Identification of a Novel Tumor Microenvironment Prognostic Signature for Advanced-Stage Serous Ovarian Cancer
Abstract
:Simple Summary
Abstract
1. Introduction
2. Materials and Methods
2.1. Data Download and Preprocessing
2.2. Identification of Molecular Subtypes Using Non-Negative Matrix Factorization (NMF) Algorithm
2.3. Comparison of Immune Scores between Clusters
2.4. Sample Preparation
2.5. Lasso Regression Analysis
2.6. Construction of Nomogram Combined with Risk Score (RS) and Clinical Features
2.7. Prediction Model Evaluation
2.8. Immunotherapy Prediction
3. Results
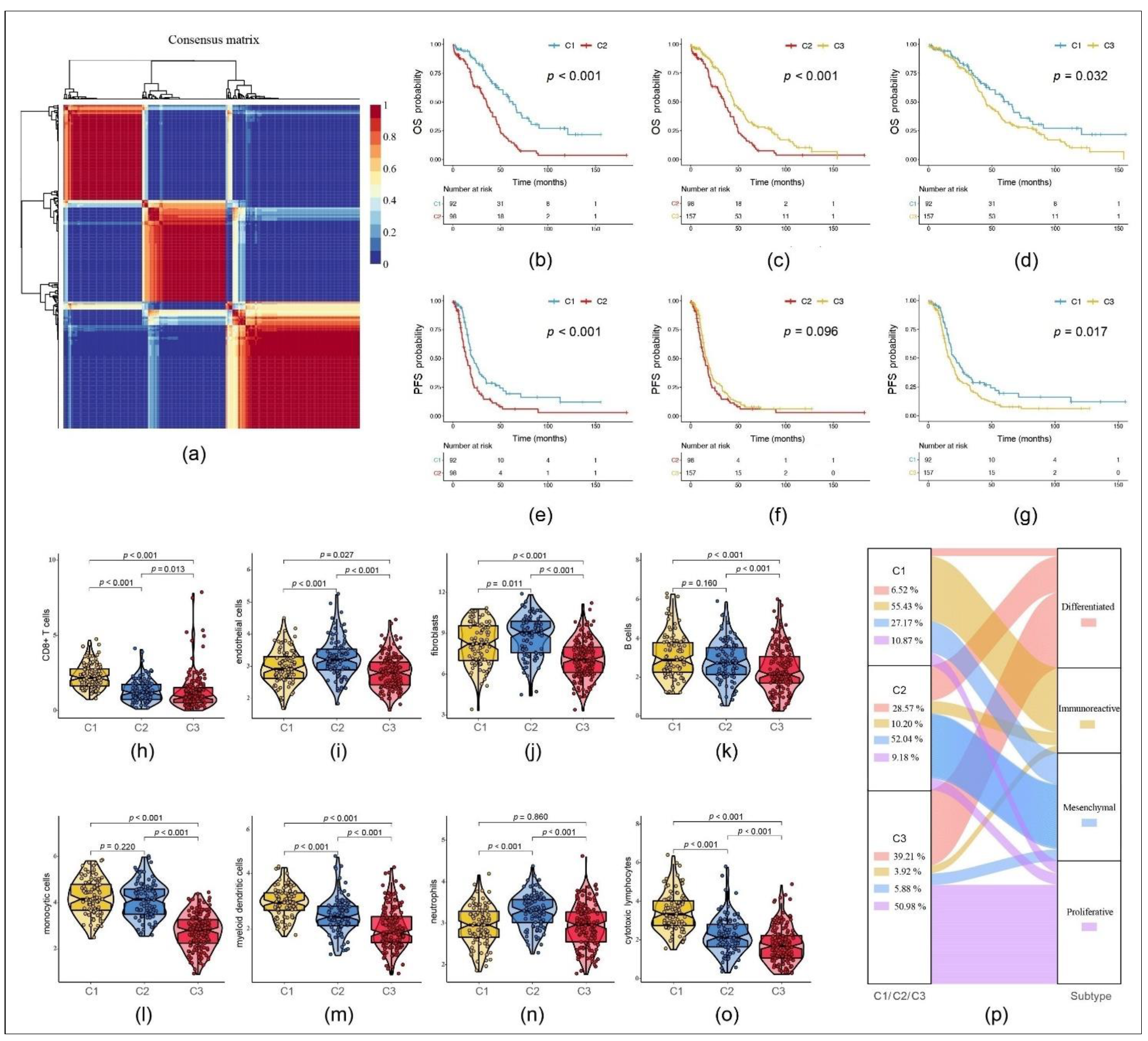
3.1. Immune Scores Indicate Different Patterns According to Molecular Subtypes in OC
3.2. Establishment of a Predictive RS Based on TME-Related Genes
−(5.9449 × SNRPA1)
− (6.9887 × CCL19)
− (4.4685 × CXCL11)
− (6.9226 × CDC5L)
− (6.1777 × APCDD1)
− (8.9229 × LPAR2)
+ (0.2541 × PI3)
+ (1.7480 × PLEKHF1)
+ (5.4819 × CCDC80)
+ (0.3243 × CPXM1)
+ (0.7416 × CTAG2).
3.3. RS Assessment in Subgoups Presenting Different Clinical Features or Mutation Statuses
3.4. TME-Related Genes Correlate with Clinical Outcome
3.5. The 11-Gene Signature Risk Model as a Novel Predictive RS in OC
3.6. The 11-Gene Signature Risk Model Validation in Another Gynecological Cancer
(NES = 0.4600, q = 0.0049),
KEGG_T_CELL_RECEPTOR_SIGNALING_PATHWAY
(NES = 0.5604, q = 0.0030),
REACTOME_ADAPTIVE_IMMUNE_SYSTEM
(NES = 0.4685, q = 0.0010),
REACTOME_CYTOKINE_SIGNALING_IN_IMMUNE_SYSTEM
(NES = 0.4100, q = 0.0030),
REACTOME_METABOLISM_OF_LIPIDS
(NES = 0.4862, q = 0.0010).
3.7. Prediction of Response to Immunotherapy Based on 11-Gene Risk Model
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
Abbreviations
| (q-value) | adjusted p-value |
| (CESC) | cervical squamous cell carcinoma |
| (CR) | complete response |
| (C-index) | concordance index |
| (CI) | confidence interval |
| (C1/2/3) | cluster 1/2/3 |
| (DCA) | decision curve analysis |
| (EMT) | epithelial–mesenchymal transition |
| (ESTIMATE) | Estimation of Stromal and Immune cells in Malignant Tumor tissues using Expression data |
| (FIGO) | Fédération Internationale de Gynécologie et d’Obstétrique |
| (HR) | hazard ratio |
| (HRG) | high-risk group |
| (IC-score) | immune cell score |
| (ICGC) | International Cancer Genome Consortium |
| (KM) | Kaplan–Meier |
| (LRG) | low-risk group |
| (MCP) | Microenvironment Cell Populations |
| (NEO) | neoantigen |
| (NMF) | non-negative matrix factorization |
| (NES) | normalized enrichment score |
| (OC) | ovarian cancer |
| (OS) | overall survival |
| (PR) | partial response |
| (PD-L1) | programmed death ligand 1 |
| (PFS) | progression-free survival |
| (PD) | progressive disease |
| (ROC) | receiver operating characteristic |
| (rss) | residual sum of squares |
| (RMS) | restricted mean survival |
| (RS) | risk score |
| (SOC) | serous ovarian cancer |
| (SD) | stable disease |
| (TCGA) | The Cancer Genome Atlas |
| (TME) | tumor microenvironment |
| (TMB) | tumor mutation burden |
References
- National Cancer Institute: Surveillance, Epidemiology, and End Results Program. Cancer Stat Facts: Ovarian Cancer. 2020. Available online: https://seer.cancer.gov/statfacts/html/ovary.html (accessed on 1 May 2021).
- Waldmann, A.; Eisemann, N.; Katalinic, A. Epidemiology of Malignant Cervical, Corpus Uteri and Ovarian Tumours—Current Data and Epidemiological Trends. Geburtshilfe Frauenheilkd 2013, 73, 123–129. [Google Scholar] [CrossRef] [Green Version]
- Menon, U.; Karpinskyj, C.; Gentry-Maharaj, A. Ovarian Cancer Prevention and Screening. Obstet. Gynecol. 2018, 131, 909–927. [Google Scholar] [CrossRef] [Green Version]
- Mok, S.C.; Bonome, T.; Vathipadiekal, V.; Bell, A.; Johnson, M.E.; Wong, K.; Park, D.C.; Hao, K.; Yip, D.K.P.; Donninger, H.; et al. A gene signature predictive for outcome in advanced ovarian cancer identifies a survival factor: Microfibril-associated glycoprotein 2. Cancer Cell 2009, 16, 521–532. [Google Scholar] [CrossRef] [Green Version]
- Lheureux, S.; Braunstein, M.; Oza, A.M. Epithelial ovarian cancer: Evolution of management in the era of precision medicine. CA Cancer J. Clin. 2019, 69, 280–304. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Lorusso, D.; Ceni, V.; Daniele, G.; Salutari, V.; Pietragalla, A.; Muratore, M.; Nero, C.; Ciccarone, F.; Scambia, G. Newly diagnosed ovarian cancer: Which first-line treatment? Cancer Treat. Rev. 2020, 91, 102111. [Google Scholar] [CrossRef]
- Kulbe, H.; Chakravarty, P.; Leinster, D.A.; Charles, K.A.; Kwong, J.; Thompson, R.G.; Coward, J.I.; Schioppa, T.; Robinson, S.C.; Gallagher, W.M.; et al. A dynamic inflammatory cytokine network in the human ovarian cancer microenvironment. Cancer Res. 2012, 72, 66–75. [Google Scholar] [CrossRef] [Green Version]
- Reinartz, S.; Schumann, T.; Finkernagel, F.; Wortmann, A.; Jansen, J.M.; Meissner, W.; Krause, M.; Schwoerer, A.M.; Wagner, U.; Mueller-Bruesselbach, S.; et al. Mixed-polarization phenotype of ascites-associated macrophages in human ovarian carcinoma: Correlation of CD163 expression, cytokine levels and early relapse. Int. J. Cancer 2014, 134, 32–42. [Google Scholar] [CrossRef] [PubMed]
- Hui, L.; Chen, Y. Tumor microenvironment: Sanctuary of the devil. Cancer Lett. 2015, 368, 7–13. [Google Scholar] [CrossRef]
- Whiteside, T.L. The tumor microenvironment and its role in promoting tumor growth. Oncogene 2008, 27, 5904–5912. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Nomelini, R.S.; Borges Junior, L.E.; de Lima, C.A.; Chiovato, A.F.C.; Micheli, D.C.; Tavares-Murta, B.M.; Murta, E.F.C. TNF-R2 in tumor microenvironment as prognostic factor in epithelial ovarian cancer. Clin. Exp. Med. 2018, 18, 547–554. [Google Scholar] [CrossRef]
- Yin, M.; Li, X.; Tan, S.; Zhou, H.J.; Ji, W.; Bellone, S.; Xu, X.; Zhang, H.; Santin, A.D.; Lou, G.; et al. Tumor-associated macrophages drive spheroid formation during early transcoelomic metastasis of ovarian cancer. J. Clin. Investig. 2016, 126, 4157–4173. [Google Scholar] [CrossRef] [Green Version]
- Pollard, J.W. Tumour-educated macrophages promote tumour progression and metastasis. Nat. Rev. Cancer 2004, 4, 71–78. [Google Scholar] [CrossRef]
- Ma, Y.; Shurin, G.V.; Peiyuan, Z.; Shurinet, M.R. Dendritic cells in the cancer microenvironment. J. Cancer 2013, 4, 36–44. [Google Scholar] [CrossRef] [Green Version]
- Feng, J.; Guo, J.; Zhao, P.; Shen, J.; Chai, B.; Wang, J. mTOR up-regulation of SNRPA1 contributes to hepatocellular carcinoma development. Biosci. Rep. 2020, 40, BSR20193815. [Google Scholar] [CrossRef] [PubMed]
- Liu, J.F.; Herold, C.; Gray, K.P.; Penson, R.T.; Horowitz, N.; Konstantinopoulos, P.A.; Castro, C.M.; Hill, S.J.; Curtis, J.; Luo, W.; et al. Assessment of Combined Nivolumab and Bevacizumab in Relapsed Ovarian Cancer: A Phase 2 Clinical Trial. JAMA Oncol. 2019, 5, 1731–1738. [Google Scholar] [CrossRef]
- Burger, R.A.; Brady, M.F.; Bookman, M.A.; Fleming, G.F.; Monk, B.J.; Huang, H.; Mannel, R.S.; Homesley, H.D.; Fowler, J.; Greer, B.E.; et al. Incorporation of bevacizumab in the primary treatment of ovarian cancer. N. Engl. J. Med. 2011, 365, 2473–2483. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- U.S. Food & Drug Administration. FDA Grants Accelerated Approval to New Treatment for Advanced Ovarian Cancer. 2016. Available online: https://www.fda.gov/news-events/press-announcements/fda-grants-accelerated-approval-new-treatment-advanced-ovarian-cancer (accessed on 1 May 2021).
- Jimenez-Sanchez, A.; Memon, D.; Pourpe, S.; Veeraraghavan, H.; Li, Y.; Vargas, H.A.; Gill, M.B.; Park, K.J.; Zivanovic, O.; Konner, J.; et al. Heterogeneous Tumor-Immune Microenvironments among Differentially Growing Metastases in an Ovarian Cancer Patient. Cell 2017, 170, 927–938.e20. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Tirosh, I.; Izar, B.; Prakadan, S.M.; Wadsworth, M.C.; Treacy, D.; Trombetta, J.J.; Rotem, A.; Rodman, C.; Lian, C.; Murphy, G.; et al. Dissecting the multicellular ecosystem of metastatic melanoma by single-cell RNA-seq. Science 2016, 352, 189–196. [Google Scholar] [CrossRef] [Green Version]
- Rooney, M.S.; Shukla, S.A.; Wu, C.J.; Getz, G.; Hacohen, N. Molecular and genetic properties of tumors associated with local immune cytolytic activity. Cell 2015, 160, 48–61. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Newman, A.M.; Liu, C.L.; Green, M.R.; Gentles, A.J.; Feng, W.; Xu, Y.; Hoang, C.D.; Diehn, M.; Alizadeh, A.A. Robust enumeration of cell subsets from tissue expression profiles. Nat. Methods 2015, 12, 453–457. [Google Scholar] [CrossRef] [Green Version]
- Li, B.; Severson, E.; Pignon, J.C.; Zhao, H.; Li, T.; Novak, J.; Jiang, P.; Shen, H.; Aster, J.C.; Rodig, S.; et al. Comprehensive analyses of tumor immunity: Implications for cancer immunotherapy. Genome Biol. 2016, 17, 174. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Chifman, J.; Pullikuth, A.; Chou, J.W.; Bedognetti, D.; Miller, L.D. Conservation of immune gene signatures in solid tumors and prognostic implications. BMC Cancer 2016, 16, 911. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Aran, D.; Hu, Z.; Butte, A.J. xCell: Digitally portraying the tissue cellular heterogeneity landscape. Genome Biol. 2017, 18, 220. [Google Scholar] [CrossRef] [Green Version]
- Becht, E.; Giraldo, N.A.; Lacroix, L.; Buttard, B.; Elarouci, N.; Petitprez, F.; Selves, J.; Laurent-Puig, P.; Sautès-Fridman, C.; Fridman, W.H.; et al. Estimating the population abundance of tissue-infiltrating immune and stromal cell populations using gene expression. Genome Biol. 2016, 17, 218. [Google Scholar] [CrossRef]
- Andersen, P.K.; Grill, R.D. Cox’s Regression Model for Counting Processes: A Large Sample Study. Ann. Stat. 1982, 10, 1100–1120. [Google Scholar] [CrossRef]
- Therneau, T.M.; Grambsch, P.M. Modeling Survival Data: Extending the Cox Model; Springer: New York, NY, USA, 2013. [Google Scholar]
- Gaujoux, R.; Seoighe, C. A flexible R package for nonnegative matrix factorization. BMC Bioinform. 2010, 11, 367. [Google Scholar] [CrossRef] [Green Version]
- Kuhn, M. Building Predictive Models in R Using the caret Package. J. Stat. Softw. 2008, 28, 1–26. [Google Scholar] [CrossRef] [Green Version]
- Breitung, K.; Ripley, B.D. Stochastic simulation. Stat. Pap. 1989, 30, 184. [Google Scholar] [CrossRef]
- Tibshirani, R. Regression shrinkage and selection via the lasso: A retrospective. J. R. Stat. Soc. Ser. B Stat. Methodol. 2011, 73, 273–282. [Google Scholar] [CrossRef]
- Friedman, J.; Hastie, T.; Tibshirani, R. Regularization Paths for Generalized Linear Models via Coordinate Descent. J. Stat. Softw. 2010, 33, 1–22. [Google Scholar] [CrossRef] [Green Version]
- Zhang, Z.; Kattan, M.W. Drawing Nomograms with R: Applications to categorical outcome and survival data. Ann. Transl. Med. 2017, 5, 211. [Google Scholar] [CrossRef] [Green Version]
- Cancer Genome Atlas Research Network. Integrated genomic analyses of ovarian carcinoma. Nature 2011, 474, 609–615. [Google Scholar] [CrossRef]
- He, C.; Zhang, Y.; Cai, Z.; Lin, X.; Li, S. Overall survival and cancer-specific survival in patients with surgically resected pancreatic head adenocarcinoma: A competing risk nomogram analysis. J. Cancer 2018, 9, 3156–3167. [Google Scholar] [CrossRef]
- Vickers, A.J.; van Calster, B.; Steyerberg, E.W. A simple, step-by-step guide to interpreting decision curve analysis. Diagn. Progn. Res. 2019, 3, 18. [Google Scholar] [CrossRef]
- Wang, R.; Ye, X.H.; Zhao, X.L.; Liu, J.L.; Zhang, C.Y. Development of a five-gene signature as a novel prognostic marker in ovarian cancer. Neoplasma 2019, 66, 343–349. [Google Scholar] [CrossRef] [PubMed]
- Yue, H.; Wang, J.; Chen, R.; Hou, X.; Li, J.; Lu, X. Gene signature characteristic of elevated stromal infiltration and activation is associated with increased risk of hematogenous and lymphatic metastasis in serous ovarian cancer. BMC Cancer 2019, 19, 1266. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Wang, L.; Wang, L.; Ma, L.; Liu, J.; Ma, S. Identifying Gene Signature for the Detection of Ovarian Cancer Based on the Achieved Related Genes. Gynecol. Obstet. Investig. 2017, 82, 361–370. [Google Scholar] [CrossRef] [PubMed]
- Sabatier, R.; Finetti, P.; Bonensea, J.; Jacquemier, J.; Adelaide, J.; Lambaudie, E.; Viens, P.; Birnbaum, D.; Bertucci, F. A seven-gene prognostic model for platinum-treated ovarian carcinomas. Br. J. Cancer 2011, 105, 304–311. [Google Scholar] [CrossRef]
- Cancer Genome Atlas Research Network. Integrated genomic and molecular characterization of cervical cancer. Nature 2017, 543, 378–384. [Google Scholar] [CrossRef]
- Pujade-Lauraine, E.; Fujiwara, K.; Ledermann, J.A.; Oza, A.M.; Kristeleit, R.S.; Ray-Coquard, I.L.; Richardson, G.E.; Sessa, C.; Yonemori, K.; Banerjee, S. Avelumab alone or in combination with pegylated liposomal doxorubicin versus pegylated liposomal doxorubicin alone in platinum-resistant or refractory epithelial ovarian cancer: Primary and biomarker analysis of the phase III JAVELIN Ovarian 200 trial. Gynecol. Oncol. 2019, 154 (Suppl. 1), 21–22. [Google Scholar] [CrossRef]
- Moore, K.N.; Bookman, M.; Sehouli, J.; Miller, A.; Anderson, C.; Scambia, G.; Myers, T.; Taskiran, C.; Robison, K.; Maeenpaeae, J.; et al. Atezolizumab, Bevacizumab, and Chemotherapy for Newly Diagnosed Stage III or IV Ovarian Cancer: Placebo-Controlled Randomized Phase III Trial (IMagyn050/GOG 3015/ENGOT-OV39). J. Clin. Oncol. 2021. online ahead of print. [Google Scholar] [CrossRef]
- Rosenberg, J.E.; Hoffman-Censits, J.; Powles, T.; van der Heijden, M.S.; Balar, A.V.; Necchi, A.; Dawson, N.; O’Donnell, P.H.; Balmanoukian, A.; Loriot, Y.; et al. Atezolizumab in patients with locally advanced and metastatic urothelial carcinoma who have progressed following treatment with platinum-based chemotherapy: A single-arm, multicentre, phase 2 trial. Lancet 2016, 387, 1909–1920. [Google Scholar] [CrossRef] [Green Version]
- Hanahan, D.; Coussens, L.M. Accessories to the crime: Functions of cells recruited to the tumor microenvironment. Cancer Cell 2012, 21, 309–322. [Google Scholar] [CrossRef] [Green Version]
- Schulz, M.; Salamero-Boix, A.; Niesel, K.; Alekseeva, T.; Sevenich, L. Microenvironmental Regulation of Tumor Progression and Therapeutic Response in Brain Metastasis. Front. Immunol. 2019, 10, 1713. [Google Scholar] [CrossRef]
- Hanahan, D.; Weinberg, R.A. Hallmarks of cancer: The next generation. Cell 2011, 144, 646–674. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Tran Janco, J.M.; Lamichhane, P.; Karyampudi, L.; Knutson, K.L. Tumor-infiltrating dendritic cells in cancer pathogenesis. J. Immunol. 2015, 194, 2985–2991. [Google Scholar] [CrossRef] [Green Version]
- Kaaks, R.; Fortner, R.T.; Husing, A.; Barrdahl, M.; Hopper, M.; Johnson, T.; Tjønneland, A.; Hansen, L.; Overvad, K.; Fournier, A.; et al. Tumor-associated autoantibodies as early detection markers for ovarian cancer? A prospective evaluation. Int. J. Cancer 2018, 143, 515–526. [Google Scholar] [CrossRef]
- Cheng, H.W.; Onder, L.; Cupovic, J.; Boesch, M.; Novkovic, M.; Pikor, N.; Tarantino, I.; Rodriguez, R.; Schneider, T.; Jochum, W.; et al. CCL19-producing fibroblastic stromal cells restrain lung carcinoma growth by promoting local antitumor T-cell responses. J. Allergy Clin. Immunol. 2018, 142, 1257–1271.e4. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Namkoong, H.; Song, M.Y.; Seo, Y.B.; Choi, D.H.; Kim, S.W.; Im, S.J.; Sung, Y.C.; Park, Y. Enhancement of antigen-specific CD8 T cell responses by co-delivery of Fc-fused CXCL11. Vaccine 2014, 32, 1205–1212. [Google Scholar] [CrossRef] [PubMed]
- Kurman, R.J.; Visvanathan, K.; Roden, R.; Wu, T.C.; Shih, L. Early detection and treatment of ovarian cancer: Shifting from early stage to minimal volume of disease based on a new model of carcinogenesis. Am. J. Obstet. Gynecol. 2008, 198, 351–356. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- De Graeff, P.; Crijns, A.P.; Ten Hoor, K.A.; Klip, H.G.; Hollema, H.; Oien, K.; Bartlett, J.M.; Wisman, G.B.; de Bock, G.H.; de Vries, E.G.; et al. The ErbB signalling pathway: Protein expression and prognostic value in epithelial ovarian cancer. Br. J. Cancer 2008, 99, 341–349. [Google Scholar] [CrossRef] [Green Version]
- Shahin, M.S.; Hughes, J.H.; Sood, A.K.; Buller, R.E. The prognostic significance of p53 tumor suppressor gene alterations in ovarian carcinoma. Cancer 2000, 89, 2006–2017. [Google Scholar] [CrossRef]
- Ahmed, A.A.; Etemadmoghadam, D.; Temple, J.; Lynch, A.G.; Riad, M.; Sharma, R.; Stewart, C.; Fereday, S.; Caldas, C.; Defazio, A.; et al. Driver mutations in TP53 are ubiquitous in high grade serous carcinoma of the ovary. J. Pathol. 2010, 221, 49–56. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Koebel, M.; Reuss, A.; du Bois, A.; Kommoss, S.; Kommoss, F.; Gao, D.; Kalloger, S.E.; Huntsman, D.G.; Gilks, C.B. The biological and clinical value of p53 expression in pelvic high-grade serous carcinomas. J. Pathol. 2010, 222, 191–198. [Google Scholar] [CrossRef]
- Previs, R.A.; Bevis, K.S.; Huh, W.; Tillmanns, T.; Perry, L.; Moore, K.; Chapman, J.; McClung, C.; Kiet, T.; Java, J.; et al. A prognostic nomogram to predict overall survival in women with recurrent ovarian cancer treated with bevacizumab and chemotherapy. Gynecol. Oncol. 2014, 132, 531–536. [Google Scholar] [CrossRef]
- Lee, C.K.; Simes, R.J.; Brown, C.; Gebski, V.; Pfisterer, J.; Swart, A.M.; Berton-Rigaud, D.; Plante, M.; Skeie-Jensen, T.; Vergote, I.; et al. A prognostic nomogram to predict overall survival in patients with platinum-sensitive recurrent ovarian cancer. Ann. Oncol. 2013, 24, 937–943. [Google Scholar] [CrossRef]
- Rose, P.G.; Java, J.J.; Salani, R.; Geller, M.A.; Secord, A.A.; Tewari, K.S.; Bender, D.P.; Mutch, D.G.; Friedlander, M.L.; Van Le, L.; et al. Nomogram for Predicting Individual Survival After Recurrence of Advanced-Stage, High-Grade Ovarian Carcinoma. Obstet. Gynecol. 2019, 133, 245–254. [Google Scholar] [CrossRef]
- van de Laar, R.; IntHout, J.; Van Gorp, T.; Verdonschot, S.; van Altena, A.M.; Gerestein, C.G.; Massuger, L.F.; Zusterzeel, P.L.; Kruitwagen, R.F. External validation of three prognostic models for overall survival in patients with advanced-stage epithelial ovarian cancer. Br. J. Cancer 2014, 110, 42–48. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Lee, C.K.; Simes, R.J.; Brown, C.; Lord, S.; Wagner, U.; Plante, M.; Vergote, I.; Pisano, C.; Parma, G.; Burges, A.; et al. Prognostic nomogram to predict progression-free survival in patients with platinum-sensitive recurrent ovarian cancer. Br. J. Cancer 2011, 105, 1144–1150. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Patel, S.A.; Minn, A.J. Combination Cancer Therapy with Immune Checkpoint Blockade: Mechanisms and Strategies. Immunity 2018, 48, 417–433. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Alexandrov, L.B.; Nik-Zainal, S.; Wedge, D.C.; Aparicio, S.A.; Behjati, S.; Biankin, A.V.; Bignell, G.R.; Bolli, N.; Borg, A.; Børresen-Dale, A.L.; et al. Signatures of mutational processes in human cancer. Nature 2013, 500, 415–421. [Google Scholar] [CrossRef] [Green Version]
- Sharma, P.; Hu-Lieskovan, S.; Wargo, J.A.; Ribas, A. Primary, Adaptive, and Acquired Resistance to Cancer Immunotherapy. Cell 2017, 168, 707–723. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Ribas, A. Adaptive Immune Resistance: How Cancer Protects from Immune Attack. Cancer Discov. 2015, 5, 915–919. [Google Scholar] [CrossRef] [Green Version]
- Koyama, S.; Akbay, E.A.; Li, Y.Y.; Herter-Sprie, G.S.; Buczkowski, K.A.; Richards, W.G.; Gandhi, L.; Redig, A.J.; Rodig, S.J.; Asahina, H.; et al. Adaptive resistance to therapeutic PD-1 blockade is associated with upregulation of alternative immune checkpoints. Nat. Commun. 2016, 7, 10501. [Google Scholar] [CrossRef] [PubMed]
- Hegde, P.S.; Karanikas, V.; Evers, S. The Where, the When, and the How of Immune Monitoring for Cancer Immunotherapies in the Era of Checkpoint Inhibition. Clin. Cancer Res. 2016, 22, 1865–1874. [Google Scholar] [CrossRef] [Green Version]
- Vento, J.; Mulgaonkar, A.; Woolford, L.; Nham, K.; Christie, A.; Bagrodia, A.; de Leon, A.D.; Hannan, R.; Bowman, I.; McKay, R.M.; et al. PD-L1 detection using 89Zr-atezolizumab immuno-PET in renal cell carcinoma tumorgrafts from a patient with favorable nivolumab response. J. Immunother. Cancer 2019, 7, 144. [Google Scholar] [CrossRef] [PubMed]
- Zheng, M.; Mullikin, H.; Hester, A.; Czogalla, B.; Heidegger, H.; Vilsmaier, T.; Vattai, A.; Chelariu-Raicu, A.; Jeschke, U.; Trillsch, F.; et al. Prognostic Model for Serous Ovarian Carcinomas Based on Lipid Metabolism Expression Profile. Int. J. Mol. Sci. 2020, 21, 9169. [Google Scholar] [CrossRef]






| Clinical Feature | TCGA | ICGC-Australian | |
|---|---|---|---|
| Event | Censored | 125 | 19 |
| Dead | 222 | 74 | |
| FIGO stage | III | 290 | 79 |
| IV | 57 | 14 | |
| Grade | G1 | 1 | |
| G2 | 35 | ||
| G3 | 302 | ||
| G4 | 1 | ||
| None | 8 | ||
| Chemotherapy | Yes | 322 | |
| No | 25 | ||
| Recurrence | Yes | 186 | |
| No | 161 | ||
| Age | ≤60 | 192 | |
| >60 | 155 | ||
| Residual disease | No | 55 | |
| 1–10 mm | 25 | ||
| 11–20 mm | 164 | ||
| >20 mm | 69 | ||
| Unknown | 34 | ||
| Clinical Feature | TCGA Training Cohort | TCGA Testing Cohort | p-Value | |
|---|---|---|---|---|
| Event | Censored | 92 | 33 | 0.3333 |
| Dead | 151 | 71 | ||
| FIGO stage | III | 201 | 89 | 0.6165 |
| IV | 42 | 15 | ||
| Grade | G1 | 1 | 0 | 0.5779 |
| G2 | 22 | 13 | ||
| G3 | 212 | 90 | ||
| G4 | 1 | 0 | ||
| None | 7 | 1 | ||
| Chemotherapy | Yes | 227 | 95 | 0.6481 |
| No | 16 | 9 | ||
| Recurrence | Yes | 128 | 58 | 0.6803 |
| No | 115 | 46 | ||
| Age | ≤60 | 140 | 52 | 0.1913 |
| >60 | 103 | 52 | ||
| Residual disease | No | 41 | 14 | 0.8823 |
| 1–10 mm | 114 | 50 | ||
| 11–20 mm | 17 | 8 | ||
| >20 mm | 46 | 23 | ||
| Unknown | 25 | 9 | ||
| Variables | Univariable Analysis | Multivariable Analysis | ||||
|---|---|---|---|---|---|---|
| HR | 95% CI | p-Value | HR | 95% CI | p-Value | |
| FIGO stage | 1.16 | 0.82–1.66 | 0.408 | 1.24 | 0.86–1.78 | 0.251 |
| Grade | 1.03 | 0.67–1.58 | 0.884 | 1.12 | 0.73–1.71 | 0.620 |
| Chemotherapy | 0.29 | 0.18–0.48 | <0.001 | 0.35 | 0.20–0.59 | <0.001 |
| Recurrence | 1.15 | 0.86–1.54 | 0.353 | 1.21 | 0.89–1.65 | 0.227 |
| Age | 0.79 | 0.60–1.04 | 0.097 | 0.79 | 0.59–1.05 | 0.101 |
| Residual disease | 2.06 | 1.29–3.26 | 0.001 | 1.70 | 1.06–2.73 | 0.027 |
| RS | 2.19 | 1.53–3.15 | <0.001 | 1.73 | 1.17–2.55 | <0.006 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Zheng, M.; Long, J.; Chelariu-Raicu, A.; Mullikin, H.; Vilsmaier, T.; Vattai, A.; Heidegger, H.H.; Batz, F.; Keckstein, S.; Jeschke, U.; et al. Identification of a Novel Tumor Microenvironment Prognostic Signature for Advanced-Stage Serous Ovarian Cancer. Cancers 2021, 13, 3343. https://doi.org/10.3390/cancers13133343
Zheng M, Long J, Chelariu-Raicu A, Mullikin H, Vilsmaier T, Vattai A, Heidegger HH, Batz F, Keckstein S, Jeschke U, et al. Identification of a Novel Tumor Microenvironment Prognostic Signature for Advanced-Stage Serous Ovarian Cancer. Cancers. 2021; 13(13):3343. https://doi.org/10.3390/cancers13133343
Chicago/Turabian StyleZheng, Mingjun, Junyu Long, Anca Chelariu-Raicu, Heather Mullikin, Theresa Vilsmaier, Aurelia Vattai, Helene Hildegard Heidegger, Falk Batz, Simon Keckstein, Udo Jeschke, and et al. 2021. "Identification of a Novel Tumor Microenvironment Prognostic Signature for Advanced-Stage Serous Ovarian Cancer" Cancers 13, no. 13: 3343. https://doi.org/10.3390/cancers13133343
APA StyleZheng, M., Long, J., Chelariu-Raicu, A., Mullikin, H., Vilsmaier, T., Vattai, A., Heidegger, H. H., Batz, F., Keckstein, S., Jeschke, U., Trillsch, F., Mahner, S., & Kaltofen, T. (2021). Identification of a Novel Tumor Microenvironment Prognostic Signature for Advanced-Stage Serous Ovarian Cancer. Cancers, 13(13), 3343. https://doi.org/10.3390/cancers13133343








