Reproducibility of CT-Based Hepatocellular Carcinoma Radiomic Features across Different Contrast Imaging Phases: A Proof of Concept on SORAMIC Trial Data
Abstract
:Simple Summary
Abstract
1. Introduction
2. Materials and Methods
2.1. Patients and Imaging Data
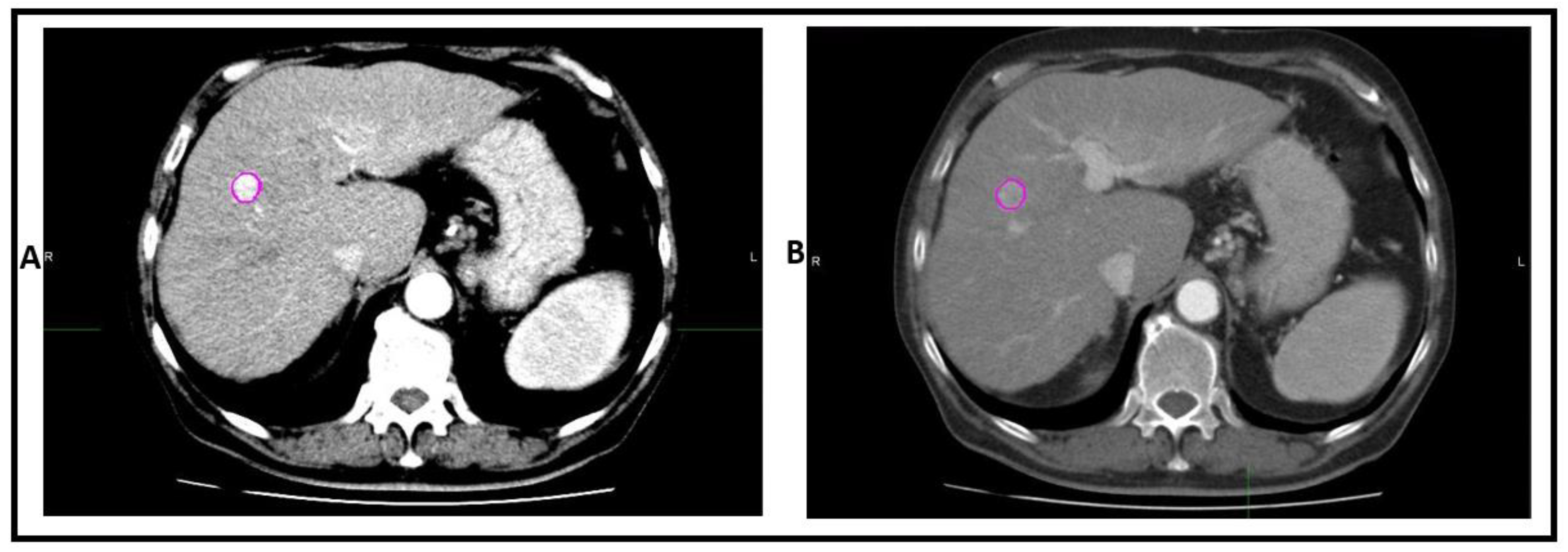
2.2. Segmentation and HRFs Extraction
2.3. ComBat Harmonization
2.4. Statistical Analysis
3. Results
3.1. Patient Characteristics
3.2. Extracted HRFs
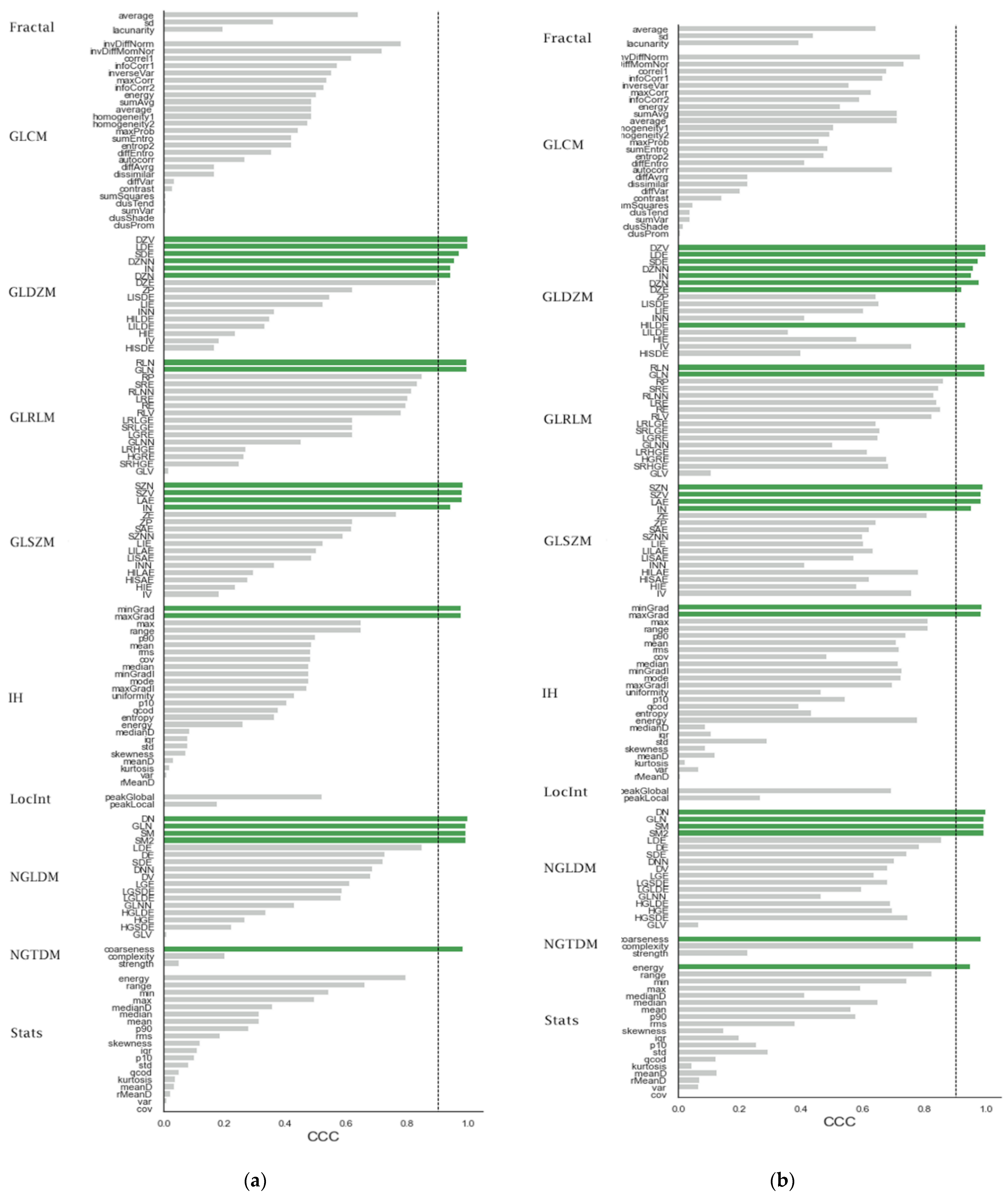
3.3. The Effects of Differences in Imaging Phase on the Reproducibility of HRFs
3.4. The Effects of ComBat on the Reproducibility of HRFs
4. Discussion
5. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Walsh, S.; de Jong, E.E.C.; van Timmeren, J.E.; Ibrahim, A.; Compter, I.; Peerlings, J.; Sanduleanu, S.; Refaee, T.; Keek, S.; Larue, R.T.H.; et al. Decision Support Systems in Oncology. JCO Clin. Cancer Inform. 2019, 3, 1–9. [Google Scholar] [CrossRef]
- Lambin, P.; Rios-Velazquez, E.; Leijenaar, R.; Carvalho, S.; van Stiphout, R.G.P.M.; Granton, P.; Zegers, C.M.L.; Gillies, R.; Boellard, R.; Dekker, A.; et al. Radiomics: Extracting more information from medical images using advanced feature analysis. Eur. J. Cancer 2012, 48, 441–446. [Google Scholar] [CrossRef] [Green Version]
- Gillies, R.J.; Kinahan, P.E.; Hricak, H. Radiomics: Images Are More than Pictures, They Are Data. Radiology 2016, 278, 563–577. [Google Scholar] [CrossRef] [Green Version]
- Refaee, T.; Wu, G.; Ibrahim, A.; Halilaj, I.; Leijenaar, R.T.H.H.; Rogers, W.; Gietema, H.A.; Hendriks, L.E.L.; Lambin, P.; Woodruff, H.C. The Emerging Role of Radiomics in COPD and Lung Cancer. Respiration 2020, 99, 99–107. [Google Scholar] [CrossRef] [PubMed]
- Aerts, H.J.W.L. The Potential of Radiomic-Based Phenotyping in Precision Medicine: A Review. JAMA Oncol. 2016, 2, 1636–1642. [Google Scholar] [CrossRef] [PubMed]
- van Timmeren, J.E.; Leijenaar, R.T.H.; van Elmpt, W.; Reymen, B.; Oberije, C.; Monshouwer, R.; Bussink, J.; Brink, C.; Hansen, O.; Lambin, P. Survival prediction of non-small cell lung cancer patients using radiomics analyses of cone-beam CT images. Radiother. Oncol. 2017, 123, 363–369. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Panth, K.M.; Leijenaar, R.T.H.; Carvalho, S.; Lieuwes, N.G.; Yaromina, A.; Dubois, L.; Lambin, P. Is there a causal relationship between genetic changes and radiomics-based image features? An in vivo preclinical experiment with doxycycline inducible GADD34 tumor cells. Radiother. Oncol. 2015, 116, 462–466. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Jethanandani, A.; Lin, T.A.; Volpe, S.; Elhalawani, H.; Mohamed, A.S.R.; Yang, P.; Fuller, C.D. Exploring Applications of Radiomics in Magnetic Resonance Imaging of Head and Neck Cancer: A Systematic Review. Front. Oncol. 2018, 8, 131. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Ursprung, S.; Beer, L.; Bruining, A.; Woitek, R.; Stewart, G.D.; Gallagher, F.A.; Sala, E. Radiomics of computed tomography and magnetic resonance imaging in renal cell carcinoma—a systematic review and meta-analysis. Eur. Radiol. 2020, 30, 3558–3566. [Google Scholar] [CrossRef] [Green Version]
- Samiei, S.; Granzier, R.W.Y.; Ibrahim, A.; Primakov, S.; Lobbes, M.B.I.; Beets-Tan, R.G.H.; van Nijnatten, T.J.A.; Engelen, S.M.E.; Woodruff, H.C.; Smidt, M.L. Dedicated Axillary MRI-Based Radiomics Analysis for the Prediction of Axillary Lymph Node Metastasis in Breast Cancer. Cancers 2021, 13, 757. [Google Scholar] [CrossRef]
- Ibrahim, A.; Vallières, M.; Woodruff, H.; Primakov, S.; Beheshti, M.; Keek, S.; Refaee, T.; Sanduleanu, S.; Walsh, S.; Morin, O.; et al. Radiomics Analysis for Clinical Decision Support in Nuclear Medicine. Semin. Nucl. Med. 2019, 49, 438–449. [Google Scholar] [CrossRef]
- Lovinfosse, P.; Visvikis, D.; Hustinx, R.; Hatt, M. FDG PET radiomics: A review of the methodological aspects. Clin. Transl. Imaging 2018, 6, 379–391. [Google Scholar] [CrossRef]
- Sung, H.; Ferlay, J.; Siegel, R.L.; Laversanne, M.; Soerjomataram, I.; Jemal, A.; Bray, F. Global Cancer Statistics 2020: GLOBOCAN Estimates of Incidence and Mortality Worldwide for 36 Cancers in 185 Countries. CA Cancer J. Clin. 2021, 71, 209–249. [Google Scholar] [CrossRef]
- Aubé, C.; Oberti, F.; Lonjon, J.; Pageaux, G.; Seror, O.; N’Kontchou, G.; Rode, A.; Radenne, S.; Cassinotto, C.; Vergniol, J.; et al. EASL and AASLD recommendations for the diagnosis of HCC to the test of daily practice. Liver Int. 2017, 37, 1515–1525. [Google Scholar] [CrossRef]
- Finn, R.S.; Qin, S.; Ikeda, M.; Galle, P.R.; Ducreux, M.; Kim, T.-Y.; Kudo, M.; Breder, V.; Merle, P.; Kaseb, A.O. Atezolizumab plus Bevacizumab in Unresectable Hepatocellular Carcinoma. N. Engl. J. Med. 2020, 382, 1894–1905. [Google Scholar] [CrossRef] [PubMed]
- Mokrane, F.-Z.; Lu, L.; Vavasseur, A.; Otal, P.; Peron, J.-M.; Luk, L.; Yang, H.; Ammari, S.; Saenger, Y.; Rousseau, H.; et al. Radiomics machine-learning signature for diagnosis of hepatocellular carcinoma in cirrhotic patients with indeterminate liver nodules. Eur. Radiol. 2020, 30, 558–570. [Google Scholar] [CrossRef]
- Wu, J.; Liu, A.; Cui, J.; Chen, A.; Song, Q.; Xie, L. Radiomics-based classification of hepatocellular carcinoma and hepatic haemangioma on precontrast magnetic resonance images. BMC Med. Imaging 2019, 19, 1–11. [Google Scholar] [CrossRef] [Green Version]
- Zhou, Y.; He, L.; Huang, Y.; Chen, S.; Wu, P.; Ye, W.; Liu, Z.; Liang, C. CT-based radiomics signature: A potential biomarker for preoperative prediction of early recurrence in hepatocellular carcinoma. Abdom. Radiol. 2017, 42, 1695–1704. [Google Scholar] [CrossRef] [PubMed]
- Wakabayashi, T.; Ouhmich, F.; Gonzalez-Cabrera, C.; Felli, E.; Saviano, A.; Agnus, V.; Savadjiev, P.; Baumert, T.F.; Pessaux, P.; Marescaux, J.; et al. Radiomics in hepatocellular carcinoma: A quantitative review. Hepatol. Int. 2019, 13, 546–559. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Yip, S.S.F.; Aerts, H.J.W.L. Applications and limitations of radiomics. Phys. Med. Biol. 2016, 61, R150–R166. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Ibrahim, A.; Primakov, S.; Beuque, M.; Woodruff, H.C.; Halilaj, I.; Wu, G.; Refaee, T.; Granzier, R.; Widaatalla, Y.; Hustinx, R.; et al. Radiomics for precision medicine: Current challenges, future prospects, and the proposal of a new framework. Methods 2021, 188, 20–29. [Google Scholar] [CrossRef]
- LaRue, R.T.H.M.; Van Timmeren, J.E.; De Jong, E.E.C.; Feliciani, G.; Leijenaar, R.T.H.H.; Schreurs, W.M.J.; Sosef, M.N.; Raat, F.H.P.J.; Van Der Zande, F.H.R.; Das, M.; et al. Influence of gray level discretization on radiomic feature stability for different CT scanners, tube currents and slice thicknesses: A comprehensive phantom study. Acta Oncol. 2017, 56, 1544–1553. [Google Scholar] [CrossRef]
- Van Timmeren, J.E.; Leijenaar, R.T.H.H.; van Elmpt, W.; Wang, J.; Zhang, Z.; Dekker, A.; Lambin, P. Test–Retest Data for Radiomics Feature Stability Analysis: Generalizable or Study-Specific? Tomography 2016, 2, 361–365. [Google Scholar] [CrossRef]
- Peerlings, J.; Woodruff, H.C.; Winfield, J.M.; Ibrahim, A.; Van Beers, B.E.; Heerschap, A.; Jackson, A.; Wildberger, J.E.; Mottaghy, F.M.; DeSouza, N.M.; et al. Stability of radiomics features in apparent diffusion coefficient maps from a multi-centre test-retest trial. Sci. Rep. 2019, 9, 4800. [Google Scholar] [CrossRef] [Green Version]
- Granzier, R.W.Y.; Verbakel, N.M.H.; Ibrahim, A.; Van Timmeren, J.E.; Van Nijnatten, T.J.A.; Leijenaar, R.T.H.H.; Lobbes, M.B.I.; Smidt, M.L.; Woodruff, H.C. MRI-based radiomics in breast cancer: Feature robustness with respect to inter-observer segmentation variability. Sci. Rep. 2020, 10, 14163. [Google Scholar] [CrossRef]
- Leijenaar, R.T.H.H.; Carvalho, S.; Velazquez, E.R.; Van Elmpt, W.J.C.; Parmar, C.; Hoekstra, O.S.; Hoekstra, C.J.; Boellaard, R.; Dekker, A.L.A.J.; Gillies, R.J.; et al. Stability of FDG-PET Radiomics features: An integrated analysis of test-retest and inter-observer variability. Acta Oncol. 2013, 52, 1391–1397. [Google Scholar] [CrossRef] [Green Version]
- Fortin, J.-P.; Parker, D.; Tunç, B.; Watanabe, T.; Elliott, M.A.; Ruparel, K.; Roalf, D.R.; Satterthwaite, T.D.; Gur, R.C.; Gur, R.E.; et al. Harmonization of Multi-Site Diffusion Tensor Imaging Data. Neuroimage 2017, 161, 149–170. [Google Scholar] [CrossRef] [PubMed]
- Fortin, J.-P.; Cullen, N.; Sheline, Y.I.; Taylor, W.D.; Aselcioglu, I.; Cook, P.A.; Adams, P.; Cooper, C.; Fava, M.; McGrath, P.J.; et al. Harmonization of Cortical Thickness Measurements across Scanners and Sites. Neuroimage 2018, 167, 104–120. [Google Scholar] [CrossRef] [PubMed]
- Da-Ano, R.; Masson, I.; Lucia, F.; Doré, M.; Robin, P.; Alfieri, J.; Rousseau, C.; Mervoyer, A.; Reinhold, C.; Castelli, J.; et al. Performance comparison of modified ComBat for harmonization of radiomic features for multicenter studies. Sci. Rep. 2020, 10, 10248. [Google Scholar] [CrossRef] [PubMed]
- Ibrahim, A.; Primakov, S.; Barufaldi, B.; Acciavatti, R.J.; Granzier, R.W.Y.; Hustinx, R.; Mottaghy, F.M.; Woodruff, H.C.; Wildberger, J.E.; Lambin, P.; et al. Reply to Orlhac, F.; Buvat, I. Comment on “Ibrahim et Al. The Effects of In-Plane Spatial Resolution on CT-Based Radiomic Features’ Stability with and without ComBat Harmonization. Cancers 2021, 13, 1848.”. Cancers 2021, 13, 3080. [Google Scholar] [CrossRef] [PubMed]
- Johnson, W.E.; Li, C.; Rabinovic, A. Adjusting batch effects in microarray expression data using empirical Bayes methods. Biostatistics 2007, 8, 118–127. [Google Scholar] [CrossRef]
- Ibrahim, A.; Refaee, T.; Leijenaar, R.T.H.H.; Primakov, S.; Hustinx, R.; Mottaghy, F.M.; Woodruff, H.C.; Maidment, A.D.A.; Lambin, P. The application of a workflow integrating the variable reproducibility and harmonizability of radiomic features on a phantom dataset. PLoS ONE 2021, 16, e0251147. [Google Scholar] [CrossRef] [PubMed]
- Ibrahim, A.; Refaee, T.; Primakov, S.; Barufaldi, B.; Acciavatti, R.J.; Granzier, R.W.Y.; Hustinx, R.; Mottaghy, F.M.; Woodruff, H.C.; Wildberger, J.E.; et al. The Effects of In-Plane Spatial Resolution on CT-Based Radiomic Features’ Stability with and without ComBat Harmonization. Cancers 2021, 13, 1848. [Google Scholar] [CrossRef] [PubMed]
- Ricke, J.; Klümpen, H.J.; Amthauer, H.; Bargellini, I.; Bartenstein, P.; de Toni, E.N.; Gasbarrini, A.; Pech, M.; Peck-Radosavljevic, M.; Popovič, P.; et al. Impact of Combined Selective Internal Radiation Therapy and Sorafenib on Survival in Advanced Hepatocellular Carcinoma. J. Hepatol. 2019, 71, 1164–1174. [Google Scholar] [CrossRef] [PubMed]
- World Medical Association. World Medical Association Declaration of Helsinki: Ethical Principles for Medical Research involving Human Subjects. JAMA 2013, 310, 2191–2194. [Google Scholar] [CrossRef] [Green Version]
- Zwanenburg, A.; Vallières, M.; Abdalah, M.A.; Aerts, H.J.W.L.; Andrearczyk, V.; Apte, A.; Ashrafinia, S.; Bakas, S.; Beukinga, R.J.; Boellaard, R.; et al. The Image Biomarker Standardization Initiative: Standardized Quantitative Radiomics for High-Throughput Image-based Phenotyping. Radiology 2020, 295, 328–338. [Google Scholar] [CrossRef] [Green Version]
- R Core Team. R Language Definition; R Foundation for Statistical Computing: Vienna, Austria, 2000. [Google Scholar]
- Gandrud, C. Reproducible Research with R and R Studio; CRC Press: Boca Raton, FL, USA, 2013; ISBN 9781466572843. [Google Scholar]
- Lawrence, I.; Lin, K. A Concordance Correlation Coefficient to Evaluate Reproducibility. Biometrics 1989, 45, 255–268. [Google Scholar] [CrossRef]
- Stevenson, M.; Stevenson, M.M.; BiasedUrn, I. Package “epiR”. Available online: https://vps.fmvz.usp.br/CRAN/web/packages/epiR/epiR.pdf (accessed on 10 March 2021).
- Aerts, H.J.W.L.; Velazquez, E.R.; Leijenaar, R.T.H.H.; Parmar, C.; Grossmann, P.; Carvalho, S.; Bussink, J.; Monshouwer, R.; Haibe-Kains, B.; Rietveld, D.; et al. Decoding tumour phenotype by noninvasive imaging using a quantitative radiomics approach. Nat. Commun. 2014, 5, 4006. [Google Scholar] [CrossRef]
- Perrin, T.; Midya, A.; Yamashita, R.; Chakraborty, J.; Saidon, T.; Jarnagin, W.R.; Gonen, M.; Simpson, A.L.; Do, R.K.G. Short-term reproducibility of radiomic features in liver parenchyma and liver malignancies on contrast-enhanced CT imaging. Abdom. Radiol. 2018, 43, 3271–3278. [Google Scholar] [CrossRef]
- Welch, M.L.; McIntosh, C.; Haibe-Kains, B.; Milosevic, M.F.; Wee, L.; Dekker, A.; Huang, S.H.; Purdie, T.G.; O’Sullivan, B.; Aerts, H.J.W.L.; et al. Vulnerabilities of radiomic signature development: The need for safeguards. Radiother. Oncol. 2019, 130, 2–9. [Google Scholar] [CrossRef] [Green Version]
- Andrearczyk, V.; Depeursinge, A.; Müller, H. Neural network training for cross-protocol radiomic feature standardization in computed tomography. J. Med. Imaging 2019, 6, 024008. [Google Scholar] [CrossRef] [PubMed]
- Bashyam, V.M.; Doshi, J.; Erus, G.; Srinivasan, D.; Abdulkadir, A.; Habes, M.; Fan, Y.; Masters, C.L.; Maruff, P.; Zhuo, C.; et al. Medical Image Harmonization Using Deep Learning Based Canonical Mapping: Toward Robust and Generalizable Learning in Imaging. arXiv 2020, arXiv:2010.05355. [Google Scholar]
- Modanwal, G.; Vellal, A.; Mazurowski, M.A. Normalization of breast MRIs using cycle-consistent generative adversarial networks. arXiv 2019, arXiv:1912.08061. [Google Scholar]
- Dewey, B.E.; Zhao, C.; Reinhold, J.C.; Carass, A.; Fitzgerald, K.C.; Sotirchos, E.S.; Saidha, S.; Oh, J.; Pham, D.L.; Calabresi, P.A.; et al. DeepHarmony: A deep learning approach to contrast harmonization across scanner changes. Magn. Reson. Imaging 2019, 64, 160–170. [Google Scholar] [CrossRef] [PubMed]

| Manufacturer | Scanner Model | X-ray Tube Current (kV) | Exposure (mAs) | Convolution Kernels | Slice Thickness (mm) | Pixel Spacing (mm2) |
|---|---|---|---|---|---|---|
| TOSHIBA | Aquilion | 50–360 | 2–300 | FC13 | 1–5 | 0.39 × 0.39 − 0.98 × 0.98 |
| Aquilion PRIME | ||||||
| Philips | Brilliance 64 | B | ||||
| GE | Discovery CT750 HD | STANDARD | ||||
| Optima CT660 | ||||||
| SIEMENS | Sensation 16 | B31f | ||||
| SOMATOM Definition AS | ||||||
| SOMATOM Definition Flash | I30f, I40f | |||||
| SOMATOM Force | Br40d |
| Characteristic | N = 61 |
|---|---|
| Gender, male (%) | 50 (81.9%) |
| Age, median (range) | 66 (48–81) |
| Cirrhosis, yes (%) | 56 (91.8%) |
| Child–Pugh grade | |
| A | 56 (91.8%) |
| B | 5 (8.2%) |
| Diameter of largest lesion, in mm, median (range) | 37 (10–220) |
| Portal vein invasion, yes (%) | 11 (18.1%) |
| Extrahepatic disease yes (%) | 7 (11.4%) |
| * BCLC staging | |
| A | 22 (36.1%) |
| B | 22 (36.1%) |
| C | 17 (27.8%) |
| ** ECOG performance | |
| 0 | 58 (95.1%) |
| 1 | 3 (4.9%) |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Ibrahim, A.; Widaatalla, Y.; Refaee, T.; Primakov, S.; Miclea, R.L.; Öcal, O.; Fabritius, M.P.; Ingrisch, M.; Ricke, J.; Hustinx, R.; et al. Reproducibility of CT-Based Hepatocellular Carcinoma Radiomic Features across Different Contrast Imaging Phases: A Proof of Concept on SORAMIC Trial Data. Cancers 2021, 13, 4638. https://doi.org/10.3390/cancers13184638
Ibrahim A, Widaatalla Y, Refaee T, Primakov S, Miclea RL, Öcal O, Fabritius MP, Ingrisch M, Ricke J, Hustinx R, et al. Reproducibility of CT-Based Hepatocellular Carcinoma Radiomic Features across Different Contrast Imaging Phases: A Proof of Concept on SORAMIC Trial Data. Cancers. 2021; 13(18):4638. https://doi.org/10.3390/cancers13184638
Chicago/Turabian StyleIbrahim, Abdalla, Yousif Widaatalla, Turkey Refaee, Sergey Primakov, Razvan L. Miclea, Osman Öcal, Matthias P. Fabritius, Michael Ingrisch, Jens Ricke, Roland Hustinx, and et al. 2021. "Reproducibility of CT-Based Hepatocellular Carcinoma Radiomic Features across Different Contrast Imaging Phases: A Proof of Concept on SORAMIC Trial Data" Cancers 13, no. 18: 4638. https://doi.org/10.3390/cancers13184638
APA StyleIbrahim, A., Widaatalla, Y., Refaee, T., Primakov, S., Miclea, R. L., Öcal, O., Fabritius, M. P., Ingrisch, M., Ricke, J., Hustinx, R., Mottaghy, F. M., Woodruff, H. C., Seidensticker, M., & Lambin, P. (2021). Reproducibility of CT-Based Hepatocellular Carcinoma Radiomic Features across Different Contrast Imaging Phases: A Proof of Concept on SORAMIC Trial Data. Cancers, 13(18), 4638. https://doi.org/10.3390/cancers13184638







