Insights into the Role of Matrix Metalloproteinases in Precancerous Conditions and in Colorectal Cancer
Simple Summary
Abstract
1. Introduction
2. CRC Pathogenesis and Molecular Classification
2.1. Molecular Mechanism of CRC
- Chromosomal instability (CIN): this abnormality is identified in 85% of CRC cases and consists of a gain (1q, 7p, 8q, 13q, 2pq) or loss (8q, 15q, 17p, 18p) of chromosomal genes, activation of proto-oncogenes (KRAS, SRC, c-MYC), and inactivation of tumor suppressor genes (P53, APC) [21].
2.2. Molecular Classification Based on Transcriptomic Analysis
- Consensus molecular subtype (CMS) classification: CMS classification provides biological insight into metastatic colorectal cancer (mCRC) carcinogenesis and predicts CRC prognosis [25].
- ◦
- CMS1 (14%) indicates MSI, CIMP, and BRAF mutation and immune activation.
- ◦
- CMS2 (37%) shows Wingless-Type MMTR integration site family member (WNT), MYC signaling activation, and epithelial involvement.
- ◦
- CMS3 (13%) demonstrates MSI, CIMP, and KRAS mutations and metabolic involvement.
- ◦
- CMS4 (23%) includes invasion, metastatic situations, and TGF-β signaling co-activation and angiogenesis. Also, epithelial-mesenchymal transition (EMT) is a crucial event in colorectal carcinogenesis and is involved in CMS4 status. EMT can result in advanced-stage CRC, poor patient survival, and worst clinical features [26,27] and CMS4 subgroup shows the most unfavorable prognosis.
- CRC intrinsic subtypes (CRIS): CRIS is a unique classification exclusively based on the cancer cell-specific transcriptome of CRC since the extrinsic factors of the stroma have not been analyzed. It classifies CRC into five novel transcriptional groups that, thus, further clarify biological understanding of CRC heterogeneity.
- ◦
- CRIS-A is enriched for BRAF-mutated MSI tumors and KRAS-mutated MSS tumors that are without targeted therapeutic options.
- ◦
- CRIS-B is related to invasive tumors with poor prognosis and high TGF-ß signaling. CRIS-B is unconnected to the CMS4 mesenchymal subtype, which also indicates aggressive tumors with TGF-ß pathway activation.
- ◦
- CRIS-C is dependent on EGFR signals and is sensitive to anti-EGFR monoclonal antibody treatment.
- ◦
- CRIS-D shows IGF2 overexpression. This occurrence has been involved in desensitization to the EGFR blockade in patients with KRAS wild-type tumors.
- ◦
- CRIS-E indicates KRAS-mutated, Paneth cell-like CIN tumors refractory to anti-EGFR antibody treatment [28].
3. Structure and Function of MMPs
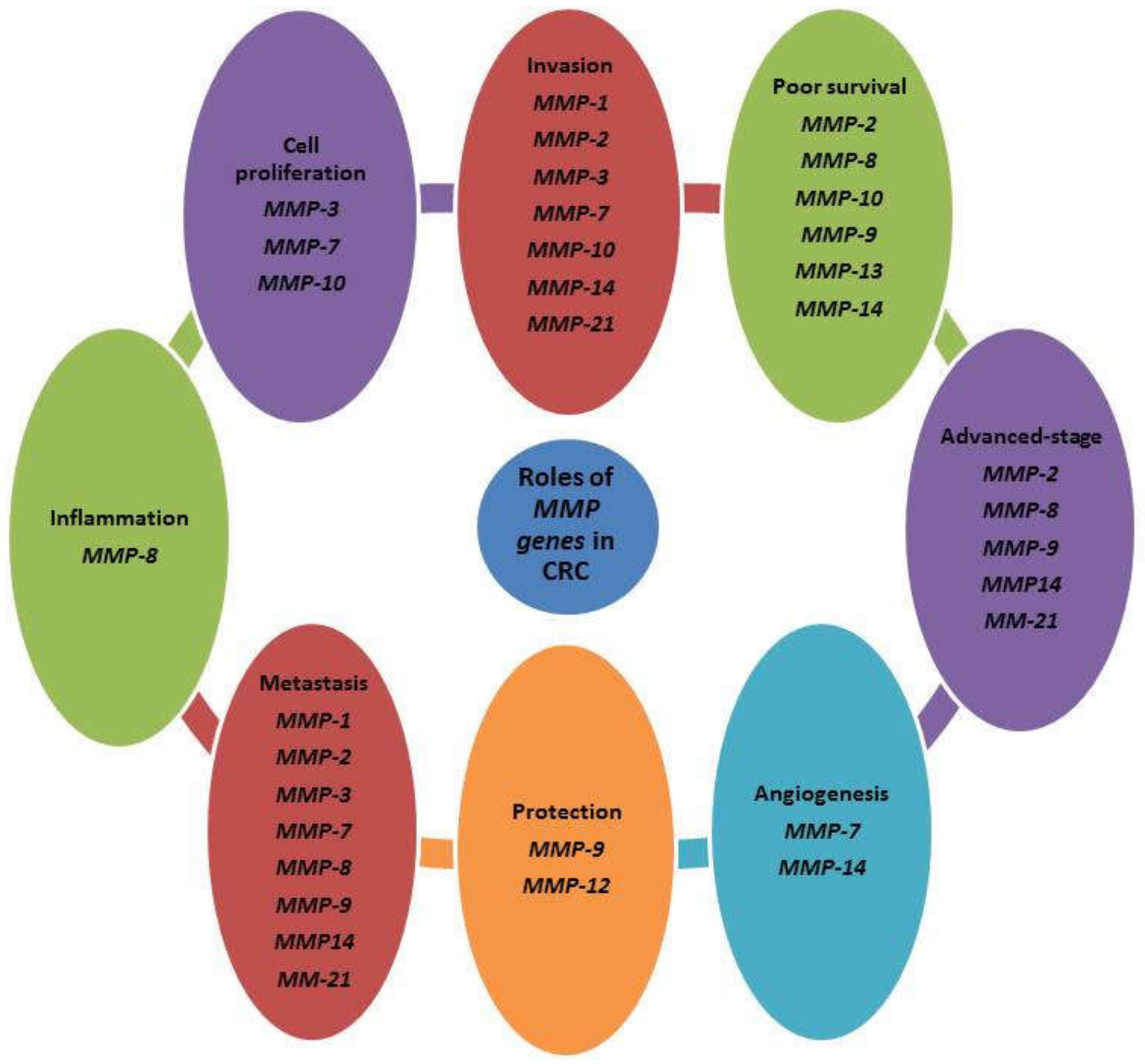
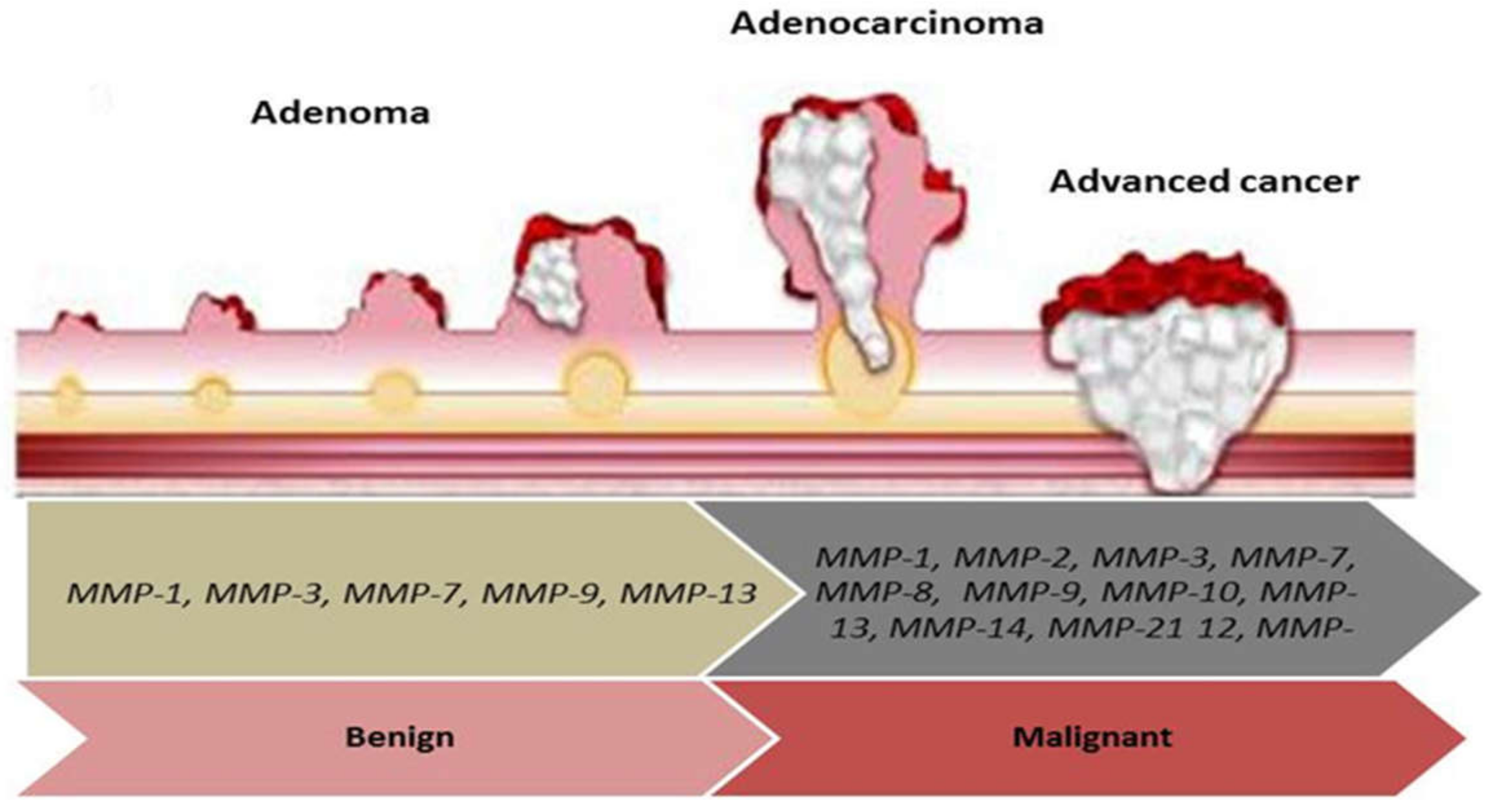
4. The Function of MMPs in Colorectal Polyps and Cancer
4.1. MMP-1, MMP-13, and MMP-8 (Collagenases)
4.2. MMP-2 and MMP-9 (Gelatinase)
4.3. MMP-3, MMP-10 (Stromelysin)
4.4. MMP-7 (Matrilysin)
4.5. MMP-12 (Metalloelastase)
4.6. MMP-21 (XMMP)
4.7. MMP-14 (MT1-MMP)
5. The Effects of Polymorphisms of MMP Genes on Colorectal Carcinogenesis
6. Targeting MMPs in CRC Treatment
6.1. Pharmacological Inhibition
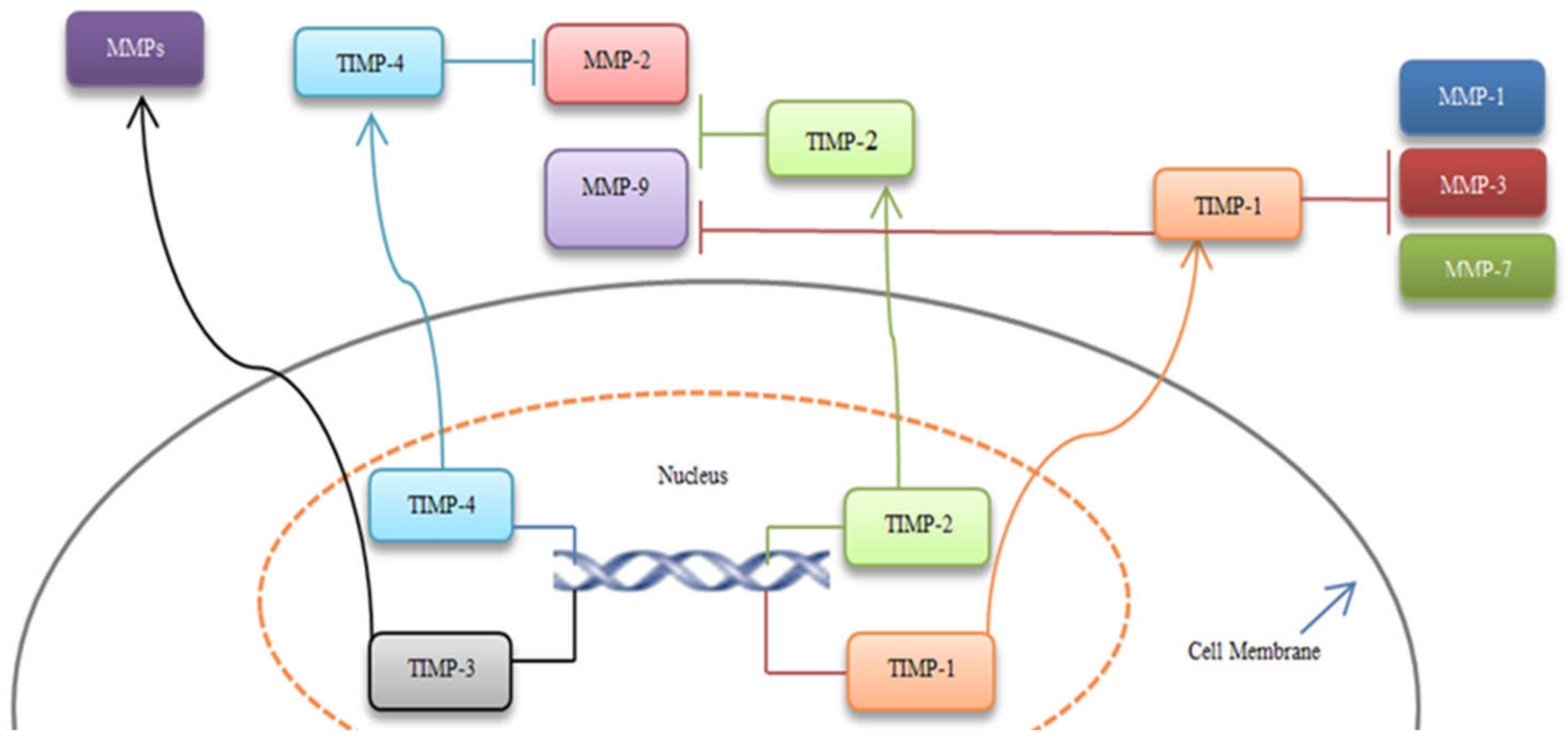
6.2. Inhibition of MMPs by TIMPs
6.3. MMPs Regulation by microRNA
6.4. MMPs Regulation by Long Non-Coding RNAs
7. Conclusions
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Sung, H.; Ferlay, J.; Siegel, R.L.; Laversanne, M.; Soerjomataram, I.; Jemal, A.; Bray, F. Global Cancer Statistics 2020: GLOBOCAN Estimates of Incidence and Mortality Worldwide for 36 Cancers in 185 Countries. CA Cancer J. Clin. 2021, 71, 209–249. [Google Scholar] [CrossRef] [PubMed]
- Vogelstein, B.; Fearon, E.R.; Hamilton, S.R.; Kern, S.E.; Preisinger, A.C.; Leppert, M.; Nakamura, Y.; White, R.; Smits, A.M.; Bos, J.L. Genetic alterations during colorectal-tumor development. N. Engl. J. Med. 1988, 319, 525–532. [Google Scholar] [CrossRef] [PubMed]
- Kasi, A.; Handa, S.; Bhatti, S.; Umar, S.; Bansal, A.; Sun, W. Molecular Pathogenesis and Classification of Colorectal Carcinoma. Curr. Colorectal Cancer Rep. 2020, 16, 97–106. [Google Scholar] [CrossRef] [PubMed]
- Huck, M.B.; Bohl, J.L. Colonic Polyps: Diagnosis and Surveillance. Clin. Colon Rectal Surg. 2016, 29, 296–305. [Google Scholar] [CrossRef]
- Smit, W.L.; Spaan, C.N.; de Boer, R.J.; Ramesh, P.; Garcia, T.M.; Meijer, B.J.; Vermeulen, J.L.M.; Lezzerini, M.; MacInnes, A.W.; Koster, J.; et al. Driver mutations of the adenoma-carcinoma sequence govern the intestinal epithelial global translational capacity. Proc. Natl. Acad. Sci. USA 2020, 117, 25560. [Google Scholar] [CrossRef]
- Wang, K.; Zheng, J.; Yu, J.; Wu, Y.; Guo, J.; Xu, Z.; Sun, X. Knockdown of MMP--1 inhibits the progression of colorectal cancer by suppressing the PI3K/Akt/c--myc signaling pathway and EMT. Oncol. Rep. 2020, 43, 1103–1112. [Google Scholar] [CrossRef]
- Yuan, X.; Sun, X.; Shi, X.; Wang, H.; Wu, G.; Jiang, C.; Yu, D.; Zhang, W.; Xue, B.; Ding, Y. USP39 promotes colorectal cancer growth and metastasis through the Wnt/β-catenin pathway. Oncol. Rep. 2017, 37, 2398–2404. [Google Scholar] [CrossRef]
- Quintero-Fabián, S.; Arreola, R.; Becerril-Villanueva, E.; Torres-Romero, J.C.; Arana-Argáez, V.; Lara-Riegos, J.; Ramírez-Camacho, M.A.; Alvarez-Sánchez, M.E. Role of Matrix Metalloproteinases in Angiogenesis and Cancer. Front. Oncol. 2019, 9, 1370. [Google Scholar] [CrossRef]
- Yadav, L.; Puri, N.; Rastogi, V.; Satpute, P.; Ahmad, R.; Kaur, G. Role of Matrix Metalloproteinases in Angiogenesis and Cancer. Asian Pac. J. Cancer Prev. 2014, 15, 1085–1091. [Google Scholar] [CrossRef]
- Ligi, D.; Mannello, F. Do matrix metalloproteinases represent reliable circulating biomarkers in colorectal cancer? Br. J. Cancer 2016, 115, 633–634. [Google Scholar] [CrossRef][Green Version]
- Yeh, Y.-C.; Sheu, B.-S. Matrix metalloproteinases and their inhibitors in the gastrointestinal cancers: Current knowledge and clinical potential. Met. Med. 2014, 1, 3–13. [Google Scholar] [CrossRef][Green Version]
- Wu, Z.; Liu, Z.; Ge, W.; Shou, J.; You, L.; Pan, H.; Han, W. Analysis of potential genes and pathways associated with the colorectal normal mucosa-adenoma-carcinoma sequence. Cancer Med. 2018, 7, 2555–2566. [Google Scholar] [CrossRef]
- Bertelson, N.L.; Kalkbrenner, K.A.; Merchea, A.; Dozois, E.J.; Landmann, R.G.; De Petris, G.; Young-Fadok, T.M.; Etzioni, D.A. Colectomy for Endoscopically Unresectable Polyps: How Often Is It Cancer? Dis. Colon Rectum 2012, 55, 1111–1116. [Google Scholar] [CrossRef]
- Shussman, N.; Wexner, S.D. Colorectal polyps and polyposis syndromes. Gastroenterol. Rep. 2014, 2, 1–15. [Google Scholar] [CrossRef]
- Liljegren, A.; Lindblom, A.; Rotstein, S.; Nilsson, B.; Rubio, C.; Jaramillo, E. Prevalence and incidence of hyperplastic polyps and adenomas in familial colorectal cancer: Correlation between the two types of colon polyps. Gut 2003, 52, 1140–1147. [Google Scholar] [CrossRef]
- Malki, A.; ElRuz, R.A.; Gupta, I.; Allouch, A.; Vranic, S.; Al Moustafa, A.E. Molecular Mechanisms of Colon Cancer Progression and Metastasis: Recent Insights and Advancements. Int. J. Mol. Sci. 2020, 22, 130. [Google Scholar] [CrossRef]
- Kato, S.; Lippman, S.M.; Flaherty, K.T.; Kurzrock, R. The Conundrum of Genetic “Drivers” in Benign Conditions. J. Natl. Cancer Inst. 2016, 108, djw036. [Google Scholar] [CrossRef]
- Mustjoki, S.; Young, N.S. Somatic Mutations in “Benign” Disease. N. Engl. J. Med. 2021, 384, 2039–2052. [Google Scholar] [CrossRef]
- Nojadeh, J.N.; Behrouz Sharif, S.; Sakhinia, E. Microsatellite instability in colorectal cancer. EXCLI J. 2018, 17, 159–168. [Google Scholar] [CrossRef]
- Arabsorkhi, Z.; Sadeghi, H.; Gharib, E.; Rejali, L.; Asadzadeh-Aghdaei, H.; Nazemalhosseini-Mojarad, E. Can hypoxia-inducible factor-1α overexpression discriminate human colorectal cancers with different microsatellite instability? Genes Genet. Syst. 2021, 96, 1–6. [Google Scholar] [CrossRef]
- Pino, M.S.; Chung, D.C. The chromosomal instability pathway in colon cancer. Gastroenterology 2010, 138, 2059–2072. [Google Scholar] [CrossRef]
- Colussi, D.; Brandi, G.; Bazzoli, F.; Ricciardiello, L. Molecular pathways involved in colorectal cancer: Implications for disease behavior and prevention. Int. J. Mol. Sci. 2013, 14, 16365–16385. [Google Scholar] [CrossRef]
- Grady, W.M.; Markowitz, S.D. The molecular pathogenesis of colorectal cancer and its potential application to colorectal cancer screening. Dig. Dis. Sci. 2015, 60, 762–772. [Google Scholar] [CrossRef]
- Sadanandam, A.; Lyssiotis, C.A.; Homicsko, K.; Collisson, E.A.; Gibb, W.J.; Wullschleger, S.; Ostos, L.C.; Lannon, W.A.; Grotzinger, C.; Del Rio, M.; et al. A colorectal cancer classification system that associates cellular phenotype and responses to therapy. Nat. Med. 2013, 19, 619–625. [Google Scholar] [CrossRef]
- Guinney, J.; Dienstmann, R.; Wang, X.; de Reyniès, A.; Schlicker, A.; Soneson, C.; Marisa, L.; Roepman, P.; Nyamundanda, G.; Angelino, P.; et al. The consensus molecular subtypes of colorectal cancer. Nat. Med. 2015, 21, 1350–1356. [Google Scholar] [CrossRef]
- Thanki, K.; Nicholls, M.E.; Gajjar, A.; Senagore, A.J.; Qiu, S.; Szabo, C.; Hellmich, M.R.; Chao, C. Consensus Molecular Subtypes of Colorectal Cancer and their Clinical Implications. Int. Biol. Biomed. J. 2017, 3, 105–111. [Google Scholar]
- McCorry, A.M.; Loughrey, M.B.; Longley, D.B.; Lawler, M.; Dunne, P.D. Epithelial-to-mesenchymal transition signature assessment in colorectal cancer quantifies tumour stromal content rather than true transition. J. Pathol. 2018, 246, 422–426. [Google Scholar] [CrossRef]
- Isella, C.; Brundu, F.; Bellomo, S.E.; Galimi, F.; Zanella, E.; Porporato, R.; Petti, C.; Fiori, A.; Orzan, F.; Senetta, R.; et al. Selective analysis of cancer-cell intrinsic transcriptional traits defines novel clinically relevant subtypes of colorectal cancer. Nat. Commun. 2017, 8, 15107. [Google Scholar] [CrossRef]
- Cui, N.; Hu, M.; Khalil, R.A. Biochemical and biological attributes of matrix metalloproteinases. Prog. Mol. Biol. Transl. Sci. 2017, 147, 1–73. [Google Scholar] [CrossRef]
- Morgunova, E.; Tuuttila, A.; Bergmann, U.; Isupov, M.; Lindqvist, Y.; Schneider, G.; Tryggvason, K. Structure of human pro-matrix metalloproteinase-2: Activation mechanism revealed. Science 1999, 284, 1667–1670. [Google Scholar] [CrossRef]
- Laronha, H.; Caldeira, J. Structure and function of human matrix metalloproteinases. Cells 2020, 9, 1076. [Google Scholar] [CrossRef] [PubMed]
- Sternlicht, M.D.; Werb, Z. How matrix metalloproteinases regulate cell behavior. Annu. Rev. Cell. Dev. Biol. 2001, 17, 463–516. [Google Scholar] [CrossRef] [PubMed]
- Baker, A.H.; Edwards, D.R.; Murphy, G. Metalloproteinase inhibitors: Biological actions and therapeutic opportunities. J. Cell Sci. 2002, 115, 3719–3727. [Google Scholar] [CrossRef] [PubMed]
- Hua, H.; Li, M.; Luo, T.; Yin, Y.; Jiang, Y. Matrix metalloproteinases in tumorigenesis: An evolving paradigm. Cell Mol. Life Sci. 2011, 68, 3853–3868. [Google Scholar] [CrossRef]
- Alaseem, A.; Alhazzani, K.; Dondapati, P.; Alobid, S.; Bishayee, A.; Rathinavelu, A. Matrix Metalloproteinases: A challenging paradigm of cancer management. Semin. Cancer Biol. 2019, 56, 100–115. [Google Scholar] [CrossRef]
- Gonzalez-Avila, G.; Sommer, B.; Mendoza-Posada, D.A.; Ramos, C.; Garcia-Hernandez, A.A.; Falfan-Valencia, R. Matrix metalloproteinases participation in the metastatic process and their diagnostic and therapeutic applications in cancer. Crit. Rev. Oncol. Hematol. 2019, 137, 57–83. [Google Scholar] [CrossRef]
- Herszényi, L.; Hritz, I.; Lakatos, G.; Varga, M.Z.; Tulassay, Z. The behavior of matrix metalloproteinases and their inhibitors in colorectal cancer. Int. J. Mol. Sci. 2012, 13, 13240–13263. [Google Scholar] [CrossRef]
- Huang, X.; Lan, Y.; Li, E.; Li, J.; Deng, Q.; Deng, X. Diagnostic values of MMP-7, MMP-9, MMP-11, TIMP-1, TIMP-2, CEA, and CA19-9 in patients with colorectal cancer. J. Int. Med. Res. 2021, 49, 1–11. [Google Scholar] [CrossRef]
- Zhou, X.; Yang, D.; Ding, X.; Xu, P. Clinical value of microRNA-135a and MMP-13 in colon cancer. Oncol. Lett. 2021, 22, 583. [Google Scholar] [CrossRef]
- Rasool, M.; Malik, A.; Waquar, S.; Ain, Q.T.; Rasool, R.; Asif, M.; Anfinan, N.; Haque, A.; Alam, H.; Ahmed, S.; et al. Assessment of clinical variables as predictive markers in the development and progression of colorectal cancer. Bioengineered 2021, 12, 2288–2298. [Google Scholar] [CrossRef]
- Barabás, L.; Hritz, I.; István, G.; Tulassay, Z.; Herszényi, L. The Behavior of MMP-2, MMP-7, MMP-9, and Their Inhibitors TIMP-1 and TIMP-2 in Adenoma-Colorectal Cancer Sequence. Dig. Dis. 2021, 39, 217–224. [Google Scholar] [CrossRef]
- Hsieh, S.L.; Hsieh, S.; Lai, P.Y.; Wang, J.J.; Li, C.C.; Wu, C.C. Carnosine Suppresses Human Colorectal Cell Migration and Intravasation by Regulating EMT and MMP Expression. Am. J. Chin. Med. 2019, 47, 477–494. [Google Scholar] [CrossRef]
- Kıyak, R.; Keles, D.; Bengi, G.; Yalcin, M.; Topalak, Ö.; Oktay, G. The Importance of Fecal and Plasma CEA, COX-2, MMP-7, and TIMP-1 in the Diagnosis of Colorectal Cancer. J. Basic Clin. Health Sci. 2018, 2, 7–14. [Google Scholar] [CrossRef]
- Eiró, N.; Gonzalez, L.; Cid, S.; Andicoechea, A.; Vizoso, F. Matrix metalloproteases expression in different histological types of colorectal polyps. Rev. Esp. Enferm. Dig. 2017, 109, 414–420. [Google Scholar] [CrossRef]
- Pezeshkian, Z.; Forouzesh, F.; Peyravian, N.; Yaghoob-Taleghani, M.; Asadzadeh-Aghdaei, H.; Zali, M.; Nazemalhosseini-Mojarad, E. Clinicopathological correlations of VEGF-A and MMP-7 genes expression in different types of colorectal adenoma polyps. WCRJ 2017, 4, e978. [Google Scholar]
- Wernicke, A.-K.; Churin, Y.; Sheridan, D.; Windhorst, A.; Tschuschner, A.; Gattenlöhner, S.; Roderfeld, M.; Roeb, E. Matrix metalloproteinase-13 refines pathological staging of precancerous colorectal lesions. Oncotarget 2016, 7, 73552–73557. [Google Scholar] [CrossRef][Green Version]
- Gimeno-García, A.Z.; Triñanes, J.; Quintero, E.; Salido, E.; Nicolás-Pérez, D.; Adrián-de-Ganzo, Z.; Alarcón-Fernández, O.; Abrante, B.; Romero, R.; Carrillo, M.; et al. Plasma matrix metalloproteinase 9 as an early surrogate biomarker of advanced colorectal neoplasia. Gastroenterol. Hepatol. 2016, 39, 433–441. [Google Scholar] [CrossRef]
- Annaházi, A.; Ábrahám, S.; Farkas, K.; Rosztóczy, A.; Inczefi, O.; Földesi, I.; Szűcs, M.; Rutka, M.; Theodorou, V.; Eutamene, H.; et al. A pilot study on faecal MMP-9: A new noninvasive diagnostic marker of colorectal cancer. Br. J. Cancer 2016, 114, 787–792. [Google Scholar] [CrossRef]
- Klupp, F.; Neumann, L.; Kahlert, C.; Diers, J.; Halama, N.; Franz, C.; Schmidt, T.; Koch, M.; Weitz, J.; Schneider, M.; et al. Serum MMP7, MMP10 and MMP12 level as negative prognostic markers in colon cancer patients. BMC Cancer 2016, 16, 494. [Google Scholar] [CrossRef]
- Otero-Estévez, O.; De Chiara, L.; Rodríguez-Girondo, M.; Rodríguez-Berrocal, F.J.; Cubiella, J.; Castro, I.; Hernández, V.; Martínez-Zorzano, V.S. Serum matrix metalloproteinase-9 in colorectal cancer family-risk population screening. Sci. Rep. 2015, 5, 13030. [Google Scholar] [CrossRef]
- Bengi, G.; Keles, D.; Topalak, Ö.; Yalçin, M.; Kiyak, R.; Oktay, G. Expressions of TIMP-1, COX-2 and MMP-7 in Colon Polyp and Colon Cancer. Euroasian J. Hepatogastroenterol. 2015, 5, 74–79. [Google Scholar] [CrossRef]
- Odabasi, M.; Yesil, A.; Ozkara, S.; Paker, N.; Ozkan, S.; Eris, C.; Yildiz, M.K.; Abuoglu, H.H.; Gunay, E.; Tekeşin, K. Role of human neutrophil gelatinase associated lipocalin (NGAL) and Matrix Metalloproteinase-9 (MMP-9) overexpression in neoplastic colon polyps. Int. J. Clin. Exp. Med. 2014, 7, 2804–2811. [Google Scholar]
- Qasim, B.J.; Ali, H.H.; Hussein, A.G. Immunohistochemical expression of matrix metalloproteinase-7 in human colorectal adenomas using specified automated cellular image analysis system: A clinicopathological study. Saudi J. Gastroenterol. 2013, 19, 23–27. [Google Scholar] [CrossRef]
- Sheth, R.A.; Kunin, A.; Stangenberg, L.; Sinnamon, M.; Hung, K.E.; Kucherlapati, R.; Mahmood, U. In vivo optical molecular imaging of matrix metalloproteinase activity following celecoxib therapy for colorectal cancer. Mol. Imaging 2012, 11, 417–425. [Google Scholar] [CrossRef] [PubMed]
- Murnane, M.J.; Cai, J.; Shuja, S.; McAneny, D.; Klepeis, V.; Willett, J.B. Active MMP-2 effectively identifies the presence of colorectal cancer. Int. J. Cancer 2009, 125, 2893–2902. [Google Scholar] [CrossRef] [PubMed]
- Jeffery, N.; McLean, M.H.; El-Omar, E.M.; Murray, G.I. The matrix metalloproteinase/tissue inhibitor of matrix metalloproteinase profile in colorectal polyp cancers. Histopathology 2009, 54, 820–828. [Google Scholar] [CrossRef] [PubMed]
- Lièvre, A.; Milet, J.; Carayol, J.; Le Corre, D.; Milan, C.; Pariente, A.; Nalet, B.; Lafon, J.; Faivre, J.; Bonithon-Kopp, C.; et al. Genetic polymorphisms of MMP1, MMP3 and MMP7 gene promoter and risk of colorectal adenoma. BMC Cancer 2006, 6, 270. [Google Scholar] [CrossRef] [PubMed]
- Tutton, M.G.; George, M.L.; Eccles, S.A.; Burton, S.; Swift, R.I.; Abulafi, A.M. Use of plasma MMP-2 and MMP-9 levels as a surrogate for tumour expression in colorectal cancer patients. Int. J. Cancer 2003, 107, 541–550. [Google Scholar] [CrossRef]
- Jonsson, A.; Falk, P.; Angenete, E.; Hjalmarsson, C.; Ivarsson, M.-L. Plasma MMP-1 Expression as a Prognostic Factor in Colon Cancer. J. Surg. Res. 2021, 266, 254–260. [Google Scholar] [CrossRef]
- Liang, Y.; Lv, Z.; Huang, G.; Qin, J.; Li, H.; Nong, F.; Wen, B. Prognostic significance of abnormal matrix collagen remodeling in colorectal cancer based on histologic and bioinformatics analysis. Oncol. Rep. 2020, 44, 1671–1685. [Google Scholar] [CrossRef]
- Sunami, E.; Tsuno, N.; Osada, T.; Saito, S.; Kitayama, J.; Tomozawa, S.; Tsuruo, T.; Shibata, Y.; Muto, T.; Nagawa, H. MMP-1 is a prognostic marker for hematogenous metastasis of colorectal cancer. Oncologist 2000, 5, 108–114. [Google Scholar] [CrossRef]
- Yamada, T.; Oshima, T.; Yoshihara, K.; Tamura, S.; Kanazawa, A.; Inagaki, D.; Yamamoto, N.; Sato, T.; Fujii, S.; Numata, K.; et al. Overexpression of MMP-13 gene in colorectal cancer with liver metastasis. Anticancer Res. 2010, 30, 2693–2699. [Google Scholar]
- Leeman, M.F.; McKay, J.A.; Murray, G.I. Matrix metalloproteinase 13 activity is associated with poor prognosis in colorectal cancer. J. Clin. Pathol. 2002, 55, 758–762. [Google Scholar] [CrossRef]
- Merchant, N.; Chalikonda, G.; Nagaraju, G.P. Role of Matrix Metalloproteinases in Colorectal Cancer. In Theranostics Approaches to Gastric and Colon Cancer; Springer: Singapore, 2020; pp. 49–59. [Google Scholar] [CrossRef]
- Yan, Q.; Yuan, Y.; Yankui, L.; Jingjie, F.; Linfang, J.; Yong, P.; Dong, H.; Xiaowei, Q. The Expression and Significance of CXCR5 and MMP-13 in Colorectal Cancer. Cell Biochem. Biophys. 2015, 73, 253–259. [Google Scholar] [CrossRef]
- Korpi, J.T.; Kervinen, V.; Mäklin, H.; Väänänen, A.; Lahtinen, M.; Läärä, E.; Ristimäki, A.; Thomas, G.; Ylipalosaari, M.; Aström, P.; et al. Collagenase-2 (matrix metalloproteinase-8) plays a protective role in tongue cancer. Br. J. Cancer 2008, 98, 766–775. [Google Scholar] [CrossRef]
- Balbín, M.; Fueyo, A.; Tester, A.M.; Pendás, A.M.; Pitiot, A.S.; Astudillo, A.; Overall, C.M.; Shapiro, S.D.; López-Otín, C. Loss of collagenase-2 confers increased skin tumor susceptibility to male mice. Nat. Genet. 2003, 35, 252–257. [Google Scholar] [CrossRef]
- Sirniö, P.; Tuomisto, A.; Tervahartiala, T.; Sorsa, T.; Klintrup, K.; Karhu, T.; Herzig, K.-H.; Mäkelä, J.; Karttunen, T.J.; Salo, T.; et al. High-serum MMP-8 levels are associated with decreased survival and systemic inflammation in colorectal cancer. Br. J. Cancer 2018, 119, 213–219. [Google Scholar] [CrossRef]
- Beutel, B.; Song, J.; Konken, C.P.; Korpos, E.; Schinor, B.; Gerwien, H.; Vidyadharan, R.; Burmeister, M.; Li, L.; Haufe, G.; et al. New in Vivo Compatible Matrix Metalloproteinase (MMP)-2 and MMP-9 Inhibitors. Bioconjugate Chem. 2018, 29, 3715–3725. [Google Scholar] [CrossRef]
- Murnane, M.J.; Cai, J.; Shuja, S.; McAneny, D.; Willett, J.B. Active matrix metalloproteinase-2 activity discriminates colonic mucosa, adenomas with and without high-grade dysplasia, and cancers. Hum. Pathol. 2011, 42, 688–701. [Google Scholar] [CrossRef]
- Gimeno-García, A.Z.; Santana-Rodríguez, A.; Jiménez, A.; Parra-Blanco, A.; Nicolás-Pérez, D.; Paz-Cabrera, C.; Díaz-González, F.; Medina, C.; Díaz-Flores, L.; Quintero, E. Up-regulation of gelatinases in the colorectal adenoma-carcinoma sequence. Eur. J. Cancer 2006, 42, 3246–3252. [Google Scholar] [CrossRef]
- Salem, N.; Kamal, I.; Al-Maghrabi, J.; Abuzenadah, A.; Peer-Zada, A.A.; Qari, Y.; Al-Ahwal, M.; Al-Qahtani, M.; Buhmeida, A. High expression of matrix metalloproteinases: MMP-2 and MMP-9 predicts poor survival outcome in colorectal carcinoma. Future Oncol. 2016, 12, 323–331. [Google Scholar] [CrossRef] [PubMed]
- Langenskiöld, M.; Holmdahl, L.; Falk, P.; Ivarsson, M.-L. Increased plasma MMP-2 protein expression in lymph node-positive patients with colorectal cancer. Int. J. Colorectal Dis. 2005, 20, 245–252. [Google Scholar] [CrossRef] [PubMed]
- Kryczka, J.; Stasiak, M.; Dziki, L.; Mik, M.; Dziki, A.; Cierniewski, C.S. Matrix metalloproteinase-2 cleavage of the β1 integrin ectodomain facilitates colon cancer cell motility. J. Biol. Chem. 2012, 287, 36556–36566. [Google Scholar] [CrossRef] [PubMed]
- Takeuchi, T.; Hisanaga, M.; Nagao, M.; Ikeda, N.; Fujii, H.; Koyama, F.; Mukogawa, T.; Matsumoto, H.; Kondo, S.; Takahashi, C.; et al. The membrane-anchored matrix metalloproteinase (MMP) regulator RECK in combination with MMP-9 serves as an informative prognostic indicator for colorectal cancer. Clin. Cancer Res. 2004, 10, 5572–5579. [Google Scholar] [CrossRef] [PubMed]
- Chen, H.; Ye, Y.; Yang, Y.; Zhong, M.; Gu, L.; Han, Z.; Qiu, J.; Liu, Z.; Qiu, X.; Zhuang, G. TIPE-mediated up-regulation of MMP-9 promotes colorectal cancer invasion and metastasis through MKK-3/p38/NF-κB pro-oncogenic signaling pathway. Signal Transduct. Target. Ther. 2020, 5, 163. [Google Scholar] [CrossRef] [PubMed]
- Garg, P.; Jeppsson, S.; Dalmasso, G.; Ghaleb, A.M.; McConnell, B.B.; Yang, V.W.; Gewirtz, A.T.; Merlin, D.; Sitaraman, S.V. Notch1 regulates the effects of matrix metalloproteinase-9 on colitis-associated cancer in mice. Gastroenterology 2011, 141, 1381–1392. [Google Scholar] [CrossRef]
- Garg, P.; Sarma, D.; Jeppsson, S.; Patel, N.R.; Gewirtz, A.T.; Merlin, D.; Sitaraman, S.V. Matrix metalloproteinase-9 functions as a tumor suppressor in colitis-associated cancer. Cancer Res. 2010, 70, 792–801. [Google Scholar] [CrossRef]
- Walter, L.; Canup, B.; Pujada, A.; Bui, T.A.; Arbasi, B.; Laroui, H.; Merlin, D.; Garg, P. Matrix metalloproteinase 9 (MMP9) limits reactive oxygen species (ROS) accumulation and DNA damage in colitis-associated cancer. Cell Death Dis. 2020, 11, 767. [Google Scholar] [CrossRef] [PubMed]
- Wei, Z.; Cao, S.; Liu, S.; Yao, Z.; Sun, T.; Li, Y.; Li, J.; Zhang, D.; Zhou, Y. Could gut microbiota serve as prognostic biomarker associated with colorectal cancer patients’ survival? A pilot study on relevant mechanism. Oncotarget 2016, 7, 46158–46172. [Google Scholar] [CrossRef]
- Sipos, F.; Germann, T.M.; Wichmann, B.; Galamb, O.; Spisák, S.; Krenács, T.; Tulassay, Z.; Molnár, B.; Műzes, G. MMP3 and CXCL1 are potent stromal protein markers of dysplasia-carcinoma transition in sporadic colorectal cancer. Eur. J. Cancer Prev. 2014, 23, 336–343. [Google Scholar] [CrossRef]
- Işlekel, H.; Oktay, G.; Terzi, C.; Canda, A.E.; Füzün, M.; Küpelioğlu, A. Matrix metalloproteinase-9,-3 and tissue inhibitor of matrix metalloproteinase-1 in colorectal cancer: Relationship to clinicopathological variables. Cell Biochem. Funct. 2007, 25, 433–441. [Google Scholar] [CrossRef] [PubMed]
- Jin, X.; Yagi, M.; Akiyama, N.; Hirosaki, T.; Higashi, S.; Lin, C.Y.; Dickson, R.B.; Kitamura, H.; Miyazaki, K. Matriptase activates stromelysin (MMP-3) and promotes tumor growth and angiogenesis. Cancer Sci. 2006, 97, 1327–1334. [Google Scholar] [CrossRef] [PubMed]
- Inuzuka, K.; Ogata, Y.; Nagase, H.; Shirouzu, K. Significance of coexpression of urokinase-type plasminogen activator, and matrix metalloproteinase 3 (stromelysin) and 9 (gelatinase B) in colorectal carcinoma. J. Surg. Res. 2000, 93, 211–218. [Google Scholar] [CrossRef] [PubMed]
- Batra, J.; Robinson, J.; Soares, A.S.; Fields, A.P.; Radisky, D.C.; Radisky, E.S. Matrix metalloproteinase-10 (MMP-10) interaction with tissue inhibitors of metalloproteinases TIMP-1 and TIMP-2: Binding studies and crystal structure. J. Biol. Chem. 2012, 287, 15935–15946. [Google Scholar] [CrossRef]
- Surlin, V.; Ioana, M.; Pleşea, I.E. Genetic patterns of metalloproteinases and their tissular inhibitors—Clinicopathologic and prognostic significance in colorectal cancer. Rom. J. Morphol. Embryol. 2011, 52, 231–236. [Google Scholar]
- Asadzadeh Aghdaei, H.; Pezeshkian, Z.; Abdollahpour-Alitappeh, M.; Nazemalhosseini Mojarad, E.; Zali, M.R. The Role of Angiogenesis in Colorectal Polyps and Cancer, a Review. Med. Lab. J. 2018, 12, 1–6. [Google Scholar] [CrossRef]
- Ii, M.; Yamamoto, H.; Adachi, Y.; Maruyama, Y.; Shinomura, Y. Role of matrix metalloproteinase-7 (matrilysin) in human cancer invasion, apoptosis, growth, and angiogenesis. Exp. Biol. Med. 2006, 231, 20–27. [Google Scholar] [CrossRef]
- Cheng, K.; Xie, G.; Raufman, J.P. Matrix metalloproteinase-7-catalyzed release of HB-EGF mediates deoxycholyltaurine-induced proliferation of a human colon cancer cell line. Biochem. Pharmacol. 2007, 73, 1001–1012. [Google Scholar] [CrossRef]
- Xie, G.; Cheng, K.; Shant, J.; Raufman, J.P. Acetylcholine-induced activation of M3 muscarinic receptors stimulates robust matrix metalloproteinase gene expression in human colon cancer cells. Am. J. Physiol. Gastrointest. Liver Physiol. 2009, 296, G755–G763. [Google Scholar] [CrossRef]
- Decock, J.; Thirkettle, S.; Wagstaff, L.; Edwards, D.R. Matrix metalloproteinases: Protective roles in cancer. J. Cell Mol. Med. 2011, 15, 1254–1265. [Google Scholar] [CrossRef]
- Asano, T.; Tada, M.; Cheng, S.; Takemoto, N.; Kuramae, T.; Abe, M.; Takahashi, O.; Miyamoto, M.; Hamada, J.; Moriuchi, T.; et al. Prognostic values of matrix metalloproteinase family expression in human colorectal carcinoma. J. Surg. Res. 2008, 146, 32–42. [Google Scholar] [CrossRef]
- Yang, W.; Arii, S.; Gorrin-Rivas, M.J.; Mori, A.; Onodera, H.; Imamura, M. Human macrophage metalloelastase gene expression in colorectal carcinoma and its clinicopathologic significance. Cancer 2001, 91, 1277–1283. [Google Scholar] [CrossRef]
- Shi, H.; Xu, J.M.; Hu, N.Z.; Wang, X.L.; Mei, Q.; Song, Y.L. Transfection of mouse macrophage metalloelastase gene into murine CT-26 colon cancer cells suppresses orthotopic tumor growth, angiogenesis and vascular endothelial growth factor expression. Cancer Lett. 2006, 233, 139–150. [Google Scholar] [CrossRef]
- Xu, Z.; Shi, H.; Li, Q.; Mei, Q.; Bao, J.; Shen, Y.; Xu, J. Mouse macrophage metalloelastase generates angiostatin from plasminogen and suppresses tumor angiogenesis in murine colon cancer. Oncol. Rep. 2008, 20, 81–88. [Google Scholar] [CrossRef]
- Beurden, P.; Von den Hoff, J. Zymographic techniques for the analysis of matrix metalloproteinases and their inhibitors. BioTechniques 2005, 38, 73–83. [Google Scholar] [CrossRef]
- Wu, T.; Li, Y.; Liu, X.; Lu, J.; He, X.; Wang, Q.; Li, J.; Du, X. Identification of high-risk stage II and stage III colorectal cancer by analysis of MMP-21 expression. J. Surg. Oncol. 2011, 104, 787–791. [Google Scholar] [CrossRef]
- Huang, Y.; Li, W.; Chu, D.; Zheng, J.; Ji, G.; Li, M.; Zhang, H.; Wang, W.; Du, J.; Li, J. Overexpression of matrix metalloproteinase-21 is associated with poor overall survival of patients with colorectal cancer. J. Gastrointest. Surg. 2011, 15, 1188–1194. [Google Scholar] [CrossRef]
- Zhang, J.; Pan, Q.; Yan, W.; Wang, Y.; He, X.; Zhao, Z. Overexpression of MMP21 and MMP28 is associated with gastric cancer progression and poor prognosis. Oncol. Lett. 2018, 15, 7776–7782. [Google Scholar] [CrossRef]
- Pahwa, S.; Stawikowski, M.J.; Fields, G.B. Monitoring and Inhibiting MT1-MMP during Cancer Initiation and Progression. Cancers 2014, 6, 416–435. [Google Scholar] [CrossRef]
- Devy, L.; Huang, L.; Naa, L.; Yanamandra, N.; Pieters, H.; Frans, N.; Chang, E.; Tao, Q.; Vanhove, M.; Lejeune, A.; et al. Selective Inhibition of Matrix Metalloproteinase-14 Blocks Tumor Growth, Invasion, and Angiogenesis. Cancer Res. 2009, 69, 1517–1526. [Google Scholar] [CrossRef]
- Duan, F.; Peng, Z.; Yin, J.; Yang, Z.; Shang, J. Expression of MMP-14 and prognosis in digestive system carcinoma: A meta-analysis and databases validation. J. Cancer 2020, 11, 1141–1150. [Google Scholar] [CrossRef]
- Yang, B.; Gao, J.; Rao, Z.; Shen, Q. Clinicopathological and prognostic significance of α5β1-integrin and MMP-14 expressions in colorectal cancer. Neoplasma 2013, 60, 254–261. [Google Scholar] [CrossRef] [PubMed]
- Cui, G.; Cai, F.; Ding, Z.; Gao, L. MMP14 predicts a poor prognosis in patients with colorectal cancer. Hum. Pathol. 2019, 83, 36–42. [Google Scholar] [CrossRef] [PubMed]
- Claesson-Welsh, L. How the matrix metalloproteinase MMP14 contributes to the progression of colorectal cancer. J. Clin. Investig. 2020, 130, 1093–1095. [Google Scholar] [CrossRef] [PubMed]
- Decock, J.; Paridaens, R.; Ye, S. Genetic polymorphisms of matrix metalloproteinases in lung, breast and colorectal cancer. Clin. Genet. 2008, 73, 197–211. [Google Scholar] [CrossRef] [PubMed]
- Langers, A.M.; Verspaget, H.W.; Hommes, D.W.; Sier, C.F. Single-nucleotide polymorphisms of matrix metalloproteinases and their inhibitors in gastrointestinal cancer. World J. Gastrointest. Oncol. 2011, 3, 79–98. [Google Scholar] [CrossRef] [PubMed]
- Hinoda, Y.; Okayama, N.; Takano, N.; Fujimura, K.; Suehiro, Y.; Hamanaka, Y.; Hazama, S.; Kitamura, Y.; Kamatani, N.; Oka, M. Association of functional polymorphisms of matrix metalloproteinase (MMP)-1 and MMP-3 genes with colorectal cancer. Int. J. Cancer 2002, 102, 526–529. [Google Scholar] [CrossRef]
- Kouhkan, F.; Motovali-Bashi, M.; Hojati, Z. The influence of interstitial collagenas-1 genotype polymorphism on colorectal cancer risk in Iranian population. Cancer Investig. 2008, 26, 836–842. [Google Scholar] [CrossRef]
- Langers, A.M.J.; Sier, C.F.M.; Hawinkels, L.J.A.C.; Kubben, F.J.G.M.; van Duijn, W.; van der Reijden, J.J.; Lamers, C.B.H.W.; Hommes, D.W.; Verspaget, H.W. MMP-2 geno-phenotype is prognostic for colorectal cancer survival, whereas MMP-9 is not. Br. J. Cancer 2008, 98, 1820–1823. [Google Scholar] [CrossRef]
- Ting, W.-C.; Chen, L.-M.; Pao, J.-B.; Yang, Y.-P.; You, B.-J.; Chang, T.-Y.; Lan, Y.-H.; Lee, H.-Z.; Bao, B.-Y. Genetic Polymorphisms of Matrix Metalloproteinases and Clinical Outcomes in Colorectal Cancer Patients. Int. J. Med. Sci. 2013, 10, 1022–1027. [Google Scholar] [CrossRef]
- Dziki, L.; Przybyłowska, K.; Majsterek, I.; Trzciński, R.; Mik, M.; Sygut, A. A/G Polymorphism of the MMP-7 Gene Promoter Region in Colorectal Cancer. Pol. Przegl. Chir. 2011, 83, 622–626. [Google Scholar] [CrossRef]
- Fang, W.-L.; Liang, W.; He, H.; Zhu, Y.; Li, S.-L.; Gao, L.-B.; Zhang, L. Association of Matrix Metalloproteinases 1, 7, and 9 Gene Polymorphisms with Genetic Susceptibility to Colorectal Carcinoma in a Han Chinese Population. DNA Cell Biol. 2010, 29, 657–661. [Google Scholar] [CrossRef]
- Park, K.S.; Kim, S.J.; Kim, K.H.; Kim, J.C. Clinical characteristics of TIMP2, MMP2, and MMP9 gene polymorphisms in colorectal cancer. J. Gastroenterol. Hepatol. 2011, 26, 391–397. [Google Scholar] [CrossRef]
- Van Nguyen, S.; Skarstedt, M.; LÖFgren, S.; Zar, N.; Andersson, R.E.; Lindh, M.; Matussek, A.; Dimberg, J.A.N. Gene Polymorphism of Matrix Metalloproteinase-12 and -13 and Association with Colorectal Cancer in Swedish Patients. Anticancer Res. 2013, 33, 3247–3250. [Google Scholar]
- Tai, J.; Sun, D.; Wang, X.; Kang, Z. Matrix metalloproteinase-8 rs11225395 polymorphism correlates with colorectal cancer risk and survival in a Chinese Han population: A case-control study. Aging 2020, 12, 19618–19627. [Google Scholar] [CrossRef]
- Vandenbroucke, R.E.; Libert, C. Is there new hope for therapeutic matrix metalloproteinase inhibition? Nat. Rev. Drug Discov. 2014, 13, 904–927. [Google Scholar] [CrossRef]
- Raeeszadeh-Sarmazdeh, M.; Do, L.D.; Hritz, B.G. Metalloproteinases and Their Inhibitors: Potential for the Development of New Therapeutics. Cells 2020, 9, 1313. [Google Scholar] [CrossRef]
- Caton, J.G. Evaluation of Periostat for patient management. Compend. Contin. Educ. Dent. 1999, 20, 451, 458–460. [Google Scholar]
- Bissett, D.; O’Byrne, K.J.; von Pawel, J.; Gatzemeier, U.; Price, A.; Nicolson, M.; Mercier, R.; Mazabel, E.; Penning, C.; Zhang, M.H.; et al. Phase III study of matrix metalloproteinase inhibitor prinomastat in non-small-cell lung cancer. J. Clin. Oncol. 2005, 23, 842–849. [Google Scholar] [CrossRef]
- Scatena, R. Prinomastat, a hydroxamate-based matrix metalloproteinase inhibitor. A novel pharmacological approach for tissue remodelling-related diseases. Expert Opin. Investig. Drugs 2000, 9, 2159–2165. [Google Scholar] [CrossRef]
- Hande, K.R.; Collier, M.; Paradiso, L.; Stuart-Smith, J.; Dixon, M.; Clendeninn, N.; Yeun, G.; Alberti, D.; Binger, K.; Wilding, G. Phase I and pharmacokinetic study of prinomastat, a matrix metalloprotease inhibitor. Clin. Cancer Res. 2004, 10, 909–915. [Google Scholar] [CrossRef] [PubMed]
- Yang, J.-S.; Lin, C.-W.; Su, S.-C.; Yang, S.-F. Pharmacodynamic considerations in the use of matrix metalloproteinase inhibitors in cancer treatment. Expert Opin. Drug Metab. Toxicol. 2015, 12, 191–200. [Google Scholar] [CrossRef] [PubMed]
- Shah, M.A.; Starodub, A.; Sharma, S.; Berlin, J.; Patel, M.; Wainberg, Z.A.; Chaves, J.; Gordon, M.; Windsor, K.; Brachmann, C.B.; et al. Andecaliximab/GS-5745 Alone and Combined with mFOLFOX6 in Advanced Gastric and Gastroesophageal Junction Adenocarcinoma: Results from a Phase I Study. Clin. Cancer Res. 2018, 24, 3829–3837. [Google Scholar] [CrossRef] [PubMed]
- Sandborn, W.J.; Bhandari, B.R.; Randall, C.; Younes, Z.H.; Romanczyk, T.; Xin, Y.; Wendt, E.; Chai, H.; McKevitt, M.; Zhao, S.; et al. Andecaliximab [Anti-matrix Metalloproteinase-9] Induction Therapy for Ulcerative Colitis: A Randomised, Double-Blind, Placebo-Controlled, Phase 2/3 Study in Patients With Moderate to Severe Disease. J. Crohn’s Colitis 2018, 12, 1021–1029. [Google Scholar] [CrossRef]
- Murphy, G. Tissue inhibitors of metalloproteinases. Genome Biol. 2011, 12, 233. [Google Scholar] [CrossRef]
- Li, K.; Tay, F.R.; Yiu, C.K.Y. The past, present and future perspectives of matrix metalloproteinase inhibitors. Pharmacol. Ther. 2020, 207, 107465. [Google Scholar] [CrossRef]
- Melendez-Zajgla, J.; Del Pozo, L.; Ceballos, G.; Maldonado, V. Tissue Inhibitor of Metalloproteinases-4. The road less traveled. Mol. Cancer 2008, 7, 85. [Google Scholar] [CrossRef]
- Hayden, D.M.; Forsyth, C.; Keshavarzian, A. The role of matrix metalloproteinases in intestinal epithelial wound healing during normal and inflammatory states. J. Surg. Res. 2011, 168, 315–324. [Google Scholar] [CrossRef]
- Song, G.; Xu, S.; Zhang, H.; Wang, Y.; Xiao, C.; Jiang, T.; Wu, L.; Zhang, T.; Sun, X.; Zhong, L.; et al. TIMP1 is a prognostic marker for the progression and metastasis of colon cancer through FAK-PI3K/AKT and MAPK pathway. J. Exp. Clin. Cancer Res. 2016, 35, 148. [Google Scholar] [CrossRef]
- Noël, A.; Jost, M.; Maquoi, E. Matrix metalloproteinases at cancer tumor-host interface. Semin. Cell Dev. Biol. 2008, 19, 52–60. [Google Scholar] [CrossRef]
- Lu, X.; Duan, L.; Xie, H.; Lu, X.; Lu, D.; Lu, D.; Jiang, N.; Chen, Y. Evaluation of MMP-9 and MMP-2 and their suppressor TIMP-1 and TIMP-2 in adenocarcinoma of esophagogastric junction. OncoTargets Ther. 2016, 9, 4343–4349. [Google Scholar] [CrossRef][Green Version]
- Groblewska, M.; Mroczko, B.; Gryko, M.; Pryczynicz, A.; Guzińska-Ustymowicz, K.; Kędra, B.; Kemona, A.; Szmitkowski, M. Serum levels and tissue expression of matrix metalloproteinase 2 (MMP-2) and tissue inhibitor of metalloproteinases 2 (TIMP-2) in colorectal cancer patients. Tumor Biol. 2014, 35, 3793–3802. [Google Scholar] [CrossRef]
- Wang, W.; Li, D.; Xiang, L.; Lv, M.; Tao, L.; Ni, T.; Deng, J.; Gu, X.; Masatara, S.; Liu, Y.; et al. TIMP-2 inhibits metastasis and predicts prognosis of colorectal cancer via regulating MMP-9. Cell Adhes. Migr. 2019, 13, 273–284. [Google Scholar] [CrossRef]
- Huang, H.-L.; Liu, Y.-M.; Sung, T.-Y.; Huang, T.-C.; Cheng, Y.-W.; Liou, J.-P.; Pan, S.-L. TIMP3 expression associates with prognosis in colorectal cancer and its novel arylsulfonamide inducer, MPT0B390, inhibits tumor growth, metastasis and angiogenesis. Theranostics 2019, 9, 6676–6689. [Google Scholar] [CrossRef]
- Lin, H.; Zhang, Y.; Wang, H.; Xu, D.; Meng, X.; Shao, Y.; Lin, C.; Ye, Y.; Qian, H.; Wang, S. Tissue inhibitor of metalloproteinases-3 transfer suppresses malignant behaviors of colorectal cancer cells. Cancer Gene Ther. 2012, 19, 845–851. [Google Scholar] [CrossRef]
- Soheilifar, M.H.; Grusch, M.; Keshmiri Neghab, H.; Amini, R.; Maadi, H.; Saidijam, M.; Wang, Z. Angioregulatory microRNAs in Colorectal Cancer. Cancers 2019, 12, 71. [Google Scholar] [CrossRef]
- Wu, J.; Wu, G.; Lv, L.; Ren, Y.; Zhang, X.; Xue, Y.; Li, G.; Lu, X.; Sun, Z.; Tang, K. MicroRNA-34a inhibits migration and invasion of colon cancer cells via targeting to Fra-1. Carcinogenesis 2012, 33, 519–528. [Google Scholar] [CrossRef]
- Abba, M.; Patil, N.; Allgayer, H. MicroRNAs in the Regulation of MMPs and Metastasis. Cancers 2014, 6, 625–645. [Google Scholar] [CrossRef]
- Shen, K.; Liang, Q.; Xu, K.; Cui, D.; Jiang, L.; Yin, P.; Lu, Y.; Li, Q.; Liu, J. MiR-139 inhibits invasion and metastasis of colorectal cancer by targeting the type I insulin-like growth factor receptor. Biochem. Pharmacol. 2012, 84, 320–330. [Google Scholar] [CrossRef]
- Tang, W.; Zhu, Y.; Gao, J.; Fu, J.; Liu, C.; Liu, Y.; Song, C.; Zhu, S.; Leng, Y.; Wang, G.; et al. MicroRNA-29a promotes colorectal cancer metastasis by regulating matrix metalloproteinase 2 and E-cadherin via KLF4. Br. J. Cancer 2014, 110, 450–458. [Google Scholar] [CrossRef]
- Leng, Y.; Chen, Z.; Ding, H.; Zhao, X.; Qin, L.; Pan, Y. Overexpression of microRNA-29b inhibits epithelial-mesenchymal transition and angiogenesis of colorectal cancer through the ETV4/ERK/EGFR axis. Cancer Cell Int. 2021, 21, 17. [Google Scholar] [CrossRef] [PubMed]
- Yu, B.; Liu, X.; Chang, H. MicroRNA-143 inhibits colorectal cancer cell proliferation by targeting MMP7. Minerva Med. 2017, 108, 13–19. [Google Scholar] [CrossRef] [PubMed]
- Schwarzmueller, L.; Bril, O.; Vermeulen, L.; Léveillé, N. Emerging Role and Therapeutic Potential of lncRNAs in Colorectal Cancer. Cancers 2020, 12, 3843. [Google Scholar] [CrossRef] [PubMed]
- Tian, L.; Zhao, Z.F.; Xie, L.; Zhu, J.P. Taurine up-regulated 1 accelerates tumorigenesis of colon cancer by regulating miR-26a-5p/MMP14/p38 MAPK/Hsp27 axis in vitro and in vivo. Life Sci. 2019, 239, 117035. [Google Scholar] [CrossRef]
- Sun, N.; Zhang, G.; Liu, Y. Long non-coding RNA XIST sponges miR-34a to promotes colon cancer progression via Wnt/β-catenin signaling pathway. Gene 2018, 665, 141–148. [Google Scholar] [CrossRef]
- Lv, H.; Zhou, D.; Liu, G. LncRNA LINC00963 promotes colorectal cancer cell proliferation and metastasis by regulating miR--1281 and TRIM65. Mol. Med. Rep. 2021, 24, 781. [Google Scholar] [CrossRef]
- Duan, Y.; Fang, Z.; Shi, Z.; Zhang, L. Knockdown of lncRNA CCEPR suppresses colorectal cancer progression. Exp. Ther. Med. 2019, 18, 3534–3542. [Google Scholar] [CrossRef]
- Pan, Y.; Zhu, L.; Pu, J.; Wang, W.; Qian, W. lncRNA PCA3 plays a key role in colon cancer occurrence and development. Arch. Med. Sci. 2020. [Google Scholar] [CrossRef]
| MMP Gene | Chromosomal Location | Enzyme | Substrate |
|---|---|---|---|
| MMP-1 | 11q22.2 | Collagenase-1 | Col I, II, III, VII, VIII, X, Gelatin |
| MMP-8 | 11q22.2 | Collagenase-2 | Col I, II, III, VII, VIII, X, Gelatin, Aggrecan |
| MMP-13 | 11q22.2 | Collagenase-3 | Col I, II, III, VII, VIII, X, Gelatin |
| MMP-2 | 16q12.2 | Gelatinase A | Gelatin, Col I, II, III, IV, VII |
| MMP-9 | 20q13.12 | Gelatinase B | Gelatin, Col IV, V |
| MMP-3 | 11q22.3 | Stromelysin-1 | Col II, III, IV, IX, X, proteoglycans, fibronectin, laminin, and elastin. |
| MMP-10 | 11q22.2 | Stromelysin-2 | Col II, III, IV, IX, X, proteoglycans, fibronectin, laminin, and elastin |
| MMP-7 | 11q22.2 | Marilysin-1 | Fibronectin, Laminin, Col I, Gelatin |
| MMP-14 | 14q11.2 | MT-MMP | Gelatin, Fibronectin, Laminin |
| MMP-12 | 11q22.2 | Metalloelastase | Gelatin, Fibronectin, Col IV |
| MMP-21 | 10q26.2 | XMMP | Aggrecan |
| References | Gene/Protein Expression | Samples | Methods | Results |
|---|---|---|---|---|
| Huang X., et al., 2021 [38] | MMP-7, MMP-9, MMP-11, TIMP-1, TIMP-2, CEA | Human polyps and tumor | Enzyme-linked immunosorbent assay | A combined detection model, including MMP-7, TIMP-1, and CEA improved both the specificity and sensitivity for detecting CRC. |
| Zhou X., et al., 2021 [39] | MMP-7, MMP-9, MMP-11, TIMP-1, TIMP-2, CEA | Human CRC | ELISA and electrochemiluminescence immunoassay | The miR 135a was downregulated and MMP 13 was increased in samples. Combined detection of the two had a good diagnostic effect on the occurrence of CRC. |
| Rasool M., et al., 2021 [40] | TGF, VEGF, TNF, ILs, MMP-2, 9, 11, and 19 | Human polyps and tumor | ELISA | Significant upregulation of MMP-2, MMP-9, MMP-11, and MMP-19 was reported in polyp and colon cancer samples compared with their MMP profile in normal samples. |
| Barabás L., et al., 2020 [41] | MMP-2, MMP-7, MMP-9, TIMP-1 and TIMP-2 | Human adenomas, and CRC | ELISA | The serum antigen concentrations of MMP-7, MMP-9, TIMP-1, and TIMP-2 were significantly increased in patients with CRC and adenomas compared with the controls. They were also activated in premalignant adenomas. |
| Hsieh S.L., et al., 2019 [42] | Study of the mechanism of carnosine, TIMP-1, and MMP-9 | Human HCT-116 CRC cell line | MTT assay and qPCR | The carnosine inhibits the migration and intravasation of human CRC cells. The regulatory mechanism may occur by suppressing NF-κB activity and modulating MMPs and EMT-related gene expression in HCT-116 cells treated with carnosine. MMP-9 mRNA and protein levels were decreased. TIMP-1 mRNA and protein levels were increased. |
| Kıyak R., et al., 2018 [43] | MMP-7, COX-2, TIMP-1, and CEA protein | Human polyps | ELISA and chemiluminescent enzyme immunometric assay (CEIA) | The plasma TIMP-1 levels were significantly elevated in cancer compared with the polyp group. The plasma MMP-7 levels were decreased in polyps compared with the control group. The plasma CEA and TIMP-1 are valuable biomarker candidates for differentiating CRC from colorectal polyps. |
| Eiró N., et al., 2017 [44] | MMP-1, 2, 7, 9, 11, 13 and 14 | Human adenomas and hyperplastic polyps | Real-time PCR and Western-blot, and | The hyperplastic polyps had the lowest levels of MMP-1 and MMP-7. Tubular polyps had high levels of both MMP-7 and MMP-14, and tubulo-villous adenomas had high levels of MMP-1, 7, and 14 compared with the normal group. |
| Pezeshkian Z., et al., 2017 [45] | MMP-7 and VEGF-A | Human adenomas | Real-time PCR in 50 biopsy samples of adenomas including villous, tubular, and tubulo-villous types, and 20 paired tissue samples | The MMP-7 mRNA expression was significantly higher in villous adenoma with high-grade dysplasia compared with the control group. MMP-7 and VEGF-A are prognostic biomarkers for colorectal adenoma polyp progression to malignancy. |
| Wernicke A.K., et al., 2016 [46] | Association between grade of dysplasia and MMP-13 expression | Human adenomas and hyperplastic polyps | Immunohistochemistry and immune-reactive score (IRS) | The MMP-13 has been identified as an excellent marker of high-grade intraepithelial neoplasia and CRC. The strength of the association between pathologic stage and immune-reactive MMP-13 scoring emphasizes its potential for diagnosis in precancerous colorectal lesions. |
| Gimeno-García A., et al., 2016 [47] | MMP-9 | Patients’ blood, adenomas, hyperplastic polyps, and CRC tissue | Luminex XMAP technology, gelatin zymography, western blot, and SNP analysis in 150 blood and tissue | There was a significant correlation between plasma and tissue levels of MMP-9. Plasma MMP-9 levels in patients with neoplastic lesions were significantly higher than in healthy controls. Also, MMP-9 in CRC was higher than in non-advanced adenomas. |
| Annaha’zi A., et al., 2016 [48] | MMP-9 | Patients′ stool samples, adenomas, hyperplastic polyps, and CRC tissue | ELISA | Stool MMP-9 was significantly increased in CRC compared with all the other groups. Stool MMP-9 may be a new noninvasive marker in CRC. |
| Klupp et al., 2016 [49] | MMP-7, MMP-10, and MMP-12 | Serum specimens of patients with colon adenocarcinoma | Luminex based multiplex assay | Expression levels of MMP-7, MMP-10, and MMP-12 in serum of colon cancer patients are different compared with serum specimens of the healthy control group. The upregulation of MMP-7, MMP-10, and MMP-12 in colon cancer patients’ serum was associated with a poor prognosis. |
| Otero-Estévez O., et al., 2015 [50] | MMP-9 | Human adenomas and CRC | non-invasive stool immunochemical test (FIT) and ELISA | The MMP-9 levels were higher in advanced adenomas and CRC compared with those reported in samples of healthy individual. Elevated MMP-9 concentration was associated with several lesions, size, and adenoma histology. |
| Bengi G., et al., 2015 [51] | MMP-7, TIMP-1, and COX-2 | Human adenomas and CRC | Real-time PCR | The expression of TIMP-1, COX-2, and MMP-7 was significantly higher in polyps compared with normal tissue. Overexpression of MMP-7, COX-2, and TIMP-1 determine an important role of these genes in the progression of colon cancer. |
| Odabasi M., et al., 2014 [52] | MMP-9 and NGAL | Human adenomas and CRC | Immunohistochemistry | The MMP-9 and NGAL overexpression in neoplastic polyps might be used as markers to separate them from non-neoplastic polyps. These genes as immune-histochemical markers determine dysplasia in the early steps of the colorectal adenoma-carcinoma sequence. |
| Qasim B.J., et al., 2013 [53] | MMP-7 | Human adenomas | Immunohistochemistry | MMP-7 was expressed in advanced colorectal adenomatous polyps with large size, severe dysplasia, and villous. |
| Sheth R.A., et al., 2012 [54] | MMP-2, and MMP-9 | Xenograft model of CRC in nude mice | The MMP enzyme activity was measured by an enzyme-activatable optical molecular probe and quantitative fluorescence colonoscopy in nude mice which received celecoxib versus vehicle | There was an apparent linear relationship between measured MMP activity and tumor growth rate. |
| Murname M.J., et al., 2009 [55] | MMP-2 and MMP-9 | Mouse models of CRC and human HT-29 CRC cell line | Gene-expression microarray and ELISA | The plotted receiver operating characteristic (ROC) curves estimated the sensitivity and specificity profiles of MMP-2 and MMP-9 for the identification of CRC. |
| Jeffery N., et al., 2009 [56] | MMP-1, 2, 3, 7, 9, 13, MT1-MMP, MT2-MMP and TIMP-1, TIMP-2, and IMP-3 | Human adenomas and CRC | Immunohistochemistry | MMP-1, MMP-2, MMP-3, TIMP-1, and TIMP-2 showed a significant increase in carcinomatous epithelium compared with adenoma epithelium. The increased expression of MMPs and TIMPs occurred at an early stage of colorectal neoplasia. |
| Lièvre A., et al., 2006 [57] | The functional gene promoter polymorphisms of MMP1, MMP3, and MMP7 | Human adenomas | Real-time PCR allelic discrimination assay | These data showed a relation between MMP-1 -1607 ins/del G and MMP-3 -1612 ins/del A combined polymorphisms and risk of small adenomas. |
| Tutton M.G., et al., 2003 [58] | MMP-2 and MMP-9 | Patients’ plasma samples, adenomas, and CRC | Immunohistochemistry, real-time PCR, and ELISA | The expression of MMP-2 and MMP-9 was significantly increased in CRC tissues compared with matched normal tissues. Plasma MMP-2 and MMP-9 levels were significantly elevated at all stages in CRC patients. Plasma levels of these enzymes may be a noninvasive indicator of invasion or metastasis in CRC. |
| MicroRNA | MMP | Result |
|---|---|---|
| miR-34a | MMP-1, MMP-9 | miR-34a overexpression prevents tumor cell proliferation, migration, and invasion [138,139]. |
| miR-139 | MMP-2 | Downregulation of miR-139 reduces proliferation, migration, and invasion [140]. |
| miR-29a | MMP-2 | Upregulation of miR-29a increases metastasis [141]. |
| miR-29b | MMP-2 | Upregulation of miR-29b increases metastasis [142]. |
| miR-143 | MMP-7 | Upregulation of miR-143 enhances tumor cell proliferation and invasion [143]. |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Pezeshkian, Z.; Nobili, S.; Peyravian, N.; Shojaee, B.; Nazari, H.; Soleimani, H.; Asadzadeh-Aghdaei, H.; Ashrafian Bonab, M.; Nazemalhosseini-Mojarad, E.; Mini, E. Insights into the Role of Matrix Metalloproteinases in Precancerous Conditions and in Colorectal Cancer. Cancers 2021, 13, 6226. https://doi.org/10.3390/cancers13246226
Pezeshkian Z, Nobili S, Peyravian N, Shojaee B, Nazari H, Soleimani H, Asadzadeh-Aghdaei H, Ashrafian Bonab M, Nazemalhosseini-Mojarad E, Mini E. Insights into the Role of Matrix Metalloproteinases in Precancerous Conditions and in Colorectal Cancer. Cancers. 2021; 13(24):6226. https://doi.org/10.3390/cancers13246226
Chicago/Turabian StylePezeshkian, Zahra, Stefania Nobili, Noshad Peyravian, Bahador Shojaee, Haniye Nazari, Hiva Soleimani, Hamid Asadzadeh-Aghdaei, Maziar Ashrafian Bonab, Ehsan Nazemalhosseini-Mojarad, and Enrico Mini. 2021. "Insights into the Role of Matrix Metalloproteinases in Precancerous Conditions and in Colorectal Cancer" Cancers 13, no. 24: 6226. https://doi.org/10.3390/cancers13246226
APA StylePezeshkian, Z., Nobili, S., Peyravian, N., Shojaee, B., Nazari, H., Soleimani, H., Asadzadeh-Aghdaei, H., Ashrafian Bonab, M., Nazemalhosseini-Mojarad, E., & Mini, E. (2021). Insights into the Role of Matrix Metalloproteinases in Precancerous Conditions and in Colorectal Cancer. Cancers, 13(24), 6226. https://doi.org/10.3390/cancers13246226










