Precision Medicine and Triple-Negative Breast Cancer: Current Landscape and Future Directions
Abstract
Simple Summary
Abstract
1. Precision Medicine: Perspective and Challenges
2. Triple-Negative Breast Cancer (TNBC)
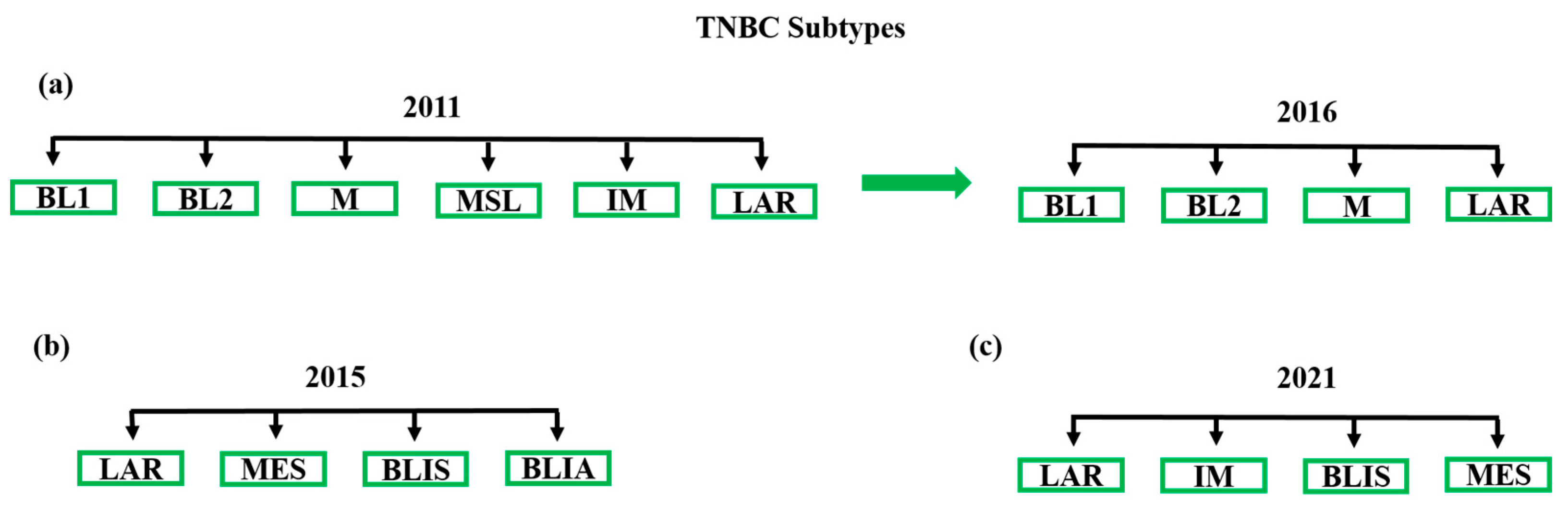
3. TNBC Subtypes and Current Treatment Options
4. Recently FDA-Approved Therapies for TNBC
4.1. Approved Checkpoint Inhibitors
Other Immune Checkpoint Inhibitors in Clinical Trials
4.2. Poly-ADP-Ribose Polymerase (PARP) Inhibitors
4.3. Antibody-Drug Conjugates (ADC)
5. Precision Medicine in TNBC: Emerging Therapies and Ongoing Studies
5.1. Receptor Tyrosine Kinases (RTKs) and Downstream Signaling Pathways
5.2. PI3K/AKT/mTOR Targeted Therapy
5.3. Notch Signaling and TNBC
5.4. Cyclin-Dependent Kinases (CDKs)
5.5. Androgen Receptor (AR) Expression and TNBC
5.6. Angiogenesis and TNBC
5.7. Investigational Antibody-Drug Conjugates (ADC)
6. Other Targeted Therapies
6.1. Cancer Vaccines
6.2. Combination Regimens
7. Design of Precision Medicine-Based Clinical Trials in TNBC: The Path Forward
8. Conclusions
Author Contributions
Funding
Conflicts of Interest
References
- Goodwin, S.; McPherson, J.D.; McCombie, W.R. Coming of age: Ten years of next-generation sequencing technologies. Nat. Rev. Genet. 2016, 17, 333–351. [Google Scholar] [CrossRef]
- Bush, W.S.; Moore, J.H. Chapter 11: Genome-wide association studies. PLoS Comput. Biol. 2012, 8, e1002822. [Google Scholar] [CrossRef]
- Ginsburg, G.S.; Phillips, K.A. Precision Medicine: From Science To Value. Health Aff. (Millwood) 2018, 37, 694–701. [Google Scholar] [CrossRef]
- Dugger, S.A.; Platt, A.; Goldstein, D.B. Drug development in the era of precision medicine. Nat. Rev. Drug Discov. 2018, 17, 183–196. [Google Scholar] [CrossRef]
- Xie, F.; Chan, J.C.; Ma, R.C. Precision medicine in diabetes prevention, classification and management. J. Diabetes Investig. 2018, 9, 998–1015. [Google Scholar] [CrossRef] [PubMed]
- Cooper, G.M.; Johnson, J.A.; Langaee, T.Y.; Feng, H.; Stanaway, I.B.; Schwarz, U.I.; Ritchie, M.D.; Stein, C.M.; Roden, D.M.; Smith, J.D.; et al. A genome-wide scan for common genetic variants with a large influence on warfarin maintenance dose. Blood 2008, 112, 1022–1027. [Google Scholar] [CrossRef] [PubMed]
- Motsinger-Reif, A.A.; Jorgenson, E.; Relling, M.V.; Kroetz, D.L.; Weinshilboum, R.; Cox, N.J.; Roden, D.M. Genome-wide association studies in pharmacogenomics: Successes and lessons. Pharm. Genom. 2013, 23, 383–394. [Google Scholar] [CrossRef] [PubMed]
- Scharfe, C.P.I.; Tremmel, R.; Schwab, M.; Kohlbacher, O.; Marks, D.S. Genetic variation in human drug-related genes. Genome Med. 2017, 9, 117. [Google Scholar] [CrossRef]
- Sim, S.C.; Kacevska, M.; Ingelman-Sundberg, M. Pharmacogenomics of drug-metabolizing enzymes: A recent update on clinical implications and endogenous effects. Pharm. J. 2013, 13, 1–11. [Google Scholar] [CrossRef]
- Liu, X.; Luo, X.; Jiang, C.; Zhao, H. Difficulties and challenges in the development of precision medicine. Clin. Genet. 2019, 95, 569–574. [Google Scholar] [CrossRef]
- Singh, R.S.; Gupta, B.P. Genes and genomes and unnecessary complexity in precision medicine. NPJ Genom. Med. 2020, 5, 21. [Google Scholar] [CrossRef]
- Chanfreau-Coffinier, C.; Peredo, J.; Russell, M.M.; Yano, E.M.; Hamilton, A.B.; Lerner, B.; Provenzale, D.; Knight, S.J.; Voils, C.I.; Scheuner, M.T. A logic model for precision medicine implementation informed by stakeholder views and implementation science. Genet. Med. 2019, 21, 1139–1154. [Google Scholar] [CrossRef]
- Reuter, C.M.; Kohler, J.N.; Bonner, D.; Zastrow, D.; Fernandez, L.; Dries, A.; Marwaha, S.; Davidson, J.; Brokamp, E.; Herzog, M.; et al. Yield of whole exome sequencing in undiagnosed patients facing insurance coverage barriers to genetic testing. J. Genet. Couns. 2019, 28, 1107–1118. [Google Scholar] [CrossRef]
- Klein, M.E.; Parvez, M.M.; Shin, J.G. Clinical Implementation of Pharmacogenomics for Personalized Precision Medicine: Barriers and Solutions. J. Pharm. Sci. 2017, 106, 2368–2379. [Google Scholar] [CrossRef]
- Nakagawa, H.; Fujita, M. Whole genome sequencing analysis for cancer genomics and precision medicine. Cancer Sci. 2018, 109, 513–522. [Google Scholar] [CrossRef]
- Slamon, D.J.; Leyland-Jones, B.; Shak, S.; Fuchs, H.; Paton, V.; Bajamonde, A.; Fleming, T.; Eiermann, W.; Wolter, J.; Pegram, M.; et al. Use of chemotherapy plus a monoclonal antibody against HER2 for metastatic breast cancer that overexpresses HER2. N. Engl. J. Med. 2001, 344, 783–792. [Google Scholar] [CrossRef] [PubMed]
- Karapetis, C.S.; Khambata-Ford, S.; Jonker, D.J.; O’Callaghan, C.J.; Tu, D.; Tebbutt, N.C.; Simes, R.J.; Chalchal, H.; Shapiro, J.D.; Robitaille, S.; et al. K-ras mutations and benefit from cetuximab in advanced colorectal cancer. N. Engl. J. Med. 2008, 359, 1757–1765. [Google Scholar] [CrossRef] [PubMed]
- Mullane, S.A.; Van Allen, E.M. Precision medicine for advanced prostate cancer. Curr. Opin. Urol. 2016, 26, 231–239. [Google Scholar] [CrossRef]
- Zhao, D.; Klempner, S.J.; Chao, J. Progress and challenges in HER2-positive gastroesophageal adenocarcinoma. J. Hematol. Oncol. 2019, 12, 50. [Google Scholar] [CrossRef]
- Hall, R.D.; Kudchadkar, R.R. BRAF mutations: Signaling, epidemiology, and clinical experience in multiple malignancies. Cancer Control. 2014, 21, 221–230. [Google Scholar] [CrossRef] [PubMed]
- Arber, N.; Levin, B. Chemoprevention of colorectal neoplasia: The potential for personalized medicine. Gastroenterology 2008, 134, 1224–1237. [Google Scholar] [CrossRef]
- Mennel, R.G. Precision medicine: Hype or hoax? Proc. (Bayl. Univ. Med. Cent.) 2015, 28, 397–400. [Google Scholar] [CrossRef][Green Version]
- Dagogo-Jack, I.; Shaw, A.T. Tumour heterogeneity and resistance to cancer therapies. Nat. Rev. Clin. Oncol. 2018, 15, 81–94. [Google Scholar] [CrossRef] [PubMed]
- Miller, C.A.; McMichael, J.; Dang, H.X.; Maher, C.A.; Ding, L.; Ley, T.J.; Mardis, E.R.; Wilson, R.K. Visualizing tumor evolution with the fishplot package for R. BMC Genom. 2016, 17, 880. [Google Scholar] [CrossRef] [PubMed]
- Jackson, S.E.; Chester, J.D. Personalised cancer medicine. Int. J. Cancer 2015, 137, 262–266. [Google Scholar] [CrossRef] [PubMed]
- Dent, R.; Trudeau, M.; Pritchard, K.I.; Hanna, W.M.; Kahn, H.K.; Sawka, C.A.; Lickley, L.A.; Rawlinson, E.; Sun, P.; Narod, S.A. Triple-negative breast cancer: Clinical features and patterns of recurrence. Clin. Cancer Res. 2007, 13, 4429–4434. [Google Scholar] [CrossRef] [PubMed]
- Aysola, K.; Desai, A.; Welch, C.; Xu, J.; Qin, Y.; Reddy, V.; Matthews, R.; Owens, C.; Okoli, J.; Beech, D.J.; et al. Triple Negative Breast Cancer—An Overview. Hered. Genet. 2013, 2013. [Google Scholar] [CrossRef]
- Yin, L.; Duan, J.J.; Bian, X.W.; Yu, S.C. Triple-negative breast cancer molecular subtyping and treatment progress. Breast Cancer Res. 2020, 22, 61. [Google Scholar] [CrossRef]
- Garrido-Castro, A.C.; Lin, N.U.; Polyak, K. Insights into Molecular Classifications of Triple-Negative Breast Cancer: Improving Patient Selection for Treatment. Cancer Discov. 2019, 9, 176–198. [Google Scholar] [CrossRef]
- Sporikova, Z.; Koudelakova, V.; Trojanec, R.; Hajduch, M. Genetic Markers in Triple-Negative Breast Cancer. Clin. Breast Cancer 2018, 18, e841–e850. [Google Scholar] [CrossRef]
- Bianchini, G.; Balko, J.M.; Mayer, I.A.; Sanders, M.E.; Gianni, L. Triple-negative breast cancer: Challenges and opportunities of a heterogeneous disease. Nat. Rev. Clin. Oncol. 2016, 13, 674–690. [Google Scholar] [CrossRef] [PubMed]
- Salas-Benito, D.; Perez-Gracia, J.L.; Ponz-Sarvise, M.; Rodriguez-Ruiz, M.E.; Martinez-Forero, I.; Castanon, E.; Lopez-Picazo, J.M.; Sanmamed, M.F.; Melero, I. Paradigms on Immunotherapy Combinations with Chemotherapy. Cancer Discov. 2021. [Google Scholar] [CrossRef]
- Singh, S.; Numan, A.; Maddiboyina, B.; Arora, S.; Riadi, Y.; Md, S.; Alhakamy, N.A.; Kesharwani, P. The emerging role of immune checkpoint inhibitors in the treatment of triple-negative breast cancer. Drug Discov. Today 2021. [Google Scholar] [CrossRef]
- Tiwari, P.; Blank, A.; Cui, C.; Schoenfelt, K.Q.; Zhou, G.; Xu, Y.; Khramtsova, G.; Olopade, F.; Shah, A.M.; Khan, S.A.; et al. Metabolically activated adipose tissue macrophages link obesity to triple-negative breast cancer. J. Exp. Med. 2019, 216, 1345–1358. [Google Scholar] [CrossRef] [PubMed]
- Teslow, E.A.; Mitrea, C.; Bao, B.; Mohammad, R.M.; Polin, L.A.; Dyson, G.; Purrington, K.S.; Bollig-Fischer, A. Obesity-induced MBD2_v2 expression promotes tumor-initiating triple-negative breast cancer stem cells. Mol. Oncol. 2019, 13, 894–908. [Google Scholar] [CrossRef]
- Bowers, L.W.; Rossi, E.L.; McDonell, S.B.; Doerstling, S.S.; Khatib, S.A.; Lineberger, C.G.; Albright, J.E.; Tang, X.; deGraffenried, L.A.; Hursting, S.D. Leptin Signaling Mediates Obesity-Associated CSC Enrichment and EMT in Preclinical TNBC Models. Mol. Cancer Res. 2018, 16, 869–879. [Google Scholar] [CrossRef]
- Dietze, E.C.; Chavez, T.A.; Seewaldt, V.L. Obesity and Triple-Negative Breast Cancer: Disparities, Controversies, and Biology. Am. J. Pathol. 2018, 188, 280–290. [Google Scholar] [CrossRef] [PubMed]
- Elstrom, R.L.; Bauer, D.E.; Buzzai, M.; Karnauskas, R.; Harris, M.H.; Plas, D.R.; Zhuang, H.; Cinalli, R.M.; Alavi, A.; Rudin, C.M.; et al. Akt stimulates aerobic glycolysis in cancer cells. Cancer Res. 2004, 64, 3892–3899. [Google Scholar] [CrossRef] [PubMed]
- Robey, R.B.; Hay, N. Is Akt the “Warburg kinase”?-Akt-energy metabolism interactions and oncogenesis. Semin. Cancer Biol. 2009, 19, 25–31. [Google Scholar] [CrossRef] [PubMed]
- Steenbrugge, J.; Vander Elst, N.; Demeyere, K.; De Wever, O.; Sanders, N.N.; Van Den Broeck, W.; Dirix, L.; Van Laere, S.; Meyer, E. Comparative Profiling of Metastatic 4T1- vs. Non-metastatic Py230-Based Mammary Tumors in an Intraductal Model for Triple-Negative Breast Cancer. Front. Immunol. 2019, 10, 2928. [Google Scholar] [CrossRef]
- Khosravi-Shahi, P.; Cabezon-Gutierrez, L.; Custodio-Cabello, S. Metastatic triple negative breast cancer: Optimizing treatment options, new and emerging targeted therapies. Asia Pac. J. Clin. Oncol. 2018, 14, 32–39. [Google Scholar] [CrossRef] [PubMed]
- Stover, D.G.; Parsons, H.A.; Ha, G.; Freeman, S.S.; Barry, W.T.; Guo, H.; Choudhury, A.D.; Gydush, G.; Reed, S.C.; Rhoades, J.; et al. Association of Cell-Free DNA Tumor Fraction and Somatic Copy Number Alterations With Survival in Metastatic Triple-Negative Breast Cancer. J. Clin. Oncol. 2018, 36, 543–553. [Google Scholar] [CrossRef] [PubMed]
- Shah, S.P.; Roth, A.; Goya, R.; Oloumi, A.; Ha, G.; Zhao, Y.; Turashvili, G.; Ding, J.; Tse, K.; Haffari, G.; et al. The clonal and mutational evolution spectrum of primary triple-negative breast cancers. Nature 2012, 486, 395–399. [Google Scholar] [CrossRef]
- Cancer Genome Atlas, N. Comprehensive molecular portraits of human breast tumours. Nature 2012, 490, 61–70. [Google Scholar] [CrossRef] [PubMed]
- Brewster, A.M.; Chavez-MacGregor, M.; Brown, P. Epidemiology, biology, and treatment of triple-negative breast cancer in women of African ancestry. Lancet Oncol. 2014, 15, e625–e634. [Google Scholar] [CrossRef]
- Cejalvo, J.M.; Martinez de Duenas, E.; Galvan, P.; Garcia-Recio, S.; Burgues Gasion, O.; Pare, L.; Antolin, S.; Martinello, R.; Blancas, I.; Adamo, B.; et al. Intrinsic Subtypes and Gene Expression Profiles in Primary and Metastatic Breast Cancer. Cancer Res. 2017, 77, 2213–2221. [Google Scholar] [CrossRef]
- Amabile, M.I.; Frusone, F.; De Luca, A.; Tripodi, D.; Imbimbo, G.; Lai, S.; D’Andrea, V.; Sorrenti, S.; Molfino, A. Locoregional Surgery in Metastatic Breast Cancer: Do Concomitant Metabolic Aspects Have a Role on the Management and Prognosis in this Setting? J. Pers. Med. 2020, 10, 227. [Google Scholar] [CrossRef]
- Ma, L.; Mi, Y.; Cui, S.; Wang, H.; Fu, P.; Yin, Y.; Jin, F.; Li, J.; Liu, Y.; Fan, Z.; et al. Role of locoregional surgery in patients with de novo stage IV breast cancer: Analysis of real-world data from China. Sci. Rep. 2020, 10, 18132. [Google Scholar] [CrossRef]
- Carey, L.; Winer, E.; Viale, G.; Cameron, D.; Gianni, L. Triple-negative breast cancer: Disease entity or title of convenience? Nat. Rev. Clin. Oncol. 2010, 7, 683–692. [Google Scholar] [CrossRef]
- Marra, A.; Trapani, D.; Viale, G.; Criscitiello, C.; Curigliano, G. Practical classification of triple-negative breast cancer: Intratumoral heterogeneity, mechanisms of drug resistance, and novel therapies. NPJ Breast Cancer 2020, 6, 54. [Google Scholar] [CrossRef]
- Rakha, E.A.; El-Sayed, M.E.; Green, A.R.; Lee, A.H.; Robertson, J.F.; Ellis, I.O. Prognostic markers in triple-negative breast cancer. Cancer 2007, 109, 25–32. [Google Scholar] [CrossRef] [PubMed]
- Lehmann, B.D.; Bauer, J.A.; Chen, X.; Sanders, M.E.; Chakravarthy, A.B.; Shyr, Y.; Pietenpol, J.A. Identification of human triple-negative breast cancer subtypes and preclinical models for selection of targeted therapies. J. Clin. Investig. 2011, 121, 2750–2767. [Google Scholar] [CrossRef] [PubMed]
- Chen, X.; Li, J.; Gray, W.H.; Lehmann, B.D.; Bauer, J.A.; Shyr, Y.; Pietenpol, J.A. TNBCtype: A Subtyping Tool for Triple-Negative Breast Cancer. Cancer Inform. 2012, 11, 147–156. [Google Scholar] [CrossRef] [PubMed]
- Burstein, M.D.; Tsimelzon, A.; Poage, G.M.; Covington, K.R.; Contreras, A.; Fuqua, S.A.; Savage, M.I.; Osborne, C.K.; Hilsenbeck, S.G.; Chang, J.C.; et al. Comprehensive genomic analysis identifies novel subtypes and targets of triple-negative breast cancer. Clin. Cancer Res. 2015, 21, 1688–1698. [Google Scholar] [CrossRef] [PubMed]
- Liu, Y.R.; Jiang, Y.Z.; Xu, X.E.; Yu, K.D.; Jin, X.; Hu, X.; Zuo, W.J.; Hao, S.; Wu, J.; Liu, G.Y.; et al. Comprehensive transcriptome analysis identifies novel molecular subtypes and subtype-specific RNAs of triple-negative breast cancer. Breast Cancer Res. 2016, 18, 33. [Google Scholar] [CrossRef] [PubMed]
- Lehmann, B.D.; Jovanovic, B.; Chen, X.; Estrada, M.V.; Johnson, K.N.; Shyr, Y.; Moses, H.L.; Sanders, M.E.; Pietenpol, J.A. Refinement of Triple-Negative Breast Cancer Molecular Subtypes: Implications for Neoadjuvant Chemotherapy Selection. PLoS ONE 2016, 11, e0157368. [Google Scholar] [CrossRef]
- Jiang, Y.Z.; Liu, Y.; Xiao, Y.; Hu, X.; Jiang, L.; Zuo, W.J.; Ma, D.; Ding, J.; Zhu, X.; Zou, J.; et al. Molecular subtyping and genomic profiling expand precision medicine in refractory metastatic triple-negative breast cancer: The FUTURE trial. Cell Res. 2021, 31, 178–186. [Google Scholar] [CrossRef]
- Gupta, G.K.; Collier, A.L.; Lee, D.; Hoefer, R.A.; Zheleva, V.; Siewertsz van Reesema, L.L.; Tang-Tan, A.M.; Guye, M.L.; Chang, D.Z.; Winston, J.S.; et al. Perspectives on Triple-Negative Breast Cancer: Current Treatment Strategies, Unmet Needs, and Potential Targets for Future Therapies. Cancers 2020, 12, 2392. [Google Scholar] [CrossRef]
- Hutchinson, L. Breast cancer: TNBC: Can we treat the untargetable? Nat. Rev. Clin. Oncol. 2014, 11, 379. [Google Scholar] [CrossRef]
- Pal, S.K.; Childs, B.H.; Pegram, M. Triple negative breast cancer: Unmet medical needs. Breast Cancer Res. Treat. 2011, 125, 627–636. [Google Scholar] [CrossRef] [PubMed]
- Sharma, P. Biology and Management of Patients With Triple-Negative Breast Cancer. Oncologist 2016, 21, 1050–1062. [Google Scholar] [CrossRef] [PubMed]
- Andreopoulou, E.; Schweber, S.J.; Sparano, J.A.; McDaid, H.M. Therapies for triple negative breast cancer. Expert Opin. Pharmacother. 2015, 16, 983–998. [Google Scholar] [CrossRef] [PubMed]
- Ciriello, G.; Gatza, M.L.; Beck, A.H.; Wilkerson, M.D.; Rhie, S.K.; Pastore, A.; Zhang, H.; McLellan, M.; Yau, C.; Kandoth, C.; et al. Comprehensive Molecular Portraits of Invasive Lobular Breast Cancer. Cell 2015, 163, 506–519. [Google Scholar] [CrossRef]
- Gradishar, W.J.; Anderson, B.O.; Balassanian, R.; Blair, S.L.; Burstein, H.J.; Cyr, A.; Elias, A.D.; Farrar, W.B.; Forero, A.; Giordano, S.H.; et al. NCCN Guidelines Insights: Breast Cancer, Version 1.2017. J. Natl. Compr. Canc. Netw. 2017, 15, 433–451. [Google Scholar] [CrossRef]
- Isakoff, S.J.; Mayer, E.L.; He, L.; Traina, T.A.; Carey, L.A.; Krag, K.J.; Rugo, H.S.; Liu, M.C.; Stearns, V.; Come, S.E.; et al. TBCRC009: A Multicenter Phase II Clinical Trial of Platinum Monotherapy With Biomarker Assessment in Metastatic Triple-Negative Breast Cancer. J. Clin. Oncol. 2015, 33, 1902–1909. [Google Scholar] [CrossRef]
- Theriault, R.L.; Litton, J.K.; Mittendorf, E.A.; Chen, H.; Meric-Bernstam, F.; Chavez-Macgregor, M.; Morrow, P.K.; Woodward, W.A.; Sahin, A.; Hortobagyi, G.N.; et al. Age and survival estimates in patients who have node-negative T1ab breast cancer by breast cancer subtype. Clin. Breast Cancer 2011, 11, 325–331. [Google Scholar] [CrossRef]
- Vaz-Luis, I.; Ottesen, R.A.; Hughes, M.E.; Mamet, R.; Burstein, H.J.; Edge, S.B.; Gonzalez-Angulo, A.M.; Moy, B.; Rugo, H.S.; Theriault, R.L.; et al. Outcomes by tumor subtype and treatment pattern in women with small, node-negative breast cancer: A multi-institutional study. J. Clin. Oncol. 2014, 32, 2142–2150. [Google Scholar] [CrossRef]
- Park, J.H.; Ahn, J.H.; Kim, S.B. How shall we treat early triple-negative breast cancer (TNBC): From the current standard to upcoming immuno-molecular strategies. ESMO Open 2018, 3, e000357. [Google Scholar] [CrossRef]
- Blum, J.L.; Flynn, P.J.; Yothers, G.; Asmar, L.; Geyer, C.E., Jr.; Jacobs, S.A.; Robert, N.J.; Hopkins, J.O.; O’Shaughnessy, J.A.; Dang, C.T.; et al. Anthracyclines in Early Breast Cancer: The ABC Trials-USOR 06-090, NSABP B-46-I/USOR 07132, and NSABP B-49 (NRG Oncology). J. Clin. Oncol. 2017, 35, 2647–2655. [Google Scholar] [CrossRef] [PubMed]
- Martin, M.; Rodriguez-Lescure, A.; Ruiz, A.; Alba, E.; Calvo, L.; Ruiz-Borrego, M.; Santaballa, A.; Rodriguez, C.A.; Crespo, C.; Abad, M.; et al. Molecular predictors of efficacy of adjuvant weekly paclitaxel in early breast cancer. Breast Cancer Res. Treat. 2010, 123, 149–157. [Google Scholar] [CrossRef]
- Arteaga, C.L.; Sliwkowski, M.X.; Osborne, C.K.; Perez, E.A.; Puglisi, F.; Gianni, L. Treatment of HER2-positive breast cancer: Current status and future perspectives. Nat. Rev. Clin. Oncol. 2011, 9, 16–32. [Google Scholar] [CrossRef]
- DeMichele, A.; Yee, D.; Berry, D.A.; Albain, K.S.; Benz, C.C.; Boughey, J.; Buxton, M.; Chia, S.K.; Chien, A.J.; Chui, S.Y.; et al. The Neoadjuvant Model Is Still the Future for Drug Development in Breast Cancer. Clin. Cancer Res. 2015, 21, 2911–2915. [Google Scholar] [CrossRef] [PubMed]
- Haddad, T.C.; Goetz, M.P. Landscape of neoadjuvant therapy for breast cancer. Ann. Surg. Oncol. 2015, 22, 1408–1415. [Google Scholar] [CrossRef]
- King, T.A.; Morrow, M. Surgical issues in patients with breast cancer receiving neoadjuvant chemotherapy. Nat. Rev. Clin. Oncol. 2015, 12, 335–343. [Google Scholar] [CrossRef] [PubMed]
- Nagayama, A.; Hayashida, T.; Jinno, H.; Takahashi, M.; Seki, T.; Matsumoto, A.; Murata, T.; Ashrafian, H.; Athanasiou, T.; Okabayashi, K.; et al. Comparative effectiveness of neoadjuvant therapy for HER2-positive breast cancer: A network meta-analysis. J. Natl. Cancer Inst. 2014, 106. [Google Scholar] [CrossRef] [PubMed]
- Prat, A.; Fan, C.; Fernandez, A.; Hoadley, K.A.; Martinello, R.; Vidal, M.; Viladot, M.; Pineda, E.; Arance, A.; Munoz, M.; et al. Response and survival of breast cancer intrinsic subtypes following multi-agent neoadjuvant chemotherapy. BMC Med. 2015, 13, 303. [Google Scholar] [CrossRef] [PubMed]
- Tevaarwerk, A.J.; Gray, R.J.; Schneider, B.P.; Smith, M.L.; Wagner, L.I.; Fetting, J.H.; Davidson, N.; Goldstein, L.J.; Miller, K.D.; Sparano, J.A. Survival in patients with metastatic recurrent breast cancer after adjuvant chemotherapy: Little evidence of improvement over the past 30 years. Cancer 2013, 119, 1140–1148. [Google Scholar] [CrossRef] [PubMed]
- Thompson, A.M.; Moulder-Thompson, S.L. Neoadjuvant treatment of breast cancer. Ann. Oncol. 2012, 23 (Suppl. 10), x231–x236. [Google Scholar] [CrossRef]
- Zardavas, D.; Baselga, J.; Piccart, M. Emerging targeted agents in metastatic breast cancer. Nat. Rev. Clin. Oncol. 2013, 10, 191–210. [Google Scholar] [CrossRef]
- Foulkes, W.D.; Smith, I.E.; Reis-Filho, J.S. Triple-negative breast cancer. N. Engl. J. Med. 2010, 363, 1938–1948. [Google Scholar] [CrossRef]
- Isakoff, S.J. Triple-negative breast cancer: Role of specific chemotherapy agents. Cancer J. 2010, 16, 53–61. [Google Scholar] [CrossRef]
- Santonja, A.; Sanchez-Munoz, A.; Lluch, A.; Chica-Parrado, M.R.; Albanell, J.; Chacon, J.I.; Antolin, S.; Jerez, J.M.; de la Haba, J.; de Luque, V.; et al. Triple negative breast cancer subtypes and pathologic complete response rate to neoadjuvant chemotherapy. Oncotarget 2018, 9, 26406–26416. [Google Scholar] [CrossRef]
- Asano, Y.; Kashiwagi, S.; Goto, W.; Takada, K.; Takahashi, K.; Hatano, T.; Noda, S.; Takashima, T.; Onoda, N.; Tomita, S.; et al. Prediction of survival after neoadjuvant chemotherapy for breast cancer by evaluation of tumor-infiltrating lymphocytes and residual cancer burden. BMC Cancer 2017, 17, 888. [Google Scholar] [CrossRef]
- Asano, Y.; Kashiwagi, S.; Goto, W.; Takada, K.; Takahashi, K.; Morisaki, T.; Fujita, H.; Takashima, T.; Tomita, S.; Ohsawa, M.; et al. Prediction of treatment responses to neoadjuvant chemotherapy in triple-negative breast cancer by analysis of immune checkpoint protein expression. J. Transl. Med. 2018, 16, 87. [Google Scholar] [CrossRef] [PubMed]
- Byrne, A.; Savas, P.; Sant, S.; Li, R.; Virassamy, B.; Luen, S.J.; Beavis, P.A.; Mackay, L.K.; Neeson, P.J.; Loi, S. Tissue-resident memory T cells in breast cancer control and immunotherapy responses. Nat. Rev. Clin. Oncol. 2020, 17, 341–348. [Google Scholar] [CrossRef] [PubMed]
- Dieci, M.V.; Criscitiello, C.; Goubar, A.; Viale, G.; Conte, P.; Guarneri, V.; Ficarra, G.; Mathieu, M.C.; Delaloge, S.; Curigliano, G.; et al. Prognostic value of tumor-infiltrating lymphocytes on residual disease after primary chemotherapy for triple-negative breast cancer: A retrospective multicenter study. Ann. Oncol. 2015, 26, 1518. [Google Scholar] [CrossRef] [PubMed]
- Dieci, M.V.; Radosevic-Robin, N.; Fineberg, S.; van den Eynden, G.; Ternes, N.; Penault-Llorca, F.; Pruneri, G.; D’Alfonso, T.M.; Demaria, S.; Castaneda, C.; et al. Update on tumor-infiltrating lymphocytes (TILs) in breast cancer, including recommendations to assess TILs in residual disease after neoadjuvant therapy and in carcinoma in situ: A report of the International Immuno-Oncology Biomarker Working Group on Breast Cancer. Semin. Cancer Biol. 2018, 52, 16–25. [Google Scholar] [CrossRef] [PubMed]
- Garcia-Martinez, E.; Gil, G.L.; Benito, A.C.; Gonzalez-Billalabeitia, E.; Conesa, M.A.; Garcia Garcia, T.; Garcia-Garre, E.; Vicente, V.; Ayala de la Pena, F. Tumor-infiltrating immune cell profiles and their change after neoadjuvant chemotherapy predict response and prognosis of breast cancer. Breast Cancer Res. 2014, 16, 488. [Google Scholar] [CrossRef] [PubMed]
- Ladoire, S.; Mignot, G.; Dabakuyo, S.; Arnould, L.; Apetoh, L.; Rebe, C.; Coudert, B.; Martin, F.; Bizollon, M.H.; Vanoli, A.; et al. In situ immune response after neoadjuvant chemotherapy for breast cancer predicts survival. J. Pathol. 2011, 224, 389–400. [Google Scholar] [CrossRef]
- Pinard, C.; Debled, M.; Ben Rejeb, H.; Velasco, V.; Tunon de Lara, C.; Hoppe, S.; Richard, E.; Brouste, V.; Bonnefoi, H.; MacGrogan, G. Residual cancer burden index and tumor-infiltrating lymphocyte subtypes in triple-negative breast cancer after neoadjuvant chemotherapy. Breast Cancer Res. Treat. 2020, 179, 11–23. [Google Scholar] [CrossRef]
- Denkert, C.; von Minckwitz, G.; Brase, J.C.; Sinn, B.V.; Gade, S.; Kronenwett, R.; Pfitzner, B.M.; Salat, C.; Loi, S.; Schmitt, W.D.; et al. Tumor-infiltrating lymphocytes and response to neoadjuvant chemotherapy with or without carboplatin in human epidermal growth factor receptor 2-positive and triple-negative primary breast cancers. J Clin Oncol 2015, 33, 983–991. [Google Scholar] [CrossRef] [PubMed]
- Filho, O.M.; Stover, D.G.; Asad, S.; Ansell, P.J.; Watson, M.; Loibl, S.; Geyer, C.E., Jr.; Bae, J.; Collier, K.; Cherian, M.; et al. Association of Immunophenotype With Pathologic Complete Response to Neoadjuvant Chemotherapy for Triple-Negative Breast Cancer: A Secondary Analysis of the BrighTNess Phase 3 Randomized Clinical Trial. JAMA Oncol. 2021. [Google Scholar] [CrossRef] [PubMed]
- Criscitiello, C.; Bayar, M.A.; Curigliano, G.; Symmans, F.W.; Desmedt, C.; Bonnefoi, H.; Sinn, B.; Pruneri, G.; Vicier, C.; Pierga, J.Y.; et al. A gene signature to predict high tumor-infiltrating lymphocytes after neoadjuvant chemotherapy and outcome in patients with triple-negative breast cancer. Ann. Oncol. 2018, 29, 162–169. [Google Scholar] [CrossRef]
- Karn, T.; Denkert, C.; Weber, K.E.; Holtrich, U.; Hanusch, C.; Sinn, B.V.; Higgs, B.W.; Jank, P.; Sinn, H.P.; Huober, J.; et al. Tumor mutational burden and immune infiltration as independent predictors of response to neoadjuvant immune checkpoint inhibition in early TNBC in GeparNuevo. Ann. Oncol. 2020, 31, 1216–1222. [Google Scholar] [CrossRef] [PubMed]
- Guo, S.; Loibl, S.; von Minckwitz, G.; Darb-Esfahani, S.; Lederer, B.; Denkert, C. PIK3CA H1047R Mutation Associated with a Lower Pathological Complete Response Rate in Triple-Negative Breast Cancer Patients Treated with Anthracycline-Taxane-Based Neoadjuvant Chemotherapy. Cancer Res. Treat. 2020, 52, 689–696. [Google Scholar] [CrossRef] [PubMed]
- Cain, H.; Macpherson, I.R.; Beresford, M.; Pinder, S.E.; Pong, J.; Dixon, J.M. Neoadjuvant Therapy in Early Breast Cancer: Treatment Considerations and Common Debates in Practice. Clin. Oncol. (R Coll Radiol) 2017, 29, 642–652. [Google Scholar] [CrossRef]
- Gass, P.; Lux, M.P.; Rauh, C.; Hein, A.; Bani, M.R.; Fiessler, C.; Hartmann, A.; Haberle, L.; Pretscher, J.; Erber, R.; et al. Prediction of pathological complete response and prognosis in patients with neoadjuvant treatment for triple-negative breast cancer. BMC Cancer 2018, 18, 1051. [Google Scholar] [CrossRef]
- Lee, J.S.; Yost, S.E.; Yuan, Y. Neoadjuvant Treatment for Triple Negative Breast Cancer: Recent Progresses and Challenges. Cancers 2020, 12, 1404. [Google Scholar] [CrossRef]
- Pelizzari, G.; Gerratana, L.; Basile, D.; Fanotto, V.; Bartoletti, M.; Liguori, A.; Fontanella, C.; Spazzapan, S.; Puglisi, F. Post-neoadjuvant strategies in breast cancer: From risk assessment to treatment escalation. Cancer Treat. Rev. 2019, 72, 7–14. [Google Scholar] [CrossRef]
- Symmans, W.F.; Peintinger, F.; Hatzis, C.; Rajan, R.; Kuerer, H.; Valero, V.; Assad, L.; Poniecka, A.; Hennessy, B.; Green, M.; et al. Measurement of residual breast cancer burden to predict survival after neoadjuvant chemotherapy. J. Clin. Oncol. 2007, 25, 4414–4422. [Google Scholar] [CrossRef]
- Symmans, W.F.; Wei, C.; Gould, R.; Yu, X.; Zhang, Y.; Liu, M.; Walls, A.; Bousamra, A.; Ramineni, M.; Sinn, B.; et al. Long-Term Prognostic Risk After Neoadjuvant Chemotherapy Associated With Residual Cancer Burden and Breast Cancer Subtype. J. Clin. Oncol. 2017, 35, 1049–1060. [Google Scholar] [CrossRef] [PubMed]
- Leon-Ferre, R.A.; Polley, M.Y.; Liu, H.; Gilbert, J.A.; Cafourek, V.; Hillman, D.W.; Elkhanany, A.; Akinhanmi, M.; Lilyquist, J.; Thomas, A.; et al. Impact of histopathology, tumor-infiltrating lymphocytes, and adjuvant chemotherapy on prognosis of triple-negative breast cancer. Breast Cancer Res. Treat. 2018, 167, 89–99. [Google Scholar] [CrossRef] [PubMed]
- Marra, A.; Viale, G.; Curigliano, G. Recent advances in triple negative breast cancer: The immunotherapy era. BMC Med. 2019, 17, 90. [Google Scholar] [CrossRef]
- Planes-Laine, G.; Rochigneux, P.; Bertucci, F.; Chretien, A.S.; Viens, P.; Sabatier, R.; Goncalves, A. PD-1/PD-L1 Targeting in Breast Cancer: The First Clinical Evidences Are Emerging. A Literature Review. Cancers 2019, 11, 1033. [Google Scholar] [CrossRef] [PubMed]
- Schmid, P.; Adams, S.; Rugo, H.S.; Schneeweiss, A.; Barrios, C.H.; Iwata, H.; Dieras, V.; Hegg, R.; Im, S.A.; Shaw Wright, G.; et al. Atezolizumab and Nab-Paclitaxel in Advanced Triple-Negative Breast Cancer. N. Engl. J. Med. 2018, 379, 2108–2121. [Google Scholar] [CrossRef]
- Schmid, P.; Rugo, H.S.; Adams, S.; Schneeweiss, A.; Barrios, C.H.; Iwata, H.; Dieras, V.; Henschel, V.; Molinero, L.; Chui, S.Y.; et al. Atezolizumab plus nab-paclitaxel as first-line treatment for unresectable, locally advanced or metastatic triple-negative breast cancer (IMpassion130): Updated efficacy results from a randomised, double-blind, placebo-controlled, phase 3 trial. Lancet Oncol. 2020, 21, 44–59. [Google Scholar] [CrossRef]
- FDA, K. FDA Grants Accelerated Approval to Pembrolizumab. Available online: https://www.fda.gov/drugs/resources-information-approved-drugs/fda-grants-accelerated-approval-pembrolizumab-locally-recurrent-unresectable-or-metastatic-triple (accessed on 18 June 2021).
- Heeke, A.L.; Tan, A.R. Checkpoint inhibitor therapy for metastatic triple-negative breast cancer. Cancer Metastasis Rev. 2021. [Google Scholar] [CrossRef] [PubMed]
- De Melo Gagliato, D.; Buzaid, A.C.; Perez-Garcia, J.; Cortes, J. Immunotherapy in Breast Cancer: Current Practice and Clinical Challenges. BioDrugs 2020, 34, 611–623. [Google Scholar] [CrossRef]
- Nakhjavani, M.; Hardingham, J.E.; Palethorpe, H.M.; Price, T.J.; Townsend, A.R. Druggable Molecular Targets for the Treatment of Triple Negative Breast Cancer. J. Breast Cancer 2019, 22, 341–361. [Google Scholar] [CrossRef] [PubMed]
- Bottai, G.; Raschioni, C.; Losurdo, A.; Di Tommaso, L.; Tinterri, C.; Torrisi, R.; Reis-Filho, J.S.; Roncalli, M.; Sotiriou, C.; Santoro, A.; et al. An immune stratification reveals a subset of PD-1/LAG-3 double-positive triple-negative breast cancers. Breast Cancer Res. 2016, 18, 121. [Google Scholar] [CrossRef]
- Du, W.; Yang, M.; Turner, A.; Xu, C.; Ferris, R.L.; Huang, J.; Kane, L.P.; Lu, B. TIM-3 as a Target for Cancer Immunotherapy and Mechanisms of Action. Int. J. Mol. Sci. 2017, 18, 645. [Google Scholar] [CrossRef]
- Gonzalez-Angulo, A.M.; Timms, K.M.; Liu, S.; Chen, H.; Litton, J.K.; Potter, J.; Lanchbury, J.S.; Stemke-Hale, K.; Hennessy, B.T.; Arun, B.K.; et al. Incidence and outcome of BRCA mutations in unselected patients with triple receptor-negative breast cancer. Clin. Cancer Res. 2011, 17, 1082–1089. [Google Scholar] [CrossRef] [PubMed]
- Cortesi, L.; Rugo, H.S.; Jackisch, C. An Overview of PARP Inhibitors for the Treatment of Breast Cancer. Target. Oncol. 2021. [Google Scholar] [CrossRef] [PubMed]
- Pop, L.; Suciu, I.; Ionescu, O.; Bacalbasa, N.; Ionescu, P. The role of novel poly (ADP-ribose) inhibitors in the treatment of locally advanced and metastatic Her-2/neu negative breast cancer with inherited germline BRCA1/2 mutations. A review of the literature. J. Med. Life 2021, 14, 17–20. [Google Scholar] [CrossRef] [PubMed]
- FDA, L. FDA Granted Regular Approval to Olaparib Tablets Lynparza. Available online: https://www.fda.gov/drugs/resources-information-approved-drugs/fda-approves-olaparib-germline-brca-mutated-metastatic-breast-cancer (accessed on 17 June 2021).
- FDA, T. FDA Approves Talazoparib. Available online: https://www.fda.gov/drugs/drug-approvals-and-databases/fda-approves-talazoparib-gbrcam-her2-negative-locally-advanced-or-metastatic-breast-cancer (accessed on 17 June 2021).
- Nagayama, A.; Vidula, N.; Ellisen, L.; Bardia, A. Novel antibody-drug conjugates for triple negative breast cancer. Ther. Adv. Med. Oncol. 2020, 12. [Google Scholar] [CrossRef] [PubMed]
- FDA, T. FDA Granted Accelerated Approval to Trodelvy. Available online: https://www.fda.gov/news-events/press-announcements/fda-approves-new-therapy-triple-negative-breast-cancer-has-spread-not-responded-other-treatments (accessed on 17 June 2021).
- FDA, S.G. FDA Grants Regular Approval to Sacituzumab Govitecan. Available online: https://www.fda.gov/drugs/resources-information-approved-drugs/fda-grants-regular-approval-sacituzumab-govitecan-triple-negative-breast-cancer (accessed on 17 June 2021).
- McGuinness, J.E.; Kalinsky, K. Antibody-drug conjugates in metastatic triple negative breast cancer: A spotlight on sacituzumab govitecan, ladiratuzumab vedotin, and trastuzumab deruxtecan. Expert Opin. Biol. Ther. 2021, 21, 903–913. [Google Scholar] [CrossRef]
- Wu, N.; Zhang, J.; Zhao, J.; Mu, K.; Zhang, J.; Jin, Z.; Yu, J.; Liu, J. Precision medicine based on tumorigenic signaling pathways for triple-negative breast cancer. Oncol. Lett. 2018, 16, 4984–4996. [Google Scholar] [CrossRef]
- Van Reesema, L.L.S.; Zheleva, V.; Winston, J.S.; Jansen, R.J.; O’Connor, C.F.; Isbell, A.J.; Bian, M.; Qin, R.; Bassett, P.T.; Hinson, V.J.; et al. SIAH and EGFR, Two RAS Pathway Biomarkers, are Highly Prognostic in Locally Advanced and Metastatic Breast Cancer. EBioMedicine 2016, 11, 183–198. [Google Scholar] [CrossRef]
- Bayraktar, S.; Gluck, S. Molecularly targeted therapies for metastatic triple-negative breast cancer. Breast Cancer Res. Treat. 2013, 138, 21–35. [Google Scholar] [CrossRef]
- Hsiao, Y.C.; Yeh, M.H.; Chen, Y.J.; Liu, J.F.; Tang, C.H.; Huang, W.C. Lapatinib increases motility of triple-negative breast cancer cells by decreasing miRNA-7 and inducing Raf-1/MAPK-dependent interleukin-6. Oncotarget 2015, 6, 37965–37978. [Google Scholar] [CrossRef] [PubMed]
- Bartholomeusz, C.; Xie, X.; Pitner, M.K.; Kondo, K.; Dadbin, A.; Lee, J.; Saso, H.; Smith, P.D.; Dalby, K.N.; Ueno, N.T. MEK Inhibitor Selumetinib (AZD6244; ARRY-142886) Prevents Lung Metastasis in a Triple-Negative Breast Cancer Xenograft Model. Mol. Cancer Ther. 2015, 14, 2773–2781. [Google Scholar] [CrossRef] [PubMed]
- Loi, S.; Dushyanthen, S.; Beavis, P.A.; Salgado, R.; Denkert, C.; Savas, P.; Combs, S.; Rimm, D.L.; Giltnane, J.M.; Estrada, M.V.; et al. Correction: RAS/MAPK Activation Is Associated with Reduced Tumor-Infiltrating Lymphocytes in Triple-Negative Breast Cancer: Therapeutic Cooperation Between MEK and PD-1/PD-L1 Immune Checkpoint Inhibitors. Clin. Cancer Res. 2019, 25, 1437. [Google Scholar] [CrossRef] [PubMed]
- Khan, M.A.; Jain, V.K.; Rizwanullah, M.; Ahmad, J.; Jain, K. PI3K/AKT/mTOR pathway inhibitors in triple-negative breast cancer: A review on drug discovery and future challenges. Drug Discov. Today 2019, 24, 2181–2191. [Google Scholar] [CrossRef]
- Cossu-Rocca, P.; Orru, S.; Muroni, M.R.; Sanges, F.; Sotgiu, G.; Ena, S.; Pira, G.; Murgia, L.; Manca, A.; Uras, M.G.; et al. Analysis of PIK3CA Mutations and Activation Pathways in Triple Negative Breast Cancer. PLoS ONE 2015, 10, e0141763. [Google Scholar] [CrossRef]
- Shrivastava, S.; Kulkarni, P.; Thummuri, D.; Jeengar, M.K.; Naidu, V.G.; Alvala, M.; Redddy, G.B.; Ramakrishna, S. Piperlongumine, an alkaloid causes inhibition of PI3 K/Akt/mTOR signaling axis to induce caspase-dependent apoptosis in human triple-negative breast cancer cells. Apoptosis 2014, 19, 1148–1164. [Google Scholar] [CrossRef]
- Killock, D. AKT inhibition improves OS in TNBC. Nat. Rev. Clin. Oncol. 2020, 17, 135. [Google Scholar] [CrossRef]
- Lin, J.; Sampath, D.; Nannini, M.A.; Lee, B.B.; Degtyarev, M.; Oeh, J.; Savage, H.; Guan, Z.; Hong, R.; Kassees, R.; et al. Targeting activated Akt with GDC-0068, a novel selective Akt inhibitor that is efficacious in multiple tumor models. Clin. Cancer Res. 2013, 19, 1760–1772. [Google Scholar] [CrossRef]
- Baselga, J.; Campone, M.; Piccart, M.; Burris, H.A., 3rd; Rugo, H.S.; Sahmoud, T.; Noguchi, S.; Gnant, M.; Pritchard, K.I.; Lebrun, F.; et al. Everolimus in postmenopausal hormone-receptor-positive advanced breast cancer. N. Engl. J. Med. 2012, 366, 520–529. [Google Scholar] [CrossRef]
- Ganesan, P.; Moulder, S.; Lee, J.J.; Janku, F.; Valero, V.; Zinner, R.G.; Naing, A.; Fu, S.; Tsimberidou, A.M.; Hong, D.; et al. Triple-negative breast cancer patients treated at MD Anderson Cancer Center in phase I trials: Improved outcomes with combination chemotherapy and targeted agents. Mol. Cancer Ther. 2014, 13, 3175–3184. [Google Scholar] [CrossRef]
- Singh, J.; Novik, Y.; Stein, S.; Volm, M.; Meyers, M.; Smith, J.; Omene, C.; Speyer, J.; Schneider, R.; Jhaveri, K.; et al. Phase 2 trial of everolimus and carboplatin combination in patients with triple negative metastatic breast cancer. Breast Cancer Res. 2014, 16, R32. [Google Scholar] [CrossRef]
- Beuvink, I.; Boulay, A.; Fumagalli, S.; Zilbermann, F.; Ruetz, S.; O’Reilly, T.; Natt, F.; Hall, J.; Lane, H.A.; Thomas, G. The mTOR inhibitor RAD001 sensitizes tumor cells to DNA-damaged induced apoptosis through inhibition of p21 translation. Cell 2005, 120, 747–759. [Google Scholar] [CrossRef] [PubMed]
- Ibrahim, Y.H.; Garcia-Garcia, C.; Serra, V.; He, L.; Torres-Lockhart, K.; Prat, A.; Anton, P.; Cozar, P.; Guzman, M.; Grueso, J.; et al. PI3K inhibition impairs BRCA1/2 expression and sensitizes BRCA-proficient triple-negative breast cancer to PARP inhibition. Cancer Discov. 2012, 2, 1036–1047. [Google Scholar] [CrossRef]
- Bhola, N.E.; Jansen, V.M.; Koch, J.P.; Li, H.; Formisano, L.; Williams, J.A.; Grandis, J.R.; Arteaga, C.L. Treatment of Triple-Negative Breast Cancer with TORC1/2 Inhibitors Sustains a Drug-Resistant and Notch-Dependent Cancer Stem Cell Population. Cancer Res. 2016, 76, 440–452. [Google Scholar] [CrossRef] [PubMed]
- Giuli, M.V.; Giuliani, E.; Screpanti, I.; Bellavia, D.; Checquolo, S. Notch Signaling Activation as a Hallmark for Triple-Negative Breast Cancer Subtype. J. Oncol. 2019, 2019, 8707053. [Google Scholar] [CrossRef] [PubMed]
- Qiu, M.; Peng, Q.; Jiang, I.; Carroll, C.; Han, G.; Rymer, I.; Lippincott, J.; Zachwieja, J.; Gajiwala, K.; Kraynov, E.; et al. Specific inhibition of Notch1 signaling enhances the antitumor efficacy of chemotherapy in triple negative breast cancer through reduction of cancer stem cells. Cancer Lett. 2013, 328, 261–270. [Google Scholar] [CrossRef]
- Zhu, H.; Bhaijee, F.; Ishaq, N.; Pepper, D.J.; Backus, K.; Brown, A.S.; Zhou, X.; Miele, L. Correlation of Notch1, pAKT and nuclear NF-kappaB expression in triple negative breast cancer. Am. J. Cancer Res. 2013, 3, 230–239. [Google Scholar] [PubMed]
- Majumder, S.; Crabtree, J.S.; Golde, T.E.; Minter, L.M.; Osborne, B.A.; Miele, L. Targeting Notch in oncology: The path forward. Nat. Rev. Drug Discov. 2021, 20, 125–144. [Google Scholar] [CrossRef] [PubMed]
- Lehal, R.; Zaric, J.; Vigolo, M.; Urech, C.; Frismantas, V.; Zangger, N.; Cao, L.; Berger, A.; Chicote, I.; Loubery, S.; et al. Pharmacological disruption of the Notch transcription factor complex. Proc. Natl. Acad. Sci. USA 2020, 117, 16292–16301. [Google Scholar] [CrossRef] [PubMed]
- Deng, J.; Wang, E.S.; Jenkins, R.W.; Li, S.; Dries, R.; Yates, K.; Chhabra, S.; Huang, W.; Liu, H.; Aref, A.R.; et al. CDK4/6 Inhibition Augments Antitumor Immunity by Enhancing T-cell Activation. Cancer Discov. 2018, 8, 216–233. [Google Scholar] [CrossRef]
- Goel, S.; DeCristo, M.J.; Watt, A.C.; BrinJones, H.; Sceneay, J.; Li, B.B.; Khan, N.; Ubellacker, J.M.; Xie, S.; Metzger-Filho, O.; et al. CDK4/6 inhibition triggers anti-tumour immunity. Nature 2017, 548, 471–475. [Google Scholar] [CrossRef]
- Bhattarai, S.; Klimov, S.; Mittal, K.; Krishnamurti, U.; Li, X.B.; Oprea-Ilies, G.; Wetherilt, C.S.; Riaz, A.; Aleskandarany, M.A.; Green, A.R.; et al. Prognostic Role of Androgen Receptor in Triple Negative Breast Cancer: A Multi-Institutional Study. Cancers 2019, 11, 995. [Google Scholar] [CrossRef] [PubMed]
- Chan, J.J.; Tan, T.J.Y.; Dent, R.A. Novel therapeutic avenues in triple-negative breast cancer: PI3K/AKT inhibition, androgen receptor blockade, and beyond. Ther. Adv. Med. Oncol. 2019, 11. [Google Scholar] [CrossRef] [PubMed]
- Ricciardi, G.R.; Adamo, B.; Ieni, A.; Licata, L.; Cardia, R.; Ferraro, G.; Franchina, T.; Tuccari, G.; Adamo, V. Androgen Receptor (AR), E-Cadherin, and Ki-67 as Emerging Targets and Novel Prognostic Markers in Triple-Negative Breast Cancer (TNBC) Patients. PLoS ONE 2015, 10, e0128368. [Google Scholar] [CrossRef]
- Bonnefoi, H.; Grellety, T.; Tredan, O.; Saghatchian, M.; Dalenc, F.; Mailliez, A.; L’Haridon, T.; Cottu, P.; Abadie-Lacourtoisie, S.; You, B.; et al. A phase II trial of abiraterone acetate plus prednisone in patients with triple-negative androgen receptor positive locally advanced or metastatic breast cancer (UCBG 12-1). Ann. Oncol. 2016, 27, 812–818. [Google Scholar] [CrossRef]
- Gucalp, A.; Tolaney, S.; Isakoff, S.J.; Ingle, J.N.; Liu, M.C.; Carey, L.A.; Blackwell, K.; Rugo, H.; Nabell, L.; Forero, A.; et al. Phase II trial of bicalutamide in patients with androgen receptor-positive, estrogen receptor-negative metastatic Breast Cancer. Clin. Cancer Res. 2013, 19, 5505–5512. [Google Scholar] [CrossRef] [PubMed]
- Madu, C.O.; Wang, S.; Madu, C.O.; Lu, Y. Angiogenesis in Breast Cancer Progression, Diagnosis, and Treatment. J. Cancer 2020, 11, 4474–4494. [Google Scholar] [CrossRef] [PubMed]
- Ribatti, D.; Nico, B.; Ruggieri, S.; Tamma, R.; Simone, G.; Mangia, A. Angiogenesis and Antiangiogenesis in Triple-Negative Breast cancer. Transl. Oncol. 2016, 9, 453–457. [Google Scholar] [CrossRef]
- Hutchinson, L. Breast cancer: BEATRICE bevacizumab trial—Every cloud has a silver lining. Nat. Rev. Clin. Oncol. 2013, 10, 548. [Google Scholar] [CrossRef]
- Barroso-Sousa, R.; Tolaney, S.M. Clinical Development of New Antibody-Drug Conjugates in Breast Cancer: To Infinity and Beyond. BioDrugs 2021, 35, 159–174. [Google Scholar] [CrossRef]
- Islam, R.; Lam, K.W. Recent progress in small molecule agents for the targeted therapy of triple-negative breast cancer. Eur. J. Med. Chem. 2020, 207, 112812. [Google Scholar] [CrossRef]
- Emens, L.A. Breast Cancer Immunotherapy: Facts and Hopes. Clin. Cancer Res. 2018, 24, 511–520. [Google Scholar] [CrossRef] [PubMed]
- Vikas, P.; Borcherding, N.; Zhang, W. The clinical promise of immunotherapy in triple-negative breast cancer. Cancer Manag. Res. 2018, 10, 6823–6833. [Google Scholar] [CrossRef]
- Zhao, H.; Yang, Q.; Hu, Y.; Zhang, J. Antitumor effects and mechanisms of olaparib in combination with carboplatin and BKM120 on human triplenegative breast cancer cells. Oncol. Rep. 2018, 40, 3223–3234. [Google Scholar] [CrossRef] [PubMed]
- Cardillo, T.M.; Sharkey, R.M.; Rossi, D.L.; Arrojo, R.; Mostafa, A.A.; Goldenberg, D.M. Synthetic Lethality Exploitation by an Anti-Trop-2-SN-38 Antibody-Drug Conjugate, IMMU-132, Plus PARP Inhibitors in BRCA1/2-wild-type Triple-Negative Breast Cancer. Clin. Cancer Res. 2017, 23, 3405–3415. [Google Scholar] [CrossRef] [PubMed]
- Bergin, A.R.T.; Loi, S. Triple-negative breast cancer: Recent treatment advances. F1000Res 2019, 8. [Google Scholar] [CrossRef]
- Higuchi, T.; Flies, D.B.; Marjon, N.A.; Mantia-Smaldone, G.; Ronner, L.; Gimotty, P.A.; Adams, S.F. CTLA-4 Blockade Synergizes Therapeutically with PARP Inhibition in BRCA1-Deficient Ovarian Cancer. Cancer Immunol. Res. 2015, 3, 1257–1268. [Google Scholar] [CrossRef] [PubMed]
- Wu, L.; Meng, F.; Dong, L.; Block, C.J.; Mitchell, A.V.; Wu, J.; Jang, H.; Chen, W.; Polin, L.; Yang, Q.; et al. Disulfiram and BKM120 in Combination with Chemotherapy Impede Tumor Progression and Delay Tumor Recurrence in Tumor Initiating Cell-Rich TNBC. Sci. Rep. 2019, 9, 236. [Google Scholar] [CrossRef]
- Hossain, F.; Sorrentino, C.; Ucar, D.A.; Peng, Y.; Matossian, M.; Wyczechowska, D.; Crabtree, J.; Zabaleta, J.; Morello, S.; Del Valle, L.; et al. Notch Signaling Regulates Mitochondrial Metabolism and NF-kappaB Activity in Triple-Negative Breast Cancer Cells via IKKalpha-Dependent Non-canonical Pathways. Front. Oncol. 2018, 8, 575. [Google Scholar] [CrossRef]
- Diluvio, G.; Del Gaudio, F.; Giuli, M.V.; Franciosa, G.; Giuliani, E.; Palermo, R.; Besharat, Z.M.; Pignataro, M.G.; Vacca, A.; d’Amati, G.; et al. NOTCH3 inactivation increases triple negative breast cancer sensitivity to gefitinib by promoting EGFR tyrosine dephosphorylation and its intracellular arrest. Oncogenesis 2018, 7, 42. [Google Scholar] [CrossRef]
- Kim, K.; Skora, A.D.; Li, Z.; Liu, Q.; Tam, A.J.; Blosser, R.L.; Diaz, L.A., Jr.; Papadopoulos, N.; Kinzler, K.W.; Vogelstein, B.; et al. Eradication of metastatic mouse cancers resistant to immune checkpoint blockade by suppression of myeloid-derived cells. Proc. Natl. Acad. Sci. USA 2014, 111, 11774–11779. [Google Scholar] [CrossRef] [PubMed]
- Pauli, C.; Hopkins, B.D.; Prandi, D.; Shaw, R.; Fedrizzi, T.; Sboner, A.; Sailer, V.; Augello, M.; Puca, L.; Rosati, R.; et al. Personalized In Vitro and In Vivo Cancer Models to Guide Precision Medicine. Cancer Discov. 2017, 7, 462–477. [Google Scholar] [CrossRef]
- Tsimberidou, A.M.; Wen, S.; Hong, D.S.; Wheler, J.J.; Falchook, G.S.; Fu, S.; Piha-Paul, S.; Naing, A.; Janku, F.; Aldape, K.; et al. Personalized medicine for patients with advanced cancer in the phase I program at MD Anderson: Validation and landmark analyses. Clin. Cancer Res. 2014, 20, 4827–4836. [Google Scholar] [CrossRef] [PubMed]
- Aref, A.R.; Campisi, M.; Ivanova, E.; Portell, A.; Larios, D.; Piel, B.P.; Mathur, N.; Zhou, C.; Coakley, R.V.; Bartels, A.; et al. 3D microfluidic ex vivo culture of organotypic tumor spheroids to model immune checkpoint blockade. Lab Chip 2018, 18, 3129–3143. [Google Scholar] [CrossRef] [PubMed]
- Low, S.K.; Zembutsu, H.; Nakamura, Y. Breast cancer: The translation of big genomic data to cancer precision medicine. Cancer Sci. 2018, 109, 497–506. [Google Scholar] [CrossRef]
- Sehgal, K.; Portell, A.; Ivanova, E.V.; Lizotte, P.H.; Mahadevan, N.R.; Greene, J.R.; Vajdi, A.; Gurjao, C.; Teceno, T.; Taus, L.J.; et al. Dynamic single-cell RNA sequencing identifies immunotherapy persister cells following PD-1 blockade. J. Clin. Investig. 2021, 131. [Google Scholar] [CrossRef] [PubMed]
- Mertins, P.; Mani, D.R.; Ruggles, K.V.; Gillette, M.A.; Clauser, K.R.; Wang, P.; Wang, X.; Qiao, J.W.; Cao, S.; Petralia, F.; et al. Proteogenomics connects somatic mutations to signalling in breast cancer. Nature 2016, 534, 55–62. [Google Scholar] [CrossRef]
- Wu, Q.; Ba-Alawi, W.; Deblois, G.; Cruickshank, J.; Duan, S.; Lima-Fernandes, E.; Haight, J.; Tonekaboni, S.A.M.; Fortier, A.M.; Kuasne, H.; et al. GLUT1 inhibition blocks growth of RB1-positive triple negative breast cancer. Nat. Commun. 2020, 11, 4205. [Google Scholar] [CrossRef] [PubMed]
| Title | Phase | Status | Age | ID |
|---|---|---|---|---|
| Evaluation of IPI-549 combined with front-line treatments in patients with TNBCr or Renal Cell Carcinoma (MARIO-3) | Phase II | Active | 18 and over | NCT03961698 |
| Testing the addition of Copanlisib to Eribulin for the Treatment of Advanced-Stage Triple Negative Breast Cancer | Phase I/II | Active | 18 and over | NCT04345913 |
| Study of Pembrolizumab (MK-3475) plus chemotherapy vs. placebo plus chemotherapy for previously untreated locally recurrent inoperable or metastatic TNBC | Phase III | Active, not recruiting | 18 and over | NCT02819518 |
| Atorvastatin in treating patients with stage IIb-III TNBC who did not achieve a PCR after receiving neoadjuvant chemotherapy | Phase II | Active | 18 and over | NCT03872388 |
| A Study of Atezolizumab in combination with Nab-Paclitaxel compared with placebo with Nab-paclitaxel for participants with previously untreated metastatic TNBC | Phase III | Active, not recruiting | 18 and over | NCT02425891 |
| Avelumab With Binimetinib, Sacituzumab Govitecan, or Liposomal Doxorubicin in treating patients with stage IV or unresectable recurrent TNBC (InCITe) | Phase II | Active | 18 and over | NCT03971409 |
| A Study of Atezolizumab and Paclitaxel versus placebo and paclitaxel in participants with previously untreated locally advanced or metastatic TNBC (Impassion131) | Phase III | Active, not recruiting | 18 and over | NCT03125902 |
| A Study evaluating the efficacy and safety of multiple immunotherapy-based treatment combinations in patients with metastatic or inoperable locally advanced TNBC (Morpheus-TNBC) | Phase I/II | Active | 18 and over | NCT03424005 |
| A Study of Cobimetinib plus paclitaxel, Cobimetinib plus Atezolizumab plus Paclitaxel, or Cobimetinib plus Atezolizumab plus Nab-Paclitaxel as initial treatment for participants with TNBC that has spread | Phase II | Active, not recruiting | 18 and over | NCT02322814 |
| Women’s MoonShot: Neoadjuvant treatment with PaCT for patients with locally advanced TNBC | Phase II | Active | 18 and over | NCT02593175 |
| A phase II study of Nivolumab in combination with Cabozantinib for metastatic TNBC | Phase II | Active, not recruiting | 18 and over | NCT03316586 |
| FUSCC Refractory TNBC Umbrella (FUTURE) | Phase I/II | Active | 18 to 75 years | NCT03805399 |
| Peri-Operative Ipilimumab + Nivolumab and Cryoablation in women with TNBC | Phase II | Active | 18 and over | NCT03546686 |
| Trilaciclib (G1T28), a CDK 4/6 Inhibitor, in Combination with Gemcitabine and Carboplatin in metastatic TNBC | Phase II | Active, not recruiting | 18 and over | NCT02978716 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Hossain, F.; Majumder, S.; David, J.; Miele, L. Precision Medicine and Triple-Negative Breast Cancer: Current Landscape and Future Directions. Cancers 2021, 13, 3739. https://doi.org/10.3390/cancers13153739
Hossain F, Majumder S, David J, Miele L. Precision Medicine and Triple-Negative Breast Cancer: Current Landscape and Future Directions. Cancers. 2021; 13(15):3739. https://doi.org/10.3390/cancers13153739
Chicago/Turabian StyleHossain, Fokhrul, Samarpan Majumder, Justin David, and Lucio Miele. 2021. "Precision Medicine and Triple-Negative Breast Cancer: Current Landscape and Future Directions" Cancers 13, no. 15: 3739. https://doi.org/10.3390/cancers13153739
APA StyleHossain, F., Majumder, S., David, J., & Miele, L. (2021). Precision Medicine and Triple-Negative Breast Cancer: Current Landscape and Future Directions. Cancers, 13(15), 3739. https://doi.org/10.3390/cancers13153739






