Hormonal Receptor Status Determines Prognostic Significance of FGFR2 in Invasive Breast Carcinoma
Simple Summary
Abstract
1. Introduction
2. Results
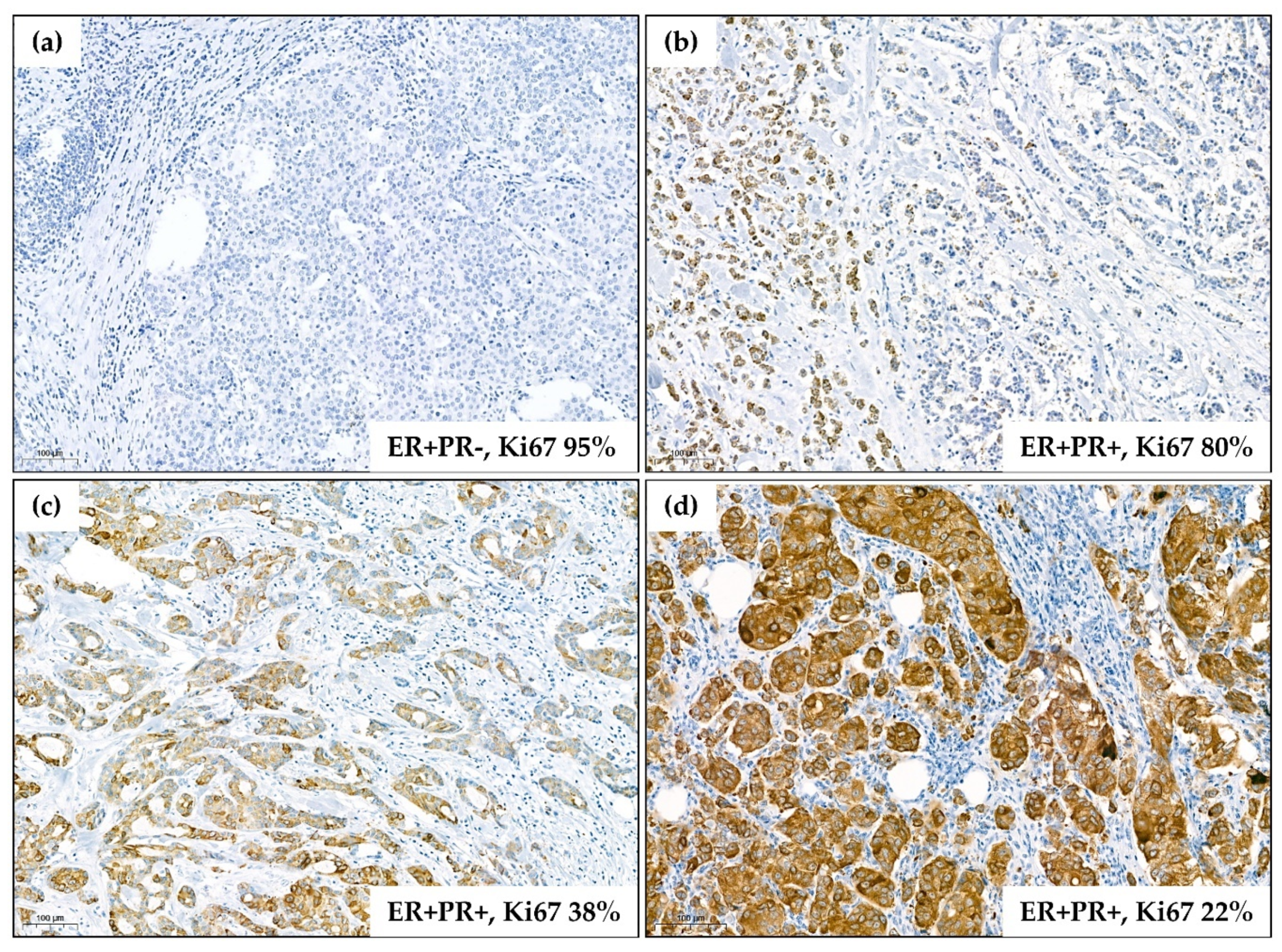
2.1. Low FGFR2 Expression Is Associated with Unfavorable Clinicopathological Characteristics Including Negative Hormonal Receptor Status
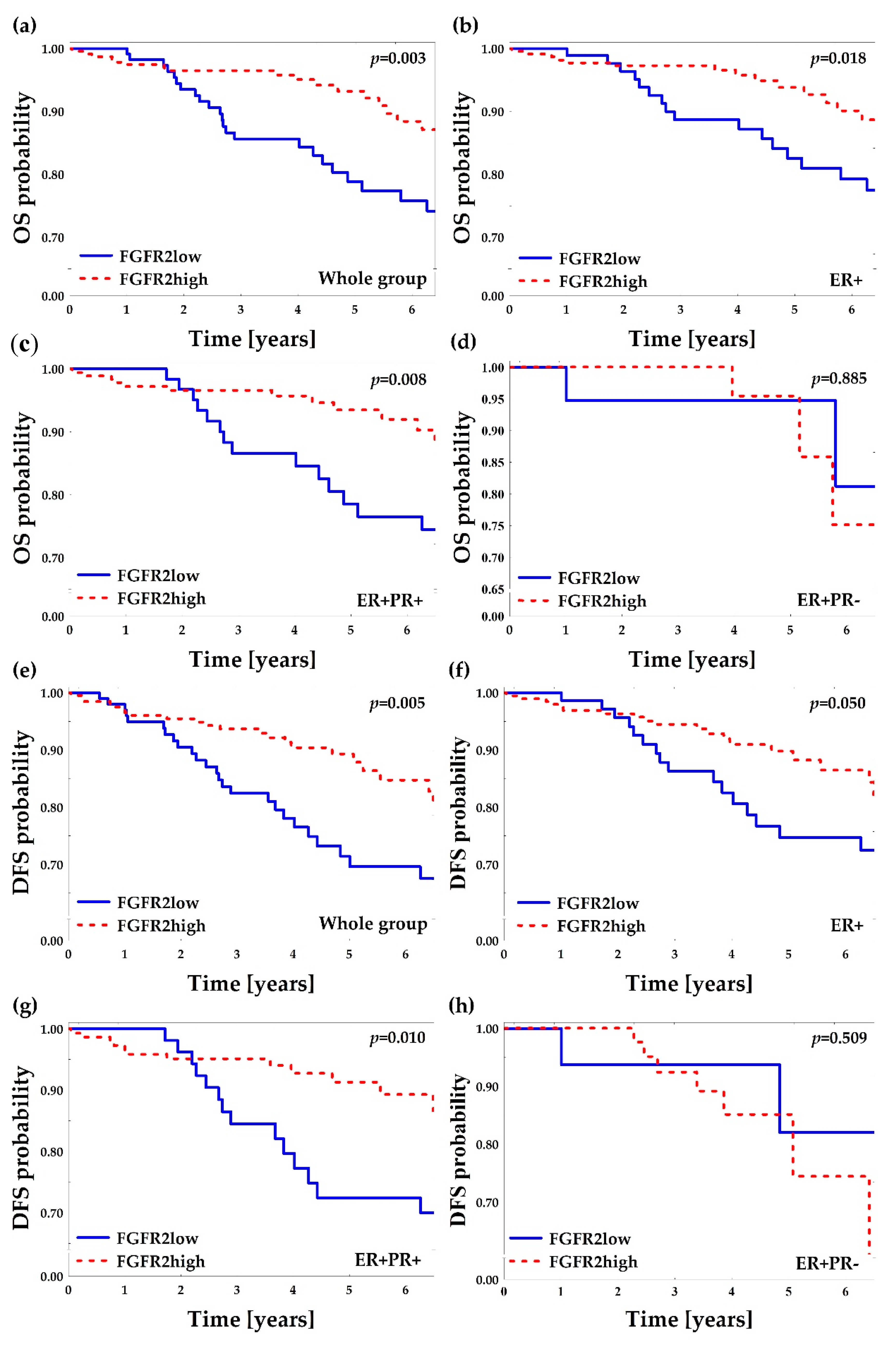
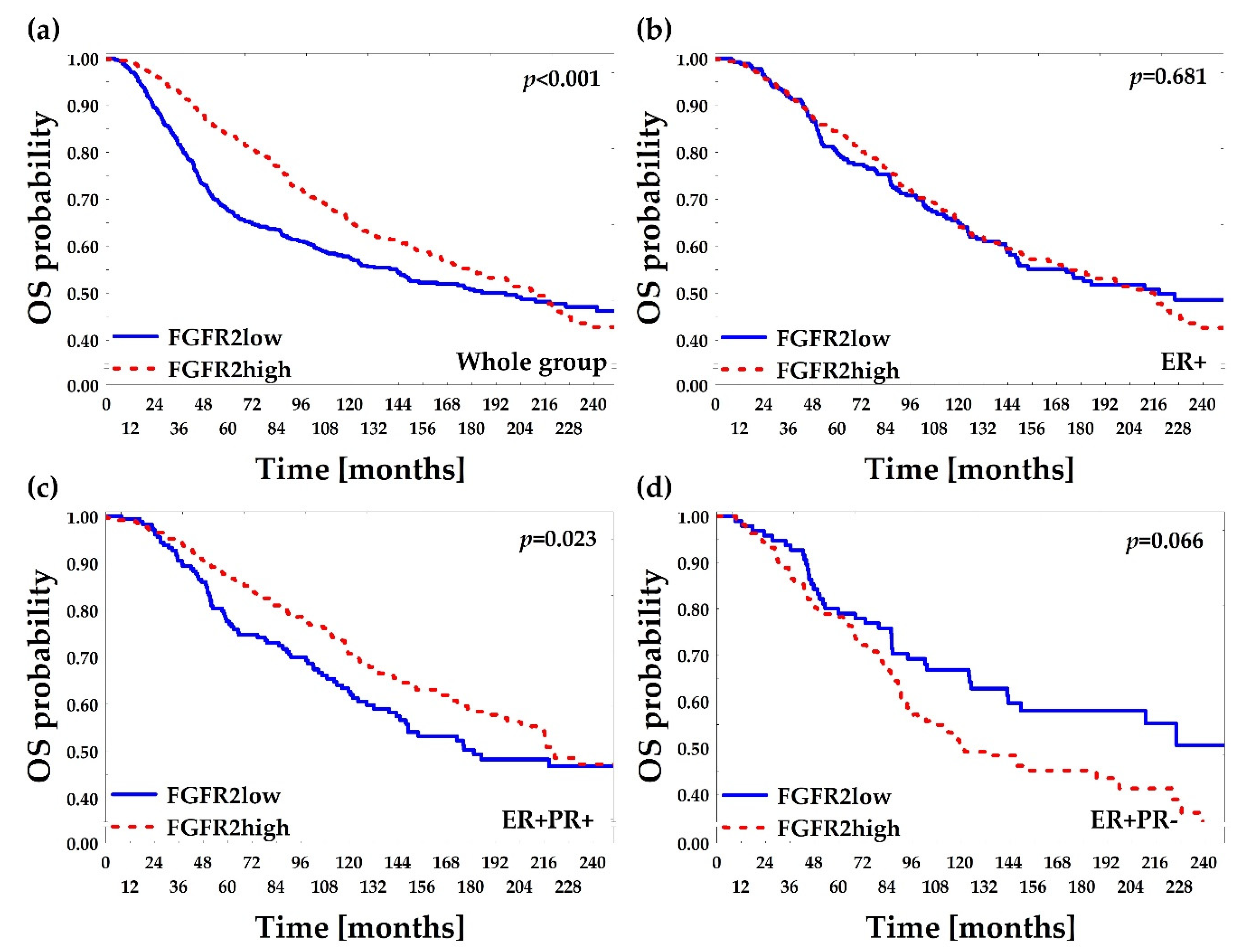
2.2. Low FGFR2 Protein Is Associated with Poor Overall and Disease-Free Survival
2.3. Subclassification of ER+ Patients by PR(mol) Supports Association of PR-Negativity with Unfavorable Clinicopathological Characteristics
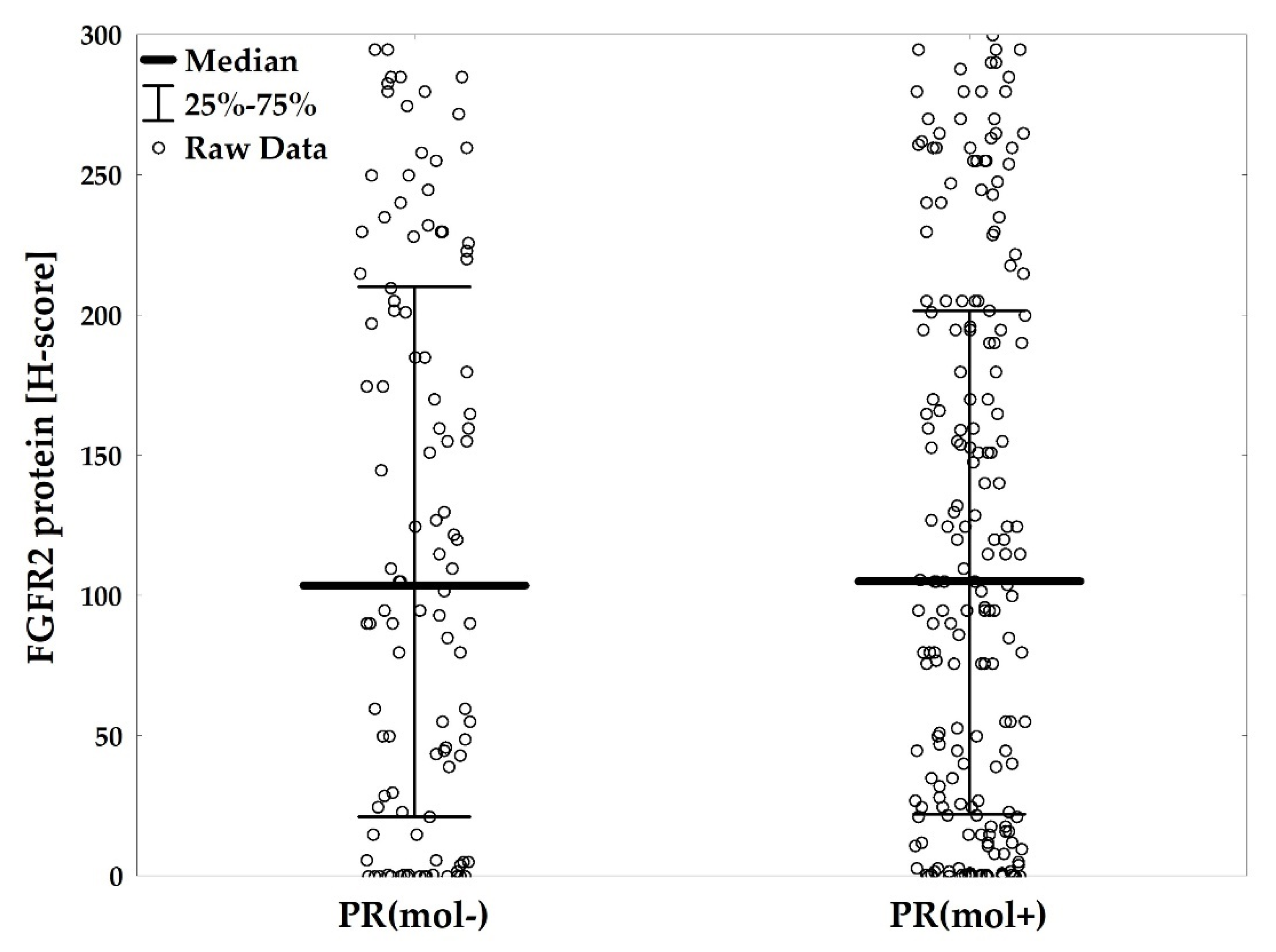
2.4. PR(mol) Status Does Not Affect Prognostic Value of FGFR2
2.5. Low FGFR2 Gene mRNA Level Is Associated with Poor Overall and Relapse-Free Survival—In Silico Confirmation
3. Discussion
4. Materials and Methods
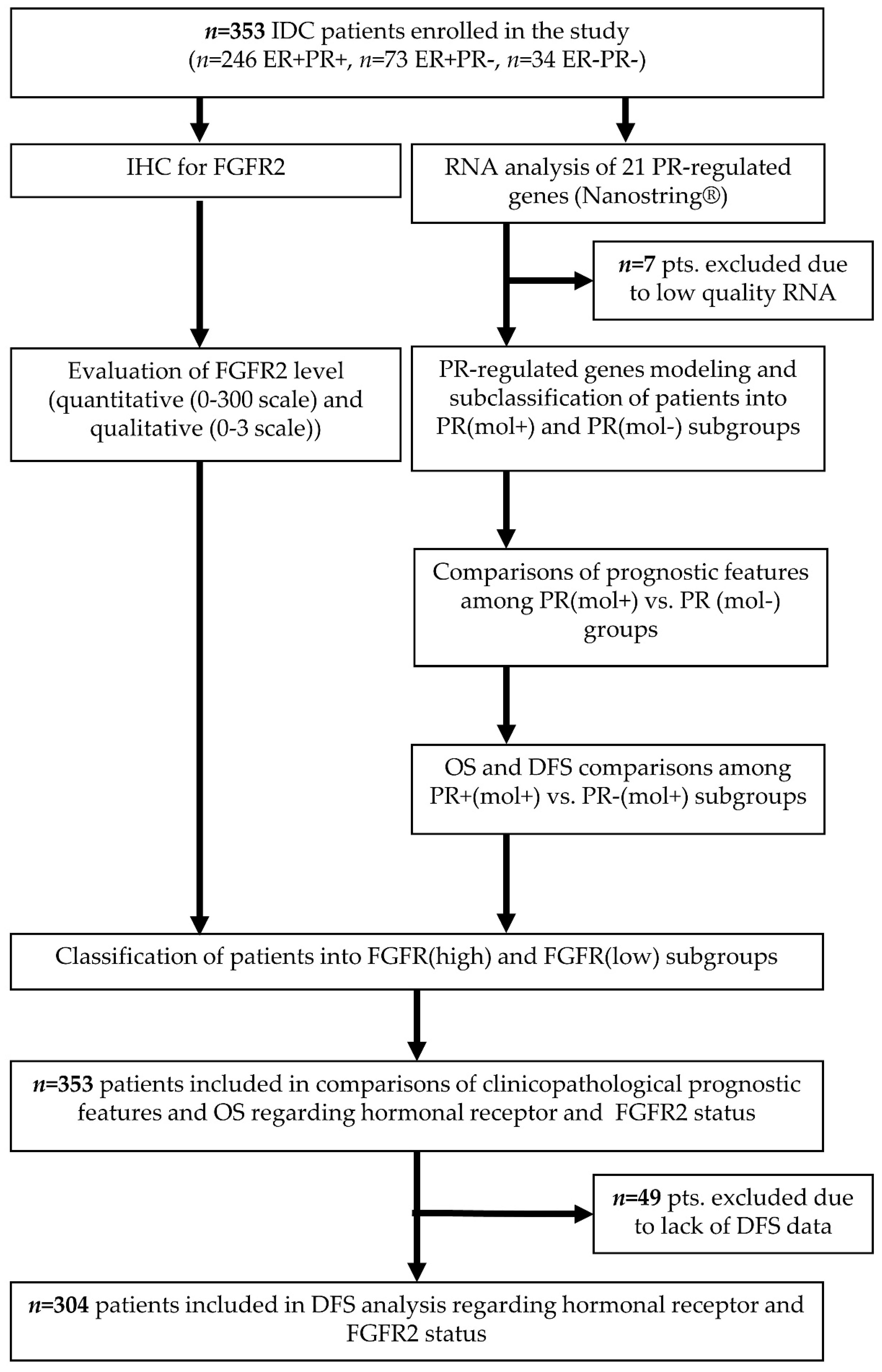
4.1. Patient Selection and Collection of Histopathological and Clinical Data
4.2. Immunohistochemistry for FGFR2
4.3. RNA Quantification
4.4. In Silico Analysis of TCGA Data
4.5. Statistical Analysis
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Place, A.E.; Jin Huh, S.; Polyak, K. The microenvironment in breast cancer progression: Biology and implications for treatment. Breast Cancer Res. 2011, 13, 227. [Google Scholar] [CrossRef] [PubMed]
- Balkwill, F.R.; Capasso, M.; Hagemann, T. The tumor microenvironment at a glance. J. Cell Sci. 2012, 125, 5591–5596. [Google Scholar] [CrossRef] [PubMed]
- Osborne, C.K.; Schiff, R. Mechanisms of Endocrine Resistance in Breast Cancer. Annu. Rev. Med. 2011, 62, 233–247. [Google Scholar] [CrossRef]
- Babina, I.S.; Turner, N.C. Advances and challenges in targeting FGFR signalling in cancer. Nat. Rev. Cancer 2017, 17, 318–332. [Google Scholar] [CrossRef]
- Piasecka, D.; Braun, M.; Kitowska, K.; Mieczkowski, K.; Kordek, R.; Sadej, R.; Romanska, H. FGFs/FGFRs-dependent signalling in regulation of steroid hormone receptors–implications for therapy of luminal breast cancer. J. Exp. Clin. Cancer Res. 2019, 38, 230. [Google Scholar] [CrossRef]
- Campbell, T.M.; Castro, M.A.A.; de Santiago, I.; Fletcher, M.N.C.; Halim, S.; Prathalingam, R.; Ponder, B.A.J.; Meyer, K.B. FGFR2 risk SNPs confer breast cancer risk by augmenting oestrogen responsiveness. Carcinogenesis 2016, 37, 741–750. [Google Scholar] [CrossRef]
- Campbell, T.M.; Castro, M.A.A.; De Oliveira, K.G.; Ponder, B.A.J.; Meyer, K.B. ERα Binding by Transcription Factors NFIB and YBX1 Enables FGFR2 Signaling to Modulate Estrogen Responsiveness in Breast Cancer. Cancer Res. 2018, 78, 410–421. [Google Scholar] [CrossRef]
- Giulianelli, S.; Cerliani, J.P.; Lamb, C.A.; Fabris, V.T.; Bottino, M.C.; Gorostiaga, M.A.; Novaro, V.; Góngora, A.; Baldi, A.; Molinolo, A.; et al. Carcinoma-associated fibroblasts activate progesterone receptors and induce hormone independent mammary tumor growth: A role for the FGF-2/FGFR-2 axis. Int. J. Cancer 2008, 123, 2518–2531. [Google Scholar] [CrossRef]
- Piasecka, D.; Kitowska, K.; Czaplinska, D.; Mieczkowski, K.; Mieszkowska, M.; Turczyk, L.; Skladanowski, A.C.; Zaczek, A.J.; Biernat, W.; Kordek, R.; et al. Fibroblast growth factor signalling induces loss of progesterone receptor in breast cancer cells. Oncotarget 2016, 7, 86011–86025. [Google Scholar] [CrossRef]
- Cerliani, J.P.; Guillardoy, T.; Giulianelli, S.; Vaque, J.P.; Gutkind, J.S.; Vanzulli, S.I.; Martins, R.; Zeitlin, E.; Lamb, C.A.; Lanari, C. Interaction between FGFR-2, STAT5, and progesterone receptors in breast cancer. Cancer Res. 2011, 71, 3720–3731. [Google Scholar] [CrossRef]
- Musgrove, E.A.; Sutherland, R.L. Biological determinants of endocrine resistance in breast cancer. Nat. Rev. Cancer 2009, 9, 631–643. [Google Scholar] [CrossRef] [PubMed]
- Daniel, A.R.; Knutson, T.P.; Lange, C.A. Signaling inputs to progesterone receptor gene regulation and promoter selectivity. Mol. Cell Endocrinol. 2009, 308, 47–52. [Google Scholar] [CrossRef] [PubMed]
- McLeskey, S.W.; Zhang, L.; El-Ashry, D.; Trock, B.J.; Lopez, C.A.; Kharbanda, S.; Tobias, C.A.; Lorant, L.A.; Hannum, R.S.; Dickson, R.B.; et al. Tamoxifen-resistant fibroblast growth factor-transfected MCF-7 cells are cross-resistant in vivo to the antiestrogen ICI 182,780 and two aromatase inhibitors. Clin. Cancer Res. 1998, 4, 697–711. [Google Scholar] [PubMed]
- Lupien, M.; Meyer, C.A.; Bailey, S.T.; Eeckhoute, J.; Cook, J.; Westerling, T.; Zhang, X.; Carroll, J.S.; Rhodes, D.R.; Liu, X.S.; et al. Growth factor stimulation induces a distinct ER(alpha) cistrome underlying breast cancer endocrine resistance. Genes Dev. 2010, 24, 2219–2227. [Google Scholar] [CrossRef] [PubMed]
- Turczyk, L.; Kitowska, K.; Mieszkowska, M.; Mieczkowski, K.; Czaplinska, D.; Piasecka, D.; Kordek, R.; Skladanowski, A.C.; Potemski, P.; Romanska, H.M.; et al. FGFR2-Driven Signaling Counteracts Tamoxifen Effect on ERalpha-Positive Breast Cancer Cells. Neoplasia 2017, 19, 791–804. [Google Scholar] [CrossRef]
- Daniel, A.R.; Faivre, E.J.; Lange, C.A. Phosphorylation-dependent antagonism of sumoylation derepresses progesterone receptor action in breast cancer cells. Mol. Endocrinol. 2007, 21, 2890–2906. [Google Scholar] [CrossRef]
- Knutson, T.P.; Daniel, A.R.; Fan, D.; Silverstein, K.A.; Covington, K.R.; Fuqua, S.A.; Lange, C.A. Phosphorylated and sumoylation-deficient progesterone receptors drive proliferative gene signatures during breast cancer progression. Breast Cancer Res. 2012, 14, 1–23. [Google Scholar] [CrossRef]
- Drago, J.Z.; Formisano, L.; Juric, D.; Niemierko, A.; Servetto, A.; Wander, S.A.; Spring, L.M.; Vidula, N.; Younger, J.; Peppercorn, J.; et al. FGFR1 gene amplification mediates endocrine resistance but retains TORC sensitivity in metastatic hormone receptor positive (HR+) breast cancer. Clin. Cancer Res. 2019, 25, 6443–6451. [Google Scholar] [CrossRef]
- Formisano, L.; Lu, Y.; Servetto, A.; Hanker, A.B.; Jansen, V.M.; Bauer, J.A.; Sudhan, D.R.; Guerrero-Zotano, A.L.; Croessmann, S.; Guo, Y.; et al. Aberrant FGFR signaling mediates resistance to CDK4/6 inhibitors in ER+ breast cancer. Nat. Commun. 2019, 10, 1–14. [Google Scholar] [CrossRef]
- Giltnane, J.M.; Hutchinson, K.E.; Stricker, T.P.; Formisano, L.; Young, C.D.; Estrada, M.V.; Nixon, M.J.; Du, L.; Sanchez, V.; Ericsson, P.G.; et al. Genomic profiling of ER+ breast cancers after short-term estrogen suppression reveals alterations associated with endocrine resistance. Sci. Transl. Med. 2017, 9, eaai7993. [Google Scholar] [CrossRef]
- Mao, P.; Cohen, O.; Kowalski, K.J.; Kusiel, J.G.; Buendia-Buendia, J.E.; Cuoco, M.S.; Exman, P.; Wander, S.A.; Waks, A.G.; Nayar, U.; et al. Acquired FGFR and FGF alterations confer resistance to estrogen receptor (ER) targeted therapy in ER+ metastatic breast cancer. bioRxiv 2019. [Google Scholar] [CrossRef]
- Lee, H.J.; Seo, A.N.; Park, S.Y.; Kim, J.Y.; Park, J.Y.; Yu, J.H.; Ahn, J.-H.; Gong, G. Low Prognostic Implication of Fibroblast Growth Factor Family Activation in Triple-negative Breast Cancer Subsets. Ann. Surg. Oncol. 2014, 21, 1561–1568. [Google Scholar] [CrossRef] [PubMed]
- Liu, G.; Xiong, D.; Xiao, R.; Huang, Z. Prognostic role of fibroblast growth factor receptor 2 in human solid tumors: A systematic review and meta-analysis. Tumor Biol. 2017, 39. [Google Scholar] [CrossRef] [PubMed]
- Czaplinska, D.; Mieczkowski, K.; Supernat, A.; Skladanowski, A.C.; Kordek, R.; Biernat, W.; Zaczek, A.J.; Romanska, H.M.; Sadej, R. Interactions between FGFR2 and RSK2—implications for breast cancer prognosis. Tumor Biol. 2016, 37, 13721–13731. [Google Scholar] [CrossRef] [PubMed]
- Sadej, R.; Lu, X.; Turczyk, L.; Novitskaya, V.; Lopez-Clavijo, A.F.; Kordek, R.; Potemski, P.; Wakelam, M.J.O.; Romanska-Knight, H.; Berditchevski, F. CD151 regulates expression of FGFR2 in breast cancer cells via PKC-dependent pathways. J. Cell Sci. 2018, 131. [Google Scholar] [CrossRef] [PubMed]
- Sun, S.; Jiang, Y.; Zhang, G.; Song, H.; Zhang, X.; Zhang, Y.; Liang, X.; Sun, Q.; Pang, D. Increased expression of fibroblastic growth factor receptor 2 is correlated with poor prognosis in patients with breast cancer. J. Surg. Oncol. 2012, 105, 773–779. [Google Scholar] [CrossRef]
- Martin, A.J.; Grant, A.; Ashfield, A.M.; Palmer, C.N.; Baker, L.; Quinlan, P.R.; Purdie, C.A.; Thompson, A.M.; Jordan, L.B.; Berg, J.N. FGFR2 protein expression in breast cancer: Nuclear localisation and correlation with patient genotype. BMC Res. Notes 2011, 4, 72. [Google Scholar] [CrossRef]
- Knutson, T.P.; Truong, T.H.; Ma, S.; Brady, N.J.; Sullivan, M.E.; Raj, G.; Schwertfeger, K.L.; Lange, C.A. Posttranslationally modified progesterone receptors direct ligand-specific expression of breast cancer stem cell-associated gene programs. J. Hematol. Oncol. 2017, 10, 89. [Google Scholar] [CrossRef]
- Richer, J.K.; Jacobsen, B.M.; Manning, N.G.; Abel, M.G.; Wolf, D.M.; Horwitz, K.B. Differential gene regulation by the two progesterone receptor isoforms in human breast cancer cells. J. Biol. Chem. 2002, 277, 5209–5218. [Google Scholar] [CrossRef]
- Pereira, B.; Chin, S.-F.; Rueda, O.M.; Vollan, H.-K.M.; Provenzano, E.; Bardwell, H.A.; Pugh, M.; Jones, L.; Russell, R.; Sammut, S.-J.; et al. The somatic mutation profiles of 2433 breast cancers refine their genomic and transcriptomic landscapes. Nat. Commun. 2016, 7, 11479. [Google Scholar] [CrossRef]
- Curtis, C.; Shah, S.P.; Chin, S.-F.; Turashvili, G.; Rueda, O.M.; Dunning, M.J.; Speed, D.; Lynch, A.G.; Samarajiwa, S.; Yuan, Y.; et al. The genomic and transcriptomic architecture of 2,000 breast tumours reveals novel subgroups. Nature 2012, 486, 346–352. [Google Scholar] [CrossRef] [PubMed]
- Györffy, B.; Lanczky, A.; Eklund, A.C.; Denkert, C.; Budczies, J.; Li, Q.; Szallasi, Z. An online survival analysis tool to rapidly assess the effect of 22,277 genes on breast cancer prognosis using microarray data of 1809 patients. Breast Cancer Res. Treat. 2010, 123, 725–731. [Google Scholar] [CrossRef] [PubMed]
- Alvarez-Fernandez, M.; Malumbres, M. Mechanisms of Sensitivity and Resistance to CDK4/6 Inhibition. Cancer Cell 2020, 37, 514–529. [Google Scholar] [CrossRef] [PubMed]
- Gartside, M.G.; Chen, H.; Ibrahimi, O.A.; Byron, S.A.; Curtis, A.V.; Wellens, C.L.; Bengston, A.; Yudt, L.M.; Eliseenkova, A.V.; Ma, J.; et al. Loss-of-Function Fibroblast Growth Factor Receptor-2 Mutations in Melanoma. Mol. Cancer Res. 2009, 7, 41–54. [Google Scholar] [CrossRef] [PubMed]
- Byron, S.A.; Gartside, M.; Powell, M.A.; Wellens, C.L.; Gao, F.; Mutch, D.G.; Goodfellow, P.J.; Pollock, P.M. FGFR2 point mutations in 466 endometrioid endometrial tumors: Relationship with MSI, KRAS, PIK3CA, CTNNB1 mutations and clinicopathological features. PLoS ONE 2012, 7, e30801. [Google Scholar] [CrossRef]
- Pollock, P.M.; Gartside, M.G.; Dejeza, L.C.; Powell, M.A.; Mallon, M.A.; Davies, H.; Mohammadi, M.; Futreal, P.A.; Stratton, M.R.; Trent, J.M.; et al. Frequent activating FGFR2 mutations in endometrial carcinomas parallel germline mutations associated with craniosynostosis and skeletal dysplasia syndromes. Oncogene 2007, 26, 7158–7162. [Google Scholar] [CrossRef]
- Grose, R.; Fantl, V.; Werner, S.; Chioni, A.-M.; Jarosz, M.; Rudling, R.; Cross, B.; Hart, I.R.; Dickson, C. The role of fibroblast growth factor receptor 2b in skin homeostasis and cancer development. EMBO J. 2007, 26, 1268–1278. [Google Scholar] [CrossRef]
- Amann, T.; Bataille, F.; Spruss, T.; Dettmer, K.; Wild, P.; Liedtke, C.; Mühlbauer, M.; Kiefer, P.; Oefner, P.J.; Trautwein, C.; et al. Reduced Expression of Fibroblast Growth Factor Receptor 2IIIb in Hepatocellular Carcinoma Induces a More Aggressive Growth. Am. J. Pathol. 2010, 176, 1433–1442. [Google Scholar] [CrossRef]
- Katoh, Y.; Katoh, M. FGFR2-related pathogenesis and FGFR2-targeted therapeutics (Review). Int. J. Mol. Med. 2009, 23. [Google Scholar] [CrossRef]
- DeSantis, C.E.; Ma, J.; Gaudet, M.M.; Newman, L.A.; Miller, K.D.; Goding Sauer, A.; Jemal, A.; Siegel, R.L. Breast cancer statistics, 2019. CA Cancer J. Clin. 2019, 69, 438–451. [Google Scholar] [CrossRef]
- Hoon Tan, P.; Ellis, I.; Allison, K.; Brogi, E.; Fox, S.B.; Lakhani, S.; Lazar, A.J.; Morris, E.A.; Sahin, A.; Salgado, R.; et al. The 2019 WHO classification of tumours of the breast. Histopathology 2020. [Google Scholar] [CrossRef] [PubMed]
- Allred, D.C.; Harvey, J.M.; Berardo, M.; Clark, G.M. Prognostic and predictive factors in breast cancer by immunohistochemical analysis. Mod. Pathol. 1998, 11, 155–168. [Google Scholar] [PubMed]
- Wolff, A.C.; Hammond, M.E.H.; Allison, K.H.; Harvey, B.E.; Mangu, P.B.; Bartlett, J.M.S.; Bilous, M.; Ellis, I.O.; Fitzgibbons, P.; Hanna, W.; et al. Human Epidermal Growth Factor Receptor 2 Testing in Breast Cancer: American Society of Clinical Oncology/College of American Pathologists Clinical Practice Guideline Focused Update. Arch. Pathol. Lab. Med. 2018, 142, 1364–1382. [Google Scholar] [CrossRef]
- Lal, P.; Salazar, P.A.; Hudis, C.A.; Ladanyi, M.; Chen, B. HER−2 testing in breast cancer using immunohistochemical analysis and fluorescence in situ hybridization: A single-institution experience of 2,279 cases and comparison of dual-color and single-color scoring. Am. J. Clin. Pathol. 2004, 121, 631–636. [Google Scholar] [CrossRef] [PubMed]
- Borchert, S.; Wessolly, M.; Schmeller, J.; Mairinger, E.; Kollmeier, J.; Hager, T.; Mairinger, T.; Herold, T.; Christoph, D.C.; Walter, R.F.H.; et al. Gene expression profiling of homologous recombination repair pathway indicates susceptibility for olaparib treatment in malignant pleural mesothelioma in vitro. BMC Cancer 2019, 19, 108. [Google Scholar] [CrossRef] [PubMed]
- Wu, C.; Mairinger, F.; Casanova, R.; Batavia, A.A.; Leblond, A.L.; Soltermann, A. Prognostic Immune Cell Profiling of Malignant Pleural Effusion Patients by Computerized Immunohistochemical and Transcriptional Analysis. Cancers 2019, 11, 1953. [Google Scholar] [CrossRef] [PubMed]
- Hudis, C.A.; Barlow, W.E.; Costantino, J.P.; Gray, R.J.; Pritchard, K.I.; Chapman, J.-A.W.; Sparano, J.A.; Hunsberger, S.; Enos, R.A.; Gelber, R.D.; et al. Proposal for Standardized Definitions for Efficacy End Points in Adjuvant Breast Cancer Trials: The STEEP System. J. Clin. Oncol. 2007, 25, 2127–2132. [Google Scholar] [CrossRef] [PubMed]

| Variable | Whole Group n = 353 (100) | ER−PR− n = 34 (9.6) | ER+PR− n = 73 (20.7) | ER+PR+ n = 246 (69.7) | p-Value |
|---|---|---|---|---|---|
| Age (years) 1 | 63.8 | 63.6 | 64.7 | 63.6 | 0.132 |
| (55.0–71.3) | (55.7–73.1) | (60.0–70.6) | (52.7–70.8) | ||
| Menopausal status 2 | 0.063 | ||||
| Pre | 37 (11.4) | 2 (6.1) | 3 (4.6) | 32 (14.1) | |
| Post | 288 (88.6) | 31 (93.9) | 62 (95.4) | 195 (85.9) | |
| Grade 2 | <0.001 * | ||||
| 1 | 42 (11.9) | 2 (5.9) | 9 (12.3) | 31 (12.6) | |
| 2 | 227 (64.3) | 12 (35.3) | 43 (58.9) | 168 (69.9) | |
| 3 | 84 (23.8) | 20 (58.8) | 21 (28.8) | 42 (17.5) | |
| Ki67 (%) 1 | 18.0 | 32.0 | 20.0 | 12.0 | 0.011 * |
| (5.0–30.0) | (30.5–35.5) | (10.0–40.0) | (5.0–25.0) | ||
| HER2 amplification positivity 2 | 45 (12.7) | 12 (35.3) | 13 (17.8) | 20.0 (8.1) | <0.001 * |
| Tumor size (mm) 1 | 20.0 | 25.0 | 25.0 | 20.0 | 0.004 * |
| (15.0–27.0) | (15.0–35.0) | (19.0–30.0) | (15.0–25.0) | ||
| T feature 2 | 0.064 | ||||
| pT1 | 143 (53.0) | 13 (40.6) | 15 (38.5) | 115 (57.8) | |
| pT2 | 116 (43.0) | 16 (50.0) | 22 (56.4) | 78 (39.2) | |
| pT3-4 | 11 (4.0) | 3 (9.4) | 2 (5.1) | 6 (3.0) | |
| Metastases present 2 | 116 (33.5) | 13 (42.4) | 25 (34.7) | 77 (32.0) | 0.475 |
| N feature 2 | 0.733 | ||||
| pN0 | 230 (66.5) | 17 (57.6) | 47 (65.3) | 164 (68.0) | |
| pN1 | 81 (23.4) | 10 (30.3) | 16 (22.2) | 55 (22.8) | |
| pN2-3 | 34 (10.1) | 4 (12.1) | 9 (12.5) | 22 (9.1) | |
| Staging 2 | 0.289 | ||||
| Very early (IA) | 135 (39.1) | 11 (34.4) | 23 (31.9) | 101 (41.9) | |
| Early (IB-IIIA) | 189 (54.8) | 17 (53.1) | 45 (62.5) | 127 (52.7) | |
| Advanced (IIIB-IV) | 21 (6.1) | 4 (12.5) | 4 (5.6) | 13 (5.4) | |
| Multifocality 2 | 39 (14.3) | 3 (8.8) | 6 (15.4) | 30 (15.0) | 0.622 |
| DCIS present 2 | 103 (29.1) | 10 (29.4) | 23 (31.5) | 70 (28.3) | 0.871 |
| Hormonotherapy 2 | 220 (81.2) | 0 (0.0) | 46 (82.1) | 174 (86.6) | <0.001 * |
| Neoadjuvant therapy 2 | 231 (75.5) | 26 (89.7) | 30 (48.4) | 175 (81.4) | <0.001 * |
| Adjuvant chemotherapy 2 | 150 (62.2) | 25 (80.7) | 38 (66.7) | 87 (56.9) | 0.032 * |
| Adjuvant radiotherapy 2 | 151 (64.8) | 11 (50.0) | 30 (61.2) | 110 (67.9) | 0.111 |
| Progression/relapse 3 | 30 (9.8) | 10 (31.3) | 8 (11.9) | 12 (5.8) | <0.001 * |
| Disease-free survival (years) | 3.9 (2.7–6.4) | 3.6 (2.2–4.5) | 3.8 (2.5–4.9) | 4.2 (2.9–6.6) | |
| Deaths 3 | 41 (11.6) | 10 (29.4) | 6 (8.2) | 25 (10.1) | <0.001 * |
| Overall survival (years) | 4.2 (2.9–6.5) | 3.6 (2.6–4.5) | 4.0 (2.7–5.5) | 4.7 (3.0–6.7) |
| Variable | FGFR2low n = 117 (33.1) | FGFR2high n = 236 (66.9) | p-Value |
|---|---|---|---|
| Age (years) 1 | 64.6 (56.5–75.2) | 63.4 (54.2–69.9) | 0.078 |
| Menopausal status 2 | 0.078 | ||
| Pre | 8 (7.1) | 29 (13.7) | |
| Post | 104 (92.9) | 183 (86.3) | |
| Grade 2 | <0.001 * | ||
| 1 | 7 (5.9) | 35 (14.8) | |
| 2 | 65 (55.6) | 162 (68.6) | |
| 3 | 45 (38.5) | 39 (16.5) | |
| Ki67 (%) 1 | 23.5 (12.0–40.0) | 15.0 (5.0–28.0) | 0.014 * |
| HER2 amplification positivity 2 | 11 (9.4) | 33 (14.0) | 0.220 |
| Hormonal status 2 | <0.001 * | ||
| ER−PR− | 25 (21.4) | 9 (3.8) | |
| ER+PR− | 22 (18.8) | 51 (21.6) | |
| ER+PR+ | 70 (59.8) | 176 (74.6) | |
| PR molecular status 2 | 0.757 | ||
| PR(mol+) | 59 (66.3) | 145 (64.4) | |
| PR(mol-) | 30 (33.7) | 80 (35.6) | |
| Tumor size (mm) 1 | 20.0 (15.0–30.0) | 20.0 (15.0–25.0) | 0.898 |
| T feature 2 | 0.731 | ||
| pT1 | 55 (51.4) | 88 (54.0) | |
| pT2 | 46 (43.0 | 69 (42.3) | |
| pT3-4 | 6 (5.6) | 6 (3.7) | |
| Metastases present 2 | 44 (38.6) | 76 (32.9) | 0.296 |
| N feature 2 | 0.682 | ||
| pN0 | 73 (64.0) | 156 (67.5) | |
| pN1 | 30 (26.3) | 51 (22.1) | |
| pN2-3 | 11 (9.6) | 24 (10.4) | |
| Staging 2 | 0.234 | ||
| Very early (IA) | 40 (35.1) | 95 (41.1) | |
| Early (IB-IIIA) | 69 (60.5) | 119 (51.5) | |
| Advanced (IIIB-IV) | 5 (4.4) | 17 (7.4) | |
| DCIS present 2 | 30 (25.6) | 72 (30.5) | 0.342 |
| DFS events 3 | 25 (24.8) | 22 (10.8) | 0.005 * |
| Disease-free survival (years) | 3.9 (2.6–6.6) | 3.9 (2.7–5.8) | |
| Deaths 3 | 23 (19.7) | 18 (7.6) | 0.003 * |
| Overall survival (years) | 4.6 (2.7–6.7) | 4.1 (2.9–6.4) |
| Outcome | HR (95% CI) Raw | HR (95% CI) Adjusted for Hormonal Status | HR (95% CI) Adjusted for Hormonal Status, Age and Tumor Size |
|---|---|---|---|
| OS (FGFR2high as reference) | 2.34 (1.26–4.34), p = 0.007 | 2.09 (1.08–4.04), p = 0.028 | 1.45 (0.73–2.90), p = 0.283 |
| DFS (FGFR2high as reference) | 2.22 (1.25–3.93), p = 0.006 | 1.92 (1.03–3.56), p = 0.038 | 1.25 (0.66–2.37), p = 0.496 |
| Hormonal Status | PR(mol+) | PR(mol−) |
|---|---|---|
| ER−PR− | 0 (0.0%) | 32 (100.0%) |
| ER+PR− | 10 (13.7%) | 63 (86.3%) |
| ER+PR+ | 194 (80.6%) | 47 (19.4%) |
| All | 204 (59.1%) | 142 (40.9%) |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Braun, M.; Piasecka, D.; Tomasik, B.; Mieczkowski, K.; Stawiski, K.; Zielinska, A.; Kopczynski, J.; Nejc, D.; Kordek, R.; Sadej, R.; et al. Hormonal Receptor Status Determines Prognostic Significance of FGFR2 in Invasive Breast Carcinoma. Cancers 2020, 12, 2713. https://doi.org/10.3390/cancers12092713
Braun M, Piasecka D, Tomasik B, Mieczkowski K, Stawiski K, Zielinska A, Kopczynski J, Nejc D, Kordek R, Sadej R, et al. Hormonal Receptor Status Determines Prognostic Significance of FGFR2 in Invasive Breast Carcinoma. Cancers. 2020; 12(9):2713. https://doi.org/10.3390/cancers12092713
Chicago/Turabian StyleBraun, Marcin, Dominika Piasecka, Bartlomiej Tomasik, Kamil Mieczkowski, Konrad Stawiski, Aleksandra Zielinska, Janusz Kopczynski, Dariusz Nejc, Radzislaw Kordek, Rafal Sadej, and et al. 2020. "Hormonal Receptor Status Determines Prognostic Significance of FGFR2 in Invasive Breast Carcinoma" Cancers 12, no. 9: 2713. https://doi.org/10.3390/cancers12092713
APA StyleBraun, M., Piasecka, D., Tomasik, B., Mieczkowski, K., Stawiski, K., Zielinska, A., Kopczynski, J., Nejc, D., Kordek, R., Sadej, R., & Romanska, H. M. (2020). Hormonal Receptor Status Determines Prognostic Significance of FGFR2 in Invasive Breast Carcinoma. Cancers, 12(9), 2713. https://doi.org/10.3390/cancers12092713








