Quercetin–Resveratrol Combination for Prostate Cancer Management in TRAMP Mice
Abstract
1. Introduction
2. Results and Discussion
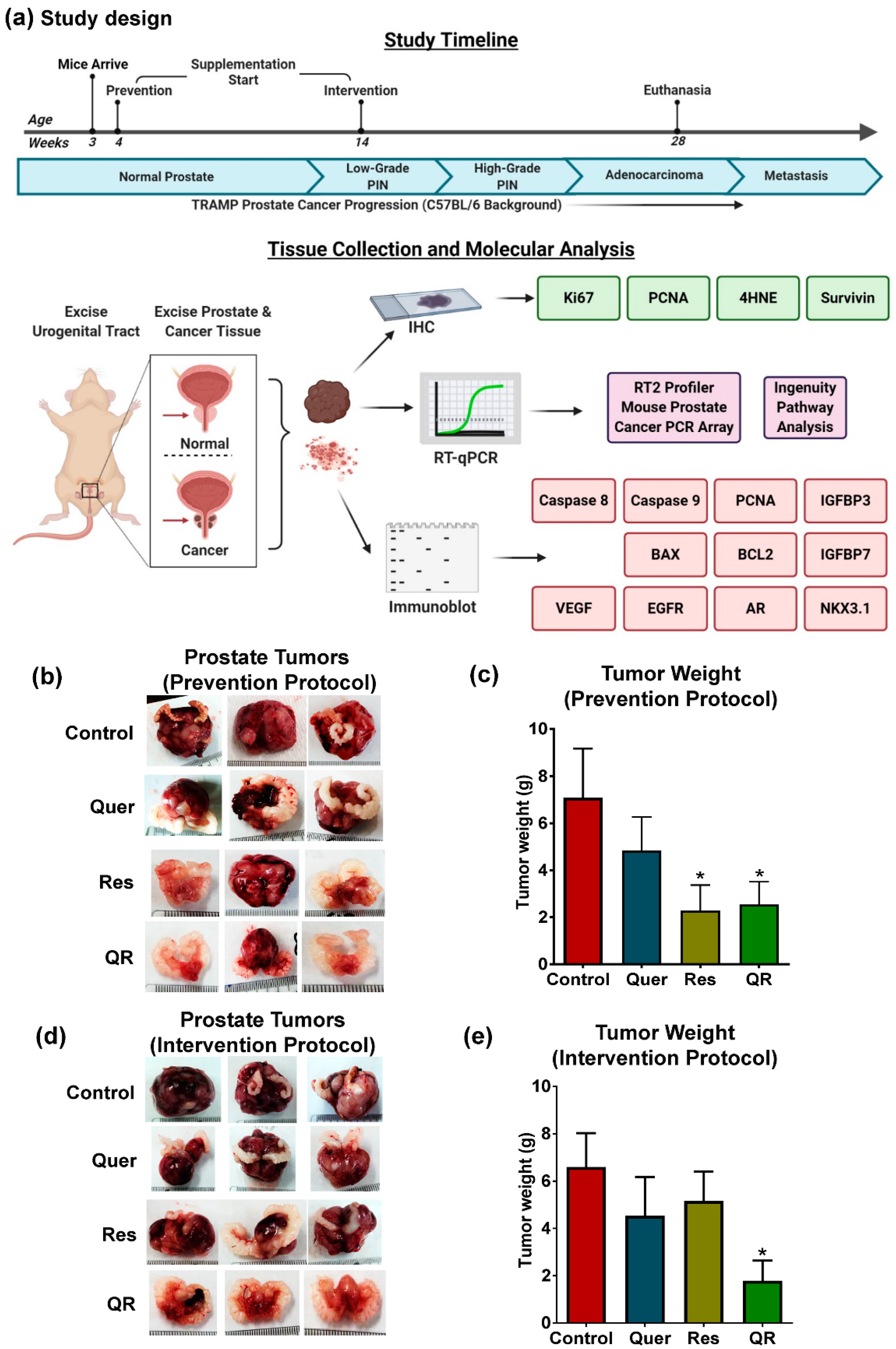
2.1. Quercetin–Resveratrol Combination Exerted Significant Antitumor Effects against PCa
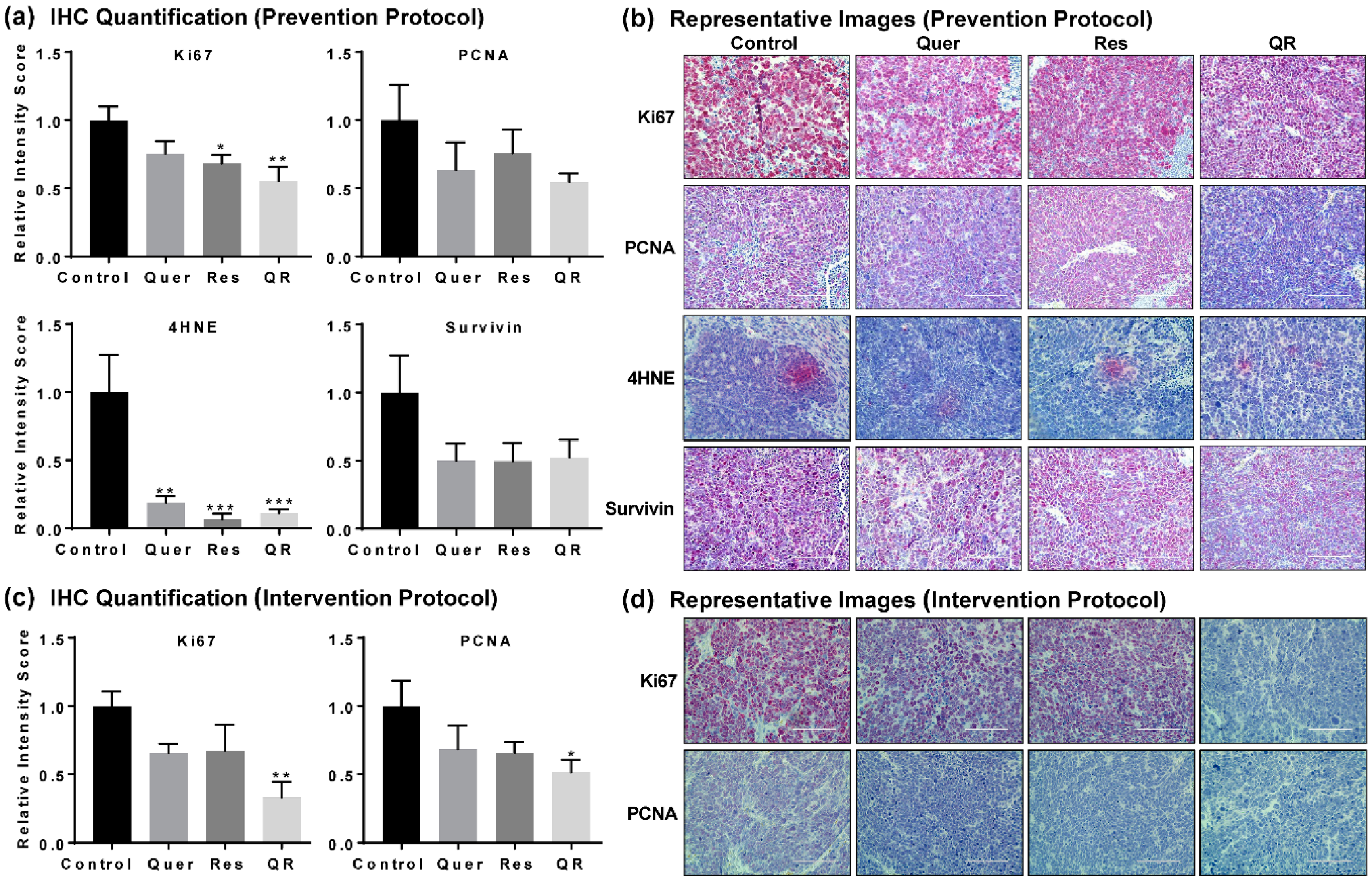
2.2. Anti-PCa Effect of Quercetin–Resveratrol Is Associated with Marked Inhibition in Markers of Cell Proliferation, Oxidative Stress, and Tumor Survival, as well as Induction of Apoptosis
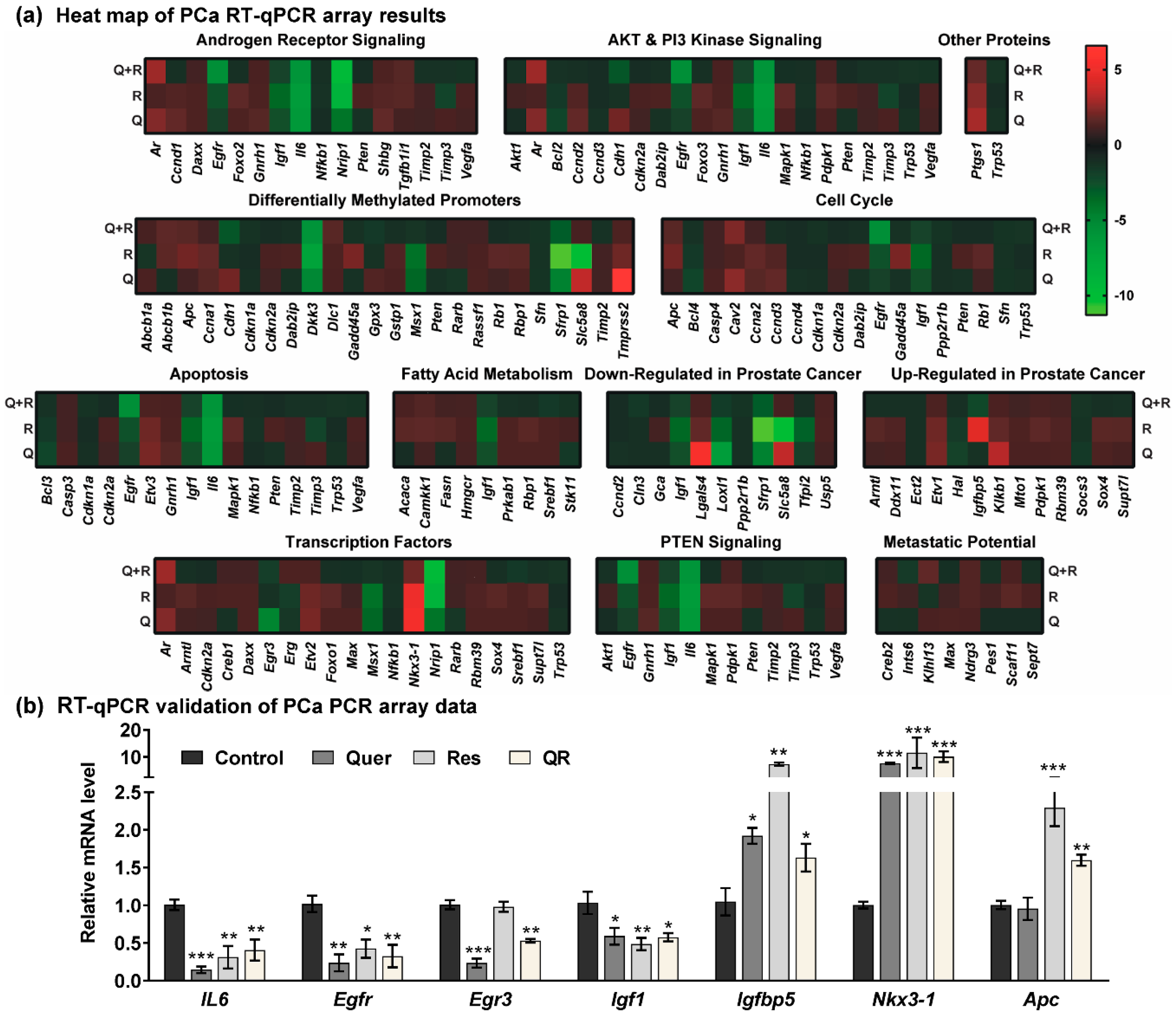
2.3. Anti-PCa Effects of Quercetin–Resveratrol Are Associated with Key PCa-Related Genes
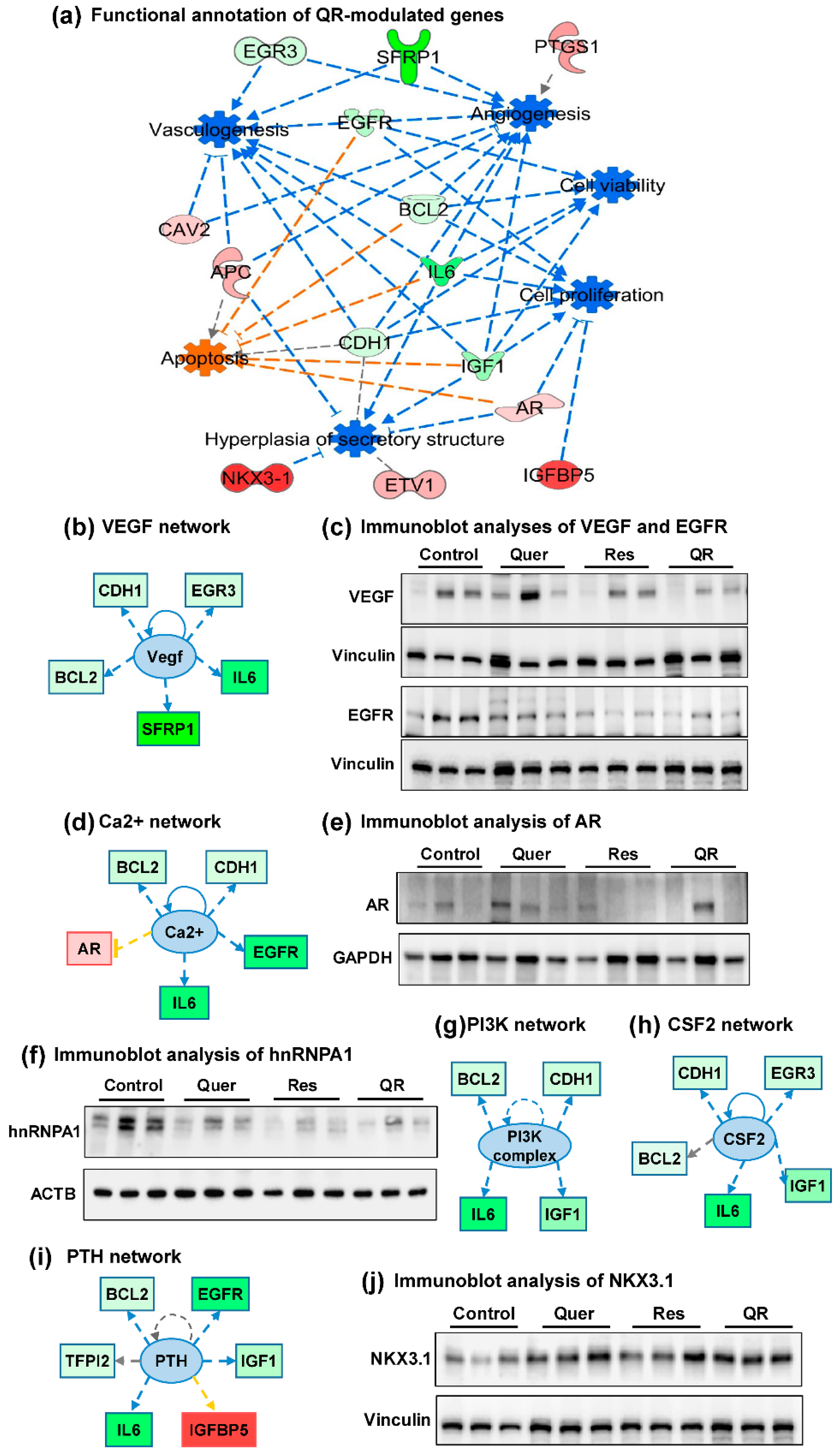
2.4. Functional Annotation of Quercetin–Resveratrol Modulated Genes Predicts Cumulative Antitumor Actions and Upstream Inhibition of Major PCa-Associated Pathways
3. Materials and Methods
3.1. TRAMP Mice
3.2. Experimental Design and Treatments
3.3. Preparation of Tumor Protein Lysates and RNA
3.4. Quantitative Reverse Transcription PCR (RT-qPCR) Analysis
3.5. Immunoblot Analysis
3.6. PCa PCR Array Analysis
3.7. Ingenuity Pathway Analysis (IPA)
3.8. Immunohistochemical (IHC) Analysis
3.9. Statistical Analysis
4. Conclusions
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Siegel, R.L.; Miller, K.D.; Jemal, A. Cancer statistics, 2020. CA Cancer J. Clin. 2020, 70, 7–30. [Google Scholar] [CrossRef] [PubMed]
- Martin, N.E.; Mucci, L.A.; Loda, M.; Depinho, R.A. Prognostic determinants in prostate cancer. Cancer J. 2011, 17, 429–437. [Google Scholar] [CrossRef] [PubMed]
- Chhabra, G.; Singh, C.K.; Ndiaye, M.A.; Fedorowicz, S.; Molot, A.; Ahmad, N. Prostate cancer chemoprevention by natural agents: Clinical evidence and potential implications. Cancer Lett. 2018, 422, 9–18. [Google Scholar] [CrossRef] [PubMed]
- Chhabra, G.; Dixit, A. Structure modeling and antidiabetic activity of a seed protein of Momordica charantia in non-obese diabetic (NOD) mice. Bioinformation 2013, 9, 766–770. [Google Scholar] [CrossRef] [PubMed]
- Mitchell, S.H.; Zhu, W.; Young, C.Y. Resveratrol inhibits the expression and function of the androgen receptor in LNCaP prostate cancer cells. Cancer Res. 1999, 59, 5892–5895. [Google Scholar] [PubMed]
- Wang, T.T.; Hudson, T.S.; Wang, T.C.; Remsberg, C.M.; Davies, N.M.; Takahashi, Y.; Kim, Y.S.; Seifried, H.; Vinyard, B.T.; Perkins, S.N.; et al. Differential effects of resveratrol on androgen-responsive LNCaP human prostate cancer cells in vitro and in vivo. Carcinogenesis 2008, 29, 2001–2010. [Google Scholar] [CrossRef] [PubMed]
- Wang, Y.; Romigh, T.; He, X.; Orloff, M.S.; Silverman, R.H.; Heston, W.D.; Eng, C. Resveratrol regulates the PTEN/AKT pathway through androgen receptor-dependent and -independent mechanisms in prostate cancer cell lines. Hum. Mol. Genet. 2010, 19, 4319–4329. [Google Scholar] [CrossRef]
- Pezzuto, J.M. Resveratrol as an Inhibitor of Carcinogenesis. Pharm. Biol. 2008, 46, 443–573. [Google Scholar] [CrossRef]
- Singh, C.K.; Chhabra, G.; Ahmad, N. Chapter 7: Resveratrol and Cancer Cell Biology. In Resveratrol: State of the Art Science and Health Applications; Wu, M.J., Hsieh, T.C., Eds.; World Scientific Publishing Co Pte Ltd.: Singapore, 2018; pp. 183–207. [Google Scholar]
- Singh, C.K.; George, J.; Ahmad, N. Resveratrol-based combinatorial strategies for cancer management. Ann. N. Y. Acad. Sci. 2013, 1290, 113–121. [Google Scholar] [CrossRef]
- Singh, C.K.; Liu, X.; Ahmad, N. Resveratrol, in its natural combination in whole grape, for health promotion and disease management. Ann. N. Y. Acad. Sci. 2015, 1348, 150–160. [Google Scholar] [CrossRef]
- Aziz, M.H.; Nihal, M.; Fu, V.X.; Jarrard, D.F.; Ahmad, N. Resveratrol-caused apoptosis of human prostate carcinoma LNCaP cells is mediated via modulation of phosphatidylinositol 3’-kinase/Akt pathway and Bcl-2 family proteins. Mol. Cancer Ther. 2006, 5, 1335–1341. [Google Scholar] [CrossRef] [PubMed]
- Harper, C.E.; Patel, B.B.; Wang, J.; Arabshahi, A.; Eltoum, I.A.; Lamartiniere, C.A. Resveratrol suppresses prostate cancer progression in transgenic mice. Carcinogenesis 2007, 28, 1946–1953. [Google Scholar] [CrossRef] [PubMed]
- Seeni, A.; Takahashi, S.; Takeshita, K.; Tang, M.; Sugiura, S.; Sato, S.Y.; Shirai, T. Suppression of prostate cancer growth by resveratrol in the transgenic rat for adenocarcinoma of prostate (TRAP) model. Asian Pac. J. Cancer Prev. 2008, 9, 7–14. [Google Scholar] [PubMed]
- Singh, C.K.; Ndiaye, M.A.; Ahmad, N. Resveratrol and cancer: Challenges for clinical translation. Biochim. Biophys. Acta 2015, 1852, 1178–1185. [Google Scholar] [CrossRef] [PubMed]
- Singh, C.K.; Siddiqui, I.A.; El-Abd, S.; Mukhtar, H.; Ahmad, N. Combination chemoprevention with grape antioxidants. Mol. Nutr. Food Res. 2016, 60, 1406–1415. [Google Scholar] [CrossRef]
- Hashemzaei, M.; Far, A.D.; Yari, A.; Heravi, R.E.; Tabrizian, K.; Taghdisi, S.M.; Sadegh, S.E.; Tsarouhas, K.; Kouretas, D.; Tzanakakis, G.; et al. Anticancer and apoptosis inducing effects of quercetin in vitro and in vivo. Oncol. Rep. 2017, 38, 819–828. [Google Scholar] [CrossRef]
- Gibellini, L.; Pinti, M.; Nasi, M.; Montagna, J.P.; De Biasi, S.; Roat, E.; Bertoncelli, L.; Cooper, E.L.; Cossarizza, A. Quercetin and cancer chemoprevention. Evid. Based Complement Alternat. Med. 2011, 2011, 591356. [Google Scholar] [CrossRef]
- Singh, C.K.; Mintie, C.A.; Ndiaye, M.A.; Chhabra, G.; Dakup, P.P.; Ye, T.; Yu, M.; Ahmad, N. Chemoprotective Effects of Dietary Grape Powder on UVB Radiation-Mediated Skin Carcinogenesis in SKH-1 Hairless Mice. J. Invest. Dermatol. 2019, 139, 552–561. [Google Scholar] [CrossRef]
- Sharmila, G.; Athirai, T.; Kiruthiga, B.; Senthilkumar, K.; Elumalai, P.; Arunkumar, R.; Arunakaran, J. Chemopreventive effect of quercetin in MNU and testosterone induced prostate cancer of Sprague-Dawley rats. Nuter. Cancer 2014, 66, 38–46. [Google Scholar] [CrossRef]
- Xing, N.; Chen, Y.; Mitchell, S.H.; Young, C.Y. Quercetin inhibits the expression and function of the androgen receptor in LNCaP prostate cancer cells. Carcinogenesis 2001, 22, 409–414. [Google Scholar] [CrossRef]
- Yang, F.; Jiang, X.; Song, L.; Wang, H.; Mei, Z.; Xu, Z.; Xing, N. Quercetin inhibits angiogenesis through thrombospondin-1 upregulation to antagonize human prostate cancer PC-3 cell growth in vitro and in vivo. Oncol. Rep. 2016, 35, 1602–1610. [Google Scholar] [CrossRef] [PubMed]
- Ward, A.B.; Mir, H.; Kapur, N.; Gales, D.N.; Carriere, P.P.; Singh, S. Quercetin inhibits prostate cancer by attenuating cell survival and inhibiting anti-apoptotic pathways. World J. Surg. Oncol. 2018, 16, 108. [Google Scholar] [CrossRef] [PubMed]
- De Santi, C.; Pietrabissa, A.; Spisni, R.; Mosca, F.; Pacifici, G.M. Sulphation of resveratrol, a natural compound present in wine, and its inhibition by natural flavonoids. Xenobiotica 2000, 30, 857–866. [Google Scholar] [CrossRef] [PubMed]
- Costello, L.C.; Franklin, R.B.; Zou, J.; Feng, P.; Bok, R.; Mark, G.S.; Kurhanewicz, J. Human prostate cancer ZIP1/zinc/citrate genetic/metabolic relationship in the TRAMP prostate cancer animal model. Cancer Biol. Ther. 2011, 12, 1078–1084. [Google Scholar] [CrossRef] [PubMed]
- Valkenburg, K.C.; Williams, B.O. Mouse models of prostate cancer. Prostate Cancer 2011, 2011, 895238. [Google Scholar] [CrossRef] [PubMed]
- Gingrich, J.R.; Barrios, R.J.; Morton, R.A.; Boyce, B.F.; DeMayo, F.J.; Finegold, M.J.; Angelopoulou, R.; Rosen, J.M.; Greenberg, N.M. Metastatic prostate cancer in a transgenic mouse. Cancer Res. 1996, 56, 4096–4102. [Google Scholar] [PubMed]
- Yeh, I.T.; Reddick, R.L.; Kumar, A.P. Malignancy arising in seminal vesicles in the transgenic adenocarcinoma of mouse prostate (TRAMP) model. Prostate 2009, 69, 755–760. [Google Scholar] [CrossRef]
- Hurwitz, A.A.; Foster, B.A.; Allison, J.P.; Greenberg, N.M.; Kwon, E.D. The TRAMP mouse as a model for prostate cancer. Curr. Protoc. Immunol. 2001. [Google Scholar] [CrossRef]
- Gelman, I.H. How the TRAMP Model Revolutionized the Study of Prostate Cancer Progression. Cancer Res. 2016, 76, 6137–6139. [Google Scholar] [CrossRef]
- Reagan-Shaw, S.; Nihal, M.; Ahmad, N. Dose translation from animal to human studies revisited. FASEB J. 2008, 22, 659–661. [Google Scholar] [CrossRef]
- Crowe, A.R.; Yue, W. Semi-quantitative Determination of Protein Expression using Immunohistochemistry Staining and Analysis: An Integrated Protocol. Bio-Protocol 2019, 9. [Google Scholar] [CrossRef] [PubMed]
- Paunesku, T.; Mittal, S.; Protic, M.; Oryhon, J.; Korolev, S.V.; Joachimiak, A.; Woloschak, G.E. Proliferating cell nuclear antigen (PCNA): Ringmaster of the genome. Int. J. Radiat. Biol. 2001, 77, 1007–1021. [Google Scholar] [CrossRef] [PubMed]
- Hata, A.N.; Engelman, J.A.; Faber, A.C. The BCL2 Family: Key Mediators of the Apoptotic Response to Targeted Anticancer Therapeutics. Cancer Discov. 2015, 5, 475–487. [Google Scholar] [CrossRef] [PubMed]
- Chen, Z.H.; Saito, Y.; Yoshida, Y.; Sekine, A.; Noguchi, N.; Niki, E. 4-Hydroxynonenal induces adaptive response and enhances PC12 cell tolerance primarily through induction of thioredoxin reductase 1 via activation of Nrf2. J. Biol. Chem. 2005, 280, 41921–41927. [Google Scholar] [CrossRef]
- Sajadimajd, S.; Khazaei, M. Oxidative Stress and Cancer: The Role of Nrf2. Curr. Cancer Drug Targets 2018, 18, 538–557. [Google Scholar] [CrossRef]
- Schumacher, F.R.; Cheng, I.; Freedman, M.L.; Mucci, L.; Allen, N.E.; Pollak, M.N.; Hayes, R.B.; Stram, D.O.; Canzian, F.; Henderson, B.E.; et al. A comprehensive analysis of common IGF1, IGFBP1 and IGFBP3 genetic variation with prospective IGF-I and IGFBP-3 blood levels and prostate cancer risk among Caucasians. Hum. Mol. Genet. 2010, 19, 3089–3101. [Google Scholar] [CrossRef]
- Roberts, C.T., Jr. IGF-1 and prostate cancer. In Biology of IGF-1: Its Interaction with Insulin in Health and Malignant States: Novartis Foundation Symposium; Bock, G., Goode, J., Eds.; John Wiley & Sons: Hoboken, NJ, USA, 2004. [Google Scholar]
- Jung, M.; Bu, S.Y.; Tak, K.H.; Park, J.E.; Kim, E. Anticarcinogenic effect of quercetin by inhibition of insulin-like growth factor (IGF)-1 signaling in mouse skin cancer. Nutr. Res. Pr. 2013, 7, 439–445. [Google Scholar] [CrossRef]
- Beattie, J.; Allan, G.J.; Lochrie, J.D.; Flint, D.J. Insulin-like growth factor-binding protein-5 (IGFBP-5): A critical member of the IGF axis. Biochem. J. 2006, 395, 1–19. [Google Scholar] [CrossRef]
- Bach, L.A. Insulin-like growth factor binding proteins 4-6. Best Pr. Res. Clin. Endocrinol. Metab. 2015, 29, 713–722. [Google Scholar] [CrossRef]
- Gregory, C.W.; Kim, D.; Ye, P.; D’Ercole, A.J.; Pretlow, T.G.; Mohler, J.L.; French, F.S. Androgen receptor up-regulates insulin-like growth factor binding protein-5 (IGFBP-5) expression in a human prostate cancer xenograft. Endocrinology 1999, 140, 2372–2381. [Google Scholar] [CrossRef] [PubMed]
- Miyake, H.; Pollak, M.; Gleave, M.E. Castration-induced up-regulation of insulin-like growth factor binding protein-5 potentiates insulin-like growth factor-I activity and accelerates progression to androgen independence in prostate cancer models. Cancer Res. 2000, 60, 3058–3064. [Google Scholar] [PubMed]
- Jin, L.; Shen, F.; Weinfeld, M.; Sergi, C. Insulin Growth Factor Binding Protein 7 (IGFBP7)-Related Cancer and IGFBP3 and IGFBP7 Crosstalk. Front. Oncol. 2020, 10, 727. [Google Scholar] [CrossRef] [PubMed]
- Singh, C.K.; Chhabra, G.; Mintie, C.A.; Ahmad, N. Grape chemopreventive agents against angiogenesis and metastasis. In Natural Products for Cancer Chemoprevention: Single Compounds and Combinations; Pezzuto, J.M., Vang, O., Eds.; Springer International Publishing: Cham, Switzerland, 2020; pp. 375–400. [Google Scholar] [CrossRef]
- Tabernero, J. The role of VEGF and EGFR inhibition: Implications for combining anti-VEGF and anti-EGFR agents. Mol. Cancer Res. 2007, 5, 203–220. [Google Scholar] [CrossRef] [PubMed]
- Day, K.C.; Lorenzatti Hiles, G.; Kozminsky, M.; Dawsey, S.J.; Paul, A.; Broses, L.J.; Shah, R.; Kunja, L.P.; Hall, C.; Palanisamy, N.; et al. HER2 and EGFR Overexpression Support Metastatic Progression of Prostate Cancer to Bone. Cancer Res. 2017, 77, 74–85. [Google Scholar] [CrossRef] [PubMed]
- Cui, C.; Merritt, R.; Fu, L.; Pan, Z. Targeting calcium signaling in cancer therapy. Acta Pharm. Sin. B 2017, 7, 3–17. [Google Scholar] [CrossRef] [PubMed]
- Flourakis, M.; Prevarskaya, N. Insights into Ca2+ homeostasis of advanced prostate cancer cells. Biochim. Biophys. Acta 2009, 1793, 1105–1109. [Google Scholar] [CrossRef]
- Coffey, K.; Robson, C.N. Regulation of the androgen receptor by post-translational modifications. J. Endocrinol. 2012, 215, 221–237. [Google Scholar] [CrossRef]
- Tummala, R.; Lou, W.; Gao, A.C.; Nadiminty, N. Quercetin Targets hnRNPA1 to Overcome Enzalutamide Resistance in Prostate Cancer Cells. Mol. Cancer 2017, 16, 2770–2779. [Google Scholar] [CrossRef]
- Ko, C.C.; Chen, Y.J.; Chen, C.T.; Liu, Y.C.; Cheng, F.C.; Hsu, K.C.; Chow, L.P. Chemical proteomics identifies heterogeneous nuclear ribonucleoprotein (hnRNP) A1 as the molecular target of quercetin in its anti-cancer effects in PC-3 cells. J. Biol. Chem. 2014, 289, 22078–22089. [Google Scholar] [CrossRef]
- Baniwal, S.K.; Khalid, O.; Gabet, Y.; Shah, R.R.; Purcell, D.J.; Mav, D.; Kohn-Gabet, A.E.; Shi, Y.; Coetzee, G.A.; Frenkel, B. Runx2 transcriptome of prostate cancer cells: Insights into invasiveness and bone metastasis. Mol. Cancer 2010, 9, 258. [Google Scholar] [CrossRef] [PubMed]
- Lin, Y.; Fukuchi, J.; Hiipakka, R.A.; Kokontis, J.M.; Xiang, J. Up-regulation of Bcl-2 is required for the progression of prostate cancer cells from an androgen-dependent to an androgen-independent growth stage. Cell Res. 2007, 17, 531–536. [Google Scholar] [CrossRef] [PubMed]
- Nguyen, D.P.; Li, J.; Tewari, A.K. Inflammation and prostate cancer: The role of interleukin 6 (IL-6). Bju Int. 2014, 113, 986–992. [Google Scholar] [CrossRef] [PubMed]
- Bowen, C.; Bubendorf, L.; Voeller, H.J.; Slack, R.; Willi, N.; Sauter, G.; Gasser, T.C.; Koivisto, P.; Lack, E.E.; Kononen, J.; et al. Loss of NKX3.1 expression in human prostate cancers correlates with tumor progression. Cancer Res. 2000, 60, 6111–6115. [Google Scholar] [PubMed]
- Le Magnen, C.; Virk, R.K.; Dutta, A.; Kim, J.Y.; Panja, S.; Lopez-Bujanda, Z.A.; Califano, A.; Drake, C.G.; Mitrofanova, A.; Abate-Shen, C. Cooperation of loss of NKX3.1 and inflammation in prostate cancer initiation. Dis. Model. Mech. 2018, 11. [Google Scholar] [CrossRef] [PubMed]
- Bhatia-Gaur, R.; Donjacour, A.A.; Sciavolino, P.J.; Kim, M.; Desai, N.; Young, P.; Norton, C.R.; Gridley, T.; Cardiff, R.D.; Cunha, G.R.; et al. Roles for Nkx3.1 in prostate development and cancer. Genes Dev. 1999, 13, 966–977. [Google Scholar] [CrossRef] [PubMed]
- Pio, R.; Jia, Z.; Baron, V.T.; Mercola, D.; UCI NCI SPECS consortium of the Strategic Partners for the Evaluation of Cancer Signatures, Prostate Cancer. Early growth response 3 (Egr3) is highly over-expressed in non-relapsing prostate cancer but not in relapsing prostate cancer. PLoS ONE 2013, 8, e54096. [Google Scholar] [CrossRef]
- Watanabe, M.; Kakiuchi, H.; Kato, H.; Shiraishi, T.; Yatani, R.; Sugimura, T.; Nagao, M. APC gene mutations in human prostate cancer. Jpn J. Clin. Oncol. 1996, 26, 77–81. [Google Scholar] [CrossRef]
- Rawla, P. Epidemiology of Prostate Cancer. World J. Oncol. 2019, 10, 63–89. [Google Scholar] [CrossRef]
- Mintie, C.A.; Musarra, A.K.; Singh, C.K.; Ndiaye, M.A.; Sullivan, R.; Eickhoff, J.C.; Ahmad, N. Protective Effects of Dietary Grape on UVB-Mediated Cutaneous Damages and Skin Tumorigenesis in SKH-1 Mice. Cancers 2020, 12, 1751. [Google Scholar] [CrossRef]
- Mintie, C.A.; Singh, C.K.; Ahmad, N. Whole Fruit Phytochemicals Combating Skin Damage and Carcinogenesis. Transl. Oncol. 2020, 13, 146–156. [Google Scholar] [CrossRef] [PubMed]


| Gene | Quercetin | Resveratrol | QR | Cellular Location 1 | Protein Type 2 | |||
|---|---|---|---|---|---|---|---|---|
| Fold Change | p Value | Fold Change | p Value | Fold Change | p Value | |||
| Apc | 1.05 | 0.2826 | 2.00 | 0.0000 | 1.54 | 0.0027 | N | Enzyme |
| Ar | 2.33 | 0.0073 | 1.19 | 0.2867 | 2.79 | 0.0013 | N | NR |
| Bcl2 | −2.19 | 0.0011 | −1.60 | 0.0206 | −1.19 | 0.4120 | C | Transporter |
| Cav2 | 2.49 | 0.0569 | 1.27 | 0.0741 | 2.23 | 0.0009 | PM | Other |
| Dkk3 | −5.10 | 0.0010 | −7.21 | 0.0008 | −5.90 | 0.0009 | ES | Cytokine |
| Egfr | −1.95 | 0.0678 | −2.67 | 0.0075 | −5.54 | 0.0021 | PM | Kinase |
| Egr3 | −4.82 | 0.0021 | −1.25 | 0.4571 | −1.85 | 0.0522 | N | TR |
| Etv1 | 2.12 | 0.0005 | 1.94 | 0.0004 | 1.25 | 0.0358 | N | TR |
| Hal | −2.01 | 0.0020 | −1.68 | 0.0099 | −1.11 | 0.2460 | C | Enzyme |
| Igf1 | −2.64 | 0.0017 | −3.56 | 0.0008 | −2.08 | 0.0039 | ES | GR |
| Igfbp5 | −1.23 | 0.4209 | 4.48 | 0.0013 | −2.29 | 0.0048 | ES | Other |
| Il6 | −6.37 | 0.0014 | −6.57 | 0.0012 | −6.06 | 0.0023 | ES | Cytokine |
| Lgals4 | 5.55 | 0.0010 | 1.71 | 0.0233 | 1.16 | 0.3417 | ES | Other |
| Loxl1 | −6.62 | 0.0060 | −3.45 | 0.0289 | −1.74 | 0.3113 | ES | Enzyme |
| Msx1 | −3.52 | 0.0003 | −3.89 | 0.0002 | −1.42 | 0.0167 | N | TR |
| Nkx3−1 | 5.34 | 0.1512 | 5.01 | 0.0440 | 1.95 | 0.1645 | N | TR |
| Nrip1 | −4.04 | 0.1678 | −8.78 | 0.0202 | −9.63 | 0.0202 | N | TR |
| Ptgs1 | 2.93 | 0.0128 | 2.44 | 0.0025 | 1.80 | 0.0517 | C | Enzyme |
| Sfrp1 | −4.68 | 0.0010 | −11.30 | 0.0000 | −3.12 | 0.0001 | PM | TmR |
| Slc5a8 | 3.52 | 0.0540 | −10.36 | 0.0019 | 1.13 | 0.6243 | PM | Transporter |
| Tfpi2 | −1.92 | 0.0003 | −3.28 | 0.0001 | −1.28 | 0.1904 | ES | Other |
| Tmprss2 | 6.56 | 0.0435 | 1.94 | 0.0134 | 1.28 | 0.3619 | PM | Peptidase |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Singh, C.K.; Chhabra, G.; Ndiaye, M.A.; Siddiqui, I.A.; Panackal, J.E.; Mintie, C.A.; Ahmad, N. Quercetin–Resveratrol Combination for Prostate Cancer Management in TRAMP Mice. Cancers 2020, 12, 2141. https://doi.org/10.3390/cancers12082141
Singh CK, Chhabra G, Ndiaye MA, Siddiqui IA, Panackal JE, Mintie CA, Ahmad N. Quercetin–Resveratrol Combination for Prostate Cancer Management in TRAMP Mice. Cancers. 2020; 12(8):2141. https://doi.org/10.3390/cancers12082141
Chicago/Turabian StyleSingh, Chandra K., Gagan Chhabra, Mary A. Ndiaye, Imtiaz A. Siddiqui, Jennifer E. Panackal, Charlotte A. Mintie, and Nihal Ahmad. 2020. "Quercetin–Resveratrol Combination for Prostate Cancer Management in TRAMP Mice" Cancers 12, no. 8: 2141. https://doi.org/10.3390/cancers12082141
APA StyleSingh, C. K., Chhabra, G., Ndiaye, M. A., Siddiqui, I. A., Panackal, J. E., Mintie, C. A., & Ahmad, N. (2020). Quercetin–Resveratrol Combination for Prostate Cancer Management in TRAMP Mice. Cancers, 12(8), 2141. https://doi.org/10.3390/cancers12082141






