Lack of Electron Acceptors Contributes to Redox Stress and Growth Arrest in Asparagine-Starved Sarcoma Cells
Abstract
Simple Summary
Abstract
1. Introduction
2. Results
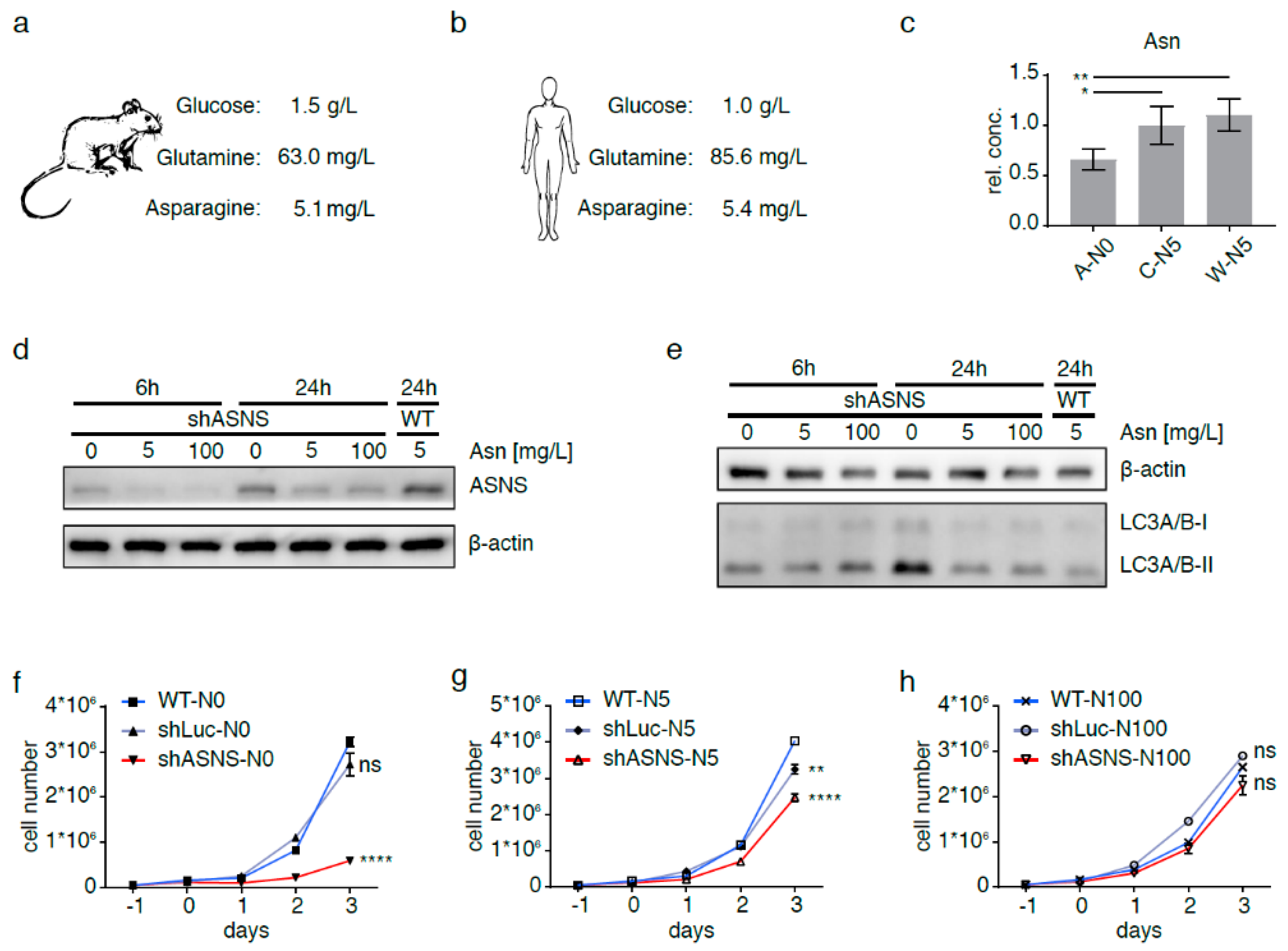
2.1. Asparagine Deprivation of Mouse Sarcoma Cells
2.2. Asparagine Starvation Impacts Proliferation, Apoptosis and Autophagy of Mouse Sarcoma Cells
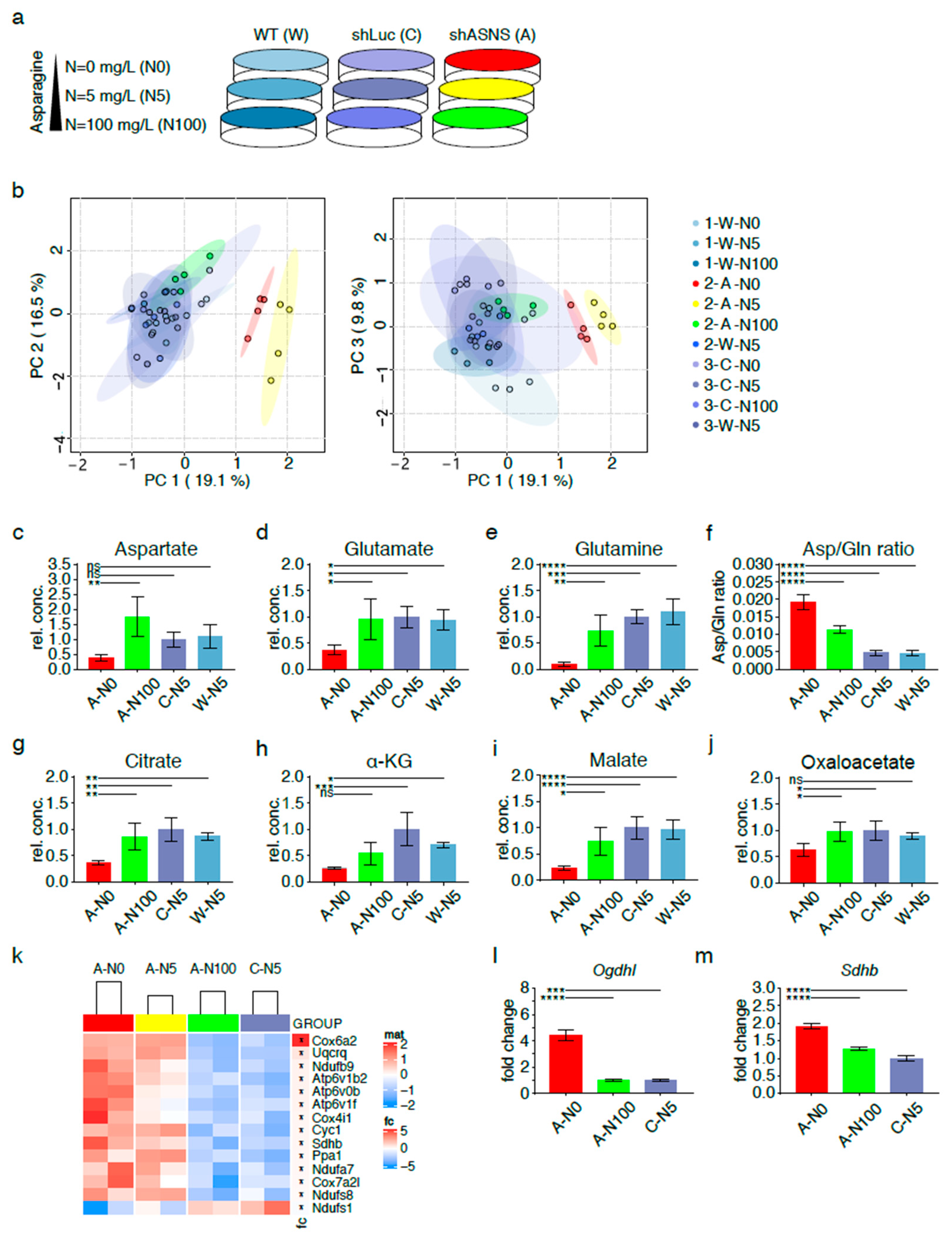
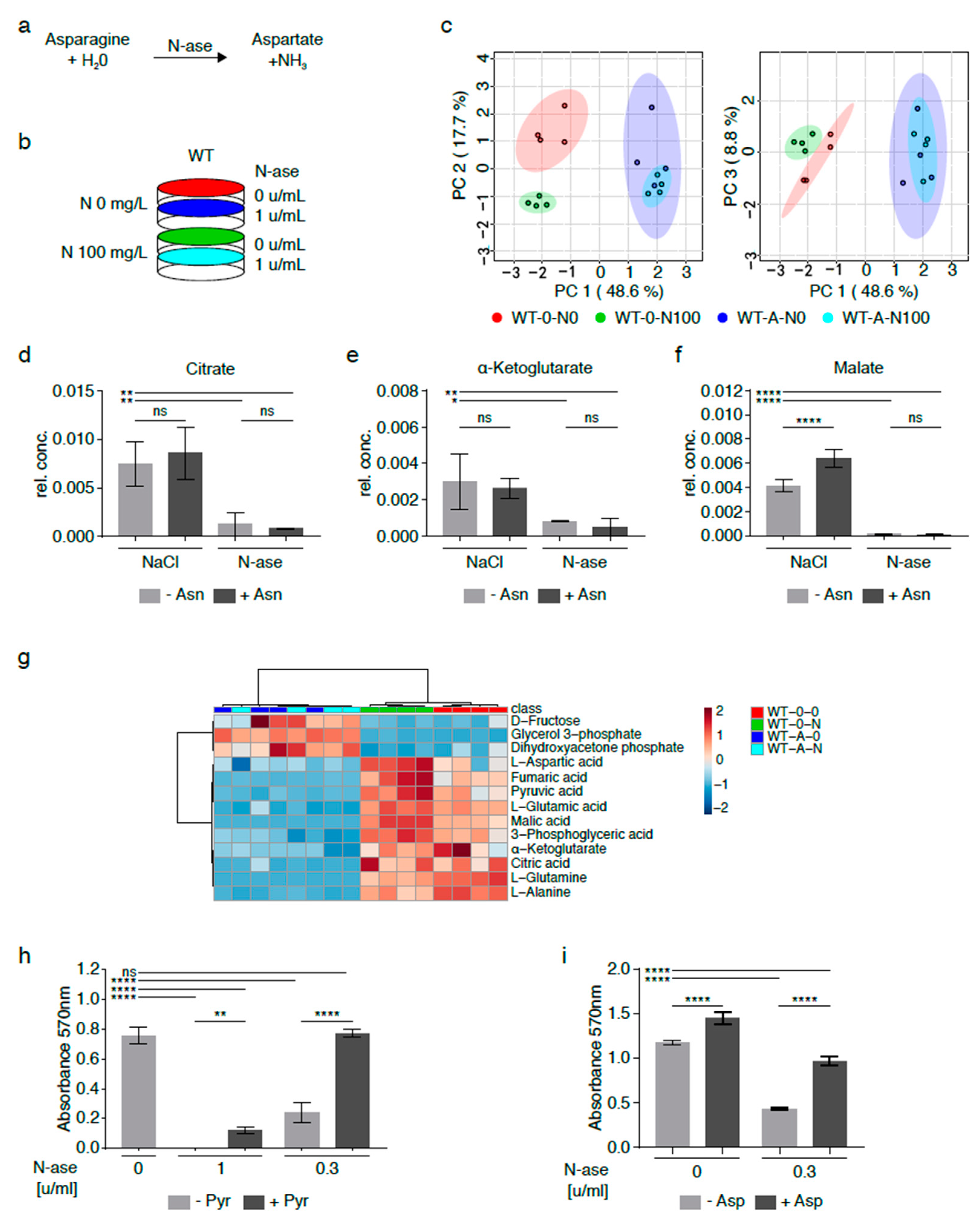
2.3. Metabolic Adaptation of Asparagine-Starved Mouse Sarcoma Cells
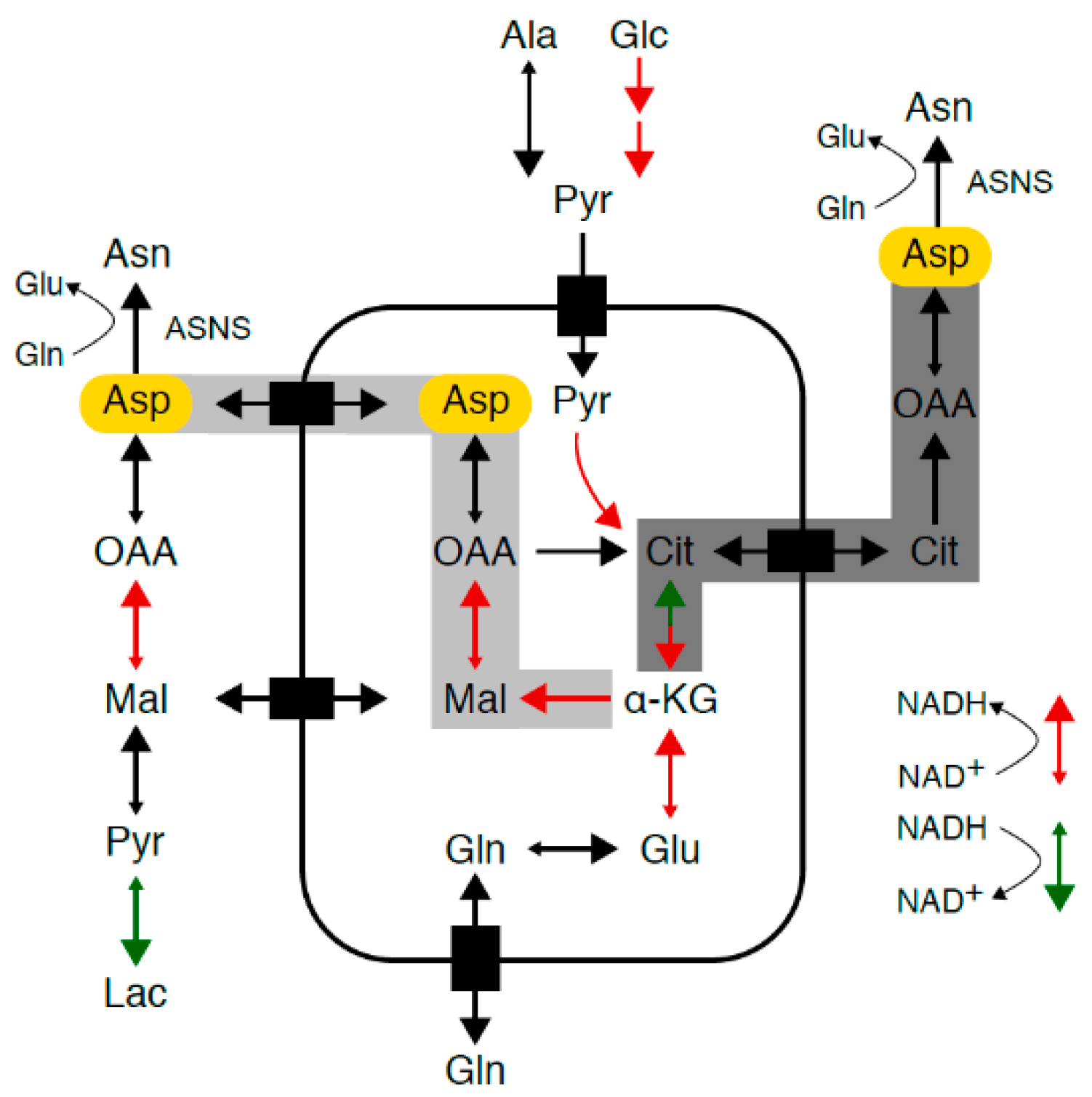
2.4. Redirection of TCA Cycle flux in Asparagine-Depleted Mouse Sarcoma Cells
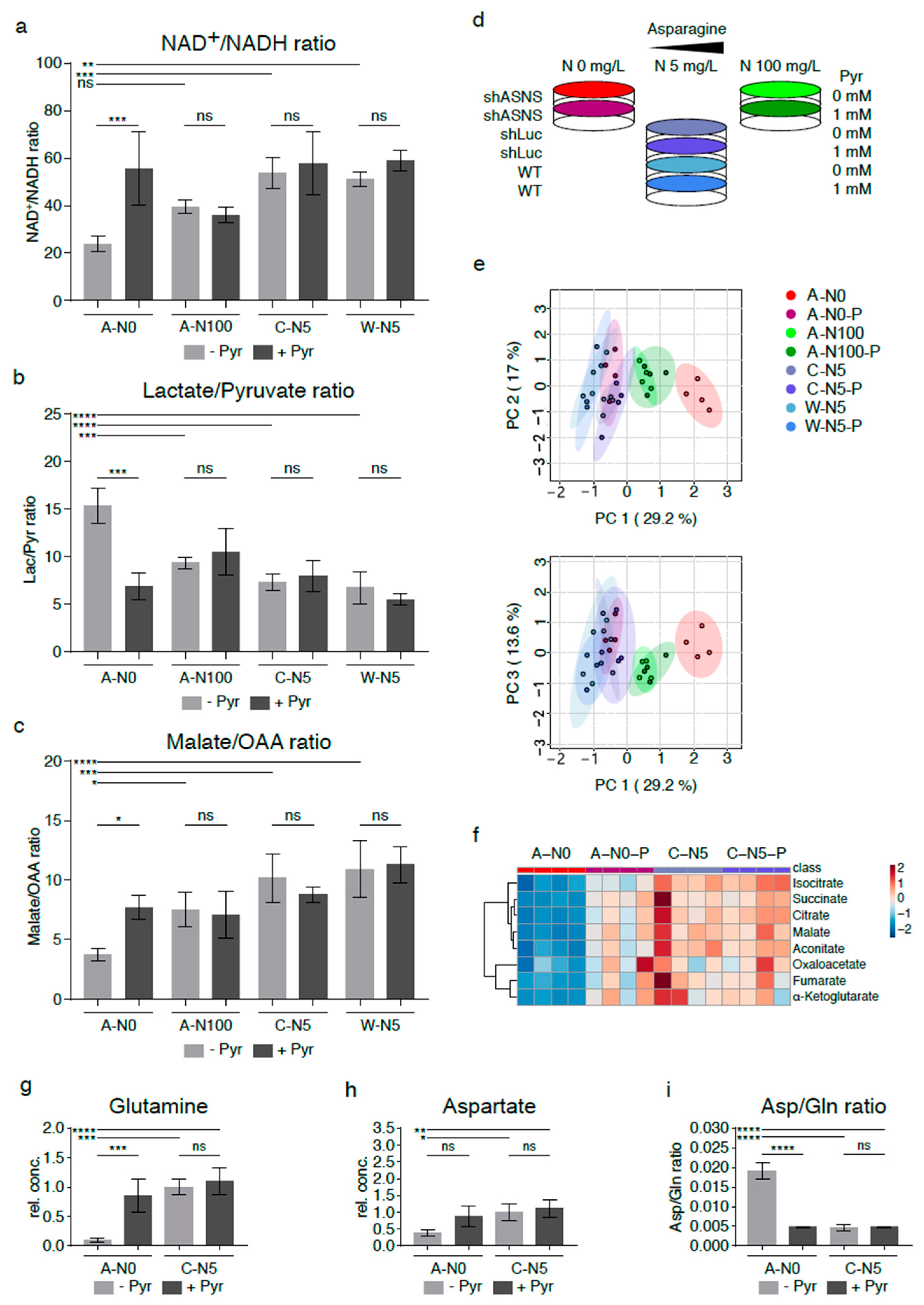
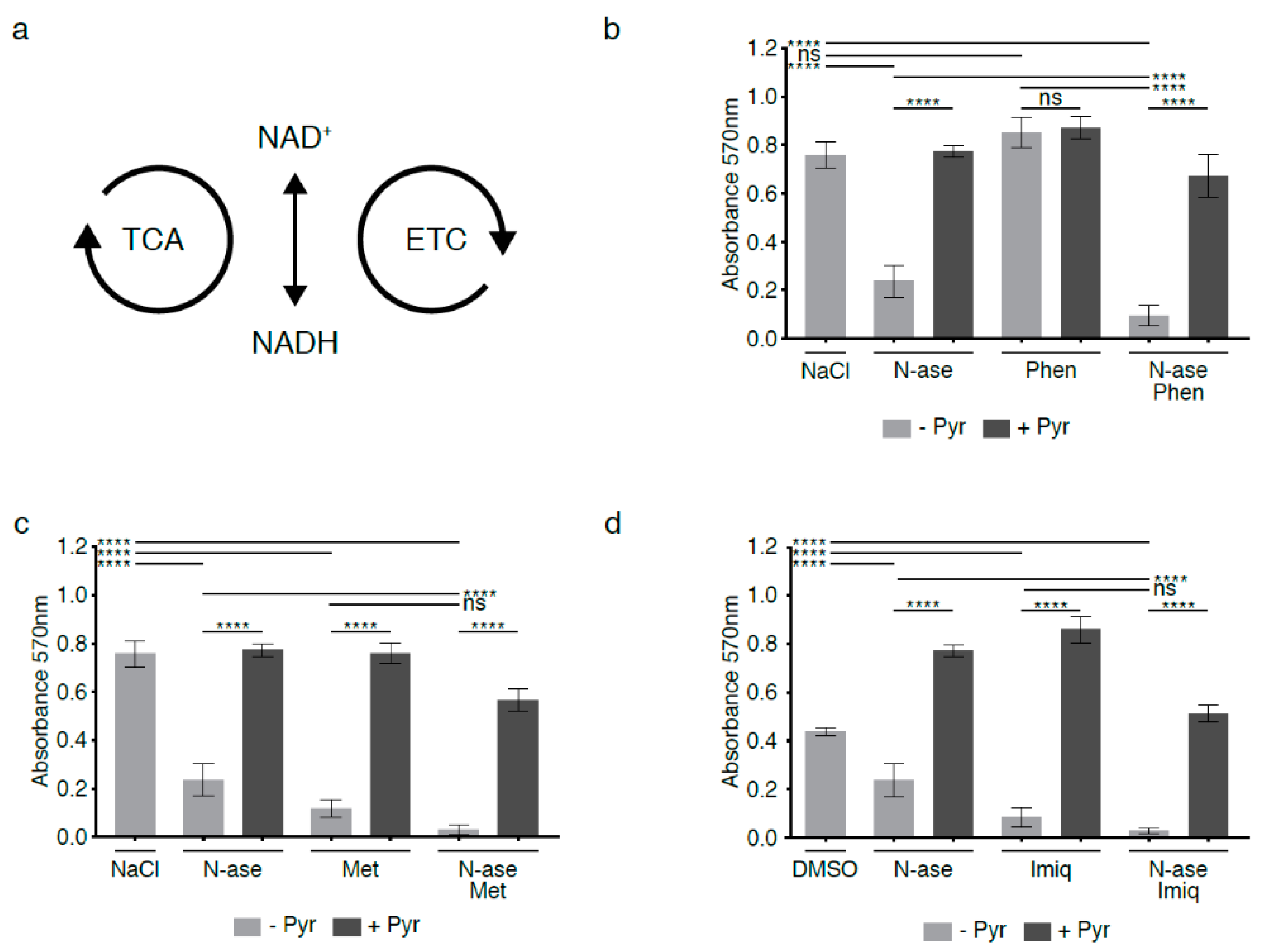
2.5. Relative Lack of the Electron Acceptor NAD+ in Asparagine-Starved Cells
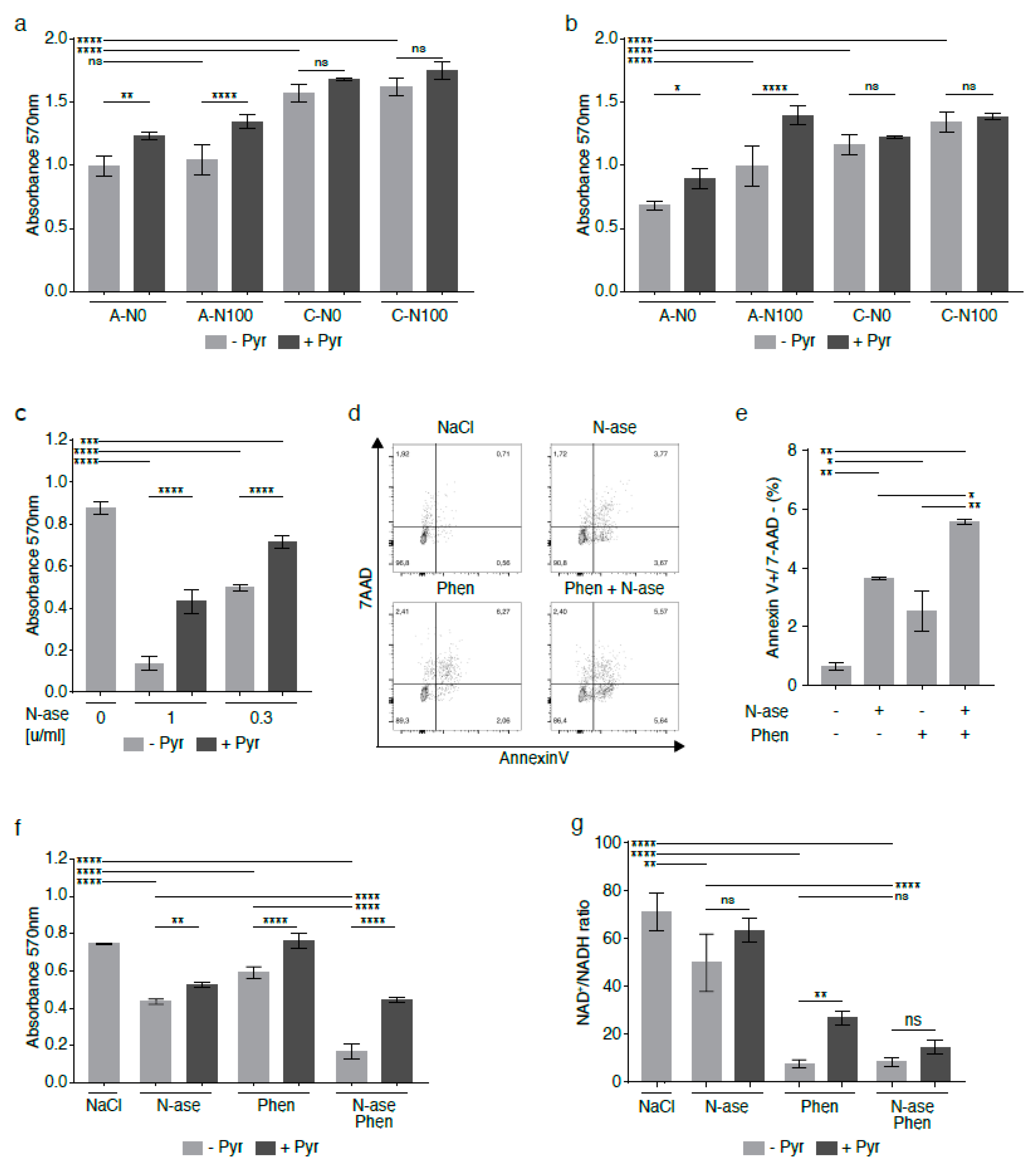
2.6. Supplementation with Pyruvate Reverses Reductive Stress and Global Metabolomic Changes in Asparagine-Starved Sarcoma Cells
2.7. Supplementation with Pyruvate Reverses the Growth-Inhibitory Effects of Asparagine Depletion in Mouse Sarcoma Cells
2.8. Complex 1 Inhibitors Augment the Anti-Proliferative Effects of Asparagine Depletion on Mouse Sarcoma Cells
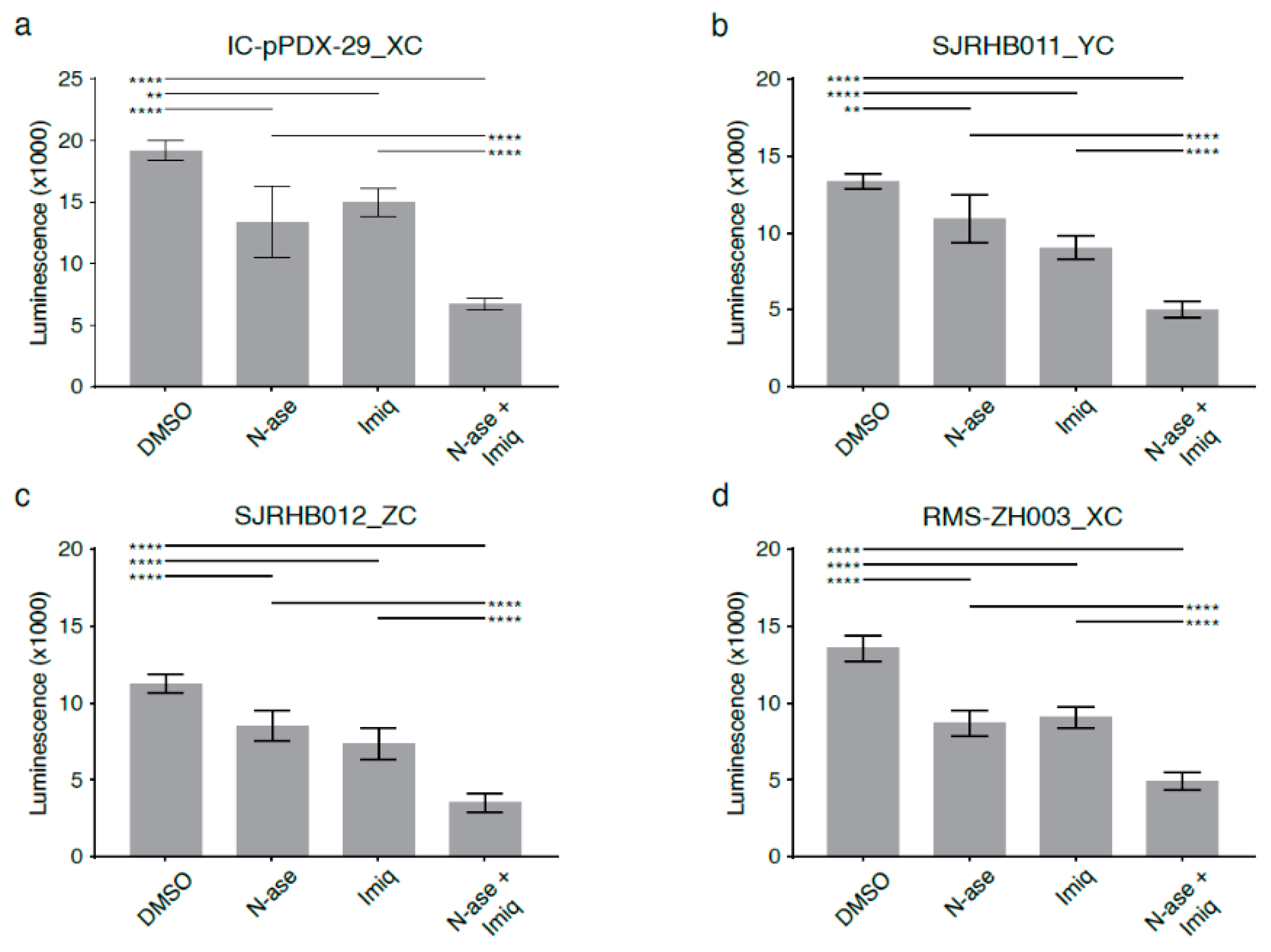
2.9. Complex 1 Inhibitors Augment Reductive Stress and Growth Inhibition Induced by Asparagine Deprivation of Human Sarcoma Cells
3. Discussion
4. Materials and Methods
4.1. Cell Lines
4.2. ASNS Silencing
4.3. Proliferation Assays
4.4. Cell Viability Assay
4.5. Annexin V Staining
4.6. Western Blots
4.7. RNA Isolation, RNA-Seq and qRT-PCR
4.8. Metabolite Extraction
4.9. Gas Chromatography–Mass Spectrometry (GC-MS)
4.10. GC-MS Data Analysis
4.11. Liquid Chromatography–Mass Spectrometry (LC-MS)
4.12. LC-MS Data Analysis
4.13. Oxygen Consumption Rate
4.14. Statistics
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Lieu, E.L.; Nguyen, T.; Rhyne, S.; Kim, J. Amino acids in cancer. Exp. Mol. Med. 2020, 52, 15–30. [Google Scholar] [CrossRef]
- DeBerardinis, R.J.; Mancuso, A.; Daikhin, E.; Nissim, I.; Yudkoff, M.; Wehrli, S.; Thompson, C.B. Beyond aerobic glycolysis: Transformed cells can engage in glutamine metabolism that exceeds the requirement for protein and nucleotide synthesis. Proc. Natl. Acad. Sci. USA 2007, 104, 19345–19350. [Google Scholar] [CrossRef]
- Hettmer, S.; Schinzel, A.C.; Tchessalova, D.; Schneider, M.; Parker, C.L.; Bronson, R.T.; Richards, N.G.; Hahn, W.C.; Wagers, A.J. Functional genomic screening reveals asparagine dependence as a metabolic vulnerability in sarcoma. Elife 2015, 4. [Google Scholar] [CrossRef]
- Gwinn, D.M.; Lee, A.G.; Briones-Martin-Del-Campo, M.; Conn, C.S.; Simpson, D.R.; Scott, A.I.; Le, A.; Cowan, T.M.; Ruggero, D.; Sweet-Cordero, E.A. Oncogenic KRAS Regulates Amino Acid Homeostasis and Asparagine Biosynthesis via ATF4 and Alters Sensitivity to L-Asparaginase. Cancer Cell 2018, 33, 91–107. [Google Scholar] [CrossRef]
- Krall, A.S.; Xu, S.; Graeber, T.G.; Braas, D.; Christofk, H.R. Asparagine promotes cancer cell proliferation through use as an amino acid exchange factor. Nat. Commun. 2016, 7, 11457. [Google Scholar] [CrossRef]
- Balasubramanian, M.N.; Butterworth, E.A.; Kilberg, M.S. Asparagine synthetase: Regulation by cell stress and involvement in tumor biology. Am. J. Physiol. Endocrinol. Metab. 2013, 304, E789–E799. [Google Scholar] [CrossRef]
- Takach, E.; O’Shea, T.; Liu, H. High-throughput quantitation of amino acids in rat and mouse biological matrices using stable isotope labeling and UPLC-MS/MS analysis. J. Chromatogr. B Analyt. Technol. Biomed. Life Sci. 2014, 964, 180–190. [Google Scholar] [CrossRef]
- Psychogios, N.; Hau, D.D.; Peng, J.; Guo, A.C.; Mandal, R.; Bouatra, S.; Sinelnikov, I.; Krishnamurthy, R.; Eisner, R.; Gautam, B.; et al. The human serum metabolome. PLoS ONE 2011, 6, e16957. [Google Scholar] [CrossRef]
- Knott, S.R.V.; Wagenblast, E.; Khan, S.; Kim, S.Y.; Soto, M.; Wagner, M.; Turgeon, M.-O.; Fish, L.; Erard, N.; Gable, A.L.; et al. Asparagine bioavailability governs metastasis in a model of breast cancer. Nature 2018, 554, 378–381. [Google Scholar] [CrossRef]
- Zhang, J.; Fan, J.; Venneti, S.; Cross, J.R.; Takagi, T.; Bhinder, B.; Djaballah, H.; Kanai, M.; Cheng, E.H.; Judkins, A.R.; et al. Asparagine plays a critical role in regulating cellular adaptation to glutamine depletion. Mol. Cell 2014, 56, 205–218. [Google Scholar] [CrossRef]
- Fine, B.M.; Kaspers, G.J.L.; Ho, M.; Loonen, A.H.; Boxer, L.M. A genome-wide view of the in vitro response to l-asparaginase in acute lymphoblastic leukemia. Cancer Res. 2005, 65, 291–299. [Google Scholar]
- Hettmer, S.; Liu, J.; Miller, C.M.; Lindsay, M.C.; Sparks, C.A.; Guertin, D.A.; Bronson, R.T.; Langenau, D.M.; Wagers, A.J. Sarcomas induced in discrete subsets of prospectively isolated skeletal muscle cells. Proc. Natl. Acad. Sci. USA 2011, 108, 20002–20007. [Google Scholar] [CrossRef]
- Birsoy, K.; Wang, T.; Chen, W.W.; Freinkman, E.; Abu-Remaileh, M.; Sabatini, D.M. An Essential Role of the Mitochondrial Electron Transport Chain in Cell Proliferation Is to Enable Aspartate Synthesis. Cell 2015, 162, 540–551. [Google Scholar] [CrossRef]
- Sullivan, L.B.; Gui, D.Y.; Hosios, A.M.; Bush, L.N.; Freinkman, E.; Vander Heiden, M.G. Supporting Aspartate Biosynthesis Is an Essential Function of Respiration in Proliferating Cells. Cell 2015, 162, 552–563. [Google Scholar] [CrossRef]
- Sullivan, L.B.; Luengo, A.; Danai, L.V.; Bush, L.N.; Diehl, F.F.; Hosios, A.M.; Lau, A.N.; Elmiligy, S.; Malstrom, S.; Lewis, C.A.; et al. Aspartate is an endogenous metabolic limitation for tumour growth. Nat. Cell Biol. 2018, 20, 782–788. [Google Scholar] [CrossRef]
- Richards, N.G.J.; Kilberg, M.S. Asparagine synthetase chemotherapy. Annu. Rev. Biochem. 2006, 75, 629–654. [Google Scholar] [CrossRef]
- Purwaha, P.; Lorenzi, P.L.; Silva, L.P.; Hawke, D.H.; Weinstein, J.N. Targeted metabolomic analysis of amino acid response to L-asparaginase in adherent cells. Metabolomics 2014, 10, 909–919. [Google Scholar] [CrossRef]
- Hinson, A.R.P.; Jones, R.; Crose, L.E.S.; Belyea, B.C.; Barr, F.G.; Linardic, C.M. Human rhabdomyosarcoma cell lines for rhabdomyosarcoma research: Utility and pitfalls. Front. Oncol. 2013, 3, 183. [Google Scholar] [CrossRef]
- DeBerardinis, R.J.; Lum, J.J.; Hatzivassiliou, G.; Thompson, C.B. The biology of cancer: Metabolic reprogramming fuels cell growth and proliferation. Cell Metab. 2008, 7, 11–20. [Google Scholar] [CrossRef]
- Teodoro, J.S.; Rolo, A.P.; Palmeira, C.M. The NAD ratio redox paradox: Why does too much reductive power cause oxidative stress? Toxicol. Mech. Methods 2013, 23, 297–302. [Google Scholar] [CrossRef]
- Toda, K.; Kawada, K.; Iwamoto, M.; Inamoto, S.; Sasazuki, T.; Shirasawa, S.; Hasegawa, S.; Sakai, Y. Metabolic Alterations Caused by KRAS Mutations in Colorectal Cancer Contribute to Cell Adaptation to Glutamine Depletion by Upregulation of Asparagine Synthetase. Neoplasia 2016, 18, 654–665. [Google Scholar] [CrossRef]
- Chen, Q.; Ye, L.; Fan, J.; Zhang, X.; Wang, H.; Liao, S.; Song, P.; Wang, Z.; Wang, S.; Li, Y.; et al. Autophagy suppression potentiates the anti-glioblastoma effect of asparaginase in vitro and in vivo. Oncotarget 2017, 8, 91052–91066. [Google Scholar] [CrossRef]
- Horton, T.M.; Gannavarapu, A.; Blaney, S.M.; D’Argenio, D.Z.; Plon, S.E.; Berg, S.L. Bortezomib interactions with chemotherapy agents in acute leukemia in vitro. Cancer Chemother. Pharmacol. 2006, 58, 13–23. [Google Scholar] [CrossRef]
- Manzella, G.; Schreck, L.D.; Breunis, W.B.; Molenaar, J.; Merks, H.; Barr, F.G.; Sun, W.; Römmele, M.; Zhang, L.; Tchinda, J.; et al. Phenotypic profiling with a living biobank of primary rhabdomyosarcoma unravels disease heterogeneity and AKT sensitivity. Nat. Commun. 2020, 11, 4629. [Google Scholar] [CrossRef]
- Bolger, A.M.; Lohse, M.; Usadel, B. Trimmomatic: A flexible trimmer for Illumina sequence data. Bioinformatics 2014, 30, 2114–2120. [Google Scholar] [CrossRef]
- Dobin, A.; Davis, C.A.; Schlesinger, F.; Drenkow, J.; Zaleski, C.; Jha, S.; Batut, P.; Chaisson, M.; Gingeras, T.R. STAR: Ultrafast universal RNA-seq aligner. Bioinformatics 2013, 29, 15–21. [Google Scholar] [CrossRef]
- Ritchie, M.E.; Phipson, B.; Wu, D.; Hu, Y.; Law, C.W.; Shi, W.; Smyth, G.K. limma powers differential expression analyses for RNA-sequencing and microarray studies. Nucleic Acids Res. 2015, 43, e47. [Google Scholar] [CrossRef]
- Luo, W.; Friedman, M.S.; Shedden, K.; Hankenson, K.D.; Woolf, P.J. GAGE: Generally applicable gene set enrichment for pathway analysis. BMC Bioinform. 2009, 10, 161. [Google Scholar] [CrossRef]
- Subramanian, A.; Tamayo, P.; Mootha, V.K.; Mukherjee, S.; Ebert, B.L.; Gillette, M.A.; Paulovich, A.; Pomeroy, S.L.; Golub, T.R.; Lander, E.S.; et al. Gene set enrichment analysis: A knowledge-based approach for interpreting genome-wide expression profiles. Proc. Natl. Acad. Sci. USA 2005, 102, 15545–15550. [Google Scholar] [CrossRef]
- Haug, K.; Cochrane, K.; Nainala, V.C.; Williams, M.; Chang, J.; Jayaseelan, K.V.; O’Donovan, C. MetaboLights: A resource evolving in response to the needs of its scientific community. Nucleic Acids Res. 2020, 48, D440–D444. [Google Scholar] [CrossRef]
- Styczynski, M.P.; Moxley, J.F.; Tong, L.V.; Walther, J.L.; Jensen, K.L.; Stephanopoulos, G.N. Systematic identification of conserved metabolites in GC/MS data for metabolomics and biomarker discovery. Anal. Chem. 2007, 79, 966–973. [Google Scholar] [CrossRef] [PubMed]
- Kind, T.; Wohlgemuth, G.; Lee, D.Y.; Lu, Y.; Palazoglu, M.; Shahbaz, S.; Fiehn, O. FiehnLib: Mass spectral and retention index libraries for metabolomics based on quadrupole and time-of-flight gas chromatography/mass spectrometry. Anal. Chem. 2009, 81, 10038–10048. [Google Scholar] [CrossRef] [PubMed]
- Babushok, V.I.; Linstrom, P.J.; Reed, J.J.; Zenkevich, I.G.; Brown, R.L.; Mallard, W.G.; Stein, S.E. Development of a database of gas chromatographic retention properties of organic compounds. J. Chromatogr. A 2007, 1157, 414–421. [Google Scholar] [CrossRef] [PubMed]
- Kopka, J.; Schauer, N.; Krueger, S.; Birkemeyer, C.; Usadel, B.; Bergmüller, E.; Dörmann, P.; Weckwerth, W.; Gibon, Y.; Stitt, M.; et al. GMD@CSB.DB: The Golm Metabolome Database. Bioinformatics 2005, 21, 1635–1638. [Google Scholar] [CrossRef] [PubMed]
- Hutschenreuther, A.; Kiontke, A.; Birkenmeier, G.; Birkemeyer, C. Comparison of extraction conditions and normalization approaches for cellular metabolomics of adherent growing cells with GC-MS. Anal. Methods 2012, 4, 1953. [Google Scholar] [CrossRef]
- Tan, B.; Lu, Z.; Dong, S.; Zhao, G.; Kuo, M.-S. Derivatization of the tricarboxylic acid intermediates with O-benzylhydroxylamine for liquid chromatography-tandem mass spectrometry detection. Anal. Biochem. 2014, 465, 134–147. [Google Scholar] [CrossRef]
- Chong, J.; Soufan, O.; Li, C.; Caraus, I.; Li, S.; Bourque, G.; Wishart, D.S.; Xia, J. MetaboAnalyst 4.0: Towards more transparent and integrative metabolomics analysis. Nucleic Acids Res. 2018, 46, W486–W494. [Google Scholar] [CrossRef]
- Chong, J.; Wishart, D.S.; Xia, J. Using MetaboAnalyst 4.0 for Comprehensive and Integrative Metabolomics Data Analysis. Curr. Protoc. Bioinform. 2019, 68, e86. [Google Scholar] [CrossRef]
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Bauer, C.; Quante, M.; Breunis, W.B.; Regina, C.; Schneider, M.; Andrieux, G.; Gorka, O.; Groß, O.; Boerries, M.; Kammerer, B.; et al. Lack of Electron Acceptors Contributes to Redox Stress and Growth Arrest in Asparagine-Starved Sarcoma Cells. Cancers 2021, 13, 412. https://doi.org/10.3390/cancers13030412
Bauer C, Quante M, Breunis WB, Regina C, Schneider M, Andrieux G, Gorka O, Groß O, Boerries M, Kammerer B, et al. Lack of Electron Acceptors Contributes to Redox Stress and Growth Arrest in Asparagine-Starved Sarcoma Cells. Cancers. 2021; 13(3):412. https://doi.org/10.3390/cancers13030412
Chicago/Turabian StyleBauer, Christoph, Meret Quante, Willemijn B. Breunis, Carla Regina, Michaela Schneider, Geoffroy Andrieux, Oliver Gorka, Olaf Groß, Melanie Boerries, Bernd Kammerer, and et al. 2021. "Lack of Electron Acceptors Contributes to Redox Stress and Growth Arrest in Asparagine-Starved Sarcoma Cells" Cancers 13, no. 3: 412. https://doi.org/10.3390/cancers13030412
APA StyleBauer, C., Quante, M., Breunis, W. B., Regina, C., Schneider, M., Andrieux, G., Gorka, O., Groß, O., Boerries, M., Kammerer, B., & Hettmer, S. (2021). Lack of Electron Acceptors Contributes to Redox Stress and Growth Arrest in Asparagine-Starved Sarcoma Cells. Cancers, 13(3), 412. https://doi.org/10.3390/cancers13030412





