Comparative Study of Organoids from Patient-Derived Normal and Tumor Colon and Rectal Tissue
Abstract
1. Introduction
2. Results
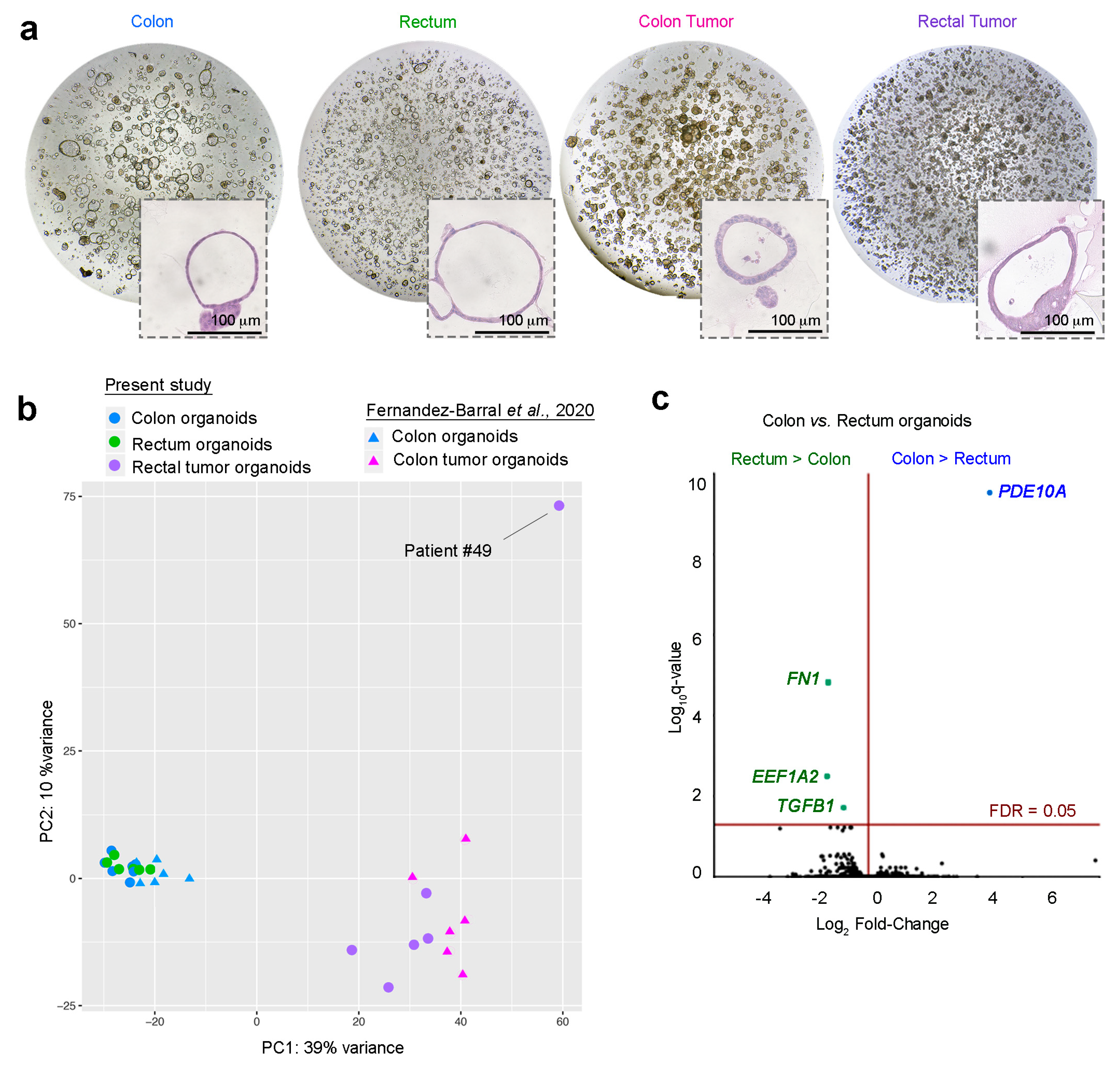
2.1. Establishment and Morphology of 3D Organoids From Endoscopic Biopsies from Rectal Cancer Patients
2.2. Gene Profiling of Normal Colon and Normal Rectum Organoids
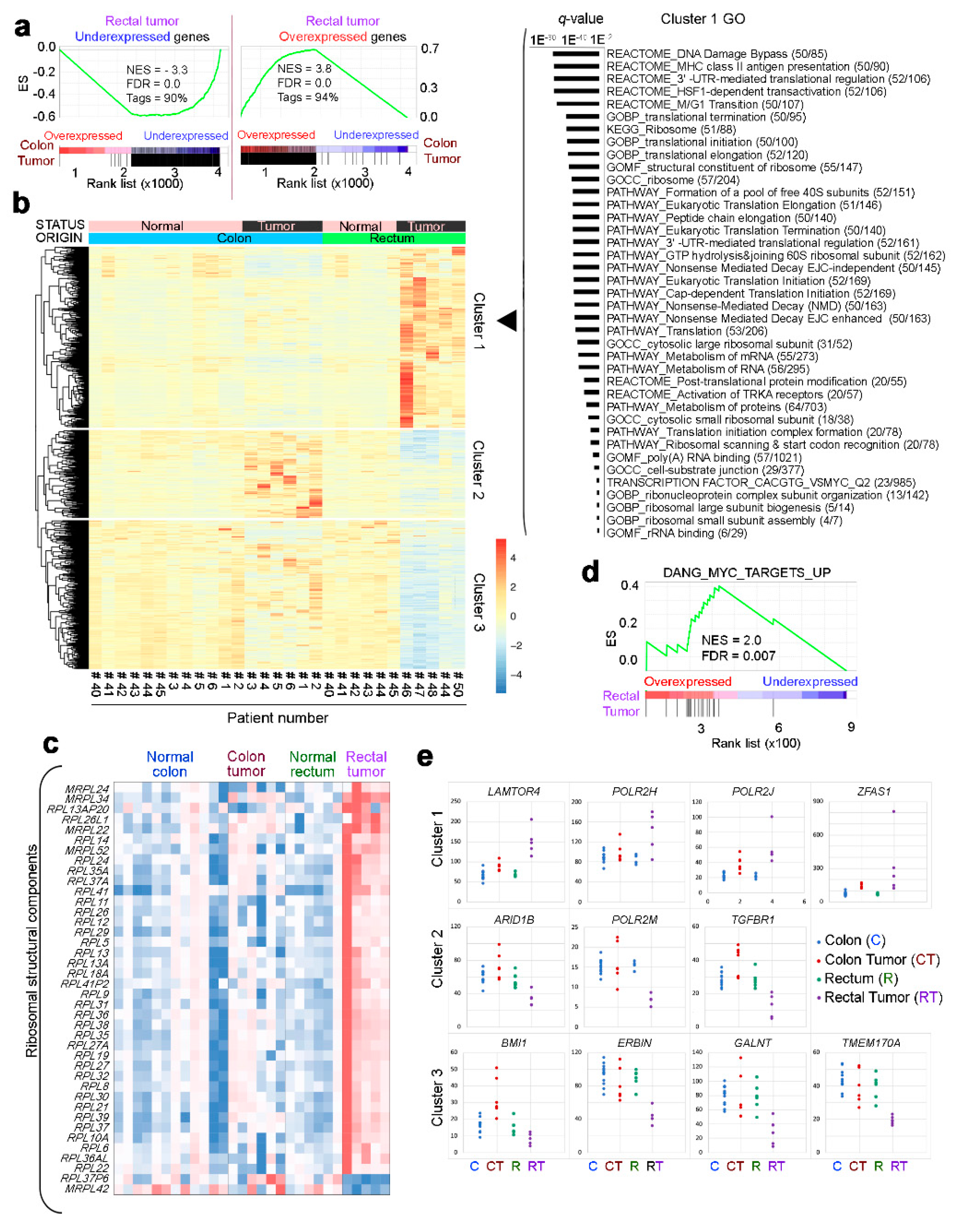
2.3. Gene Profiling of Normal Rectum and Rectal Tumor Organoids
2.4. Transcriptomic Profiles of Rectal Tumor and Colon Tumor Organoids
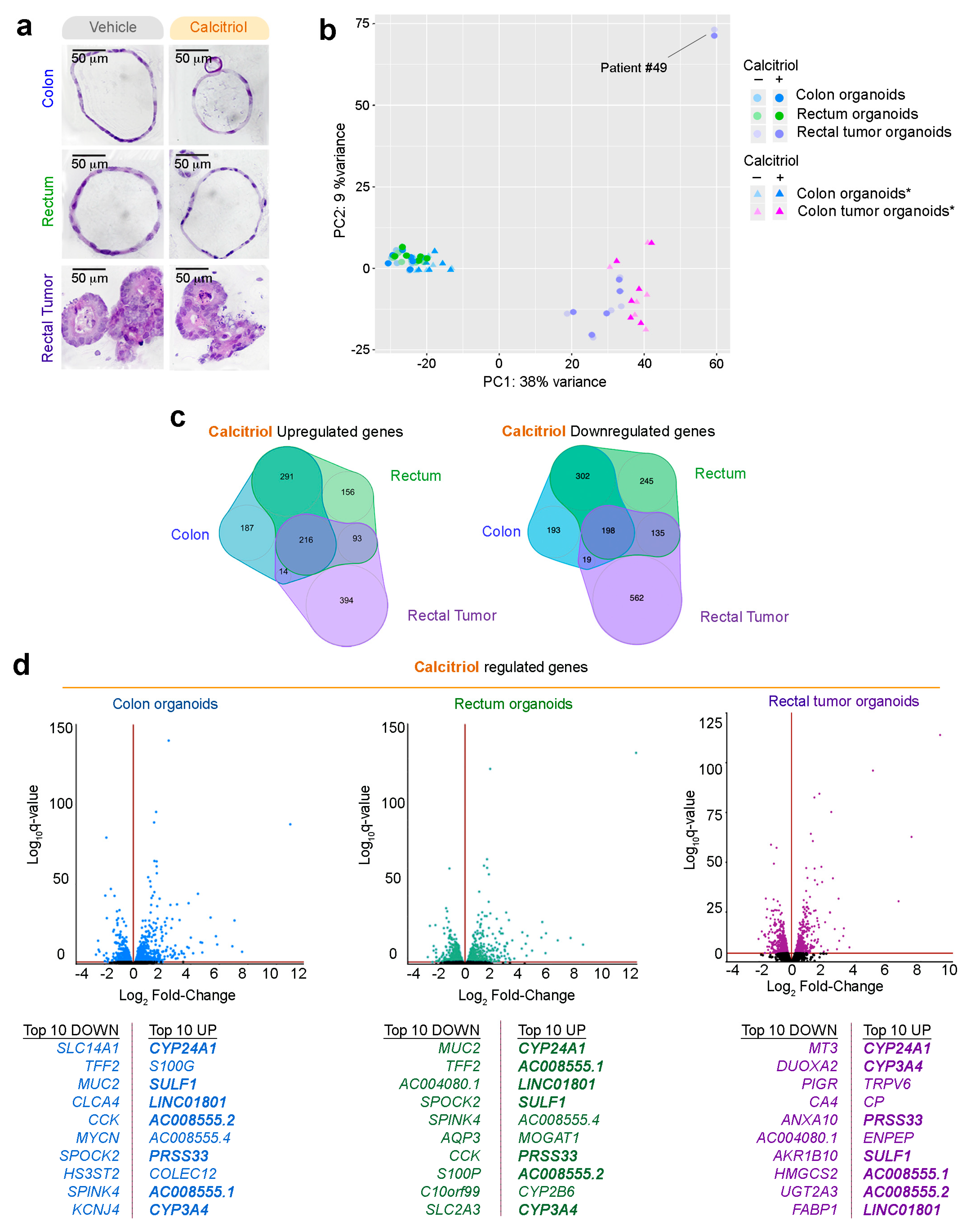
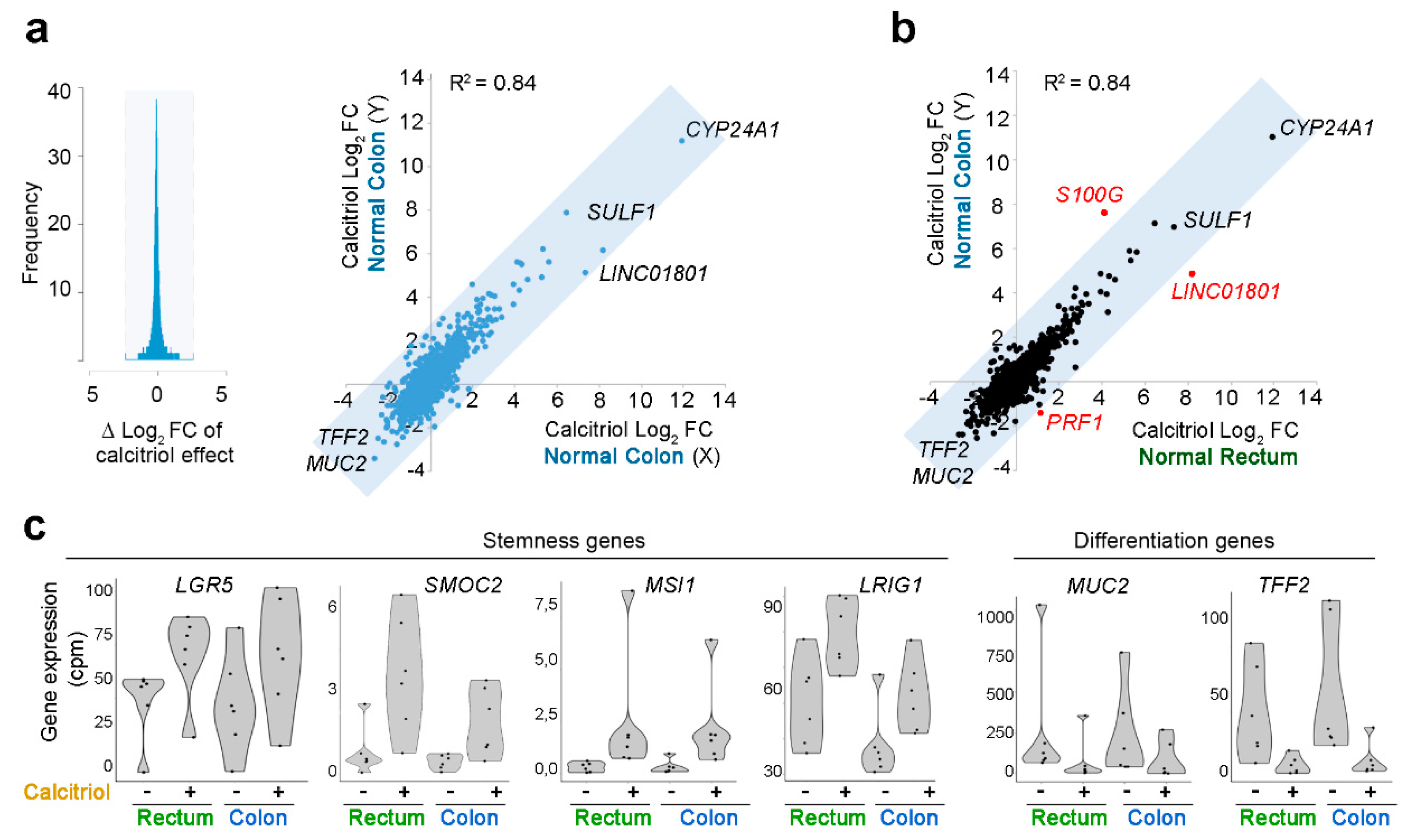
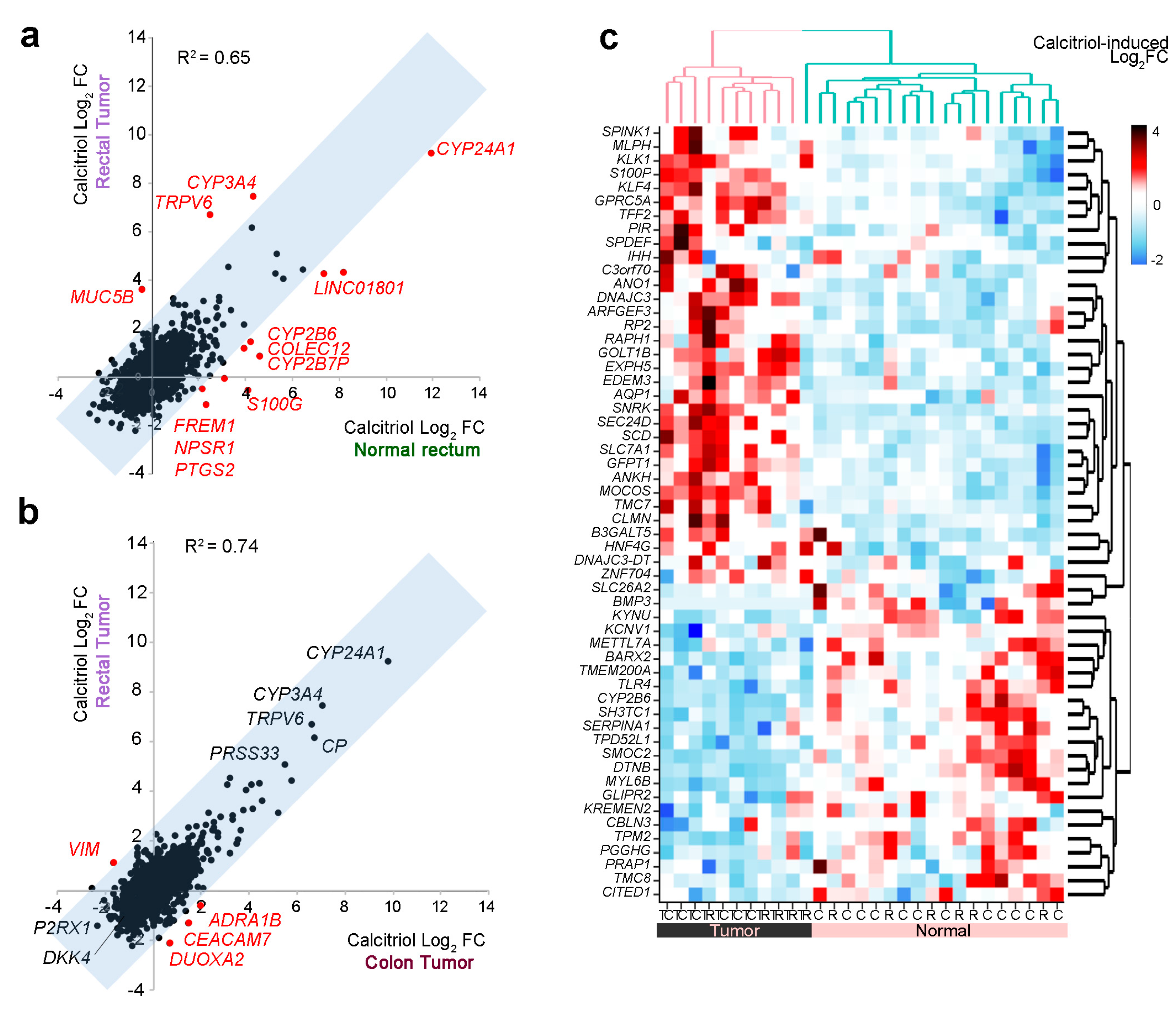
2.5. Effect of Calcitriol on Organoids from Normal and Tumor Tissues
3. Discussion
4. Materials and Methods
4.1. Human Samples and Ethical Guidelines
4.2. Establishment of 3D Organoid Cultures from Endoscopy Biopsies
4.3. Growth and Expansion of Organoid Cultures
4.4. Histology and Imaging
4.5. Mutational Status
4.6. RNA-Sequencing
4.7. Coefficient of Determination and Correlation Zone
4.8. Gene Set Enrichment Analysis (GSEA)
4.9. Data Availability
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Yamauchi, M.; Lochhead, P.; Morikawa, T.; Huttenhower, C.; Chan, A.T.; Giovannucci, E.; Fuchs, C.; Ogino, S. Colorectal cancer: A tale of two sides or a continuum? Gut 2012, 61, 794–797. [Google Scholar] [CrossRef] [PubMed]
- Hjartaker, A.; Aagnes, B.; Robsahm, T.E.; Langseth, H.; Bray, F.; Larsen, I.K. Subsite-specific dietary risk factors for colorectal cancer: A review of cohort studies. J. Oncol. 2013, 2013, 703854. [Google Scholar] [CrossRef] [PubMed]
- Paschke, S.; Jafarov, S.; Staib, L.; Kreuser, E.D.; Maulbecker-Armstrong, C.; Roitman, M.; Holm, T.; Harris, C.C.; Link, K.H.; Kornmann, M. Are Colon and Rectal Cancer Two Different Tumor Entities? A Proposal to Abandon the Term Colorectal Cancer. Int. J. Mol. Sci. 2018, 19, 2577. [Google Scholar] [CrossRef] [PubMed]
- Murphy, C.C.; Wallace, K.; Sandler, R.S.; Baron, J.A. Racial Disparities in Incidence of Young-Onset Colorectal Cancer and Patient Survival. Gastroenterology 2019, 156, 958–965. [Google Scholar] [CrossRef] [PubMed]
- Rezaianzadeh, A.; Rahimikazerooni, S.; Khazraei, H.; Tadayon, S.M.K.; Akool, M.A.; Rahimi, M.; Hosseini, S.V. Do clinicopathologic features of rectal and colon cancer guide us towards distinct malignancies? J. Gastrointest. Oncol. 2019, 10, 203–208. [Google Scholar] [CrossRef]
- Kornmann, M.; Staib, L.; Wiegel, T.; Kron, M.; Henne-Bruns, D.; Link, K.H.; Formentini, A.; Study Group Oncology of Gastrointestinal Tumors. Long-term results of 2 adjuvant trials reveal differences in chemosensitivity and the pattern of metastases between colon cancer and rectal cancer. Clin. Colorectal Cancer 2013, 12, 54–61. [Google Scholar] [CrossRef]
- van der Sijp, M.P.; Bastiaannet, E.; Mesker, W.E.; van der Geest, L.G.; Breugom, A.J.; Steup, W.H.; Marinelli, A.W.; Tseng, L.N.; Tollenaar, R.A.; van de Velde, C.J.; et al. Differences between colon and rectal cancer in complications, short-term survival and recurrences. Int. J. Colorectal Dis. 2016, 31, 1683–1691. [Google Scholar] [CrossRef]
- Guraya, S.Y. Pattern, Stage, and Time of Recurrent Colorectal Cancer After Curative Surgery. Clin. Colorectal Cancer 2019, 18, e223–e228. [Google Scholar] [CrossRef]
- Iqbal, A.; George, T.J. Randomized Clinical Trials in Colon and Rectal Cancer. Surg. Oncol. Clin. 2017, 26, 689–704. [Google Scholar] [CrossRef]
- Delattre, O.; Olschwang, S.; Law, D.J.; Melot, T.; Remvikos, Y.; Salmon, R.J.; Sastre, X.; Validire, P.; Feinberg, A.P.; Thomas, G. Multiple genetic alterations in distal and proximal colorectal cancer. Lancet 1989, 2, 353–356. [Google Scholar] [CrossRef]
- Bauer, K.M.; Hummon, A.B.; Buechler, S. Right-side and left-side colon cancer follow different pathways to relapse. Mol. Carcinog. 2012, 51, 411–421. [Google Scholar] [CrossRef] [PubMed]
- Guinney, J.; Dienstmann, R.; Wang, X.; de Reynies, A.; Schlicker, A.; Soneson, C.; Marisa, L.; Roepman, P.; Nyamundanda, G.; Angelino, P.; et al. The consensus molecular subtypes of colorectal cancer. Nat. Med. 2015, 21, 1350–1356. [Google Scholar] [CrossRef] [PubMed]
- Yaeger, R.; Chatila, W.K.; Lipsyc, M.D.; Hechtman, J.F.; Cercek, A.; Sanchez-Vega, F.; Jayakumaran, G.; Middha, S.; Zehir, A.; Donoghue, M.T.A.; et al. Clinical Sequencing Defines the Genomic Landscape of Metastatic Colorectal Cancer. Cancer Cell 2018, 33, 125–136. [Google Scholar] [CrossRef] [PubMed]
- Sanz-Pamplona, R.; Cordero, D.; Berenguer, A.; Lejbkowicz, F.; Rennert, H.; Salazar, R.; Biondo, S.; Sanjuan, X.; Pujana, M.A.; Rozek, L.; et al. Gene expression differences between colon and rectum tumors. Clin. Cancer Res. 2011, 17, 7303–7312. [Google Scholar] [CrossRef]
- Wei, C.; Chen, J.; Pande, M.; Lynch, P.M.; Frazier, M.L. A pilot study comparing protein expression in different segments of the normal colon and rectum and in normal colon versus adenoma in patients with Lynch syndrome. J. Cancer Res. Clin. Oncol. 2013, 139, 1241–1250. [Google Scholar] [CrossRef][Green Version]
- Barker, N.; Ridgway, R.A.; van Es, J.H.; van de Wetering, M.; Begthel, H.; van den Born, M.; Danenberg, E.; Clarke, A.R.; Sansom, O.J.; Clevers, H. Crypt stem cells as the cells-of-origin of intestinal cancer. Nature 2009, 457, 608–611. [Google Scholar] [CrossRef]
- Barbachano, A.; Fernandez-Barral, A.; Ferrer-Mayorga, G.; Costales-Carrera, A.; Larriba, M.J.; Munoz, A. The endocrine vitamin D system in the gut. Mol. Cell Endocrinol. 2017, 453, 79–87. [Google Scholar] [CrossRef]
- Lee, J.E.; Li, H.; Chan, A.T.; Hollis, B.W.; Lee, I.M.; Stampfer, M.J.; Wu, K.; Giovannucci, E.; Ma, J. Circulating levels of vitamin D and colon and rectal cancer: The Physicians’ Health Study and a meta-analysis of prospective studies. Cancer Prev. Res. 2011, 4, 735–743. [Google Scholar] [CrossRef]
- Touvier, M.; Chan, D.S.; Lau, R.; Aune, D.; Vieira, R.; Greenwood, D.C.; Kampman, E.; Riboli, E.; Hercberg, S.; Norat, T. Meta-analyses of vitamin D intake, 25-hydroxyvitamin D status, vitamin D receptor polymorphisms, and colorectal cancer risk. Cancer Epidemiol. Biomark. Prev. 2011, 20, 1003–1016. [Google Scholar] [CrossRef]
- Heath, A.K.; Hodge, A.M.; Ebeling, P.R.; Eyles, D.W.; Kvaskoff, D.; Buchanan, D.D.; Giles, G.G.; Williamson, E.J.; English, D.R. Circulating 25-Hydroxyvitamin D Concentration and Risk of Breast, Prostate, and Colorectal Cancers: The Melbourne Collaborative Cohort Study. Cancer Epidemiol. Biomark. Prev. 2019, 28, 900–908. [Google Scholar] [CrossRef]
- Ma, Y.; Zhang, P.; Wang, F.; Yang, J.; Liu, Z.; Qin, H. Association between vitamin D and risk of colorectal cancer: A systematic review of prospective studies. J. Clin. Oncol. 2011, 29, 3775–3782. [Google Scholar] [CrossRef] [PubMed]
- Zgaga, L.; Theodoratou, E.; Farrington, S.M.; Din, F.V.; Ooi, L.Y.; Glodzik, D.; Johnston, S.; Tenesa, A.; Campbell, H.; Dunlop, M.G. Plasma vitamin D concentration influences survival outcome after a diagnosis of colorectal cancer. J. Clin. Oncol. 2014, 32, 2430–2439. [Google Scholar] [CrossRef]
- Weinstein, S.J.; Purdue, M.P.; Smith-Warner, S.A.; Mondul, A.M.; Black, A.; Ahn, J.; Huang, W.Y.; Horst, R.L.; Kopp, W.; Rager, H.; et al. Serum 25-hydroxyvitamin D, vitamin D binding protein and risk of colorectal cancer in the Prostate, Lung, Colorectal and Ovarian Cancer Screening Trial. Int. J. Cancer 2015, 136, E654–E664. [Google Scholar] [CrossRef] [PubMed]
- McCullough, M.L.; Zoltick, E.S.; Weinstein, S.J.; Fedirko, V.; Wang, M.; Cook, N.R.; Eliassen, A.H.; Zeleniuch-Jacquotte, A.; Agnoli, C.; Albanes, D.; et al. Circulating Vitamin D and Colorectal Cancer Risk: An International Pooling Project of 17 Cohorts. J. Nat. Cancer Inst. 2019, 111, 158–169. [Google Scholar] [CrossRef] [PubMed]
- Vaughan-Shaw, P.G.; Zgaga, L.; Theodoratou, E.; Blackmur, J.P.; Dunlop, M.G. Whether vitamin D supplementation protects against colorectal cancer risk remains an open question. Eur. J. Cancer 2019, 115, 1–3. [Google Scholar] [CrossRef]
- Zhu, K.; Knuiman, M.; Divitini, M.; Hung, J.; Lim, E.M.; Cooke, B.R.; Walsh, J.P. Lower serum 25-hydroxyvitamin D is associated with colorectal and breast cancer, but not overall cancer risk: A 20-year cohort study. Nutr. Res. 2019, 67, 100–107. [Google Scholar] [CrossRef]
- Abrahamsson, H.; Porojnicu, A.C.; Lindstrom, J.C.; Dueland, S.; Flatmark, K.; Hole, K.H.; Seierstad, T.; Moan, J.; Redalen, K.R.; Meltzer, S.; et al. High level of circulating vitamin D during neoadjuvant therapy may lower risk of metastatic progression in high-risk rectal cancer. BMC Cancer 2019, 19, 488. [Google Scholar] [CrossRef]
- Markotic, A.; Langer, S.; Kelava, T.; Vucic, K.; Turcic, P.; Tokic, T.; Stefancic, L.; Radetic, E.; Farrington, S.; Timofeeva, M.; et al. Higher Post-Operative Serum Vitamin D Level is Associated with Better Survival Outcome in Colorectal Cancer Patients. Nutr. Cancer 2019, 71, 1078–1085. [Google Scholar] [CrossRef]
- Ng, K.; Nimeiri, H.S.; McCleary, N.J.; Abrams, T.A.; Yurgelun, M.B.; Cleary, J.M.; Rubinson, D.A.; Schrag, D.; Miksad, R.; Bullock, A.J.; et al. Effect of High-Dose vs Standard-Dose Vitamin D3 Supplementation on Progression-Free Survival Among Patients With Advanced or Metastatic Colorectal Cancer: The SUNSHINE Randomized Clinical Trial. JAMA 2019, 321, 1370–1379. [Google Scholar] [CrossRef]
- Calderwood, A.H.; Baron, J.A.; Mott, L.A.; Ahnen, D.J.; Bostick, R.M.; Figueiredo, J.C.; Passarelli, M.N.; Rees, J.R.; Robertson, D.J.; Barry, E.L. No Evidence for Posttreatment Effects of Vitamin D and Calcium Supplementation on Risk of Colorectal Adenomas in a Randomized Trial. Cancer Prev. Res. 2019, 12, 295–304. [Google Scholar] [CrossRef]
- Thomas, M.G.; Tebbutt, S.; Williamson, R.C. Vitamin D and its metabolites inhibit cell proliferation in human rectal mucosa and a colon cancer cell line. Gut 1992, 33, 1660–1663. [Google Scholar] [CrossRef] [PubMed]
- Palmer, H.G.; Gonzalez-Sancho, J.M.; Espada, J.; Berciano, M.T.; Puig, I.; Baulida, J.; Quintanilla, M.; Cano, A.; de Herreros, A.G.; Lafarga, M.; et al. Vitamin D(3) promotes the differentiation of colon carcinoma cells by the induction of E-cadherin and the inhibition of beta-catenin signaling. J. Cell Biol. 2001, 154, 369–387. [Google Scholar] [CrossRef] [PubMed]
- Fernandez-Barral, A.; Costales-Carrera, A.; Buira, S.P.; Jung, P.; Ferrer-Mayorga, G.; Larriba, M.J.; Bustamante-Madrid, P.; Dominguez, O.; Real, F.X.; Guerra-Pastrian, L.; et al. Vitamin D differentially regulates colon stem cells in patient-derived normal and tumor organoids. FEBS J. 2020, 287, 53–72. [Google Scholar] [CrossRef] [PubMed]
- Ganesh, K.; Wu, C.; O’Rourke, K.P.; Szeglin, B.C.; Zheng, Y.; Sauve, C.G.; Adileh, M.; Wasserman, I.; Marco, M.R.; Kim, A.S.; et al. A rectal cancer organoid platform to study individual responses to chemoradiation. Nat. Med. 2019, 25, 1607–1614. [Google Scholar] [CrossRef]
- Yao, Y.; Xu, X.; Yang, L.; Zhu, J.; Wan, J.; Shen, L.; Xia, F.; Fu, G.; Deng, Y.; Pan, M.; et al. Patient-Derived Organoids Predict Chemoradiation Responses of Locally Advanced Rectal Cancer. Cell Stem Cell 2020, 26, 17–26. [Google Scholar] [CrossRef]
- Lee, K.; Lindsey, A.S.; Li, N.; Gary, B.; Andrews, J.; Keeton, A.B.; Piazza, G.A. beta-catenin nuclear translocation in colorectal cancer cells is suppressed by PDE10A inhibition, cGMP elevation, and activation of PKG. Oncotarget 2016, 7, 5353–5365. [Google Scholar] [CrossRef]
- Romano, G.; Santi, L.; Bianco, M.R.; Giuffre, M.R.; Pettinato, M.; Bugarin, C.; Garanzini, C.; Savarese, L.; Leoni, S.; Cerrito, M.G.; et al. The TGF-beta pathway is activated by 5-fluorouracil treatment in drug resistant colorectal carcinoma cells. Oncotarget 2016, 7, 22077–22091. [Google Scholar] [CrossRef]
- Hassan, M.K.; Kumar, D.; Naik, M.; Dixit, M. The expression profile and prognostic significance of eukaryotic translation elongation factors in different cancers. PLoS ONE 2018, 13, e0191377. [Google Scholar] [CrossRef]
- Losada, A.; Munoz-Alonso, M.J.; Martinez-Diez, M.; Gago, F.; Dominguez, J.M.; Martinez-Leal, J.F.; Galmarini, C.M. Binding of eEF1A2 to the RNA-dependent protein kinase PKR modulates its activity and promotes tumour cell survival. Br. J. Cancer 2018, 119, 1410–1420. [Google Scholar] [CrossRef]
- Losada, A.; Munoz-Alonso, M.J.; Garcia, C.; Sanchez-Murcia, P.A.; Martinez-Leal, J.F.; Dominguez, J.M.; Lillo, M.P.; Gago, F.; Galmarini, C.M. Translation Elongation Factor eEF1A2 is a Novel Anticancer Target for the Marine Natural Product Plitidepsin. Sci. Rep. 2016, 6, 35100. [Google Scholar] [CrossRef]
- van Riggelen, J.; Yetil, A.; Felsher, D.W. MYC as a regulator of ribosome biogenesis and protein synthesis. Nat. Rev. Cancer 2010, 10, 301–309. [Google Scholar] [CrossRef] [PubMed]
- Muncan, V.; Sansom, O.J.; Tertoolen, L.; Phesse, T.J.; Begthel, H.; Sancho, E.; Cole, A.M.; Gregorieff, A.; de Alboran, I.M.; Clevers, H.; et al. Rapid loss of intestinal crypts upon conditional deletion of the Wnt/Tcf-4 target gene c-Myc. Mol. Cell Biol. 2006, 26, 8418–8426. [Google Scholar] [CrossRef] [PubMed]
- Piazzi, M.; Bavelloni, A.; Gallo, A.; Faenza, I.; Blalock, W.L. Signal Transduction in Ribosome Biogenesis: A Recipe to Avoid Disaster. Int. J. Mol. Sci. 2019, 20, 2718. [Google Scholar] [CrossRef] [PubMed]
- Daniels, D.L.; Mendez, J.; Mosley, A.L.; Ramisetty, S.R.; Murphy, N.; Benink, H.; Wood, K.V.; Urh, M.; Washburn, M.P. Examining the complexity of human RNA polymerase complexes using HaloTag technology coupled to label free quantitative proteomics. J. Proteome Res. 2012, 11, 564–575. [Google Scholar] [CrossRef]
- Proshkin, S.A.; Shematorova, E.K.; Shpakovski, G.V. The Human Isoform of RNA Polymerase II Subunit hRPB11balpha Specifically Interacts with Transcription Factor ATF4. Int. J. Mol. Sci. 2019, 21, 135. [Google Scholar] [CrossRef]
- Proshkin, S.A.; Shematorova, E.K.; Souslova, E.A.; Proshkina, G.M.; Shpakovski, G.V. A minor isoform of the human RNA polymerase II subunit hRPB11 (POLR2J) interacts with several components of the translation initiation factor eIF3. Biochem. 2011, 76, 976–980. [Google Scholar] [CrossRef]
- Jishage, M.; Yu, X.; Shi, Y.; Ganesan, S.J.; Chen, W.Y.; Sali, A.; Chait, B.T.; Asturias, F.J.; Roeder, R.G. Architecture of Pol II(G) and molecular mechanism of transcription regulation by Gdown1. Nat. Struct. Mol. Biol. 2018, 25, 859–867. [Google Scholar] [CrossRef]
- Stevens, P.D.; Wen, Y.A.; Xiong, X.; Zaytseva, Y.Y.; Li, A.T.; Wang, C.; Stevens, A.T.; Farmer, T.N.; Gan, T.; Weiss, H.L.; et al. Erbin Suppresses KSR1-Mediated RAS/RAF Signaling and Tumorigenesis in Colorectal Cancer. Cancer Res. 2018, 78, 4839–4852. [Google Scholar] [CrossRef]
- Mu, Z.; Wang, L.; Deng, W.; Wang, J.; Wu, G. Structural insight into the Ragulator complex which anchors mTORC1 to the lysosomal membrane. Cell Discov. 2017, 3, 17049. [Google Scholar] [CrossRef]
- Mo, D.; Liu, W.; Li, Y.; Cui, W. Long Non-coding RNA Zinc Finger Antisense 1 (ZFAS1) Regulates Proliferation, Migration, Invasion, and Apoptosis by Targeting MiR-7-5p in Colorectal Cancer. Med. Sci. Monit. 2019, 25, 5150–5158. [Google Scholar] [CrossRef]
- Neme, A.; Seuter, S.; Malinen, M.; Nurmi, T.; Tuomainen, T.P.; Virtanen, J.K.; Carlberg, C. In vivo transcriptome changes of human white blood cells in response to vitamin D. J. Steroid Biochem. Mol. Biol. 2019, 188, 71–76. [Google Scholar] [CrossRef] [PubMed]
- Lou, X.; Sun, B.; Song, J.; Wang, Y.; Jiang, J.; Xu, Y.; Ren, Z.; Su, C. Human sulfatase 1 exerts anti-tumor activity by inhibiting the AKT/ CDK4 signaling pathway in melanoma. Oncotarget 2016, 7, 84486–84495. [Google Scholar] [CrossRef][Green Version]
- Lee, H.Y.; Yeh, B.W.; Chan, T.C.; Yang, K.F.; Li, W.M.; Huang, C.N.; Ke, H.L.; Li, C.C.; Yeh, H.C.; Liang, P.I.; et al. Sulfatase-1 overexpression indicates poor prognosis in urothelial carcinoma of the urinary bladder and upper tract. Oncotarget 2017, 8, 47216–47229. [Google Scholar] [CrossRef] [PubMed]
- Krishnakumar, K.; Chakravorty, I.; Foy, W.; Allen, S.; Justo, T.; Mukherjee, A.; Dhoot, G.K. Multi-tasking Sulf1/Sulf2 enzymes do not only facilitate extracellular cell signalling but also participate in cell cycle related nuclear events. Exp. Cell Res. 2018, 364, 16–27. [Google Scholar] [CrossRef] [PubMed]
- Barbáchano, A.; Larriba, M.J.; Ferrer-Mayorga, G.; González-Sancho, J.M.; Muñoz, A. Vitamin D and Colon Cancer. In Vitamin D, 4th ed.; Feldman, D., Pike, J.W., Bouillon, R., Giovannucci, E., Goltzman, D., Hewison, M., Eds.; Elsevier: London, UK; Academic Press: San Diego, CA, USA, 2018; Volume 2, pp. 837–862. [Google Scholar] [CrossRef]
- Pendas-Franco, N.; Garcia, J.M.; Pena, C.; Valle, N.; Palmer, H.G.; Heinaniemi, M.; Carlberg, C.; Jimenez, B.; Bonilla, F.; Munoz, A.; et al. DICKKOPF-4 is induced by TCF/beta-catenin and upregulated in human colon cancer, promotes tumour cell invasion and angiogenesis and is repressed by 1alpha,25-dihydroxyvitamin D3. Oncogene 2008, 27, 4467–4477. [Google Scholar] [CrossRef] [PubMed]
- Wang, T.; Qin, Y.; Lai, H.; Wei, W.; Li, Z.; Yang, Y.; Huang, M.; Chen, J. The prognostic value of ADRA1 subfamily genes in gastric carcinoma. Oncol. Lett. 2019, 18, 3150–3158. [Google Scholar] [CrossRef]
- Chakraborty, S.; Kumar, A.; Faheem, M.M.; Katoch, A.; Kumar, A.; Jamwal, V.L.; Nayak, D.; Golani, A.; Rasool, R.U.; Ahmad, S.M.; et al. Vimentin activation in early apoptotic cancer cells errands survival pathways during DNA damage inducer CPT treatment in colon carcinoma model. Cell Death Dis. 2019, 10, 467. [Google Scholar] [CrossRef]
- van de Wetering, M.; Francies, H.E.; Francis, J.M.; Bounova, G.; Iorio, F.; Pronk, A.; van Houdt, W.; van Gorp, J.; Taylor-Weiner, A.; Kester, L.; et al. Prospective derivation of a living organoid biobank of colorectal cancer patients. Cell 2015, 161, 933–945. [Google Scholar] [CrossRef]
- Fujii, M.; Shimokawa, M.; Date, S.; Takano, A.; Matano, M.; Nanki, K.; Ohta, Y.; Toshimitsu, K.; Nakazato, Y.; Kawasaki, K.; et al. A Colorectal Tumor Organoid Library Demonstrates Progressive Loss of Niche Factor Requirements during Tumorigenesis. Cell Stem Cell 2016, 18, 827–838. [Google Scholar] [CrossRef]
- Knight, J.M.; Kim, E.; Ivanov, I.; Davidson, L.A.; Goldsby, J.S.; Hullar, M.A.; Randolph, T.W.; Kaz, A.M.; Levy, L.; Lampe, J.W.; et al. Comprehensive site-specific whole genome profiling of stromal and epithelial colonic gene signatures in human sigmoid colon and rectal tissue. Physiol. Genom. 2016, 48, 651–659. [Google Scholar] [CrossRef]
- Cramer, J.M.; Thompson, T.; Geskin, A.; LaFramboise, W.; Lagasse, E. Distinct human stem cell populations in small and large intestine. PLoS ONE 2015, 10, e0118792. [Google Scholar] [CrossRef] [PubMed]
- Middendorp, S.; Schneeberger, K.; Wiegerinck, C.L.; Mokry, M.; Akkerman, R.D.; van Wijngaarden, S.; Clevers, H.; Nieuwenhuis, E.E. Adult stem cells in the small intestine are intrinsically programmed with their location-specific function. Stem Cells 2014, 32, 1083–1091. [Google Scholar] [CrossRef] [PubMed]
- Wang, X.; Yamamoto, Y.; Wilson, L.H.; Zhang, T.; Howitt, B.E.; Farrow, M.A.; Kern, F.; Ning, G.; Hong, Y.; Khor, C.C.; et al. Cloning and variation of ground state intestinal stem cells. Nature 2015, 522, 173–178. [Google Scholar] [CrossRef] [PubMed]
- Atlas, N. Comprehensive molecular characterization of human colon and rectal cancer. Nature 2012, 487, 330–337. [Google Scholar] [CrossRef]
- Penzo, M.; Montanaro, L.; Trere, D.; Derenzini, M. The Ribosome Biogenesis-Cancer Connection. Cells 2019, 8, 55. [Google Scholar] [CrossRef] [PubMed]
- Morral, C.; Stanisavljevic, J.; Hernando-Momblona, X.; Mereu, E.; Alvarez-Varela, A.; Cortina, C.; Stork, D.; Slebe, F.; Turon, G.; Whissell, G.; et al. Zonation of Ribosomal DNA Transcription Defines a Stem Cell Hierarchy in Colorectal Cancer. Cell Stem Cell 2020, 26, 845–861.e12. [Google Scholar] [CrossRef]
- Sarkissyan, M.; Wu, Y.; Chen, Z.; Mishra, D.K.; Sarkissyan, S.; Giannikopoulos, I.; Vadgama, J.V. Vitamin D receptor FokI gene polymorphisms may be associated with colorectal cancer among African American and Hispanic participants. Cancer 2014, 120, 1387–1393. [Google Scholar] [CrossRef]
- Takeshige, N.; Yin, G.; Ohnaka, K.; Kono, S.; Ueki, T.; Tanaka, M.; Maehara, Y.; Okamura, T.; Ikejiri, K.; Maekawa, T.; et al. Associations between vitamin D receptor (VDR) gene polymorphisms and colorectal cancer risk and effect modifications of dietary calcium and vitamin D in a Japanese population. Asian Pac. J. Cancer Prev. 2015, 16, 2019–2026. [Google Scholar] [CrossRef]
- Berger, M.D.; Stintzing, S.; Heinemann, V.; Cao, S.; Yang, D.; Sunakawa, Y.; Matsusaka, S.; Ning, Y.; Okazaki, S.; Miyamoto, Y.; et al. A Polymorphism within the Vitamin D Transporter Gene Predicts Outcome in Metastatic Colorectal Cancer Patients Treated with FOLFIRI/Bevacizumab or FOLFIRI/Cetuximab. Clin. Cancer Res. 2018, 24, 784–793. [Google Scholar] [CrossRef]
- Zhu, Y.; Wang, P.P.; Zhai, G.; Bapat, B.; Savas, S.; Woodrow, J.R.; Sharma, I.; Li, Y.; Zhou, X.; Yang, N.; et al. Vitamin D receptor and calcium-sensing receptor polymorphisms and colorectal cancer survival in the Newfoundland population. Br. J. Cancer 2017, 117, 898–906. [Google Scholar] [CrossRef][Green Version]
- Cho, Y.A.; Lee, J.; Oh, J.H.; Chang, H.J.; Sohn, D.K.; Shin, A.; Kim, J. Vitamin D receptor FokI polymorphism and the risks of colorectal cancer, inflammatory bowel disease, and colorectal adenoma. Sci. Rep. 2018, 8, 12899. [Google Scholar] [CrossRef] [PubMed]
- Yuan, C.; Sato, K.; Hollis, B.W.; Zhang, S.; Niedzwiecki, D.; Ou, F.S.; Chang, I.W.; O’Neil, B.H.; Innocenti, F.; Lenz, H.J.; et al. Plasma 25-Hydroxyvitamin D Levels and Survival in Patients with Advanced or Metastatic Colorectal Cancer: Findings from CALGB/SWOG 80405 (Alliance). Clin. Cancer Res. 2019, 25, 7497–7505. [Google Scholar] [CrossRef] [PubMed]
- Cristobal, A.; van den Toorn, H.W.P.; van de Wetering, M.; Clevers, H.; Heck, A.J.R.; Mohammed, S. Personalized Proteome Profiles of Healthy and Tumor Human Colon Organoids Reveal Both Individual Diversity and Basic Features of Colorectal Cancer. Cell Rep. 2017, 18, 263–274. [Google Scholar] [CrossRef] [PubMed]
- Michels, B.E.; Mosa, M.H.; Grebbin, B.M.; Yepes, D.; Darvishi, T.; Hausmann, J.; Urlaub, H.; Zeuzem, S.; Kvasnicka, H.M.; Oellerich, T.; et al. Human colon organoids reveal distinct physiologic and oncogenic Wnt responses. J. Exp. Med. 2019, 216, 704–720. [Google Scholar] [CrossRef]
- Bolhaqueiro, A.C.F.; Ponsioen, B.; Bakker, B.; Klaasen, S.J.; Kucukkose, E.; van Jaarsveld, R.H.; Vivie, J.; Verlaan-Klink, I.; Hami, N.; Spierings, D.C.J.; et al. Ongoing chromosomal instability and karyotype evolution in human colorectal cancer organoids. Nat. Genet. 2019, 51, 824–834. [Google Scholar] [CrossRef]
- Kim, D.; Pertea, G.; Trapnell, C.; Pimentel, H.; Kelley, R.; Salzberg, S.L. TopHat2: Accurate alignment of transcriptomes in the presence of insertions, deletions and gene fusions. Genome Biol. 2013, 14, R36. [Google Scholar] [CrossRef]
- Love, M.I.; Huber, W.; Anders, S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 2014, 15, 550. [Google Scholar] [CrossRef]
- Perez-Silva, J.G.; Araujo-Voces, M.; Quesada, V. nVenn: Generalized, quasi-proportional Venn and Euler diagrams. Bioinformatics 2018, 34, 2322–2324. [Google Scholar] [CrossRef]

| Patient | Gender | Age (y) | Sample | T a | N b | M c |
|---|---|---|---|---|---|---|
| #40 | Female | 65 | Normal: Colon + Rectum | pTis | pN0 | cM0 |
| #41 | Male | 88 | Normal: Colon + Rectum | cT3, ypT0 | cN2a, ypN0 | cM1a |
| #42 | Male | 70 | Normal: Colon + Rectum | cT2-3, ypTis | cN1a, ypN0 | cM0 |
| #43 | Male | 64 | Normal: Colon + Rectum | pT4b | pN0 | cM0 |
| #44 | Male | 65 | Normal: Colon + Rectum Tumor: Lower rectum | cT3, ypT2 | cN2, ypN0 | cM0 |
| #45 | Male | 72 | Normal: Colon + Rectum | cT3, ypT3 | cN2, ypN0 | cM0 |
| #46 | Male | 69 | Tumor: Lower rectum | cT3, ypT2 | cN1-2, ypN0 | cM0 |
| #47 | Male | 88 | Tumor: Lower rectum | cT3-4, ypT3 | cN1, ypN0 | cM0 |
| #48 | Female | 69 | Tumor: Medial rectum | cT4 | cN2b | cM1 |
| #49 | Female | 51 | Tumor: Upper rectum * | NA | cN+ | cM1 |
| #50 | Female | 88 | Tumor: Upper rectum | pT3 | pN1b | cM0 |
| Buffer | Reagent | Concentration | Supplier |
|---|---|---|---|
| Washing buffer | Advanced DMEM/F12 | 100% | Thermo Fisher Scientific |
| Hepes | 10 mM | Thermo Fisher Scientific | |
| Glutamax | 10 mM | Thermo Fisher Scientific | |
| Normal culture medium | Advanced DMEM/F12 | 50% | Thermo Fisher Scientific |
| Wnt3a-Conditioned medium | 50% | Previously described [33] | |
| Hepes | 10 mM | Thermo Fisher Scientific | |
| Glutamax | 10 mM | Thermo Fisher Scientific | |
| Nicotinamide | 10 mM | Sigma-Aldrich, MD, USA | |
| N2 | 1× | Thermo Fisher Scientific | |
| B27 | 1× | Thermo Fisher Scientific | |
| N-acetyl-L-cysteine | 1 mM | Sigma-Aldrich | |
| Primocin | 1:500 | Invivogen | |
| Noggin | 0.1 μg/mL | Peprotech, NJ, USA | |
| Gastrin | 1 μg/mL | Tocris, Bristol, UK | |
| Y-27632 | 10 μM | Tocris | |
| RSPO1 | 1 μg/mL | Sinobiological, Beijin, China | |
| EGF | 50 ng/mL | Peprotech | |
| PGE2 | 0.02 μM | Sigma-Aldrich | |
| LY-2157299 | 1 μM | Axon-Medchem, Groningen, The Netherlands | |
| SB-202190 | 10 μM | Sigma-Aldrich | |
| Lysis buffer | TrisHCL pH 8.0 | 50 mM | Sigma-Aldrich |
| EDTA pH 8.0 | 100 mM | Sigma-Aldrich | |
| NaCl | 100 mM | Sigma-Aldrich | |
| SDS | 1% | Sigma-Aldrich | |
| Proteinase K | 20 mg/mL | Merck-Millipore, MA, USA |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Costales-Carrera, A.; Fernández-Barral, A.; Bustamante-Madrid, P.; Domínguez, O.; Guerra-Pastrián, L.; Cantero, R.; del Peso, L.; Burgos, A.; Barbáchano, A.; Muñoz, A. Comparative Study of Organoids from Patient-Derived Normal and Tumor Colon and Rectal Tissue. Cancers 2020, 12, 2302. https://doi.org/10.3390/cancers12082302
Costales-Carrera A, Fernández-Barral A, Bustamante-Madrid P, Domínguez O, Guerra-Pastrián L, Cantero R, del Peso L, Burgos A, Barbáchano A, Muñoz A. Comparative Study of Organoids from Patient-Derived Normal and Tumor Colon and Rectal Tissue. Cancers. 2020; 12(8):2302. https://doi.org/10.3390/cancers12082302
Chicago/Turabian StyleCostales-Carrera, Alba, Asunción Fernández-Barral, Pilar Bustamante-Madrid, Orlando Domínguez, Laura Guerra-Pastrián, Ramón Cantero, Luis del Peso, Aurora Burgos, Antonio Barbáchano, and Alberto Muñoz. 2020. "Comparative Study of Organoids from Patient-Derived Normal and Tumor Colon and Rectal Tissue" Cancers 12, no. 8: 2302. https://doi.org/10.3390/cancers12082302
APA StyleCostales-Carrera, A., Fernández-Barral, A., Bustamante-Madrid, P., Domínguez, O., Guerra-Pastrián, L., Cantero, R., del Peso, L., Burgos, A., Barbáchano, A., & Muñoz, A. (2020). Comparative Study of Organoids from Patient-Derived Normal and Tumor Colon and Rectal Tissue. Cancers, 12(8), 2302. https://doi.org/10.3390/cancers12082302






