Radiosensitizing Pancreatic Cancer with PARP Inhibitor and Gemcitabine: An In Vivo and a Whole-Transcriptome Analysis after Proton or Photon Irradiation
Abstract
Simple Summary
Abstract
1. Introduction
2. Results
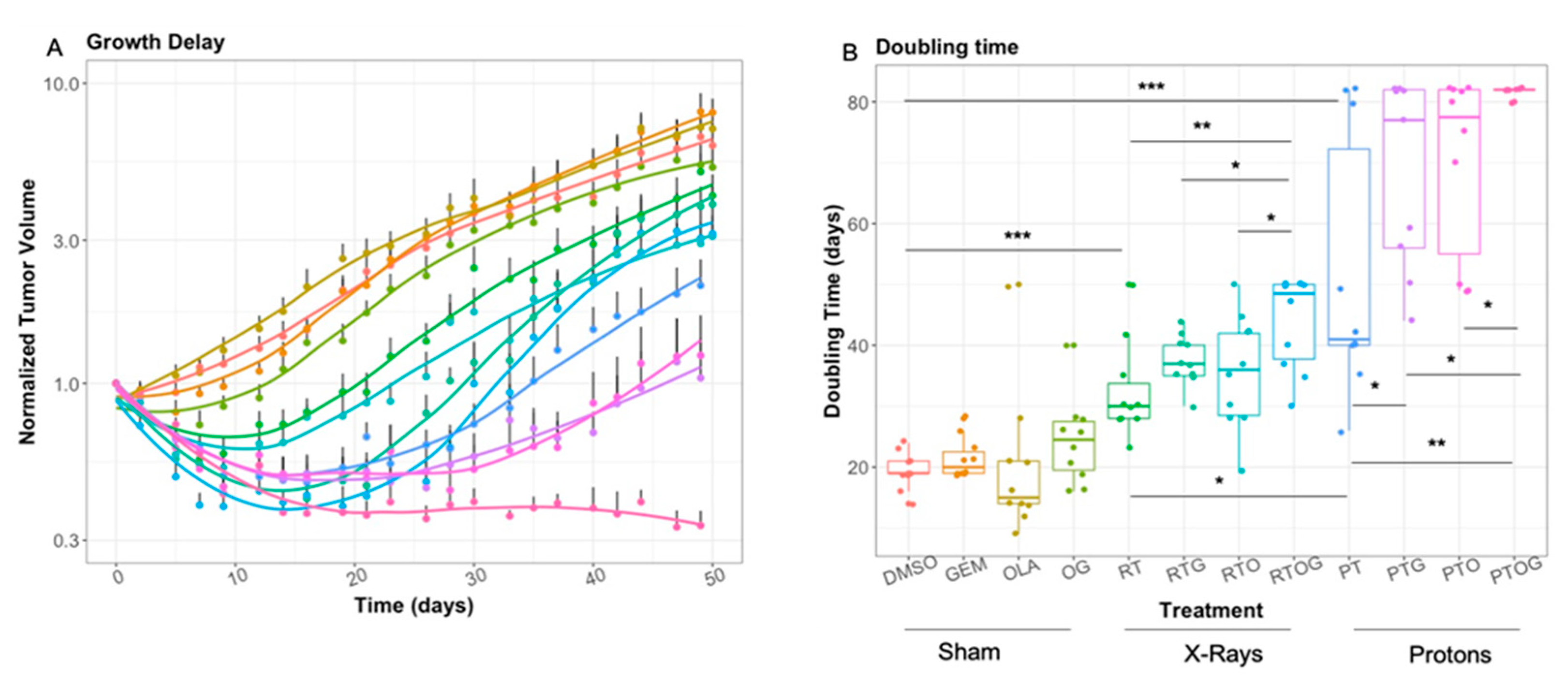
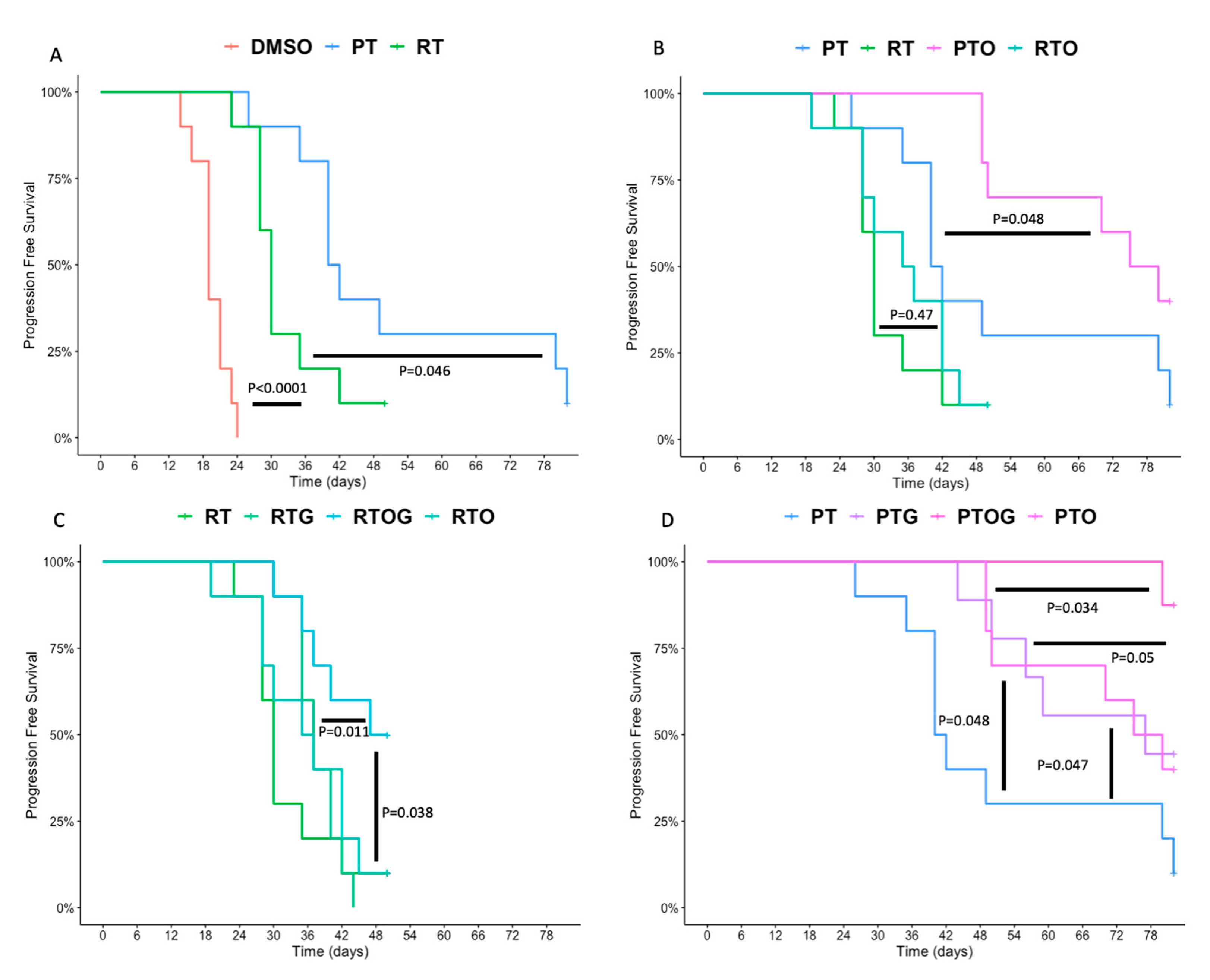
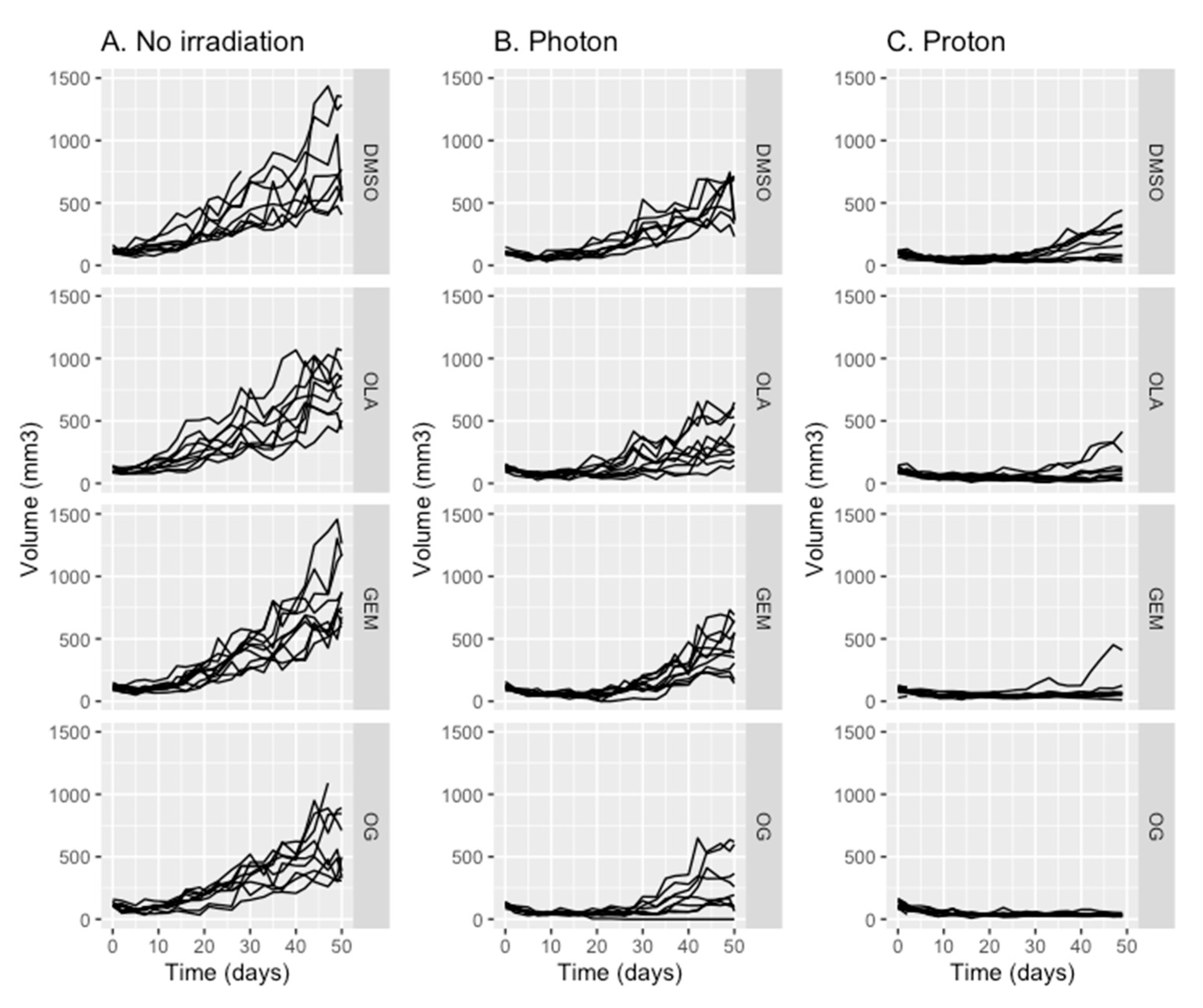
2.1. Olaparib, Gemcitabine and Proton Therapy Significantly Inhibit Tumor Growth In Vivo
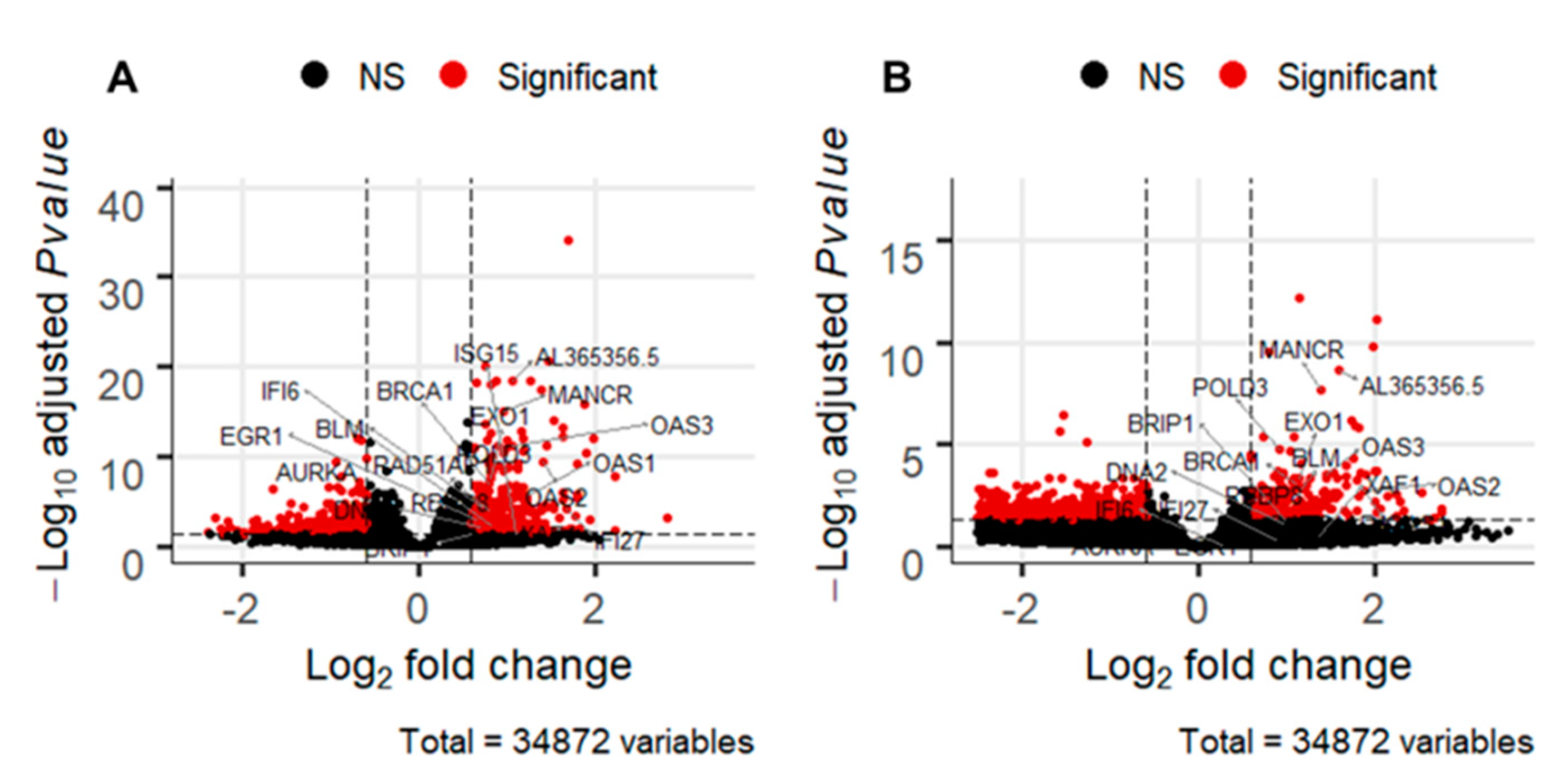
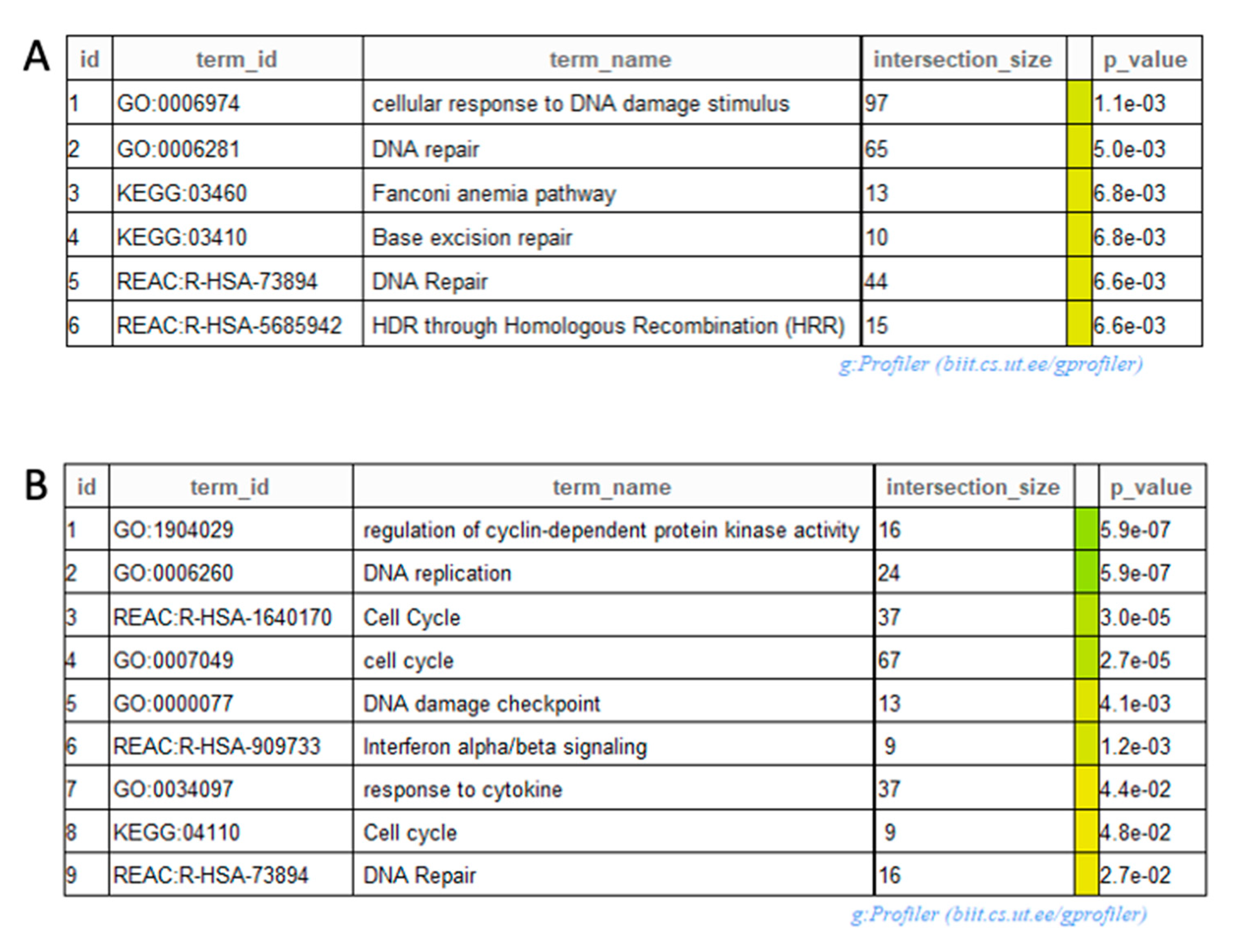
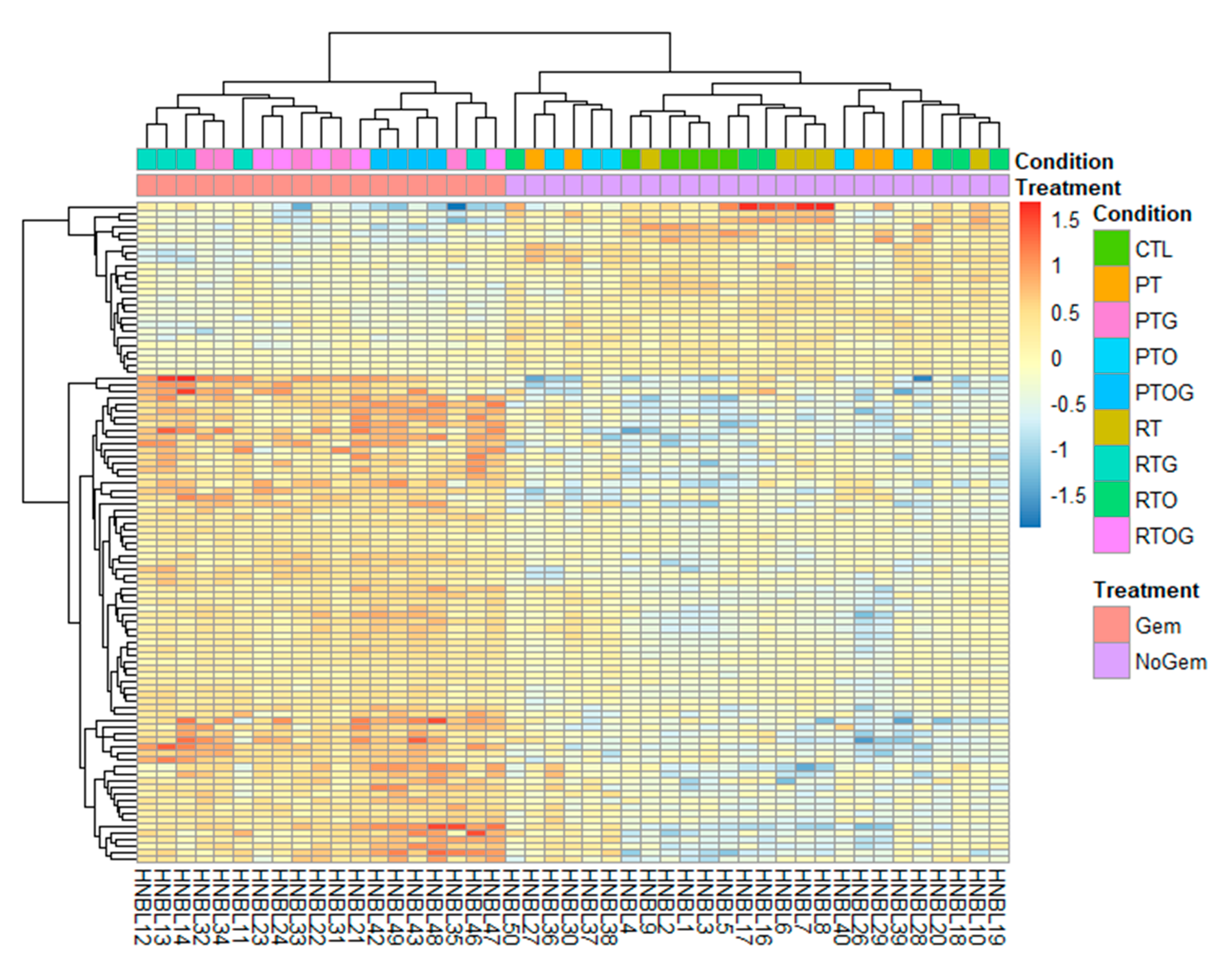
2.2. RNA Sequencing Identifies Pathways Involved in Radiosenstization
3. Discussion
4. Materials and Methods
4.1. Cell Line and Mouse Model
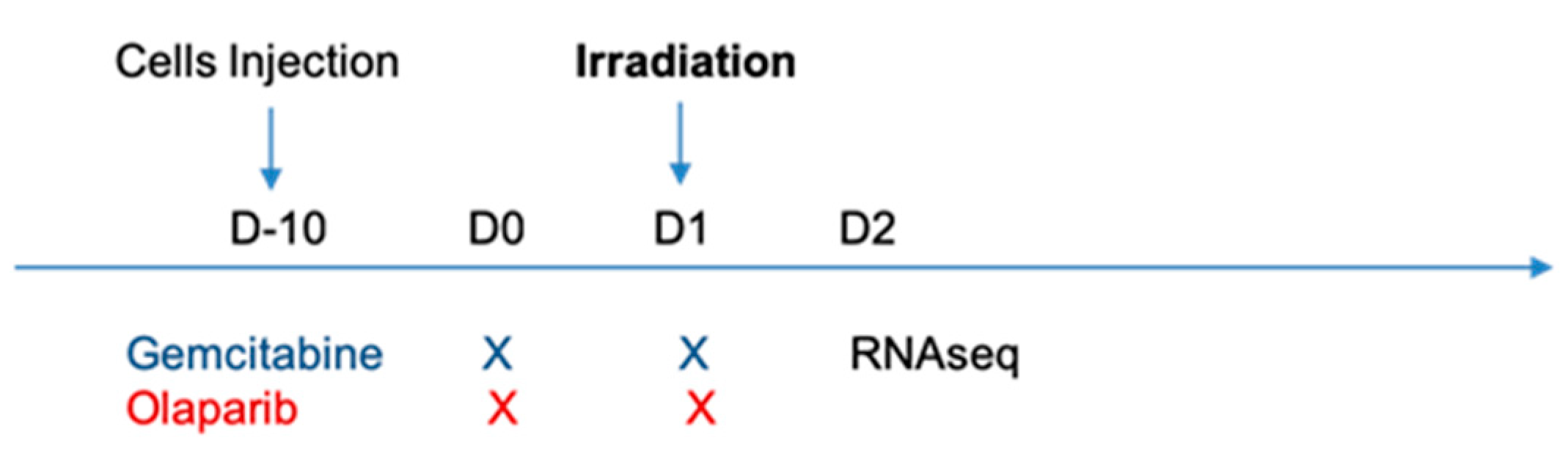
4.2. Treatment Protocols
4.3. Irradiation Protocols
4.3.1. Photon Irradiation
4.3.2. Proton Irradiation
4.4. RNA Extraction, RNA-Seq Profiling and Gene Enrichment Analysis
4.5. Statistical Analysis
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Vincent, A.; Herman, J.; Schulick, R.; Hruban, R.H.; Goggins, M. Pancreatic cancer. Lancet 2011, 378, 607–620. [Google Scholar] [CrossRef]
- Suker, M.; Beumer, B.R.; Sadot, E.; Marthey, L.; Faris, J.E.; Mellon, E.A.; El-Rayes, B.F.; Wang-Gillam, A.; Lacy, J.; Hosein, P.J.; et al. FOLFIRINOX for locally advanced pancreatic cancer: A systematic review and patient-level meta-analysis. Lancet Oncol. 2016, 17, 801–810. [Google Scholar] [CrossRef]
- Huguet, F.; Mukherjee, S.; Javle, M. Locally advanced pancreatic cancer: The role of definitive chemoradiotherapy. Clin. Oncol. 2014, 26, 560–568. [Google Scholar] [CrossRef] [PubMed]
- Rutenberg, M.S.; Nichols, R.C. Proton beam radiotherapy for pancreas cancer. J. Gastrointest. Oncol. 2020, 11, 166–175. [Google Scholar] [CrossRef] [PubMed]
- Tommasino, F.; Durante, M. Proton radiobiology. Cancers 2015, 7, 353–381. [Google Scholar] [CrossRef] [PubMed]
- Woodward, W.A.; Amos, R.A. Proton radiation biology considerations for radiation oncologists. Int. J. Radiat. Oncol. Biol. Phys. 2016, 95, 59–61. [Google Scholar] [CrossRef] [PubMed]
- Nichols, R.C.; George, T.J.; Zaiden, R.A.; Awad, Z.T.; Asbun, H.J.; Huh, S.; Ho, M.W.; Mendenhall, N.P.; Morris, C.G.; Hoppe, B.S. Proton therapy with concomitant capecitabine for pancreatic and ampullary cancers is associated with a low incidence of gastrointestinal toxicity. Acta Oncol. 2013, 52, 498–505. [Google Scholar] [CrossRef]
- Alvarellos, M.L.; Lamba, J.; Sangkuhl, K.; Thorn, C.F.; Wang, L.; Klein, D.J.; Altman, R.B.; Klein, T.E. PharmGKB summary: Gemcitabine pathway. Pharm. Genom. 2014, 24, 564–574. [Google Scholar] [CrossRef]
- Im, M.M.; Flanagan, S.A.; Ackroyd, J.J.; Shewach, D.S. Drug metabolism and homologous recombination repair in radiosensitization with gemcitabine. Radiat. Res. 2015, 183, 114–123. [Google Scholar] [CrossRef]
- Curtin, N.J.; Szabo, C. Therapeutic applications of PARP inhibitors: Anticancer therapy and beyond. Mol. Asp. Med. 2013, 34, 1217–1256. [Google Scholar] [CrossRef]
- Noël, G.; Godon, C.; Fernet, M.; Giocanti, N.; Mégnin-Chanet, F.; Favaudon, V. Radiosensitization by the poly(ADP-ribose) polymerase inhibitor 4-amino-1,8-naphthalimide is specific of the S phase of the cell cycle and involves arrest of DNA synthesis. Mol. Cancer Ther. 2006, 5, 564–574. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Paganetti, H. Relative biological effectiveness (RBE) values for proton beam therapy. Variations as a function of biological endpoint, dose, and linear energy transfer. Phys. Med. Biol. 2014, 59, R419–R472. [Google Scholar] [CrossRef] [PubMed]
- Newhauser, W. International commission on radiation units and measurements report 78: Prescribing, recording and reporting proton-beam therapy. Radiat. Prot. Dosim. 2009, 133, 60–62. [Google Scholar] [CrossRef]
- Karnak, D.; Engelke, C.G.; Parsels, L.A.; Kausar, T.; Wei, D.; Robertson, J.R.; Marsh, K.B.; Davis, M.A.; Zhao, L.; Maybaum, J.; et al. Combined inhibition of Wee1 and PARP1/2 for radiosensitization in pancreatic cancer. Clin. Cancer Res. 2014, 20, 5085–5096. [Google Scholar] [CrossRef]
- Lohse, I.; Kumareswaran, R.; Cao, P.; Pitcher, B.; Gallinger, S.; Bristow, R.G.; Hedley, D.W. Effects of combined treatment with ionizing radiation and the PARP inhibitor olaparib in BRCA mutant and wild type patient-derived pancreatic cancer xenografts. PLoS ONE 2016, 11, e0167272. [Google Scholar] [CrossRef]
- Hirai, T.; Saito, S.; Fujimori, H.; Matsushita, K.; Nishio, T.; Okayasu, R.; Masutani, M. Radiosensitization by PARP inhibition to proton beam irradiation in cancer cells. Biochem. Biophys. Res. Commun. 2016, 478, 234–240. [Google Scholar] [CrossRef]
- Keszenman, D.J.; Sutherland, B.M. Yields of clustered DNA damage induced by charged-particle radiations of similar kinetic energy per nucleon: LET dependence in different DNA microenvironments. Radiat. Res. 2010, 174, 238–250. [Google Scholar] [CrossRef]
- Chaudhary, P.; Marshall, T.I.; Currell, F.J.; Kacperek, A.; Schettino, G.; Prise, K.M. Variations in the processing of DNA double-strand breaks along 60-MeV therapeutic proton beams. Int. J. Radiat. Oncol. Biol. Phys. 2016, 95, 86–94. [Google Scholar] [CrossRef]
- Carter, R.J.; Nickson, C.M.; Thompson, J.M.; Kacperek, A.; Hill, M.A.; Parsons, J.L. Complex DNA damage induced by high linear energy transfer alpha-particles and protons triggers a specific cellular DNA damage response. Int. J. Radiat. Oncol. Biol. Phys. 2018, 100, 776–784. [Google Scholar] [CrossRef]
- Hirai, T.; Shirai, H.; Fujimori, H.; Okayasu, R.; Sasai, K.; Masutani, M. Radiosensitization effect of poly(ADP-ribose) polymerase inhibition in cells exposed to low and high liner energy transfer radiation. Cancer Sci. 2012, 103, 1045–1050. [Google Scholar] [CrossRef]
- Kausar, T.; Schreiber, J.S.; Karnak, D.; Parsels, L.A.; Parsels, J.D.; Davis, M.A.; Zhao, L.; Maybaum, J.; Lawrence, T.S.; Morgan, M.A. Sensitization of pancreatic cancers to gemcitabine chemoradiation by WEE1 kinase inhibition depends on homologous recombination repair. Neoplasia 2015, 17, 757–766. [Google Scholar] [CrossRef] [PubMed]
- Engelke, C.G.; Parsels, L.A.; Qian, Y.; Zhang, Q.; Karnak, D.; Robertson, J.R.; Tanska, D.M.; Wei, D.; Davis, M.A.; Parsels, J.D.; et al. Sensitization of pancreatic cancer to chemoradiation by the Chk1 inhibitor MK8776. Clin. Cancer Res. 2013, 19, 4412–4421. [Google Scholar] [CrossRef] [PubMed]
- Fokas, E.; Prevo, R.; Pollard, J.R.; Reaper, P.M.; Charlton, P.A.; Cornelissen, B.; Vallis, K.A.; Hammond, E.M.; Olcina, M.M.; Gillies McKenna, W.; et al. Targeting ATR in vivo using the novel inhibitor VE-822 results in selective sensitization of pancreatic tumors to radiation. Cell Death Dis. 2012, 3, e441. [Google Scholar] [CrossRef] [PubMed]
- Görte, J.; Beyreuther, E.; Danen, E.H.J.; Cordes, N. Comparative proton and photon irradiation combined with pharmacological inhibitors in 3D pancreatic cancer cultures. Cancers 2020, 12, 3216. [Google Scholar] [CrossRef] [PubMed]
- Tuli, R.; Shiao, S.L.; Nissen, N.; Tighiouart, M.; Kim, S.; Osipov, A.; Bryant, M.; Ristow, L.; Placencio-Hickok, V.R.; Hoffman, D.; et al. A phase 1 study of veliparib, a PARP-1/2 inhibitor, with gemcitabine and radiotherapy in locally advanced pancreatic cancer. EBioMedicine 2019. [Google Scholar] [CrossRef]
- Engreitz, J.M.; Ollikainen, N.; Guttman, M. Long non-coding RNAs: Spatial amplifiers that control nuclear structure and gene expression. Nat. Rev. Mol. Cell Biol. 2016, 17, 756–770. [Google Scholar] [CrossRef]
- Dragomir, M.; Chen, B.; Calin, G.A. Exosomal lncRNAs as new players in cell-to-cell communication. Transl. Cancer Res. 2018, 7, S243–S252. [Google Scholar] [CrossRef]
- Bhan, A.; Soleimani, M.; Mandal, S.S. Long noncoding RNA and cancer: A new paradigm. Cancer Res. 2017, 77, 3965–3981. [Google Scholar] [CrossRef]
- Nagasawa, M.; Tomimatsu, K.; Terada, K.; Kondo, K.; Miyazaki, K.; Miyazaki, M.; Motooka, D.; Okuzaki, D.; Yoshida, T.; Kageyama, S.; et al. Long non-coding RNA MANCR is a target of BET bromodomain protein BRD4 and plays a critical role in cellular migration and invasion abilities of prostate cancer. Biochem. Biophys. Res. Commun. 2020, 526, 128–134. [Google Scholar] [CrossRef]
- Tracy, K.M.; Tye, C.E.; Ghule, P.N.; Malaby, H.L.H.; Stumpff, J.; Stein, J.L.; Stein, G.S.; Lian, J.B. Mitotically-Associated lncRNA (MANCR) affects genomic stability and cell division in aggressive breast cancer. Mol. Cancer Res. 2018, 16, 587–598. [Google Scholar] [CrossRef]
- Wen, S.; Zeng, M.; Li, Y.; Hu, X.; Li, S.; Liang, X.; Zhu, L.; Yang, S. Downregulation of MANCR inhibits cancer cell proliferation in mantle cell lymphoma possibly by interacting with RUNX2. Acta Biochim. Biophys. Sin. 2019, 51, 1142–1147. [Google Scholar] [CrossRef] [PubMed]
- Yao, L.; Yan, J.; Gan, L.; Huang, S.; Cheng, F.; Fang, N. Upregulation of MANCR predicts poor survival in patients with gastric cancer. Oncol. Lett. 2019, 18, 6801–6806. [Google Scholar] [CrossRef] [PubMed]
- Fritz, J.; Karakhanova, S.; Brecht, R.; Nachtigall, I.; Werner, J.; Bazhin, A.V. In vitro immunomodulatory properties of gemcitabine alone and in combination with interferon-alpha. Immunol. Lett. 2015, 168, 111–119. [Google Scholar] [CrossRef] [PubMed]
- Martinez-Bosch, N.; Vinaixa, J.; Navarro, P. Immune evasion in pancreatic cancer: From mechanisms to therapy. Cancers 2018, 10, 6. [Google Scholar] [CrossRef] [PubMed]
- Césaire, M.; Thariat, J.; Candéias, S.M.; Stefan, D.; Saintigny, Y.; Chevalier, F. Combining PARP inhibition, radiation, and immunotherapy: A possible strategy to improve the treatment of cancer? Int. J. Mol. Sci. 2018, 19, 3793. [Google Scholar] [CrossRef]
- Castle, K.D.; Chen, M.; Wisdom, A.J.; Kirsch, D.G. Genetically engineered mouse models for studying radiation biology. Transl. Cancer Res. 2017, 6, S900–S913. [Google Scholar] [CrossRef]
- Kersten, K.; de Visser, K.E.; van Miltenburg, M.H.; Jonkers, J. Genetically engineered mouse models in oncology research and cancer medicine. EMBO Mol. Med. 2017, 9, 137–153. [Google Scholar] [CrossRef]
- Baba, Y.; Nosho, K.; Shima, K.; Irahara, N.; Kure, S.; Toyoda, S.; Kirkner, G.J.; Goel, A.; Fuchs, C.S.; Ogino, S. Aurora-A expression is independently associated with chromosomal instability in colorectal cancer. Neoplasia 2009, 11, 418–425. [Google Scholar] [CrossRef]
- Vanstalle, M.; Constanzo, J.; Karakaya, Y.; Finck, C.; Rousseau, M.; Brasse, D. Analytical dose modeling for preclinical proton irradiation of millimetric targets. Med. Phys. 2018, 45, 470–478. [Google Scholar] [CrossRef]
- Constanzo, J.; Vanstalle, M.; Finck, C.; Brasse, D.; Rousseau, M. Dosimetry and characterization of a 25-MeV proton beam line for preclinical radiobiology research. Med. Phys. 2019, 46, 2356–2362. [Google Scholar] [CrossRef]
- Dobin, A.; Davis, C.A.; Schlesinger, F.; Drenkow, J.; Zaleski, C.; Jha, S.; Batut, P.; Chaisson, M.; Gingeras, T.R. STAR: Ultrafast universal RNA-seq aligner. Bioinformatics 2013, 29, 15–21. [Google Scholar] [CrossRef] [PubMed]
- Anders, S.; Pyl, P.T.; Huber, W. HTSeq—A Python framework to work with high-throughput sequencing data. Bioinformatics 2015, 31, 166–169. [Google Scholar] [CrossRef] [PubMed]
- Love, M.I.; Huber, W.; Anders, S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome. Biol. 2014, 15. [Google Scholar] [CrossRef] [PubMed]
- Blighe, K.; Rana, S.; Lewis, M. EnhancedVolcano: Publication-Ready Volcano Plots with Enhanced Colouring and Labeling; Bioconductor, Version: 1.8.0; Available online: https://github.com/kevinblighe/EnhancedVolcano (accessed on 30 January 2021).
- Wickham, H. Ggplot2: Elegant Graphics for Data Analysis; Use R; Springer: New York, NY, USA, 2009; ISBN 978-0-387-98140-6. [Google Scholar]
- Yu, G.; Wang, L.-G.; Han, Y.; He, Q.-Y. clusterProfiler: An R package for comparing biological themes among gene clusters. OMICS J. Integr. Biol. 2012, 16, 284–287. [Google Scholar] [CrossRef]
| Comparison (A vs. B) | Number of DE Transcripts | Number of lncRNA |
|---|---|---|
| CTL vs. RT | 9 | 1 (11.1%) |
| RTG vs. RT | 150 | 10 (6.67%) |
| PT vs. RT | 417 | 29 (6.95%) |
| RTO vs. RT | 0 | 0 (0%) |
| RTOG vs. RT | 266 | 17 (6.39%) |
| RTG vs. RTOG | 39 | 1 (2.6%) |
| RTO vs. RTOG | 38 | 4 (10.5%) |
| CTL vs. PT | 778 | 46 (5.91%) |
| PT vs. PTG | 128 | 9 (7.03%) |
| PTO vs. PT | 657 | 8 (1.2%) |
| PTOG vs. PT | 1679 | 61 (3.63%) |
| PTG vs. PTOG | 0 | 0 (0%) |
| PTO vs. PTOG | 44 | 6 (13.63%) |
| RTOG vs. PTOG | 0 | 0 (0%) |
| PTO vs. RTO | 33 | 1 (0.03%) |
| PTG vs. RTG | 306 | 20 (6.53%) |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Waissi, W.; Nicol, A.; Jung, M.; Rousseau, M.; Jarnet, D.; Noel, G.; Burckel, H. Radiosensitizing Pancreatic Cancer with PARP Inhibitor and Gemcitabine: An In Vivo and a Whole-Transcriptome Analysis after Proton or Photon Irradiation. Cancers 2021, 13, 527. https://doi.org/10.3390/cancers13030527
Waissi W, Nicol A, Jung M, Rousseau M, Jarnet D, Noel G, Burckel H. Radiosensitizing Pancreatic Cancer with PARP Inhibitor and Gemcitabine: An In Vivo and a Whole-Transcriptome Analysis after Proton or Photon Irradiation. Cancers. 2021; 13(3):527. https://doi.org/10.3390/cancers13030527
Chicago/Turabian StyleWaissi, Waisse, Anaïs Nicol, Matthieu Jung, Marc Rousseau, Delphine Jarnet, Georges Noel, and Hélène Burckel. 2021. "Radiosensitizing Pancreatic Cancer with PARP Inhibitor and Gemcitabine: An In Vivo and a Whole-Transcriptome Analysis after Proton or Photon Irradiation" Cancers 13, no. 3: 527. https://doi.org/10.3390/cancers13030527
APA StyleWaissi, W., Nicol, A., Jung, M., Rousseau, M., Jarnet, D., Noel, G., & Burckel, H. (2021). Radiosensitizing Pancreatic Cancer with PARP Inhibitor and Gemcitabine: An In Vivo and a Whole-Transcriptome Analysis after Proton or Photon Irradiation. Cancers, 13(3), 527. https://doi.org/10.3390/cancers13030527





