TOM40 Inhibits Ovarian Cancer Cell Growth by Modulating Mitochondrial Function Including Intracellular ATP and ROS Levels
Abstract
1. Introduction
2. Results
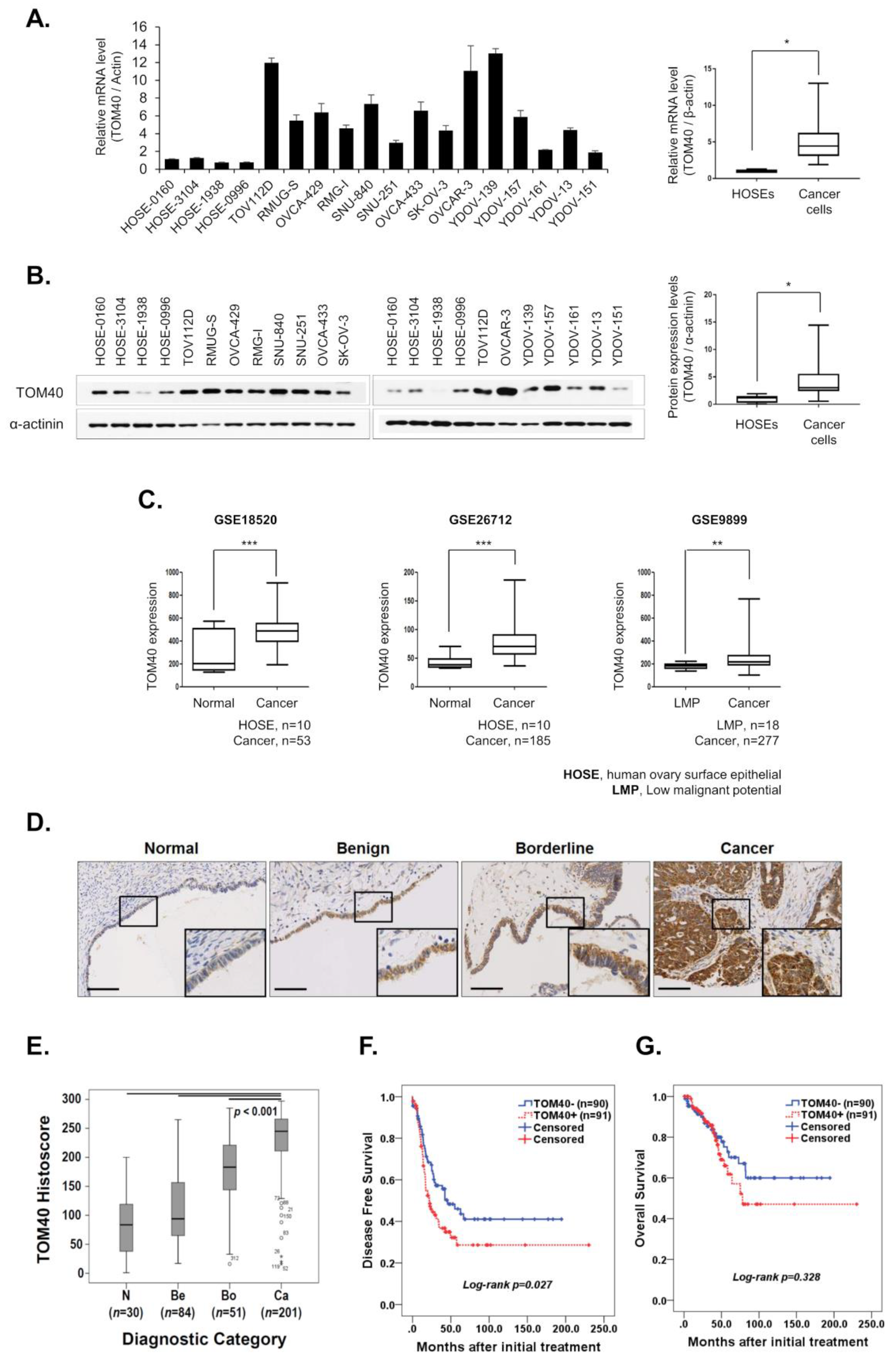
2.1. TOM40 Overexpression Is Negatively Correlated with Disease-Free Survival in Ovarian Cancer Patients
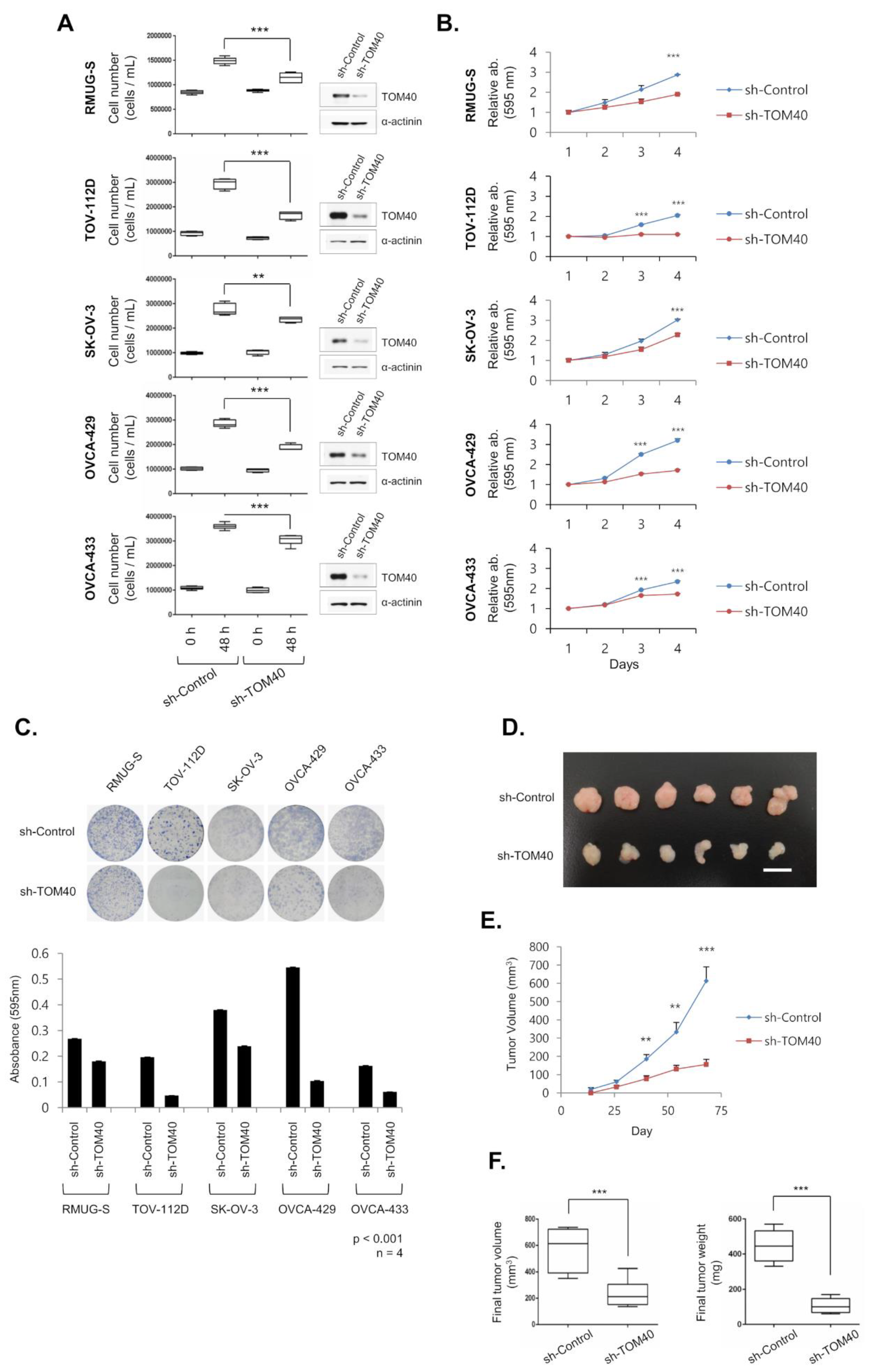
2.2. TOM40 Knockdown Suppresses the Growth of EOC Cell Lines and Xenografts
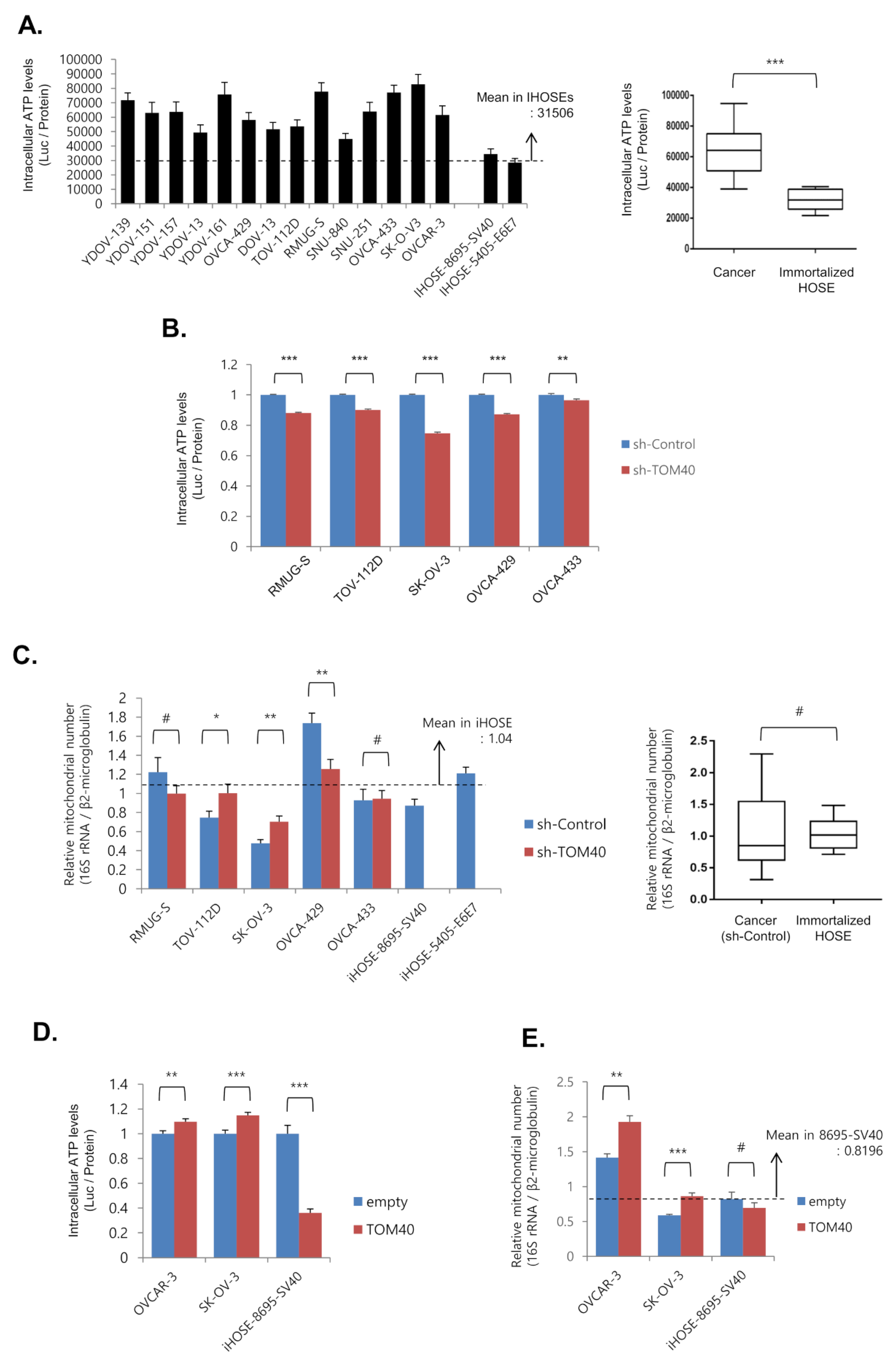
2.3. TOM40 Expression Is Positively Correlated with Intracellular ATP Levels
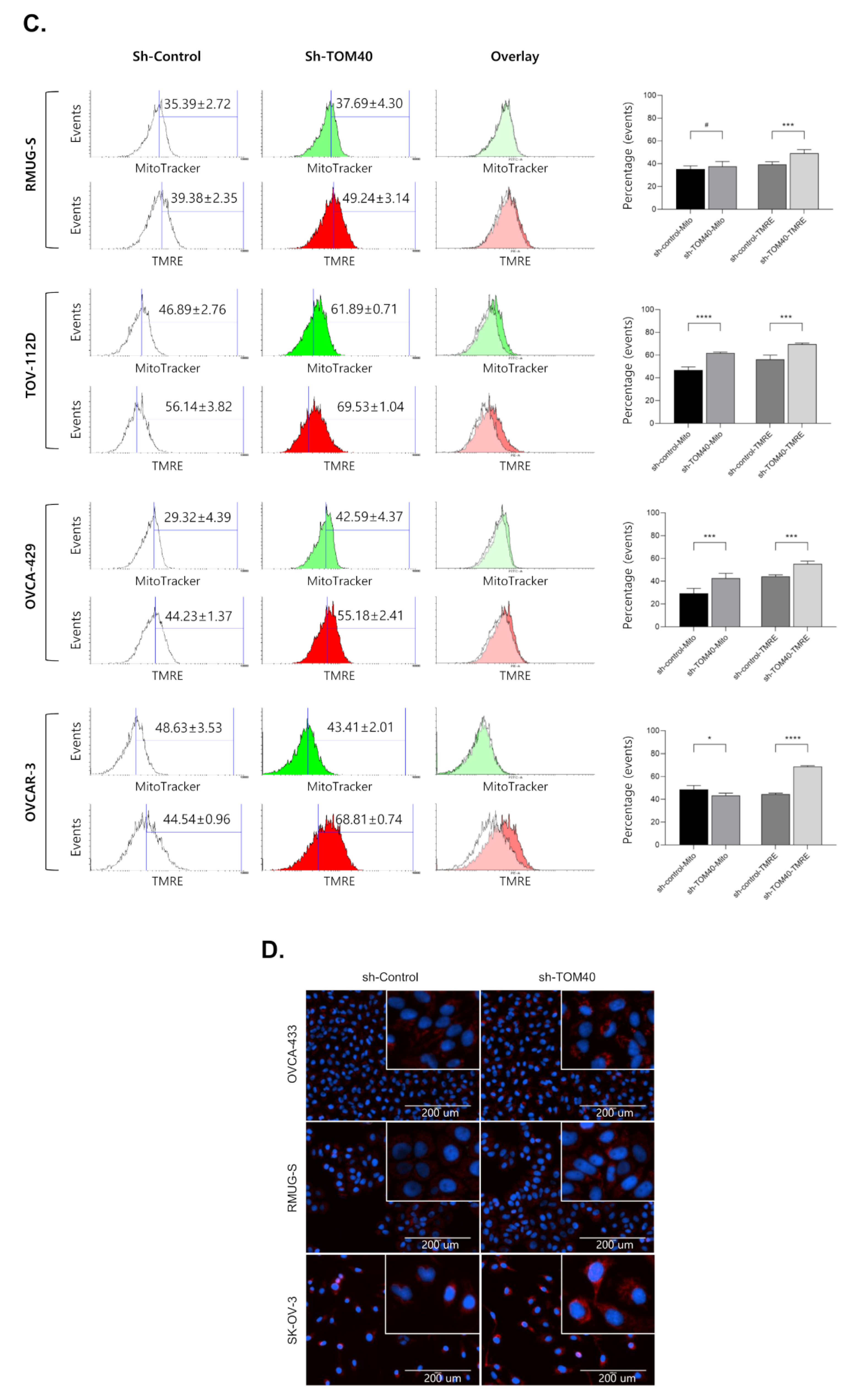
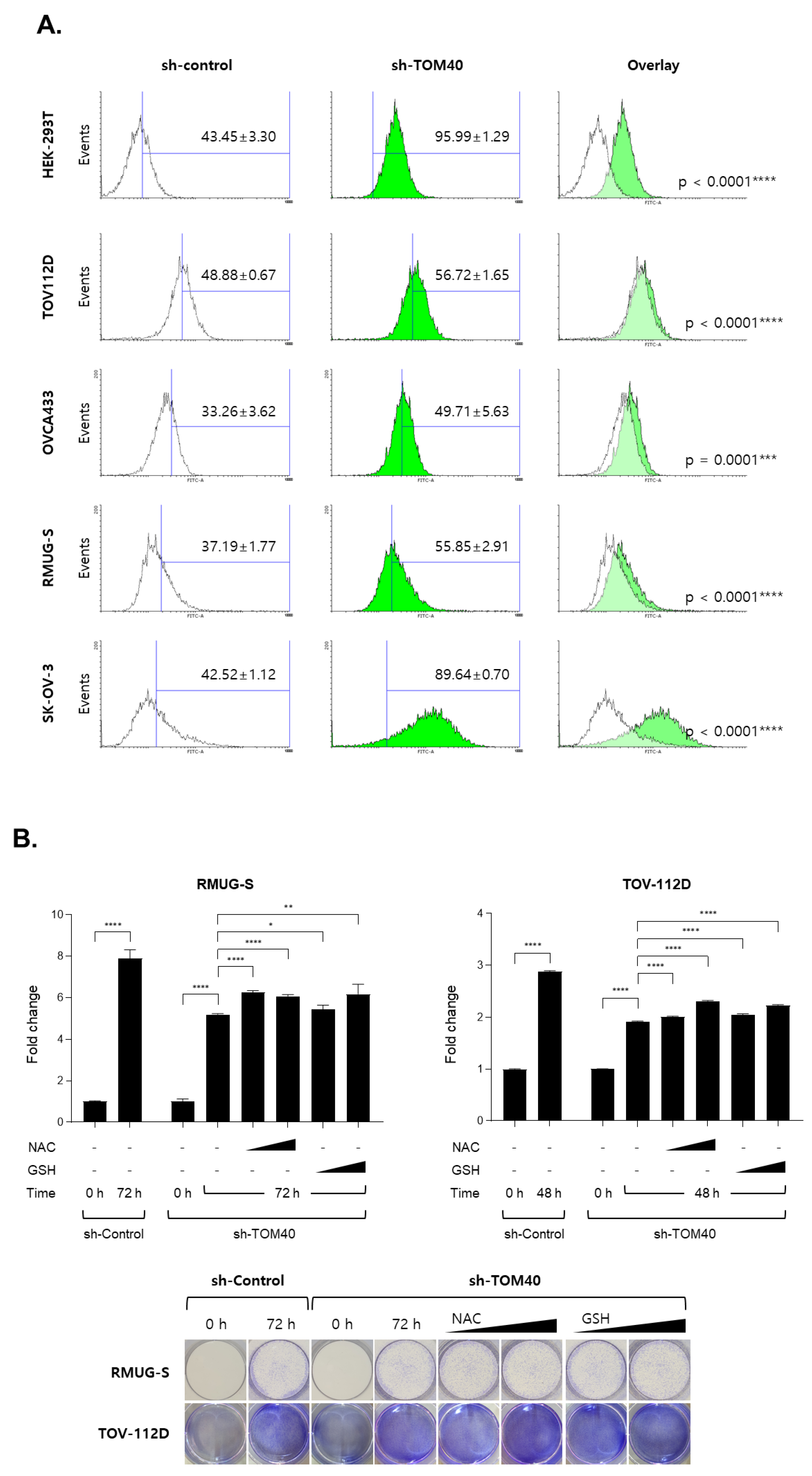
2.4. TOM40 Expression Knockdown in EOC Cells Increases Mitochondrial Membrane Potential and Intercellular Reactive Oxygen Species
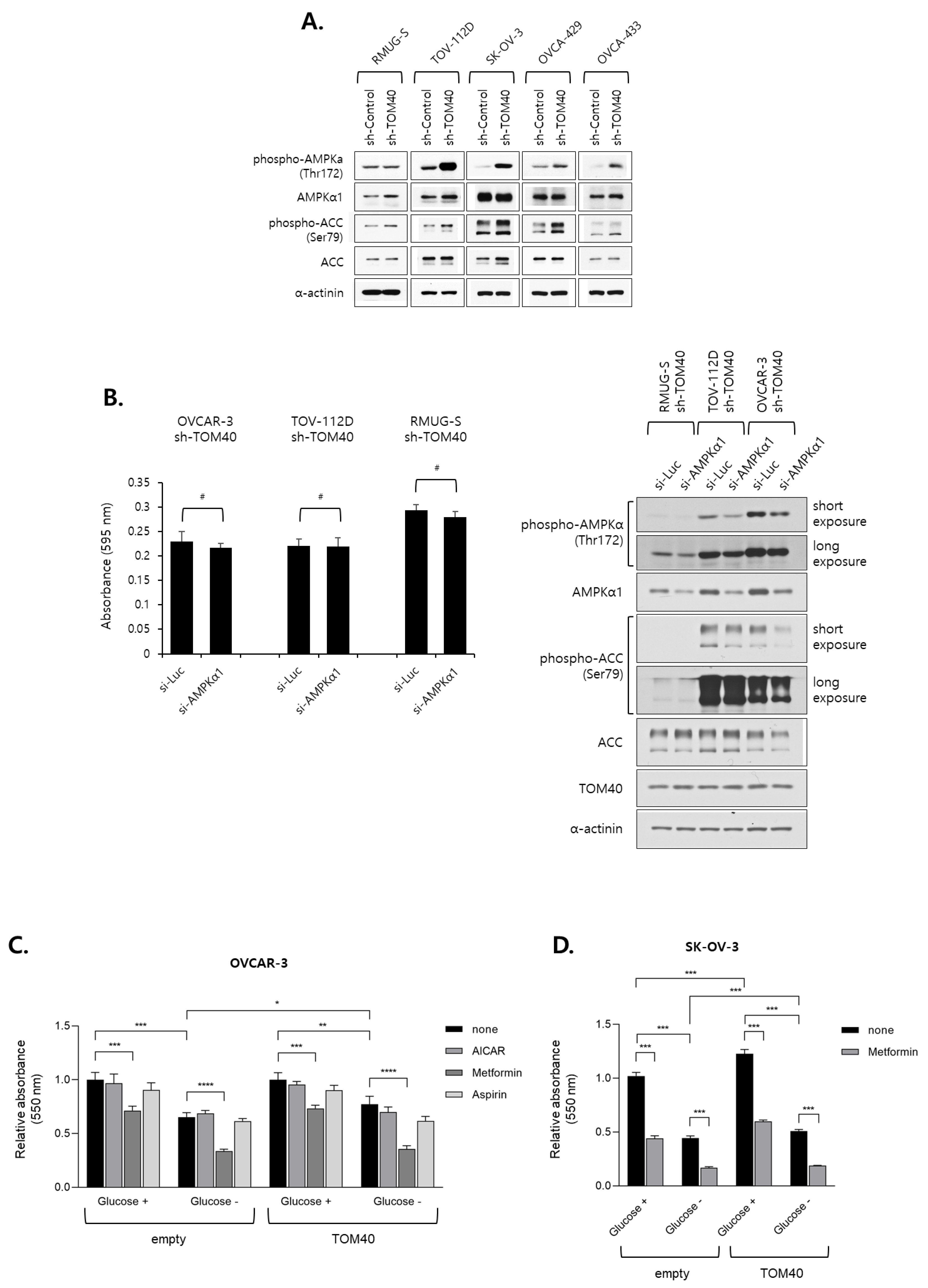
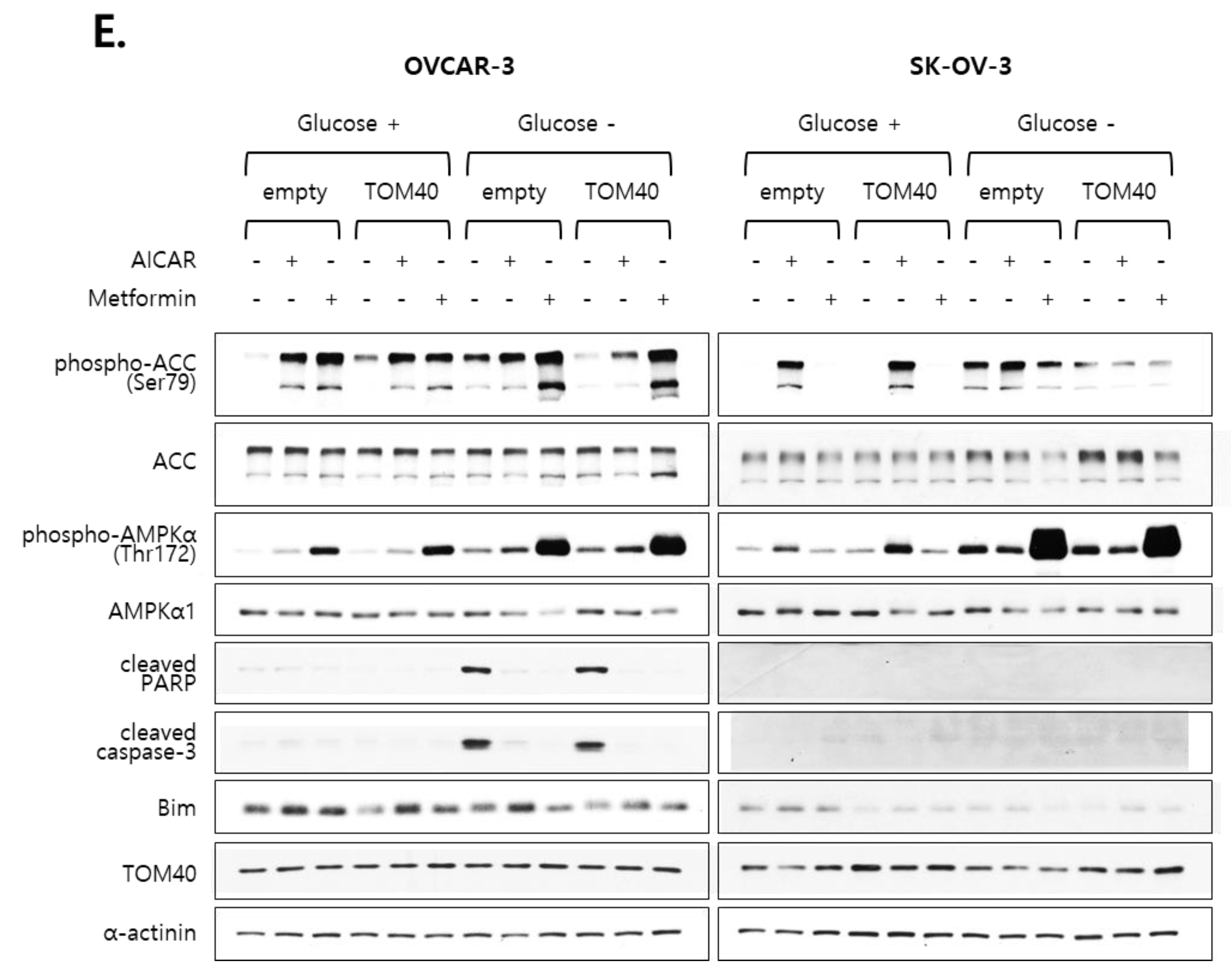
2.5. Metformin Inhibits the Proliferation of TOM40-Overexpressing EOC Cells in an AMPK-Independent Manner
3. Discussion
4. Materials and Methods
4.1. Patients, Tumor Samples, and Tissue Microarrays
4.2. Cell Culture and Antibodies
4.3. SYBR Green Real-Time PCR
4.4. Gene Expression Omnibus Dataset Analysis
4.5. Engineering Cell Lines That Stably Express TOM40 or sh-TOM40
4.6. Proliferation and Colony Formation Assays
4.7. Xenograft Mouse Model of Ovarian Cancer
4.8. Measurement of Intracellular ATP Levels
4.9. Determination of Mitochondrial DNA Copy Number
4.10. siRNA Transfection
4.11. Mitochondrial Membrane Potential Assay
4.12. Measurement of Intracellular Reactive Oxygen Species
4.13. Cytochrome Oxidase Activity Assay
4.14. Statistical Analysis
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Chacinska, A.; Koehler, C.M.; Milenkovic, D.; Lithgow, T.; Pfanner, N. Importing mitochondrial proteins: Machineries and mechanisms. Cell 2009, 138, 628–644. [Google Scholar] [CrossRef]
- Pagliarini, D.J.; Calvo, S.E.; Chang, B.; Sheth, S.A.; Vafai, S.B.; Ong, S.E.; Walford, G.A.; Sugiana, C.; Boneh, A.; Chen, W.K.; et al. A mitochondrial protein compendium elucidates complex I disease biology. Cell 2008, 134, 112–123. [Google Scholar] [CrossRef]
- Perocchi, F.; Jensen, L.J.; Gagneur, J.; Ahting, U.; von Mering, C.; Bork, P.; Prokisch, H.; Steinmetz, L.M. Assessing systems properties of yeast mitochondria through an interaction map of the organelle. PLoS Genet. 2006, 2, e170. [Google Scholar] [CrossRef]
- Bolender, N.; Sickmann, A.; Wagner, R.; Meisinger, C.; Pfanner, N. Multiple pathways for sorting mitochondrial precursor proteins. EMBO Rep. 2008, 9, 42–49. [Google Scholar] [CrossRef]
- Becker, L.; Bannwarth, M.; Meisinger, C.; Hill, K.; Model, K.; Krimmer, T.; Casadio, R.; Truscott, K.N.; Schulz, G.E.; Pfanner, N.; et al. Preprotein translocase of the outer mitochondrial membrane: Reconstituted Tom40 forms a characteristic TOM pore. J. Mol. Biol. 2005, 353, 1011–1020. [Google Scholar] [CrossRef]
- Kuszak, A.J.; Jacobs, D.; Gurnev, P.A.; Shiota, T.; Louis, J.M.; Lithgow, T.; Bezrukov, S.M.; Rostovtseva, T.K.; Buchanan, S.K. Evidence of Distinct Channel Conformations and Substrate Binding Affinities for the Mitochondrial Outer Membrane Protein Translocase Pore Tom40. J. Biol. Chem. 2015, 290, 26204–26217. [Google Scholar] [CrossRef]
- Gottschalk, W.K.; Lutz, M.W.; He, Y.T.; Saunders, A.M.; Burns, D.K.; Roses, A.D.; Chiba-Falek, O. The Broad Impact of TOM40 on Neurodegenerative Diseases in Aging. J. Parkinson’s Dis. Alzheimer’s Dis. 2014, 1, 12. [Google Scholar] [CrossRef]
- Baker, M.J.; Frazier, A.E.; Gulbis, J.M.; Ryan, M.T. Mitochondrial protein-import machinery: Correlating structure with function. Trends Cell Biol. 2007, 17, 456–464. [Google Scholar] [CrossRef]
- Esaki, M.; Kanamori, T.; Nishikawa, S.; Shin, I.; Schultz, P.G.; Endo, T. Tom40 protein import channel binds to non-native proteins and prevents their aggregation. Nat. Struct. Biol. 2003, 10, 988–994. [Google Scholar] [CrossRef]
- Billing, O.; Kao, G.; Naredi, P. Mitochondrial function is required for secretion of DAF-28/insulin in C. elegans. PLoS ONE 2011, 6, e14507. [Google Scholar] [CrossRef]
- Bertolin, G.; Ferrando-Miguel, R.; Jacoupy, M.; Traver, S.; Grenier, K.; Greene, A.W.; Dauphin, A.; Waharte, F.; Bayot, A.; Salamero, J.; et al. The TOMM machinery is a molecular switch in PINK1 and PARK2/PARKIN-dependent mitochondrial clearance. Autophagy 2013, 9, 1801–1817. [Google Scholar] [CrossRef]
- Zen, R. Neurological and molecular biological characterization of the mutant mouse line Tom40. Ph.D. Thesis, Techische University Munich, Munich, Germany, 2013; p. 243. [Google Scholar]
- Bender, A.; Desplats, P.; Spencer, B.; Rockenstein, E.; Adame, A.; Elstner, M.; Laub, C.; Mueller, S.; Koob, A.O.; Mante, M.; et al. TOM40 mediates mitochondrial dysfunction induced by alpha-synuclein accumulation in Parkinson’s disease. PLoS ONE 2013, 8, e62277. [Google Scholar] [CrossRef]
- Vyas, S.; Zaganjor, E.; Haigis, M.C. Mitochondria and Cancer. Cell 2016, 166, 555–566. [Google Scholar] [CrossRef]
- Vuda, M.; Kamath, A. Drug induced mitochondrial dysfunction: Mechanisms and adverse clinical consequences. Mitochondrion 2016, 31, 63–74. [Google Scholar] [CrossRef]
- Zhao, X.; Zeng, Z.; Gaur, U.; Fang, J.; Peng, T.; Li, S.; Zheng, W. Metformin protects PC12 cells and hippocampal neurons from H2 O 2 -induced oxidative damage through activation of AMPK pathway. J. Cell Physiol. 2019. [Google Scholar] [CrossRef]
- Gadducci, A.; Biglia, N.; Tana, R.; Cosio, S.; Gallo, M. Metformin use and gynecological cancers: A novel treatment option emerging from drug repositioning. Crit. Rev. Oncol./Hematol. 2016, 105, 73–83. [Google Scholar] [CrossRef]
- Zhuang, Y.; Chan, D.K.; Haugrud, A.B.; Miskimins, W.K. Mechanisms by which low glucose enhances the cytotoxicity of metformin to cancer cells both in vitro and in vivo. PLoS ONE 2014, 9, e108444. [Google Scholar] [CrossRef]
- Gwak, H.; Kim, Y.; An, H.; Dhanasekaran, D.N.; Song, Y.S. Metformin induces degradation of cyclin D1 via AMPK/GSK3beta axis in ovarian cancer. Mol. Carcinog. 2017, 56, 349–358. [Google Scholar] [CrossRef]
- Tang, G.; Guo, J.; Zhu, Y.; Huang, Z.; Liu, T.; Cai, J.; Yu, L.; Wang, Z. Metformin inhibits ovarian cancer via decreasing H3K27 trimethylation. Int. J. Oncol. 2018, 52, 1899–1911. [Google Scholar] [CrossRef]
- Tebbe, C.; Chhina, J.; Dar, S.A.; Sarigiannis, K.; Giri, S.; Munkarah, A.R.; Rattan, R. Metformin limits the adipocyte tumor-promoting effect on ovarian cancer. Oncotarget 2014, 5, 4746–4764. [Google Scholar] [CrossRef]
- Lee, J.; An, S.; Jung, J.H.; Kim, K.; Kim, J.Y.; An, I.S.; Bae, S. MUL1 E3 ligase regulates the antitumor effects of metformin in chemoresistant ovarian cancer cells via AKT degradation. Int. J. Oncol. 2019, 54, 1833–1842. [Google Scholar] [CrossRef]
- Zhang, H.H.; Guo, X.L. Combinational strategies of metformin and chemotherapy in cancers. Cancer Chemother. Pharmacol. 2016, 78, 13–26. [Google Scholar] [CrossRef]
- Hijaz, M.; Chhina, J.; Mert, I.; Taylor, M.; Dar, S.; Al-Wahab, Z.; Ali-Fehmi, R.; Buekers, T.; Munkarah, A.R.; Rattan, R. Preclinical evaluation of olaparib and metformin combination in BRCA1 wildtype ovarian cancer. Gynecol. Oncol. 2016, 142, 323–331. [Google Scholar] [CrossRef]
- Piel, S.; Ehinger, J.K.; Elmer, E.; Hansson, M.J. Metformin induces lactate production in peripheral blood mononuclear cells and platelets through specific mitochondrial complex I inhibition. Acta Physiol. (Oxf. Engl.) 2015, 213, 171–180. [Google Scholar] [CrossRef]
- Protti, A.; Lecchi, A.; Fortunato, F.; Artoni, A.; Greppi, N.; Vecchio, S.; Fagiolari, G.; Moggio, M.; Comi, G.P.; Mistraletti, G.; et al. Metformin overdose causes platelet mitochondrial dysfunction in humans. Crit. Care (Lond. Engl.) 2012, 16, R180. [Google Scholar] [CrossRef]
- Andrzejewski, S.; Gravel, S.P.; Pollak, M.; St-Pierre, J. Metformin directly acts on mitochondria to alter cellular bioenergetics. Cancer Metab. 2014, 2, 12. [Google Scholar] [CrossRef]
- Cho, H.; Lee, Y.S.; Kim, J.; Chung, J.Y.; Kim, J.H. Overexpression of glucose transporter-1 (GLUT-1) predicts poor prognosis in epithelial ovarian cancer. Cancer Investig. 2013, 31, 607–615. [Google Scholar] [CrossRef]
- Chay, D.; Cho, H.; Lim, B.J.; Kang, E.S.; Oh, Y.J.; Choi, S.M.; Kim, B.W.; Kim, Y.T.; Kim, J.H. ER-60 (PDIA3) is highly expressed in a newly established serous ovarian cancer cell line, YDOV-139. Int. J. Oncol. 2010, 37, 399–412. [Google Scholar]
- Cho, H.; Kang, E.S.; Hong, S.W.; Oh, Y.J.; Choi, S.M.; Kim, S.W.; Kim, S.H.; Kim, Y.T.; Lee, K.S.; Choi, Y.K.; et al. Genomic and proteomic characterization of YDOV-157, a newly established human epithelial ovarian cancer cell line. Mol. Cell. Biochem. 2008, 319, 189–201. [Google Scholar] [CrossRef]
- Hinchy, E.C.; Gruszczyk, A.V.; Willows, R.; Navaratnam, N.; Hall, A.R.; Bates, G.; Bright, T.P.; Krieg, T.; Carling, D.; Murphy, M.P. Mitochondria-derived ROS activate AMP-activated protein kinase (AMPK) indirectly. J. Biol. Chem. 2018, 293, 17208–17217. [Google Scholar] [CrossRef]
- Yang, W.; Park, I.J.; Yun, H.; Im, D.U.; Ock, S.; Kim, J.; Seo, S.M.; Shin, H.Y.; Viollet, B.; Kang, I.; et al. AMP-activated protein kinase alpha2 and E2F1 transcription factor mediate doxorubicin-induced cytotoxicity by forming a positive signal loop in mouse embryonic fibroblasts and non-carcinoma cells. J. Biol. Chem. 2014, 289, 4839–4852. [Google Scholar] [CrossRef]
- Warburg, O. On the origin of cancer cells. Science 1956, 123, 309–314. [Google Scholar] [CrossRef]
- Mathupala, S.P.; Ko, Y.H.; Pedersen, P.L. Hexokinase-2 bound to mitochondria: cancer’s stygian link to the “Warburg Effect” and a pivotal target for effective therapy. Semin. Cancer Biol. 2009, 19, 17–24. [Google Scholar] [CrossRef]
- Carew, J.S.; Huang, P. Mitochondrial defects in cancer. Mol. Cancer 2002, 1, 9. [Google Scholar] [CrossRef]
- Moreno-Sanchez, R.; Rodriguez-Enriquez, S.; Marin-Hernandez, A.; Saavedra, E. Energy metabolism in tumor cells. FEBS J. 2007, 274, 1393–1418. [Google Scholar] [CrossRef]
- Weinberg, F.; Chandel, N.S. Mitochondrial metabolism and cancer. Ann. N. Y. Acad. Sci. 2009, 1177, 66–73. [Google Scholar] [CrossRef]
- Zeitlow, K.; Charlambous, L.; Ng, I.; Gagrani, S.; Mihovilovic, M.; Luo, S.; Rock, D.L.; Saunders, A.; Roses, A.D.; Gottschalk, W.K. The biological foundation of the genetic association of TOMM40 with late-onset Alzheimer’s disease. Biochim. Biophys. Acta Mol. Basis Dis. 2017, 1863, 2973–2986. [Google Scholar] [CrossRef]
- Namba, T. BAP31 regulates mitochondrial function via interaction with Tom40 within ER-mitochondria contact sites. Sci. Adv. 2019, 5, eaaw1386. [Google Scholar] [CrossRef]
- Harbauer, A.B.; Opalinska, M.; Gerbeth, C.; Herman, J.S.; Rao, S.; Schonfisch, B.; Guiard, B.; Schmidt, O.; Pfanner, N.; Meisinger, C. Cell cycle-dependent regulation of mitochondrial preprotein translocase. Science 2014, 346, 1109–1113. [Google Scholar] [CrossRef]
- Zorova, L.D.; Popkov, V.A.; Plotnikov, E.Y.; Silachev, D.N.; Pevzner, I.B.; Jankauskas, S.S.; Babenko, V.A.; Zorov, S.D.; Balakireva, A.V.; Juhaszova, M.; et al. Mitochondrial membrane potential. Anal. Biochem. 2018, 552, 50–59. [Google Scholar] [CrossRef]
- Korshunov, S.S.; Skulachev, V.P.; Starkov, A.A. High protonic potential actuates a mechanism of production of reactive oxygen species in mitochondria. FEBS Lett. 1997, 416, 15–18. [Google Scholar] [CrossRef]
- Starkov, A.A.; Fiskum, G. Regulation of brain mitochondrial H2O2 production by membrane potential and NAD(P)H redox state. J. Neurochem. 2003, 86, 1101–1107. [Google Scholar] [CrossRef]
- Skulachev, V.P. Role of uncoupled and non-coupled oxidations in maintenance of safely low levels of oxygen and its one-electron reductants. Q. Rev. Biophys. 1996, 29, 169–202. [Google Scholar] [CrossRef]
- Kemp, B.E.; Stapleton, D.; Campbell, D.J.; Chen, Z.P.; Murthy, S.; Walter, M.; Gupta, A.; Adams, J.J.; Katsis, F.; van Denderen, B.; et al. AMP-activated protein kinase, super metabolic regulator. Biochem. Soc. Trans. 2003, 31, 162–168. [Google Scholar] [CrossRef]
- Viollet, B.; Athea, Y.; Mounier, R.; Guigas, B.; Zarrinpashneh, E.; Horman, S.; Lantier, L.; Hebrard, S.; Devin-Leclerc, J.; Beauloye, C.; et al. AMPK: Lessons from transgenic and knockout animals. Front. Biosci. 2009, 14, 19–44. [Google Scholar] [CrossRef]
- Li, C.; Liu, V.W.; Chiu, P.M.; Chan, D.W.; Ngan, H.Y. Over-expressions of AMPK subunits in ovarian carcinomas with significant clinical implications. BMC Cancer 2012, 12, 357. [Google Scholar] [CrossRef]
- Henry, W.S.; Laszewski, T.; Tsang, T.; Beca, F.; Beck, A.H.; McAllister, S.S.; Toker, A. Aspirin Suppresses Growth in PI3K-Mutant Breast Cancer by Activating AMPK and Inhibiting mTORC1 Signaling. Cancer Res. 2017, 77, 790–801. [Google Scholar] [CrossRef]
- Ducommun, S.; Ford, R.J.; Bultot, L.; Deak, M.; Bertrand, L.; Kemp, B.E.; Steinberg, G.R.; Sakamoto, K. Enhanced activation of cellular AMPK by dual-small molecule treatment: AICAR and A769662. Am. J. Physiol. Endocrinol. Metab. 2014, 306, E688–E696. [Google Scholar] [CrossRef]
- Xu, S.; Yang, Z.; Jin, P.; Yang, X.; Li, X.; Wei, X.; Wang, Y.; Long, S.; Zhang, T.; Chen, G.; et al. Metformin Suppresses Tumor Progression by Inactivating Stromal Fibroblasts in Ovarian Cancer. Mol. Cancer Ther. 2018, 17, 1291–1302. [Google Scholar] [CrossRef]
- Rattan, R.; Graham, R.P.; Maguire, J.L.; Giri, S.; Shridhar, V. Metformin suppresses ovarian cancer growth and metastasis with enhancement of cisplatin cytotoxicity in vivo. Neoplasia 2011, 13, 483–491. [Google Scholar] [CrossRef]
- Stine, J.E.; Bae-Jump, V. Metformin and gynecologic cancers. Obs. Gynecol. Surv. 2014, 69, 477–489. [Google Scholar] [CrossRef] [PubMed]
- Franciosi, M.; Lucisano, G.; Lapice, E.; Strippoli, G.F.; Pellegrini, F.; Nicolucci, A. Metformin therapy and risk of cancer in patients with type 2 diabetes: Systematic review. PLoS ONE 2013, 8, e71583. [Google Scholar] [CrossRef] [PubMed]
- Martin-Rodriguez, S.; de Pablos-Velasco, P.; Calbet, J.A.L. Mitochondrial Complex I Inhibition by Metformin: Drug-Exercise Interactions. Trends Endocrinol. Metab. 2020, 31, 269–271. [Google Scholar] [CrossRef] [PubMed]
- Dier, U.; Shin, D.H.; Hemachandra, L.P.; Uusitalo, L.M.; Hempel, N. Bioenergetic analysis of ovarian cancer cell lines: Profiling of histological subtypes and identification of a mitochondria-defective cell line. PLoS ONE 2014, 9, e98479. [Google Scholar] [CrossRef] [PubMed]
- Wookyeom, Y.; Hanbyoul, C.; Ha-Yeon, S.; Joon-Yong, C.; Kang, E.S.; Eun-Ju, L.; Jae-Hoon, K. Accumulation of cytoplasmic Cdk1 is associated with cancer growth and survival rate in epithelial ovarian cancer. Oncotarget 2016, 7, 49481. [Google Scholar] [CrossRef]
- Choi, C.H.; Chung, J.Y.; Kim, J.H.; Kim, B.G.; Hewitt, S.M. Expression of fibroblast growth factor receptor family members is associated with prognosis in early stage cervical cancer patients. J. Transl. Med. 2016, 14, 124. [Google Scholar] [CrossRef]
- Shin, H.Y.; Yang, W.; Lee, E.J.; Han, G.H.; Cho, H.; Chay, D.B.; Kim, J.H. Establishment of five immortalized human ovarian surface epithelial cell lines via SV40 T antigen or HPV E6/E7 expression. PLoS ONE 2018, 13, e0205297. [Google Scholar] [CrossRef]

| Gene | TOM40 | ||
|---|---|---|---|
| Parameter | No. | Mean IHC a Score (95% CI) | p-Value |
| All study subjects | 366 | 184 (176–192) | |
| Diagnostic Category | <0.001 | ||
| Normal | 30 | 82 (63–101) | |
| Benign | 84 | 112 (98–126) | |
| Borderline | 51 | 176 (158–194) | |
| Cancer | 201 | 231 (225–238) | |
| Age | 0.412 | ||
| ≤50 | 93 | 228 (218–239) | |
| >50 | 108 | 234 (225–243) | |
| FIGO stage b | 0.441 | ||
| I–II | 57 | 226 (210–241) | |
| III–IV | 124 | 235 (227–243) | |
| Recurrence | 20 | 226 (211–241) | |
| Cell type | 0.101 | ||
| Serous | 137 | 235 (227–243) | |
| Others | 64 | 223 (210–237) | |
| Tumor grade | 0.070 | ||
| Well + Moderate | 86 | 226 (214–238) | |
| Poor | 100 | 239 (232–247) | |
| Cancer type | 0.005 | ||
| Type I tumor c | 81 | 219 (207–232) | |
| Type II tumor d | 114 | 241 (233–249) | |
| CA125 | 0.264 | ||
| Negative | 32 | 241 (227–254) | |
| Positive (>35 U/mL) | 166 | 230 (222–238) | |
| Chemosensitivity | 0.600 | ||
| Sensitive | 141 | 233 (225–241) | |
| Resistant | 41 | 237 (227–248) | |
| Disease-Free Survival | Overall Survival | ||||||||
|---|---|---|---|---|---|---|---|---|---|
| Variables | Univariate Analysis | Multivariate Analysis | Variables | Univariate Analysis | Multivariate Analysis | ||||
| Hazard Ratio (95%CI) | p-Value | Hazard Ratio (95%CI) | p-Value | Hazard Ratio (95%CI) | p-Value | Hazard Ratio (95%CI) | p-Value | ||
| Age (>50) | 1.62 (1.07–2.45) | 0.022 | 1.14 (0.73–1.77) | 0.554 | Age (>50) | 2.34 (1.22–4.47) | 0.01 | 1.76 (0.91–3.39) | 0.091 |
| FIGO stage (≥III) | 6.51 (3.26–13.00) | <0.001 | 4.75 (2.33–9.68) | <0.001 | FIGO stage (≥ III) | 4.45 (1.75–11.31) | 0.002 | 2.62 (1.01–6.81) | 0.047 |
| Cell type (serous) | 2.98 (1.74–5.11) | <0.001 | 1.68 (0.91–3.07) | 0.093 | Cell type (serous) | 4.08 (1.60–10.35) | 0.003 | 2.35 (0.91–6.07) | 0.077 |
| Grade (poor) | 1.98 (1.29–3.04) | 0.002 | 1.70 (1.10–2.62) | 0.017 | Grade (poor) | 2.27 (1.21–4.25) | 0.01 | 2.12 (1.12–4.00) | 0.021 |
| CA125+ (>35 U/mL) | 2.30 (1.11–4.76) | 0.024 | 1.01 (0.46–2.24) | 0.966 | CA125+ (>35 U/mL) | 1.91 (0.68–5.35) | 0.216 | NA | |
| TOM40+a | 1.57 (1.04–2.36) | 0.028 | 1.73 (1.12–2.67) | 0.012 | TOM40+a | 1.34 (0.74–2.43) | 0.33 | NA | |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Yang, W.; Shin, H.-Y.; Cho, H.; Chung, J.-Y.; Lee, E.-j.; Kim, J.-H.; Kang, E.-S. TOM40 Inhibits Ovarian Cancer Cell Growth by Modulating Mitochondrial Function Including Intracellular ATP and ROS Levels. Cancers 2020, 12, 1329. https://doi.org/10.3390/cancers12051329
Yang W, Shin H-Y, Cho H, Chung J-Y, Lee E-j, Kim J-H, Kang E-S. TOM40 Inhibits Ovarian Cancer Cell Growth by Modulating Mitochondrial Function Including Intracellular ATP and ROS Levels. Cancers. 2020; 12(5):1329. https://doi.org/10.3390/cancers12051329
Chicago/Turabian StyleYang, Wookyeom, Ha-Yeon Shin, Hanbyoul Cho, Joon-Yong Chung, Eun-ju Lee, Jae-Hoon Kim, and Eun-Suk Kang. 2020. "TOM40 Inhibits Ovarian Cancer Cell Growth by Modulating Mitochondrial Function Including Intracellular ATP and ROS Levels" Cancers 12, no. 5: 1329. https://doi.org/10.3390/cancers12051329
APA StyleYang, W., Shin, H.-Y., Cho, H., Chung, J.-Y., Lee, E.-j., Kim, J.-H., & Kang, E.-S. (2020). TOM40 Inhibits Ovarian Cancer Cell Growth by Modulating Mitochondrial Function Including Intracellular ATP and ROS Levels. Cancers, 12(5), 1329. https://doi.org/10.3390/cancers12051329







