Targeting Protein Synthesis in Colorectal Cancer
Abstract
1. Introduction
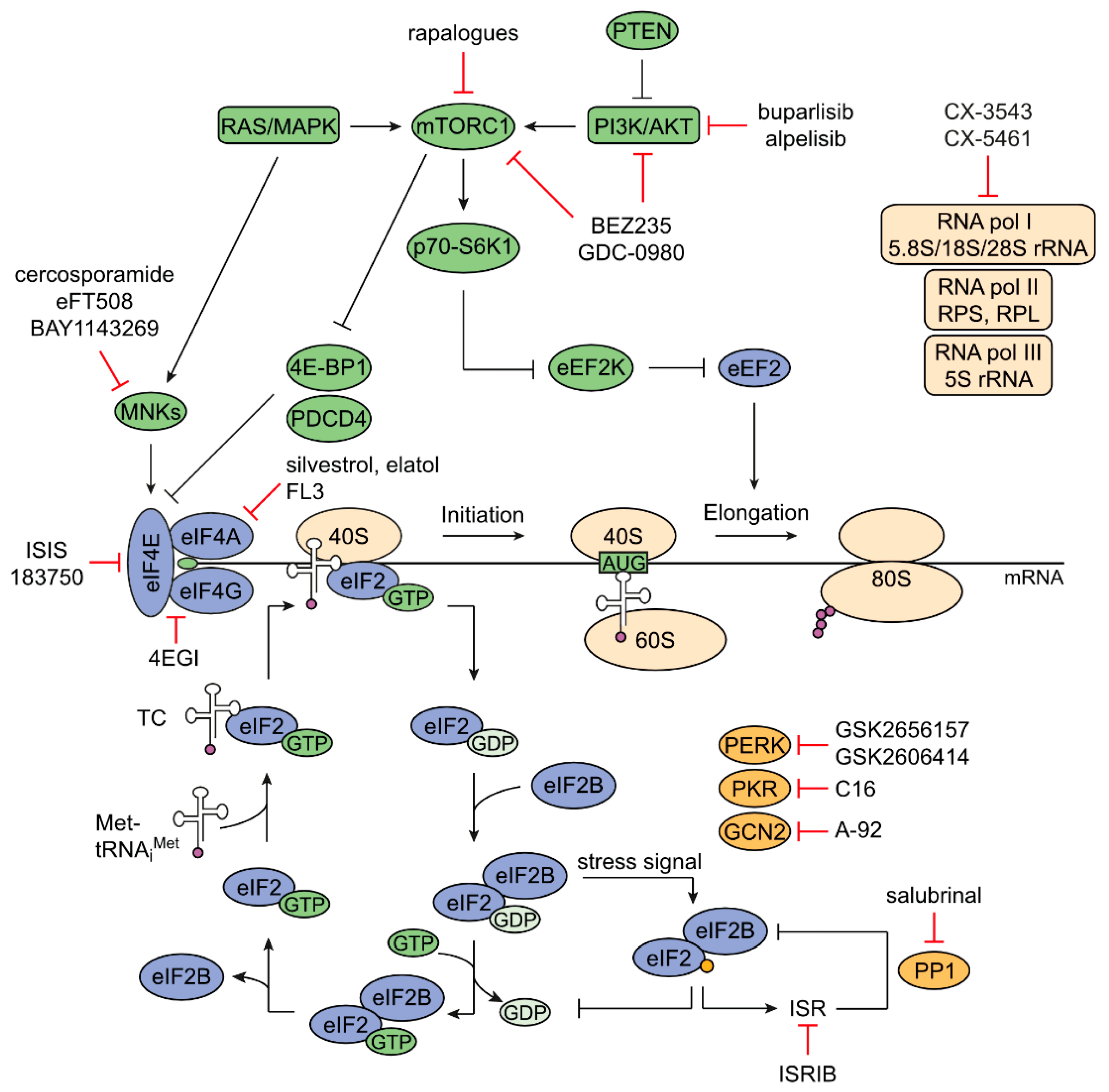
2. Mechanisms of Regulation of Protein Synthesis
2.1. Translation Initiation
2.2. Regulation and Alternative Pathways of Initiation
2.3. Elongation and Termination
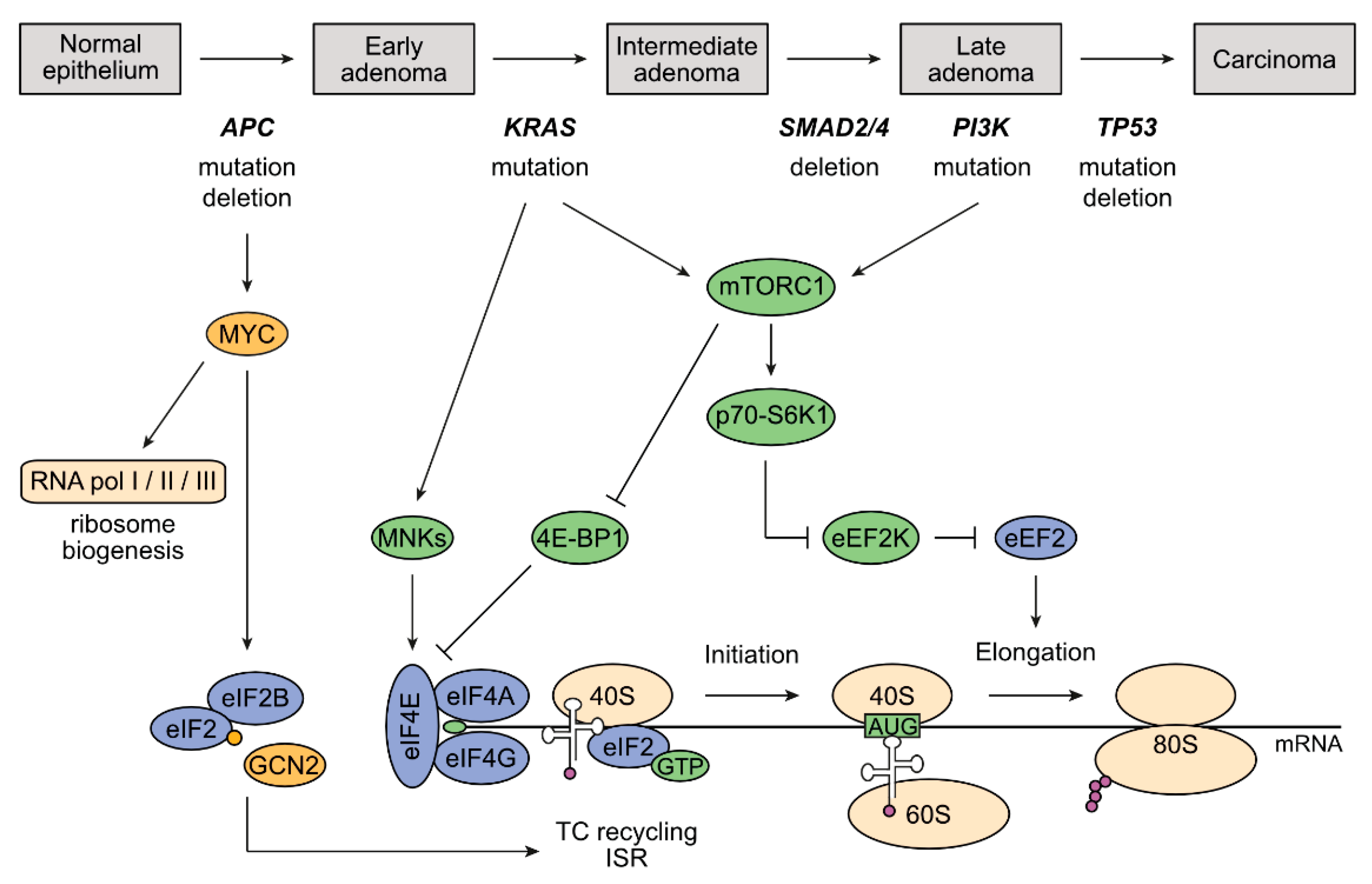
3. The Adenoma-Carcinoma Sequence and Its Impact on Deregulation of Protein Synthesis in CRC
4. Deregulation of Protein Synthesis in CRC and Potential Therapeutic Strategies
4.1. Deregulation of Ribosome Biogenesis in CRC
Targeting Ribosome Biogenesis in CRC
4.2. Deregulation of mTOR Signaling and Translation Factors in CRC
4.3. Targeting mTOR Signaling and Translation Factors in CRC
4.4. Deregulation of the Translation Initiation Factors eIF2/eIF2B and the ISR in CRC
4.5. Targeting the Translation Initiation Factors eIF2/eIF2B and the ISR in CRC
5. Clinical Advances in Targeting Protein Synthesis in CRC
5.1. Clinical Data for Targeting mTOR Signaling in CRC
5.1.1. mTORC1 Inhibitors—Rapalogues
5.1.2. Single PI3K and Dual PI3K/mTOR Inhibitors
5.2. MNK1/2 Inhibitors in CRC
5.3. Antisense Oligonucleotides Against eIF4E
6. Conclusions
Author Contributions
Funding
Conflicts of Interest
References
- Ferlay, J.; Soerjomataram, I.; Dikshit, R.; Eser, S.; Mathers, C.; Rebelo, M.; Parkin, D.M.; Forman, D.; Bray, F. Cancer incidence and mortality worldwide: Sources, methods and major patterns in GLOBOCAN 2012. Int. J. Cancer 2015, 136, E359–E386. [Google Scholar] [CrossRef] [PubMed]
- Siegel, R.L.; Miller, K.D.; Fedewa, S.A.; Ahnen, D.J.; Meester, R.G.S.; Barzi, A.; Jemal, A. Colorectal cancer statistics, 2017. CA Cancer J. Clin. 2017, 67, 177–193. [Google Scholar] [CrossRef] [PubMed]
- Wiegering, A.; Isbert, C.; Dietz, U.A.; Kunzmann, V.; Ackermann, S.; Kerscher, A.; Maeder, U.; Flentje, M.; Schlegel, N.; Reibetanz, J.; et al. Multimodal therapy in treatment of rectal cancer is associated with improved survival and reduced local recurrence—A retrospective analysis over two decades. BMC Cancer 2014, 14, 816. [Google Scholar] [CrossRef] [PubMed]
- McQuade, R.M.; Stojanovska, V.; Bornstein, J.C.; Nurgali, K. Colorectal Cancer Chemotherapy: The Evolution of Treatment and New Approaches. Curr. Med. Chem. 2017, 24, 1537–1557. [Google Scholar] [CrossRef]
- Fearon, E.R.; Vogelstein, B. A genetic model for colorectal tumorigenesis. Cell 1990, 61, 759–767. [Google Scholar] [CrossRef]
- Cancer Genome Atlas, N. Comprehensive molecular characterization of human colon and rectal cancer. Nature 2012, 487, 330–337. [Google Scholar] [CrossRef]
- Hagland, H.R.; Berg, M.; Jolma, I.W.; Carlsen, A.; Soreide, K. Molecular pathways and cellular metabolism in colorectal cancer. Dig. Surg. 2013, 30, 12–25. [Google Scholar] [CrossRef]
- Chu, J.; Cargnello, M.; Topisirovic, I.; Pelletier, J. Translation Initiation Factors: Reprogramming Protein Synthesis in Cancer. Trends Cell Biol. 2016, 26, 918–933. [Google Scholar] [CrossRef]
- Truitt, M.L.; Ruggero, D. New frontiers in translational control of the cancer genome. Nat. Rev. Cancer 2016, 16, 288–304. [Google Scholar] [CrossRef]
- Kim, H.J. Cell Fate Control by Translation: mRNA Translation Initiation as a Therapeutic Target for Cancer Development and Stem Cell Fate Control. Biomolecules 2019, 9, 665. [Google Scholar] [CrossRef]
- Faller, W.J.; Jackson, T.J.; Knight, J.R.; Ridgway, R.A.; Jamieson, T.; Karim, S.A.; Jones, C.; Radulescu, S.; Huels, D.J.; Myant, K.B.; et al. mTORC1-mediated translational elongation limits intestinal tumour initiation and growth. Nature 2015, 517, 497–500. [Google Scholar] [CrossRef] [PubMed]
- Schmidt, S.; Gay, D.; Uthe, F.W.; Denk, S.; Paauwe, M.; Matthes, N.; Diefenbacher, M.E.; Bryson, S.; Warrander, F.C.; Erhard, F.; et al. A MYC-GCN2-eIF2alpha negative feedback loop limits protein synthesis to prevent MYC-dependent apoptosis in colorectal cancer. Nat. Cell Biol. 2019, 21, 1413–1424. [Google Scholar] [CrossRef] [PubMed]
- Heys, S.D.; Park, K.G.; McNurlan, M.A.; Keenan, R.A.; Miller, J.D.; Eremin, O.; Garlick, P.J. Protein synthesis rates in colon and liver: Stimulation by gastrointestinal pathologies. Gut 1992, 33, 976–981. [Google Scholar] [CrossRef] [PubMed]
- Kafri, M.; Metzl-Raz, E.; Jona, G.; Barkai, N. The Cost of Protein Production. Cell Rep. 2016, 14, 22–31. [Google Scholar] [CrossRef] [PubMed]
- Jackson, R.J.; Hellen, C.U.; Pestova, T.V. The mechanism of eukaryotic translation initiation and principles of its regulation. Nat. Rev. Mol. Cell Biol. 2010, 11, 113–127. [Google Scholar] [CrossRef] [PubMed]
- Lomakin, I.B.; Kolupaeva, V.G.; Marintchev, A.; Wagner, G.; Pestova, T.V. Position of eukaryotic initiation factor eIF1 on the 40S ribosomal subunit determined by directed hydroxyl radical probing. Genes Dev. 2003, 17, 2786–2797. [Google Scholar] [CrossRef]
- Siridechadilok, B.; Fraser, C.S.; Hall, R.J.; Doudna, J.A.; Nogales, E. Structural roles for human translation factor eIF3 in initiation of protein synthesis. Science 2005, 310, 1513–1515. [Google Scholar] [CrossRef]
- Passmore, L.A.; Schmeing, T.M.; Maag, D.; Applefield, D.J.; Acker, M.G.; Algire, M.A.; Lorsch, J.R.; Ramakrishnan, V. The eukaryotic translation initiation factors eIF1 and eIF1A induce an open conformation of the 40S ribosome. Mol. Cell 2007, 26, 41–50. [Google Scholar] [CrossRef]
- Pelletier, J.; Graff, J.; Ruggero, D.; Sonenberg, N. Targeting the eIF4F translation initiation complex: A critical nexus for cancer development. Cancer Res. 2015, 75, 250–263. [Google Scholar] [CrossRef]
- LeFebvre, A.K.; Korneeva, N.L.; Trutschl, M.; Cvek, U.; Duzan, R.D.; Bradley, C.A.; Hershey, J.W.; Rhoads, R.E. Translation initiation factor eIF4G-1 binds to eIF3 through the eIF3e subunit. J. Biol. Chem. 2006, 281, 22917–22932. [Google Scholar] [CrossRef]
- Kozlov, G.; Trempe, J.F.; Khaleghpour, K.; Kahvejian, A.; Ekiel, I.; Gehring, K. Structure and function of the C-terminal PABC domain of human poly(A)-binding protein. Proc. Natl. Acad. Sci. USA 2001, 98, 4409–4413. [Google Scholar] [CrossRef] [PubMed]
- Deo, R.C.; Sonenberg, N.; Burley, S.K. X-ray structure of the human hyperplastic discs protein: An ortholog of the C-terminal domain of poly(A)-binding protein. Proc. Natl. Acad. Sci. USA 2001, 98, 4414–4419. [Google Scholar] [CrossRef] [PubMed]
- Algire, M.A.; Maag, D.; Lorsch, J.R. Pi release from eIF2, not GTP hydrolysis, is the step controlled by start-site selection during eukaryotic translation initiation. Mol. Cell 2005, 20, 251–262. [Google Scholar] [CrossRef]
- Lind, C.; Aqvist, J. Principles of start codon recognition in eukaryotic translation initiation. Nucleic Acids Res. 2016, 44, 8425–8432. [Google Scholar] [CrossRef]
- Kapp, L.D.; Lorsch, J.R. GTP-dependent recognition of the methionine moiety on initiator tRNA by translation factor eIF2. J. Mol. Biol. 2004, 335, 923–936. [Google Scholar] [CrossRef]
- Unbehaun, A.; Borukhov, S.I.; Hellen, C.U.; Pestova, T.V. Release of initiation factors from 48S complexes during ribosomal subunit joining and the link between establishment of codon-anticodon base-pairing and hydrolysis of eIF2-bound GTP. Genes Dev. 2004, 18, 3078–3093. [Google Scholar] [CrossRef] [PubMed]
- Pestova, T.V.; Lomakin, I.B.; Lee, J.H.; Choi, S.K.; Dever, T.E.; Hellen, C.U. The joining of ribosomal subunits in eukaryotes requires eIF5B. Nature 2000, 403, 332–335. [Google Scholar] [CrossRef] [PubMed]
- Haghighat, A.; Mader, S.; Pause, A.; Sonenberg, N. Repression of cap-dependent translation by 4E-binding protein 1: Competition with p220 for binding to eukaryotic initiation factor-4E. EMBO J. 1995, 14, 5701–5709. [Google Scholar] [CrossRef] [PubMed]
- Siddiqui, N.; Sonenberg, N. Signalling to eIF4E in cancer. Biochem. Soc. Trans. 2015, 43, 763–772. [Google Scholar] [CrossRef]
- Pavitt, G.D. eIF2B, a mediator of general and gene-specific translational control. Biochem. Soc. Trans. 2005, 33, 1487–1492. [Google Scholar] [CrossRef]
- Ryoo, H.D.; Vasudevan, D. Two distinct nodes of translational inhibition in the Integrated Stress Response. BMB Rep. 2017, 50, 539–545. [Google Scholar] [CrossRef] [PubMed]
- Rodnina, M.V.; Wintermeyer, W. Recent mechanistic insights into eukaryotic ribosomes. Curr. Opin. Cell Biol. 2009, 21, 435–443. [Google Scholar] [CrossRef] [PubMed]
- Dever, T.E.; Green, R. The elongation, termination, and recycling phases of translation in eukaryotes. Cold Spring Harb. Perspect. Biol. 2012, 4, a013706. [Google Scholar] [CrossRef] [PubMed]
- Vogelstein, B.; Fearon, E.R.; Hamilton, S.R.; Kern, S.E.; Preisinger, A.C.; Leppert, M.; Nakamura, Y.; White, R.; Smits, A.M.; Bos, J.L. Genetic alterations during colorectal-tumor development. New Engl. J. Med. 1988, 319, 525–532. [Google Scholar] [CrossRef]
- Fearon, E.R. Molecular genetics of colorectal cancer. Annu. Rev. Pathol. 2011, 6, 479–507. [Google Scholar] [CrossRef]
- Sansom, O.J.; Meniel, V.S.; Muncan, V.; Phesse, T.J.; Wilkins, J.A.; Reed, K.R.; Vass, J.K.; Athineos, D.; Clevers, H.; Clarke, A.R. Myc deletion rescues Apc deficiency in the small intestine. Nature 2007, 446, 676–679. [Google Scholar] [CrossRef]
- Grandori, C.; Gomez-Roman, N.; Felton-Edkins, Z.A.; Ngouenet, C.; Galloway, D.A.; Eisenman, R.N.; White, R.J. c-Myc binds to human ribosomal DNA and stimulates transcription of rRNA genes by RNA polymerase I. Nat. Cell Biol. 2005, 7, 311–318. [Google Scholar] [CrossRef]
- Van Riggelen, J.; Yetil, A.; Felsher, D.W. MYC as a regulator of ribosome biogenesis and protein synthesis. Nat. Rev. Cancer 2010, 10, 301–309. [Google Scholar] [CrossRef]
- Campbell, K.J.; White, R.J. MYC regulation of cell growth through control of transcription by RNA polymerases I and III. Cold Spring Harb. Perspect. Med. 2014, 4. [Google Scholar] [CrossRef]
- Oskarsson, T.; Trumpp, A. The Myc trilogy: Lord of RNA polymerases. Nat. Cell Biol. 2005, 7, 215–217. [Google Scholar] [CrossRef]
- Gomez-Roman, N.; Grandori, C.; Eisenman, R.N.; White, R.J. Direct activation of RNA polymerase III transcription by c-Myc. Nature 2003, 421, 290–294. [Google Scholar] [CrossRef]
- Lai, M.D.; Xu, J. Ribosomal proteins and colorectal cancer. Curr. Genom. 2007, 8, 43–49. [Google Scholar] [CrossRef]
- Berkel, H.J.; Turbat-Herrera, E.A.; Shi, R.; de Benedetti, A. Expression of the translation initiation factor eIF4E in the polyp-cancer sequence in the colon. Cancer Epidemiol. Biomark. Prev. 2001, 10, 663–666. [Google Scholar]
- Xu, T.; Zong, Y.; Peng, L.; Kong, S.; Zhou, M.; Zou, J.; Liu, J.; Miao, R.; Sun, X.; Li, L. Overexpression of eIF4E in colorectal cancer patients is associated with liver metastasis. Onco Targets Ther. 2016, 9, 815–822. [Google Scholar] [CrossRef] [PubMed]
- Gao, M.; Zhang, X.; Li, D.; He, P.; Tian, W.; Zeng, B. Expression analysis and clinical significance of eIF4E, VEGF-C, E-cadherin and MMP-2 in colorectal adenocarcinoma. Oncotarget 2016, 7, 85502–85514. [Google Scholar] [CrossRef] [PubMed]
- Roux, P.P.; Topisirovic, I. Signaling Pathways Involved in the Regulation of mRNA Translation. Mol. Cell. Biol. 2018, 38. [Google Scholar] [CrossRef]
- Lu, Q.; Wang, J.; Yu, G.; Guo, T.; Hu, C.; Ren, P. Expression and clinical significance of mammalian target of rapamycin/P70 ribosomal protein S6 kinase signaling pathway in human colorectal carcinoma tissue. Oncol. Lett. 2015, 10, 277–282. [Google Scholar] [CrossRef]
- Miao, Y.; Chen, L.; Shi, C.; Fan, R.; Chen, P.; Liu, H.; Xia, A.; Qian, H. Increased phosphorylation of 4Ebinding protein 1 predicts poor prognosis for patients with colorectal cancer. Mol. Med. Rep. 2017, 15, 3099–3104. [Google Scholar] [CrossRef]
- Chen, Y.; Wang, J.; Fan, H.; Xie, J.; Xu, L.; Zhou, B. Phosphorylated 4E-BP1 is associated with tumor progression and adverse prognosis in colorectal cancer. Neoplasma 2017, 64, 787–794. [Google Scholar] [CrossRef]
- Fujishita, T.; Aoki, K.; Lane, H.A.; Aoki, M.; Taketo, M.M. Inhibition of the mTORC1 pathway suppresses intestinal polyp formation and reduces mortality in ApcDelta716 mice. Proc. Natl. Acad. Sci. USA 2008, 105, 13544–13549. [Google Scholar] [CrossRef]
- Khatter, H.; Myasnikov, A.G.; Natchiar, S.K.; Klaholz, B.P. Structure of the human 80S ribosome. Nature 2015, 520, 640–645. [Google Scholar] [CrossRef] [PubMed]
- Chester, K.A.; Robson, L.; Begent, R.H.; Talbot, I.C.; Pringle, J.H.; Primrose, L.; Macpherson, A.J.; Boxer, G.; Southall, P.; Malcolm, A.D. Identification of a human ribosomal protein mRNA with increased expression in colorectal tumours. Biochim. Biophys. Acta 1989, 1009, 297–300. [Google Scholar] [CrossRef]
- Elvin, P.; Kerr, I.B.; McArdle, C.S.; Birnie, G.D. Isolation and preliminary characterisation of cDNA clones representing mRNAs associated with tumour progression and metastasis in colorectal cancer. Br. J. Cancer 1988, 57, 36–42. [Google Scholar] [CrossRef] [PubMed]
- Pogue-Geile, K.; Geiser, J.R.; Shu, M.; Miller, C.; Wool, I.G.; Meisler, A.I.; Pipas, J.M. Ribosomal protein genes are overexpressed in colorectal cancer: Isolation of a cDNA clone encoding the human S3 ribosomal protein. Mol. Cell. Biol. 1991, 11, 3842–3849. [Google Scholar] [CrossRef] [PubMed]
- Guimaraes, J.C.; Zavolan, M. Patterns of ribosomal protein expression specify normal and malignant human cells. Genome Biol. 2016, 17, 236. [Google Scholar] [CrossRef]
- Wang, Y.; Sui, J.; Li, X.; Cao, F.; He, J.; Yang, B.; Zhu, X.; Sun, Y.; Pu, Y.D. RPS24 knockdown inhibits colorectal cancer cell migration and proliferation in vitro. Gene 2015, 571, 286–291. [Google Scholar] [CrossRef]
- Dong, Z.; Jiang, H.; Liang, S.; Wang, Y.; Jiang, W.; Zhu, C. Ribosomal Protein L15 is involved in Colon Carcinogenesis. Int. J. Med. Sci. 2019, 16, 1132–1141. [Google Scholar] [CrossRef]
- Lindeboom, R.G.; van Voorthuijsen, L.; Oost, K.C.; Rodriguez-Colman, M.J.; Luna-Velez, M.V.; Furlan, C.; Baraille, F.; Jansen, P.W.; Ribeiro, A.; Burgering, B.M.; et al. Integrative multi-omics analysis of intestinal organoid differentiation. Mol. Syst. Biol. 2018, 14, e8227. [Google Scholar] [CrossRef]
- Goudarzi, K.M.; Lindstrom, M.S. Role of ribosomal protein mutations in tumor development (Review). Int. J. Oncol. 2016, 48, 1313–1324. [Google Scholar] [CrossRef]
- Nieminen, T.T.; O’Donohue, M.F.; Wu, Y.; Lohi, H.; Scherer, S.W.; Paterson, A.D.; Ellonen, P.; Abdel-Rahman, W.M.; Valo, S.; Mecklin, J.P.; et al. Germline mutation of RPS20, encoding a ribosomal protein, causes predisposition to hereditary nonpolyposis colorectal carcinoma without DNA mismatch repair deficiency. Gastroenterology 2014, 147, 595–598.e5. [Google Scholar] [CrossRef]
- Ferreira, A.M.; Tuominen, I.; van Dijk-Bos, K.; Sanjabi, B.; van der Sluis, T.; van der Zee, A.G.; Hollema, H.; Zazula, M.; Sijmons, R.H.; Aaltonen, L.A.; et al. High frequency of RPL22 mutations in microsatellite-unstable colorectal and endometrial tumors. Hum. Mutat. 2014, 35, 1442–1445. [Google Scholar] [CrossRef] [PubMed]
- Gaviraghi, M.; Vivori, C.; Tonon, G. How Cancer Exploits Ribosomal RNA Biogenesis: A Journey beyond the Boundaries of rRNA Transcription. Cells 2019, 8, 1098. [Google Scholar] [CrossRef] [PubMed]
- Gomez-Roman, N.; Felton-Edkins, Z.A.; Kenneth, N.S.; Goodfellow, S.J.; Athineos, D.; Zhang, J.; Ramsbottom, B.A.; Innes, F.; Kantidakis, T.; Kerr, E.R.; et al. Activation by c-Myc of transcription by RNA polymerases I, II and III. Biochem. Soc. Symp. 2006, 141–154. [Google Scholar] [CrossRef]
- Schaub, F.X.; Dhankani, V.; Berger, A.C.; Trivedi, M.; Richardson, A.B.; Shaw, R.; Zhao, W.; Zhang, X.; Ventura, A.; Liu, Y.; et al. Pan-cancer Alterations of the MYC Oncogene and Its Proximal Network across the Cancer Genome Atlas. Cell Syst. 2018, 6, 282–300.e2. [Google Scholar] [CrossRef]
- Morcelle, C.; Menoyo, S.; Moron-Duran, F.D.; Tauler, A.; Kozma, S.C.; Thomas, G.; Gentilella, A. Oncogenic MYC Induces the Impaired Ribosome Biogenesis Checkpoint and Stabilizes p53 Independent of Increased Ribosome Content. Cancer Res. 2019, 79, 4348–4359. [Google Scholar] [CrossRef]
- Tsoi, H.; Lam, K.C.; Dong, Y.; Zhang, X.; Lee, C.K.; Zhang, J.; Ng, S.C.; Ng, S.S.M.; Zheng, S.; Chen, Y.; et al. Pre-45s rRNA promotes colon cancer and is associated with poor survival of CRC patients. Oncogene 2017, 36, 6109–6118. [Google Scholar] [CrossRef]
- Telechea-Fernandez, M.; Rodriguez-Fernandez, L.; Garcia, C.; Zaragoza, R.; Vina, J.; Cervantes, A.; Garcia-Trevijano, E.R. New localization and function of calpain-2 in nucleoli of colorectal cancer cells in ribosomal biogenesis: Effect of KRAS status. Oncotarget 2018, 9, 9100–9113. [Google Scholar] [CrossRef][Green Version]
- Charitou, T.; Srihari, S.; Lynn, M.A.; Jarboui, M.A.; Fasterius, E.; Moldovan, M.; Shirasawa, S.; Tsunoda, T.; Ueffing, M.; Xie, J.; et al. Transcriptional and metabolic rewiring of colorectal cancer cells expressing the oncogenic KRAS(G13D) mutation. Br. J. Cancer 2019, 121, 37–50. [Google Scholar] [CrossRef]
- Wang, X.W.; Zhang, Y.J. Targeting mTOR network in colorectal cancer therapy. World J. Gastroenterol. 2014, 20, 4178–4188. [Google Scholar] [CrossRef]
- Greenhalgh, D.A.; Parish, J.H. Effect of 5-fluorouracil combination therapy on RNA processing in human colonic carcinoma cells. Br. J. Cancer 1990, 61, 415–419. [Google Scholar] [CrossRef]
- Bash-Imam, Z.; Therizols, G.; Vincent, A.; Laforets, F.; Polay Espinoza, M.; Pion, N.; Macari, F.; Pannequin, J.; David, A.; Saurin, J.C.; et al. Translational reprogramming of colorectal cancer cells induced by 5-fluorouracil through a miRNA-dependent mechanism. Oncotarget 2017, 8, 46219–46233. [Google Scholar] [CrossRef] [PubMed]
- Bruno, P.M.; Liu, Y.; Park, G.Y.; Murai, J.; Koch, C.E.; Eisen, T.J.; Pritchard, J.R.; Pommier, Y.; Lippard, S.J.; Hemann, M.T. A subset of platinum-containing chemotherapeutic agents kills cells by inducing ribosome biogenesis stress. Nat. Med. 2017, 23, 461–471. [Google Scholar] [CrossRef] [PubMed]
- Burger, K.; Muhl, B.; Harasim, T.; Rohrmoser, M.; Malamoussi, A.; Orban, M.; Kellner, M.; Gruber-Eber, A.; Kremmer, E.; Holzel, M.; et al. Chemotherapeutic drugs inhibit ribosome biogenesis at various levels. J. Biol. Chem. 2010, 285, 12416–12425. [Google Scholar] [CrossRef] [PubMed]
- Ghoshal, K.; Jacob, S.T. An alternative molecular mechanism of action of 5-fluorouracil, a potent anticancer drug. Biochem. Pharmacol. 1997, 53, 1569–1575. [Google Scholar] [CrossRef]
- Rixe, O.; Ortuzar, W.; Alvarez, M.; Parker, R.; Reed, E.; Paull, K.; Fojo, T. Oxaliplatin, tetraplatin, cisplatin, and carboplatin: Spectrum of activity in drug-resistant cell lines and in the cell lines of the National Cancer Institute’s Anticancer Drug Screen panel. Biochem. Pharmacol. 1996, 52, 1855–1865. [Google Scholar] [CrossRef]
- Machover, D.; Diaz-Rubio, E.; de Gramont, A.; Schilf, A.; Gastiaburu, J.J.; Brienza, S.; Itzhaki, M.; Metzger, G.; N’Daw, D.; Vignoud, J.; et al. Two consecutive phase II studies of oxaliplatin (L-OHP) for treatment of patients with advanced colorectal carcinoma who were resistant to previous treatment with fluoropyrimidines. Ann. Oncol. 1996, 7, 95–98. [Google Scholar] [CrossRef]
- Drygin, D.; Lin, A.; Bliesath, J.; Ho, C.B.; O’Brien, S.E.; Proffitt, C.; Omori, M.; Haddach, M.; Schwaebe, M.K.; Siddiqui-Jain, A.; et al. Targeting RNA polymerase I with an oral small molecule CX-5461 inhibits ribosomal RNA synthesis and solid tumor growth. Cancer Res. 2011, 71, 1418–1430. [Google Scholar] [CrossRef]
- Drygin, D.; Siddiqui-Jain, A.; O’Brien, S.; Schwaebe, M.; Lin, A.; Bliesath, J.; Ho, C.B.; Proffitt, C.; Trent, K.; Whitten, J.P.; et al. Anticancer activity of CX-3543: A direct inhibitor of rRNA biogenesis. Cancer Res. 2009, 69, 7653–7661. [Google Scholar] [CrossRef]
- Haddach, M.; Schwaebe, M.K.; Michaux, J.; Nagasawa, J.; O’Brien, S.E.; Whitten, J.P.; Pierre, F.; Kerdoncuff, P.; Darjania, L.; Stansfield, R.; et al. Discovery of CX-5461, the First Direct and Selective Inhibitor of RNA Polymerase I, for Cancer Therapeutics. ACS Med. Chem. Lett. 2012, 3, 602–606. [Google Scholar] [CrossRef]
- Pelletier, J.; Thomas, G.; Volarevic, S. Corrigendum: Ribosome biogenesis in cancer: New players and therapeutic avenues. Nat. Rev. Cancer 2018, 18, 134. [Google Scholar] [CrossRef]
- Pelletier, J.; Thomas, G.; Volarevic, S. Ribosome biogenesis in cancer: New players and therapeutic avenues. Nat. Rev. Cancer 2018, 18, 51–63. [Google Scholar] [CrossRef] [PubMed]
- Ismael, M.; Webb, R.; Ajaz, M.; Kirkby, K.J.; Coley, H.M. The Targeting of RNA Polymerase I Transcription Using CX-5461 in Combination with Radiation Enhances Tumour Cell Killing Effects in Human Solid Cancers. Cancers 2019, 11, 1429. [Google Scholar] [CrossRef] [PubMed]
- Xu, H.; Di Antonio, M.; McKinney, S.; Mathew, V.; Ho, B.; O’Neil, N.J.; Santos, N.D.; Silvester, J.; Wei, V.; Garcia, J.; et al. CX-5461 is a DNA G-quadruplex stabilizer with selective lethality in BRCA1/2 deficient tumours. Nat. Commun. 2017, 8, 14432. [Google Scholar] [CrossRef] [PubMed]
- Reilly, N.M.; Novara, L.; Di Nicolantonio, F.; Bardelli, A. Exploiting DNA repair defects in colorectal cancer. Mol. Oncol. 2019, 13, 681–700. [Google Scholar] [CrossRef] [PubMed]
- Stedman, A.; Beck-Cormier, S.; Le Bouteiller, M.; Raveux, A.; Vandormael-Pournin, S.; Coqueran, S.; Lejour, V.; Jarzebowski, L.; Toledo, F.; Robine, S.; et al. Ribosome biogenesis dysfunction leads to p53-mediated apoptosis and goblet cell differentiation of mouse intestinal stem/progenitor cells. Cell Death Differ. 2015, 22, 1865–1876. [Google Scholar] [CrossRef]
- Francipane, M.G.; Lagasse, E. mTOR pathway in colorectal cancer: An update. Oncotarget 2014, 5, 49–66. [Google Scholar] [CrossRef]
- Johnson, S.M.; Gulhati, P.; Rampy, B.A.; Han, Y.; Rychahou, P.G.; Doan, H.Q.; Weiss, H.L.; Evers, B.M. Novel expression patterns of PI3K/Akt/mTOR signaling pathway components in colorectal cancer. J. Am. Coll. Surg. 2010, 210, 767–776. [Google Scholar] [CrossRef]
- Caputo, F.; Santini, C.; Bardasi, C.; Cerma, K.; Casadei-Gardini, A.; Spallanzani, A.; Andrikou, K.; Cascinu, S.; Gelsomino, F. BRAF-Mutated Colorectal Cancer: Clinical and Molecular Insights. Int. J. Mol. Sci. 2019, 20, 5369. [Google Scholar] [CrossRef]
- Grabiner, B.C.; Nardi, V.; Birsoy, K.; Possemato, R.; Shen, K.; Sinha, S.; Jordan, A.; Beck, A.H.; Sabatini, D.M. A diverse array of cancer-associated MTOR mutations are hyperactivating and can predict rapamycin sensitivity. Cancer Discov. 2014, 4, 554–563. [Google Scholar] [CrossRef]
- Sato, T.; Nakashima, A.; Guo, L.; Coffman, K.; Tamanoi, F. Single amino-acid changes that confer constitutive activation of mTOR are discovered in human cancer. Oncogene 2010, 29, 2746–2752. [Google Scholar] [CrossRef]
- She, Q.B.; Halilovic, E.; Ye, Q.; Zhen, W.; Shirasawa, S.; Sasazuki, T.; Solit, D.B.; Rosen, N. 4E-BP1 is a key effector of the oncogenic activation of the AKT and ERK signaling pathways that integrates their function in tumors. Cancer Cell 2010, 18, 39–51. [Google Scholar] [CrossRef] [PubMed]
- Showkat, M.; Beigh, M.A.; Andrabi, K.I. mTOR Signaling in Protein Translation Regulation: Implications in Cancer Genesis and Therapeutic Interventions. Mol. Biol. Int. 2014, 2014, 686984. [Google Scholar] [CrossRef] [PubMed]
- Wiesweg, M.; Reis, H.; Koster, T.; Goetz, M.; Worm, K.; Herold, T.; Paul, A.; Dechene, A.; Schumacher, B.; Markus, P.; et al. Phosphorylation of p70 Ribosomal Protein S6 Kinase beta-1 is an Independent Prognostic Parameter in Metastatic Colorectal Cancer. Clin. Colorectal Cancer 2018, 17, e331–e352. [Google Scholar] [CrossRef]
- Dorrello, N.V.; Peschiaroli, A.; Guardavaccaro, D.; Colburn, N.H.; Sherman, N.E.; Pagano, M. S6K1- and betaTRCP-mediated degradation of PDCD4 promotes protein translation and cell growth. Science 2006, 314, 467–471. [Google Scholar] [CrossRef] [PubMed]
- Wang, X.; Li, W.; Williams, M.; Terada, N.; Alessi, D.R.; Proud, C.G. Regulation of elongation factor 2 kinase by p90(RSK1) and p70 S6 kinase. EMBO J. 2001, 20, 4370–4379. [Google Scholar] [CrossRef]
- Long, J.; Yin, Y.; Guo, H.; Li, S.; Sun, Y.; Zeng, C.; Zhu, W. The mechanisms and clinical significance of PDCD4 in colorectal cancer. Gene 2019, 680, 59–64. [Google Scholar] [CrossRef]
- Wiegering, A.; Uthe, F.W.; Jamieson, T.; Ruoss, Y.; Huttenrauch, M.; Kuspert, M.; Pfann, C.; Nixon, C.; Herold, S.; Walz, S.; et al. Targeting Translation Initiation Bypasses Signaling Crosstalk Mechanisms That Maintain High MYC Levels in Colorectal Cancer. Cancer Discov. 2015, 5, 768–781. [Google Scholar] [CrossRef]
- Mudduluru, G.; Medved, F.; Grobholz, R.; Jost, C.; Gruber, A.; Leupold, J.H.; Post, S.; Jansen, A.; Colburn, N.H.; Allgayer, H. Loss of programmed cell death 4 expression marks adenoma-carcinoma transition, correlates inversely with phosphorylated protein kinase B, and is an independent prognostic factor in resected colorectal cancer. Cancer 2007, 110, 1697–1707. [Google Scholar] [CrossRef]
- Yang, H.S.; Jansen, A.P.; Komar, A.A.; Zheng, X.; Merrick, W.C.; Costes, S.; Lockett, S.J.; Sonenberg, N.; Colburn, N.H. The transformation suppressor Pdcd4 is a novel eukaryotic translation initiation factor 4A binding protein that inhibits translation. Mol. Cell. Biol. 2003, 23, 26–37. [Google Scholar] [CrossRef]
- Ryazanov, A.G.; Shestakova, E.A.; Natapov, P.G. Phosphorylation of elongation factor 2 by EF-2 kinase affects rate of translation. Nature 1988, 334, 170–173. [Google Scholar] [CrossRef]
- Ng, T.H.; Sham, K.W.Y.; Xie, C.M.; Ng, S.S.M.; To, K.F.; Tong, J.H.M.; Liu, W.Y.Z.; Zhang, L.; Chan, M.T.V.; Wu, W.K.K.; et al. Eukaryotic elongation factor-2 kinase expression is an independent prognostic factor in colorectal cancer. BMC Cancer 2019, 19, 649. [Google Scholar] [CrossRef]
- Xie, C.M.; Liu, X.Y.; Sham, K.W.; Lai, J.M.; Cheng, C.H. Silencing of EEF2K (eukaryotic elongation factor-2 kinase) reveals AMPK-ULK1-dependent autophagy in colon cancer cells. Autophagy 2014, 10, 1495–1508. [Google Scholar] [CrossRef]
- Nakamura, J.; Aoyagi, S.; Nanchi, I.; Nakatsuka, S.; Hirata, E.; Shibata, S.; Fukuda, M.; Yamamoto, Y.; Fukuda, I.; Tatsumi, N.; et al. Overexpression of eukaryotic elongation factor eEF2 in gastrointestinal cancers and its involvement in G2/M progression in the cell cycle. Int. J. Oncol. 2009, 34, 1181–1189. [Google Scholar]
- Oji, Y.; Tatsumi, N.; Fukuda, M.; Nakatsuka, S.; Aoyagi, S.; Hirata, E.; Nanchi, I.; Fujiki, F.; Nakajima, H.; Yamamoto, Y.; et al. The translation elongation factor eEF2 is a novel tumorassociated antigen overexpressed in various types of cancers. Int. J. Oncol. 2014, 44, 1461–1469. [Google Scholar] [CrossRef]
- Hassan, M.K.; Kumar, D.; Naik, M.; Dixit, M. The expression profile and prognostic significance of eukaryotic translation elongation factors in different cancers. PLoS ONE 2018, 13, e0191377. [Google Scholar] [CrossRef]
- Ali, M.U.; Ur Rahman, M.S.; Jia, Z.; Jiang, C. Eukaryotic translation initiation factors and cancer. Tumour Biol. 2017, 39, 1010428317709805. [Google Scholar] [CrossRef]
- Yang, S.X.; Hewitt, S.M.; Steinberg, S.M.; Liewehr, D.J.; Swain, S.M. Expression levels of eIF4E, VEGF, and cyclin D1, and correlation of eIF4E with VEGF and cyclin D1 in multi-tumor tissue microarray. Oncol. Rep. 2007, 17, 281–287. [Google Scholar] [CrossRef]
- Fan, S.; Ramalingam, S.S.; Kauh, J.; Xu, Z.; Khuri, F.R.; Sun, S.Y. Phosphorylated eukaryotic translation initiation factor 4 (eIF4E) is elevated in human cancer tissues. Cancer Biol. Ther. 2009, 8, 1463–1469. [Google Scholar] [CrossRef]
- Waskiewicz, A.J.; Flynn, A.; Proud, C.G.; Cooper, J.A. Mitogen-activated protein kinases activate the serine/threonine kinases Mnk1 and Mnk2. EMBO J. 1997, 16, 1909–1920. [Google Scholar] [CrossRef]
- Hou, J.; Lam, F.; Proud, C.; Wang, S. Targeting Mnks for cancer therapy. Oncotarget 2012, 3, 118–131. [Google Scholar] [CrossRef]
- Lu, W.T.; Wilczynska, A.; Smith, E.; Bushell, M. The diverse roles of the eIF4A family: You are the company you keep. Biochem. Soc. Trans. 2014, 42, 166–172. [Google Scholar] [CrossRef]
- Raza, F.; Waldron, J.A.; Quesne, J.L. Translational dysregulation in cancer: eIF4A isoforms and sequence determinants of eIF4A dependence. Biochem. Soc. Trans. 2015, 43, 1227–1233. [Google Scholar] [CrossRef]
- Li, W.; Chen, A.; Xiong, L.; Chen, T.; Tao, F.; Lu, Y.; He, Q.; Zhao, L.; Ou, R.; Xu, Y. miR-133a acts as a tumor suppressor in colorectal cancer by targeting eIF4A1. Tumour Biol. 2017, 39. [Google Scholar] [CrossRef]
- Meijer, H.A.; Kong, Y.W.; Lu, W.T.; Wilczynska, A.; Spriggs, R.V.; Robinson, S.W.; Godfrey, J.D.; Willis, A.E.; Bushell, M. Translational repression and eIF4A2 activity are critical for microRNA-mediated gene regulation. Science 2013, 340, 82–85. [Google Scholar] [CrossRef]
- Wilczynska, A.; Gillen, S.L.; Schmidt, T.; Meijer, H.A.; Jukes-Jones, R.; Langlais, C.; Kopra, K.; Lu, W.T.; Godfrey, J.D.; Hawley, B.R.; et al. eIF4A2 drives repression of translation at initiation by Ccr4-Not through purine-rich motifs in the 5′UTR. Genome Biol. 2019, 20, 262. [Google Scholar] [CrossRef]
- Galicia-Vazquez, G.; Chu, J.; Pelletier, J. eIF4AII is dispensable for miRNA-mediated gene silencing. RNA 2015, 21, 1826–1833. [Google Scholar] [CrossRef]
- Chen, Z.H.; Qi, J.J.; Wu, Q.N.; Lu, J.H.; Liu, Z.X.; Wang, Y.; Hu, P.S.; Li, T.; Lin, J.F.; Wu, X.Y.; et al. Eukaryotic initiation factor 4A2 promotes experimental metastasis and oxaliplatin resistance in colorectal cancer. J. Exp. Clin. Cancer Res. 2019, 38, 196. [Google Scholar] [CrossRef]
- Golob-Schwarzl, N.; Schweiger, C.; Koller, C.; Krassnig, S.; Gogg-Kamerer, M.; Gantenbein, N.; Toeglhofer, A.M.; Wodlej, C.; Bergler, H.; Pertschy, B.; et al. Separation of low and high grade colon and rectum carcinoma by eukaryotic translation initiation factors 1, 5 and 6. Oncotarget 2017, 8, 101224–101243. [Google Scholar] [CrossRef]
- Wu, D.; Matsushita, K.; Matsubara, H.; Nomura, F.; Tomonaga, T. An alternative splicing isoform of eukaryotic initiation factor 4H promotes tumorigenesis in vivo and is a potential therapeutic target for human cancer. Int. J. Cancer 2011, 128, 1018–1030. [Google Scholar] [CrossRef]
- Haybaeck, J.; O’Connor, T.; Spilka, R.; Spizzo, G.; Ensinger, C.; Mikuz, G.; Brunhuber, T.; Vogetseder, A.; Theurl, I.; Salvenmoser, W.; et al. Overexpression of p150, a part of the large subunit of the eukaryotic translation initiation factor 3, in colon cancer. Anticancer Res. 2010, 30, 1047–1055. [Google Scholar]
- Shi, J.; Kahle, A.; Hershey, J.W.; Honchak, B.M.; Warneke, J.A.; Leong, S.P.; Nelson, M.A. Decreased expression of eukaryotic initiation factor 3f deregulates translation and apoptosis in tumor cells. Oncogene 2006, 25, 4923–4936. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Tomlinson, I.P.; Webb, E.; Carvajal-Carmona, L.; Broderick, P.; Howarth, K.; Pittman, A.M.; Spain, S.; Lubbe, S.; Walther, A.; Sullivan, K.; et al. A genome-wide association study identifies colorectal cancer susceptibility loci on chromosomes 10p14 and 8q23.3. Nat. Genet. 2008, 40, 623–630. [Google Scholar] [CrossRef] [PubMed]
- Goh, S.H.; Hong, S.H.; Hong, S.H.; Lee, B.C.; Ju, M.H.; Jeong, J.S.; Cho, Y.R.; Kim, I.H.; Lee, Y.S. eIF3m expression influences the regulation of tumorigenesis-related genes in human colon cancer. Oncogene 2011, 30, 398–409. [Google Scholar] [CrossRef] [PubMed]
- Zhu, W.; Cai, M.Y.; Tong, Z.T.; Dong, S.S.; Mai, S.J.; Liao, Y.J.; Bian, X.W.; Lin, M.C.; Kung, H.F.; Zeng, Y.X.; et al. Overexpression of EIF5A2 promotes colorectal carcinoma cell aggressiveness by upregulating MTA1 through C-myc to induce epithelial-mesenchymaltransition. Gut 2012, 61, 562–575. [Google Scholar] [CrossRef]
- Sanvito, F.; Vivoli, F.; Gambini, S.; Santambrogio, G.; Catena, M.; Viale, E.; Veglia, F.; Donadini, A.; Biffo, S.; Marchisio, P.C. Expression of a highly conserved protein, p27BBP, during the progression of human colorectal cancer. Cancer Res. 2000, 60, 510–516. [Google Scholar]
- Lam, F.F.; Jankova, L.; Dent, O.F.; Molloy, M.P.; Kwun, S.Y.; Clarke, C.; Chapuis, P.; Robertson, G.; Beale, P.; Clarke, S.; et al. Identification of distinctive protein expression patterns in colorectal adenoma. Proteom. Clin. Appl. 2010, 4, 60–70. [Google Scholar] [CrossRef]
- Zhang, Y.J.; Dai, Q.; Sun, D.F.; Xiong, H.; Tian, X.Q.; Gao, F.H.; Xu, M.H.; Chen, G.Q.; Han, Z.G.; Fang, J.Y. mTOR signaling pathway is a target for the treatment of colorectal cancer. Ann. Surg. Oncol. 2009, 16, 2617–2628. [Google Scholar] [CrossRef]
- Gulhati, P.; Cai, Q.; Li, J.; Liu, J.; Rychahou, P.G.; Qiu, S.; Lee, E.Y.; Silva, S.R.; Bowen, K.A.; Gao, T.; et al. Targeted inhibition of mammalian target of rapamycin signaling inhibits tumorigenesis of colorectal cancer. Clin. Cancer Res. 2009, 15, 7207–7216. [Google Scholar] [CrossRef]
- He, K.; Zheng, X.; Li, M.; Zhang, L.; Yu, J. mTOR inhibitors induce apoptosis in colon cancer cells via CHOP-dependent DR5 induction on 4E-BP1 dephosphorylation. Oncogene 2016, 35, 148–157. [Google Scholar] [CrossRef]
- Kaneko, M.; Nozawa, H.; Hiyoshi, M.; Tada, N.; Murono, K.; Nirei, T.; Emoto, S.; Kishikawa, J.; Iida, Y.; Sunami, E.; et al. Temsirolimus and chloroquine cooperatively exhibit a potent antitumor effect against colorectal cancer cells. J. Cancer Res. Clin. Oncol. 2014, 140, 769–781. [Google Scholar] [CrossRef]
- Nozawa, H.; Watanabe, T.; Nagawa, H. Phosphorylation of ribosomal p70 S6 kinase and rapamycin sensitivity in human colorectal cancer. Cancer Lett. 2007, 251, 105–113. [Google Scholar] [CrossRef]
- Faes, S.; Demartines, N.; Dormond, O. Resistance to mTORC1 Inhibitors in Cancer Therapy: From Kinase Mutations to Intratumoral Heterogeneity of Kinase Activity. Oxid. Med. Cell Longev. 2017, 2017, 1726078. [Google Scholar] [CrossRef]
- Maira, S.M.; Stauffer, F.; Brueggen, J.; Furet, P.; Schnell, C.; Fritsch, C.; Brachmann, S.; Chene, P.; De Pover, A.; Schoemaker, K.; et al. Identification and characterization of NVP-BEZ235, a new orally available dual phosphatidylinositol 3-kinase/mammalian target of rapamycin inhibitor with potent in vivo antitumor activity. Mol. Cancer Ther. 2008, 7, 1851–1863. [Google Scholar] [CrossRef]
- Hua, H.; Kong, Q.; Zhang, H.; Wang, J.; Luo, T.; Jiang, Y. Targeting mTOR for cancer therapy. J. Hematol. Oncol. 2019, 12, 71. [Google Scholar] [CrossRef]
- Bahrami, A.; Khazaei, M.; Hasanzadeh, M.; ShahidSales, S.; Joudi Mashhad, M.; Farazestanian, M.; Sadeghnia, H.R.; Rezayi, M.; Maftouh, M.; Hassanian, S.M.; et al. Therapeutic Potential of Targeting PI3K/AKT Pathway in Treatment of Colorectal Cancer: Rational and Progress. J. Cell. Biochem. 2018, 119, 2460–2469. [Google Scholar] [CrossRef]
- Roper, J.; Richardson, M.P.; Wang, W.V.; Richard, L.G.; Chen, W.; Coffee, E.M.; Sinnamon, M.J.; Lee, L.; Chen, P.C.; Bronson, R.T.; et al. The dual PI3K/mTOR inhibitor NVP-BEZ235 induces tumor regression in a genetically engineered mouse model of PIK3CA wild-type colorectal cancer. PLoS ONE 2011, 6, e25132. [Google Scholar] [CrossRef]
- Foley, T.M.; Payne, S.N.; Pasch, C.A.; Yueh, A.E.; Van De Hey, D.R.; Korkos, D.P.; Clipson, L.; Maher, M.E.; Matkowskyj, K.A.; Newton, M.A.; et al. Dual PI3K/mTOR Inhibition in Colorectal Cancers with APC and PIK3CA Mutations. Mol. Cancer Res. 2017, 15, 317–327. [Google Scholar] [CrossRef]
- Bordeleau, M.E.; Robert, F.; Gerard, B.; Lindqvist, L.; Chen, S.M.; Wendel, H.G.; Brem, B.; Greger, H.; Lowe, S.W.; Porco, J.A., Jr.; et al. Therapeutic suppression of translation initiation modulates chemosensitivity in a mouse lymphoma model. J. Clin. Investig. 2008, 118, 2651–2660. [Google Scholar] [CrossRef]
- Chen, W.L.; Pan, L.; Kinghorn, A.D.; Swanson, S.M.; Burdette, J.E. Silvestrol induces early autophagy and apoptosis in human melanoma cells. BMC Cancer 2016, 16, 17. [Google Scholar] [CrossRef]
- Peters, T.L.; Tillotson, J.; Yeomans, A.M.; Wilmore, S.; Lemm, E.; Jimenez-Romero, C.; Amador, L.A.; Li, L.; Amin, A.D.; Pongtornpipat, P.; et al. Target-Based Screening against eIF4A1 Reveals the Marine Natural Product Elatol as a Novel Inhibitor of Translation Initiation with In Vivo Antitumor Activity. Clin. Cancer Res. 2018, 24, 4256–4270. [Google Scholar] [CrossRef]
- Thuaud, F.; Bernard, Y.; Turkeri, G.; Dirr, R.; Aubert, G.; Cresteil, T.; Baguet, A.; Tomasetto, C.; Svitkin, Y.; Sonenberg, N.; et al. Synthetic analogue of rocaglaol displays a potent and selective cytotoxicity in cancer cells: Involvement of apoptosis inducing factor and caspase-12. J. Med. Chem. 2009, 52, 5176–5187. [Google Scholar] [CrossRef] [PubMed]
- Wang, S.; Darini, C.; Desaubry, L.; Koromilas, A.E. STAT1 Promotes KRAS Colon Tumor Growth and Susceptibility to Pharmacological Inhibition of Translation Initiation Factor eIF4A. Mol. Cancer Ther. 2016, 15, 3055–3063. [Google Scholar] [CrossRef] [PubMed]
- Moerke, N.J.; Aktas, H.; Chen, H.; Cantel, S.; Reibarkh, M.Y.; Fahmy, A.; Gross, J.D.; Degterev, A.; Yuan, J.; Chorev, M.; et al. Small-molecule inhibition of the interaction between the translation initiation factors eIF4E and eIF4G. Cell 2007, 128, 257–267. [Google Scholar] [CrossRef] [PubMed]
- Sekiyama, N.; Arthanari, H.; Papadopoulos, E.; Rodriguez-Mias, R.A.; Wagner, G.; Leger-Abraham, M. Molecular mechanism of the dual activity of 4EGI-1: Dissociating eIF4G from eIF4E but stabilizing the binding of unphosphorylated 4E-BP1. Proc. Natl. Acad. Sci. USA 2015, 112, E4036–E4045. [Google Scholar] [CrossRef] [PubMed]
- Liu, Y.; Deguchi, Y.; Tian, R.; Wei, D.; Wu, L.; Chen, W.; Xu, W.; Xu, M.; Liu, F.; Gao, S.; et al. Pleiotropic Effects of PPARD Accelerate Colorectal Tumorigenesis, Progression, and Invasion. Cancer Res. 2019, 79, 954–969. [Google Scholar] [CrossRef]
- Matassa, D.S.; Amoroso, M.R.; Agliarulo, I.; Maddalena, F.; Sisinni, L.; Paladino, S.; Romano, S.; Romano, M.F.; Sagar, V.; Loreni, F.; et al. Translational control in the stress adaptive response of cancer cells: A novel role for the heat shock protein TRAP1. Cell Death Dis. 2013, 4, e851. [Google Scholar] [CrossRef]
- Ueda, T.; Watanabe-Fukunaga, R.; Fukuyama, H.; Nagata, S.; Fukunaga, R. Mnk2 and Mnk1 are essential for constitutive and inducible phosphorylation of eukaryotic initiation factor 4E but not for cell growth or development. Mol. Cell. Biol. 2004, 24, 6539–6549. [Google Scholar] [CrossRef]
- Konicek, B.W.; Stephens, J.R.; McNulty, A.M.; Robichaud, N.; Peery, R.B.; Dumstorf, C.A.; Dowless, M.S.; Iversen, P.W.; Parsons, S.; Ellis, K.E.; et al. Therapeutic inhibition of MAP kinase interacting kinase blocks eukaryotic initiation factor 4E phosphorylation and suppresses outgrowth of experimental lung metastases. Cancer Res. 2011, 71, 1849–1857. [Google Scholar] [CrossRef]
- Reich, S.H.; Sprengeler, P.A.; Chiang, G.G.; Appleman, J.R.; Chen, J.; Clarine, J.; Eam, B.; Ernst, J.T.; Han, Q.; Goel, V.K.; et al. Structure-based Design of Pyridone-Aminal eFT508 Targeting Dysregulated Translation by Selective Mitogen-activated Protein Kinase Interacting Kinases 1 and 2 (MNK1/2) Inhibition. J. Med. Chem. 2018, 61, 3516–3540. [Google Scholar] [CrossRef]
- Yuan, X.; Wu, H.; Bu, H.; Zheng, P.; Zhou, J.; Zhang, H. Design, synthesis and biological evaluation of pyridone-aminal derivatives as MNK1/2 inhibitors. Bioorg. Med. Chem. 2019, 27, 1211–1225. [Google Scholar] [CrossRef]
- Xu, Y.; Poggio, M.; Jin, H.Y.; Shi, Z.; Forester, C.M.; Wang, Y.; Stumpf, C.R.; Xue, L.; Devericks, E.; So, L.; et al. Translation control of the immune checkpoint in cancer and its therapeutic targeting. Nat. Med. 2019, 25, 301–311. [Google Scholar] [CrossRef]
- Kenner, L.R.; Anand, A.A.; Nguyen, H.C.; Myasnikov, A.G.; Klose, C.J.; McGeever, L.A.; Tsai, J.C.; Miller-Vedam, L.E.; Walter, P.; Frost, A. eIF2B-catalyzed nucleotide exchange and phosphoregulation by the integrated stress response. Science 2019, 364, 491–495. [Google Scholar] [CrossRef]
- Kashiwagi, K.; Yokoyama, T.; Nishimoto, M.; Takahashi, M.; Sakamoto, A.; Yonemochi, M.; Shirouzu, M.; Ito, T. Structural basis for eIF2B inhibition in integrated stress response. Science 2019, 364, 495–499. [Google Scholar] [CrossRef]
- Pakos-Zebrucka, K.; Koryga, I.; Mnich, K.; Ljujic, M.; Samali, A.; Gorman, A.M. The integrated stress response. EMBO Rep. 2016, 17, 1374–1395. [Google Scholar] [CrossRef]
- Tameire, F.; Verginadis, I.I.; Koumenis, C. Cell intrinsic and extrinsic activators of the unfolded protein response in cancer: Mechanisms and targets for therapy. Semin. Cancer Biol. 2015, 33, 3–15. [Google Scholar] [CrossRef]
- Cubillos-Ruiz, J.R.; Bettigole, S.E.; Glimcher, L.H. Tumorigenic and Immunosuppressive Effects of Endoplasmic Reticulum Stress in Cancer. Cell 2017, 168, 692–706. [Google Scholar] [CrossRef]
- Koromilas, A.E. Roles of the translation initiation factor eIF2alpha serine 51 phosphorylation in cancer formation and treatment. Biochim. Biophys. Acta 2015, 1849, 871–880. [Google Scholar] [CrossRef]
- McQuiston, A.; Diehl, J.A. Recent insights into PERK-dependent signaling from the stressed endoplasmic reticulum. F1000Research 2017, 6, 1897. [Google Scholar] [CrossRef]
- Feng, Y.X.; Sokol, E.S.; Del Vecchio, C.A.; Sanduja, S.; Claessen, J.H.; Proia, T.A.; Jin, D.X.; Reinhardt, F.; Ploegh, H.L.; Wang, Q.; et al. Epithelial-to-mesenchymal transition activates PERK-eIF2alpha and sensitizes cells to endoplasmic reticulum stress. Cancer Discov. 2014, 4, 702–715. [Google Scholar] [CrossRef]
- Bi, M.; Naczki, C.; Koritzinsky, M.; Fels, D.; Blais, J.; Hu, N.; Harding, H.; Novoa, I.; Varia, M.; Raleigh, J.; et al. ER stress-regulated translation increases tolerance to extreme hypoxia and promotes tumor growth. EMBO J. 2005, 24, 3470–3481. [Google Scholar] [CrossRef]
- Tian, M.; Wan, Y.; Tang, J.; Li, H.; Yu, G.; Zhu, J.; Ji, S.; Guo, H.; Zhang, N.; Li, W.; et al. Depletion of tissue factor suppresses hepatic metastasis and tumor growth in colorectal cancer via the downregulation of MMPs and the induction of autophagy and apoptosis. Cancer Biol. Ther. 2011, 12, 896–907. [Google Scholar] [CrossRef] [PubMed]
- Chern, Y.J.; Wong, J.C.T.; Cheng, G.S.W.; Yu, A.; Yin, Y.; Schaeffer, D.F.; Kennecke, H.F.; Morin, G.; Tai, I.T. The interaction between SPARC and GRP78 interferes with ER stress signaling and potentiates apoptosis via PERK/eIF2alpha and IRE1alpha/XBP-1 in colorectal cancer. Cell Death Dis. 2019, 10, 504. [Google Scholar] [CrossRef] [PubMed]
- Ranganathan, A.C.; Ojha, S.; Kourtidis, A.; Conklin, D.S.; Aguirre-Ghiso, J.A. Dual function of pancreatic endoplasmic reticulum kinase in tumor cell growth arrest and survival. Cancer Res. 2008, 68, 3260–3268. [Google Scholar] [CrossRef]
- Heijmans, J.; van Lidth de Jeude, J.F.; Koo, B.K.; Rosekrans, S.L.; Wielenga, M.C.; van de Wetering, M.; Ferrante, M.; Lee, A.S.; Onderwater, J.J.; Paton, J.C.; et al. ER stress causes rapid loss of intestinal epithelial stemness through activation of the unfolded protein response. Cell Rep. 2013, 3, 1128–1139. [Google Scholar] [CrossRef]
- Spaan, C.N.; Smit, W.L.; van Lidth de Jeude, J.F.; Meijer, B.J.; Muncan, V.; van den Brink, G.R.; Heijmans, J. Expression of UPR effector proteins ATF6 and XBP1 reduce colorectal cancer cell proliferation and stemness by activating PERK signaling. Cell Death Dis. 2019, 10, 490. [Google Scholar] [CrossRef]
- Dong, J.; Qiu, H.; Garcia-Barrio, M.; Anderson, J.; Hinnebusch, A.G. Uncharged tRNA activates GCN2 by displacing the protein kinase moiety from a bipartite tRNA-binding domain. Mol. Cell 2000, 6, 269–279. [Google Scholar] [CrossRef]
- Ye, J.; Kumanova, M.; Hart, L.S.; Sloane, K.; Zhang, H.; De Panis, D.N.; Bobrovnikova-Marjon, E.; Diehl, J.A.; Ron, D.; Koumenis, C. The GCN2-ATF4 pathway is critical for tumour cell survival and proliferation in response to nutrient deprivation. EMBO J. 2010, 29, 2082–2096. [Google Scholar] [CrossRef]
- Garcia, M.A.; Gil, J.; Ventoso, I.; Guerra, S.; Domingo, E.; Rivas, C.; Esteban, M. Impact of protein kinase PKR in cell biology: From antiviral to antiproliferative action. Microbiol. Mol. Biol. Rev. 2006, 70, 1032–1060. [Google Scholar] [CrossRef]
- Dabo, S.; Meurs, E.F. dsRNA-dependent protein kinase PKR and its role in stress, signaling and HCV infection. Viruses 2012, 4, 2598–2635. [Google Scholar] [CrossRef]
- Brunelli, C.; Amici, C.; Angelini, M.; Fracassi, C.; Belardo, G.; Santoro, M.G. The non-steroidal anti-inflammatory drug indomethacin activates the eIF2alpha kinase PKR, causing a translational block in human colorectal cancer cells. Biochem. J. 2012, 443, 379–386. [Google Scholar] [CrossRef]
- Kim, S.H.; Gunnery, S.; Choe, J.K.; Mathews, M.B. Neoplastic progression in melanoma and colon cancer is associated with increased expression and activity of the interferon-inducible protein kinase, PKR. Oncogene 2002, 21, 8741–8748. [Google Scholar] [CrossRef] [PubMed]
- De Simone, V.; Bevivino, G.; Sedda, S.; Izzo, R.; Laudisi, F.; Dinallo, V.; Franze, E.; Colantoni, A.; Ortenzi, A.; Salvatori, S.; et al. Smad7 knockdown activates protein kinase RNA-associated eIF2alpha pathway leading to colon cancer cell death. Cell Death Dis. 2017, 8, e2681. [Google Scholar] [CrossRef] [PubMed]
- Yoon, C.H.; Lee, E.S.; Lim, D.S.; Bae, Y.S. PKR, a p53 target gene, plays a crucial role in the tumor-suppressor function of p53. Proc. Natl. Acad. Sci. USA 2009, 106, 7852–7857. [Google Scholar] [CrossRef] [PubMed]
- Singh, C.; Haines, G.K.; Talamonti, M.S.; Radosevich, J.A. Expression of p68 in human colon cancer. Tumour Biol. 1995, 16, 281–289. [Google Scholar] [CrossRef]
- Garcia, M.A.; Carrasco, E.; Aguilera, M.; Alvarez, P.; Rivas, C.; Campos, J.M.; Prados, J.C.; Calleja, M.A.; Esteban, M.; Marchal, J.A.; et al. The chemotherapeutic drug 5-fluorouracil promotes PKR-mediated apoptosis in a p53-independent manner in colon and breast cancer cells. PLoS ONE 2011, 6, e23887. [Google Scholar] [CrossRef]
- Treton, X.; Pedruzzi, E.; Cazals-Hatem, D.; Grodet, A.; Panis, Y.; Groyer, A.; Moreau, R.; Bouhnik, Y.; Daniel, F.; Ogier-Denis, E. Altered endoplasmic reticulum stress affects translation in inactive colon tissue from patients with ulcerative colitis. Gastroenterology 2011, 141, 1024–1035. [Google Scholar] [CrossRef]
- Stidham, R.W.; Higgins, P.D.R. Colorectal Cancer in Inflammatory Bowel Disease. Clin. Colon Rectal Surg. 2018, 31, 168–178. [Google Scholar] [CrossRef]
- Lobo, M.V.; Martin, M.E.; Perez, M.I.; Alonso, F.J.; Redondo, C.; Alvarez, M.I.; Salinas, M. Levels, phosphorylation status and cellular localization of translational factor eIF2 in gastrointestinal carcinomas. Histochem. J. 2000, 32, 139–150. [Google Scholar] [CrossRef]
- Rosenwald, I.B.; Wang, S.; Savas, L.; Woda, B.; Pullman, J. Expression of translation initiation factor eIF-2alpha is increased in benign and malignant melanocytic and colonic epithelial neoplasms. Cancer 2003, 98, 1080–1088. [Google Scholar] [CrossRef]
- Gallagher, J.W.; Kubica, N.; Kimball, S.R.; Jefferson, L.S. Reduced eukaryotic initiation factor 2Bepsilon-subunit expression suppresses the transformed phenotype of cells overexpressing the protein. Cancer Res. 2008, 68, 8752–8760. [Google Scholar] [CrossRef]
- Balachandran, S.; Barber, G.N. Defective translational control facilitates vesicular stomatitis virus oncolysis. Cancer Cell 2004, 5, 51–65. [Google Scholar] [CrossRef]
- Sidrauski, C.; Acosta-Alvear, D.; Khoutorsky, A.; Vedantham, P.; Hearn, B.R.; Li, H.; Gamache, K.; Gallagher, C.M.; Ang, K.K.; Wilson, C.; et al. Pharmacological brake-release of mRNA translation enhances cognitive memory. Elife 2013, 2, e00498. [Google Scholar] [CrossRef] [PubMed]
- Sidrauski, C.; McGeachy, A.M.; Ingolia, N.T.; Walter, P. The small molecule ISRIB reverses the effects of eIF2alpha phosphorylation on translation and stress granule assembly. Elife 2015, 4, e05033. [Google Scholar] [CrossRef]
- Sidrauski, C.; Tsai, J.C.; Kampmann, M.; Hearn, B.R.; Vedantham, P.; Jaishankar, P.; Sokabe, M.; Mendez, A.S.; Newton, B.W.; Tang, E.L.; et al. Pharmacological dimerization and activation of the exchange factor eIF2B antagonizes the integrated stress response. Elife 2015, 4, e07314. [Google Scholar] [CrossRef]
- Tsai, J.C.; Miller-Vedam, L.E.; Anand, A.A.; Jaishankar, P.; Nguyen, H.C.; Renslo, A.R.; Frost, A.; Walter, P. Structure of the nucleotide exchange factor eIF2B reveals mechanism of memory-enhancing molecule. Science 2018, 359. [Google Scholar] [CrossRef] [PubMed]
- Rabouw, H.H.; Langereis, M.A.; Anand, A.A.; Visser, L.J.; de Groot, R.J.; Walter, P.; van Kuppeveld, F.J.M. Small molecule ISRIB suppresses the integrated stress response within a defined window of activation. Proc. Natl. Acad. Sci. USA 2019, 116, 2097–2102. [Google Scholar] [CrossRef] [PubMed]
- Anand, A.A.; Walter, P. Structural insights into ISRIB, a memory-enhancing inhibitor of the integrated stress response. FEBS J. 2020, 287, 239–245. [Google Scholar] [CrossRef]
- Wong, Y.L.; LeBon, L.; Edalji, R.; Lim, H.B.; Sun, C.; Sidrauski, C. The small molecule ISRIB rescues the stability and activity of Vanishing White Matter Disease eIF2B mutant complexes. Elife 2018, 7, e32733. [Google Scholar] [CrossRef]
- Nguyen, H.G.; Conn, C.S.; Kye, Y.; Xue, L.; Forester, C.M.; Cowan, J.E.; Hsieh, A.C.; Cunningham, J.T.; Truillet, C.; Tameire, F.; et al. Development of a stress response therapy targeting aggressive prostate cancer. Sci. Transl. Med. 2018, 10. [Google Scholar] [CrossRef]
- Chou, A.; Krukowski, K.; Jopson, T.; Zhu, P.J.; Costa-Mattioli, M.; Walter, P.; Rosi, S. Inhibition of the integrated stress response reverses cognitive deficits after traumatic brain injury. Proc. Natl. Acad. Sci. USA 2017, 114, E6420–E6426. [Google Scholar] [CrossRef]
- Halliday, M.; Radford, H.; Sekine, Y.; Moreno, J.; Verity, N.; le Quesne, J.; Ortori, C.A.; Barrett, D.A.; Fromont, C.; Fischer, P.M.; et al. Partial restoration of protein synthesis rates by the small molecule ISRIB prevents neurodegeneration without pancreatic toxicity. Cell Death Dis. 2015, 6, e1672. [Google Scholar] [CrossRef] [PubMed]
- Moreno, J.A.; Halliday, M.; Molloy, C.; Radford, H.; Verity, N.; Axten, J.M.; Ortori, C.A.; Willis, A.E.; Fischer, P.M.; Barrett, D.A.; et al. Oral treatment targeting the unfolded protein response prevents neurodegeneration and clinical disease in prion-infected mice. Sci. Transl. Med. 2013, 5, 206ra138. [Google Scholar] [CrossRef] [PubMed]
- Novoa, I.; Zeng, H.; Harding, H.P.; Ron, D. Feedback inhibition of the unfolded protein response by GADD34-mediated dephosphorylation of eIF2alpha. J. Cell Biol. 2001, 153, 1011–1022. [Google Scholar] [CrossRef] [PubMed]
- Connor, J.H.; Weiser, D.C.; Li, S.; Hallenbeck, J.M.; Shenolikar, S. Growth arrest and DNA damage-inducible protein GADD34 assembles a novel signaling complex containing protein phosphatase 1 and inhibitor 1. Mol. Cell. Biol. 2001, 21, 6841–6850. [Google Scholar] [CrossRef]
- Jousse, C.; Oyadomari, S.; Novoa, I.; Lu, P.; Zhang, Y.; Harding, H.P.; Ron, D. Inhibition of a constitutive translation initiation factor 2alpha phosphatase, CReP, promotes survival of stressed cells. J. Cell Biol. 2003, 163, 767–775. [Google Scholar] [CrossRef]
- Reid, D.W.; Tay, A.S.; Sundaram, J.R.; Lee, I.C.; Chen, Q.; George, S.E.; Nicchitta, C.V.; Shenolikar, S. Complementary Roles of GADD34- and CReP-Containing Eukaryotic Initiation Factor 2alpha Phosphatases during the Unfolded Protein Response. Mol. Cell. Biol. 2016, 36, 1868–1880. [Google Scholar] [CrossRef]
- Boyce, M.; Bryant, K.F.; Jousse, C.; Long, K.; Harding, H.P.; Scheuner, D.; Kaufman, R.J.; Ma, D.; Coen, D.M.; Ron, D.; et al. A selective inhibitor of eIF2alpha dephosphorylation protects cells from ER stress. Science 2005, 307, 935–939. [Google Scholar] [CrossRef]
- Wielenga, M.C.B.; Colak, S.; Heijmans, J.; van Lidth de Jeude, J.F.; Rodermond, H.M.; Paton, J.C.; Paton, A.W.; Vermeulen, L.; Medema, J.P.; van den Brink, G.R. ER-Stress-Induced Differentiation Sensitizes Colon Cancer Stem Cells to Chemotherapy. Cell Rep. 2015, 13, 489–494. [Google Scholar] [CrossRef]
- Atkins, C.; Liu, Q.; Minthorn, E.; Zhang, S.Y.; Figueroa, D.J.; Moss, K.; Stanley, T.B.; Sanders, B.; Goetz, A.; Gaul, N.; et al. Characterization of a novel PERK kinase inhibitor with antitumor and antiangiogenic activity. Cancer Res. 2013, 73, 1993–2002. [Google Scholar] [CrossRef]
- Axten, J.M.; Romeril, S.P.; Shu, A.; Ralph, J.; Medina, J.R.; Feng, Y.; Li, W.H.; Grant, S.W.; Heerding, D.A.; Minthorn, E.; et al. Discovery of GSK2656157: An Optimized PERK Inhibitor Selected for Preclinical Development. ACS Med. Chem. Lett. 2013, 4, 964–968. [Google Scholar] [CrossRef]
- Brazeau, J.F.; Rosse, G. Triazolo [4,5-d] pyrimidine Derivatives as Inhibitors of GCN2. ACS Med. Chem. Lett. 2014, 5, 282–283. [Google Scholar] [CrossRef] [PubMed]
- Fujimoto, J.; Kurasawa, O.; Takagi, T.; Liu, X.; Banno, H.; Kojima, T.; Asano, Y.; Nakamura, A.; Nambu, T.; Hata, A.; et al. Identification of Novel, Potent, and Orally Available GCN2 Inhibitors with Type I Half Binding Mode. ACS Med. Chem. Lett. 2019, 10, 1498–1503. [Google Scholar] [CrossRef] [PubMed]
- Nakamura, A.; Nambu, T.; Ebara, S.; Hasegawa, Y.; Toyoshima, K.; Tsuchiya, Y.; Tomita, D.; Fujimoto, J.; Kurasawa, O.; Takahara, C.; et al. Inhibition of GCN2 sensitizes ASNS-low cancer cells to asparaginase by disrupting the amino acid response. Proc. Natl. Acad. Sci. USA 2018, 115, E7776–E7785. [Google Scholar] [CrossRef] [PubMed]
- Lough, L.; Sherman, D.; Beccera-Flores, M.; Lavinda, O.; Ni, E.; Wang, H.; Tibes, R.; Cardozo, T. Triazolo [4,5-d] pyrimidines as Validated General Control Nonderepressible 2 (GCN2) Protein Kinase Inhibitors Reduce Growth of Leukemia Cells. Comput. Struct. Biotechnol. J. 2018, 16, 350–360. [Google Scholar] [CrossRef]
- Jammi, N.V.; Whitby, L.R.; Beal, P.A. Small molecule inhibitors of the RNA-dependent protein kinase. Biochem. Biophys. Res. Commun. 2003, 308, 50–57. [Google Scholar] [CrossRef]
- Chen, H.M.; Wang, L.; D’Mello, S.R. A chemical compound commonly used to inhibit PKR, {8-(imidazol-4-ylmethylene)-6H-azolidino [5,4-g] benzothiazol-7-one}, protects neurons by inhibiting cyclin-dependent kinase. Eur. J. Neurosci. 2008, 28, 2003–2016. [Google Scholar] [CrossRef]
- Ali, U.; Naveed, M.; Ullah, A.; Ali, K.; Shah, S.A.; Fahad, S.; Mumtaz, A.S. L-asparaginase as a critical component to combat Acute Lymphoblastic Leukaemia (ALL): A novel approach to target ALL. Eur. J. Pharmacol. 2016, 771, 199–210. [Google Scholar] [CrossRef]
- Egler, R.A.; Ahuja, S.P.; Matloub, Y. L-asparaginase in the treatment of patients with acute lymphoblastic leukemia. J. Pharmacol. Pharmacother. 2016, 7, 62–71. [Google Scholar] [CrossRef]
- Tameire, F.; Verginadis, I.I.; Leli, N.M.; Polte, C.; Conn, C.S.; Ojha, R.; Salinas, C.S.; Chinga, F.; Monroy, A.M.; Fu, W.; et al. ATF4 couples MYC-dependent translational activity to bioenergetic demands during tumour progression. Nat. Cell Biol. 2019, 21, 889–899. [Google Scholar] [CrossRef]
- Tameire, F.; Verginadis, I.I.; Leli, N.M.; Polte, C.; Conn, C.S.; Ojha, R.; Salinas, C.S.; Chinga, F.; Monroy, A.M.; Fu, W.; et al. Author Correction: ATF4 couples MYC-dependent translational activity to bioenergetic demands during tumour progression. Nat. Cell Biol. 2019, 21, 1052. [Google Scholar] [CrossRef]
- Lehman, S.L.; Ryeom, S.; Koumenis, C. Signaling through alternative Integrated Stress Response pathways compensates for GCN2 loss in a mouse model of soft tissue sarcoma. Sci. Rep. 2015, 5, 11781. [Google Scholar] [CrossRef] [PubMed]
- Hamanaka, R.B.; Bennett, B.S.; Cullinan, S.B.; Diehl, J.A. PERK and GCN2 contribute to eIF2alpha phosphorylation and cell cycle arrest after activation of the unfolded protein response pathway. Mol. Biol. Cell 2005, 16, 5493–5501. [Google Scholar] [CrossRef] [PubMed]
- Liu, Y.; Laszlo, C.; Liu, Y.; Liu, W.; Chen, X.; Evans, S.C.; Wu, S. Regulation of G(1) arrest and apoptosis in hypoxia by PERK and GCN2-mediated eIF2alpha phosphorylation. Neoplasia 2010, 12, 61–68. [Google Scholar] [CrossRef] [PubMed]
- Roobol, A.; Roobol, J.; Bastide, A.; Knight, J.R.; Willis, A.E.; Smales, C.M. p58IPK is an inhibitor of the eIF2alpha kinase GCN2 and its localization and expression underpin protein synthesis and ER processing capacity. Biochem. J. 2015, 465, 213–225. [Google Scholar] [CrossRef] [PubMed]
- Alqurashi, N.; Hashimi, S.M.; Alowaidi, F.; Ivanovski, S.; Wei, M.Q. Dual mTOR/PI3K inhibitor NVPBEZ235 arrests colorectal cancer cell growth and displays differential inhibition of 4EBP1. Oncol. Rep. 2018, 40, 1083–1092. [Google Scholar] [CrossRef] [PubMed]
- Ng, K.; Tabernero, J.; Hwang, J.; Bajetta, E.; Sharma, S.; Del Prete, S.A.; Arrowsmith, E.R.; Ryan, D.P.; Sedova, M.; Jin, J.; et al. Phase II study of everolimus in patients with metastatic colorectal adenocarcinoma previously treated with bevacizumab-, fluoropyrimidine-, oxaliplatin-, and irinotecan-based regimens. Clin. Cancer Res. 2013, 19, 3987–3995. [Google Scholar] [CrossRef]
- Altomare, I.; Hurwitz, H. Everolimus in colorectal cancer. Expert Opin. Pharmacother. 2013, 14, 505–513. [Google Scholar] [CrossRef]
- Spindler, K.L.; Sorensen, M.M.; Pallisgaard, N.; Andersen, R.F.; Havelund, B.M.; Ploen, J.; Lassen, U.; Jakobsen, A.K. Phase II trial of temsirolimus alone and in combination with irinotecan for KRAS mutant metastatic colorectal cancer: Outcome and results of KRAS mutational analysis in plasma. Acta Oncol. 2013, 52, 963–970. [Google Scholar] [CrossRef]
- Buijsen, J.; van den Bogaard, J.; Jutten, B.; Belgers, E.; Sosef, M.; Leijtens, J.W.; Beets, G.L.; Jansen, R.L.; Riedl, R.G.; Clarijs, R.; et al. A phase I-II study on the combination of rapamycin and short course radiotherapy in rectal cancer. Radiother. Oncol. 2015, 116, 214–220. [Google Scholar] [CrossRef]
- Maira, S.M.; Pecchi, S.; Huang, A.; Burger, M.; Knapp, M.; Sterker, D.; Schnell, C.; Guthy, D.; Nagel, T.; Wiesmann, M.; et al. Identification and characterization of NVP-BKM120, an orally available pan-class I PI3-kinase inhibitor. Mol. Cancer Ther. 2012, 11, 317–328. [Google Scholar] [CrossRef]
- Piha-Paul, S.A.; Knost, J.A.; Nikolinakos, P.; Schwartzberg, L.S.; Yardley, D.A.; Slosberg, E.D.; Kang, B.; White, T.; Lebedinsky, C. Colorectal cancer (CRC) cohort analysis of patients treated with buparlisib in a distinctive tissue-agnostic trial model for molecularly preselected tumors. J. Clin. Oncol. 2015, 33. [Google Scholar] [CrossRef]
- Baranda, J.C.; Reed, G.; Williamson, S.K.; Perez, R.P.; Stoltz, M.L.; Mackay, C.; Madan, R.; Pessetto, Z.Y.; Godwin, A.K. Irinotecan (Iri) and buparlisib (B) in previously treated patients (pts) with metastatic colorectal cancer (mCRC). J. Clin. Oncol. 2015, 33. [Google Scholar] [CrossRef]
- Yang, X.D.; Jia, X.C.; Corvalan, J.R.; Wang, P.; Davis, C.G. Development of ABX-EGF, a fully human anti-EGF receptor monoclonal antibody, for cancer therapy. Crit. Rev. Oncol. Hematol. 2001, 38, 17–23. [Google Scholar] [CrossRef]
- Fritsch, C.; Huang, A.; Chatenay-Rivauday, C.; Schnell, C.; Reddy, A.; Liu, M.; Kauffmann, A.; Guthy, D.; Erdmann, D.; De Pover, A.; et al. Characterization of the novel and specific PI3Kalpha inhibitor NVP-BYL719 and development of the patient stratification strategy for clinical trials. Mol. Cancer Ther. 2014, 13, 1117–1129. [Google Scholar] [CrossRef] [PubMed]
- Andre, F.; Ciruelos, E.; Rubovszky, G.; Campone, M.; Loibl, S.; Rugo, H.S.; Iwata, H.; Conte, P.; Mayer, I.A.; Kaufman, B.; et al. Alpelisib for PIK3CA-Mutated, Hormone Receptor-Positive Advanced Breast Cancer. New Engl. J. Med. 2019, 380, 1929–1940. [Google Scholar] [CrossRef] [PubMed]
- Juric, D.; Rodon, J.; Tabernero, J.; Janku, F.; Burris, H.A.; Schellens, J.H.; Middleton, M.R.; Berlin, J.; Schuler, M.; Gil-Martin, M.; et al. Phosphatidylinositol 3-Kinase -Selective Inhibition With Alpelisib (BYL719) in PIK3CA-Altered Solid Tumors: Results From the First-in-Human Study. J. Clin. Oncol. 2018, 36, 1291. [Google Scholar] [CrossRef]
- Van Geel, R.M.J.M.; Tabernero, J.; Elez, E.; Bendell, J.C.; Spreafico, A.; Schuler, M.; Yoshino, T.; Delord, J.P.; Yamada, Y.; Lolkema, M.P.; et al. A Phase Ib Dose-Escalation Study of Encorafenib and Cetuximab with or without Alpelisib in Metastatic BRAF-Mutant Colorectal Cancer. Cancer Discov. 2017, 7, 610–619. [Google Scholar] [CrossRef]
- Tabernero, J.; Van Geel, R.; Guren, T.K.; Yaeger, R.D.; Spreafico, A.; Faris, J.E.; Yoshino, T.; Yamada, Y.; Kim, T.W.; Bendell, J.C.; et al. Phase 2 results: Encorafenib (ENCO) and cetuximab (CETUX) with or without alpelisib (ALP) in patients with advanced BRAF-mutant colorectal cancer (BRAFm CRC). J. Clin. Oncol. 2016, 34. [Google Scholar] [CrossRef]
- Dolly, S.O.; Wagner, A.J.; Bendell, J.C.; Kindler, H.L.; Krug, L.M.; Seiwert, T.Y.; Zauderer, M.G.; Lolkema, M.P.; Apt, D.; Yeh, R.F.; et al. Phase I Study of Apitolisib (GDC-0980), Dual Phosphatidylinositol-3-Kinase and Mammalian Target of Rapamycin Kinase Inhibitor, in Patients with Advanced Solid Tumors. Clin. Cancer Res. 2016, 22, 2874–2884. [Google Scholar] [CrossRef]
- Roseri, L.; Goldman, J.; Hubbard, J.M.; Roos, M.; Capdevila, J.; Maynes, J.; Lin, W.; O’Keeffe, B.; Lackner, M.; Spoerke, J.; et al. Phase Ib study of oral dual-PI3K/mTOR inhibitor GDC-0980 in combination with capecitabine and mFOLFOX6+bevacizumab in patients with advanced solid tumors and colorectal cancer. Eur. J. Cancer 2014, 50, 122–123. [Google Scholar] [CrossRef]
- Hubbard, J.M.; Patel, M.R.; Bekaii-Saab, T.S.; Falchook, G.S.; Freilich, B.L.; Dasari, A.; Knisely, B.T.; Anderson, M.; Chiang, G.G.; Webster, K.R.; et al. A phase II, open label, randomized, noncomparative study of eFT508 (tomivosertib) alone or in combination with avelumab in subjects with relapsed/refractory microsatellite stable colorectal cancer (MSS CRC). J. Clin. Oncol. 2019, 37. [Google Scholar] [CrossRef]
- Santag, S.; Siegel, F.; Wengner, A.M.; Lange, C.; Bomer, U.; Eis, K.; Puhler, F.; Lienau, P.; Bergemann, L.; Michels, M.; et al. BAY 1143269, a novel MNK1 inhibitor, targets oncogenic protein expression and shows potent anti-tumor activity. Cancer Lett. 2017, 390, 21–29. [Google Scholar] [CrossRef]
- Duffy, A.G.; Makarova-Rusher, O.V.; Ulahannan, S.V.; Rahma, O.E.; Fioravanti, S.; Walker, M.; Abdullah, S.; Raffeld, M.; Anderson, V.; Abi-Jaoudeh, N.; et al. Modulation of tumor eIF4E by antisense inhibition: A phase I/II translational clinical trial of ISIS 183750-an antisense oligonucleotide against eIF4E-in combination with irinotecan in solid tumors and irinotecan-refractory colorectal cancer. Int. J. Cancer 2016, 139, 1648–1657. [Google Scholar] [CrossRef] [PubMed]
- Van de Wetering, M.; Francies, H.E.; Francis, J.M.; Bounova, G.; Iorio, F.; Pronk, A.; van Houdt, W.; van Gorp, J.; Taylor-Weiner, A.; Kester, L.; et al. Prospective derivation of a living organoid biobank of colorectal cancer patients. Cell 2015, 161, 933–945. [Google Scholar] [CrossRef] [PubMed]
- Ooft, S.N.; Weeber, F.; Dijkstra, K.K.; McLean, C.M.; Kaing, S.; van Werkhoven, E.; Schipper, L.; Hoes, L.; Vis, D.J.; van de Haar, J.; et al. Patient-derived organoids can predict response to chemotherapy in metastatic colorectal cancer patients. Sci. Transl. Med. 2019, 11. [Google Scholar] [CrossRef] [PubMed]
- Fujii, M.; Shimokawa, M.; Date, S.; Takano, A.; Matano, M.; Nanki, K.; Ohta, Y.; Toshimitsu, K.; Nakazato, Y.; Kawasaki, K.; et al. A Colorectal Tumor Organoid Library Demonstrates Progressive Loss of Niche Factor Requirements during Tumorigenesis. Cell Stem Cell 2016, 18, 827–838. [Google Scholar] [CrossRef] [PubMed]
- Vlachogiannis, G.; Hedayat, S.; Vatsiou, A.; Jamin, Y.; Fernandez-Mateos, J.; Khan, K.; Lampis, A.; Eason, K.; Huntingford, I.; Burke, R.; et al. Patient-derived organoids model treatment response of metastatic gastrointestinal cancers. Science 2018, 359, 920–926. [Google Scholar] [CrossRef] [PubMed]
| Regulators of mRNA Translation | Deregulation in CRC | Impact on mRNA Translation | |
|---|---|---|---|
| Ribosomal Components | RPL15 | upregulation | enhanced ribosome biogenesis |
| RPL22 | mutation, downregulation | potentially deregulated translation of pro-apoptotic proteins and metastasis-related proteins | |
| RPS20 | mutation | defect in pre-ribosomal RNA maturation | |
| RPS24 | upregulation | enhanced ribosome biogenesis | |
| ribosomal RNAs | upregulation via MYC-mediated deregulation of RNA pol I and III activity | enhanced ribosome biogenesis | |
| Signaling Pathways and Associated Factors | RAS/MAPK signaling | mutation and hyperactivation | hyperactivation of mTORC1 and subsequent activation of p70-S6K1 and inhibition of 4E-BPs leading to enhanced translation initiation |
| PI3K/AKT signaling | mutation and hyperactivation, upregulation | hyperactivation of mTORC1 and subsequent activation of p70-S6K1 and inhibition of 4E-BPs leading to enhanced translation initiation | |
| PTEN | deletion | upregulation of PI3K/AKT signaling | |
| mTORC1 | mutation and hyperactivation, overexpression, increased phosphorylation of mTOR | activation of p70-S6K1 and inhibition of 4E-BPs leading to enhanced translation initiation | |
| 4E-BPs | increased phosphorylation | release of eIF4E and enhanced translation initiation | |
| PDCD4 | downregulation | enhanced eIF4A activity and translation initiation | |
| p70-S6K1 | increased phosphorylation | phosphorylation and inactivation of PDCD4 and eEF2K and enhanced translation initiation and elongation | |
| Translation Elongation Factors | eEF2K | downregulation | enhanced activity of eEF2 and translation elongation |
| eEF2 | upregulation | enhanced translation elongation | |
| Translation Initiation Factors | eIF4E | upregulation, increased phosphorylation at S209 | enhanced translation initiation |
| eIF4A1 | upregulation | enhanced translation initiation | |
| eIF2α | upregulation, increased phosphorylation at S51 | sequestration of eIF2B in an inactive complex, thereby limiting high translation rates | |
| eIF2B complex | upregulation | enhanced complex formation with p-eIF2α | |
| Stress-related Kinase | GCN2 | increased activity | increased phosphorylation of eIF2α |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Schmidt, S.; Denk, S.; Wiegering, A. Targeting Protein Synthesis in Colorectal Cancer. Cancers 2020, 12, 1298. https://doi.org/10.3390/cancers12051298
Schmidt S, Denk S, Wiegering A. Targeting Protein Synthesis in Colorectal Cancer. Cancers. 2020; 12(5):1298. https://doi.org/10.3390/cancers12051298
Chicago/Turabian StyleSchmidt, Stefanie, Sarah Denk, and Armin Wiegering. 2020. "Targeting Protein Synthesis in Colorectal Cancer" Cancers 12, no. 5: 1298. https://doi.org/10.3390/cancers12051298
APA StyleSchmidt, S., Denk, S., & Wiegering, A. (2020). Targeting Protein Synthesis in Colorectal Cancer. Cancers, 12(5), 1298. https://doi.org/10.3390/cancers12051298





