CUX1 Enhances Pancreatic Cancer Formation by Synergizing with KRAS and Inducing MEK/ERK-Dependent Proliferation
Abstract
Simple Summary
Abstract
1. Introduction
2. Materials and Methods
2.1. Cell Culture
2.2. Retroviral Plasmids and Infection
2.3. Western Blot
2.4. Quantitative Real-Time PCR (qPCR)
2.5. Flow Cytometry
2.6. Colony Formation Assay
2.7. Mouse Experiments
2.8. Histology and Immunohistochemistry
2.9. Statistical Analysis
3. Results
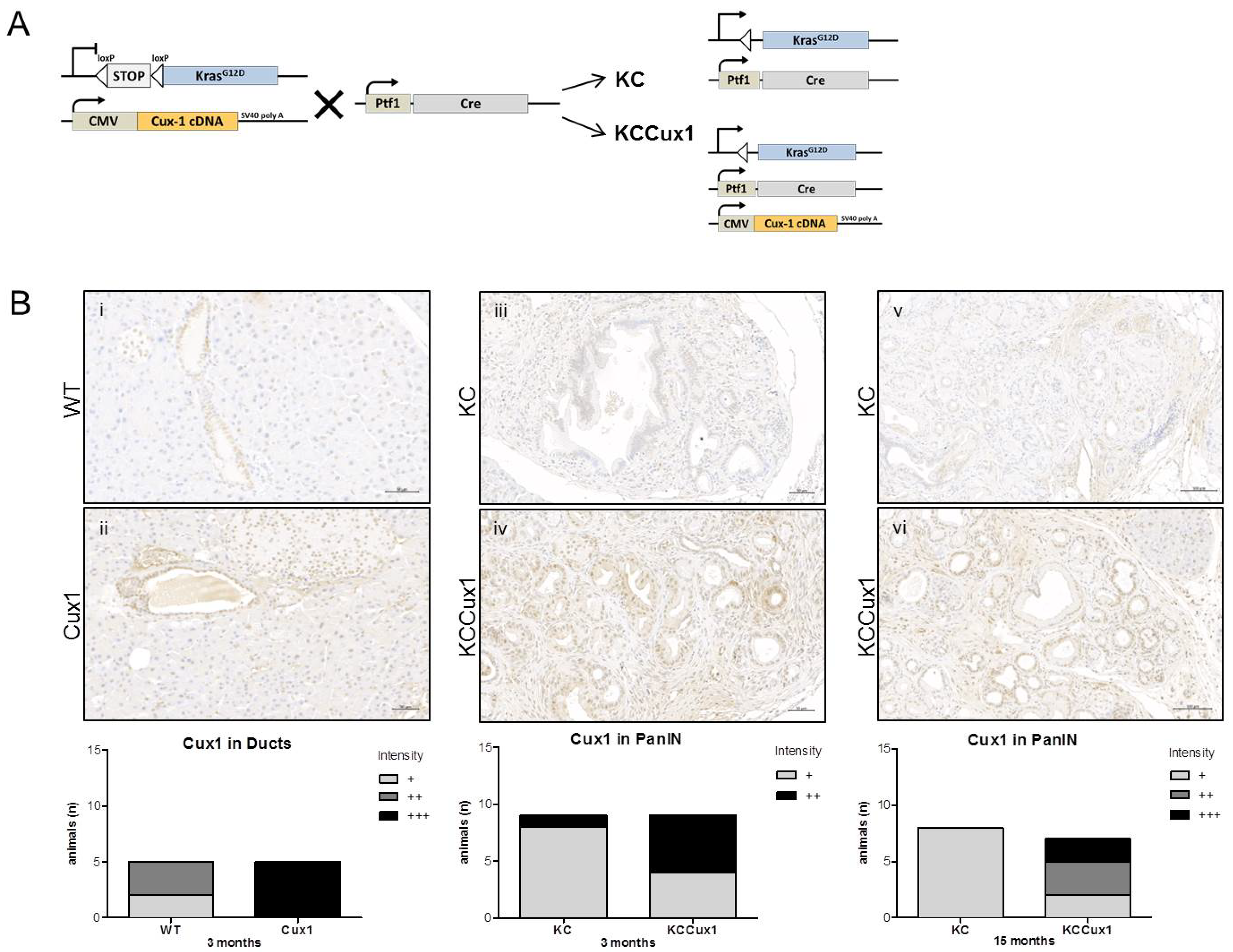
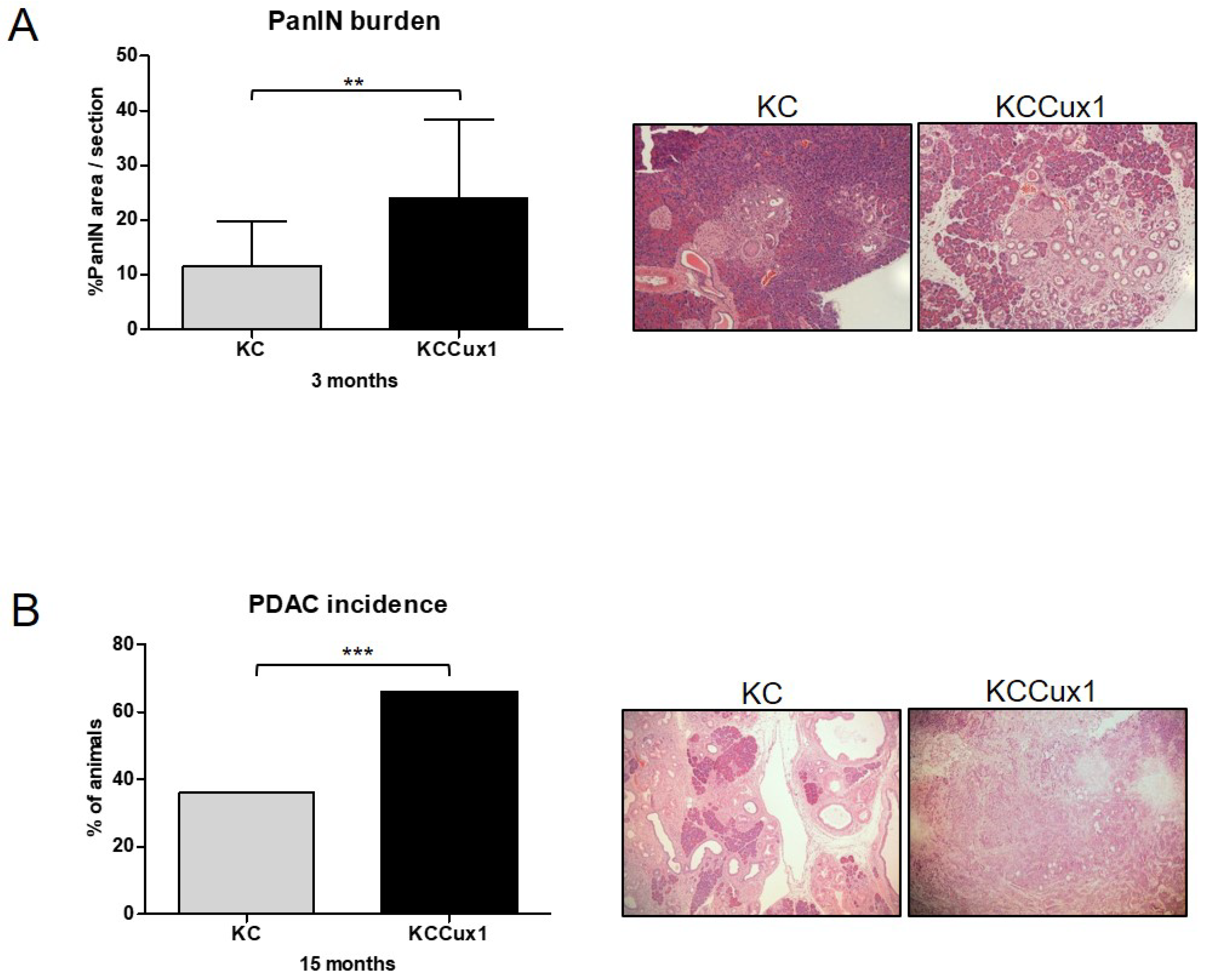
3.1. CUX1 Accelerates Pancreatic Tumorigenesis in KrasG12D Mice
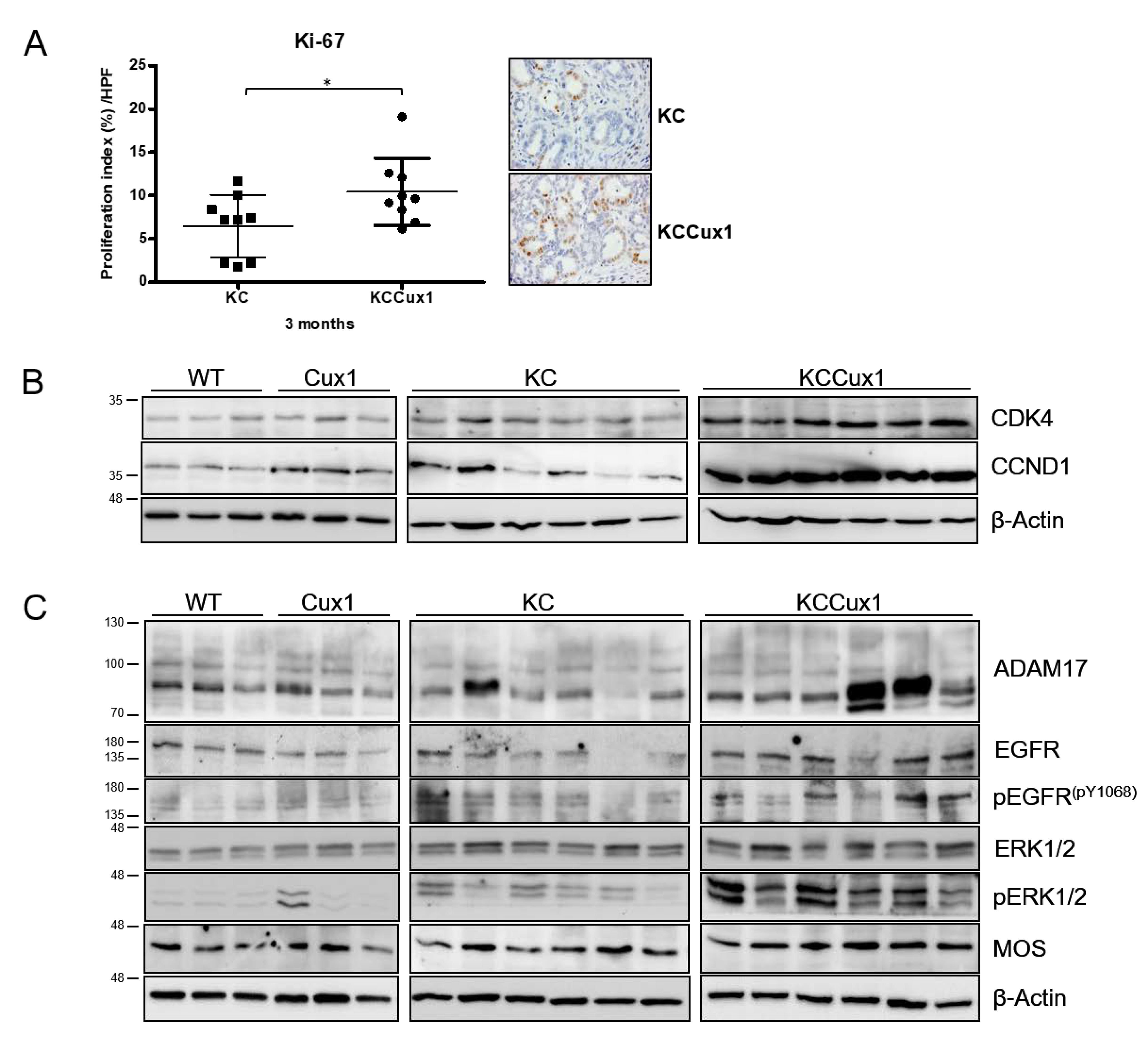
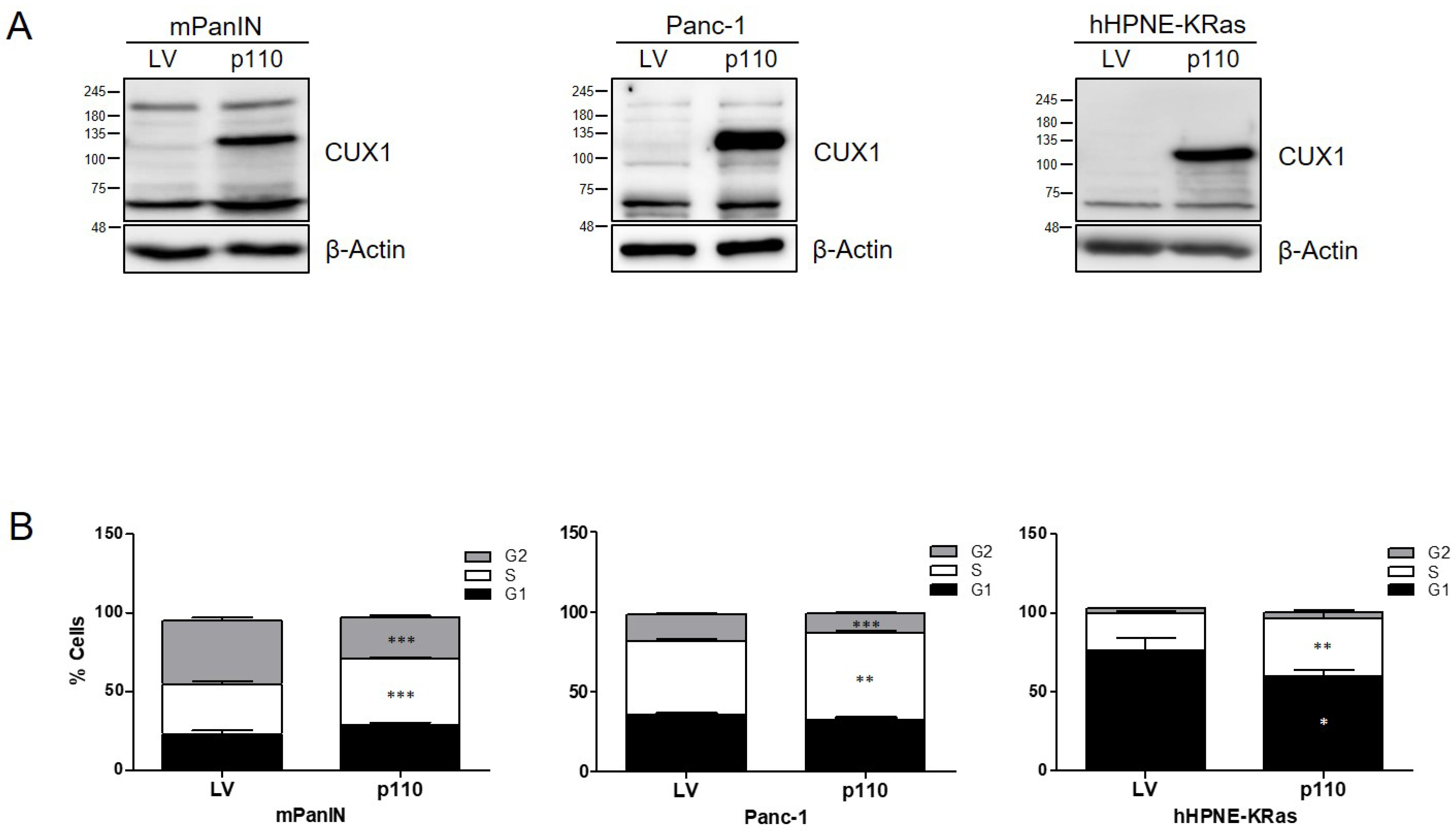
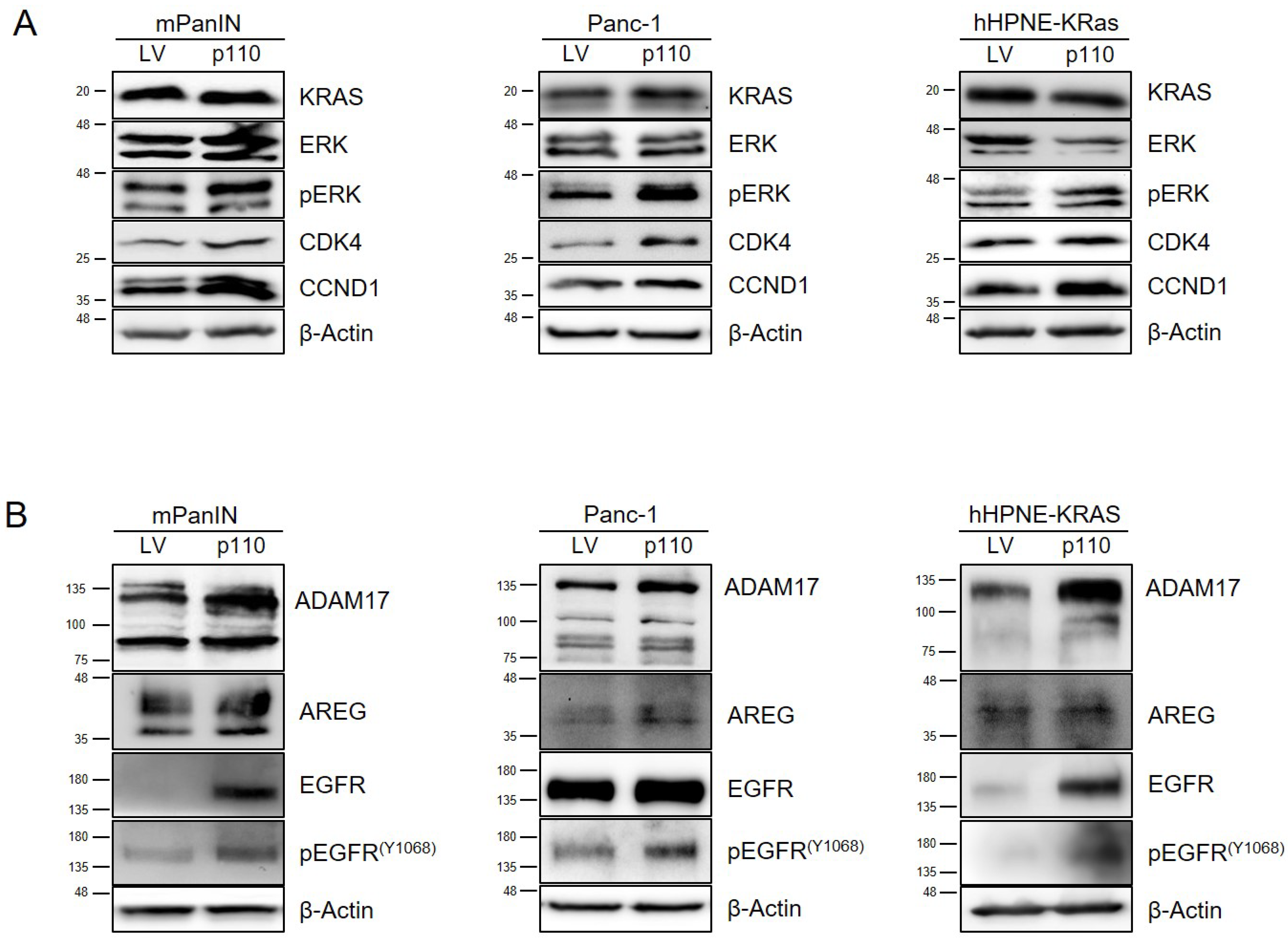
3.2. CUX1 Enhances Proliferation Pathways during Tumorigenesis
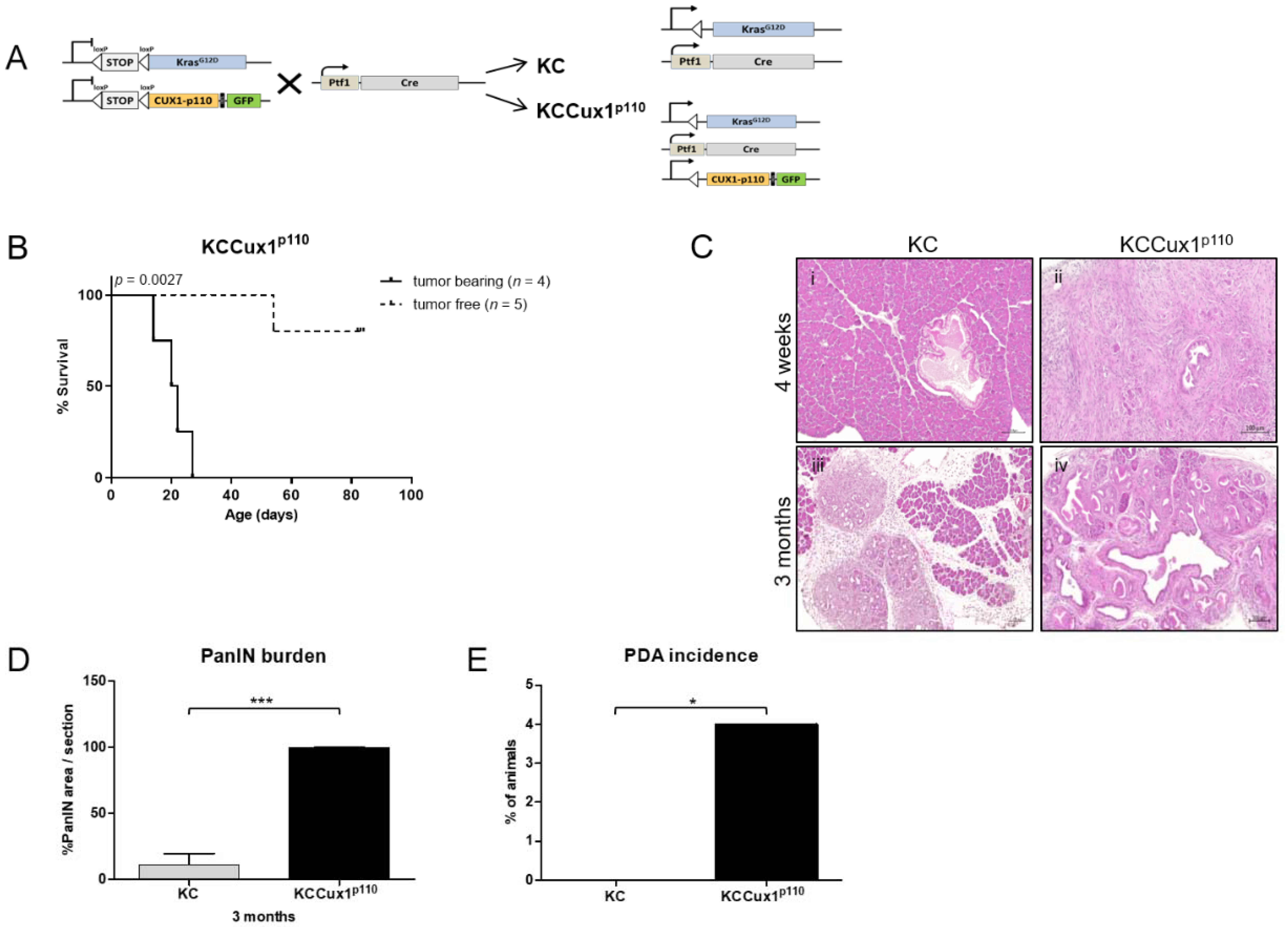
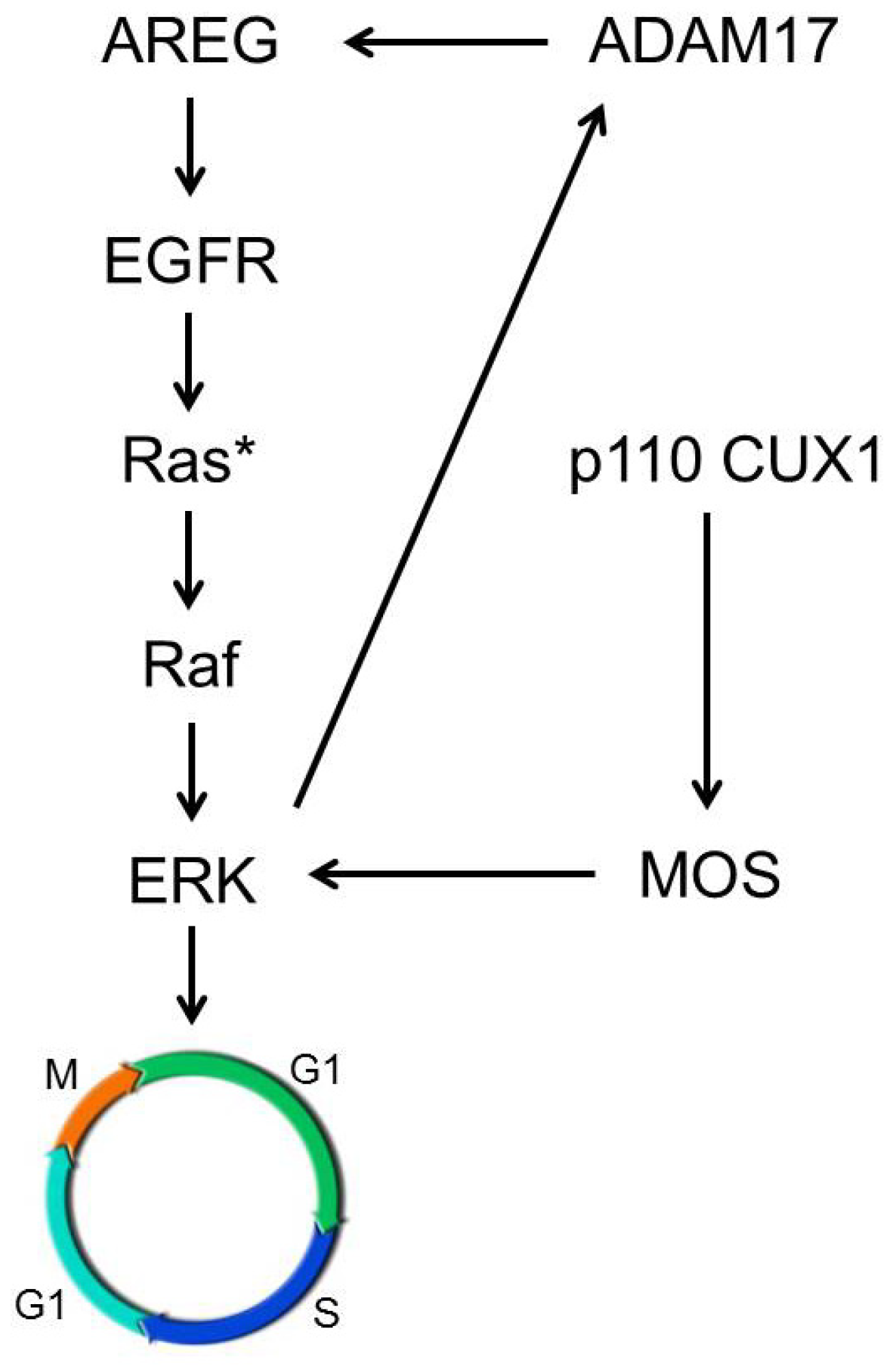
3.3. p110 CUX1 Is Crucial for Enhancing KRAS-Driven Tumorigenesis
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Siegel, R.L.; Miller, K.D.; Jemal, A. Cancer statistics, 2018. CA Cancer J. Clin. 2018, 68, 7–30. [Google Scholar] [CrossRef]
- Hingorani, S.R.; Petricoin, E.F.; Maitra, A.; Rajapakse, V.; King, C.; Jacobetz, M.A.; Ross, S.; Conrads, T.P.; Veenstra, T.D.; Hitt, B.A.; et al. Preinvasive and invasive ductal pancreatic cancer and its early detection in the mouse. Cancer Cell 2003, 4, 437–450. [Google Scholar] [CrossRef]
- Luttges, J.; Reinecke-Luthge, A.; Mollmann, B.; Menke, M.A.; Clemens, A.; Klimpfinger, M.; Sipos, B.; Kloppel, G. Duct changes and K-ras mutations in the disease-free pancreas: Analysis of type, age relation and spatial distribution. Virchows Arch. 1999, 435, 461–468. [Google Scholar] [CrossRef] [PubMed]
- Truscott, M.; Harada, R.; Vadnais, C.; Robert, F.; Nepveu, A. p110 CUX1 cooperates with E2F transcription factors in the transcriptional activation of cell cycle-regulated genes. Mol. Cell Biol. 2008, 28, 3127–3138. [Google Scholar] [CrossRef] [PubMed]
- Hulea, L.; Nepveu, A. CUX1 transcription factors: From biochemical activities and cell-based assays to mouse models and human diseases. Gene 2012, 497, 18–26. [Google Scholar] [CrossRef] [PubMed]
- Boultwood, J. CUX1 in leukemia: Dosage matters. Blood 2013, 121, 869–871. [Google Scholar] [CrossRef]
- Michl, P.; Ramjaun, A.R.; Pardo, O.E.; Warne, P.H.; Wagner, M.; Poulsom, R.; D’Arrigo, C.; Ryder, K.; Menke, A.; Gress, T.; et al. CUTL1 is a target of TGF(beta) signaling that enhances cancer cell motility and invasiveness. Cancer Cell 2005, 7, 521–532. [Google Scholar] [CrossRef]
- Ripka, S.; Neesse, A.; Riedel, J.; Bug, E.; Aigner, A.; Poulsom, R.; Fulda, S.; Neoptolemos, J.; Greenhalf, W.; Barth, P.; et al. CUX1: Target of Akt signalling and mediator of resistance to apoptosis in pancreatic cancer. Gut 2010, 59, 1101–1110. [Google Scholar] [CrossRef]
- Nepveu, A. Role of the multifunctional CDP/Cut/Cux homeodomain transcription factor in regulating differentiation, cell growth and development. Gene 2001, 270, 1–15. [Google Scholar] [CrossRef]
- Aufiero, B.; Neufeld, E.J.; Orkin, S.H. Sequence-specific DNA binding of individual cut repeats of the human CCAAT displacement/cut homeodomain protein. Proc. Natl. Acad. Sci. USA 1994, 91, 7757–7761. [Google Scholar] [CrossRef]
- Harada, R.; Dufort, D.; Denis-Larose, C.; Nepveu, A. Conserved cut repeats in the human cut homeodomain protein function as DNA binding domains. J. Biol. Chem. 1994, 269, 2062–2067. [Google Scholar] [CrossRef]
- Neufeld, E.J.; Skalnik, D.G.; Lievens, P.M.; Orkin, S.H. Human CCAAT displacement protein is homologous to the Drosophila homeoprotein, cut. Nat. Genet. 1992, 1, 50–55. [Google Scholar] [CrossRef] [PubMed]
- Andres, V.; Chiara, M.D.; Mahdavi, V. A new bipartite DNA-binding domain: Cooperative interaction between the cut repeat and homeo domain of the cut homeo proteins. Genes Dev. 1994, 8, 245–257. [Google Scholar] [CrossRef][Green Version]
- Moon, N.S.; Berube, G.; Nepveu, A. CCAAT displacement activity involves CUT repeats 1 and 2, not the CUT homeodomain. J. Biol. Chem. 2000, 275, 31325–31334. [Google Scholar] [CrossRef] [PubMed]
- Goulet, B.; Baruch, A.; Moon, N.S.; Poirier, M.; Sansregret, L.L.; Erickson, A.; Bogyo, M.; Nepveu, A. A cathepsin L isoform that is devoid of a signal peptide localizes to the nucleus in S phase and processes the CDP/Cux transcription factor. Mol. Cell 2004, 14, 207–219. [Google Scholar] [CrossRef]
- Harada, R.; Vadnais, C.; Sansregret, L.; Leduy, L.; Berube, G.; Robert, F.; Nepveu, A. Genome-wide location analysis and expression studies reveal a role for p110 CUX1 in the activation of DNA replication genes. Nucleic Acids Res. 2008, 36, 189–202. [Google Scholar] [CrossRef][Green Version]
- Kedinger, V.; Sansregret, L.; Harada, R.; Vadnais, C.; Cadieux, C.; Fathers, K.; Park, M.; Nepveu, A. p110 CUX1 homeodomain protein stimulates cell migration and invasion in part through a regulatory cascade culminating in the repression of E-cadherin and occludin. J. Biol. Chem. 2009, 284, 27701–27711. [Google Scholar] [CrossRef]
- Moon, N.S.; Premdas, P.; Truscott, M.; Leduy, L.; Berube, G.; Nepveu, A. S phase-specific proteolytic cleavage is required to activate stable DNA binding by the CDP/Cut homeodomain protein. Mol. Cell Biol. 2001, 21, 6332–6345. [Google Scholar] [CrossRef] [PubMed]
- Sansregret, L.; Goulet, B.; Harada, R.; Wilson, B.; Leduy, L.; Bertoglio, J.; Nepveu, A. The p110 isoform of the CDP/Cux transcription factor accelerates entry into S phase. Mol. Cell Biol. 2006, 26, 2441–2455. [Google Scholar] [CrossRef]
- Truscott, M.; Denault, J.B.; Goulet, B.; Leduy, L.; Salvesen, G.S.; Nepveu, A. Carboxyl-terminal proteolytic processing of CUX1 by a caspase enables transcriptional activation in proliferating cells. J. Biol. Chem. 2007, 282, 30216–30226. [Google Scholar] [CrossRef]
- Ledford, A.W.; Brantley, J.G.; Kemeny, G.; Foreman, T.L.; Quaggin, S.E.; Igarashi, P.; Oberhaus, S.M.; Rodova, M.; Calvet, J.P.; Vanden Heuvel, G.B. Deregulated expression of the homeobox gene Cux-1 in transgenic mice results in downregulation of p27(kip1) expression during nephrogenesis, glomerular abnormalities, and multiorgan hyperplasia. Dev. Biol. 2002, 245, 157–171. [Google Scholar] [CrossRef] [PubMed]
- Vadnais, C.; Davoudi, S.; Afshin, M.; Harada, R.; Dudley, R.; Clermont, P.L.; Drobetsky, E.; Nepveu, A. CUX1 transcription factor is required for optimal ATM/ATR-mediated responses to DNA damage. Nucleic Acids Res. 2012, 40, 4483–4495. [Google Scholar] [CrossRef] [PubMed]
- Sharma, M.; Fopma, A.; Brantley, J.G.; Vanden Heuvel, G.B. Coexpression of Cux-1 and Notch signaling pathway components during kidney development. Dev. Dyn. 2004, 231, 828–838. [Google Scholar] [CrossRef] [PubMed]
- Cubelos, B.; Sebastian-Serrano, A.; Beccari, L.; Calcagnotto, M.E.; Cisneros, E.; Kim, S.; Dopazo, A.; Alvarez-Dolado, M.; Redondo, J.M.; Bovolenta, P.; et al. Cux1 and Cux2 regulate dendritic branching, spine morphology, and synapses of the upper layer neurons of the cortex. Neuron 2010, 66, 523–535. [Google Scholar] [CrossRef]
- Ellis, T.; Gambardella, L.; Horcher, M.; Tschanz, S.; Capol, J.; Bertram, P.; Jochum, W.; Barrandon, Y.; Busslinger, M. The transcriptional repressor CDP (Cutl1) is essential for epithelial cell differentiation of the lung and the hair follicle. Genes Dev. 2001, 15, 2307–2319. [Google Scholar] [CrossRef]
- Sinclair, A.M.; Lee, J.A.; Goldstein, A.; Xing, D.; Liu, S.; Ju, R.; Tucker, P.W.; Neufeld, E.J.; Scheuermann, R.H. Lymphoid apoptosis and myeloid hyperplasia in CCAAT displacement protein mutant mice. Blood 2001, 98, 3658–3667. [Google Scholar] [CrossRef]
- Skalnik, D.G.; Strauss, E.C.; Orkin, S.H. CCAAT displacement protein as a repressor of the myelomonocytic-specific gp91-phox gene promoter. J. Biol. Chem. 1991, 266, 16736–16744. [Google Scholar] [CrossRef]
- Lievens, P.M.; Donady, J.J.; Tufarelli, C.; Neufeld, E.J. Repressor activity of CCAAT displacement protein in HL-60 myeloid leukemia cells. J. Biol. Chem. 1995, 270, 12745–12750. [Google Scholar] [CrossRef]
- Brantley, J.G.; Sharma, M.; Alcalay, N.I.; Heuvel, G.B. Cux-1 transgenic mice develop glomerulosclerosis and interstitial fibrosis. Kidney Int. 2003, 63, 1240–1248. [Google Scholar] [CrossRef]
- Vanden Heuvel, G.B.; Brantley, J.G.; Alcalay, N.I.; Sharma, M.; Kemeny, G.; Warolin, J.; Ledford, A.W.; Pinson, D.M. Hepatomegaly in transgenic mice expressing the homeobox gene Cux-1. Mol. Carcinog. 2005, 43, 18–30. [Google Scholar] [CrossRef]
- Ramdzan, Z.M.; Vadnais, C.; Pal, R.; Vandal, G.; Cadieux, C.; Leduy, L.; Davoudi, S.; Hulea, L.; Yao, L.; Karnezis, A.N.; et al. RAS transformation requires CUX1-dependent repair of oxidative DNA damage. PLoS Biol. 2014, 12, e1001807. [Google Scholar] [CrossRef] [PubMed]
- Cadieux, C.; Kedinger, V.; Yao, L.; Vadnais, C.; Drossos, M.; Paquet, M.; Nepveu, A. Mouse mammary tumor virus p75 and p110 CUX1 transgenic mice develop mammary tumors of various histologic types. Cancer Res. 2009, 69, 7188–7197. [Google Scholar] [CrossRef] [PubMed]
- Michl, P.; Knobel, B.; Downward, J. CUTL1 is phosphorylated by protein kinase A, modulating its effects on cell proliferation and motility. J. Biol. Chem. 2006, 281, 15138–15144. [Google Scholar] [CrossRef]
- Ardito, C.M.; Gruner, B.M.; Takeuchi, K.K.; Lubeseder-Martellato, C.; Teichmann, N.; Mazur, P.K.; Delgiorno, K.E.; Carpenter, E.S.; Halbrook, C.J.; Hall, J.C.; et al. EGF receptor is required for KRAS-induced pancreatic tumorigenesis. Cancer Cell 2012, 22, 304–317. [Google Scholar] [CrossRef]
- Shaul, Y.D.; Seger, R. The MEK/ERK cascade: From signaling specificity to diverse functions. Biochim. Biophys Acta. 2007, 1773, 1213–1226. [Google Scholar] [CrossRef] [PubMed]
- Okazaki, K.; Sagata, N. The Mos/MAP kinase pathway stabilizes c-Fos by phosphorylation and augments its transforming activity in NIH 3T3 cells. EMBO J. 1995, 14, 5048–5059. [Google Scholar] [CrossRef] [PubMed]
- Siam, R.; Harada, R.; Cadieux, C.; Battat, R.; Vadnais, C.; Nepveu, A. Transcriptional activation of the Lats1 tumor suppressor gene in tumors of CUX1 transgenic mice. Mol. Cancer 2009, 8, e60. [Google Scholar] [CrossRef]
- Burton, L.J.; Hawsawi, O.; Sweeney, J.; Bowen, N.; Hudson, T.; Odero-Marah, V. CCAAT-displacement protein/cut homeobox transcription factor (CUX1) represses estrogen receptor-alpha (ER-α) in triple-negative breast cancer cells and can be antagonized by muscadine grape skin extract (MSKE). PLoS ONE 2019, 14, e0214844. [Google Scholar] [CrossRef]
- Wang, L.; Zhao, Y.; Xiong, Y.; Wang, W.; Fei, Y.; Tan, C.; Liang, Z. K-ras mutation promotes ionizing radiation-induced invasion and migration of lung cancer in part via the Cathepsin L/CUX1 pathway. Exp. Cell Res. 2018, 362, 424–435. [Google Scholar] [CrossRef]
- Blasco, M.T.; Navas, C.; Martin-Serrano, G.; Grana-Castro, O.; Lechuga, C.G.; Martin-Diaz, L.; Djurec, M.; Li, J.; Morales-Cacho, L.; Esteban-Burgos, L.; et al. Complete Regression of Advanced Pancreatic Ductal Adenocarcinomas upon Combined Inhibition of EGFR and C-RAF. Cancer Cell 2019, 35, 573–587 e576. [Google Scholar] [CrossRef]
- Wu, X.; Feng, F.; Yang, C.; Zhang, M.; Cheng, Y.; Zhao, Y.; Wang, Y.; Che, F.; Zhang, J.; Heng, X. Upregulated Expression of CUX1 Correlates with Poor Prognosis in Glioma Patients: A Bioinformatic Analysis. J. Mol. Neurosci. 2019, 69, 527–537. [Google Scholar] [CrossRef] [PubMed]
- Li, H.; Yang, F.; Hu, A.; Wang, X.; Fang, E.; Chen, Y.; Li, D.; Song, H.; Wang, J.; Guo, Y.; et al. Therapeutic targeting of circ-CUX1/EWSR1/MAZ axis inhibits glycolysis and neuroblastoma progression. EMBO Mol. Med. 2019, 11, e10835. [Google Scholar] [CrossRef] [PubMed]
- Wong, C.C.; Martincorena, I.; Rust, A.G.; Rashid, M.; Alifrangis, C.; Alexandrov, L.B.; Tiffen, J.C.; Kober, C. Chronic Myeloid Disorders Working Group of the International Cancer Genome, C.; Green, A.R.; et al. Inactivating CUX1 mutations promote tumorigenesis. Nat. Genet. 2014, 46, 33–38. [Google Scholar] [CrossRef]
- Singh, N.; Das, P.; Datta Gupta, S.; Sahni, P.; Pandey, R.M.; Gupta, S.; Chauhan, S.S.; Saraya, A. Prognostic significance of extracellular matrix degrading enzymes-cathepsin L and matrix metalloproteases-2 [MMP-2] in human pancreatic cancer. Cancer Investig. 2013, 31, 461–471. [Google Scholar] [CrossRef] [PubMed]
- Burton, L.J.; Dougan, J.; Jones, J.; Smith, B.N.; Randle, D.; Henderson, V.; Odero-Marah, V.A. Targeting the Nuclear Cathepsin L CCAAT Displacement Protein/Cut Homeobox Transcription Factor-Epithelial Mesenchymal Transition Pathway in Prostate and Breast Cancer Cells with the Z-FY-CHO Inhibitor. Mol. Cell Biol. 2017, 37. [Google Scholar] [CrossRef]
- Fei, Y.; Xiong, Y.; Shen, X.; Zhao, Y.; Zhu, Y.; Wang, L.; Liang, Z. Cathepsin L promotes ionizing radiation-induced U251 glioma cell migration and invasion through regulating the GSK-3beta/CUX1 pathway. Cell Signal. 2018, 44, 62–71. [Google Scholar] [CrossRef] [PubMed]
- Dorris, E.R.; O’Neill, A.; Treacy, A.; Klocker, H.; Teltsh, O.; Kay, E.; Watson, R.W. The transcription factor CUX1 negatively regulates invasion in castrate resistant prostate cancer. Oncotarget 2020, 11, 846–857. [Google Scholar] [CrossRef][Green Version]
- Chen, N.M.; Singh, G.; Koenig, A.; Liou, G.Y.; Storz, P.; Zhang, J.S.; Regul, L.; Nagarajan, S.; Kuhnemuth, B.; Johnsen, S.A.; et al. NFATc1 Links EGFR Signaling to Induction of Sox9 Transcription and Acinar-Ductal Transdifferentiation in the Pancreas. Gastroenterology 2015, 148, 1024–1034 e1029. [Google Scholar] [CrossRef]
- Diersch, S.; Wirth, M.; Schneeweis, C.; Jors, S.; Geisler, F.; Siveke, J.T.; Rad, R.; Schmid, R.M.; Saur, D.; Rustgi, A.K.; et al. Kras(G12D) induces EGFR-MYC cross signaling in murine primary pancreatic ductal epithelial cells. Oncogene 2016, 35, 3880–3886. [Google Scholar] [CrossRef]
- Dusterhoft, S.; Lokau, J.; Garbers, C. The metalloprotease ADAM17 in inflammation and cancer. Pathol. Res. Pract. 2019, 215, e152410. [Google Scholar] [CrossRef]
- Sharifnia, T.; Rusu, V.; Piccioni, F.; Bagul, M.; Imielinski, M.; Cherniack, A.D.; Pedamallu, C.S.; Wong, B.; Wilson, F.H.; Garraway, L.A.; et al. Genetic modifiers of EGFR dependence in non-small cell lung cancer. Proc. Natl. Acad. Sci. USA 2014, 111, 18661–18666. [Google Scholar] [CrossRef] [PubMed]
- Zacharatos, P.; Kotsinas, A.; Tsantoulis, P.; Evangelou, K.; Kletsas, D.; Asimacopoulos, P.J.; Doussis-Anagnostopoulou, I.; Pezzella, F.; Gatter, K.; Papavassiliou, A.G.; et al. Relationship of the K-ras/c-mos expression patterns with angiogenesis in non-small cell lung carcinomas. Mol. Med. 2001, 7, 590–597. [Google Scholar] [CrossRef] [PubMed]
- Higgy, N.A.; Tarnasky, H.A.; Valarche, I.; Nepveu, A.; van der Hoorn, F.A. Cux/CDP homeodomain protein binds to an enhancer in the rat c-mos locus and represses its activity. Biochim. Biophys Acta 1997, 1351, 313–324. [Google Scholar] [CrossRef]

Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Griesmann, H.; Mühl, S.; Riedel, J.; Theuerkorn, K.; Sipos, B.; Esposito, I.; Vanden Heuvel, G.B.; Michl, P. CUX1 Enhances Pancreatic Cancer Formation by Synergizing with KRAS and Inducing MEK/ERK-Dependent Proliferation. Cancers 2021, 13, 2462. https://doi.org/10.3390/cancers13102462
Griesmann H, Mühl S, Riedel J, Theuerkorn K, Sipos B, Esposito I, Vanden Heuvel GB, Michl P. CUX1 Enhances Pancreatic Cancer Formation by Synergizing with KRAS and Inducing MEK/ERK-Dependent Proliferation. Cancers. 2021; 13(10):2462. https://doi.org/10.3390/cancers13102462
Chicago/Turabian StyleGriesmann, Heidi, Sebastian Mühl, Jan Riedel, Katharina Theuerkorn, Bence Sipos, Irene Esposito, Gregory B. Vanden Heuvel, and Patrick Michl. 2021. "CUX1 Enhances Pancreatic Cancer Formation by Synergizing with KRAS and Inducing MEK/ERK-Dependent Proliferation" Cancers 13, no. 10: 2462. https://doi.org/10.3390/cancers13102462
APA StyleGriesmann, H., Mühl, S., Riedel, J., Theuerkorn, K., Sipos, B., Esposito, I., Vanden Heuvel, G. B., & Michl, P. (2021). CUX1 Enhances Pancreatic Cancer Formation by Synergizing with KRAS and Inducing MEK/ERK-Dependent Proliferation. Cancers, 13(10), 2462. https://doi.org/10.3390/cancers13102462








