Differential Regulation of Lacto-/Neolacto- Glycosphingolipid Biosynthesis Pathway Reveals Transcription Factors as Potential Candidates in Triple-Negative Breast Cancer
Abstract
Simple Summary
Abstract
1. Introduction
2. Materials and Methods
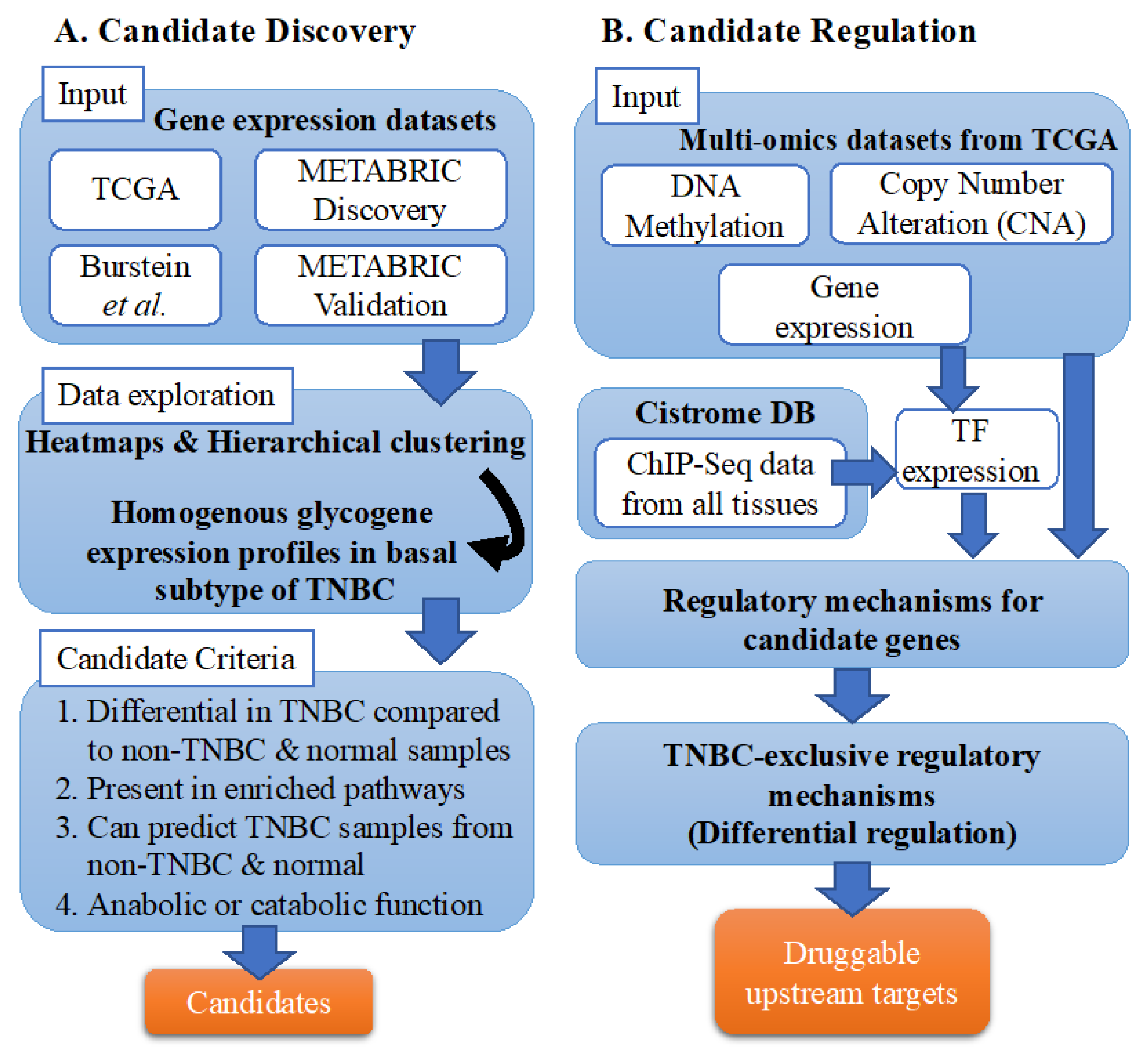
2.1. Study Design
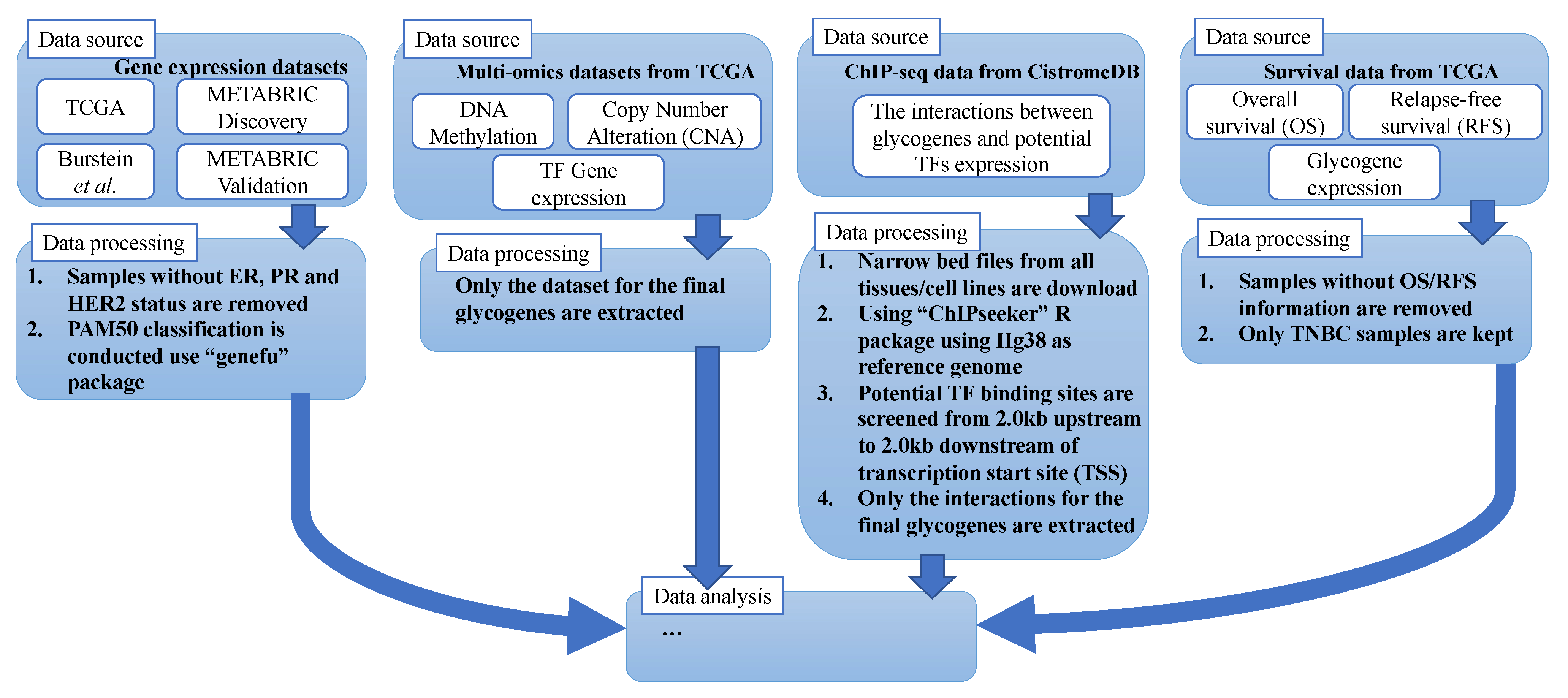
2.2. Data Sources and Processing
2.3. Data Exploration and Candidate Identification
2.4. Regulatory Mechanism Analysis
3. Results
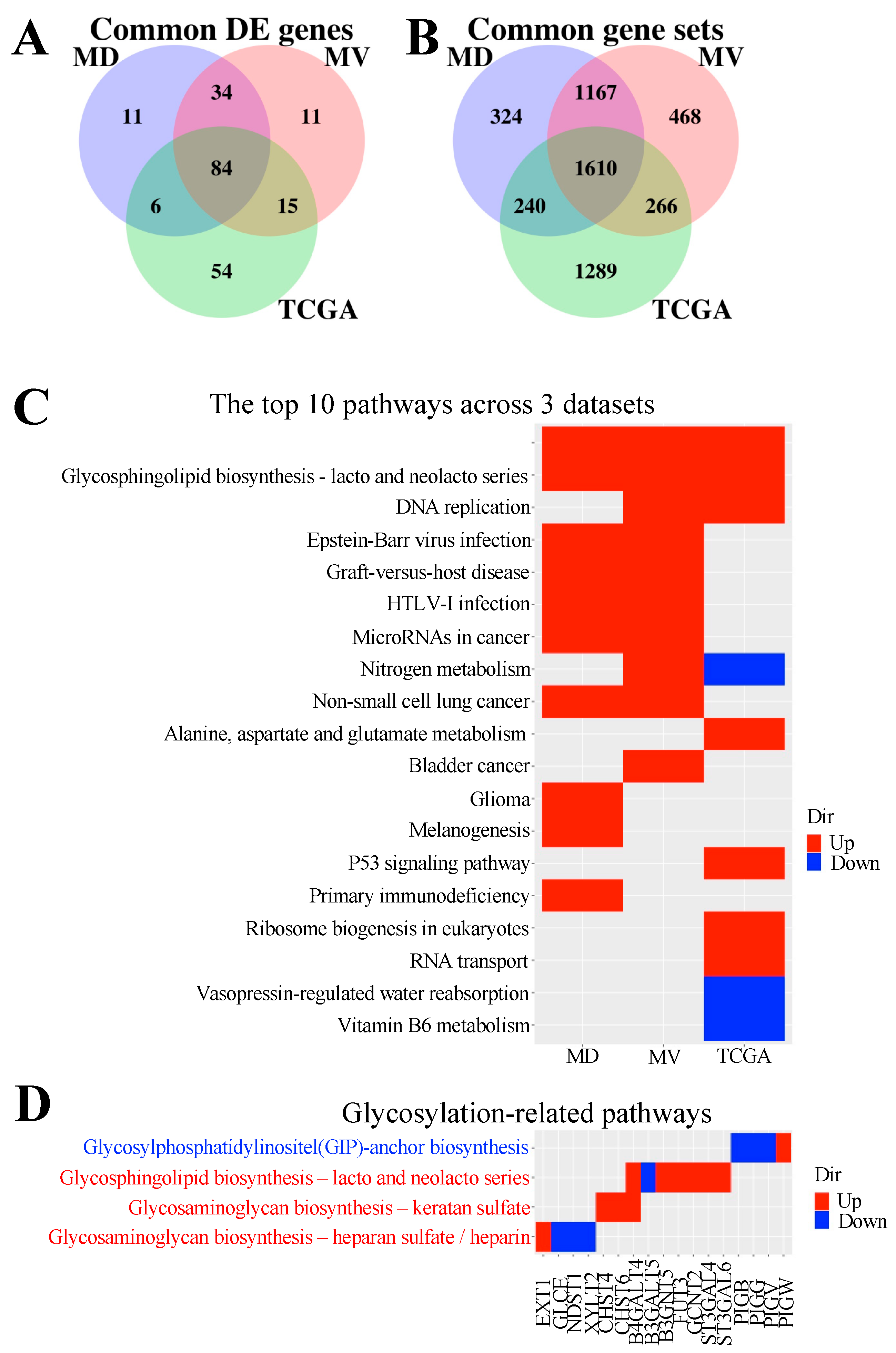
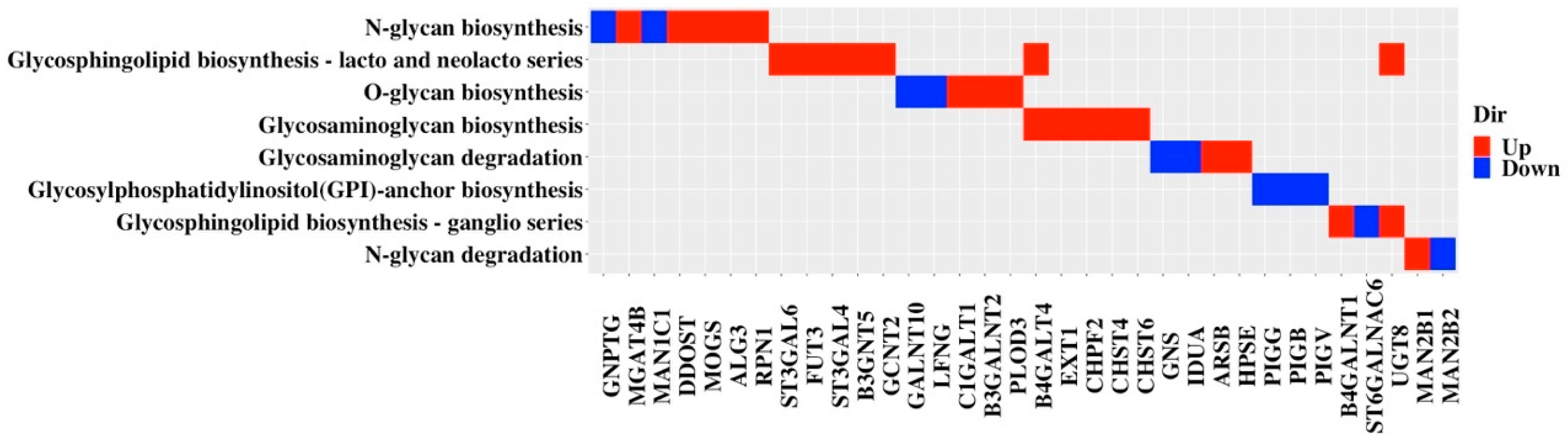
3.1. Candidate Discovery
3.2. Association of Candidate Glycogenes with Survival in TNBC
3.3. Candidate Glycogene Regulatory Mechanisms
3.4. Discovery of TFs That Regulate Candidate Glycogenes in TNBC
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- World Health Organization. International Agency for Research on Cancer, WHO Cancer Mortality Database. Available online: http://gco.iarc.fr/ (accessed on 1 April 2021).
- Prat, A.; Adamo, B.; Cheang, M.C.; Anders, C.K.; Carey, L.A.; Perou, C.M. Molecular characterization of basal-like and non-basal-like triple-negative breast cancer. Oncologist 2013, 18, 123–133. [Google Scholar] [CrossRef]
- Lehmann, B.D.; Bauer, J.A.; Chen, X.; Sanders, M.E.; Chakravarthy, A.B.; Shyr, Y.; Pietenpol, J.A. Identification of human triple-negative breast cancer subtypes and preclinical models for selection of targeted therapies. J. Clin. Investig. 2011, 121, 2750–2767. [Google Scholar] [CrossRef]
- Yin, L.; Duan, J.J.; Bian, X.W.; Yu, S.C. Triple-negative breast cancer molecular subtyping and treatment progress. Breast Cancer Res. 2020, 22, 61. [Google Scholar] [CrossRef]
- Dent, R.; Trudeau, M.; Pritchard, K.I.; Hanna, W.M.; Kahn, H.K.; Sawka, C.A.; Lickley, L.A.; Rawlinson, E.; Sun, P.; Narod, S.A. Triple-negative breast cancer: Clinical features and patterns of recurrence. Clin. Cancer Res. 2007, 13, 4429–4434. [Google Scholar] [CrossRef]
- Carey, L.A.; Dees, E.C.; Sawyer, L.; Gatti, L.; Moore, D.T.; Collichio, F.; Ollila, D.W.; Sartor, C.I.; Graham, M.L.; Perou, C.M. The triple negative paradox: Primary tumor chemosensitivity of breast cancer subtypes. Clin. Cancer Res. 2007, 13, 2329–2334. [Google Scholar] [CrossRef] [PubMed]
- Coughlin, S.S.; Ekwueme, D.U. Breast cancer as a global health concern. Cancer Epidemiol. 2009, 33, 315–318. [Google Scholar] [CrossRef]
- Pinho, S.S.; Reis, C.A. Glycosylation in cancer: Mechanisms and clinical implications. Nat. Rev. Cancer 2015, 15, 540–555. [Google Scholar] [CrossRef] [PubMed]
- Gendoo, D.M.; Ratanasirigulchai, N.; Schroder, M.S.; Pare, L.; Parker, J.S.; Prat, A.; Haibe-Kains, B. Genefu: An R/Bioconductor package for computation of gene expression-based signatures in breast cancer. Bioinformatics 2016, 32, 1097–1099. [Google Scholar] [CrossRef] [PubMed]
- Goldman, M.J.; Craft, B.; Hastie, M.; Repecka, K.; McDade, F.; Kamath, A.; Banerjee, A.; Luo, Y.; Rogers, D.; Brooks, A.N.; et al. Visualizing and interpreting cancer genomics data via the Xena platform. Nat. Biotechnol. 2020, 38, 675–678. [Google Scholar] [CrossRef]
- Huang, W.Y.; Hsu, S.D.; Huang, H.Y.; Sun, Y.M.; Chou, C.H.; Weng, S.L.; Huang, H.D. MethHC: A database of DNA methylation and gene expression in human cancer. Nucleic Acids Res. 2015, 43, D856–D861. [Google Scholar] [CrossRef]
- Prat, A.; Perou, C.M. Deconstructing the molecular portraits of breast cancer. Mol. Oncol. 2011, 5, 5–23. [Google Scholar] [CrossRef]
- Dawood, S.; Broglio, K.; Esteva, F.J.; Yang, W.; Kau, S.W.; Islam, R.; Albarracin, C.; Yu, T.K.; Green, M.; Hortobagyi, G.N.; et al. Survival among women with triple receptor-negative breast cancer and brain metastases. Ann. Oncol. 2009, 20, 621–627. [Google Scholar] [CrossRef] [PubMed]
- Weigelt, B.; Baehner, F.L.; Reis-Filho, J.S. The contribution of gene expression profiling to breast cancer classification, prognostication and prediction: A retrospective of the last decade. J. Pathol. 2010, 220, 263–280. [Google Scholar] [CrossRef] [PubMed]
- Yehia, L.; Boulos, F.; Jabbour, M.; Mahfoud, Z.; Fakhruddin, N.; El-Sabban, M. Expression of HIF-1alpha and Markers of Angiogenesis Are Not Significantly Different in Triple Negative Breast Cancer Compared to Other Breast Cancer Molecular Subtypes: Implications for Future Therapy. PLoS ONE 2015, 10, e0129356. [Google Scholar] [CrossRef] [PubMed]
- Vagia, E.; Mahalingam, D.; Cristofanilli, M. The Landscape of Targeted Therapies in TNBC. Cancers 2020, 12, 916. [Google Scholar] [CrossRef]
- Cortes, J.; Cescon, D.W.; Rugo, H.S.; Nowecki, Z.; Im, S.A.; Yusof, M.M.; Gallardo, C.; Lipatov, O.; Barrios, C.H.; Holgado, E.; et al. Pembrolizumab plus chemotherapy versus placebo plus chemotherapy for previously untreated locally recurrent inoperable or metastatic triple-negative breast cancer (KEYNOTE-355): A randomised, placebo-controlled, double-blind, phase 3 clinical trial. Lancet 2020, 396, 1817–1828. [Google Scholar] [CrossRef]
- Schmid, P.; Adams, S.; Rugo, H.S.; Schneeweiss, A.; Barrios, C.H.; Iwata, H.; Dieras, V.; Hegg, R.; Im, S.A.; Shaw Wright, G.; et al. Atezolizumab and Nab-Paclitaxel in Advanced Triple-Negative Breast Cancer. N. Engl. J. Med. 2018, 379, 2108–2121. [Google Scholar] [CrossRef]
- Fuster, M.M.; Esko, J.D. The sweet and sour of cancer: Glycans as novel therapeutic targets. Nat. Rev. Cancer 2005, 5, 526–542. [Google Scholar] [CrossRef] [PubMed]
- Mereiter, S.; Balmana, M.; Campos, D.; Gomes, J.; Reis, C.A. Glycosylation in the Era of Cancer-Targeted Therapy: Where Are We Heading? Cancer Cell 2019, 36, 6–16. [Google Scholar] [CrossRef]
- Lee, H.H.; Wang, Y.N.; Xia, W.; Chen, C.H.; Rau, K.M.; Ye, L.; Wei, Y.; Chou, C.K.; Wang, S.C.; Yan, M.; et al. Removal of N-Linked Glycosylation Enhances PD-L1 Detection and Predicts Anti-PD-1/PD-L1 Therapeutic Efficacy. Cancer Cell 2019, 36, 168–178 e164. [Google Scholar] [CrossRef]
- Li, C.W.; Lim, S.O.; Chung, E.M.; Kim, Y.S.; Park, A.H.; Yao, J.; Cha, J.H.; Xia, W.; Chan, L.C.; Kim, T.; et al. Eradication of Triple-Negative Breast Cancer Cells by Targeting Glycosylated PD-L1. Cancer Cell 2018, 33, 187–201.e10. [Google Scholar] [CrossRef]
- Scott, D.A.; Drake, R.R. Glycosylation and its implications in breast cancer. Expert Rev. Proteom. 2019, 16, 665–680. [Google Scholar] [CrossRef]
- Cancer Genome Atlas, N. Comprehensive molecular portraits of human breast tumours. Nature 2012, 490, 61–70. [Google Scholar] [CrossRef]
- Karolchik, D.; Baertsch, R.; Diekhans, M.; Furey, T.S.; Hinrichs, A.; Lu, Y.T.; Roskin, K.M.; Schwartz, M.; Sugnet, C.W.; Thomas, D.J.; et al. The UCSC Genome Browser Database. Nucleic Acids Res. 2003, 31, 51–54. [Google Scholar] [CrossRef] [PubMed]
- Curtis, C.; Shah, S.P.; Chin, S.F.; Turashvili, G.; Rueda, O.M.; Dunning, M.J.; Speed, D.; Lynch, A.G.; Samarajiwa, S.; Yuan, Y.; et al. The genomic and transcriptomic architecture of 2,000 breast tumours reveals novel subgroups. Nature 2012, 486, 346–352. [Google Scholar] [CrossRef] [PubMed]
- Burstein, M.D.; Tsimelzon, A.; Poage, G.M.; Covington, K.R.; Contreras, A.; Fuqua, S.A.; Savage, M.I.; Osborne, C.K.; Hilsenbeck, S.G.; Chang, J.C.; et al. Comprehensive genomic analysis identifies novel subtypes and targets of triple-negative breast cancer. Clin. Cancer Res. 2015, 21, 1688–1698. [Google Scholar] [CrossRef] [PubMed]
- Mei, S.; Qin, Q.; Wu, Q.; Sun, H.; Zheng, R.; Zang, C.; Zhu, M.; Wu, J.; Shi, X.; Taing, L.; et al. Cistrome Data Browser: A data portal for ChIP-Seq and chromatin accessibility data in human and mouse. Nucleic Acids Res. 2017, 45, D658–D662. [Google Scholar] [CrossRef]
- Alhamdoosh, M.; Law, C.W.; Tian, L.; Sheridan, J.M.; Ng, M.; Ritchie, M.E. Easy and efficient ensemble gene set testing with EGSEA. F1000Research 2017, 6, 2010. [Google Scholar] [CrossRef]
- Brody, B.L.; Gamst, A.C.; Williams, R.A.; Smith, A.R.; Lau, P.W.; Dolnak, D.; Rapaport, M.H.; Kaplan, R.M.; Brown, S.I. Depression, visual acuity, comorbidity, and disability associated with age-related macular degeneration. Ophthalmology 2001, 108, 1893–1900. [Google Scholar] [CrossRef]
- Ashkani, J.; Naidoo, K.J. Glycosyltransferase Gene Expression Profiles Classify Cancer Types and Propose Prognostic Subtypes. Sci. Rep. 2016, 6, 26451. [Google Scholar] [CrossRef]
- Potapenko, I.O.; Luders, T.; Russnes, H.G.; Helland, A.; Sorlie, T.; Kristensen, V.N.; Nord, S.; Lingjaerde, O.C.; Borresen-Dale, A.L.; Haakensen, V.D. Glycan-related gene expression signatures in breast cancer subtypes; relation to survival. Mol. Oncol. 2015, 9, 861–876. [Google Scholar] [CrossRef] [PubMed]
- Kanehisa, M.; Goto, S. KEGG: Kyoto encyclopedia of genes and genomes. Nucleic Acids Res. 2000, 28, 27–30. [Google Scholar] [CrossRef]
- GlycoGene DataBase. Available online: http://acgg.asia/ggdb2/ (accessed on 1 March 2017).
- Li, X.; Yu, X.; Zhou, D.; Chen, B.; Li, W.; Zheng, X.; Zeng, H.; Long, L.; Zhou, W. CCT020312 Inhibits Triple-Negative Breast Cancer through PERK Pathway-Mediated G1 Phase Cell Cycle Arrest and Apoptosis. Front. Pharmacol. 2020, 11, 737. [Google Scholar] [CrossRef] [PubMed]
- Liu, C.; Rohart, F.; Simpson, P.T.; Khanna, K.K.; Ragan, M.A.; Le Cao, K.A. Integrating Multi-omics Data to Dissect Mechanisms of DNA repair Dysregulation in Breast Cancer. Sci. Rep. 2016, 6, 34000. [Google Scholar] [CrossRef]
- Cipriano, E.; Mesquita, A. Emerging Therapeutic Drugs in Metastatic Triple-Negative Breast Cancer. Breast Cancer 2021, 15. [Google Scholar] [CrossRef]
- Traina, T.A.; Miller, K.; Yardley, D.A.; Eakle, J.; Schwartzberg, L.S.; O’Shaughnessy, J.; Gradishar, W.; Schmid, P.; Winer, E.; Kelly, C.; et al. Enzalutamide for the Treatment of Androgen Receptor-Expressing Triple-Negative Breast Cancer. J. Clin. Oncol. 2018, 36, 884–890. [Google Scholar] [CrossRef]
- Chuang, P.K.; Hsiao, M.; Hsu, T.L.; Chang, C.F.; Wu, C.Y.; Chen, B.R.; Huang, H.W.; Liao, K.S.; Chen, C.C.; Chen, C.L.; et al. Signaling pathway of globo-series glycosphingolipids and beta1,3-galactosyltransferase V (beta3GalT5) in breast cancer. Proc. Natl. Acad. Sci. USA 2019, 116, 3518–3523. [Google Scholar] [CrossRef]
- Regina Todeschini, A.; Hakomori, S.I. Functional role of glycosphingolipids and gangliosides in control of cell adhesion, motility, and growth, through glycosynaptic microdomains. Biochim. Biophys. Acta 2008, 1780, 421–433. [Google Scholar] [CrossRef]
- Lingwood, C.A. Glycosphingolipid functions. Cold Spring Harb. Perspect. Biol. 2011, 3. [Google Scholar] [CrossRef]
- Liang, Y.J.; Kuo, H.H.; Lin, C.H.; Chen, Y.Y.; Yang, B.C.; Cheng, Y.Y.; Yu, A.L.; Khoo, K.H.; Yu, J. Switching of the core structures of glycosphingolipids from globo- and lacto- to ganglio-series upon human embryonic stem cell differentiation. Proc. Natl. Acad. Sci. USA 2010, 107, 22564–22569. [Google Scholar] [CrossRef]
- Hakomori, S.I. Glycosynaptic microdomains controlling tumor cell phenotype through alteration of cell growth, adhesion, and motility. FEBS Lett. 2010, 584, 1901–1906. [Google Scholar] [CrossRef] [PubMed]
- Jacob, F.; Anugraham, M.; Pochechueva, T.; Tse, B.W.; Alam, S.; Guertler, R.; Bovin, N.V.; Fedier, A.; Hacker, N.F.; Huflejt, M.E.; et al. The glycosphingolipid P(1) is an ovarian cancer-associated carbohydrate antigen involved in migration. Br. J. Cancer 2014, 111, 1634–1645. [Google Scholar] [CrossRef] [PubMed]
- Wang, Z.; Wen, L.; Ma, X.; Chen, Z.; Yu, Y.; Zhu, J.; Wang, Y.; Liu, Z.; Liu, H.; Wu, D.; et al. High expression of lactotriaosylceramide, a differentiation-associated glycosphingolipid, in the bone marrow of acute myeloid leukemia patients. Glycobiology 2012, 22, 930–938. [Google Scholar] [CrossRef] [PubMed]
- Etcheverry, A.; Aubry, M.; de Tayrac, M.; Vauleon, E.; Boniface, R.; Guenot, F.; Saikali, S.; Hamlat, A.; Riffaud, L.; Menei, P.; et al. DNA methylation in glioblastoma: Impact on gene expression and clinical outcome. BMC Genom. 2010, 11, 701. [Google Scholar] [CrossRef] [PubMed]
- Horakova, D.; Bouchalova, K.; Cwiertka, K.; Stepanek, L.; Vlckova, J.; Kollarova, H. Risks and Protective Factors for Triple Negative Breast Cancer with a Focus on Micronutrients and Infections. Biomed. Pap. Med. Fac. Univ. Palacky. Olomouc. Czech. Repub. 2018, 162, 83–89. [Google Scholar] [CrossRef] [PubMed]
- Kaur, S.; Elkahloun, A.G.; Singh, S.P.; Chen, Q.R.; Meerzaman, D.M.; Song, T.; Manu, N.; Wu, W.; Mannan, P.; Garfield, S.H.; et al. A function-blocking CD47 antibody suppresses stem cell and EGF signaling in triple-negative breast cancer. Oncotarget 2016, 7, 10133–10152. [Google Scholar] [CrossRef] [PubMed]
- Albuquerque, A.P.B.; Balmana, M.; Mereiter, S.; Pinto, F.; Reis, C.A.; Beltrao, E.I.C. Hypoxia and serum deprivation induces glycan alterations in triple negative breast cancer cells. Biol. Chem. 2018, 399, 661–672. [Google Scholar] [CrossRef]
- Cao, Q.; Chen, X.; Wu, X.; Liao, R.; Huang, P.; Tan, Y.; Wang, L.; Ren, G.; Huang, J.; Dong, C. Inhibition of UGT8 suppresses basal-like breast cancer progression by attenuating sulfatide-alphaVbeta5 axis. J. Exp. Med. 2018, 215, 1679–1692. [Google Scholar] [CrossRef]
- Rampurwala, M.; Wisinski, K.B.; O’Regan, R. Role of the androgen receptor in triple-negative breast cancer. Clin. Adv. Hematol. Oncol. 2016, 14, 186–193. [Google Scholar] [PubMed]
- Dai, X.; Li, T.; Bai, Z.; Yang, Y.; Liu, X.; Zhan, J.; Shi, B. Breast cancer intrinsic subtype classification, clinical use and future trends. Am. J. Cancer Res. 2015, 5, 2929–2943. [Google Scholar]
- Thike, A.A.; Yong-Zheng Chong, L.; Cheok, P.Y.; Li, H.H.; Wai-Cheong Yip, G.; Huat Bay, B.; Tse, G.M.; Iqbal, J.; Tan, P.H. Loss of androgen receptor expression predicts early recurrence in triple-negative and basal-like breast cancer. Mod. Pathol. 2014, 27, 352–360. [Google Scholar] [CrossRef]
- Astvatsaturyan, K.; Yue, Y.; Walts, A.E.; Bose, S. Androgen receptor positive triple negative breast cancer: Clinicopathologic, prognostic, and predictive features. PLoS ONE 2018, 13, e0197827. [Google Scholar] [CrossRef]
- Asano, Y.; Kashiwagi, S.; Onoda, N.; Kurata, K.; Morisaki, T.; Noda, S.; Takashima, T.; Ohsawa, M.; Kitagawa, S.; Hirakawa, K. Clinical verification of sensitivity to preoperative chemotherapy in cases of androgen receptor-expressing positive breast cancer. Br. J. Cancer 2016, 114, 14–20. [Google Scholar] [CrossRef]
- Aguilera, T.A.; Rafat, M.; Castellini, L.; Shehade, H.; Kariolis, M.S.; Hui, A.B.; Stehr, H.; von Eyben, R.; Jiang, D.; Ellies, L.G.; et al. Reprogramming the immunological microenvironment through radiation and targeting Axl. Nat. Commun. 2016, 7, 13898. [Google Scholar] [CrossRef]
- Maeda, T.; Nakanishi, Y.; Hirotani, Y.; Fuchinoue, F.; Enomoto, K.; Sakurai, K.; Amano, S.; Nemoto, N. Immunohistochemical co-expression status of cytokeratin 5/6, androgen receptor, and p53 as prognostic factors of adjuvant chemotherapy for triple negative breast cancer. Med. Mol. Morphol. 2016, 49, 11–21. [Google Scholar] [CrossRef]
- Gasparini, P.; Fassan, M.; Cascione, L.; Guler, G.; Balci, S.; Irkkan, C.; Paisie, C.; Lovat, F.; Morrison, C.; Zhang, J.; et al. Androgen receptor status is a prognostic marker in non-basal triple negative breast cancers and determines novel therapeutic options. PLoS ONE 2014, 9, e88525. [Google Scholar] [CrossRef]
- Bhattarai, S.; Klimov, S.; Mittal, K.; Krishnamurti, U.; Li, X.B.; Oprea-Ilies, G.; Wetherilt, C.S.; Riaz, A.; Aleskandarany, M.A.; Green, A.R.; et al. Prognostic Role of Androgen Receptor in Triple Negative Breast Cancer: A Multi-Institutional Study. Cancers 2019, 11, 995. [Google Scholar] [CrossRef]
- Qu, Q.; Mao, Y.; Fei, X.C.; Shen, K.W. The impact of androgen receptor expression on breast cancer survival: A retrospective study and meta-analysis. PLoS ONE 2013, 8, e82650. [Google Scholar] [CrossRef] [PubMed]
- Gucalp, A.; Traina, T.A. Triple-negative breast cancer: Role of the androgen receptor. Cancer J. 2010, 16, 62–65. [Google Scholar] [CrossRef] [PubMed]
- Santonja, A.; Sanchez-Munoz, A.; Lluch, A.; Chica-Parrado, M.R.; Albanell, J.; Chacon, J.I.; Antolin, S.; Jerez, J.M.; de la Haba, J.; de Luque, V.; et al. Triple negative breast cancer subtypes and pathologic complete response rate to neoadjuvant chemotherapy. Oncotarget 2018, 9, 26406–26416. [Google Scholar] [CrossRef] [PubMed]
- Querzoli, P.; Pedriali, M.; Rinaldi, R.; Secchiero, P.; Rossi, P.G.; Kuhn, E. GATA3 as an Adjunct Prognostic Factor in Breast Cancer Patients with Less Aggressive Disease: A Study with a Review of the Literature. Diagnostics 2021, 11, 604. [Google Scholar] [CrossRef]
- Cimino-Mathews, A. Novel uses of immunohistochemistry in breast pathology: Interpretation and pitfalls. Mod. Pathol. 2021, 34, 62–77. [Google Scholar] [CrossRef]
- Aphivatanasiri, C.; Li, J.; Chan, R.; Jamidi, S.K.; Tsang, J.Y.; Poon, I.K.; Shao, Y.; Tong, J.; To, K.F.; Chan, S.K.; et al. Combined SOX10 GATA3 is most sensitive in detecting primary and metastatic breast cancers: A comparative study of breast markers in multiple tumors. Breast Cancer Res. Treat. 2020, 184, 11–21. [Google Scholar] [CrossRef] [PubMed]
- Qazi, M.S.; McGregor, S.M. Combined use of SOX10 and GATA3 in mammary carcinoma. Pathol. Res. Pract. 2020, 216, 152801. [Google Scholar] [CrossRef] [PubMed]
- Statz, E.; Jorns, J.M. Cytokeratin 7, GATA3, and SOX-10 is a Comprehensive Panel in Diagnosing Triple Negative Breast Cancer Brain Metastases. Int. J. Surg. Pathol. 2021. [Google Scholar] [CrossRef] [PubMed]
- Kawaji, H.; Kubo, M.; Yamashita, N.; Yamamoto, H.; Kai, M.; Kajihara, A.; Yamada, M.; Kurata, K.; Kaneshiro, K.; Harada, Y.; et al. Comprehensive molecular profiling broadens treatment options for breast cancer patients. Cancer Med. 2021, 10, 529–539. [Google Scholar] [CrossRef]
- Seong, H.A.; Kim, K.T.; Ha, H. Enhancement of B-MYB transcriptional activity by ZPR9, a novel zinc finger protein. J. Biol. Chem. 2003, 278, 9655–9662. [Google Scholar] [CrossRef] [PubMed]
- Seong, H.A.; Jung, H.; Manoharan, R.; Ha, H. Positive regulation of apoptosis signal-regulating kinase 1 signaling by ZPR9 protein, a zinc finger protein. J. Biol. Chem. 2011, 286, 31123–31135. [Google Scholar] [CrossRef]
- Zhou, Q.; Ren, J.; Hou, J.; Wang, G.; Ju, L.; Xiao, Y.; Gong, Y. Co-expression network analysis identified candidate biomarkers in association with progression and prognosis of breast cancer. J. Cancer Res. Clin. Oncol. 2019, 145, 2383–2396. [Google Scholar] [CrossRef] [PubMed]
- Fiscon, G.; Pegoraro, S.; Conte, F.; Manfioletti, G.; Paci, P. Gene network analysis using SWIM reveals interplay between the transcription factor-encoding genes HMGA1, FOXM1, and MYBL2 in triple-negative breast cancer. FEBS Lett. 2021. [Google Scholar] [CrossRef]
- Kleivi, K.; Lind, G.E.; Diep, C.B.; Meling, G.I.; Brandal, L.T.; Nesland, J.M.; Myklebost, O.; Rognum, T.O.; Giercksky, K.E.; Skotheim, R.I.; et al. Gene expression profiles of primary colorectal carcinomas, liver metastases, and carcinomatoses. Mol. Cancer 2007, 6, 2. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Perets, R.; Kaplan, T.; Stein, I.; Hidas, G.; Tayeb, S.; Avraham, E.; Ben-Neriah, Y.; Simon, I.; Pikarsky, E. Genome-wide analysis of androgen receptor targets reveals COUP-TF1 as a novel player in human prostate cancer. PLoS ONE 2012, 7, e46467. [Google Scholar] [CrossRef] [PubMed]



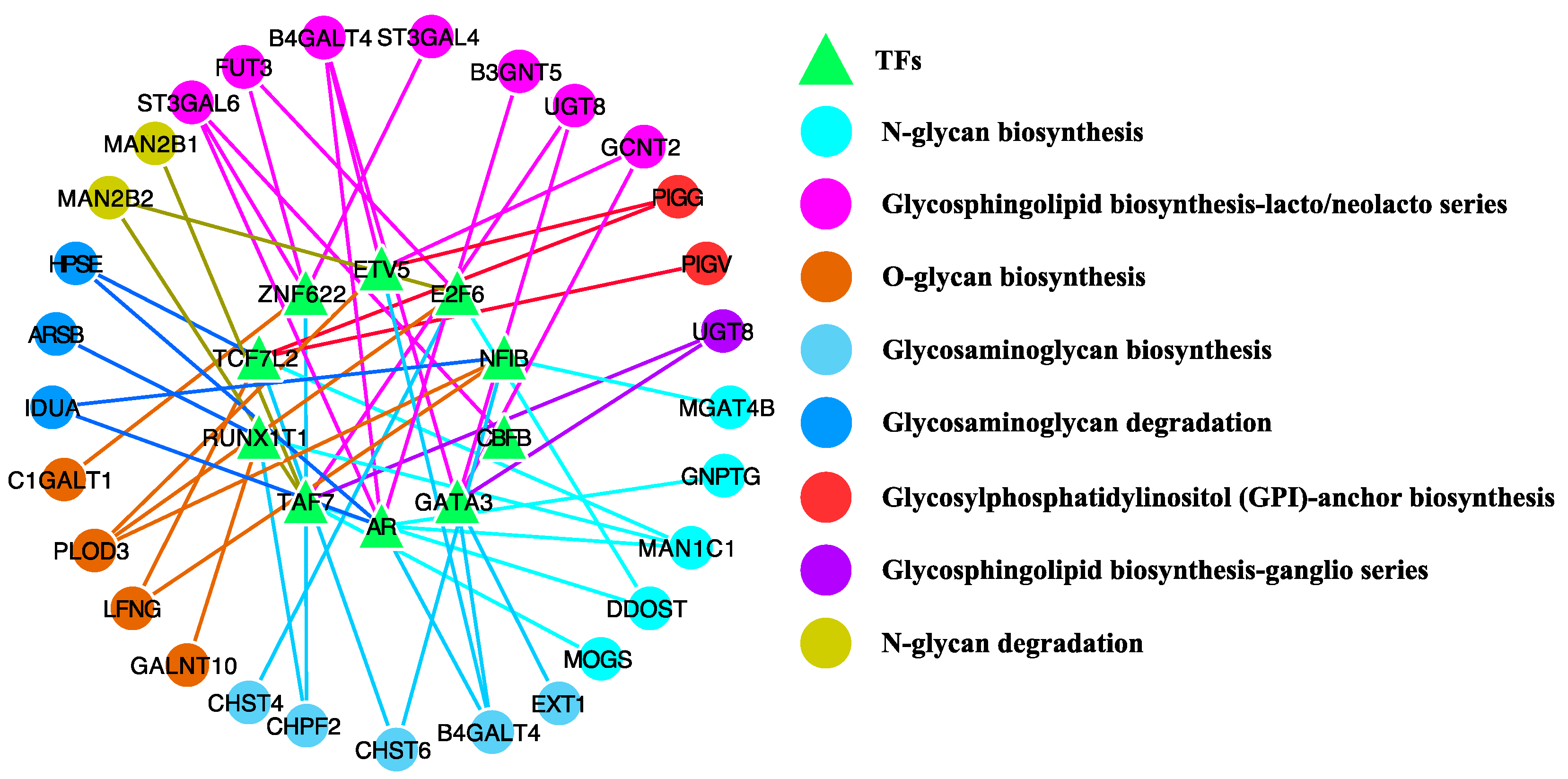
| TFs | Target Glycogenes | Number of Targets |
|---|---|---|
| AR | ST3GAL6, HPSE, IDUA, MAN1C1, B3GNT5, GNPTG, B4GALT4 | 7 |
| TCF7L2 | HPSE, LFNG, CHST6, PIGG, PIGV, MAN1C1 | 6 |
| NFIB | MGAT4B, IDUA, LFNG, CHST6, PLOD3 | 5 |
| TAF7 | DDOST, MAN2B2, MAN2B1, UGT8, MOGS | 5 |
| ZNF622 | ST3GAL6, FUT3, CHPF2, C1GALT1, ST3GAL4 | 5 |
| E2F6 | CHST4, DDOST, MAN2B2, FUT3, PLOD3 | 5 |
| GATA3 | EXT1, GCNT2, UGT8, B4GALT4 | 4 |
| RUNX1T1 | ARSB, CHPF2, GALNT10, MAN1C1 | 4 |
| ETV5 | GCNT2, PIGG, B4GALT4, PLOD3 | 4 |
| CBFB | ST3GAL6, IDUA, CHST6, UGT8 | 4 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Zeng, R.; Mohamed, A.; Khanna, K.K.; Hill, M.M. Differential Regulation of Lacto-/Neolacto- Glycosphingolipid Biosynthesis Pathway Reveals Transcription Factors as Potential Candidates in Triple-Negative Breast Cancer. Cancers 2021, 13, 3330. https://doi.org/10.3390/cancers13133330
Zeng R, Mohamed A, Khanna KK, Hill MM. Differential Regulation of Lacto-/Neolacto- Glycosphingolipid Biosynthesis Pathway Reveals Transcription Factors as Potential Candidates in Triple-Negative Breast Cancer. Cancers. 2021; 13(13):3330. https://doi.org/10.3390/cancers13133330
Chicago/Turabian StyleZeng, Ruichao, Ahmed Mohamed, Kum Kum Khanna, and Michelle M. Hill. 2021. "Differential Regulation of Lacto-/Neolacto- Glycosphingolipid Biosynthesis Pathway Reveals Transcription Factors as Potential Candidates in Triple-Negative Breast Cancer" Cancers 13, no. 13: 3330. https://doi.org/10.3390/cancers13133330
APA StyleZeng, R., Mohamed, A., Khanna, K. K., & Hill, M. M. (2021). Differential Regulation of Lacto-/Neolacto- Glycosphingolipid Biosynthesis Pathway Reveals Transcription Factors as Potential Candidates in Triple-Negative Breast Cancer. Cancers, 13(13), 3330. https://doi.org/10.3390/cancers13133330








