A Multidisciplinary Review of the Roles of Cripto in the Scientific Literature Through a Bibliometric Analysis of its Biological Roles
Abstract
1. Introduction
2. Results
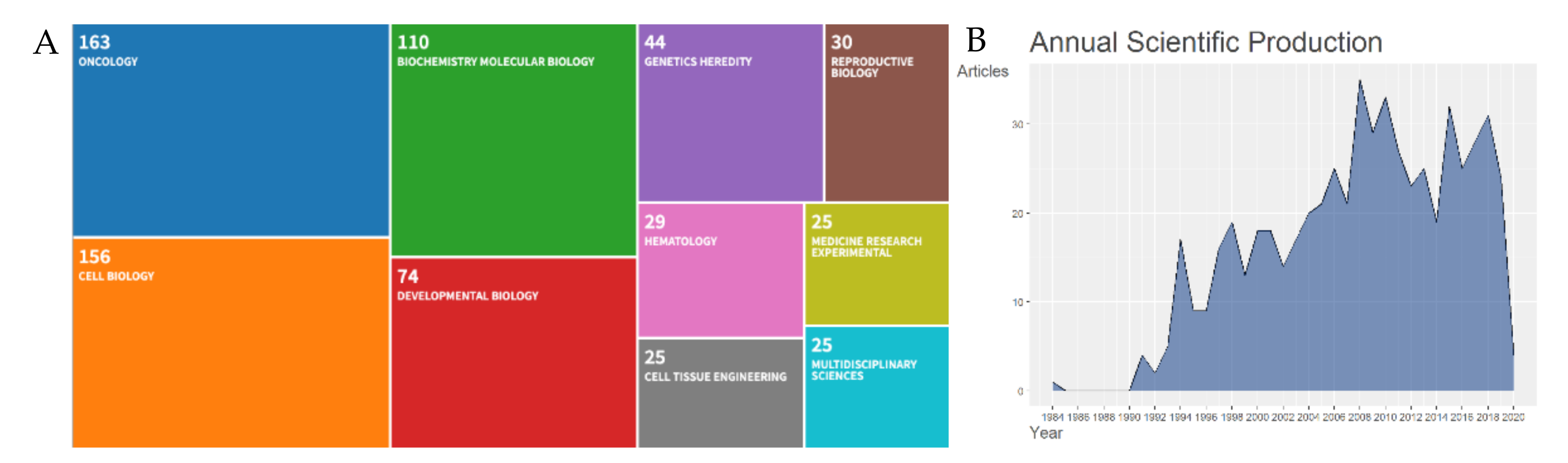
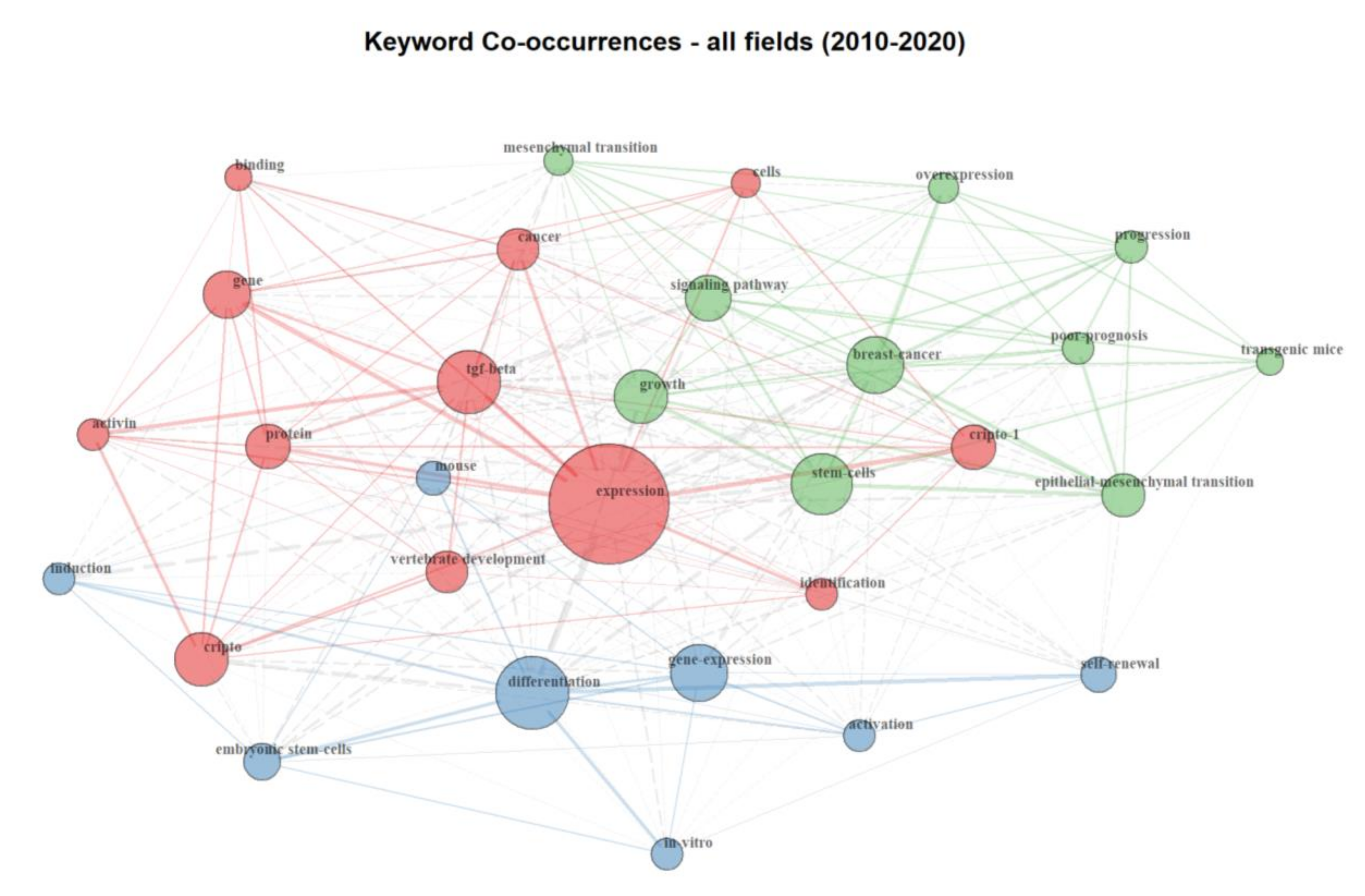
2.1. Bibliometric Analysis
2.2. Cripto in Developmental Biology
2.3. Cripto in Reproductive Biology
2.4. Cripto Signaling in Biochemistry and Molecular Biology
2.5. Cripto in Oncology
2.6. Cripto in Experimental Medical Research: Therapeutic and Translational Applications
3. Materials and Methods
3.1. Search Strategy
3.2. Citation and Publication Data
3.3. Data Analysis
4. Conclusions
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Klauzinska, M.; Castro, N.P.; Rangel, M.C.; Spike, B.T.; Gray, P.C.; Bertolette, D.; Cuttitta, F.; Salomon, D. The multifaceted role of the embryonic gene Cripto-1 in cancer, stem cells and epithelial-mesenchymal transition. Semin. Cancer Biol. 2014, 29, 51–58. [Google Scholar] [CrossRef] [PubMed]
- Zoni, E.; Chen, L.; Karkampouna, S.; Granchi, Z.; Verhoef, E.I.; La Manna, F.; Kelber, J.; Pelger, R.C.M.; Henry, M.D.; Snaar-Jagalska, E.; et al. CRIPTO and its signaling partner GRP78 drive the metastatic phenotype in human osteotropic prostate cancer. Oncogene 2017, 36, 4739–4749. [Google Scholar] [CrossRef]
- Qi, C.F.; Liscia, D.S.; Normanno, N.; Merlo, G.; Johnson, G.R.; Gullick, W.J.; Ciardiello, F.; Saeki, T.; Brandt, R.; Kim, N. Expression of transforming growth factor alpha, amphiregulin and cripto-1 in human breast carcinomas. Br. J. Cancer 1994, 69, 903–910. [Google Scholar] [CrossRef]
- Bianco, C.; Strizzi, L.; Mancino, M.; Rehman, A.; Hamada, S.; Watanabe, K.; De Luca, A.; Jones, B.; Balogh, G.; Russo, J.; et al. Identification of cripto-1 as a novel serologic marker for breast and colon cancer. Clin. Cancer Res. 2006, 12, 5158–5164. [Google Scholar] [CrossRef]
- Karkampouna, S.; van der Helm, D.; Gray, P.C.; Chen, L.; Klima, I.; Grosjean, J.; Burgmans, M.C.; Farina-Sarasqueta, A.; Snaar-Jagalska, E.B.; Stroka, D.M.; et al. CRIPTO promotes an aggressive tumour phenotype and resistance to treatment in hepatocellular carcinoma. J. Pathol. 2018, 245, 297–310. [Google Scholar] [CrossRef]
- Pilgaard, L.; Mortensen, J.H.; Henriksen, M.; Olesen, P.; Sørensen, P.; Laursen, R.; Vyberg, M.; Agger, R.; Zachar, V.; Moos, T.; et al. Cripto-1 expression in glioblastoma multiforme. Brain Pathol. Zurich Switz. 2014, 24, 360–370. [Google Scholar] [CrossRef] [PubMed]
- Xu, C.H.; Wang, Y.; Qian, L.H.; Yu, L.K.; Zhang, X.W.; Wang, Q.B. Serum Cripto-1 is a novel biomarker for non-small cell lung cancer diagnosis and prognosis. Clin. Respir. J. 2017, 11, 765–771. [Google Scholar] [CrossRef]
- Gray, P.C.; Harrison, C.A.; Vale, W. Cripto forms a complex with activin and type II activin receptors and can block activin signaling. Proc. Natl. Acad. Sci. USA 2003, 100, 5193–5198. [Google Scholar] [CrossRef] [PubMed]
- Adkins, H.B.; Bianco, C.; Schiffer, S.G.; Rayhorn, P.; Zafari, M.; Cheung, A.E.; Orozco, O.; Olson, D.; Luca, A.D.; Chen, L.L.; et al. Antibody blockade of the Cripto CFC domain suppresses tumor cell growth in vivo. J. Clin. Investig. 2003, 112, 575–587. [Google Scholar] [CrossRef]
- Gray, P.C.; Shani, G.; Aung, K.; Kelber, J.; Vale, W. Cripto Binds Transforming Growth Factor β (TGF-β) and Inhibits TGF-β Signaling. Mol. Cell. Biol. 2006, 26, 9268–9278. [Google Scholar] [CrossRef]
- Kelber, J.A.; Panopoulos, A.D.; Shani, G.; Booker, E.C.; Belmonte, J.C.; Vale, W.W.; Gray, P.C. Blockade of Cripto binding to cell surface GRP78 inhibits oncogenic Cripto signaling via MAPK/PI3K and Smad2/3 pathways. Oncogene 2009, 28, 2324–2336. [Google Scholar] [CrossRef]
- Gray, P.C.; Vale, W. Cripto/GRP78 modulation of the TGF-β pathway in development and oncogenesis. FEBS Lett. 2012, 586, 1836–1845. [Google Scholar] [CrossRef] [PubMed]
- Shen, M.M. Nodal signaling: developmental roles and regulation. Dev. Camb. Engl. 2007, 134, 1023–1034. [Google Scholar] [CrossRef] [PubMed]
- Cheng, S.K.; Olale, F.; Bennett, J.T.; Brivanlou, A.H.; Schier, A.F. EGF-CFC proteins are essential coreceptors for the TGF-beta signals Vg1 and GDF1. Genes Dev. 2003, 17, 31–36. [Google Scholar] [CrossRef]
- Chen, C.; Ware, S.M.; Sato, A.; Houston-Hawkins, D.E.; Habas, R.; Matzuk, M.M.; Shen, M.M.; Brown, C.W. The Vg1-related protein Gdf3 acts in a Nodal signaling pathway in the pre-gastrulation mouse embryo. Dev. Camb. Engl. 2006, 133, 319–329. [Google Scholar] [CrossRef]
- Shani, G.; Fischer, W.H.; Justice, N.J.; Kelber, J.A.; Vale, W.; Gray, P.C. GRP78 and Cripto form a complex at the cell surface and collaborate to inhibit transforming growth factor beta signaling and enhance cell growth. Mol. Cell. Biol. 2008, 28, 666–677. [Google Scholar] [CrossRef] [PubMed]
- Nagaoka, T.; Karasawa, H.; Turbyville, T.; Rangel, M.-C.; Castro, N.P.; Gonzales, M.; Baker, A.; Seno, M.; Lockett, S.; Greer, Y.E.; et al. Cripto-1 enhances the canonical Wnt/β-catenin signaling pathway by binding to LRP5 and LRP6 co-receptors. Cell. Signal. 2013, 25, 178–189. [Google Scholar] [CrossRef]
- Gao, Y.; Ge, L.; Shi, S.; Sun, Y.; Liu, M.; Wang, B.; Shang, Y.; Wu, J.; Tian, J. Global trends and future prospects of e-waste research: a bibliometric analysis. Environ. Sci. Pollut. Res. Int. 2019, 26, 17809–17820. [Google Scholar] [CrossRef]
- He, L.; Fang, H.; Chen, C.; Wu, Y.; Wang, Y.; Ge, H.; Wang, L.; Wan, Y.; He, H. Metastatic castration-resistant prostate cancer: Academic insights and perspectives through bibliometric analysis. Medicine (Baltimore) 2020, 99, e19760. [Google Scholar] [CrossRef]
- Lou, J.; Tian, S.-J.; Niu, S.-M.; Kang, X.-Q.; Lian, H.-X.; Zhang, L.-X.; Zhang, J.-J. Coronavirus disease 2019: A bibliometric analysis and review. Eur. Rev. Med. Pharmacol. Sci. 2020, 24, 3411–3421. [Google Scholar] [CrossRef]
- Van Eck, N.J.; Waltman, L. Citation-based clustering of publications using CitNetExplorer and VOSviewer. Scientometrics 2017, 111, 1053–1070. [Google Scholar] [CrossRef]
- Synnestvedt, M.B.; Chen, C.; Holmes, J.H. CiteSpace II: visualization and knowledge discovery in bibliographic databases. AMIA Annu. Symp. Proc. AMIA Symp. 2005, 724–728. [Google Scholar]
- Van Eck, N.J.; Waltman, L. Software survey: VOSviewer, a computer program for bibliometric mapping. Scientometrics 2010, 84, 523–538. [Google Scholar] [CrossRef] [PubMed]
- Aria, M.; Cuccurullo, C. bibliometrix: An R-tool for comprehensive science mapping analysis. J. Informetr. 2017, 11, 959–975. [Google Scholar] [CrossRef]
- Ciccodicola, A.; Dono, R.; Obici, S.; Simeone, A.; Zollo, M.; Persico, M.G. Molecular characterization of a gene of the “EGF family” expressed in undifferentiated human NTERA2 teratocarcinoma cells. EMBO J. 1989, 8, 1987–1991. [Google Scholar] [CrossRef]
- Ciardiello, F.; Dono, R.; Kim, N.; Persico, M.G.; Salomon, D.S. Expression of cripto, a novel gene of the epidermal growth factor gene family, leads to in vitro transformation of a normal mouse mammary epithelial cell line. Cancer Res. 1991, 51, 1051–1054. [Google Scholar]
- Brennan, J.; Lu, C.C.; Norris, D.P.; Rodriguez, T.A.; Beddington, R.S.; Robertson, E.J. Nodal signalling in the epiblast patterns the early mouse embryo. Nature 2001, 411, 965–969. [Google Scholar] [CrossRef]
- Guzman-Ayala, M.; Ben-Haim, N.; Beck, S.; Constam, D.B. Nodal protein processing and fibroblast growth factor 4 synergize to maintain a trophoblast stem cell microenvironment. Proc. Natl. Acad. Sci. USA 2004, 101, 15656–15660. [Google Scholar] [CrossRef] [PubMed]
- Mesnard, D. Nodal specifies embryonic visceral endoderm and sustains pluripotent cells in the epiblast before overt axial patterning. Development 2006, dev.02413. [Google Scholar] [CrossRef]
- Blakeley, P.; Fogarty, N.M.E.; del Valle, I.; Wamaitha, S.E.; Hu, T.X.; Elder, K.; Snell, P.; Christie, L.; Robson, P.; Niakan, K.K. Defining the three cell lineages of the human blastocyst by single-cell RNA-seq. Dev. Camb. Engl. 2015, 142, 3151–3165. [Google Scholar] [CrossRef]
- Schier, A.F. Nodal signaling in vertebrate development. Annu. Rev. Cell Dev. Biol. 2003, 19, 589–621. [Google Scholar] [CrossRef] [PubMed]
- Schier, A.F.; Shen, M.M. Nodal signalling in vertebrate development. Nature 2000, 403, 385–389. [Google Scholar] [CrossRef]
- Whitman, M. Nodal signaling in early vertebrate embryos: themes and variations. Dev. Cell 2001, 1, 605–617. [Google Scholar] [CrossRef]
- Fiorenzano, A.; Pascale, E.; D’Aniello, C.; Acampora, D.; Bassalert, C.; Russo, F.; Andolfi, G.; Biffoni, M.; Francescangeli, F.; Zeuner, A.; et al. Cripto is essential to capture mouse epiblast stem cell and human embryonic stem cell pluripotency. Nat. Commun. 2016, 7, 12589. [Google Scholar] [CrossRef]
- Hall, V.J.; Hyttel, P. Breaking down pluripotency in the porcine embryo reveals both a premature and reticent stem cell state in the inner cell mass and unique expression profiles of the naive and primed stem cell states. Stem Cells Dev. 2014, 23, 2030–2045. [Google Scholar] [CrossRef]
- Van Leeuwen, J.; Berg, D.K.; Pfeffer, P.L. Morphological and Gene Expression Changes in Cattle Embryos from Hatched Blastocyst to Early Gastrulation Stages after Transfer of In Vitro Produced Embryos. PLoS ONE 2015, 10, e0129787. [Google Scholar] [CrossRef]
- Bianco, C.; Cotten, C.; Lonardo, E.; Strizzi, L.; Baraty, C.; Mancino, M.; Gonzales, M.; Watanabe, K.; Nagaoka, T.; Berry, C.; et al. Cripto-1 is required for hypoxia to induce cardiac differentiation of mouse embryonic stem cells. Am. J. Pathol. 2009, 175, 2146–2158. [Google Scholar] [CrossRef]
- Kruithof-de Julio, M.; Alvarez, M.J.; Galli, A.; Chu, J.; Price, S.M.; Califano, A.; Shen, M.M. Regulation of extra-embryonic endoderm stem cell differentiation by Nodal and Cripto signaling. Dev. Camb. Engl. 2011, 138, 3885–3895. [Google Scholar] [CrossRef]
- Takaoka, K.; Nishimura, H.; Hamada, H. Both Nodal signalling and stochasticity select for prospective distal visceral endoderm in mouse embryos. Nat. Commun. 2017, 8, 1492. [Google Scholar] [CrossRef]
- Chu, J.; Shen, M.M. Functional redundancy of EGF-CFC genes in epiblast and extraembryonic patterning during early mouse embryogenesis. Dev. Biol. 2010, 342, 63–73. [Google Scholar] [CrossRef] [PubMed]
- Kimura, C.; Shen, M.M.; Takeda, N.; Aizawa, S.; Matsuo, I. Complementary functions of Otx2 and Cripto in initial patterning of mouse epiblast. Dev. Biol. 2001, 235, 12–32. [Google Scholar] [CrossRef] [PubMed]
- Conlon, F.L.; Lyons, K.M.; Takaesu, N.; Barth, K.S.; Kispert, A.; Herrmann, B.; Robertson, E.J. A primary requirement for nodal in the formation and maintenance of the primitive streak in the mouse. Dev. Camb. Engl. 1994, 120, 1919–1928. [Google Scholar]
- Liguori, G.L.; Echevarría, D.; Improta, R.; Signore, M.; Adamson, E.; Martínez, S.; Persico, M.G. Anterior neural plate regionalization in cripto null mutant mouse embryos in the absence of node and primitive streak. Dev. Biol. 2003, 264, 537–549. [Google Scholar] [CrossRef]
- Perea-Gomez, A.; Vella, F.D.J.; Shawlot, W.; Oulad-Abdelghani, M.; Chazaud, C.; Meno, C.; Pfister, V.; Chen, L.; Robertson, E.; Hamada, H.; et al. Nodal antagonists in the anterior visceral endoderm prevent the formation of multiple primitive streaks. Dev. Cell 2002, 3, 745–756. [Google Scholar] [CrossRef]
- Rivera-Pérez, J.A.; Magnuson, T. Primitive streak formation in mice is preceded by localized activation of Brachyury and Wnt3. Dev. Biol. 2005, 288, 363–371. [Google Scholar] [CrossRef] [PubMed]
- Ding, J.; Yang, L.; Yan, Y.T.; Chen, A.; Desai, N.; Wynshaw-Boris, A.; Shen, M.M. Cripto is required for correct orientation of the anterior-posterior axis in the mouse embryo. Nature 1998, 395, 702–707. [Google Scholar] [CrossRef]
- Varlet, I.; Collignon, J.; Norris, D.P.; Robertson, E.J. Nodal signaling and axis formation in the mouse. Cold Spring Harb. Symp. Quant. Biol. 1997, 62, 105–113. [Google Scholar]
- Jin, J.-Z.; Zhu, Y.; Warner, D.; Ding, J. Analysis of extraembryonic mesodermal structure formation in the absence of morphological primitive streak. Dev. Growth Differ. 2016, 58, 522–529. [Google Scholar] [CrossRef]
- Roessler, E.; Ouspenskaia, M.V.; Karkera, J.D.; Vélez, J.I.; Kantipong, A.; Lacbawan, F.; Bowers, P.; Belmont, J.W.; Towbin, J.A.; Goldmuntz, E.; et al. Reduced NODAL signaling strength via mutation of several pathway members including FOXH1 is linked to human heart defects and holoprosencephaly. Am. J. Hum. Genet. 2008, 83, 18–29. [Google Scholar] [CrossRef]
- Lin, X.; Zhao, W.; Jia, J.; Lin, T.; Xiao, G.; Wang, S.; Lin, X.; Liu, Y.; Chen, L.; Qin, Y.; et al. Ectopic expression of Cripto-1 in transgenic mouse embryos causes hemorrhages, fatal cardiac defects and embryonic lethality. Sci. Rep. 2016, 6, 34501. [Google Scholar] [CrossRef]
- Osório, L.; Wu, X.; Wang, L.; Jiang, Z.; Neideck, C.; Sheng, G.; Zhou, Z. ISM1 regulates NODAL signaling and asymmetric organ morphogenesis during development. J. Cell Biol. 2019, 218, 2388–2402. [Google Scholar] [CrossRef]
- Lee, G.-H.; Fujita, M.; Takaoka, K.; Murakami, Y.; Fujihara, Y.; Kanzawa, N.; Murakami, K.-I.; Kajikawa, E.; Takada, Y.; Saito, K.; et al. A GPI processing phospholipase A2, PGAP6, modulates Nodal signaling in embryos by shedding CRIPTO. J. Cell Biol. 2016, 215, 705–718. [Google Scholar] [CrossRef] [PubMed]
- Farina, A.; D’Aniello, C.; Severino, V.; Hochstrasser, D.F.; Parente, A.; Minchiotti, G.; Chambery, A. Temporal proteomic profiling of embryonic stem cell secretome during cardiac and neural differentiation. Proteomics 2011, 11, 3972–3982. [Google Scholar] [CrossRef]
- Feldman, B.; Gates, M.A.; Egan, E.S.; Dougan, S.T.; Rennebeck, G.; Sirotkin, H.I.; Schier, A.F.; Talbot, W.S. Zebrafish organizer development and germ-layer formation require nodal-related signals. Nature 1998, 395, 181–185. [Google Scholar] [CrossRef] [PubMed]
- Schier, A.F.; Talbot, W.S. Nodal signaling and the zebrafish organizer. Int. J. Dev. Biol. 2001, 45, 289–297. [Google Scholar] [PubMed]
- Shah, S.B.; Skromne, I.; Hume, C.R.; Kessler, D.S.; Lee, K.J.; Stern, C.D.; Dodd, J. Misexpression of chick Vg1 in the marginal zone induces primitive streak formation. Dev. Camb. Engl. 1997, 124, 5127–5138. [Google Scholar]
- Sirotkin, H.I.; Dougan, S.T.; Schier, A.F.; Talbot, W.S. bozozok and squint act in parallel to specify dorsal mesoderm and anterior neuroectoderm in zebrafish. Dev. Camb. Engl. 2000, 127, 2583–2592. [Google Scholar]
- Levin, M.; Johnson, R.L.; Stern, C.D.; Kuehn, M.; Tabin, C. A molecular pathway determining left-right asymmetry in chick embryogenesis. Cell 1995, 82, 803–814. [Google Scholar] [CrossRef]
- Whitman, M.; Mercola, M. TGF-beta superfamily signaling and left-right asymmetry. Sci. STKE Signal Transduct. Knowl. Environ. 2001, 2001, re1. [Google Scholar] [CrossRef]
- Fleming, B.M.; Yelin, R.; James, R.G.; Schultheiss, T.M. A role for Vg1/Nodal signaling in specification of the intermediate mesoderm. Dev. Camb. Engl. 2013, 140, 1819–1829. [Google Scholar] [CrossRef]
- Porokh, V.; Vaňhara, P.; Bárta, T.; Jurečková, L.; Bohačiaková, D.; Pospíšilová, V.; Mináriková, D.; Holubcová, Z.; Pelková, V.; Souček, K.; et al. Soluble Cripto-1 Induces Accumulation of Supernumerary Centrosomes and Formation of Aberrant Mitoses in Human Embryonic Stem Cells. Stem Cells Dev. 2018, 27, 1077–1084. [Google Scholar] [CrossRef] [PubMed]
- Hamdi, M.; Sánchez-Calabuig, M.J.; Rodríguez-Alonso, B.; Bagés Arnal, S.; Roussi, K.; Sturmey, R.; Gutiérrez-Adán, A.; Lonergan, P.; Rizos, D. Gene expression and metabolic response of bovine oviduct epithelial cells to the early embryo. Reprod. Camb. Engl. 2019, 158, 85–94. [Google Scholar] [CrossRef] [PubMed]
- Clemente, M.; Lopez-Vidriero, I.; O’Gaora, P.; Mehta, J.P.; Forde, N.; Gutierrez-Adan, A.; Lonergan, P.; Rizos, D. Transcriptome changes at the initiation of elongation in the bovine conceptus. Biol. Reprod. 2011, 85, 285–295. [Google Scholar] [CrossRef]
- Papageorgiou, I.; Nicholls, P.K.; Wang, F.; Lackmann, M.; Makanji, Y.; Salamonsen, L.A.; Robertson, D.M.; Harrison, C.A. Expression of nodal signalling components in cycling human endometrium and in endometrial cancer. Reprod. Biol. Endocrinol. RBE 2009, 7, 122. [Google Scholar] [CrossRef]
- Carrarelli, P.; Yen, C.-F.; Arcuri, F.; Funghi, L.; Tosti, C.; Wang, T.-H.; Huang, J.S.; Petraglia, F. Myostatin, follistatin and activin type II receptors are highly expressed in adenomyosis. Fertil. Steril. 2015, 104, 744–752. [Google Scholar] [CrossRef]
- Cruz, C.D.; Del Puerto, H.L.; Rocha, A.L.L.; Cavallo, I.K.; Clarizia, A.D.; Petraglia, F.; Reis, F.M. Expression of Nodal, Cripto, SMAD3, phosphorylated SMAD3, and SMAD4 in the proliferative endometrium of women with endometriosis. Reprod. Sci. Thousand Oaks Calif 2015, 22, 527–533. [Google Scholar] [CrossRef]
- Banz, C.; Ungethuem, U.; Kuban, R.-J.; Diedrich, K.; Lengyel, E.; Hornung, D. The molecular signature of endometriosis-associated endometrioid ovarian cancer differs significantly from endometriosis-independent endometrioid ovarian cancer. Fertil. Steril. 2010, 94, 1212–1217. [Google Scholar] [CrossRef]
- Torricelli, M.; Voltolini, C.; Novembri, R.; Bocchi, C.; Di Tommaso, M.; Severi, F.M.; Petraglia, F. Activin A and its regulatory molecules in placenta and fetal membranes of women with preterm premature rupture of the membranes associated with acute chorioamnionitis. Am. J. Reprod. Immunol. 2012, 68, 392–399. [Google Scholar] [CrossRef] [PubMed]
- Bandeira, C.L.; Urban Borbely, A.; Pulcineli Vieira Francisco, R.; Schultz, R.; Zugaib, M.; Bevilacqua, E. Tumorigenic factor CRIPTO-1 is immunolocalized in extravillous cytotrophoblast in placenta creta. BioMed Res. Int. 2014, 2014, 892856. [Google Scholar] [CrossRef]
- Kelber, J.A.; Shani, G.; Booker, E.C.; Vale, W.W.; Gray, P.C. Cripto is a noncompetitive activin antagonist that forms analogous signaling complexes with activin and nodal. J. Biol. Chem. 2008, 283, 4490–4500. [Google Scholar] [CrossRef]
- Ciarmela, P.; Bloise, E.; Gray, P.C.; Carrarelli, P.; Islam, M.S.; De Pascalis, F.; Severi, F.M.; Vale, W.; Castellucci, M.; Petraglia, F. Activin-A and Myostatin Response and Steroid Regulation in Human Myometrium: Disruption of Their Signalling in Uterine Fibroid. J. Clin. Endocrinol. Metab. 2011, 96, 755–765. [Google Scholar] [CrossRef]
- Kouznetsova, V.L.; Hu, H.; Teigen, K.; Zanetti, M.; Tsigelny, I.F. Cripto stabilizes GRP78 on the cell membrane. Protein Sci. Publ. Protein Soc. 2018, 27, 653–661. [Google Scholar] [CrossRef] [PubMed]
- Hamada, S.; Watanabe, K.; Hirota, M.; Bianco, C.; Strizzi, L.; Mancino, M.; Gonzales, M.; Salomon, D.S. beta-Catenin/TCF/LEF regulate expression of the short form human Cripto-1. Biochem. Biophys. Res. Commun. 2007, 355, 240–244. [Google Scholar] [CrossRef]
- Mancino, M.; Strizzi, L.; Wechselberger, C.; Watanabe, K.; Gonzales, M.; Hamada, S.; Normanno, N.; Salomon, D.S.; Bianco, C. Regulation of human Cripto-1 gene expression by TGF-beta1 and BMP-4 in embryonal and colon cancer cells. J. Cell. Physiol. 2008, 215, 192–203. [Google Scholar] [CrossRef]
- Pilli, V.S.; Gupta, K.; Kotha, B.P.; Aradhyam, G.K. Snail-mediated Cripto-1 repression regulates the cell cycle and epithelial-mesenchymal transition-related gene expression. FEBS Lett. 2015, 589, 1249–1256. [Google Scholar] [CrossRef][Green Version]
- Bianco, C.; Castro, N.P.; Baraty, C.; Rollman, K.; Held, N.; Rangel, M.C.; Karasawa, H.; Gonzales, M.; Strizzi, L.; Salomon, D.S. Regulation of human Cripto-1 expression by nuclear receptors and DNA promoter methylation in human embryonal and breast cancer cells. J. Cell. Physiol. 2013, 228, 1174–1188. [Google Scholar] [CrossRef] [PubMed]
- Su, R.; Cao, S.; Ma, J.; Liu, Y.; Liu, X.; Zheng, J.; Chen, J.; Liu, L.; Cai, H.; Li, Z.; et al. Knockdown of SOX2OT inhibits the malignant biological behaviors of glioblastoma stem cells via up-regulating the expression of miR-194-5p and miR-122. Mol. Cancer 2017, 16, 171. [Google Scholar] [CrossRef]
- Loying, P.; Manhas, J.; Sen, S.; Bose, B. Autoregulation and heterogeneity in expression of human Cripto-1. PLoS ONE 2015, 10, e0116748. [Google Scholar] [CrossRef][Green Version]
- Chen, F.; Hou, S.; Fan, H.; Liu, Y. MiR-15a-16 represses Cripto and inhibits NSCLC cell progression. Mol. Cell. Biochem. 2014, 391, 11–19. [Google Scholar] [CrossRef]
- Sun, G.; Yan, S.-S.; Shi, L.; Wan, Z.-Q.; Jiang, N.; Fu, L.-S.; Li, M.; Guo, J. MicroRNA-15b suppresses the growth and invasion of glioma cells through targeted inhibition of cripto-1 expression. Mol. Med. Rep. 2016, 13, 4897–4903. [Google Scholar] [CrossRef] [PubMed]
- Lee, S.I.; Jeon, M.; Kim, J.S.; Jeon, I.-S.; Byun, S.J. The gga-let-7 family post-transcriptionally regulates TGFBR1 and LIN28B during the differentiation process in early chick development: R EGULATORY R OLE OF gga-let-7 F AMILY IN E ARLY C HICK D EVELOPMENT. Mol. Reprod. Dev. 2015, 82, 967–975. [Google Scholar] [CrossRef] [PubMed]
- Hoover, M.; Runa, F.; Booker, E.; Diedrich, J.K.; Duell, E.; Williams, B.; Arellano-Garcia, C.; Uhlendorf, T.; La Kim, S.; Fischer, W.; et al. Identification of myosin II as a cripto binding protein and regulator of cripto function in stem cells and tissue regeneration. Biochem. Biophys. Res. Commun. 2019, 509, 69–75. [Google Scholar] [CrossRef]
- Guardiola, O.; Lafuste, P.; Brunelli, S.; Iaconis, S.; Touvier, T.; Mourikis, P.; De Bock, K.; Lonardo, E.; Andolfi, G.; Bouché, A.; et al. Cripto regulates skeletal muscle regeneration and modulates satellite cell determination by antagonizing myostatin. Proc. Natl. Acad. Sci. USA 2012, 109, E3231–E3240. [Google Scholar] [CrossRef]
- Kemaladewi, D.U.; de Gorter, D.J.J.; Aartsma-Rus, A.; van Ommen, G.-J.; ten Dijke, P.; ’t Hoen, P.A.C.; Hoogaars, W.M. Cell-type specific regulation of myostatin signaling. FASEB J. 2012, 26, 1462–1472. [Google Scholar] [CrossRef] [PubMed]
- Iavarone, F.; Guardiola, O.; Scagliola, A.; Andolfi, G.; Esposito, F.; Serrano, A.; Perdiguero, E.; Brunelli, S.; Muñoz-Cánoves, P.; Minchiotti, G. Cripto shapes macrophage plasticity and restricts EndMT in injured and diseased skeletal muscle. EMBO Rep. 2020, 21, e49075. [Google Scholar] [CrossRef]
- De Castro, N.P.; Rangel, M.C.; Nagaoka, T.; Salomon, D.S.; Bianco, C. Cripto-1: an embryonic gene that promotes tumorigenesis. Future Oncol. Lond. Engl. 2010, 6, 1127–1142. [Google Scholar] [CrossRef] [PubMed]
- Tysnes, B.B.; Satran, H.A.; Mork, S.J.; Margaryan, N.V.; Eide, G.E.; Petersen, K.; Strizzi, L.; Hendrix, M.J.C. Age-Dependent Association between Protein Expression of the Embryonic Stem Cell Marker Cripto-1 and Survival of Glioblastoma Patients. Transl. Oncol. 2013, 6, 732–741. [Google Scholar] [CrossRef]
- De Luca, A.; Lamura, L.; Strizzi, L.; Roma, C.; D’Antonio, A.; Margaryan, N.; Pirozzi, G.; Hsu, M.-Y.; Botti, G.; Mari, E.; et al. Expression and functional role of CRIPTO-1 in cutaneous melanoma. Br. J. Cancer 2011, 105, 1030–1038. [Google Scholar] [CrossRef]
- Yoon, H.-J.; Hong, J.-S.; Shin, W.-J.; Lee, Y.-J.; Hong, K.-O.; Lee, J.-I.; Hong, S.-P.; Hong, S.-D. The role of Cripto-1 in the tumorigenesis and progression of oral squamous cell carcinoma. Oral Oncol. 2011, 47, 1023–1031. [Google Scholar] [CrossRef]
- Lo, R.C.-L.; Leung, C.O.-N.; Chan, K.K.-S.; Ho, D.W.-H.; Wong, C.-M.; Lee, T.K.-W.; Ng, I.O.-L. Cripto-1 contributes to stemness in hepatocellular carcinoma by stabilizing Dishevelled-3 and activating Wnt/β-catenin pathway. Cell Death Differ. 2018, 25, 1426–1441. [Google Scholar] [CrossRef]
- Sun, C.; Sun, L.; Jiang, K.; Gao, D.-M.; Kang, X.-N.; Wang, C.; Zhang, S.; Huang, S.; Qin, X.; Li, Y.; et al. NANOG promotes liver cancer cell invasion by inducing epithelial-mesenchymal transition through NODAL/SMAD3 signaling pathway. Int. J. Biochem. Cell Biol. 2013, 45, 1099–1108. [Google Scholar] [CrossRef]
- Wang, J.-H.; Wei, W.; Xu, J.; Guo, Z.-X.; Xiao, C.-Z.; Zhang, Y.-F.; Jian, P.-E.; Wu, X.-L.; Shi, M.; Guo, R.-P. Elevated expression of Cripto-1 correlates with poor prognosis in hepatocellular carcinoma. Oncotarget 2015, 6, 35116–35128. [Google Scholar] [CrossRef][Green Version]
- Strizzi, L.; Bianco, C.; Normanno, N.; Seno, M.; Wechselberger, C.; Wallace-Jones, B.; Khan, N.I.; Hirota, M.; Sun, Y.; Sanicola, M.; et al. Epithelial mesenchymal transition is a characteristic of hyperplasias and tumors in mammary gland from MMTV-Cripto-1 transgenic mice. J. Cell. Physiol. 2004, 201, 266–276. [Google Scholar] [CrossRef] [PubMed]
- Witt, K.; Ligtenberg, M.A.; Conti, L.; Lanzardo, S.; Ruiu, R.; Wallmann, T.; Tufvesson-Stiller, H.; Chambers, B.J.; Rolny, C.; Lladser, A.; et al. Cripto-1 Plasmid DNA Vaccination Targets Metastasis and Cancer Stem Cells in Murine Mammary Carcinoma. Cancer Immunol. Res. 2018, 6, 1417–1425. [Google Scholar] [CrossRef]
- Castro, N.P.; Fedorova-Abrams, N.D.; Merchant, A.S.; Rangel, M.C.; Nagaoka, T.; Karasawa, H.; Klauzinska, M.; Hewitt, S.M.; Biswas, K.; Sharan, S.K.; et al. Cripto-1 as a novel therapeutic target for triple negative breast cancer. Oncotarget 2015, 6, 11910–11929. [Google Scholar] [CrossRef]
- Ligtenberg, M.A.; Witt, K.; Galvez-Cancino, F.; Sette, A.; Lundqvist, A.; Lladser, A.; Kiessling, R. Cripto-1 vaccination elicits protective immunity against metastatic melanoma. Oncoimmunology 2016, 5, e1128613. [Google Scholar] [CrossRef][Green Version]
- Zhang, H.; Zhang, B.; Gao, L.; Zhang, L.; Zhu, K.; Cheng, R.; Wang, C. Clinical significance of cripto-1 expression in lung adenocarcinoma. Oncotarget 2017, 8, 79087–79098. [Google Scholar] [CrossRef]
- Park, K.-S.; Raffeld, M.; Moon, Y.W.; Xi, L.; Bianco, C.; Pham, T.; Lee, L.C.; Mitsudomi, T.; Yatabe, Y.; Okamoto, I.; et al. CRIPTO1 expression in EGFR-mutant NSCLC elicits intrinsic EGFR-inhibitor resistance. J. Clin. Investig. 2014, 124, 3003–3015. [Google Scholar] [CrossRef] [PubMed]
- Watanabe, K.; Meyer, M.J.; Strizzi, L.; Lee, J.M.; Gonzales, M.; Bianco, C.; Nagaoka, T.; Farid, S.S.; Margaryan, N.; Hendrix, M.J.C.; et al. Cripto-1 is a cell surface marker for a tumorigenic, undifferentiated subpopulation in human embryonal carcinoma cells. Stem Cells Dayt. Ohio 2010, 28, 1303–1314. [Google Scholar] [CrossRef]
- Castro, N.P.; Rangel, M.C.; Merchant, A.S.; MacKinnon, G.; Cuttitta, F.; Salomon, D.S.; Kim, Y.S. Sulforaphane Suppresses the Growth of Triple-negative Breast Cancer Stem-like Cells In vitro and In vivo. Cancer Prev. Res. Phila. Pa 2019, 12, 147–158. [Google Scholar] [CrossRef] [PubMed]
- Cocciadiferro, L.; Miceli, V.; Kang, K.-S.; Polito, L.M.; Trosko, J.E.; Carruba, G. Profiling cancer stem cells in androgen-responsive and refractory human prostate tumor cell lines. Ann. N. Y. Acad. Sci. 2009, 1155, 257–262. [Google Scholar] [CrossRef]
- Liu, Q.; Cui, X.; Yu, X.; Bian, B.-S.-J.; Qian, F.; Hu, X.-G.; Ji, C.-D.; Yang, L.; Ren, Y.; Cui, W.; et al. Cripto-1 acts as a functional marker of cancer stem-like cells and predicts prognosis of the patients in esophageal squamous cell carcinoma. Mol. Cancer 2017, 16, 81. [Google Scholar] [CrossRef] [PubMed]
- Strizzi, L.; Hardy, K.M.; Margaryan, N.V.; Hillman, D.W.; Seftor, E.A.; Chen, B.; Geiger, X.J.; Thompson, E.A.; Lingle, W.L.; Andorfer, C.A.; et al. Potential for the embryonic morphogen Nodal as a prognostic and predictive biomarker in breast cancer. Breast Cancer Res. BCR 2012, 14, R75. [Google Scholar] [CrossRef] [PubMed]
- Gong, Y.P.; Yarrow, P.M.; Carmalt, H.L.; Kwun, S.Y.; Kennedy, C.W.; Lin, B.P.C.; Xing, P.X.; Gillett, D.J. Overexpression of Cripto and its prognostic significance in breast cancer: A study with long-term survival. Eur. J. Surg. Oncol. 2007, 33, 438–443. [Google Scholar] [CrossRef] [PubMed]
- Quail, D.F.; Zhang, G.; Walsh, L.A.; Siegers, G.M.; Dieters-Castator, D.Z.; Findlay, S.D.; Broughton, H.; Putman, D.M.; Hess, D.A.; Postovit, L.-M. Embryonic morphogen nodal promotes breast cancer growth and progression. PLoS ONE 2012, 7, e48237. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Yu, L.; Harms, P.W.; Pouryazdanparast, P.; Kim, D.S.; Ma, L.; Fullen, D.R. Expression of the embryonic morphogen Nodal in cutaneous melanocytic lesions. Mod. Pathol. 2010, 23, 1209–1214. [Google Scholar] [CrossRef]
- Strizzi, L.; Margaryan, N.V.; Gerami, P.; Haghighat, Z.; Harms, P.W.; Madonna, G.; Botti, G.; Ascierto, P.A.; Hendrix, M.J.C. Translational significance of Nodal, Cripto-1 and Notch4 in adult nevi. Oncol. Lett. 2016, 12, 1349–1354. [Google Scholar] [CrossRef]
- Kalyan, A.; Carneiro, B.A.; Chandra, S.; Kaplan, J.; Chae, Y.K.; Matsangou, M.; Hendrix, M.J.C.; Giles, F. Nodal Signaling as a Developmental Therapeutics Target in Oncology. Mol. Cancer Ther. 2017, 16, 787–792. [Google Scholar] [CrossRef]
- Strizzi, L.; Hardy, K.M.; Kirschmann, D.A.; Ahrlund-Richter, L.; Hendrix, M.J.C. Nodal expression and detection in cancer: experience and challenges. Cancer Res. 2012, 72, 1915–1920. [Google Scholar] [CrossRef]
- Aykul, S.; Ni, W.; Mutatu, W.; Martinez-Hackert, E. Human Cerberus prevents nodal-receptor binding, inhibits nodal signaling, and suppresses nodal-mediated phenotypes. PLoS ONE 2015, 10, e0114954. [Google Scholar] [CrossRef]
- Giorgio, E.; Liguoro, A.; D’Orsi, L.; Mancinelli, S.; Barbieri, A.; Palma, G.; Arra, C.; Liguori, G.L. Cripto haploinsufficiency affects in vivo colon tumor development. Int. J. Oncol. 2014, 45, 31–40. [Google Scholar] [CrossRef] [PubMed]
- Costa, A.L.; Moreira-Barbosa, C.; Lobo, J.; Vilela-Salgueiro, B.; Cantante, M.; Guimarães, R.; Lopes, P.; Braga, I.; Oliveira, J.; Antunes, L.; et al. DNA methylation profiling as a tool for testicular germ cell tumors subtyping. Epigenomics 2018, 10, 1511–1523. [Google Scholar] [CrossRef] [PubMed]
- Ruggiero, D.; Nappo, S.; Nutile, T.; Sorice, R.; Talotta, F.; Giorgio, E.; Bellenguez, C.; Leutenegger, A.-L.; Liguori, G.L.; Ciullo, M. Genetic variants modulating CRIPTO serum levels identified by genome-wide association study in Cilento isolates. PLoS Genet. 2015, 11, e1004976. [Google Scholar] [CrossRef][Green Version]
- Hass, H.G.; Vogel, U.; Scheurlen, M.; Jobst, J. Use of Gene Expression Analysis for Discrimination of Primary and Secondary Adenocarcinoma of the Liver. Oncology 2018, 95, 211–219. [Google Scholar] [CrossRef] [PubMed]
- Ledda, M.; Megiorni, F.; Pozzi, D.; Giuliani, L.; D’Emilia, E.; Piccirillo, S.; Mattei, C.; Grimaldi, S.; Lisi, A. Non ionising radiation as a non chemical strategy in regenerative medicine: Ca(2+)-ICR “In Vitro” effect on neuronal differentiation and tumorigenicity modulation in NT2 cells. PLoS ONE 2013, 8, e61535. [Google Scholar] [CrossRef] [PubMed]
- Spike, B.T.; Kelber, J.A.; Booker, E.; Kalathur, M.; Rodewald, R.; Lipianskaya, J.; La, J.; He, M.; Wright, T.; Klemke, R.; et al. CRIPTO/GRP78 signaling maintains fetal and adult mammary stem cells ex vivo. Stem Cell Rep. 2014, 2, 427–439. [Google Scholar] [CrossRef][Green Version]
- Miharada, K.; Karlsson, G.; Rehn, M.; Rörby, E.; Siva, K.; Cammenga, J.; Karlsson, S. Cripto regulates hematopoietic stem cells as a hypoxic-niche-related factor through cell surface receptor GRP78. Cell Stem Cell 2011, 9, 330–344. [Google Scholar] [CrossRef]
- Gao, L.R.; Zhang, N.K.; Ding, Q.A.; Chen, H.Y.; Hu, X.; Jiang, S.; Li, T.C.; Chen, Y.; Wang, Z.G.; Ye, Y.; et al. Common expression of stemness molecular markers and early cardiac transcription factors in human Wharton’s jelly-derived mesenchymal stem cells and embryonic stem cells. Cell Transplant. 2013, 22, 1883–1900. [Google Scholar] [CrossRef]
- Hosseinpour, B.; Bakhtiarizadeh, M.R.; Khosravi, P.; Ebrahimie, E. Predicting distinct organization of transcription factor binding sites on the promoter regions: a new genome-based approach to expand human embryonic stem cell regulatory network. Gene 2013, 531, 212–219. [Google Scholar] [CrossRef]
- Teshigawara, R.; Hirano, K.; Nagata, S.; Huang, D.; Tada, T. Visualization of sequential conversion of human intermediately reprogrammed stem cells into iPS cells. Genes Cells Devoted Mol. Cell. Mech. 2019, 24, 667–673. [Google Scholar] [CrossRef]
- Miharada, K.; Karlsson, G.; Rehn, M.; Rörby, E.; Siva, K.; Cammenga, J.; Karlsson, S. Hematopoietic stem cells are regulated by Cripto, as an intermediary of HIF-1α in the hypoxic bone marrow niche. Ann. N. Y. Acad. Sci. 2012, 1266, 55–62. [Google Scholar] [CrossRef]
- Lonardo, E.; Parish, C.L.; Ponticelli, S.; Marasco, D.; Ribeiro, D.; Ruvo, M.; De Falco, S.; Arenas, E.; Minchiotti, G. A small synthetic cripto blocking Peptide improves neural induction, dopaminergic differentiation, and functional integration of mouse embryonic stem cells in a rat model of Parkinson’s disease. Stem Cells Dayt. Ohio 2010, 28, 1326–1337. [Google Scholar] [CrossRef]
- Kang, R.; Zhou, Y.; Tan, S.; Zhou, G.; Aagaard, L.; Xie, L.; Bünger, C.; Bolund, L.; Luo, Y. Mesenchymal stem cells derived from human induced pluripotent stem cells retain adequate osteogenicity and chondrogenicity but less adipogenicity. Stem Cell Res. Ther. 2015, 6, 144. [Google Scholar] [CrossRef] [PubMed]
- Focà, G.; Iaccarino, E.; Focà, A.; Sanguigno, L.; Untiveros, G.; Cuevas-Nunez, M.; Strizzi, L.; Leonardi, A.; Ruvo, M.; Sandomenico, A. Development of conformational antibodies targeting Cripto-1 with neutralizing effects in vitro. Biochimie 2019, 158, 246–256. [Google Scholar] [CrossRef]
- Gudbergsson, J.M.; Duroux, M. An evaluation of different Cripto-1 antibodies and their variable results. J. Cell. Biochem. 2020, 121, 545–556. [Google Scholar] [CrossRef]
- Adam, A.; Ras, R.; Bhattu, A.S.; Raman, A.; Perera, M. “Researching the Research” in Prostate Cancer: A Comparative Bibliometric Analysis of the Top 100 Cited Articles in the Field of Prostate Cancer. Curr. Urol. 2017, 11, 26–35. [Google Scholar] [CrossRef]
- R Core Team: R: A Language and Environment for Statistical Computing; R Foundation for Statistical Computing: Vienna, Austria, 2019; Available online: http://www.R-project.org/ (accessed on 9 April 2020).
- R Studio. R Studio: Integrated Development for R; R Studio: Boston, MA, USA, 2016. [Google Scholar]
- Koseoglu, M.A. Mapping the institutional collaboration network of strategic management research: 1980–2014. Scientometrics 2016, 109, 203–226. [Google Scholar] [CrossRef]
- Elango, B.; Rajendran, P. Authorship trends and collaboration pattern in the marine sciences literature: a scientometric study. Int. J. Inf. Dissemin. Technol. 2012, 2, 166–169. [Google Scholar]
- Kumar, S.; Kumar, S. Collaboration in research productivity in oil seed research institutes of India. In Proceedings of the Fourth International Conference on Webometrics, Informetrics and Scientometrics & Ninth COLLNET Meeting, Proceedings of WIS 2008, Berlin, Germany, 1 August 2008. [Google Scholar]
- Radhakrishnan, S.; Erbis, S.; Isaacs, J.A.; Kamarthi, S. Novel keyword co-occurrence network-based methods to foster systematic reviews of scientific literature. PLoS ONE 2017, 12, e0172778. [Google Scholar] [CrossRef]
- Fellows, I. Word Clouds. 2018. Available online: https://cran.r-project.org/web/packages/wordcloud/index.html (accessed on 9 April 2020).
- Falagas, M.E.; Pitsouni, E.I.; Malietzis, G.A.; Pappas, G. Comparison of PubMed, Scopus, Web of Science, and Google Scholar: strengths and weaknesses. FASEB J. 2008, 22, 338–342. [Google Scholar] [CrossRef] [PubMed]
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Rodrigues Sousa, E.; Zoni, E.; Karkampouna, S.; La Manna, F.; Gray, P.C.; De Menna, M.; Kruithof-de Julio, M. A Multidisciplinary Review of the Roles of Cripto in the Scientific Literature Through a Bibliometric Analysis of its Biological Roles. Cancers 2020, 12, 1480. https://doi.org/10.3390/cancers12061480
Rodrigues Sousa E, Zoni E, Karkampouna S, La Manna F, Gray PC, De Menna M, Kruithof-de Julio M. A Multidisciplinary Review of the Roles of Cripto in the Scientific Literature Through a Bibliometric Analysis of its Biological Roles. Cancers. 2020; 12(6):1480. https://doi.org/10.3390/cancers12061480
Chicago/Turabian StyleRodrigues Sousa, Elisa, Eugenio Zoni, Sofia Karkampouna, Federico La Manna, Peter C. Gray, Marta De Menna, and Marianna Kruithof-de Julio. 2020. "A Multidisciplinary Review of the Roles of Cripto in the Scientific Literature Through a Bibliometric Analysis of its Biological Roles" Cancers 12, no. 6: 1480. https://doi.org/10.3390/cancers12061480
APA StyleRodrigues Sousa, E., Zoni, E., Karkampouna, S., La Manna, F., Gray, P. C., De Menna, M., & Kruithof-de Julio, M. (2020). A Multidisciplinary Review of the Roles of Cripto in the Scientific Literature Through a Bibliometric Analysis of its Biological Roles. Cancers, 12(6), 1480. https://doi.org/10.3390/cancers12061480




