Development and Validation of an Interpretable Artificial Intelligence Model to Predict 10-Year Prostate Cancer Mortality
Abstract
Simple Summary
Abstract
1. Introduction
2. Materials and Methods
2.1. Data
2.2. Feature Selection
- (1)
- Prostate cancer: PSA, T, N, M stage, Gleason score, and initial treatment (if performed)
- (2)
- Medical history: Age, height, weight, current smoking status, smoking pack-years, daily alcohol consumption, history of prostatitis, nocturia, arthritis, bronchitis, diabetes, emphysema, heart attack, hypertension, liver disease, osteoporosis, stroke, elevated cholesterol.
- (3)
- Physical activity: Activity at least once a month during the last year, physical activity at work
- (4)
- Socio-economic status: Family income, education
- (5)
- Hormonal status: Hair pattern at age 45, weight gain pattern
2.3. Predictions
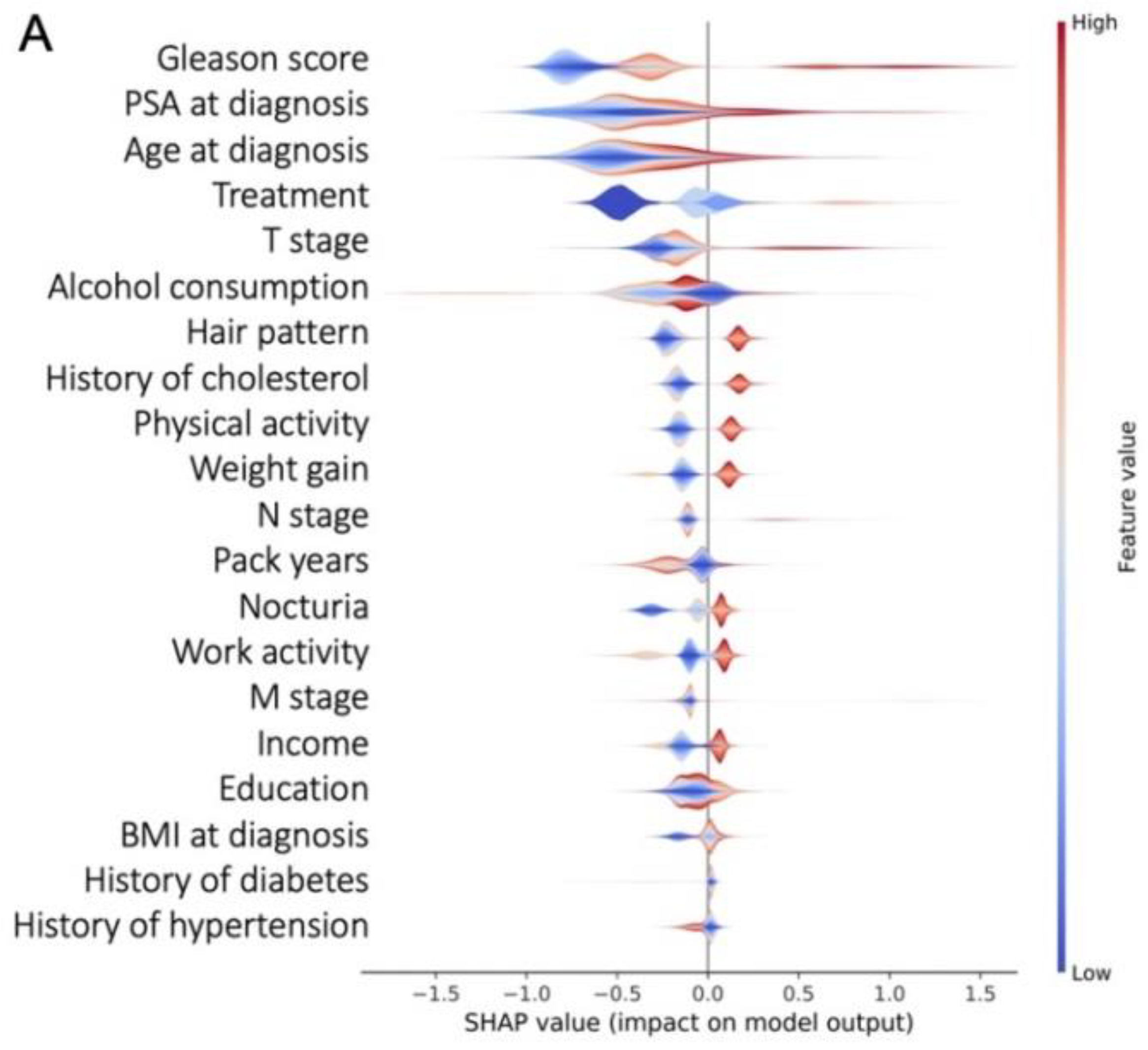
2.4. Model Interpretability
2.5. Online Model Deployment
3. Results
3.1. Cohort Description
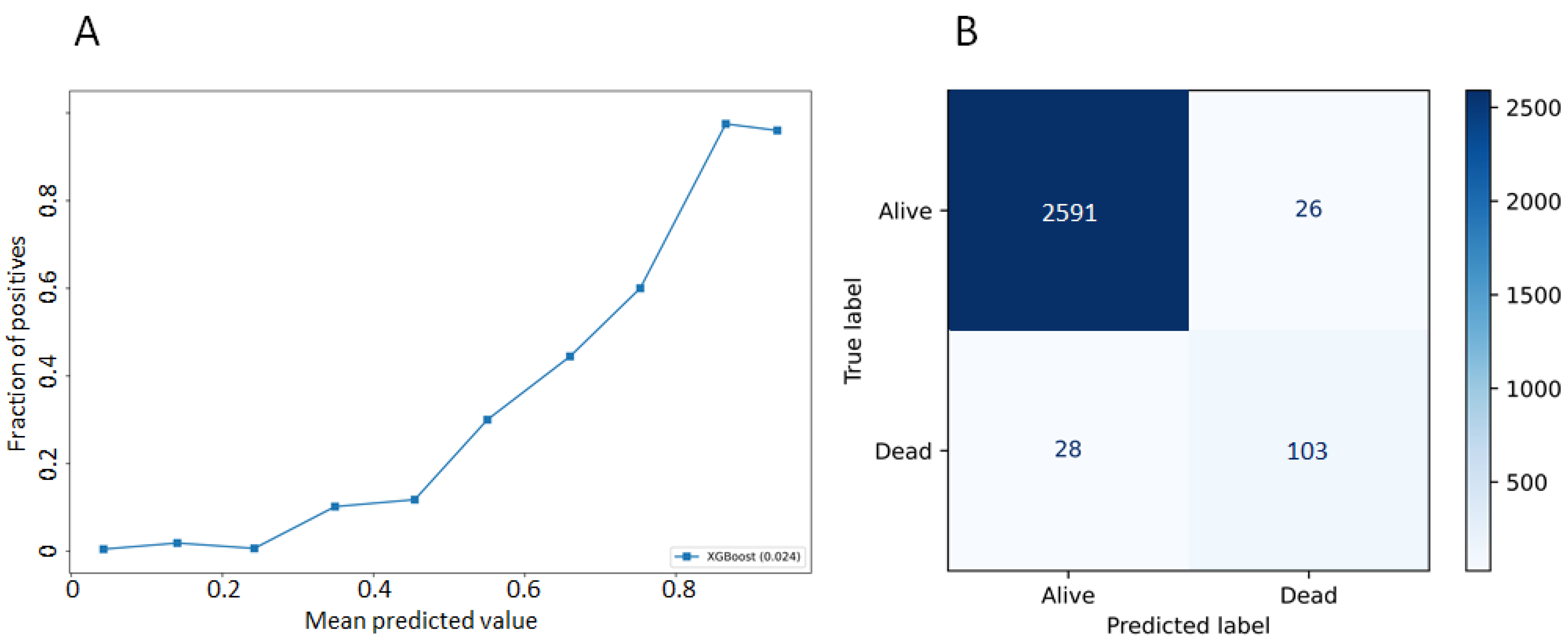
3.2. Model Performances
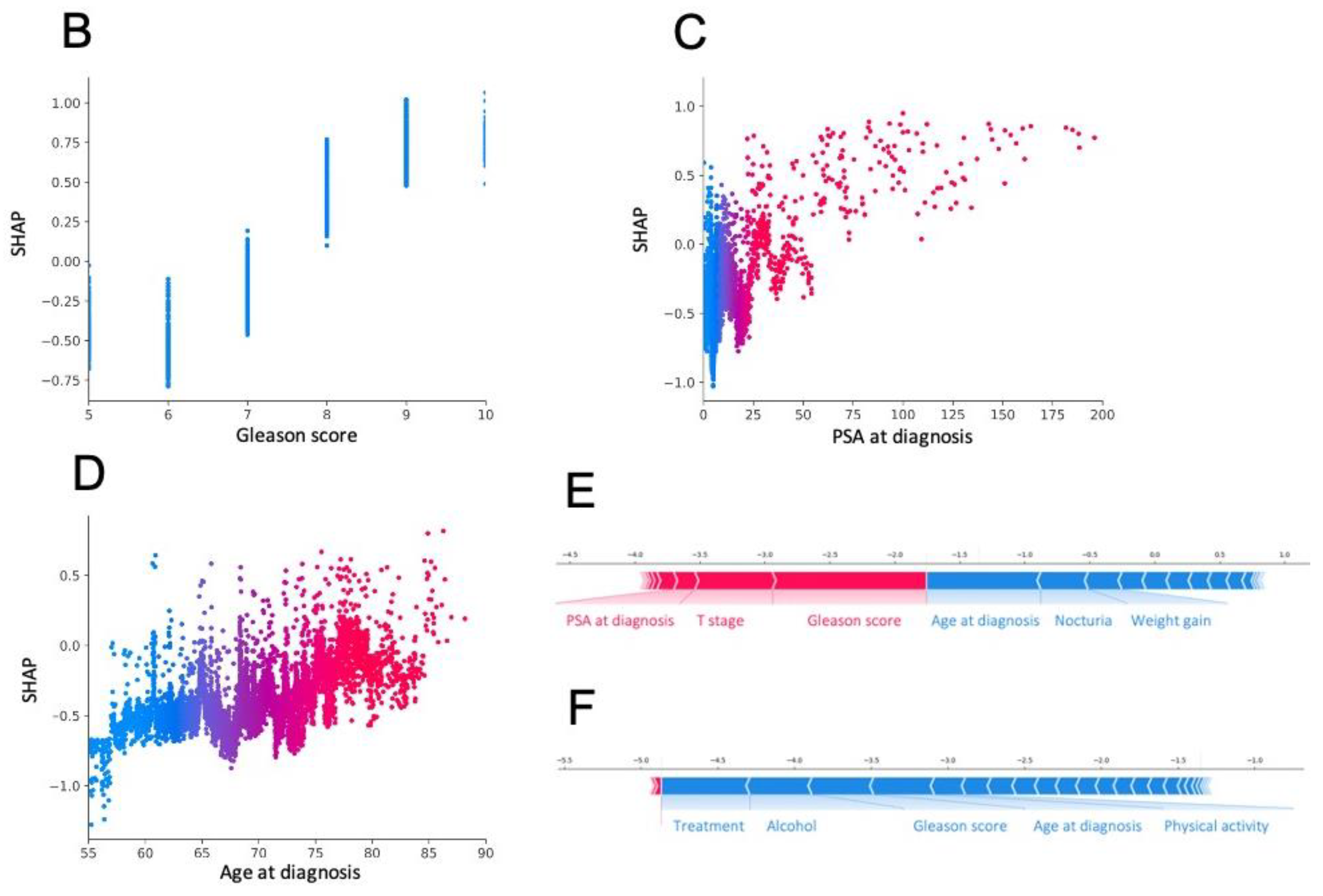
3.3. Most Important Features Explaining the Prediction
3.4. Model Deployment Online and Interpretation at the Individual Scale
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Lancet, T. Discuss prostate cancer screening with your doctor. Lancet 2017, 389, 1582. [Google Scholar] [CrossRef]
- Getaneh, A.M.; Heijnsdijk, E.A.M.; Roobol, M.J.; de Koning, H.J. Assessment of harms, benefits, and cost-effectiveness of prostate cancer screening: A micro-simulation study of 230 scenarios. Cancer Med. 2020, 9, 7742–7750. [Google Scholar] [CrossRef]
- Barnett, R. Prostate cancer. Lancet 2018, 392, 908. [Google Scholar] [CrossRef]
- Braillon, A.; Dubois, G. Re: Prostate cancer screening in the randomized prostate, lung, colorectal, and ovarian cancer screening trial: Mortality results after 13 years of follow-up. J. Natl. Cancer Inst. 2012, 104, 793–794. [Google Scholar] [CrossRef][Green Version]
- Andriole, G.L.; Crawford, E.D.; Grubb, R.L.; Buys, S.S.; Chia, D.; Church, T.R.; Fouad, M.N.; Gelmann, E.P.; Kvale, P.A.; Reding, D.J.; et al. Mortality results from a randomized prostate-cancer screening trial. N. Engl. J. Med. 2009, 360, 1310–1319. [Google Scholar] [CrossRef] [PubMed]
- Andriole, G.L.; Crawford, E.D.; Grubb, R.L.; Buys, S.S.; Chia, D.; Church, T.R.; Fouad, M.N.; Isaacs, C.; Kvale, P.A.; Reding, D.J.; et al. Prostate cancer screening in the randomized prostate, lung, colorectal, and ovarian cancer screening trial: Mortality results after 13 years of follow-up. J. Natl. Cancer Inst. 2012, 104, 125–132. [Google Scholar] [CrossRef] [PubMed]
- Pinsky, P.F.; Prorok, P.C.; Yu, K.; Kramer, B.S.; Black, A.; Gohagan, J.K.; Crawford, E.D.; Grubb, R.L.; Andriole, G.L. Extended mortality results for prostate cancer screening in the PLCO trial with median follow-up of 15 years. Cancer 2017, 123, 592–599. [Google Scholar] [CrossRef] [PubMed]
- Hamdy, F.C.; Donovan, J.L.; Lane, J.A.; Mason, M.; Metcalfe, C.; Holding, P.; Davis, M.; Peters, T.J.; Turner, E.L.; Martin, R.M.; et al. 10-Year outcomes after monitoring, surgery, or radiotherapy for localized prostate cancer. N. Engl. J. Med. 2016, 375, 1415–1424. [Google Scholar] [CrossRef]
- Donovan, J.L.; Hamdy, F.C.; Lane, J.A.; Mason, M.; Metcalfe, C.; Walsh, E.; Blazeby, J.M.; Peters, T.J.; Holding, P.; Bonnington, S.; et al. Patient-Reported outcomes after monitoring, surgery, or radiotherapy for prostate cancer. N. Engl. J. Med. 2016, 375, 1425–1437. [Google Scholar] [CrossRef]
- PLCO—The Cancer Data Access System. Available online: https://cdas.cancer.gov/plco/ (accessed on 22 January 2020).
- Cooperberg, M.R.; Hilton, J.F.; Carroll, P.R. The CAPRA-S Score: A straightforward tool for improved prediction of outcomes after radical prostatectomy. Cancer 2011, 117, 5039–5046. [Google Scholar] [CrossRef]
- Guinney, J.; Wang, T.; Laajala, T.D.; Winner, K.K.; Bare, J.C.; Neto, E.C.; Khan, S.A.; Peddinti, G.; Airola, A.; Pahikkala, T.; et al. Prediction of overall survival for patients with metastatic castration-resistant prostate cancer: Development of a prognostic model through a crowdsourced challenge with open clinical trial data. Lancet Oncol. 2017, 18, 132–142. [Google Scholar] [CrossRef]
- Thurtle, D.R.; Greenberg, D.C.; Lee, L.S.; Huang, H.H.; Pharoah, P.D.; Gnanapragasam, V.J. Individual prognosis at diagnosis in nonmetastatic prostate cancer: Development and external validation of the PREDICT prostate multivariable model. PLoS Med. 2019, 16, e1002758. [Google Scholar] [CrossRef] [PubMed]
- Cooperberg, M.R.; Broering, J.M.; Carroll, P.R. Risk assessment for prostate cancer metastasis and mortality at the time of diagnosis. J. Natl. Cancer Inst. 2009, 101, 878–887. [Google Scholar] [CrossRef] [PubMed]
- Shariat, S.F.; Karakiewicz, P.I.; Roehrborn, C.G.; Kattan, M.W. An updated catalog of prostate cancer predictive tools. Cancer 2008, 113, 3075–3099. [Google Scholar] [CrossRef] [PubMed]
- Riviere, P.; Tokeshi, C.; Hou, J.; Nalawade, V.; Sarkar, R.; Paravati, A.J.; Schiaffino, M.; Rose, B.; Xu, R.; Murphy, J.D. Claims-Based approach to predict cause-specific survival in men with prostate cancer. JCO Clin. Cancer Inform. 2019, 1–7. [Google Scholar] [CrossRef]
- Chen, T.; Guestrin, C. XGBoost: A scalable tree boosting system. In Proceedings of the 22nd ACM SIGKDD International Conference on Knowledge Discovery and Data Mining—KDD ’16, San Francisco, CA, USA, 13–17 August 2016; pp. 785–794. [Google Scholar] [CrossRef]
- Josse, J.; Prost, N.; Scornet, E.; Varoquaux, G. On the consistency of supervised learning with missing values. arXiv 2019, arXiv:1902.06931. [Google Scholar]
- Fernando, N. Fmfn/BayesianOptimization. Available online: https://github.com/fmfn/BayesianOptimization (accessed on 18 June 2021).
- Lundberg, S.; Lee, S.-I. A Unified Approach to Interpreting Model Predictions. arXiv 2017, arXiv:1705.07874. [Google Scholar]
- Lundberg, S. Slundberg/Shap. Available online: https://github.com/slundberg/shap (accessed on 18 June 2021).
- Prostate Cancer Nomograms|Memorial Sloan Kettering Cancer Center. Available online: https://www.mskcc.org/nomograms/prostate (accessed on 23 January 2020).
- Goldenberg, S.L.; Nir, G.; Salcudean, S.E. A New Era: Artificial intelligence and machine learning in prostate cancer. Nat. Rev. Urol. 2019, 16, 391–403. [Google Scholar] [CrossRef] [PubMed]
- Hameed, B.M.; Dhavileswarapu, S.; Aiswarya, V.L.; Raza, S.Z.; Karimi, H.; Khanuja, H.S.; Shetty, D.K.; Ibrahim, S.; Shah, M.J.; Naik, N.; et al. Artificial intelligence and its impact on urological diseases and management: A comprehensive review of the literature. J. Clin. Med. 2021, 10, 1864. [Google Scholar] [CrossRef]
- Yuan, Y.; Qin, W.; Buyyounouski, M.; Ibragimov, B.; Hancock, S.; Han, B.; Xing, L. Prostate cancer classification with multiparametric MRI transfer learning model. Med. Phys. 2019, 46, 756–765. [Google Scholar] [CrossRef]
- Campanella, G.; Hanna, M.G.; Geneslaw, L.; Miraflor, A.; Werneck Krauss Silva, V.; Busam, K.J.; Brogi, E.; Reuter, V.E.; Klimstra, D.S.; Fuchs, T.J. Clinical-Grade computational pathology using weakly supervised deep learning on whole slide images. Nat. Med. 2019, 25, 1301–1309. [Google Scholar] [CrossRef] [PubMed]
- Li, D.; Zang, P.; Chai, X.; Cui, Y.; Li, R.; Xing, L. Automatic multiorgan segmentation in CT images of the male pelvis using region-specific hierarchical appearance cluster models. Med. Phys. 2016, 43, 5426. [Google Scholar] [CrossRef] [PubMed]
- Bibault, J.-E.; Giraud, P.; Housset, M.; Durdux, C.; Taieb, J.; Berger, A.; Coriat, R.; Chaussade, S.; Dousset, B.; Nordlinger, B. Deep learning and radiomics predict complete response after neo-adjuvant chemoradiation for locally advanced rectal cancer. Sci. Rep. 2018, 8, 1–8. [Google Scholar]
- Wang, H.; Dong, P.; Liu, H.; Xing, L. Development of an autonomous treatment planning strategy for radiation therapy with effective use of population-based prior data. Med. Phys. 2017, 44, 389–396. [Google Scholar] [CrossRef] [PubMed]
- Zhao, W.; Han, B.; Yang, Y.; Buyyounouski, M.; Hancock, S.L.; Bagshaw, H.; Xing, L. Incorporating imaging information from deep neural network layers into Image Guided Radiation Therapy (IGRT). Radiother. Oncol. 2019, 140, 167–174. [Google Scholar] [CrossRef] [PubMed]
| Characteristic | No. (%) All Patients | No. (%) Patients Who Died from PCa |
|---|---|---|
| Age | ||
| Under 65 years old | 1990 (22.7) | 109 (15.9) |
| Between 65 and 75 years old | 5181 (59) | 283 (41.3) |
| Over 75 years old | 1605 (18.3) | 293 (42.8) |
| Prostate Cancer | ||
| Localized PCa | 7668 (87.4) | 436 (63.6) |
| Low-risk | 2940 (33.5) | 199 (29.1) |
| Intermediate-risk | 3476 (39.6) | 105 (15.3) |
| High risk | 1252 (14.3) | 132 (19.3) |
| Locally advanced PCa | 913 (10.4) | 122 (17.8) |
| Metastatic PCa | 195 (2.2) | 127 (18.5) |
| PSA | ||
| <10 ng/mL | 6516 (74.2) | 254 (37.1) |
| 10–20 ng/mL | 1137 (13) | 94 (13.7) |
| >20 ng/ml | 1123 (12.8) | 337 (49.2) |
| Gleason score | ||
| Gleason ≤ 6 | 4744 (54.1) | 353 (51.5) |
| Gleason 7 | 2842 (32.4) | 158 (23.1) |
| Gleason 8 | 607 (6.9) | 95 (13.9 |
| Gleason ≤ 9 | 455 (5.2) | 48 (7) |
| N/A | 128 (1.5) | 31 (4.5) |
| Treatment | ||
| Surgery | 3212 (36.6) | 114 (16.6) |
| Radiotherapy | 3607 (41.1) | 201 (29.3) |
| Chemotherapy | 1067 (12.2) | 54 (7.9) |
| Hormonotherapy | 654 (7.5) | 161 (23.5) |
| N/A | 236 (2.7) | 155 (22.6) |
| Metric | Definition | Result |
|---|---|---|
| Accuracy | Number of correct predictions/total number of input samples | 0.98 (±0.01) |
| Precision | Number of correct positive predictions/number of positive predictions | 0.80 (±0.1) |
| Recall | Number of correct positive predictions/number of all positive samples | 0.60 (±0.08) |
| f1-score | Harmonic mean of the precision and the recall | 0.66 (±0.07) |
| auROC | Area under the curve of the true positive rate and false positive rate at various thresholds | 0.80 (±0.04) |
| prAUC | Area under the curve of precision and recall at various thresholds | 0.54 (±0.07) |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Bibault, J.-E.; Hancock, S.; Buyyounouski, M.K.; Bagshaw, H.; Leppert, J.T.; Liao, J.C.; Xing, L. Development and Validation of an Interpretable Artificial Intelligence Model to Predict 10-Year Prostate Cancer Mortality. Cancers 2021, 13, 3064. https://doi.org/10.3390/cancers13123064
Bibault J-E, Hancock S, Buyyounouski MK, Bagshaw H, Leppert JT, Liao JC, Xing L. Development and Validation of an Interpretable Artificial Intelligence Model to Predict 10-Year Prostate Cancer Mortality. Cancers. 2021; 13(12):3064. https://doi.org/10.3390/cancers13123064
Chicago/Turabian StyleBibault, Jean-Emmanuel, Steven Hancock, Mark K. Buyyounouski, Hilary Bagshaw, John T. Leppert, Joseph C. Liao, and Lei Xing. 2021. "Development and Validation of an Interpretable Artificial Intelligence Model to Predict 10-Year Prostate Cancer Mortality" Cancers 13, no. 12: 3064. https://doi.org/10.3390/cancers13123064
APA StyleBibault, J.-E., Hancock, S., Buyyounouski, M. K., Bagshaw, H., Leppert, J. T., Liao, J. C., & Xing, L. (2021). Development and Validation of an Interpretable Artificial Intelligence Model to Predict 10-Year Prostate Cancer Mortality. Cancers, 13(12), 3064. https://doi.org/10.3390/cancers13123064








