194. [Google Scholar] [CrossRef]
- Biller, L.H.; Syngal, S.; Yurgelun, M.B. Recent Advances in Lynch Syndrome. Fam. Cancer 2019, 18, 211–219. [Google Scholar] [CrossRef]
- Stelloo, E.; Jansen, A.M.L.; Osse, E.M.; Nout, R.A.; Creutzberg, C.L.; Ruano, D.; Church, D.N.; Morreau, H.; Smit, V.T.H.B.M.; van Wezel, T.; et al. Practical Guidance for Mismatch Repair-Deficiency Testing in Endometrial Cancer. Ann. Oncol. 2017, 28, 96–102. [Google Scholar] [CrossRef] [PubMed]
- Kandoth, C.; McLellan, M.D.; Vandin, F.; Ye, K.; Niu, B.; Lu, C.; Xie, M.; Zhang, Q.; McMichael, J.F.; Wyczalkowski, M.A.; et al. Mutational Landscape and Significance across 12 Major Cancer Types. Nature 2013, 502, 333–339. [Google Scholar] [CrossRef]
- Mantovani, F.; Collavin, L.; Del Sal, G. Mutant P53 as a Guardian of the Cancer Cell. Cell Death Differ. 2019, 26, 199–212. [Google Scholar] [CrossRef]
- Edmondson, R.J.; Crosbie, E.J.; Nickkho-Amiry, M.; Kaufmann, A.; Stelloo, E.; Nijman, H.W.; Leary, A.; Auguste, A.; Mileshkin, L.; Pollock, P.; et al. Markers of the P53 Pathway Further Refine Molecular Profiling in High-Risk Endometrial Cancer: A TransPORTEC Initiative. Gynecol. Oncol. 2017, 146, 327–333. [Google Scholar] [CrossRef] [PubMed]
- Kommoss, S.; McConechy, M.K.; Kommoss, F.; Leung, S.; Bunz, A.; Magrill, J.; Britton, H.; Kommoss, F.; Grevenkamp, F.; Karnezis, A.; et al. Final Validation of the ProMisE Molecular Classifier for Endometrial Carcinoma in a Large Population-Based Case Series. Ann. Oncol. 2018, 29, 1180–1188. [Google Scholar] [CrossRef] [PubMed]
- Stelloo, E.; Bosse, T.; Nout, R.A.; MacKay, H.J.; Church, D.N.; Nijman, H.W.; Leary, A.; Edmondson, R.J.; Powell, M.E.; Crosbie, E.J.; et al. Refining Prognosis and Identifying Targetable Pathways for High-Risk Endometrial Cancer; a TransPORTEC Initiative. Mod. Pathol. 2015, 28, 836–844. [Google Scholar] [CrossRef] [PubMed]
- León-Castillo, A.; de Boer, S.M.; Powell, M.E.; Mileshkin, L.R.; Mackay, H.J.; Leary, A.; Nijman, H.W.; Singh, N.; Pollock, P.M.; Bessette, P.; et al. Molecular Classification of the PORTEC-3 Trial for High-Risk Endometrial Cancer: Impact on Prognosis and Benefit From Adjuvant Therapy. JCO 2020, 38, 3388–3397. [Google Scholar] [CrossRef] [PubMed]
- Kommoss, F.K.; Karnezis, A.N.; Kommoss, F.; Talhouk, A.; Taran, F.-A.; Staebler, A.; Gilks, C.B.; Huntsman, D.G.; Krämer, B.; Brucker, S.Y.; et al. L1CAM Further Stratifies Endometrial Carcinoma Patients with No Specific Molecular Risk Profile. Br. J. Cancer 2018, 119, 480–486. [Google Scholar] [CrossRef]
- Raffone, A.; Travaglino, A.; Mascolo, M.; Carbone, L.; Guida, M.; Insabato, L.; Zullo, F. TCGA Molecular Groups of Endometrial Cancer: Pooled Data about Prognosis. Gynecol. Oncol. 2019, 155, 374–383. [Google Scholar] [CrossRef] [PubMed]
- León-Castillo, A.; Gilvazquez, E.; Nout, R.; Smit, V.T.; McAlpine, J.N.; McConechy, M.; Kommoss, S.; Brucker, S.Y.; Carlson, J.W.; Epstein, E.; et al. Clinicopathological and Molecular Characterisation of ‘Multiple-Classifier’ Endometrial Carcinomas. J. Pathol. 2020, 250, 312–322. [Google Scholar] [CrossRef] [PubMed]
- Wortman, B.G.; Bosse, T.; Nout, R.A.; Lutgens, L.C.H.W.; van der Steen-Banasik, E.M.; Westerveld, H.; van den Berg, H.; Slot, A.; De Winter, K.a.J.; Verhoeven-Adema, K.W.; et al. Molecular-Integrated Risk Profile to Determine Adjuvant Radiotherapy in Endometrial Cancer: Evaluation of the Pilot Phase of the PORTEC-4a Trial. Gynecol. Oncol. 2018, 151, 69–75. [Google Scholar] [CrossRef]
| Risk Group | Molecular Classification Unknown | Molecular Classification Known |
|---|---|---|
| Low |
|
|
| Intermediate |
|
|
| High-intermediate |
|
|
| High |
|
|
| Advanced |
|
|
| Metastatic |
|
|
| Subtype (Synonyms) | POLE-Mutant POLE EDM | MMRd MSI | p53 wt, MSS, CN Low NSMP | p53 Abn CN High |
|---|---|---|---|---|
| Mutational frequency | >100 mutations/Mb | 100–10 mutations/Mb | <10 mutations/Mb | <10 mutations/Mb |
| Somatic copy-number alterations | Very low | Low | Low | High |
| Top five recurrent gene mutations (%) | POLE (100%) DMD (100%) CSMD1 (100%) FAT4 (100%) PTEN (94%) | PTEN (88%) PIK3CA (54%) PIK3R1 (42%) RPL22 (37%) ARID1A (37%) | PTEN (77%) PIK3CA (53%) CTNNB1 (52%) ARID1A (42%) PIK3R1 (33%) | TP53 (92%) PIK3CA (47%) FBXW7 (22%) PPP2R1A (22%) PTEN (10%) |
| Associated histological feature | Endometrioid Grade 3 Ambiguous morphology Broad front invasion TILs, peri-tumoral Lymphocytes Giant tumoral cells | Endometrioid Grade 3 LVSI substantial MELF-type invasion TILs, Crohn’s-like peri-tumoral reaction lower uterine segment involvement | Endometrioid Grade 1–2 Squamous differentiation ER/PR expression | Serous Grade 3 LVSI Destructive invasion High cytonuclear atypia Giant tumoral cells Hobnailing, Slit-like spaces |
| Associated clinical features | Lower BMI Early Stage (IA/IB)Early onset | Higher BMI Lynch Syndrome | Higher BMI | Lower BMI Advanced stageLate onset |
| Prognosis in early stage (I–II) | Excellent | Intermediate | Excellent/intermediate/poor | Poor |
| Diagnostic test | Sanger/NGS Tumor mutation burden | MMR-IHC (MLH1, MSH2, MSH6, PMS2) MSI assay Tumor mutation burden | p53-IHC NGS Somatic copy-number aberrations |
| Author | Patient Cohort | Number of Patients | FIGO Stages | Subtypes | HR OS Multivariable | HR RFS Multivariable | |||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| IA | IB | II | III | IV | POLE | MMRd | p53 wt | p53 abn | POLE | MMRd | p53 wt | p35 abn | POLE | MMRd | p53 wt | p53 abn | |||
| Stello 2015 | Portec 3 criteria | 116 | 36.2% | 18.1% | 35.3% | 9.5% | 12.1% | 16.4% | 37.9% | 33.6% | |||||||||
| Talhouk 2015 | “discovery” | 143 | 71.3% | 28.7% | 8.4% | 28.7% | 44.1% | 17.5% | 0.28 (0.00–3.01) | 0.90 (0.31–2.73) | 1.00 | 4.28 (0.95–18.34) | 0.15 (0.00–1.94) | 0.32 (0.10–1.03) | 1.00 | 1.64 (0.32–7.06) | |||
| Stello 2016 | PORTEC 1 & 2 | 834 | n/a | n/a | 5.9% | 26.3% | 59.0% | 8.9% | 1.105 (0.394–3.101) | 1.879 (1.307–2.700) | 1.00 | 3.777 (2.364–6.037) | |||||||
| Talhouk 2017 | “confirmation” | 319 | 69.3% | 29.5% | 9.4% | 20.1% | 27.0% | 43.6% | 1.01 (0.26–2.99) | 1.90 (0.88-4.04) | 1.00 | 2.61 (1.27–5.72) | 0.19 (0.02–0.81) | 0.64 (0.25–1.60) | 1.00 | 1.75 (0.84–3.96) | |||
| Bosse 2018 | Grade 3 EEC | 381 | 44.9% | 31.5% | 30.2% | 13.1% | 2.9% | 12.9% | 36.2% | 30.2% | 20.7% | 0.56 (0.27–1.15) | 0.84 (0.57-1.25) | 1.00 | 1.37(0.9–2.09) | 0.23 (0.07–0.77) | 0.61 (0.37–1.00) | 1.00 | 1.92 (1.20–3.07) |
| Cosgrove 2018 | NRG/GOG GOG210 | 982 | 74,5% | 9.3% | 14.4% | 1,8% | 4.0% | 38.6% | 48.9% | 8.6% | 0.19 (0.03–1.35) | 1.04 (0.70–1.56) | 1.00 | 1.61 (0.93–2.78) | 0.26 (0.06–1.05) | 1.08 (0.78–1.50) | 1.00 | 1.56 (0.99–2.48) | |
| Kommoss 2018 | “validation” | 452 | 61.1% | 19.7% | 5.8% | 12.2% | 1.3% | 9.3% | 28.1% | 50.4% | 12.2% | 0.95 (0.30–2.36 | 1.41 (0.82–2.41) | 1.00 | 2.29 (1.12–4.65) | 0.15 (0.00–n/a) | 1.54 (0.73–3.24) | 1.00 | 3.40 (1.30–8.81) |
| León-Castillo 2020 | PORTEC 3 | 423 | 13.2% | 17.8% | 25.6% | 43.4% | 12.4% | 33.4% | 31.5% | 22.7% | 0.118 (0.016–0.868) | 1.00 | 0.547 (0.302–0.993) | 2.298 (1.418–3.726) | 0.079 (0.011–0.576) | 1.00 | 0.976 (0.620–1.537) | 2.517 (1.621–3.907) | |
| Total | 3650 | 7.9% | 30.9% | 46.5% | 14.7% | ||||||||||||||
| Total | POLE EDM | MMRd | p53 wt | p53 abn | p-Value | ||||||
|---|---|---|---|---|---|---|---|---|---|---|---|
| n | % | n | % | n | % | n | % | n | % | ||
| Adjuvant treatment | |||||||||||
| Any | 1283 | 47.3 | 87 | 51.2 | 385 | 46.5 | 624 | 44.6 | 187 | 58.8 | <0.001 |
| None | 1432 | 52.7 | 83 | 48.8 | 443 | 53.5 | 775 | 55.4 | 131 | 41.2 | |
| Stage | |||||||||||
| I | 1838 | 68.7 | 187 | 84.6 | 581 | 65.7 | 831 | 72.3 | 241 | 56.9 | <0.001 |
| II–IV | 838 | 31.3 | 34 | 15.4 | 302 | 34.3 | 319 | 27.7 | 181 | 43.1 | |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Alexa, M.; Hasenburg, A.; Battista, M.J. The TCGA Molecular Classification of Endometrial Cancer and Its Possible Impact on Adjuvant Treatment Decisions. Cancers 2021, 13, 1478. https://doi.org/10.3390/cancers13061478
Alexa M, Hasenburg A, Battista MJ. The TCGA Molecular Classification of Endometrial Cancer and Its Possible Impact on Adjuvant Treatment Decisions. Cancers. 2021; 13(6):1478. https://doi.org/10.3390/cancers13061478
Chicago/Turabian StyleAlexa, Matthias, Annette Hasenburg, and Marco Johannes Battista. 2021. "The TCGA Molecular Classification of Endometrial Cancer and Its Possible Impact on Adjuvant Treatment Decisions" Cancers 13, no. 6: 1478. https://doi.org/10.3390/cancers13061478
APA StyleAlexa, M., Hasenburg, A., & Battista, M. J. (2021). The TCGA Molecular Classification of Endometrial Cancer and Its Possible Impact on Adjuvant Treatment Decisions. Cancers, 13(6), 1478. https://doi.org/10.3390/cancers13061478





