Potential Onco-Suppressive Role of miR122 and miR144 in Uveal Melanoma through ADAM10 and C-Met Inhibition
Abstract
1. Introduction
2. Results
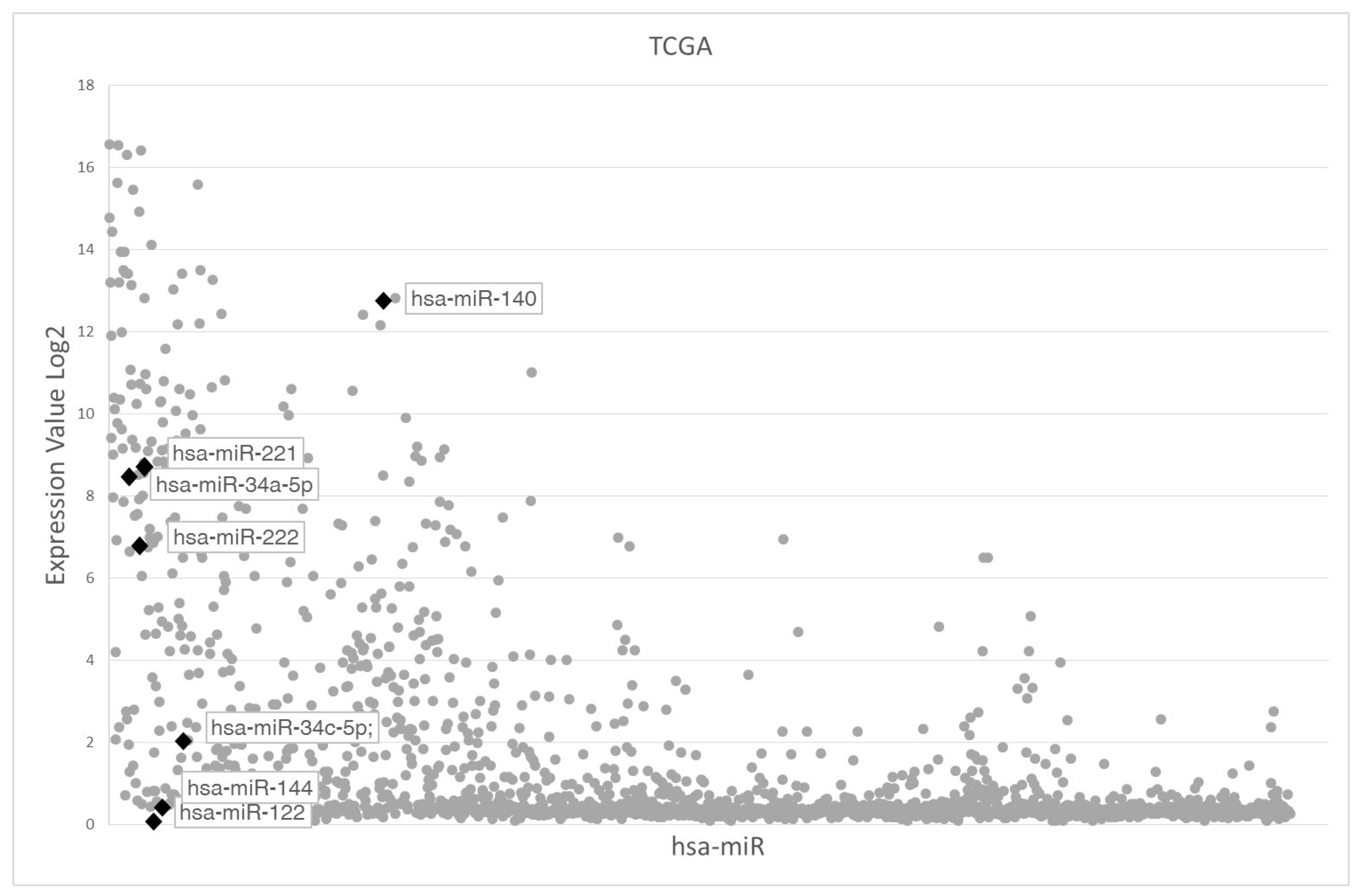
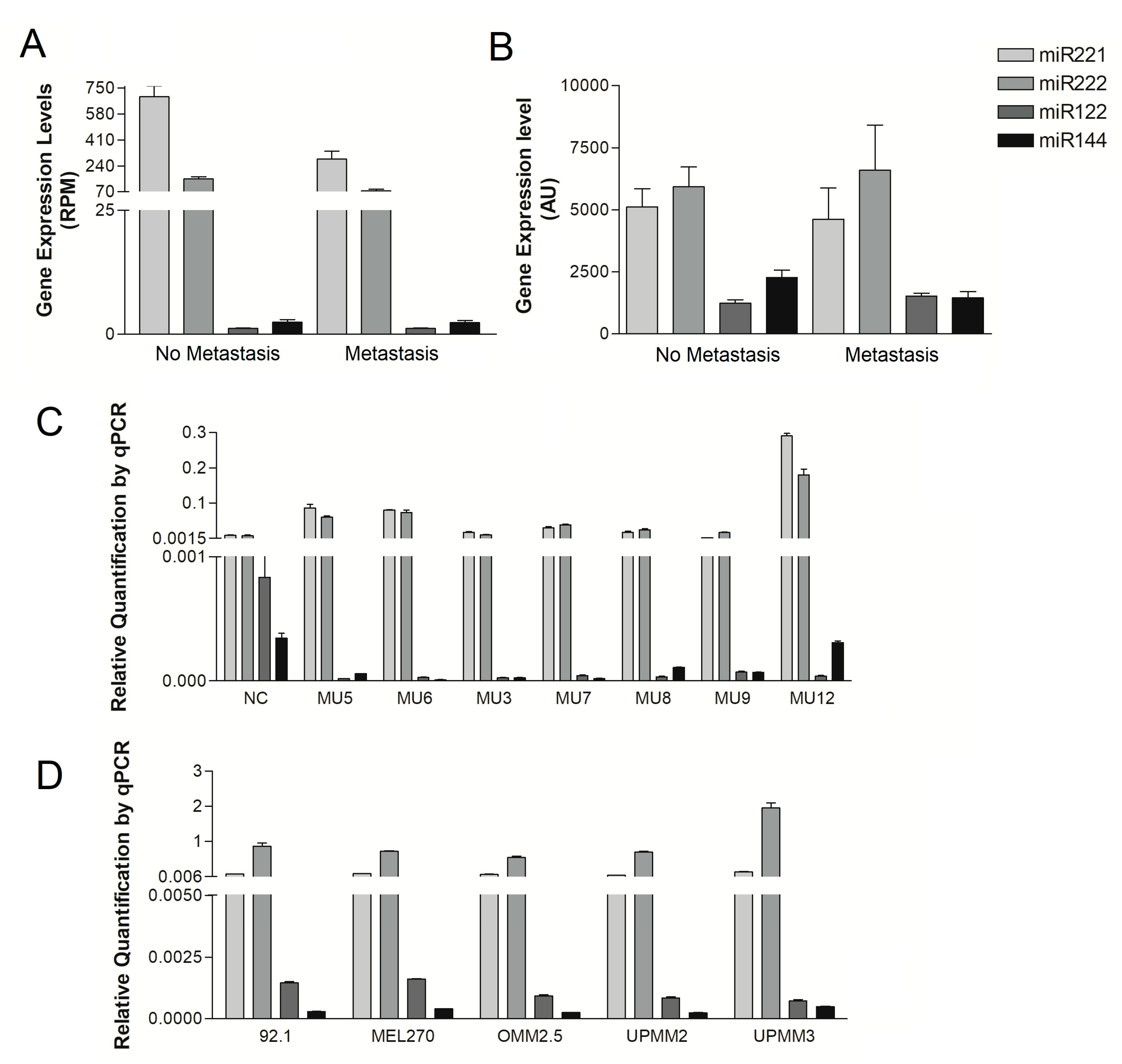
2.1. Analysis of miR122 and miR144 Expression in Primary UM Tumors and UM Cell Lines
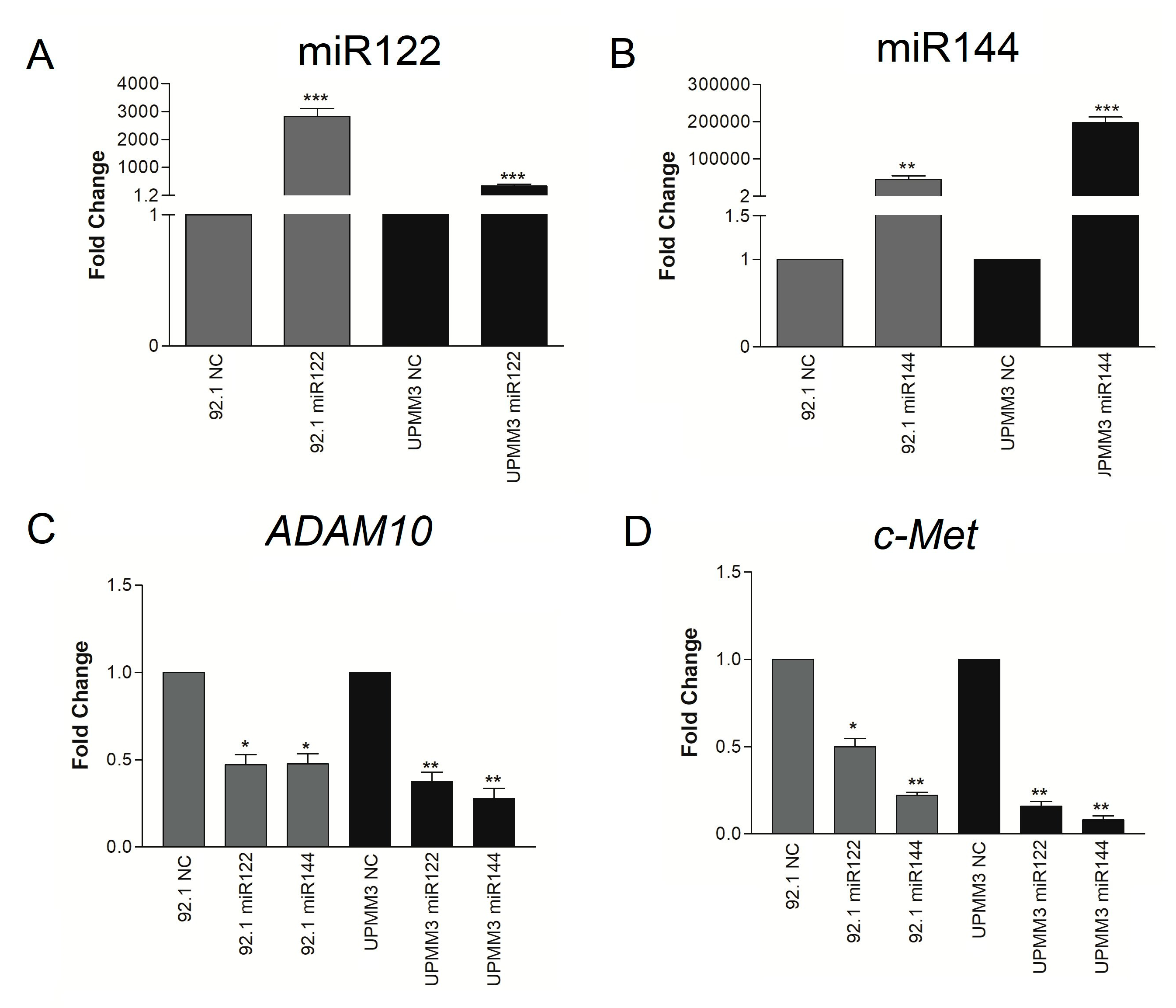
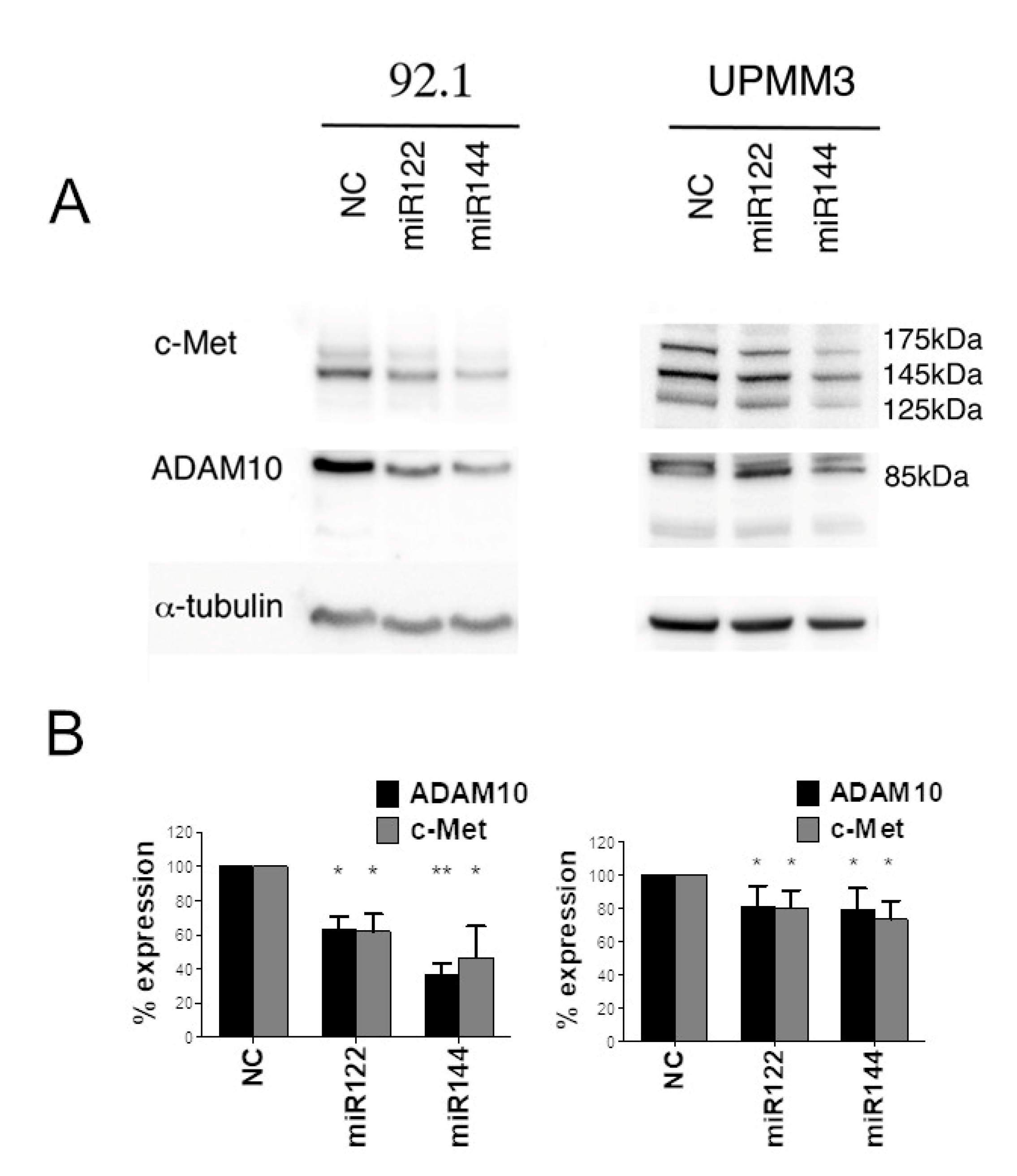
2.2. Effect of Synthetic miR122 and miR144 Transfection on ADAM10 and c-Met Expression
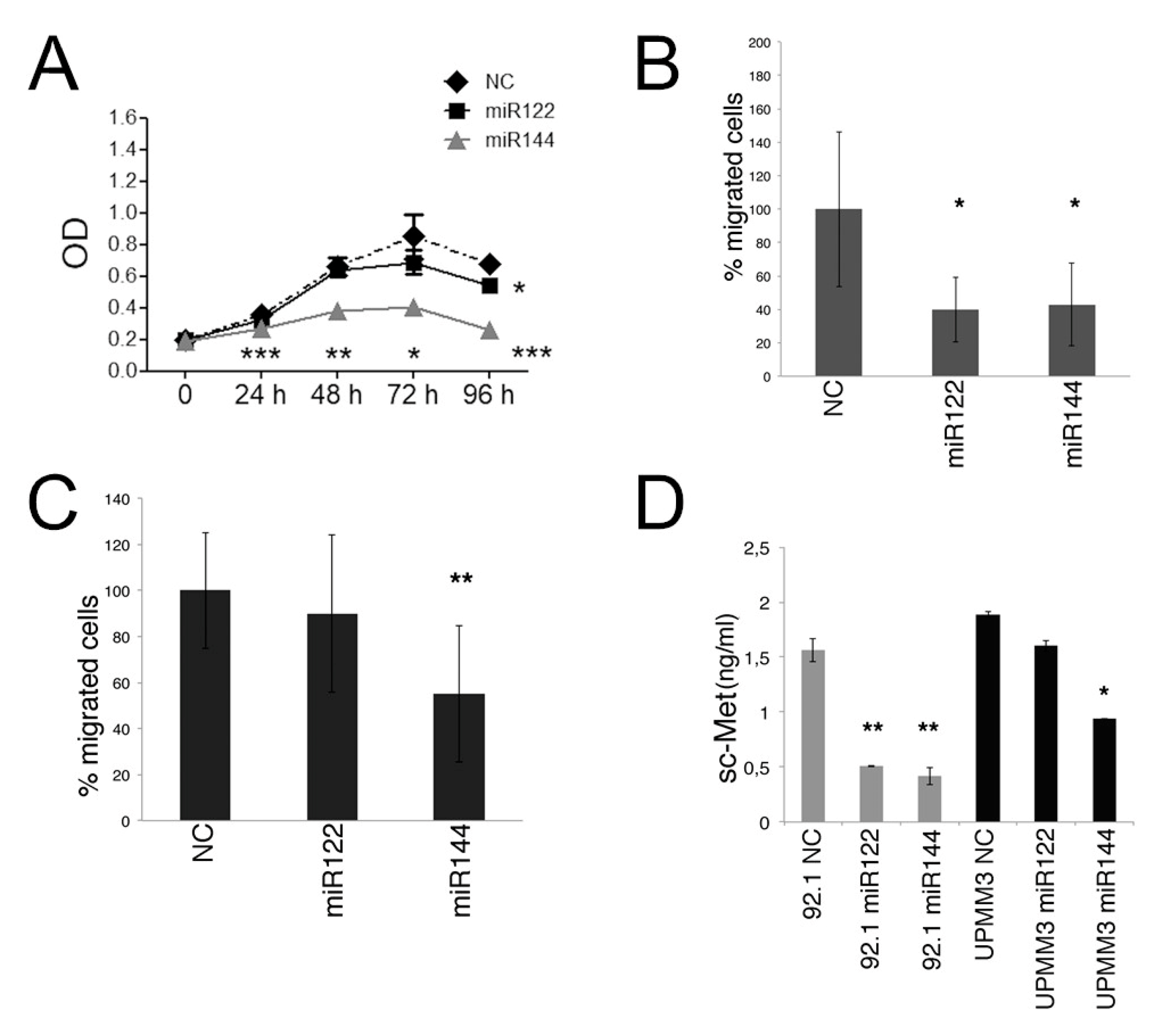
2.3. Effects of miR122 and miR144 on UM Cell Proliferation and Migration In Vitro
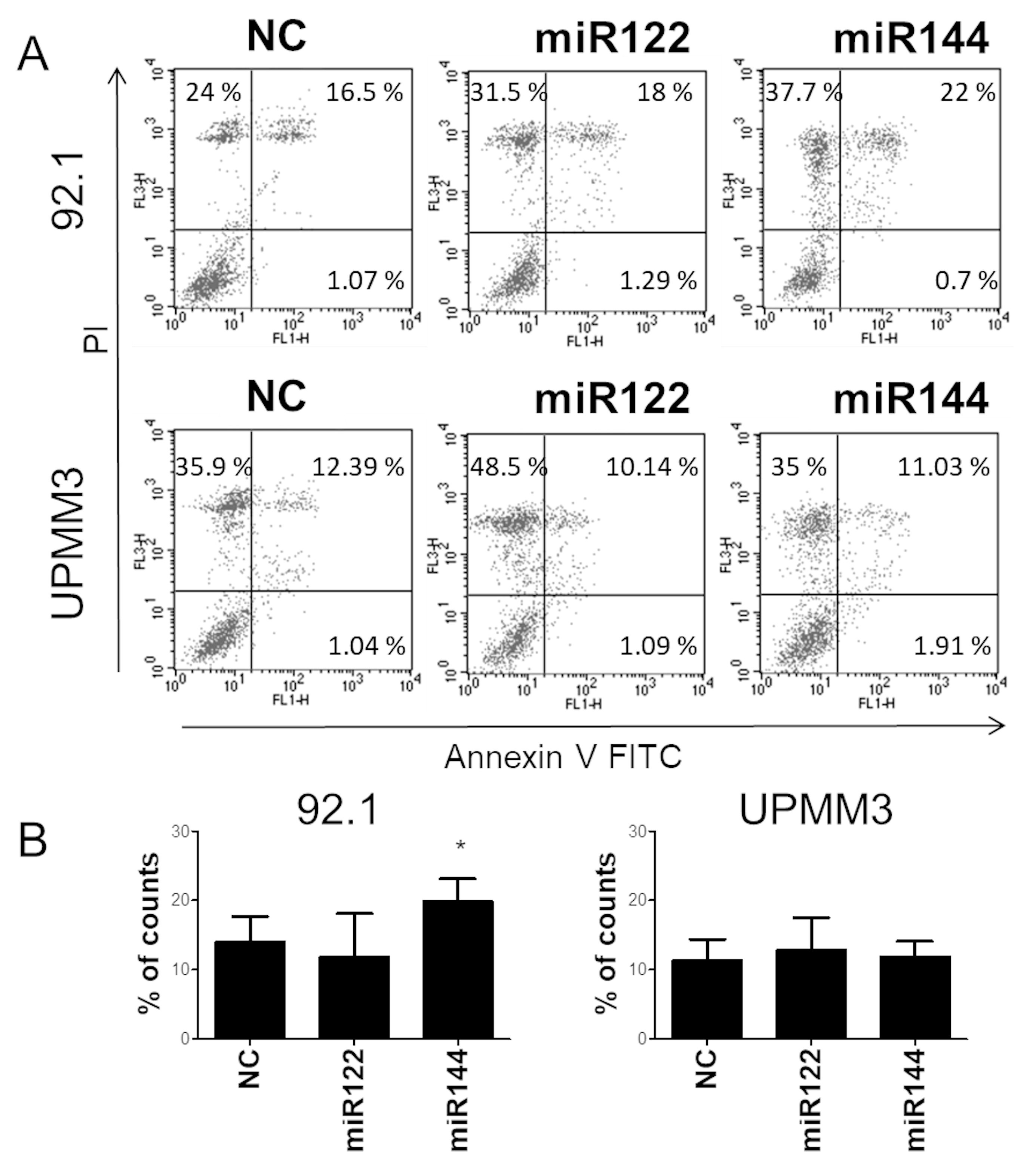
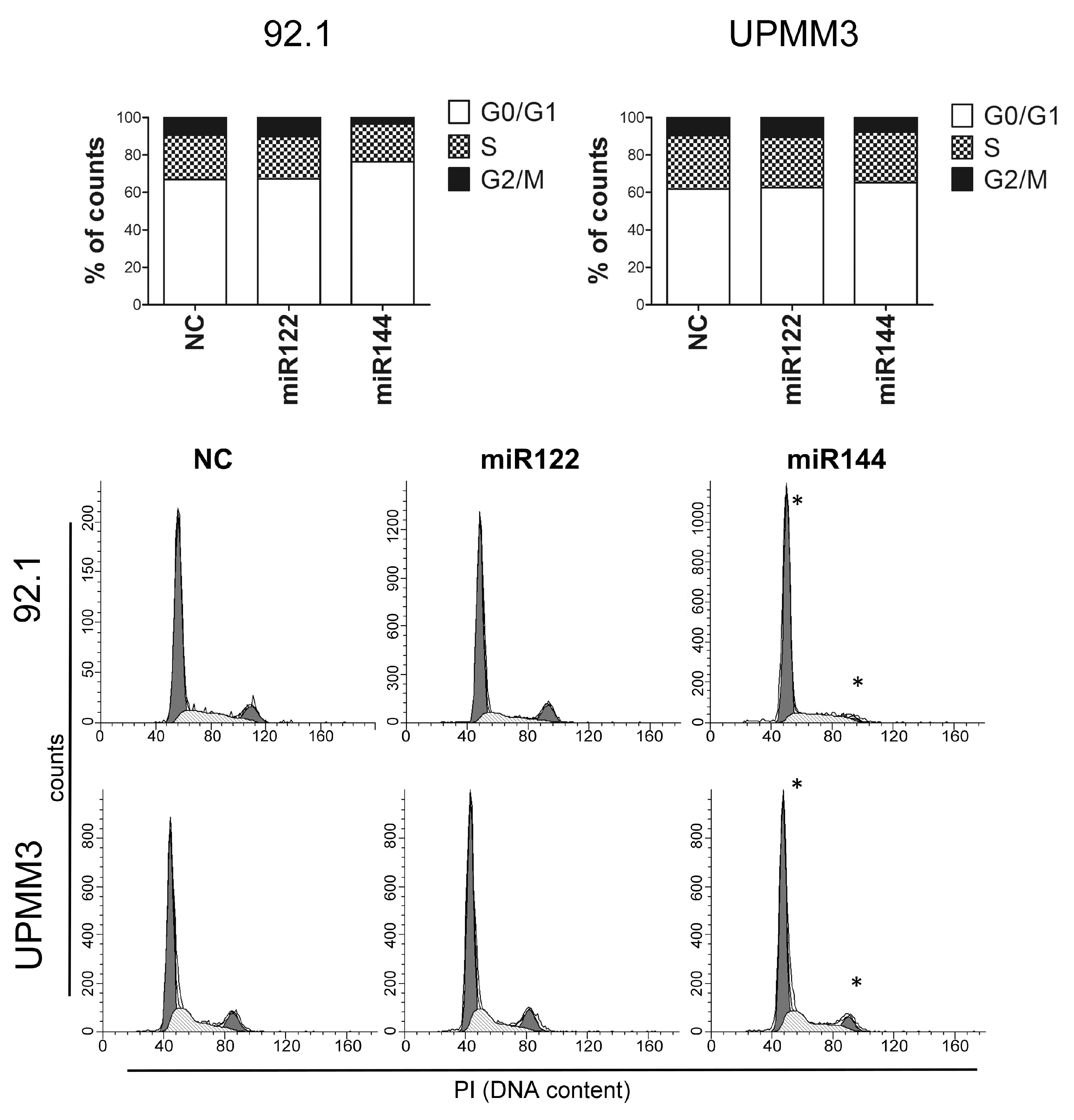
2.4. Effects of miR122 and miR144 on UM Apoptosis and Cell Cycle
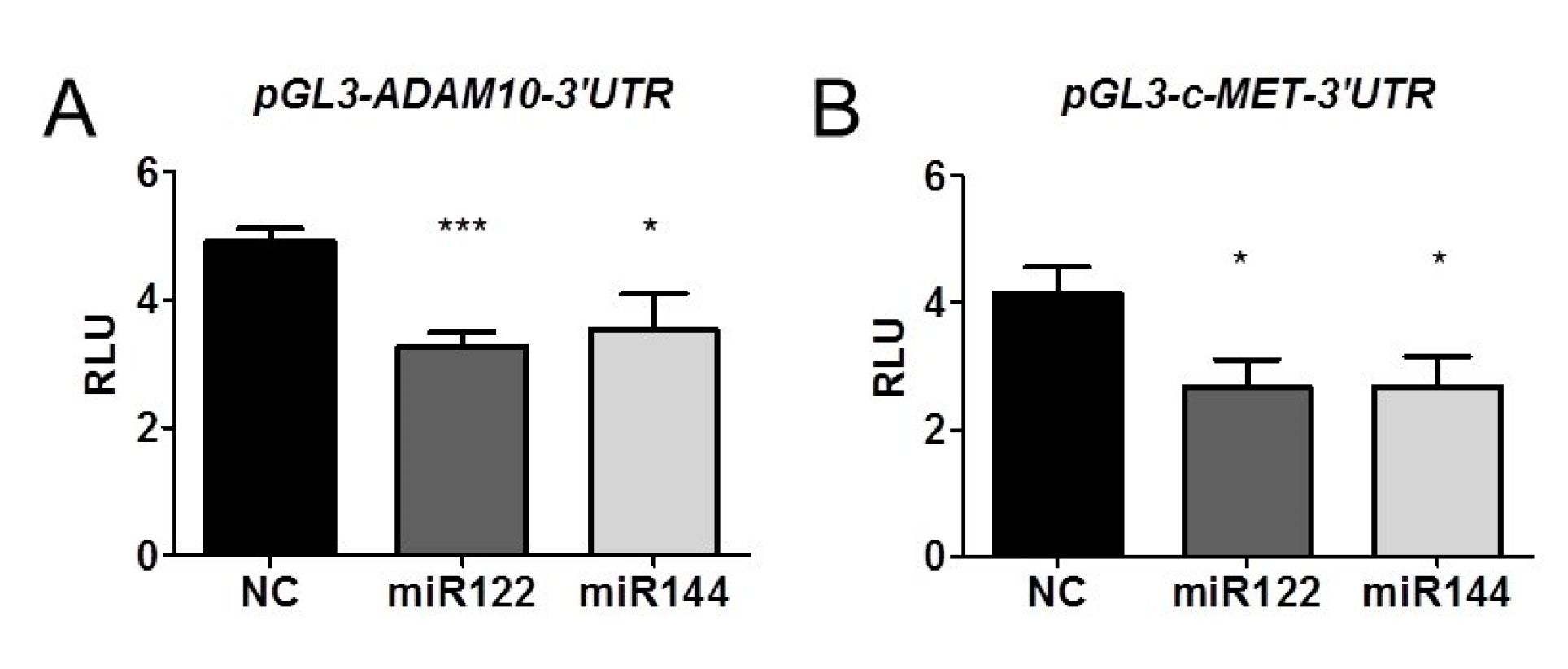
2.5. miR122 and miR144 Directly Target 3’UTR Region of Both ADAM10 and c-Met
3. Discussion
4. Methods
4.1. Cell Lines
4.2. Patients and Tissue Samples
4.3. RT-qPCR Analyses
4.4. Microarray and miRs Expression Analyses
4.5. MTT Assay
4.6. MiRs Transfection
4.7. Western Blot
4.8. Migration Assay
4.9. Detection of Soluble c-Met
4.10. Annexin V-FITC/PI Staining Assay
4.11. Cell Cycle Analysis
4.12. Luciferase Assay
4.13. Statistical Analyses
4.14. Ethical Approval and Informed Consent
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Amaro, A.; Gangemi, R.; Piaggio, F.; Angelini, G.; Barisione, G.; Ferrini, S.; Pfeffer, U. The biology of uveal melanoma. Cancer Metastasis Rev. 2017, 36, 109–140. [Google Scholar] [CrossRef] [PubMed]
- Damato, B.E.; Heimann, H.; Kalirai, H.; Coupland, S.E. Age, survival predictors, and metastatic death in patients with choroidal melanoma: Tentative evidence of a therapeutic effect on survival. JAMA Ophthalmol. 2014, 132, 605–613. [Google Scholar] [CrossRef] [PubMed]
- Niederkorn, A.; Wackernagel, W.; Artl, M.; Schwantzer, G.; Aigner, B.; Richtig, E. Response of patients with metastatic uveal melanoma to combined treatment with fotemustine and sorafenib. Acta Ophthalmol. 2014, 92, 696–697. [Google Scholar] [CrossRef] [PubMed]
- Van Raamsdonk, C.D.; Bezrookove, V.; Green, G.; Bauer, J.; Gaugler, L.; O’Brien, J.M.; Simpson, E.M.; Barsh, G.S.; Bastian, B.C. Frequent somatic mutations of GNAQ in uveal melanoma and blue naevi. Nature 2009, 457, 599–602. [Google Scholar] [CrossRef]
- Van Raamsdonk, C.D.; Griewank, K.G.; Crosby, M.B.; Garrido, M.C.; Vemula, S.; Wiesner, T.; Obenauf, A.C.; Wackernagel, W.; Green, G.; Bouvier, N.; et al. Mutations in GNA11 in uveal melanoma. N. Engl. J. Med. 2010, 363, 2191–2199. [Google Scholar] [CrossRef]
- Robertson, A.G.; Shih, J.; Yau, C.; Gibb, E.A.; Oba, J.; Mungall, K.L.; Hess, J.M.; Uzunangelov, V.; Walter, V.; Danilova, L.; et al. Integrative Analysis Identifies Four Molecular and Clinical Subsets in Uveal Melanoma. Cancer Cell 2017, 32, 204–220. [Google Scholar] [CrossRef]
- Triozzi, P.L.; Achberger, S.; Aldrich, W.; Crabb, J.W.; Saunthararajah, Y.; Singh, A.D. Association of tumor and plasma microRNA expression with tumor monosomy-3 in patients with uveal melanoma. Clin. Epigenetics 2016, 8, 80. [Google Scholar] [CrossRef]
- Damato, B.; Duke, C.; Coupland, S.E.; Hiscott, P.; Smith, P.A.; Campbell, I.; Douglas, A.; Howard, P. Cytogenetics of uveal melanoma: A 7-year clinical experience. Ophthalmology 2007, 114, 1925–1931. [Google Scholar] [CrossRef]
- Harbour, J.W.; Onken, M.D.; Roberson, E.D.O.; Duan, S.; Cao, L.; Worley, L.A.; Council, M.L.; Matatall, K.A.; Helms, C.; Bowcock, A.M. Frequent mutation of BAP1 in metastasizing uveal melanomas. Science 2010, 330, 1410–1413. [Google Scholar] [CrossRef]
- Kalirai, H.; Dodson, A.; Faqir, S.; Damato, B.E.; Coupland, S.E. Lack of BAP1 protein expression in uveal melanoma is associated with increased metastatic risk and has utility in routine prognostic testing. Br. J. Cancer 2014, 111, 1373–1380. [Google Scholar] [CrossRef]
- Van Essen, T.H.; van Pelt, S.I.; Versluis, M.; Bronkhorst, I.H.G.; van Duinen, S.G.; Marinkovic, M.; Kroes, W.G.M.; Ruivenkamp, C.A.L.; Shukla, S.; de Klein, A.; et al. Prognostic parameters in uveal melanoma and their association with BAP1 expression. Br. J. Ophthalmol. 2014, 98, 1738–1743. [Google Scholar] [CrossRef] [PubMed]
- Davis, E.J.; Johnson, D.B.; Sosman, J.A.; Chandra, S. Melanoma: What do all the mutations mean? Cancer 2018, 124, 3490–3499. [Google Scholar] [CrossRef] [PubMed]
- Hayward, N.K.; Wilmott, J.S.; Waddell, N.; Johansson, P.A.; Field, M.A.; Nones, K.; Patch, A.-M.; Kakavand, H.; Alexandrov, L.B.; Burke, H.; et al. Whole-genome landscapes of major melanoma subtypes. Nature 2017, 545, 175–180. [Google Scholar] [CrossRef] [PubMed]
- Gangemi, R.; Amaro, A.; Gino, A.; Barisione, G.; Fabbi, M.; Pfeffer, U.; Brizzolara, A.; Queirolo, P.; Salvi, S.; Boccardo, S.; et al. ADAM10 correlates with uveal melanoma metastasis and promotes in vitro invasion. Pigment. Cell Melanoma Res. 2014, 27, 1138–1148. [Google Scholar] [CrossRef]
- Surriga, O.; Rajasekhar, V.K.; Ambrosini, G.; Dogan, Y.; Huang, R.; Schwartz, G.K. Crizotinib, a c-Met inhibitor, prevents metastasis in a metastatic uveal melanoma model. Mol. Cancer Ther. 2013, 12, 2817–2826. [Google Scholar] [CrossRef]
- Topcu-Yilmaz, P.; Kiratli, H.; Saglam, A.; Söylemezoglu, F.; Hascelik, G. Correlation of clinicopathological parameters with HGF, c-Met, EGFR, and IGF-1R expression in uveal melanoma. Melanoma Res. 2010, 20, 126–132. [Google Scholar] [CrossRef]
- Miller, M.A.; Sullivan, R.J.; Lauffenburger, D.A. Molecular Pathways: Receptor Ectodomain Shedding in Treatment, Resistance, and Monitoring of Cancer. Clin. Cancer Res. 2017, 23, 623–629. [Google Scholar] [CrossRef]
- Boccaccio, C.; Comoglio, P.M. MET, a driver of invasive growth and cancer clonal evolution under therapeutic pressure. Curr. Opin. Cell Biol. 2014, 31, 98–105. [Google Scholar] [CrossRef]
- Barisione, G.; Fabbi, M.; Gino, A.; Queirolo, P.; Orgiano, L.; Spano, L.; Picasso, V.; Pfeffer, U.; Mosci, C.; Jager, M.J.; et al. Potential Role of Soluble c-Met as a New Candidate Biomarker of Metastatic Uveal Melanoma. JAMA Ophthalmol. 2015, 133, 1013–1021. [Google Scholar] [CrossRef]
- Drusco, A.; Croce, C.M. MicroRNAs and Cancer: A Long Story for Short RNAs. Adv. Cancer Res. 2017, 135, 1–24. [Google Scholar] [CrossRef]
- Ergün, S.; Ulasli, M.; Igci, Y.Z.; Igci, M.; Kırkbes, S.; Borazan, E.; Balik, A.; Yumrutaş, Ö.; Camci, C.; Cakmak, E.A.; et al. The association of the expression of miR-122-5p and its target ADAM10 with human breast cancer. Mol. Biol. Rep. 2015, 42, 497–505. [Google Scholar] [CrossRef] [PubMed]
- Bai, S.; Nasser, M.W.; Wang, B.; Hsu, S.-H.; Datta, J.; Kutay, H.; Yadav, A.; Nuovo, G.; Kumar, P.; Ghoshal, K. MicroRNA-122 inhibits tumorigenic properties of hepatocellular carcinoma cells and sensitizes these cells to sorafenib. J. Biol. Chem. 2009, 284, 32015–32027. [Google Scholar] [CrossRef] [PubMed]
- Cheng, C.; Li, W.; Zhang, Z.; Yoshimura, S.; Hao, Q.; Zhang, C.; Wang, Z. MicroRNA-144 is regulated by activator protein-1 (AP-1) and decreases expression of Alzheimer disease-related a disintegrin and metalloprotease 10 (ADAM10). J. Biol. Chem. 2013, 288, 13748–13761. [Google Scholar] [CrossRef] [PubMed]
- Lagos-Quintana, M.; Rauhut, R.; Yalcin, A.; Meyer, J.; Lendeckel, W.; Tuschl, T. Identification of tissue-specific microRNAs from mouse. Curr. Biol. 2002, 12, 735–739. [Google Scholar] [CrossRef]
- Tsai, W.-C.; Hsu, P.W.-C.; Lai, T.-C.; Chau, G.-Y.; Lin, C.-W.; Chen, C.-M.; Lin, C.-D.; Liao, Y.-L.; Wang, J.-L.; Chau, Y.-P.; et al. MicroRNA-122, a tumor suppressor microRNA that regulates intrahepatic metastasis of hepatocellular carcinoma. Hepatology 2009, 49, 1571–1582. [Google Scholar] [CrossRef]
- Qin, H.; Sha, J.; Jiang, C.; Gao, X.; Qu, L.; Yan, H.; Xu, T.; Jiang, Q.; Gao, H. miR-122 inhibits metastasis and epithelial-mesenchymal transition of non-small-cell lung cancer cells. OncoTargets Ther. 2015, 8, 3175–3184. [Google Scholar] [CrossRef][Green Version]
- Lu, W.; Zhang, Y.; Zhou, L.; Wang, X.; Mu, J.; Jiang, L.; Hu, Y.; Dong, P.; Liu, Y. miR-122 inhibits cancer cell malignancy by targeting PKM2 in gallbladder carcinoma. Tumour Biol. 2015, 37, 15615–15625. [Google Scholar] [CrossRef]
- Wang, Y.; Xing, Q.-F.; Liu, X.-Q.; Guo, Z.-J.; Li, C.-Y.; Sun, G. MiR-122 targets VEGFC in bladder cancer to inhibit tumor growth and angiogenesis. Am. J. Transl. Res. 2016, 8, 3056–3066. [Google Scholar]
- Meng, L.; Chen, Z.; Jiang, Z.; Huang, T.; Hu, J.; Luo, P.; Zhang, H.; Huang, M.; Huang, L.; Chen, Y.; et al. MiR-122-5p suppresses the proliferation, migration, and invasion of gastric cancer cells by targeting LYN. Acta Biochim. Biophys. Sin. 2020, 52, 49–57. [Google Scholar] [CrossRef]
- Rao, M.; Zhu, Y.; Zhou, Y.; Cong, X.; Feng, L. MicroRNA-122 inhibits proliferation and invasion in gastric cancer by targeting CREB1. Am. J. Cancer Res. 2017, 7, 323–333. [Google Scholar]
- Yang, Y.M.; Lee, C.G.; Koo, J.H.; Kim, T.H.; Lee, J.M.; An, J.; Kim, K.M.; Kim, S.G. Gα12 overexpressed in hepatocellular carcinoma reduces microRNA-122 expression via HNF4α inactivation, which causes c-Met induction. Oncotarget 2015, 6, 19055–19069. [Google Scholar] [CrossRef] [PubMed]
- Liu, S.; Suo, J.; Wang, C.; Sun, X.; Wang, D.; He, L.; Zhang, Y.; Li, W. Prognostic significance of low miR-144 expression in gastric cancer. Cancer Biomark 2017, 20, 547–552. [Google Scholar] [CrossRef] [PubMed]
- Pan, Y.; Zhang, J.; Fu, H.; Shen, L. miR-144 functions as a tumor suppressor in breast cancer through inhibiting ZEB1/2-mediated epithelial mesenchymal transition process. OncoTargets Ther. 2016, 9, 6247–6255. [Google Scholar] [CrossRef] [PubMed]
- Yu, M.; Lin, Y.; Zhou, Y.; Jin, H.; Hou, B.; Wu, Z.; Li, Z.; Jian, Z.; Sun, J. MiR-144 suppresses cell proliferation, migration, and invasion in hepatocellular carcinoma by targeting SMAD4. OncoTargets Ther. 2016, 9, 4705–4714. [Google Scholar] [CrossRef]
- Wu, J.; Zhao, Y.; Li, F.; Qiao, B. MiR-144-3p: A novel tumor suppressor targeting MAPK6 in cervical cancer. J. Physiol. Biochem. 2019, 75, 143–152. [Google Scholar] [CrossRef]
- Sun, L.; Bian, G.; Meng, Z.; Dang, G.; Shi, D.; Mi, S. MiR-144 Inhibits Uveal Melanoma Cell Proliferation and Invasion by Regulating c-Met Expression. PLoS ONE 2015, 10, e0124428. [Google Scholar] [CrossRef]
- Li, Z.; Yu, X.; Shen, J.; Jiang, Y. MicroRNA dysregulation in uveal melanoma: A new player enters the game. Oncotarget 2015, 6, 4562–4568. [Google Scholar] [CrossRef]
- Venkatesan, N.; Kanwar, J.; Deepa, P.R.; Khetan, V.; Crowley, T.M.; Raguraman, R.; Sugneswari, G.; Rishi, P.; Natarajan, V.; Biswas, J.; et al. Clinico-Pathological Association of Delineated miRNAs in Uveal Melanoma with Monosomy 3/Disomy 3 Chromosomal Aberrations. PLoS ONE 2016, 11, e0146128. [Google Scholar] [CrossRef]
- Worley, L.A.; Long, M.D.; Onken, M.D.; Harbour, J.W. Micro-RNAs associated with metastasis in uveal melanoma identified by multiplexed microarray profiling. Melanoma Res. 2008, 18, 184–190. [Google Scholar] [CrossRef]
- Ibarrola-Villava, M.; Llorca-Cardeñosa, M.J.; Tarazona, N.; Mongort, C.; Fleitas, T.; Perez-Fidalgo, J.A.; Roselló, S.; Navarro, S.; Ribas, G.; Cervantes, A. Deregulation of ARID1A, CDH1, cMET and PIK3CA and target-related microRNA expression in gastric cancer. Oncotarget 2015, 6, 26935–26945. [Google Scholar] [CrossRef]
- Dong, F.; Lou, D. MicroRNA-34b/c suppresses uveal melanoma cell proliferation and migration through multiple targets. Mol. Vis. 2012, 18, 537–546. [Google Scholar]
- Guo, H.; Yang, S.; Li, S.; Yan, M.; Li, L.; Zhang, H. LncRNA SNHG20 promotes cell proliferation and invasion via miR-140-5p-ADAM10 axis in cervical cancer. Biomed. Pharmacother. 2018, 102, 749–757. [Google Scholar] [CrossRef] [PubMed]
- Liao, Y.; Yin, X.; Deng, Y.; Peng, X. MiR-140-5p suppresses retinoblastoma cell growth via inhibiting c-Met/AKT/mTOR pathway. Biosci. Rep. 2018, 38, 38. [Google Scholar] [CrossRef] [PubMed]
- Felicetti, F.; Errico, M.C.; Bottero, L.; Segnalini, P.; Stoppacciaro, A.; Biffoni, M.; Felli, N.; Mattia, G.; Petrini, M.; Colombo, M.P.; et al. The promyelocytic leukemia zinc finger-microRNA-221/-222 pathway controls melanoma progression through multiple oncogenic mechanisms. Cancer Res. 2008, 68, 2745–2754. [Google Scholar] [CrossRef] [PubMed]
- De Waard-Siebinga, I.; Blom, D.J.; Griffioen, M.; Schrier, P.I.; Hoogendoorn, E.; Beverstock, G.; Danen, E.H.; Jager, M.J. Establishment and characterization of an uveal-melanoma cell line. Int. J. Cancer 1995, 62, 155–161. [Google Scholar] [CrossRef] [PubMed]
- Verbik, D.J.; Murray, T.G.; Tran, J.M.; Ksander, B.R. Melanomas that develop within the eye inhibit lymphocyte proliferation. Int. J. Cancer 1997, 73, 470–478. [Google Scholar] [CrossRef]
- Nareyeck, G.; Zeschnigk, M.; Bornfeld, N.; Anastassiou, G. Novel cell lines derived by long-term culture of primary uveal melanomas. Ophthalmologica 2009, 223, 196–201. [Google Scholar] [CrossRef]
- Zhang, L.; He, X.; Li, F.; Pan, H.; Huang, X.; Wen, X.; Zhang, H.; Li, B.; Ge, S.; Xu, X.; et al. The miR-181 family promotes cell cycle by targeting CTDSPL, a phosphatase-like tumor suppressor in uveal melanoma. J. Exp. Clin. Cancer Res. 2018, 37, 15. [Google Scholar] [CrossRef]
- Liu, N.; Sun, Q.; Chen, J.; Li, J.; Zeng, Y.; Zhai, S.; Li, P.; Wang, B.; Wang, X. MicroRNA-9 suppresses uveal melanoma cell migration and invasion through the NF-κB1 pathway. Oncol. Rep. 2012, 28, 961–968. [Google Scholar] [CrossRef]
- Yan, D.; Dong, X.D.; Chen, X.; Yao, S.; Wang, L.; Wang, J.; Wang, C.; Hu, D.-N.; Qu, J.; Tu, L. Role of microRNA-182 in posterior uveal melanoma: Regulation of tumor development through MITF, BCL2 and cyclin D2. PLoS ONE 2012, 7, e40967. [Google Scholar] [CrossRef]
- Ghini, F.; Rubolino, C.; Climent, M.; Simeone, I.; Marzi, M.J.; Nicassio, F. Endogenous transcripts control miRNA levels and activity in mammalian cells by target-directed miRNA degradation. Nat. Commun. 2018, 9, 3119. [Google Scholar] [CrossRef] [PubMed]
- Esposito, C.L.; Cerchia, L.; Catuogno, S.; De Vita, G.; Dassie, J.P.; Santamaria, G.; Swiderski, P.; Condorelli, G.; Giangrande, P.H.; de Franciscis, V. Multifunctional aptamer-miRNA conjugates for targeted cancer therapy. Mol. Ther. 2014, 22, 1151–1163. [Google Scholar] [CrossRef] [PubMed]
- Tivnan, A.; Orr, W.S.; Gubala, V.; Nooney, R.; Williams, D.E.; McDonagh, C.; Prenter, S.; Harvey, H.; Domingo-Fernández, R.; Bray, I.M.; et al. Inhibition of neuroblastoma tumor growth by targeted delivery of microRNA-34a using anti-disialoganglioside GD2 coated nanoparticles. PLoS ONE 2012, 7, e38129. [Google Scholar] [CrossRef] [PubMed]
- Kronski, E.; Fiori, M.E.; Barbieri, O.; Astigiano, S.; Mirisola, V.; Killian, P.H.; Bruno, A.; Pagani, A.; Rovera, F.; Pfeffer, U.; et al. miR181b is induced by the chemopreventive polyphenol curcumin and inhibits breast cancer metastasis via down-regulation of the inflammatory cytokines CXCL1 and -2. Mol. Oncol. 2014, 8, 581–595. [Google Scholar] [CrossRef] [PubMed]
- Gangemi, R.; Mirisola, V.; Barisione, G.; Fabbi, M.; Brizzolara, A.; Lanza, F.; Mosci, C.; Salvi, S.; Gualco, M.; Truini, M.; et al. Mda-9/syntenin is expressed in uveal melanoma and correlates with metastatic progression. PLoS ONE 2012, 7, e29989. [Google Scholar] [CrossRef]
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Amaro, A.; Croce, M.; Ferrini, S.; Barisione, G.; Gualco, M.; Perri, P.; Pfeffer, U.; Jager, M.J.; Coupland, S.E.; Mosci, C.; et al. Potential Onco-Suppressive Role of miR122 and miR144 in Uveal Melanoma through ADAM10 and C-Met Inhibition. Cancers 2020, 12, 1468. https://doi.org/10.3390/cancers12061468
Amaro A, Croce M, Ferrini S, Barisione G, Gualco M, Perri P, Pfeffer U, Jager MJ, Coupland SE, Mosci C, et al. Potential Onco-Suppressive Role of miR122 and miR144 in Uveal Melanoma through ADAM10 and C-Met Inhibition. Cancers. 2020; 12(6):1468. https://doi.org/10.3390/cancers12061468
Chicago/Turabian StyleAmaro, Adriana, Michela Croce, Silvano Ferrini, Gaia Barisione, Marina Gualco, Patrizia Perri, Ulrich Pfeffer, Martine J. Jager, Sarah E. Coupland, Carlo Mosci, and et al. 2020. "Potential Onco-Suppressive Role of miR122 and miR144 in Uveal Melanoma through ADAM10 and C-Met Inhibition" Cancers 12, no. 6: 1468. https://doi.org/10.3390/cancers12061468
APA StyleAmaro, A., Croce, M., Ferrini, S., Barisione, G., Gualco, M., Perri, P., Pfeffer, U., Jager, M. J., Coupland, S. E., Mosci, C., Filaci, G., Fabbi, M., Queirolo, P., & Gangemi, R. (2020). Potential Onco-Suppressive Role of miR122 and miR144 in Uveal Melanoma through ADAM10 and C-Met Inhibition. Cancers, 12(6), 1468. https://doi.org/10.3390/cancers12061468






