Discovery of Prognostic Markers for Early-Stage High-Grade Serous Ovarian Cancer by Maldi-Imaging
Abstract
1. Introduction
2. Results
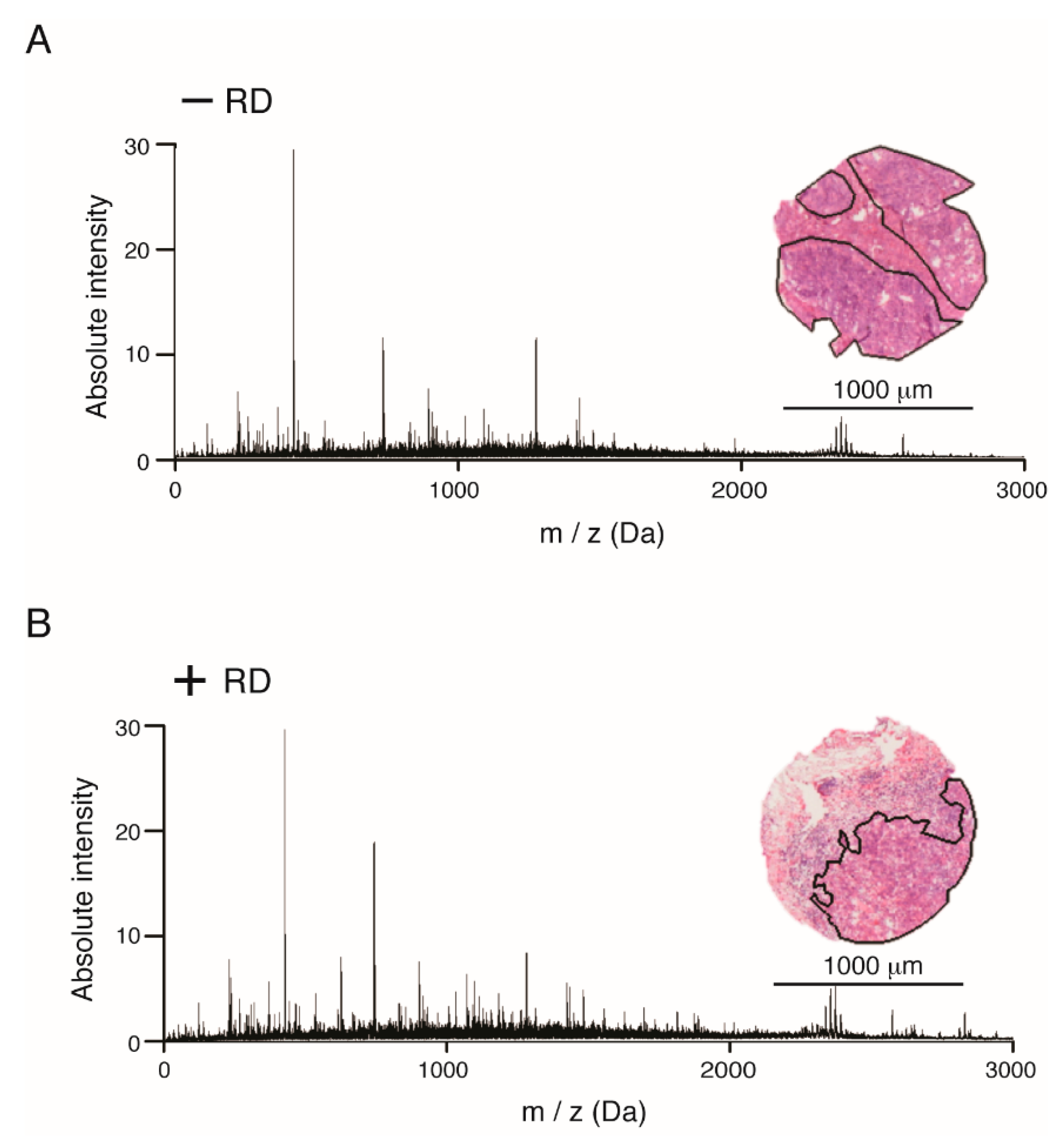
2.1. Accumulation of Proteomics Data by MALDI-IMS
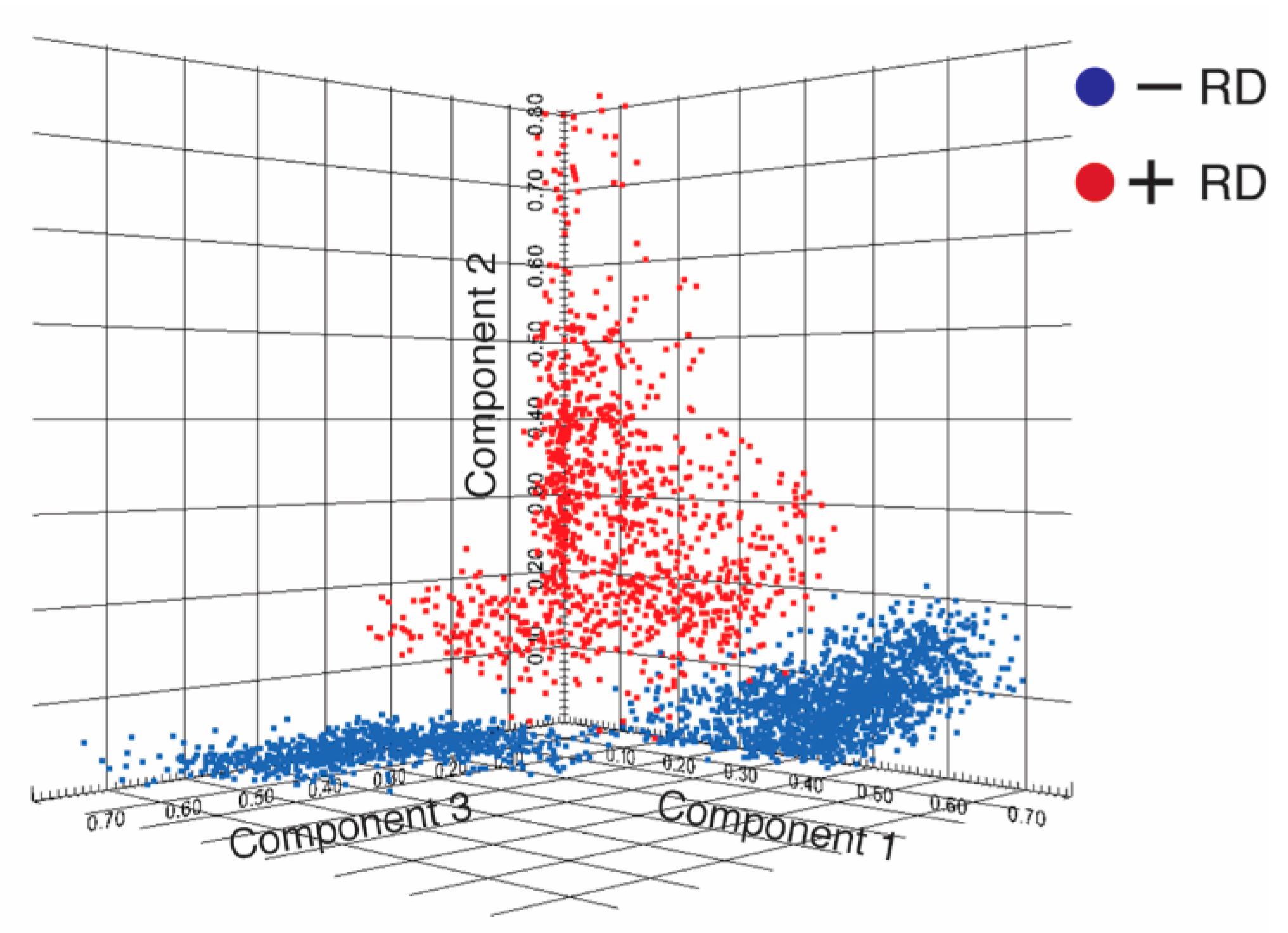
2.2. Discovery of Discriminative Peptide Signatures
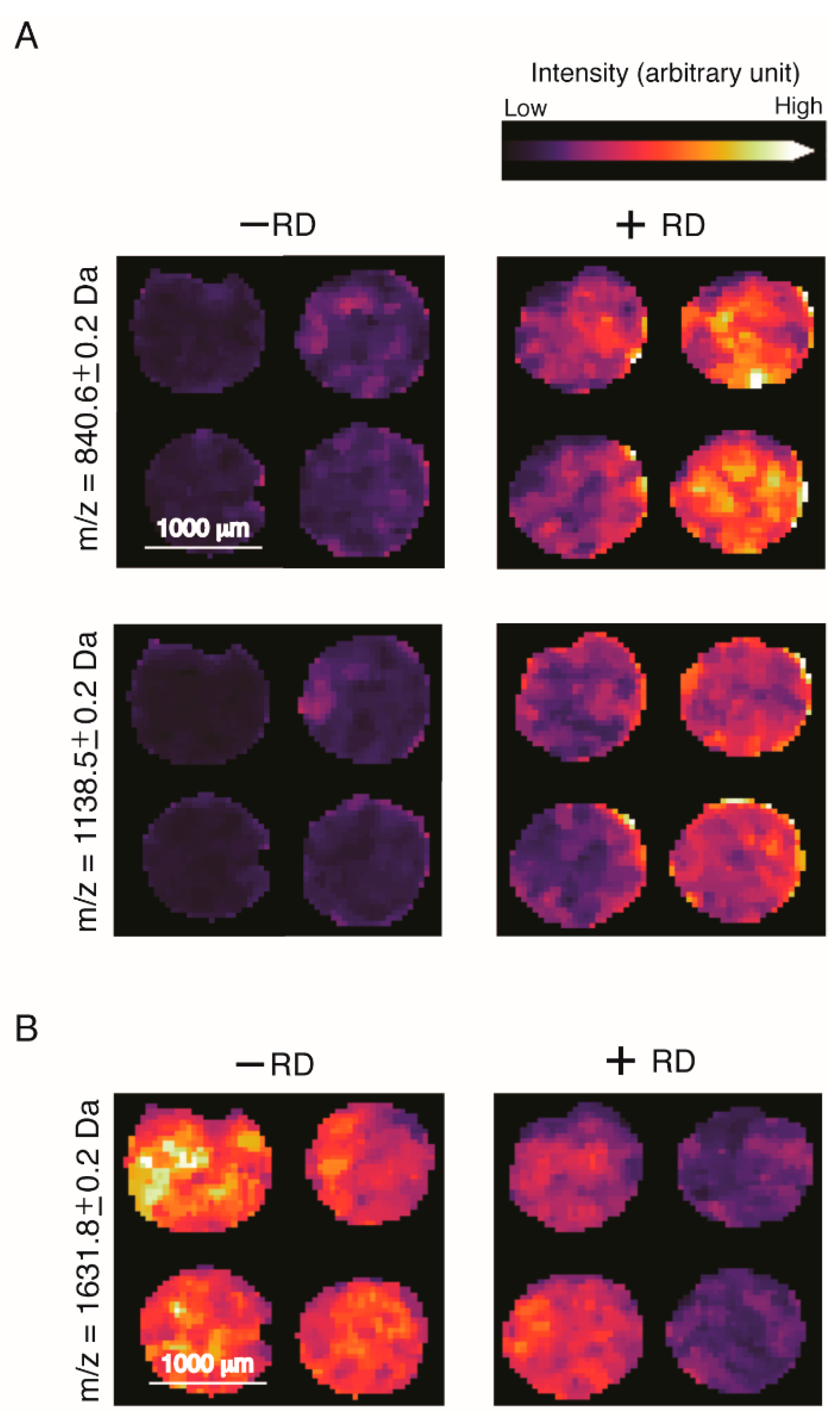
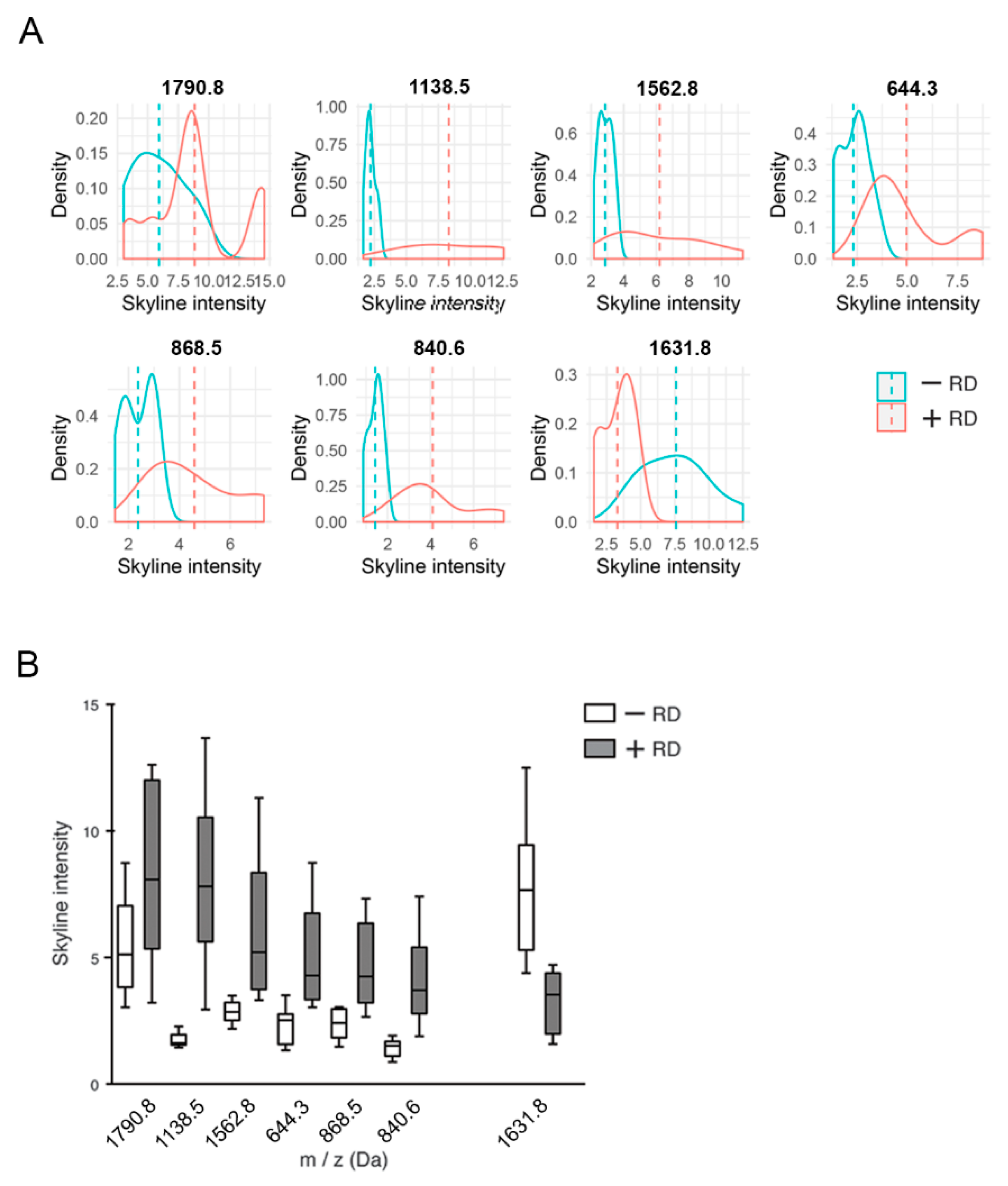
2.3. Determination of Characteristic m/z Values of HGSOC Patients
2.4. Identification of Differentially Expressed Proteins
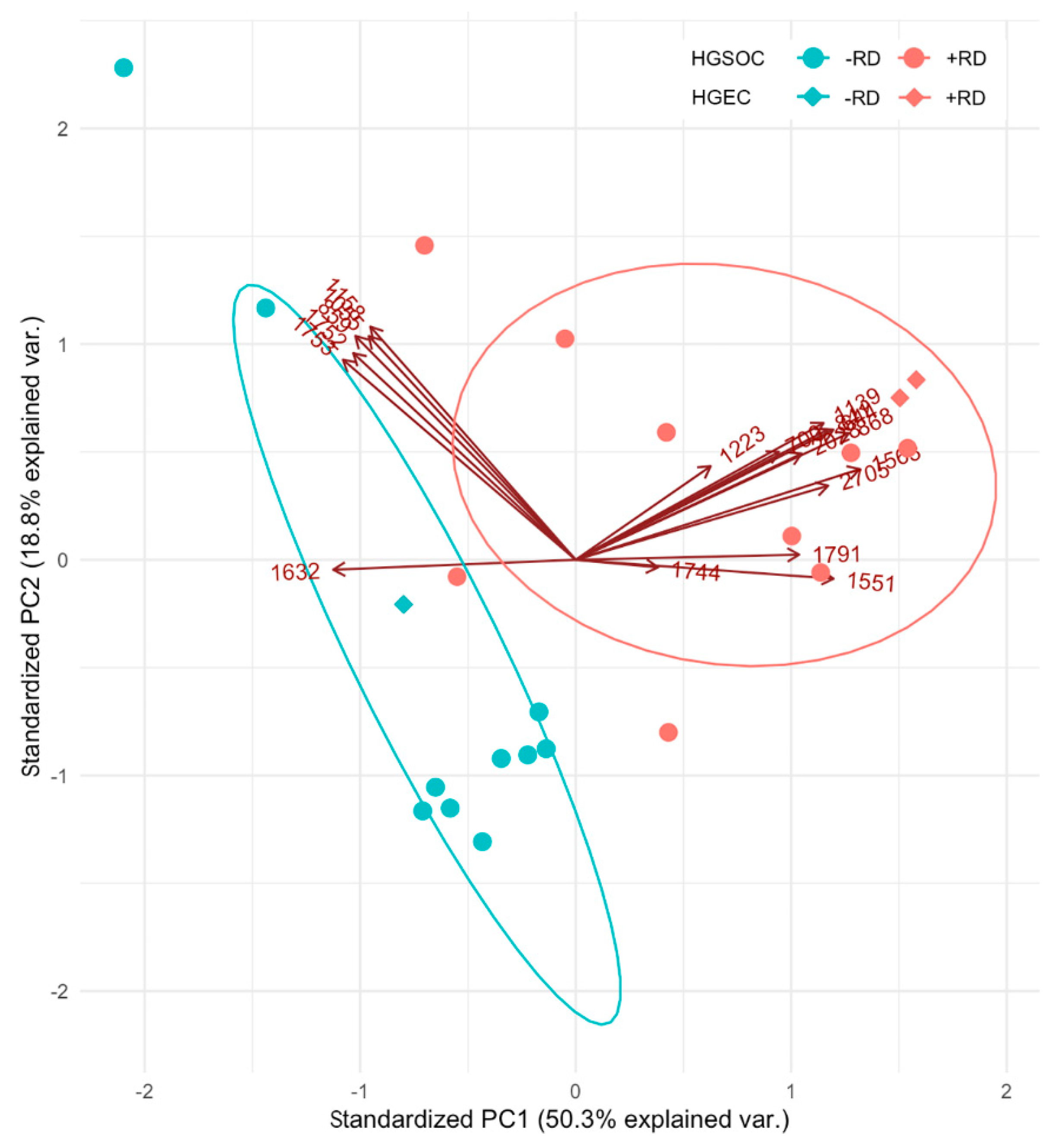
2.5. Relatedness between Patients with RD and between Patients without RD
3. Discussion
4. Materials and Methods
4.1. Clinicopathological Parameters of Patient Cohort
4.2. Procedure of MALDI-Imaging
4.3. MALDI Imaging Analysis
4.4. Data Processing
4.5. Identification of Peptides by “Bottom-Up”-Nhplc Mass Spectrometry
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Vaughan, S.; Coward, J.I.; Bast, R.C., Jr.; Berchuck, A.; Berek, J.S.; Brenton, J.D.; Coukos, G.; Crum, C.C.; Drapkin, R.; Etemadmoghadam, D.; et al. Rethinking ovarian cancer: Recommendations for improving outcomes. Nat. Rev. Cancer 2011, 11, 719–725. [Google Scholar] [CrossRef] [PubMed]
- Siegel, R.L.; Miller, K.D.; Jemal, A. Cancer statistics, 2016. CA A cancer J. Clin. 2016, 66, 7–30. [Google Scholar] [CrossRef] [PubMed]
- Prat, J.; FIGO Committee on Gynecologic Oncology. Staging classification for cancer of the ovary, fallopian tube, and peritoneum. Int. J. Gynaecol. Obstet. 2014, 124, 1–5. [Google Scholar] [CrossRef] [PubMed]
- Ahmed, F.Y.; Wiltshaw, E.; A’Hern, R.P.; Nicol, B.; Shepherd, J.; Blake, P.; Fisher, C.; Gore, M.E. Natural history and prognosis of untreated stage I epithelial ovarian carcinoma. J. Clin. Oncol. 1996, 14, 2968–2975. [Google Scholar] [CrossRef] [PubMed]
- Duggan, M.A.; Anderson, W.F.; Altekruse, S.; Penberthy, L.; Sherman, M.E. The Surveillance, Epidemiology, and End Results (SEER) Program and Pathology: Toward Strengthening the Critical Relationship. Am. J. Surg. Pathol. 2016, 40, e94–e102. [Google Scholar] [CrossRef]
- Hoskins, P.J.; Swenerton, K.D.; Manji, M.; Wong, F.; O’Reilly, S.E.; McMurtrie, E.J.; Le, N.; Acker, B.; Le Richer, J. ‘Moderate-risk’ ovarian cancer (stage I, grade 2; stage II, grade 1 or 2) treated with cisplatin chemotherapy (single agent or combination) and pelvi-abdominal irradiation. Int. J. Gynecol. Cancer 1994, 4, 272–278. [Google Scholar] [CrossRef]
- Chan, J.K.; Tian, C.; Monk, B.J.; Herzog, T.; Kapp, D.S.; Bell, J.; Young, R.C.; Gynecologic Oncology, G. Prognostic factors for high-risk early-stage epithelial ovarian cancer: A Gynecologic Oncology Group study. Cancer 2008, 112, 2202–2210. [Google Scholar] [CrossRef]
- Colombo, N.; Peiretti, M.; Parma, G.; Lapresa, M.; Mancari, R.; Carinelli, S.; Sessa, C.; Castiglione, M.; Group, E.G.W. Newly diagnosed and relapsed epithelial ovarian carcinoma: ESMO Clinical Practice Guidelines for diagnosis, treatment and follow-up. Ann. Oncol. 2010, 21, v23–v30. [Google Scholar] [CrossRef]
- Morice, P.; Denschlag, D.; Rodolakis, A.; Reed, N.; Schneider, A.; Kesic, V.; Colombo, N.; Fertility Task Force of the European Society of Gynecologic Oncology. Recommendations of the Fertility Task Force of the European Society of Gynecologic Oncology about the conservative management of ovarian malignant tumors. Int. J. Gynecol. Cancer 2011, 21, 951–963. [Google Scholar] [CrossRef]
- Fruscio, R.; Corso, S.; Ceppi, L.; Garavaglia, D.; Garbi, A.; Floriani, I.; Franchi, D.; Cantu, M.G.; Bonazzi, C.M.; Milani, R.; et al. Conservative management of early-stage epithelial ovarian cancer: Results of a large retrospective series. Ann. Oncol. 2013, 24, 138–144. [Google Scholar] [CrossRef]
- Lee, C.L.; Kusunoki, S.; Huang, C.Y.; Wu, K.Y.; Lee, P.S.; Huang, K.G. Surgical and survival outcomes of laparoscopic staging surgery for patients with stage I ovarian cancer. Taiwan J. Obstet. Gynecol. 2018, 57, 7–12. [Google Scholar] [CrossRef]
- Wei, W.; Li, N.; Sun, Y.; Li, B.; Xu, L.; Wu, L. Clinical outcome and prognostic factors of patients with early-stage epithelial ovarian cancer. Oncotarget 2017, 8, 23862–23870. [Google Scholar] [CrossRef] [PubMed]
- Leblanc, E.; Querleu, D.; Narducci, F.; Chauvet, M.P.; Chevalier, A.; Lesoin, A.; Vennin, P.; Taieb, S. Surgical staging of early invasive epithelial ovarian tumors. Semin. Surg. Oncol. 2000, 19, 36–41. [Google Scholar] [CrossRef]
- Bolis, G.; Colombo, N.; Pecorelli, S.; Torri, V.; Marsoni, S.; Bonazzi, C.; Chiari, S.; Favalli, G.; Mangili, G.; Presti, M.; et al. Adjuvant treatment for early epithelial ovarian cancer: Results of two randomised clinical trials comparing cisplatin to no further treatment or chromic phosphate (32P). G.I.C.O.G.: Gruppo Interregionale Collaborativo in Ginecologia Oncologica. Ann. Oncol. 1995, 6, 887–893. [Google Scholar] [CrossRef] [PubMed]
- Vergote, I.B.; Vergote-De Vos, L.N.; Abeler, V.M.; Aas, M.; Lindegaard, M.W.; Kjorstad, K.E.; Trope, C.G. Randomized trial comparing cisplatin with radioactive phosphorus or whole-abdomen irradiation as adjuvant treatment of ovarian cancer. Cancer 1992, 69, 741–749. [Google Scholar] [CrossRef]
- Chen, C.A.; Chiang, C.J.; Chen, Y.Y.; You, S.L.; Hsieh, S.F.; Tang, C.H.; Cheng, W.F. Survival benefit of patients with early-stage ovarian carcinoma treated with paclitaxel chemotherapeutic regimens. J. Gynecol. Oncol. 2018, 29, e16. [Google Scholar] [CrossRef]
- Meding, S.; Nitsche, U.; Balluff, B.; Elsner, M.; Rauser, S.; Schone, C.; Nipp, M.; Maak, M.; Feith, M.; Ebert, M.P.; et al. Tumor classification of six common cancer types based on proteomic profiling by MALDI imaging. J. Proteome Res. 2012, 11, 1996–2003. [Google Scholar] [CrossRef]
- Walch, A.; Rauser, S.; Deininger, S.O.; Hofler, H. MALDI imaging mass spectrometry for direct tissue analysis: A new frontier for molecular histology. Histochem. Cell Biol. 2008, 130, 421–434. [Google Scholar] [CrossRef]
- Klein, O.; Strohschein, K.; Nebrich, G.; Oetjen, J.; Trede, D.; Thiele, H.; Alexandrov, T.; Giavalisco, P.; Duda, G.N.; von Roth, P.; et al. MALDI imaging mass spectrometry: Discrimination of pathophysiological regions in traumatized skeletal muscle by characteristic peptide signatures. Proteomics 2014, 14, 2249–2260. [Google Scholar] [CrossRef]
- Klein, O.; Kanter, F.; Kulbe, H.; Jank, P.; Denkert, C.; Nebrich, G.; Schmitt, W.D.; Wu, Z.; Kunze, C.A.; Sehouli, J.; et al. MALDI-Imaging for Classification of Epithelial Ovarian Cancer Histotypes from a Tissue Microarray Using Machine Learning Methods. Proteomics Clin. Appl. 2019, 13, e1700181. [Google Scholar] [CrossRef]
- Longuespee, R.; Boyon, C.; Castellier, C.; Jacquet, A.; Desmons, A.; Kerdraon, O.; Vinatier, D.; Fournier, I.; Day, R.; Salzet, M. The C-terminal fragment of the immunoproteasome PA28S (Reg alpha) as an early diagnosis and tumor-relapse biomarker: Evidence from mass spectrometry profiling. Histochem. Cell Biol. 2012, 138, 141–154. [Google Scholar] [CrossRef] [PubMed]
- Mainini, V.; Angel, P.M.; Magni, F.; Caprioli, R.M. Detergent enhancement of on-tissue protein analysis by matrix-assisted laser desorption/ionization imaging mass spectrometry. Rapid Commun. Mass Spectrom 2011, 25, 199–204. [Google Scholar] [CrossRef] [PubMed]
- Gustafsson, J.O.; Oehler, M.K.; McColl, S.R.; Hoffmann, P. Citric acid antigen retrieval (CAAR) for tryptic peptide imaging directly on archived formalin-fixed paraffin-embedded tissue. J. Proteome Res. 2010, 9, 4315–4328. [Google Scholar] [CrossRef] [PubMed]
- Aichler, M.; Walch, A. MALDI Imaging mass spectrometry: Current frontiers and perspectives in pathology research and practice. Lab. Invest. 2015, 95, 422–431. [Google Scholar] [CrossRef] [PubMed]
- Ly, A.; Longuespee, R.; Casadonte, R.; Wandernoth, P.; Schwamborn, K.; Bollwein, C.; Marsching, C.; Kriegsmann, K.; Hopf, C.; Weichert, W.; et al. Site-to-Site Reproducibility and Spatial Resolution in MALDI-MSI of Peptides from Formalin-Fixed Paraffin-Embedded Samples. Proteomics Clin. Appl. 2019, 13, e1800029. [Google Scholar] [CrossRef]
- Hermann, J.; Noels, H.; Theelen, W.; Lellig, M.; Orth-Alampour, S.; Boor, P.; Jankowski, V.; Jankowski, J. Sample preparation of formalin-fixed paraffin-embedded tissue sections for MALDI-mass spectrometry imaging. Anal. Bioanal. Chem. 2020, 412, 1263–1275. [Google Scholar] [CrossRef]
- Lazova, R.; Seeley, E.H.; Kutzner, H.; Scolyer, R.A.; Scott, G.; Cerroni, L.; Fried, I.; Kozovska, M.E.; Rosenberg, A.S.; Prieto, V.G.; et al. Imaging mass spectrometry assists in the classification of diagnostically challenging atypical Spitzoid neoplasms. J. Am. Acad. Dermatol. 2016, 75, 1176–1186. [Google Scholar] [CrossRef]
- McDonnell, L.A.; Rompp, A.; Balluff, B.; Heeren, R.M.; Albar, J.P.; Andren, P.E.; Corthals, G.L.; Walch, A.; Stoeckli, M. Discussion point: Reporting guidelines for mass spectrometry imaging. Anal. Bioanal. Chem. 2015, 407, 2035–2045. [Google Scholar] [CrossRef]
- Meding, S.; Martin, K.; Gustafsson, O.J.; Eddes, J.S.; Hack, S.; Oehler, M.K.; Hoffmann, P. Tryptic peptide reference data sets for MALDI imaging mass spectrometry on formalin-fixed ovarian cancer tissues. J. Proteome Res. 2013, 12, 308–315. [Google Scholar] [CrossRef]
- Groseclose, M.R.; Massion, P.P.; Chaurand, P.; Caprioli, R.M. High-throughput proteomic analysis of formalin-fixed paraffin-embedded tissue microarrays using MALDI imaging mass spectrometry. Proteomics 2008, 8, 3715–3724. [Google Scholar] [CrossRef]
- Dmello, C.; Srivastava, S.S.; Tiwari, R.; Chaudhari, P.R.; Sawant, S.; Vaidya, M.M. Multifaceted role of keratins in epithelial cell differentiation and transformation. J. Biosci. 2019, 44, 33. [Google Scholar] [CrossRef] [PubMed]
- Keller, B.O.; Sui, J.; Young, A.B.; Whittal, R.M. Interferences and contaminants encountered in modern mass spectrometry. Anal. Chim. Acta 2008, 627, 71–81. [Google Scholar] [CrossRef] [PubMed]
- Kang, S.; Maeng, H.; Kim, B.G.; Qing, G.M.; Choi, Y.P.; Kim, H.Y.; Kim, P.S.; Kim, Y.; Kim, Y.H.; Choi, Y.D.; et al. In situ identification and localization of IGHA2 in the breast tumor microenvironment by mass spectrometry. J. Proteome Res. 2012, 11, 4567–4574. [Google Scholar] [CrossRef] [PubMed]
- Angel, P.M.; Schwamborn, K.; Comte-Walters, S.; Clift, C.L.; Ball, L.E.; Mehta, A.S.; Drake, R.R. Extracellular Matrix Imaging of Breast Tissue Pathologies by MALDI-Imaging Mass Spectrometry. Proteomics Clin. Appl. 2019, 13, e1700152. [Google Scholar] [CrossRef]
- Januchowski, R.; Swierczewska, M.; Sterzynska, K.; Wojtowicz, K.; Nowicki, M.; Zabel, M. Increased Expression of Several Collagen Genes is Associated with Drug Resistance in Ovarian Cancer Cell Lines. J. Cancer 2016, 7, 1295–1310. [Google Scholar] [CrossRef] [PubMed]
- Harrel, F.E. Regression Modeling Strategies; Springer: New York, NY, USA, 2001. [Google Scholar]
- Casadonte, R.; Caprioli, R.M. Proteomic analysis of formalin-fixed paraffin-embedded tissue by MALDI imaging mass spectrometry. Nat. Protoc. 2011, 6, 1695–1709. [Google Scholar] [CrossRef]
- Gustafsson, O.J.; Briggs, M.T.; Condina, M.R.; Winderbaum, L.J.; Pelzing, M.; McColl, S.R.; Everest-Dass, A.V.; Packer, N.H.; Hoffmann, P. MALDI imaging mass spectrometry of N-linked glycans on formalin-fixed paraffin-embedded murine kidney. Anal. Bioanal. Chem. 2015, 407, 2127–2139. [Google Scholar] [CrossRef]
- Alexandrov, T.; Becker, M.; Guntinas-Lichius, O.; Ernst, G.; von Eggeling, F. MALDI-imaging segmentation is a powerful tool for spatial functional proteomic analysis of human larynx carcinoma. J. Cancer Res. Clin. Oncol. 2013, 139, 85–95. [Google Scholar] [CrossRef]
- Alexandrov, T.; Becker, M.; Deininger, S.O.; Ernst, G.; Wehder, L.; Grasmair, M.; von Eggeling, F.; Thiele, H.; Maass, P. Spatial segmentation of imaging mass spectrometry data with edge-preserving image denoising and clustering. J. proteome Res. 2010, 9, 6535–6546. [Google Scholar] [CrossRef]
- Trede, D.; Schiffler, S.; Becker, M.; Wirtz, S.; Steinhorst, K.; Strehlow, J.; Aichler, M.; Kobarg, J.H.; Oetjen, J.; Dyatlov, A.; et al. Exploring three-dimensional matrix-assisted laser desorption/ionization imaging mass spectrometry data: Three-dimensional spatial segmentation of mouse kidney. Anal. Chem. 2012, 84, 6079–6087. [Google Scholar] [CrossRef]
- Hanselmann, M.; Kirchner, M.; Renard, B.Y.; Amstalden, E.R.; Glunde, K.; Heeren, R.M.; Hamprecht, F.A. Concise representation of mass spectrometry images by probabilistic latent semantic analysis. Anal. Chem. 2008, 80, 9649–9658. [Google Scholar] [CrossRef] [PubMed]
- Klerk, L.A.; Dankers, P.Y.; Popa, E.R.; Bosman, A.W.; Sanders, M.E.; Reedquist, K.A.; Heeren, R.M. TOF-secondary ion mass spectrometry imaging of polymeric scaffolds with surrounding tissue after in vivo implantation. Anal. Chem. 2010, 82, 4337–4343. [Google Scholar] [CrossRef] [PubMed]
- Chambers, M.C.; Maclean, B.; Burke, R.; Amodei, D.; Ruderman, D.L.; Neumann, S.; Gatto, L.; Fischer, B.; Pratt, B.; Egertson, J.; et al. A cross-platform toolkit for mass spectrometry and proteomics. Nat. Biotechnol. 2012, 30, 918–920. [Google Scholar] [CrossRef] [PubMed]
- Cillero-Pastor, B.; Heeren, R.M. Matrix-Assisted Laser Desorption Ionization Mass Spectrometry Imaging for Peptide and Protein Analyses: A Critical Review of On-Tissue Digestion. J. proteome res. 2013, 13, 325–335. [Google Scholar] [CrossRef]
| Centroid [m/z] | IMS Mr [m/z] [Da] | Tumor +RD vs −RD (AUC) | LC-MS Mr [Da] | Δ [Da] | Ascension | Protein | HGNC Symbol |
|---|---|---|---|---|---|---|---|
| 2705.026 | 2704.0181 | 0.7547 | 2704.1538 | 0.1358 | K1C9_HUMAN | Keratin, type I cytoskeletal 9 | KRT9 |
| 1791.698 | 1790.6901 | 0.6250 | 1790.7204 | 0.0304 | K1C9_HUMAN | Keratin, type I cytoskeletal 9 | KRT9 |
| 644.336 | 643.3281 | 0.7470 | 643.3653 | 0.0373 | ACTB_HUMAN | Actin, cytoplasmic 1 | ACTB |
| 840.564 | 839.5561 | 0.7407 | 839.4613 | 0.0947 | CO1A2_HUMAN | Collagen alpha-2(I) chain | COL1A2 |
| 868.467 | 867.4591 | 0.7331 | 867.4563 | 0.0028 | CO1A2_HUMAN | Collagen alpha-2(I) chain | COL1A2 |
| 2027.831 | 2026.8231 | 0.7008 | 2026.0093 | 0.8138 | CO1A2_HUMAN | Collagen alpha-2(I) chain | COL1A2 |
| 1562.765 | 1561.7571 | 0.6930 | 1561.7849 | 0.0278 | CO1A2_HUMAN | Collagen alpha-2(I) chain | COL1A2 |
| 1223.417 | 1222.4091 | 0.6262 | 1222.6054 | 0.1964 | CO1A2_HUMAN | Collagen alpha-2(I) chain | COL1A2 |
| 700.444 | 699.4361 | 0.6388 | 699.4643 | 0.0282 | RL37A_HUMAN | 60S ribosomal protein L37a | RPL37A |
| 1790.797 | 1789.7891 | 0.6253 | 1789.8846 | 0.0956 | ACTB_HUMAN | Actin, cytoplasmic 1 | ACTB |
| 1743.691 | 1742.6831 | 0.6055 | 1742.8120 | 0.1290 | H2B1N_HUMAN | Histone H2B type 1-N | HIST1H2BN |
| 1550.764 | 1549.7561 | 0.6016 | 1549.8100 | 0.0540 | ANXA1_HUMAN | Annexin A1 | ANXA1 |
| 858.566 | 857.5581 | 0.3975 | 857.4607 | 0.0974 | CALD1_HUMAN | Caldesmon | CALD1 |
| 1157.708 | 1156.7001 | 0.3782 | 1156.6200 | 0.0800 | APOA1_HUMAN | Apolipoprotein A-I | APOA1 |
| 1631.775 | 1630.7671 | 0.3682 | 1630.8236 | 0.0566 | TBB5_HUMAN | Tubulin beta chain | TUBB |
| 1751.792 | 1750.7841 | 0.3554 | 1750.0353 | 0.7488 | H2B1K_HUMAN | Histone H2B type 1-K | HIST1H2BK |
| 1055.394 | 1054.3861 | 0.3460 | 1054.5196 | 0.1335 | 4_HUMAN | Histone H4 | HIST1H4A |
| 1752.992 | 1751.9841 | 0.3159 | 1751.8551 | 0.1290 | LMNA_HUMAN | Prelamin-A/C | LMNA |
| Patients (−RD) | ||||
| Age | 68 | 60 | 68 | 67 |
| FIGO stage | IA | IC | IA | IC |
| Grade | G3 | G2 | G3 | G3 |
| Presence of ascites | <500 mL | <500 mL | <500 mL | no |
| Number of cycles | 6 | 6 | 4 | 6 |
| Recurrence (months) | NA | NA | NA | NA |
| Patients (+RD) | ||||
| Age | 44 | 52 | 67 | 57 |
| FIGO stage | IIB | IIA | IA | IIA |
| Grade | G3 | G3 | G3 | G3 |
| Presence of ascites | >500 mL | no | No | no |
| Number of cycles | 6 | 9 | 6 | 6 |
| Recurrence (months) | 13 | 12 | 54 | 16 |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Kulbe, H.; Klein, O.; Wu, Z.; Taube, E.T.; Kassuhn, W.; Horst, D.; Darb-Esfahani, S.; Jank, P.; Abobaker, S.; Ringel, F.; et al. Discovery of Prognostic Markers for Early-Stage High-Grade Serous Ovarian Cancer by Maldi-Imaging. Cancers 2020, 12, 2000. https://doi.org/10.3390/cancers12082000
Kulbe H, Klein O, Wu Z, Taube ET, Kassuhn W, Horst D, Darb-Esfahani S, Jank P, Abobaker S, Ringel F, et al. Discovery of Prognostic Markers for Early-Stage High-Grade Serous Ovarian Cancer by Maldi-Imaging. Cancers. 2020; 12(8):2000. https://doi.org/10.3390/cancers12082000
Chicago/Turabian StyleKulbe, Hagen, Oliver Klein, Zhiyang Wu, Eliane T. Taube, Wanja Kassuhn, David Horst, Silvia Darb-Esfahani, Paul Jank, Salem Abobaker, Frauke Ringel, and et al. 2020. "Discovery of Prognostic Markers for Early-Stage High-Grade Serous Ovarian Cancer by Maldi-Imaging" Cancers 12, no. 8: 2000. https://doi.org/10.3390/cancers12082000
APA StyleKulbe, H., Klein, O., Wu, Z., Taube, E. T., Kassuhn, W., Horst, D., Darb-Esfahani, S., Jank, P., Abobaker, S., Ringel, F., du Bois, A., Heitz, F., Sehouli, J., & Braicu, E. I. (2020). Discovery of Prognostic Markers for Early-Stage High-Grade Serous Ovarian Cancer by Maldi-Imaging. Cancers, 12(8), 2000. https://doi.org/10.3390/cancers12082000





