Oncobiosis and Microbial Metabolite Signaling in Pancreatic Adenocarcinoma
Abstract
1. Pancreatic Adenocarcinoma, an Unmet Medical Need
2. The Oncobiotic Transformation of the Microbiome
3. The Oncobiome in Pancreatic Adenocarcinoma
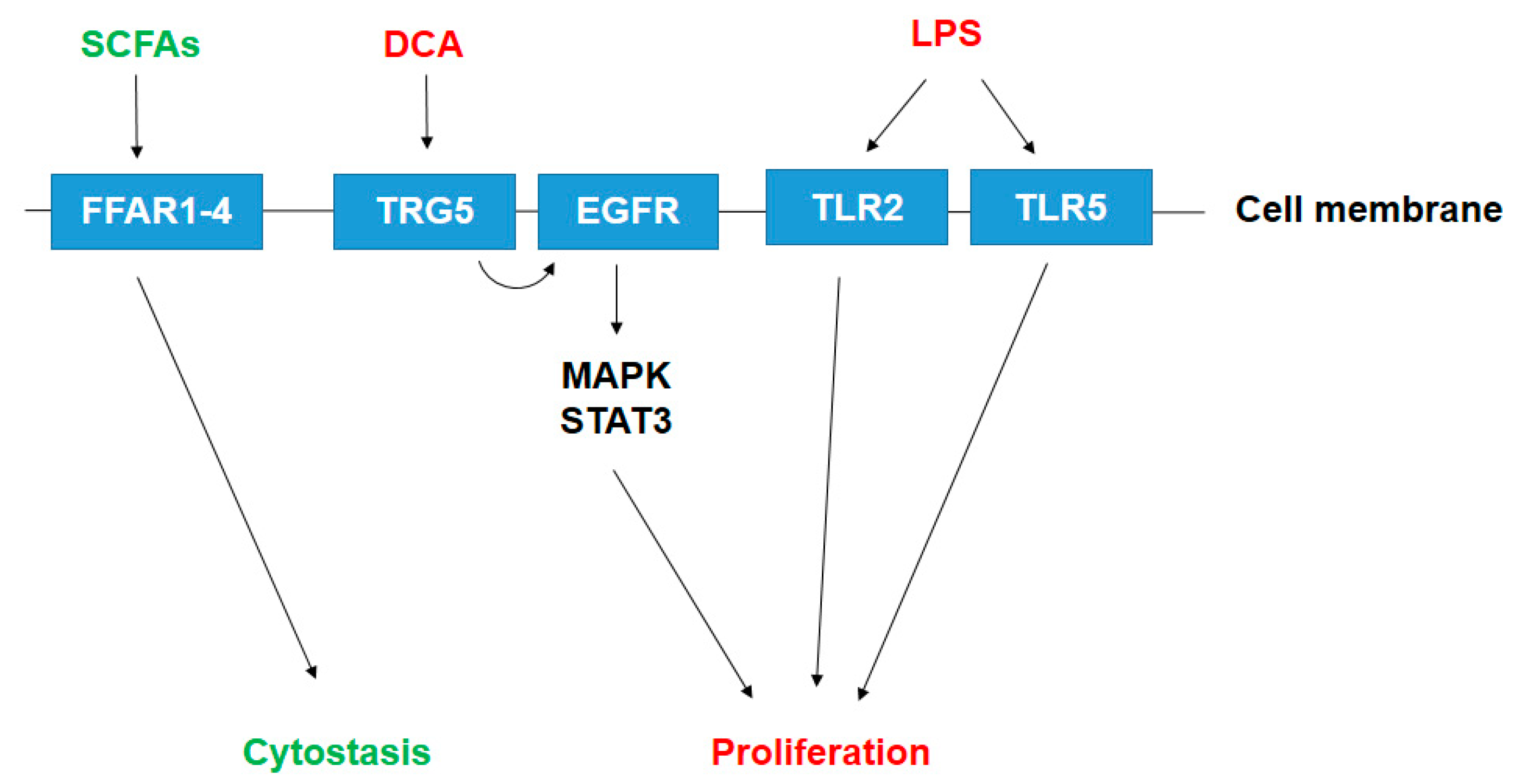
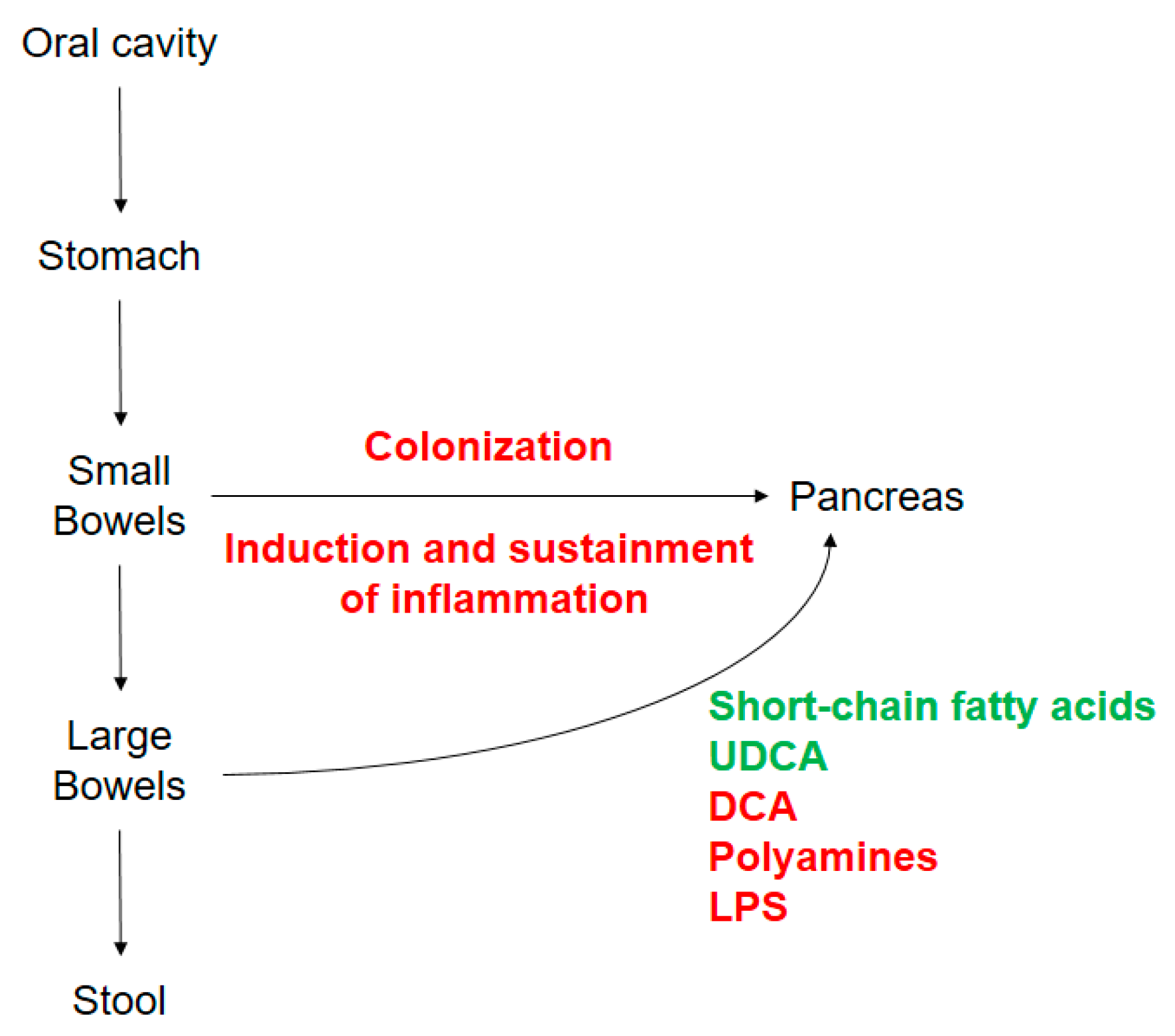
4. Bacterial Metabolites Playing Role in Pancreatic Adenocarcinoma
4.1. Short Chain Fatty Acids (SFCA)
4.2. Secondary Bile Acids
4.3. Polyamines
4.4. Bacterial Lipopolysaccharide (LPS)
4.5. Tryptophan Metabolites
- (1)
- The decarboxylation of tryptophan yields tryptamine. Clostridium sporogenes and Ruminococcus gnavus possess enzymes for tryptophan decarboxylation [133].
- (2)
- Tryptophanase deaminates tryptophan to indole pyruvic acid, which is then metabolized to indole. Indole can be further oxidized and the subsequent conjugation of sulphate yields indican. Tryptophanase, denoted as TnaA, can be found in the tryptophanase operon [211]. Tryptophanase expression is widespread among bacteria [212,213].
- (3)
- Indole pyruvic acid can be decarboxylated to indole acetaldehyde. Indole acetaldehyde can be converted to tryptophol or indole acetic acid. Indole acetic acid can be decarboxylated to yield skatole or conjugated with glutamine to yield indole acetic acid-glutamine. The main genera for this pathway are Lactobacillus, Clostridium, and Bacteroides [133].
- (4)
- The reduction of indole pyruvic acid yields indole lactate, the dehydration of which yields indole acrylic acid. This compound can be reduced to indole propionic acid. Indole propionic acid can be further converted by human enzymes.
4.6. Other Metabolites
5. Supporting Clinical Decision Making, Diagnostic Applications
6. Future Directions
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Ducreux, M.; Cuhna, A.S.; Caramella, C.; Hollebecque, A.; Burtin, P.; Goere, D.; Seufferlein, T.; Haustermans, K.; Van Laethem, J.L.; Conroy, T.; et al. Cancer of the pancreas: ESMO Clinical Practice Guidelines for diagnosis, treatment and follow-up. Ann. Oncol. 2015, 26, 56–68. [Google Scholar] [CrossRef]
- Rawla, P.; Sunkara, T.; Gaduputi, V. Epidemiology of Pancreatic Cancer: Global Trends, Etiology and Risk Factors. World J. Oncol. 2019, 10, 10–27. [Google Scholar] [CrossRef]
- Pei, X.; Song, F.; Wang, Z. Emerging incidence trends and application of curative treatments of pancreatic cancer in the USA. Medicine 2019, 98, e17175. [Google Scholar] [CrossRef]
- Rahib, L.; Smith, B.D.; Aizenberg, R.; Rosenzweig, A.B.; Fleshman, J.M.; Matrisian, L.M. Projecting cancer incidence and deaths to 2030: The unexpected burden of thyroid, liver, and pancreas cancers in the United States. Cancer Res. 2014, 74, 2913–2921. [Google Scholar] [CrossRef]
- Chun, Y.S.; Pawlik, T.M.; Vauthey, J.N. 8th Edition of the AJCC Cancer Staging Manual: Pancreas and Hepatobiliary Cancers. Ann. Surg. Oncol. 2018, 25, 845–847. [Google Scholar] [CrossRef] [PubMed]
- Lowery, M.A.; Kelsen, D.P.; Capanu, M.; Smith, S.C.; Lee, J.W.; Stadler, Z.K.; Moore, M.J.; Kindler, H.L.; Golan, T.; Segal, A.; et al. Phase II trial of veliparib in patients with previously treated BRCA-mutated pancreas ductal adenocarcinoma. Eur. J. Cancer 2018, 89, 19–26. [Google Scholar] [CrossRef] [PubMed]
- Silvestris, N.; Longo, V.; Cellini, F.; Reni, M.; Bittoni, A.; Cataldo, I.; Partelli, S.; Falconi, M.; Scarpa, A.; Brunetti, O.; et al. Neoadjuvant multimodal treatment of pancreatic ductal adenocarcinoma. Crit. Rev. Oncol. Hematol. 2016, 98, 309–324. [Google Scholar] [CrossRef] [PubMed]
- Yeo, T.P. Demographics, epidemiology, and inheritance of pancreatic ductal adenocarcinoma. Semin. Oncol. 2015, 42, 8–18. [Google Scholar] [CrossRef] [PubMed]
- Waddell, N.; Pajic, M.; Patch, A.M.; Chang, D.K.; Kassahn, K.S.; Bailey, P.; Johns, A.L.; Miller, D.; Nones, K.; Quek, K.; et al. Whole genomes redefine the mutational landscape of pancreatic cancer. Nature 2015, 518, 495–501. [Google Scholar] [CrossRef] [PubMed]
- Hashimoto, D.; Arima, K.; Yokoyama, N.; Chikamoto, A.; Taki, K.; Inoue, R.; Kaida, T.; Higashi, T.; Nitta, H.; Ohmuraya, M.; et al. Heterogeneity of KRAS Mutations in Pancreatic Ductal Adenocarcinoma. Pancreas 2016, 45, 1111–1114. [Google Scholar] [CrossRef]
- Schwabe, R.F.; Jobin, C. The microbiome and cancer. Nat. Rev. Cancer 2013, 13, 800–812. [Google Scholar] [CrossRef] [PubMed]
- Flores, R.; Shi, J.; Fuhrman, B.; Xu, X.; Veenstra, T.D.; Gail, M.H.; Gajer, P.; Ravel, J.; Goedert, J.J. Fecal microbial determinants of fecal and systemic estrogens and estrogen metabolites: A cross-sectional study. J. Transl. Med. 2012, 10, 253. [Google Scholar] [CrossRef] [PubMed]
- Goedert, J.J.; Jones, G.; Hua, X.; Xu, X.; Yu, G.; Flores, R.; Falk, R.T.; Gail, M.H.; Shi, J.; Ravel, J.; et al. Investigation of the association between the fecal microbiota and breast cancer in postmenopausal women: A population-based case-control pilot study. J. Natl. Cancer Inst. 2015, 107, 147. [Google Scholar] [CrossRef] [PubMed]
- Robinson, K.M.; Crabtree, J.; Mattick, J.S.; Anderson, K.E.; Dunning Hotopp, J.C. Distinguishing potential bacteria-tumor associations from contamination in a secondary data analysis of public cancer genome sequence data. Microbiome 2017. [Google Scholar] [CrossRef] [PubMed]
- Chen, J.; Pitmon, E.; Wang, K. Microbiome, inflammation and colorectal cancer. Semin. Immunol. 2017, 32, 43–53. [Google Scholar] [CrossRef] [PubMed]
- Picardo, S.L.; Coburn, B.; Hansen, A.R. The microbiome and cancer for clinicians. Crit. Rev. Oncol. Hematol. 2019, 141, 1–12. [Google Scholar] [CrossRef]
- Saus, E.; Iraola-Guzman, S.; Willis, J.R.; Brunet-Vega, A.; Gabaldon, T. Microbiome and colorectal cancer: Roles in carcinogenesis and clinical potential. Mol. Asp. Med. 2019. [Google Scholar] [CrossRef]
- De Martel, C.; Ferlay, J.; Franceschi, S.; Vignat, J.; Bray, F.; Forman, D.; Plummer, M. Global burden of cancers attributable to infections in 2008: A review and synthetic analysis. Lancet Oncol. 2012, 13, 607–615. [Google Scholar] [CrossRef]
- Miko, E.; Vida, A.; Bai, P. Translational aspects of the microbiome-to be exploited. Cell Biol. Toxicol. 2016, 32, 153–156. [Google Scholar] [CrossRef]
- Hall, A.; Versalovic, J. Microbial Metabolism in the Mammalian Gut: Molecular Mechanisms and Clinical Implications. J. Pediatr. Gastroenterol. Nutr. 2018, 66, S72–S79. [Google Scholar] [CrossRef]
- Zitvogel, L.; Ayyoub, M.; Routy, B.; Kroemer, G. Microbiome and Anticancer Immunosurveillance. Cell 2016, 165, 276–287. [Google Scholar] [CrossRef] [PubMed]
- Kovács, T.; Mikó, E.; Vida, A.; Sebő, É.; Toth, J.; Csonka, T.; Boratkó, A.; Ujlaki, G.; Lente, G.; Kovács, P.; et al. Cadaverine, a metabolite of the microbiome, reduces breast cancer aggressiveness through trace amino acid receptors. Sci. Rep. 2019, 9, 1300. [Google Scholar] [CrossRef] [PubMed]
- Miko, E.; Vida, A.; Kovacs, T.; Ujlaki, G.; Trencsenyi, G.; Marton, J.; Sari, Z.; Kovacs, P.; Boratko, A.; Hujber, Z.; et al. Lithocholic acid, a bacterial metabolite reduces breast cancer cell proliferation and aggressiveness. Biochim. Biophys. Acta 2018, 1859, 958–974. [Google Scholar] [CrossRef] [PubMed]
- Buchta Rosean, C.; Bostic, R.R.; Ferey, J.C.M.; Feng, T.Y.; Azar, F.N.; Tung, K.S.; Dozmorov, M.G.; Smirnova, E.; Bos, P.D.; Rutkowski, M.R. Preexisting Commensal Dysbiosis Is a Host-Intrinsic Regulator of Tissue Inflammation and Tumor Cell Dissemination in Hormone Receptor-Positive Breast Cancer. Cancer Res. 2019, 79, 3662–3675. [Google Scholar] [CrossRef]
- Ingman, W.V. The Gut Microbiome: A New Player in Breast Cancer Metastasis. Cancer Res. 2019, 79, 3539–3541. [Google Scholar] [CrossRef]
- Viaud, S.; Daillere, R.; Boneca, I.G.; Lepage, P.; Pittet, M.J.; Ghiringhelli, F.; Trinchieri, G.; Goldszmid, R.; Zitvogel, L. Harnessing the intestinal microbiome for optimal therapeutic immunomodulation. Cancer Res. 2014, 74, 4217–4221. [Google Scholar] [CrossRef]
- Routy, B.; Le Chatelier, E.; Derosa, L.; Duong, C.P.M.; Alou, M.T.; Daillere, R.; Fluckiger, A.; Messaoudene, M.; Rauber, C.; Roberti, M.P.; et al. Gut microbiome influences efficacy of PD-1-based immunotherapy against epithelial tumors. Science 2018, 359, 91–97. [Google Scholar] [CrossRef]
- Elkrief, A.; Derosa, L.; Kroemer, G.; Zitvogel, L.; Routy, B. The negative impact of antibiotics on outcomes in cancer patients treated with immunotherapy: A new independent prognostic factor? Ann. Oncol. 2019, 30, 1572–1579. [Google Scholar] [CrossRef]
- Kovacs, T.; Miko, E.; Ujlaki, G.; Sari, Z.; Bai, P. The Microbiome as a Component of the Tumor Microenvironment. Adv. Exp. Med. Biol. 2020, 1225, 137–153. [Google Scholar]
- Miko, E.; Kovacs, T.; Sebo, E.; Toth, J.; Csonka, T.; Ujlaki, G.; Sipos, A.; Szabo, J.; Mehes, G.; Bai, P. Microbiome-Microbial Metabolome-Cancer Cell Interactions in Breast Cancer-Familiar, but Unexplored. Cells 2019, 8, 293. [Google Scholar] [CrossRef]
- Hanahan, D.; Weinberg, R.A. Hallmarks of cancer: The next generation. Cell 2011, 144, 646–674. [Google Scholar] [CrossRef] [PubMed]
- Hanahan, D.; Weinberg, R.A. The hallmarks of cancer. Cell 2000, 100, 57–70. [Google Scholar] [CrossRef]
- Zitvogel, L.; Daillere, R.; Roberti, M.P.; Routy, B.; Kroemer, G. Anticancer effects of the microbiome and its products. Nat. Rev. Microbiol. 2017, 15, 465–478. [Google Scholar] [CrossRef] [PubMed]
- Kovács, P.; Csonka, T.; Kovács, T.; Sári, Z.; Ujlaki, G.; Sipos, A.; Karányi, Z.; Szeőcs, D.; Hegedűs, C.; Uray, K.; et al. Lithocholic acid, a metabolite of the microbiome, increases oxidative stress in breast cancer. Cancers 2019, 11, 1255. [Google Scholar] [CrossRef] [PubMed]
- Vergara, D.; Simeone, P.; Damato, M.; Maffia, M.; Lanuti, P.; Trerotola, M. The Cancer Microbiota: EMT and Inflammation as Shared Molecular Mechanisms Associated with Plasticity and Progression. J. Oncol. 2019. [Google Scholar] [CrossRef] [PubMed]
- Lau, A.; Villeneuve, N.F.; Sun, Z.; Wong, P.K.; Zhang, D.D. Dual roles of Nrf2 in cancer. Pharmacol. Res. 2008, 58, 262–270. [Google Scholar] [CrossRef] [PubMed]
- Jezierska-Drutel, A.; Rosenzweig, S.A.; Neumann, C.A. Role of oxidative stress and the microenvironment in breast cancer development and progression. Adv. Cancer Res. 2013, 119, 107–125. [Google Scholar]
- Smolková, K.; Mikó, E.; Kovács, T.; Leguina-Ruzzi, A.; Sipos, A.; Bai, P. NRF2 in regulating cancer metabolism. Antioxid. Redox Signal. 2020. [Google Scholar] [CrossRef]
- Pagliari, D.; Saviano, A.; Newton, E.E.; Serricchio, M.L.; Dal Lago, A.A.; Gasbarrini, A.; Cianci, R. Gut Microbiota-Immune System Crosstalk and Pancreatic Disorders. Mediators Inflamm. 2018, 2018, 7946431. [Google Scholar] [CrossRef]
- Ochi, A.; Nguyen, A.H.; Bedrosian, A.S.; Mushlin, H.M.; Zarbakhsh, S.; Barilla, R.; Zambirinis, C.P.; Fallon, N.C.; Rehman, A.; Pylayeva-Gupta, Y.; et al. MyD88 inhibition amplifies dendritic cell capacity to promote pancreatic carcinogenesis via Th2 cells. J. Exp. Med. 2012, 209, 1671–1687. [Google Scholar] [CrossRef]
- Pushalkar, S.; Hundeyin, M.; Daley, D.; Zambirinis, C.P.; Kurz, E.; Mishra, A.; Mohan, N.; Aykut, B.; Usyk, M.; Torres, L.E.; et al. The Pancreatic Cancer Microbiome Promotes Oncogenesis by Induction of Innate and Adaptive Immune Suppression. Cancer Discov. 2018, 8, 403–416. [Google Scholar] [CrossRef] [PubMed]
- Sethi, V.; Kurtom, S.; Tarique, M.; Lavania, S.; Malchiodi, Z.; Hellmund, L.; Zhang, L.; Sharma, U.; Giri, B.; Garg, B.; et al. Gut Microbiota Promotes Tumor Growth in Mice by Modulating Immune Response. Gastroenterology 2018, 155, 33–37. [Google Scholar] [CrossRef] [PubMed]
- Ren, Z.; Jiang, J.; Xie, H.; Li, A.; Lu, H.; Xu, S.; Zhou, L.; Zhang, H.; Cui, G.; Chen, X.; et al. Gut microbial profile analysis by MiSeq sequencing of pancreatic carcinoma patients in China. Oncotarget 2017, 8, 95176–95191. [Google Scholar] [CrossRef] [PubMed]
- Gopalakrishnan, V.; Spencer, C.N.; Nezi, L.; Reuben, A.; Andrews, M.C.; Karpinets, T.V.; Prieto, P.A.; Vicente, D.; Hoffman, K.; Wei, S.C.; et al. Gut microbiome modulates response to anti-PD-1 immunotherapy in melanoma patients. Science 2018, 359, 97–103. [Google Scholar] [CrossRef] [PubMed]
- Buchta Rosean, C.; Feng, T.Y.; Azar, F.N.; Rutkowski, M.R. Impact of the microbiome on cancer progression and response to anti-cancer therapies. Adv. Cancer Res. 2019, 143, 255–294. [Google Scholar] [CrossRef] [PubMed]
- Valastyan, S.; Weinberg, R.A. Tumor metastasis: Molecular insights and evolving paradigms. Cell 2011, 147, 275–292. [Google Scholar] [CrossRef] [PubMed]
- Kalluri, R.; Weinberg, R.A. The basics of epithelial-mesenchymal transition. J. Clin. Investig. 2009, 119, 1420–1428. [Google Scholar] [CrossRef]
- Mani, S.A.; Guo, W.; Liao, M.J.; Eaton, E.N.; Ayyanan, A.; Zhou, A.Y.; Brooks, M.; Reinhard, F.; Zhang, C.C.; Shipitsin, M.; et al. The epithelial-mesenchymal transition generates cells with properties of stem cells. Cell 2008, 133, 704–715. [Google Scholar] [CrossRef]
- Biedermann, L.; Zeitz, J.; Mwinyi, J.; Sutter-Minder, E.; Rehman, A.; Ott, S.J.; Steurer-Stey, C.; Frei, A.; Frei, P.; Scharl, M.; et al. Smoking cessation induces profound changes in the composition of the intestinal microbiota in humans. PLoS ONE 2013, 8, e59260. [Google Scholar] [CrossRef]
- Schulz, M.D.; Atay, C.; Heringer, J.; Romrig, F.K.; Schwitalla, S.; Aydin, B.; Ziegler, P.K.; Varga, J.; Reindl, W.; Pommerenke, C.; et al. High-fat-diet-mediated dysbiosis promotes intestinal carcinogenesis independently of obesity. Nature 2014, 514, 508–512. [Google Scholar] [CrossRef]
- Zarrinpar, A.; Chaix, A.; Panda, S. Daily Eating Patterns and Their Impact on Health and Disease. Trends Endocrinol. Metab. 2016, 27, 69–83. [Google Scholar] [CrossRef] [PubMed]
- Zarrinpar, A.; Chaix, A.; Yooseph, S.; Panda, S. Diet and feeding pattern affect the diurnal dynamics of the gut microbiome. Cell Metab. 2014, 20, 1006–1017. [Google Scholar] [CrossRef]
- Paschos, G.K.; FitzGerald, G.A. Circadian Clocks and Metabolism: Implications for Microbiome and Aging. Trends Genet. 2017, 33, 760–769. [Google Scholar] [CrossRef] [PubMed]
- Zhang, H.; Cherian, R.; Jin, K. Systemic milieu and age-related deterioration. Geroscience 2019, 41, 275–284. [Google Scholar] [CrossRef] [PubMed]
- Saffrey, M.J. Aging of the mammalian gastrointestinal tract: A complex organ system. Age 2014, 36, 9603. [Google Scholar] [CrossRef]
- Singh, H.; Torralba, M.G.; Moncera, K.J.; DiLello, L.; Petrini, J.; Nelson, K.E.; Pieper, R. Gastro-intestinal and oral microbiome signatures associated with healthy aging. Geroscience 2019. [Google Scholar] [CrossRef]
- Devaraj, S.; Hemarajata, P.; Versalovic, J. The human gut microbiome and body metabolism: Implications for obesity and diabetes. Clin. Chem. 2013, 59, 617–628. [Google Scholar] [CrossRef]
- Ticinesi, A.; Lauretani, F.; Tana, C.; Nouvenne, A.; Ridolo, E.; Meschi, T. Exercise and immune system as modulators of intestinal microbiome: Implications for the gut-muscle axis hypothesis. Exerc. Immunol. Rev. 2019, 25, 84–95. [Google Scholar]
- Friedman, G.D.; Oestreicher, N.; Chan, J.; Quesenberry, C.P., Jr.; Udaltsova, N.; Habel, L.A. Antibiotics and risk of breast cancer: Up to 9 years of follow-up of 2.1 million women. Cancer Epidemiol. Biomark. Prev. 2006, 15, 2102–2106. [Google Scholar] [CrossRef]
- Mendoza, L. Potential effect of probiotics in the treatment of breast cancer. Oncol. Rev. 2019, 13, 422. [Google Scholar] [CrossRef]
- Ranjbar, S.; Seyednejad, S.A.; Azimi, H.; Rezaeizadeh, H.; Rahimi, R. Emerging Roles of Probiotics in Prevention and Treatment of Breast Cancer: A Comprehensive Review of Their Therapeutic Potential. Nutr. Cancer 2019, 71, 1–12. [Google Scholar] [CrossRef]
- Lindkvist, B.; Johansen, D.; Borgstrom, A.; Manjer, J. A prospective study of Helicobacter pylori in relation to the risk for pancreatic cancer. BMC Cancer 2008. [Google Scholar] [CrossRef] [PubMed]
- Farrell, J.J.; Zhang, L.; Zhou, H.; Chia, D.; Elashoff, D.; Akin, D.; Paster, B.J.; Joshipura, K.; Wong, D.T. Variations of oral microbiota are associated with pancreatic diseases including pancreatic cancer. Gut 2012, 61, 582–588. [Google Scholar] [CrossRef] [PubMed]
- Mei, Q.X.; Huang, C.L.; Luo, S.Z.; Zhang, X.M.; Zeng, Y.; Lu, Y.Y. Characterization of the duodenal bacterial microbiota in patients with pancreatic head cancer vs. healthy controls. Pancreatology 2018, 18, 438–445. [Google Scholar] [CrossRef] [PubMed]
- Thomas, R.M.; Gharaibeh, R.Z.; Gauthier, J.; Beveridge, M.; Pope, J.L.; Guijarro, M.V.; Yu, Q.; He, Z.; Ohland, C.; Newsome, R.; et al. Intestinal microbiota enhances pancreatic carcinogenesis in preclinical models. Carcinogenesis 2018, 39, 1068–1078. [Google Scholar] [CrossRef]
- Archibugi, L.; Signoretti, M.; Capurso, G. The Microbiome and Pancreatic Cancer: An Evidence-based Association? J. Clin. Gastroenterol. 2018, 52, S82–S85. [Google Scholar] [CrossRef]
- Mendez, R.; Kesh, K.; Arora, N.; Di Martino, L.; McAllister, F.; Merchant, N.; Banerjee, S.; Banerjee, S. Microbial dysbiosis and polyamine metabolism as predictive markers for early detection of pancreatic cancer. Carcinogenesis 2019, 1. [Google Scholar] [CrossRef]
- Aykut, B.; Pushalkar, S.; Chen, R.; Li, Q.; Abengozar, R.; Kim, J.I.; Shadaloey, S.A.; Wu, D.; Preiss, P.; Verma, N.; et al. The fungal mycobiome promotes pancreatic oncogenesis via activation of MBL. Nature 2019, 574, 264–267. [Google Scholar] [CrossRef]
- Frulloni, L.; Lunardi, C.; Simone, R.; Dolcino, M.; Scattolini, C.; Falconi, M.; Benini, L.; Vantini, I.; Corrocher, R.; Puccetti, A. Identification of a novel antibody associated with autoimmune pancreatitis. N. Engl. J. Med. 2009, 361, 2135–2142. [Google Scholar] [CrossRef]
- Risch, H.A.; Lu, L.; Kidd, M.S.; Wang, J.; Zhang, W.; Ni, Q.; Gao, Y.T.; Yu, H. Helicobacter pylori seropositivities and risk of pancreatic carcinoma. Cancer Epidemiol. Biomark. Prev. 2014, 23, 172–178. [Google Scholar] [CrossRef]
- Wang, Y.; Zhang, F.C.; Wang, Y.J. Helicobacter pylori and pancreatic cancer risk: A meta- analysis based on 2049 cases and 2861 controls. Asian Pac. J. Cancer Prev. 2014, 15, 4449–4454. [Google Scholar] [CrossRef] [PubMed]
- Rabelo-Goncalves, E.M.; Roesler, B.M.; Zeitune, J.M. Extragastric manifestations of Helicobacter pylori infection: Possible role of bacterium in liver and pancreas diseases. World J. Hepatol. 2015, 7, 2968–2979. [Google Scholar] [CrossRef] [PubMed]
- Chen, X.Z.; Wang, R.; Chen, H.N.; Hu, J.K. Cytotoxin-Associated Gene A-Negative Strains of Helicobacter pylori as a Potential Risk Factor of Pancreatic Cancer: A Meta-Analysis Based on Nested Case-Control Studies. Pancreas 2015, 44, 1340–1344. [Google Scholar] [CrossRef] [PubMed]
- Maisonneuve, P.; Lowenfels, A.B. Risk factors for pancreatic cancer: A summary review of meta-analytical studies. Int. J. Epidemiol. 2015, 44, 186–198. [Google Scholar] [CrossRef] [PubMed]
- Ertz-Archambault, N.; Keim, P.; Von Hoff, D. Microbiome and pancreatic cancer: A comprehensive topic review of literature. World J. Gastroenterol. 2017, 23, 1899–1908. [Google Scholar] [CrossRef] [PubMed]
- Tenti, P.; Aguzzi, A.; Riva, C.; Usellini, L.; Zappatore, R.; Bara, J.; Samloff, I.M.; Solcia, E. Ovarian mucinous tumors frequently express markers of gastric, intestinal, and pancreatobiliary epithelial cells. Cancer 1992, 69, 2131–2142. [Google Scholar] [CrossRef]
- Ogrendik, M. Periodontal Pathogens in the Etiology of Pancreatic Cancer. Gastrointest. Tumors. 2017, 3, 125–127. [Google Scholar] [CrossRef]
- Wei, M.Y.; Shi, S.; Liang, C.; Meng, Q.C.; Hua, J.; Zhang, Y.Y.; Liu, J.; Zhang, B.; Xu, J.; Yu, X.J. The microbiota and microbiome in pancreatic cancer: More influential than expected. Mol. Cancer 2019, 18, 97. [Google Scholar] [CrossRef]
- Zambirinis, C.P.; Pushalkar, S.; Saxena, D.; Miller, G. Pancreatic cancer, inflammation, and microbiome. Cancer J. 2014, 20, 195–202. [Google Scholar] [CrossRef]
- Mima, K.; Nakagawa, S.; Sawayama, H.; Ishimoto, T.; Imai, K.; Iwatsuki, M.; Hashimoto, D.; Baba, Y.; Yamashita, Y.I.; Yoshida, N.; et al. The microbiome and hepatobiliary-pancreatic cancers. Cancer Lett. 2017, 402, 9–15. [Google Scholar] [CrossRef]
- Hujoel, P.P.; Drangsholt, M.; Spiekerman, C.; Weiss, N.S. An exploration of the periodontitis-cancer association. Ann. Epidemiol. 2003, 13, 312–316. [Google Scholar] [CrossRef]
- Michaud, D.S.; Izard, J.; Wilhelm-Benartzi, C.S.; You, D.H.; Grote, V.A.; Tjonneland, A.; Dahm, C.C.; Overvad, K.; Jenab, M.; Fedirko, V.; et al. Plasma antibodies to oral bacteria and risk of pancreatic cancer in a large European prospective cohort study. Gut 2013, 62, 1764–1770. [Google Scholar] [CrossRef] [PubMed]
- Torres, P.J.; Fletcher, E.M.; Gibbons, S.M.; Bouvet, M.; Doran, K.S.; Kelley, S.T. Characterization of the salivary microbiome in patients with pancreatic cancer. PeerJ 2015, 3, e1373. [Google Scholar] [CrossRef]
- Nieminen, M.T.; Listyarifah, D.; Hagstrom, J.; Haglund, C.; Grenier, D.; Nordstrom, D.; Uitto, V.J.; Hernandez, M.; Yucel-Lindberg, T.; Tervahartiala, T.; et al. Treponema denticola chymotrypsin-like proteinase may contribute to orodigestive carcinogenesis through immunomodulation. Br. J. Cancer 2018, 118, 428–434. [Google Scholar] [CrossRef] [PubMed]
- Fan, X.; Alekseyenko, A.V.; Wu, J.; Peters, B.A.; Jacobs, E.J.; Gapstur, S.M.; Purdue, M.P.; Abnet, C.C.; Stolzenberg-Solomon, R.; Miller, G.; et al. Human oral microbiome and prospective risk for pancreatic cancer: A population-based nested case-control study. Gut 2018, 67, 120–127. [Google Scholar] [CrossRef] [PubMed]
- Lu, H.; Ren, Z.; Li, A.; Li, J.; Xu, S.; Zhang, H.; Jiang, J.; Yang, J.; Luo, Q.; Zhou, K.; et al. Tongue coating microbiome data distinguish patients with pancreatic head cancer from healthy controls. J. Oral Microbiol. 2019, 11, 1563409. [Google Scholar] [CrossRef]
- Gaiser, R.A.; Halimi, A.; Alkharaan, H.; Lu, L.; Davanian, H.; Healy, K.; Hugerth, L.W.; Ateeb, Z.; Valente, R.; Fernandez Moro, C.; et al. Enrichment of oral microbiota in early cystic precursors to invasive pancreatic cancer. Gut 2019, 68, 2186–2194. [Google Scholar] [CrossRef]
- Vogtmann, E.; Han, Y.; Caporaso, J.G.; Bokulich, N.; Mohamadkhani, A.; Moayyedkazemi, A.; Hua, X.; Kamangar, F.; Wan, Y.; Suman, S.; et al. Oral microbial community composition is associated with pancreatic cancer: A case-control study in Iran. Cancer Med. 2020, 9, 797–806. [Google Scholar] [CrossRef]
- Mitsuhashi, K.; Nosho, K.; Sukawa, Y.; Matsunaga, Y.; Ito, M.; Kurihara, H.; Kanno, S.; Igarashi, H.; Naito, T.; Adachi, Y.; et al. Association of Fusobacterium species in pancreatic cancer tissues with molecular features and prognosis. Oncotarget 2015, 6, 7209–7220. [Google Scholar] [CrossRef]
- Maekawa, T.; Fukaya, R.; Takamatsu, S.; Itoyama, S.; Fukuoka, T.; Yamada, M.; Hata, T.; Nagaoka, S.; Kawamoto, K.; Eguchi, H.; et al. Possible involvement of Enterococcus infection in the pathogenesis of chronic pancreatitis and cancer. Biochem. Biophys. Res. Commun. 2018, 506, 962–969. [Google Scholar] [CrossRef]
- Riquelme, E.; Zhang, Y.; Zhang, L.; Montiel, M.; Zoltan, M.; Dong, W.; Quesada, P.; Sahin, I.; Chandra, V.; San Lucas, A.; et al. Tumor Microbiome Diversity and Composition Influence Pancreatic Cancer Outcomes. Cell 2019, 178, 795–806. [Google Scholar] [CrossRef] [PubMed]
- Swan, C.D.; Nahm, C.; Samra, J.S.; Mittal, A.; Figtree, M. Microbiology of pancreatoduodenectomy and recommendations for antimicrobial prophylaxis. ANZ J. Surg. 2019, 19, 15560. [Google Scholar] [CrossRef] [PubMed]
- Di Carlo, P.; Serra, N.; D’Arpa, F.; Agrusa, A.; Gulotta, G.; Fasciana, T.; Rodolico, V.; Giammanco, A.; Sergi, C. The microbiota of the bilio-pancreatic system: A cohort, STROBE-compliant study. Infect. Drug Resist. 2019, 12, 1513–1527. [Google Scholar] [CrossRef] [PubMed]
- Del Castillo, E.; Meier, R.; Chung, M.; Koestler, D.C.; Chen, T.; Paster, B.J.; Charpentier, K.P.; Kelsey, K.T.; Izard, J.; Michaud, D.S. The Microbiomes of Pancreatic and Duodenum Tissue Overlap and Are Highly Subject Specific but Differ between Pancreatic Cancer and Noncancer Subjects. Cancer Epidemiol. Biomark. Prev. 2019, 28, 370–383. [Google Scholar] [CrossRef] [PubMed]
- Boursi, B.; Mamtani, R.; Haynes, K.; Yang, Y.X. Recurrent antibiotic exposure may promote cancer formation--Another step in understanding the role of the human microbiota? Eur. J. Cancer 2015, 51, 2655–2664. [Google Scholar] [CrossRef]
- Tsuji, Y.; Watanabe, T.; Kudo, M.; Arai, H.; Strober, W.; Chiba, T. Sensing of commensal organisms by the intracellular sensor NOD1 mediates experimental pancreatitis. Immunity 2012, 37, 326–338. [Google Scholar] [CrossRef] [PubMed]
- Leal-Lopes, C.; Velloso, F.J.; Campopiano, J.C.; Sogayar, M.C.; Correa, R.G. Roles of Commensal Microbiota in Pancreas Homeostasis and Pancreatic Pathologies. J. Diabetes Res. 2015, 2015, 284680. [Google Scholar] [CrossRef]
- Metagenomics. Alpha and Beta Diversity. Available online: http://www.metagenomics.wiki/pdf/definition/alpha-beta-diversity (accessed on 20 April 2020).
- Vida, A.; Bodrogi, B.L.; Balogh, B.; Bai, P. Taxamat: Automated biodiversity data management tool – Implications for microbiome studies. Physiol. Int. 2020. [Google Scholar] [CrossRef]
- Half, E.; Keren, N.; Reshef, L.; Dorfman, T.; Lachter, I.; Kluger, Y.; Reshef, N.; Knobler, H.; Maor, Y.; Stein, A.; et al. Fecal microbiome signatures of pancreatic cancer patients. Sci. Rep. 2019, 9, 16801. [Google Scholar] [CrossRef]
- Michaud, D.S. Role of bacterial infections in pancreatic cancer. Carcinogenesis 2013, 34, 2193–2197. [Google Scholar] [CrossRef]
- Poore, G.D.; Kopylova, E.; Zhu, Q.; Carpenter, C.; Fraraccio, S.; Wandro, S.; Kosciolek, T.; Janssen, S.; Metcalf, J.; Song, S.J.; et al. Microbiome analyses of blood and tissues suggest cancer diagnostic approach. Nature 2020, 579, 567–574. [Google Scholar] [CrossRef] [PubMed]
- Hujoel, P.P.; Drangsholt, M.; Spiekerman, C.; DeRouen, T.A. Periodontitis-systemic disease associations in the presence of smoking--causal or coincidental? Periodontol 2002, 30, 51–60. [Google Scholar] [CrossRef] [PubMed]
- Grimmig, T.; Moench, R.; Kreckel, J.; Haack, S.; Rueckert, F.; Rehder, R.; Tripathi, S.; Ribas, C.; Chandraker, A.; Germer, C.T.; et al. Toll Like Receptor 2, 4, and 9 Signaling Promotes Autoregulative Tumor Cell Growth and VEGF/PDGF Expression in Human Pancreatic Cancer. Int. J. Mol. Sci. 2016, 17, 2060. [Google Scholar] [CrossRef] [PubMed]
- Yoshimoto, S.; Loo, T.M.; Atarashi, K.; Kanda, H.; Sato, S.; Oyadomari, S.; Iwakura, Y.; Oshima, K.; Morita, H.; Hattori, M.; et al. Obesity-induced gut microbial metabolite promotes liver cancer through senescence secretome. Nature 2013, 499, 97–101. [Google Scholar] [CrossRef]
- Ridlon, J.M.; Bajaj, J.S. The human gut sterolbiome: Bile acid-microbiome endocrine aspects and therapeutics. Acta Pharm. Sin. B 2015, 5, 99–105. [Google Scholar] [CrossRef]
- Zeng, H.; Umar, S.; Rust, B.; Lazarova, D.; Bordonaro, M. Secondary Bile Acids and Short Chain Fatty Acids in the Colon: A Focus on Colonic Microbiome, Cell Proliferation, Inflammation, and Cancer. Int. J. Mol. Sci. 2019, 20, 1214. [Google Scholar] [CrossRef]
- Sittipo, P.; Shim, J.W.; Lee, Y.K. Microbial Metabolites Determine Host Health and the Status of Some Diseases. Int. J. Mol. Sci. 2019, 20, 5296. [Google Scholar] [CrossRef]
- Tan, J.; McKenzie, C.; Potamitis, M.; Thorburn, A.N.; Mackay, C.R.; Macia, L. The role of short-chain Fatty acids in health and disease. Adv. Immunol. 2014, 121, 91–119. [Google Scholar]
- Morrison, D.J.; Preston, T. Formation of short chain fatty acids by the gut microbiota and their impact on human metabolism. Gut Microbes. 2016, 7, 189–200. [Google Scholar] [CrossRef]
- Den Besten, G.; van Eunen, K.; Groen, A.K.; Venema, K.; Reijngoud, D.J.; Bakker, B.M. The role of short-chain fatty acids in the interplay between diet, gut microbiota, and host energy metabolism. J. Lipid. Res. 2013, 54, 2325–2340. [Google Scholar] [CrossRef]
- Lin, Y.H.; Chen, Y.; Smith, T.C., II; Karna, S.L.R.; Seshu, J. Short-Chain Fatty Acids Alter Metabolic and Virulence Attributes of Borrelia burgdorferi. Infect. Immun. 2018, 86. [Google Scholar] [CrossRef] [PubMed]
- Cherrington, C.A.; Hinton, M.; Pearson, G.R.; Chopra, I. Short-chain organic acids at ph 5.0 kill Escherichia coli and Salmonella spp. without causing membrane perturbation. J. Appl. Bacteriol. 1991, 70, 161–165. [Google Scholar] [CrossRef] [PubMed]
- Duncan, S.H.; Louis, P.; Thomson, J.M.; Flint, H.J. The role of pH in determining the species composition of the human colonic microbiota. Environ. Microbiol. 2009, 11, 2112–2122. [Google Scholar] [CrossRef] [PubMed]
- Prohaszka, L.; Jayarao, B.M.; Fabian, A.; Kovacs, S. The role of intestinal volatile fatty acids in the Salmonella shedding of pigs. Zentralbl. Veterinarmed. B 1990, 37, 570–574. [Google Scholar] [CrossRef] [PubMed]
- Fachi, J.L.; Secca, C.; Rodrigues, P.B.; Mato, F.C.P.; Di Luccia, B.; Felipe, J.S.; Pral, L.P.; Rungue, M.; Rocha, V.M.; Sato, F.T.; et al. Acetate coordinates neutrophil and ILC3 responses against C. difficile through FFAR2. J. Exp. Med. 2020, 217, 133544. [Google Scholar] [CrossRef]
- Levy, M.; Thaiss, C.A.; Elinav, E. Metabolites: Messengers between the microbiota and the immune system. Genes Dev. 2016, 30, 1589–1597. [Google Scholar] [CrossRef]
- Derrien, M.; Vaughan, E.E.; Plugge, C.M.; de Vos, W.M. Akkermansia muciniphila gen. nov., sp. nov., a human intestinal mucin-degrading bacterium. Int. J. Syst. Evol. Microbiol. 2004, 54, 1469–1476. [Google Scholar] [CrossRef]
- Reichardt, N.; Duncan, S.H.; Young, P.; Belenguer, A.; McWilliam Leitch, C.; Scott, K.P.; Flint, H.J.; Louis, P. Phylogenetic distribution of three pathways for propionate production within the human gut microbiota. ISME J. 2014, 8, 1323–1335. [Google Scholar] [CrossRef]
- Fernández, J.; Redondo-Blanco, S.; Gutiérrez-del-Río, I.; Miguélez, E.M.; Villar, C.J.; Lombó, F. Colon microbiota fermentation of dietary prebiotics towards short-chain fatty acids and their roles as anti-inflammatory and antitumour agents: A review. J. Funct. Foods 2016, 25, 511–522. [Google Scholar] [CrossRef]
- Koh, A.; De Vadder, F.; Kovatcheva-Datchary, P.; Backhed, F. From Dietary Fiber to Host Physiology: Short-Chain Fatty Acids as Key Bacterial Metabolites. Cell 2016, 165, 1332–1345. [Google Scholar] [CrossRef]
- Louis, P.; Young, P.; Holtrop, G.; Flint, H.J. Diversity of human colonic butyrate-producing bacteria revealed by analysis of the butyryl-CoA:acetate CoA-transferase gene. Environ. Microbiol. 2010, 12, 304–314. [Google Scholar] [CrossRef] [PubMed]
- Kirkup, B.; McKee, A.; Makin, K.; Paveley, J.; Caim, S.; Alcon-Giner, C.; Leclaire, C.; Dalby, M.; Le Gall, G.; Andrusaite, A.; et al. Perturbation of the gut microbiota by antibiotics results in accelerated breast tumour growth and metabolic dysregulation. BioRxiv 2019. [Google Scholar] [CrossRef]
- Gerard, P. Metabolism of cholesterol and bile acids by the gut microbiota. Pathogens 2013, 3, 14–24. [Google Scholar] [CrossRef] [PubMed]
- Ridlon, J.M.; Harris, S.C.; Bhowmik, S.; Kang, D.J.; Hylemon, P.B. Consequences of bile salt biotransformations by intestinal bacteria. Gut Microbes. 2016, 7, 22–39. [Google Scholar] [CrossRef] [PubMed]
- Long, S.L.; Gahan, C.G.M.; Joyce, S.A. Interactions between gut bacteria and bile in health and disease. Mol. Asp. Med. 2017, 56, 54–65. [Google Scholar] [CrossRef] [PubMed]
- Hirano, S.; Masuda, N. Transformation of bile acids by Eubacterium lentum. Appl. Environ. Microbiol. 1981, 42, 912–915. [Google Scholar] [CrossRef]
- Hirano, S.; Masuda, N.; Mukai, H.; Hirakawa, K.; Imamura, T. Transformation of bile acids by Bacteroides fragilis strains isolated from the human intestine (author’s transl). Nihon Saikingaku Zasshi 1979, 34, 403–411. [Google Scholar] [CrossRef]
- Salvioli, G.; Salati, R.; Bondi, M.; Fratalocchi, A.; Sala, B.M.; Gibertini, A. Bile acid transformation by the intestinal flora and cholesterol saturation in bile. Effects of Streptococcus faecium administration. Digestion 1982, 23, 80–88. [Google Scholar] [CrossRef]
- Goodwin, A.C.; Destefano Shields, C.E.; Wu, S.; Huso, D.L.; Wu, X.; Murray-Stewart, T.R.; Hacker-Prietz, A.; Rabizadeh, S.; Woster, P.M.; Sears, C.L.; et al. Polyamine catabolism contributes to enterotoxigenic Bacteroides fragilis-induced colon tumorigenesis. Proc. Natl. Acad. Sci. USA 2011, 108, 15354–15359. [Google Scholar] [CrossRef]
- Michael, A.J. Polyamine function in archaea and bacteria. J. Biol. Chem. 2018, 293, 18693–18701. [Google Scholar] [CrossRef]
- De las Rivas, B.; Marcobal, A.; Carrascosa, A.V.; Munoz, R. PCR detection of foodborne bacteria producing the biogenic amines histamine, tyramine, putrescine, and cadaverine. J. Food Prot. 2006, 69, 2509–2514. [Google Scholar] [CrossRef] [PubMed]
- Gao, J.; Xu, K.; Liu, H.; Liu, G.; Bai, M.; Peng, C.; Li, T.; Yin, Y. Impact of the Gut Microbiota on Intestinal Immunity Mediated by Tryptophan Metabolism. Front. Cell. Infect. Microbiol. 2018. [Google Scholar] [CrossRef] [PubMed]
- Clausen, M.R.; Mortensen, P.B.; Bendtsen, F. Serum levels of short-chain fatty acids in cirrhosis and hepatic coma. Hepatology 1991, 14, 1040–1045. [Google Scholar] [CrossRef] [PubMed]
- Jakobsdottir, G.; Bjerregaard, J.H.; Skovbjerg, H.; Nyman, M. Fasting serum concentration of short-chain fatty acids in subjects with microscopic colitis and celiac disease: No difference compared with controls, but between genders. Scand. J. Gastroenterol. 2013, 48, 696–701. [Google Scholar] [CrossRef] [PubMed]
- Ktsoyan, Z.A.; Mkrtchyan, M.S.; Zakharyan, M.K.; Mnatsakanyan, A.A.; Arakelova, K.A.; Gevorgyan, Z.U.; Sedrakyan, A.M.; Hovhannisyan, A.I.; Arakelyan, A.A.; Aminov, R.I. Systemic Concentrations of Short Chain Fatty Acids Are Elevated in Salmonellosis and Exacerbation of Familial Mediterranean Fever. Front. Microbiol. 2016, 7, 776. [Google Scholar] [CrossRef] [PubMed]
- Pryde, S.E.; Duncan, S.H.; Hold, G.L.; Stewart, C.S.; Flint, H.J. The microbiology of butyrate formation in the human colon. FEMS Microbiol. Lett. 2002, 217, 133–139. [Google Scholar] [CrossRef]
- Ivan, J.; Major, E.; Sipos, A.; Kovacs, K.; Horvath, D.; Tamas, I.; Bay, P.; Dombradi, V.; Lontay, B. The Short-Chain Fatty Acid Propionate Inhibits Adipogenic Differentiation of Human Chorion-Derived Mesenchymal Stem Cells Through the Free Fatty Acid Receptor 2. Stem Cells Dev. 2017, 26, 1724–1733. [Google Scholar] [CrossRef]
- Fruge, A.D.; Van der Pol, W.; Rogers, L.Q.; Morrow, C.D.; Tsuruta, Y.; Demark-Wahnefried, W. Fecal Akkermansia muciniphila Is Associated with Body Composition and Microbiota Diversity in Overweight and Obese Women with Breast Cancer Participating in a Presurgical Weight Loss Trial. J. Acad. Nutr. Diet 2018, 9, 164. [Google Scholar] [CrossRef]
- Fellows, R.; Varga-Weisz, P. Chromatin dynamics and histone modifications in intestinal microbiota-host crosstalk. Mol. Metab. 2019, 27, 30951–30956. [Google Scholar] [CrossRef]
- Shimazu, T.; Hirschey, M.D.; Newman, J.; He, W.; Shirakawa, K.; Le Moan, N.; Grueter, C.A.; Lim, H.; Saunders, L.R.; Stevens, R.D.; et al. Suppression of oxidative stress by beta-hydroxybutyrate, an endogenous histone deacetylase inhibitor. Science 2013, 339, 211–214. [Google Scholar] [CrossRef]
- Menzies, K.J.; Zhang, H.; Katsyuba, E.; Auwerx, J. Protein acetylation in metabolism—Metabolites and cofactors. Nat. Rev. Endocrinol. 2016, 12, 43–60. [Google Scholar] [CrossRef] [PubMed]
- Haase, S.; Haghikia, A.; Wilck, N.; Muller, D.N.; Linker, R.A. Impacts of microbiome metabolites on immune regulation and autoimmunity. Immunology 2018, 154, 230–238. [Google Scholar] [CrossRef] [PubMed]
- Ratajczak, W.; Ryl, A.; Mizerski, A.; Walczakiewicz, K.; Sipak, O.; Laszczynska, M. Immunomodulatory potential of gut microbiome-derived short-chain fatty acids (SCFAs). Acta Biochim. Pol. 2019, 66, 1–12. [Google Scholar] [CrossRef] [PubMed]
- Kanika, G.; Khan, S.; Jena, G. Sodium Butyrate Ameliorates L-Arginine-Induced Pancreatitis and Associated Fibrosis in Wistar Rat: Role of Inflammation and Nitrosative Stress. J. Biochem. Mol. Toxicol. 2015, 29, 349–359. [Google Scholar] [CrossRef]
- Bhagat, T.D.; Von Ahrens, D.; Dawlaty, M.; Zou, Y.; Baddour, J.; Achreja, A.; Zhao, H.; Yang, L.; Patel, B.; Kwak, C.; et al. Lactate-mediated epigenetic reprogramming regulates formation of human pancreatic cancer-associated fibroblasts. Elife 2019. [Google Scholar] [CrossRef]
- Mullins, T.D.; Kern, H.F.; Metzgar, R.S. Ultrastructural differentiation of sodium butyrate-treated human pancreatic adenocarcinoma cell lines. Pancreas 1991, 6, 578–587. [Google Scholar] [CrossRef]
- Pellizzaro, C.; Speranza, A.; Zorzet, S.; Crucil, I.; Sava, G.; Scarlata, I.; Cantoni, S.; Fedeli, M.; Coradini, D. Inhibition of human pancreatic cell line MIA PaCa2 proliferation by HA-But, a hyaluronic butyric ester: A preliminary report. Pancreas 2008, 36, e15–e23. [Google Scholar] [CrossRef]
- Iwahashi, S.; Ishibashi, H.; Utsunomiya, T.; Morine, Y.; Ochir, T.L.; Hanaoka, J.; Mori, H.; Ikemoto, T.; Imura, S.; Shimada, M. Effect of histone deacetylase inhibitor in combination with 5-fluorouracil on pancreas cancer and cholangiocarcinoma cell lines. J. Med. Investig. 2011, 58, 106–109. [Google Scholar] [CrossRef]
- Norlin, M.; Wikvall, K. Enzymes in the conversion of cholesterol into bile acids. Curr. Mol. Med. 2007, 7, 199–218. [Google Scholar] [CrossRef]
- Ridlon, J.M.; Kang, D.J.; Hylemon, P.B. Bile salt biotransformations by human intestinal bacteria. J. Lipid. Res. 2006, 47, 241–259. [Google Scholar] [CrossRef]
- Wahlstrom, A.; Sayin, S.I.; Marschall, H.U.; Backhed, F. Intestinal Crosstalk between Bile Acids and Microbiota and Its Impact on Host Metabolism. Cell Metab. 2016, 24, 41–50. [Google Scholar] [CrossRef] [PubMed]
- Midtvedt, T. Microbial bile acid transformation. Am. J. Clin. Nutr. 1974, 27, 1341–1347. [Google Scholar] [CrossRef] [PubMed]
- Marshall, W.J.; Lapsley, M.; Day, A. Clinical Chemistry, 8th ed.; Elsevier: Amsterdam, The Netherlands, 2016. [Google Scholar]
- Watanabe, M.; Houten, S.M.; Mataki, C.; Christoffolete, M.A.; Kim, B.W.; Sato, H.; Messaddeq, N.; Harney, J.W.; Ezaki, O.; Kodama, T.; et al. Bile acids induce energy expenditure by promoting intracellular thyroid hormone activation. Nature 2006, 439, 484–489. [Google Scholar] [CrossRef]
- MahmoudianDehkordi, S.; Arnold, M.; Nho, K.; Ahmad, S.; Jia, W.; Xie, G.; Louie, G.; Kueider-Paisley, A.; Moseley, M.A.; Thompson, J.W.; et al. Altered bile acid profile associates with cognitive impairment in Alzheimer’s disease—An emerging role for gut microbiome. Alzheimers Dement. 2019, 15, 76–92. [Google Scholar] [CrossRef] [PubMed]
- Sarin, S.K.; Pande, A.; Schnabl, B. Microbiome as a therapeutic target in alcohol-related liver disease. J. Hepatol. 2019, 70, 260–272. [Google Scholar] [CrossRef] [PubMed]
- McIlvride, S.; Dixon, P.H.; Williamson, C. Bile acids and gestation. Mol. Asp. Med. 2017, 56, 90–100. [Google Scholar] [CrossRef] [PubMed]
- Houten, S.M.; Watanabe, M.; Auwerx, J. Endocrine functions of bile acids. EMBO J. 2006, 25, 1419–1425. [Google Scholar] [CrossRef]
- Tsuei, J.; Chau, T.; Mills, D.; Wan, Y.J. Bile acid dysregulation, gut dysbiosis, and gastrointestinal cancer. Exp. Biol. Med. 2014, 239, 1489–1504. [Google Scholar] [CrossRef]
- Rowland, I.R. (Ed.) The Role of the Gut Flora in Toxicity and Cancer; Academic Press: Carshalton, UK, 1988. [Google Scholar]
- Goldberg, A.A.; Titorenko, V.I.; Beach, A.; Sanderson, J.T. Bile acids induce apoptosis selectively in androgen-dependent and -independent prostate cancer cells. PeerJ 2013, 1, e122. [Google Scholar] [CrossRef]
- Gafar, A.A.; Draz, H.M.; Goldberg, A.A.; Bashandy, M.A.; Bakry, S.; Khalifa, M.A.; AbuShair, W.; Titorenko, V.I.; Sanderson, J.T. Lithocholic acid induces endoplasmic reticulum stress, autophagy and mitochondrial dysfunction in human prostate cancer cells. PeerJ 2016, 4, e2445. [Google Scholar] [CrossRef]
- Goldberg, A.A.; Beach, A.; Davies, G.F.; Harkness, T.A.; Leblanc, A.; Titorenko, V.I. Lithocholic bile acid selectively kills neuroblastoma cells, while sparing normal neuronal cells. Oncotarget 2011, 2, 761–782. [Google Scholar] [CrossRef] [PubMed]
- Blanchet, M.; Brunel, J.M. Bile Acid Derivatives: From Old Molecules to a New Potent Therapeutic Use: An Overview. Curr. Med. Chem. 2018, 25, 3613–3636. [Google Scholar] [CrossRef] [PubMed]
- Phelan, J.P.; Reen, F.J.; Dunphy, N.; O’Connor, R.; O’Gara, F. Bile acids destabilise HIF-1alpha and promote anti-tumour phenotypes in cancer cell models. BMC Cancer 2016. [Google Scholar] [CrossRef] [PubMed]
- Phelan, J.P.; Reen, F.J.; Caparros-Martin, J.A.; O’Connor, R.; O’Gara, F. Rethinking the bile acid/gut microbiome axis in cancer. Oncotarget 2017, 8, 115736–115747. [Google Scholar] [CrossRef]
- Shellman, Z.; Aldhahrani, A.; Verdon, B.; Mather, M.; Paleri, V.; Wilson, J.; Pearson, J.; Ward, C.; Powell, J. Bile acids: A potential role in the pathogenesis of pharyngeal malignancy. Clin. Otolaryngol. 2017, 42, 969–973. [Google Scholar] [CrossRef]
- Luu, T.H.; Bard, J.M.; Carbonnelle, D.; Chaillou, C.; Huvelin, J.M.; Bobin-Dubigeon, C.; Nazih, H. Lithocholic bile acid inhibits lipogenesis and induces apoptosis in breast cancer cells. Cell. Oncol. 2018, 41, 13–24. [Google Scholar] [CrossRef]
- Merritt, M.E.; Donaldson, J.R. Effect of bile salts on the DNA and membrane integrity of enteric bacteria. J. Med. Microbiol. 2009, 58, 1533–1541. [Google Scholar] [CrossRef]
- Garcia-Quintanilla, M.; Prieto, A.I.; Barnes, L.; Ramos-Morales, F.; Casadesus, J. Bile-induced curing of the virulence plasmid in Salmonella enterica serovar Typhimurium. J. Bacteriol. 2006, 188, 7963–7965. [Google Scholar] [CrossRef]
- Prieto, A.I.; Ramos-Morales, F.; Casadesus, J. Repair of DNA damage induced by bile salts in Salmonella enterica. Genetics 2006, 174, 575–584. [Google Scholar] [CrossRef]
- Prieto, A.I.; Ramos-Morales, F.; Casadesus, J. Bile-induced DNA damage in Salmonella enterica. Genetics 2004, 168, 1787–1794. [Google Scholar] [CrossRef]
- Kandell, R.L.; Bernstein, C. Bile salt/acid induction of DNA damage in bacterial and mammalian cells: Implications for colon cancer. Nutr. Cancer 1991, 16, 227–238. [Google Scholar] [CrossRef] [PubMed]
- Schaffler, H.; Breitruck, A. Clostridium difficile—From Colonization to Infection. Front. Microbiol. 2018, 9. [Google Scholar] [CrossRef] [PubMed]
- Sorg, J.A.; Sonenshein, A.L. Inhibiting the initiation of Clostridium difficile spore germination using analogs of chenodeoxycholic acid, a bile acid. J. Bacteriol. 2010, 192, 4983–4990. [Google Scholar] [CrossRef] [PubMed]
- Slocum, M.M.; Sittig, K.M.; Specian, R.D.; Deitch, E.A. Absence of intestinal bile promotes bacterial translocation. Am. Surg. 1992, 58, 305–310. [Google Scholar]
- Rees, D.O.; Crick, P.J.; Jenkins, G.J.; Wang, Y.; Griffiths, W.J.; Brown, T.H.; Al-Sarireh, B. Comparison of the composition of bile acids in bile of patients with adenocarcinoma of the pancreas and benign disease. J. Steroid. Biochem. Mol. Biol. 2017, 174, 290–295. [Google Scholar] [CrossRef]
- Feng, H.Y.; Chen, Y.C. Role of bile acids in carcinogenesis of pancreatic cancer: An old topic with new perspective. World J. Gastroenterol. 2016, 22, 7463–7477. [Google Scholar] [CrossRef]
- Gandhi, D.; Ojili, V.; Nepal, P.; Nagar, A.; Hernandez-Delima, F.J.; Bajaj, D.; Choudhary, G.; Gupta, N.; Sharma, P. A pictorial review of gall stones and its associated complications. Clin. Imaging 2019, 60, 228–236. [Google Scholar] [CrossRef]
- Fu, H.; Li, Y.; Bai, G.; Yin, R.; Yin, C.; Shi, W.; Zhang, L.; Li, R.; Zhao, R. Persistent cholestasis resulting from duodenal papillary carcinoma in an adolescent male: A case report. Medicine 2019, 98, e15708. [Google Scholar] [CrossRef]
- Thomas, R.M.; Jobin, C. Microbiota in pancreatic health and disease: The next frontier in microbiome research. Nat. Rev. Gastroenterol. Hepatol. 2020, 17, 53–64. [Google Scholar] [CrossRef]
- Bernstein, C.; Bernstein, H.; Payne, C.M.; Dvorak, K.; Garewal, H. Field defects in progression to gastrointestinal tract cancers. Cancer Lett. 2008, 260, 1–10. [Google Scholar] [CrossRef]
- Tucker, O.N.; Dannenberg, A.J.; Yang, E.Y.; Fahey Iii, T.J. Bile acids induce cyclooxygenase-2 expression in human pancreatic cancer cell lines. Carcinogenesis 2004, 25, 419–423. [Google Scholar] [CrossRef] [PubMed]
- Adachi, T.; Tajima, Y.; Kuroki, T.; Mishima, T.; Kitasato, A.; Fukuda, K.; Tsutsumi, R.; Kanematsu, T. Bile-reflux into the pancreatic ducts is associated with the development of intraductal papillary carcinoma in hamsters. J. Surg. Res. 2006, 136, 106–111. [Google Scholar] [CrossRef] [PubMed]
- Ura, H.; Makino, T.; Ito, S.; Tsutsumi, M.; Kinugasa, T.; Kamano, T.; Ichimiya, H.; Konishi, Y. Combined effects of cholecystectomy and lithocholic acid on pancreatic carcinogenesis of N-nitrosobis(2-hydroxypropyl)amine in Syrian golden hamsters. Cancer Res. 1986, 46, 4782–4786. [Google Scholar] [PubMed]
- Nagathihalli, N.S.; Beesetty, Y.; Lee, W.; Washington, M.K.; Chen, X.; Lockhart, A.C.; Merchant, N.B. Novel mechanistic insights into ectodomain shedding of EGFR Ligands Amphiregulin and TGF-α: Impact on gastrointestinal cancers driven by secondary bile acids. Cancer Res. 2014, 74, 2062–2072. [Google Scholar] [CrossRef] [PubMed]
- Kim, Y.J.; Jeong, S.H.; Kim, E.K.; Kim, E.J.; Cho, J.H. Ursodeoxycholic acid suppresses epithelial-mesenchymal transition and cancer stem cell formation by reducing the levels of peroxiredoxin II and reactive oxygen species in pancreatic cancer cells. Oncol. Rep. 2017, 38, 3632–3638. [Google Scholar] [CrossRef] [PubMed]
- Hummel, D.; Aggarwal, A.; Borka, K.; Bajna, E.; Kallay, E.; Horvath, H.C. The vitamin D system is deregulated in pancreatic diseases. J. Steroid. Biochem. Mol. Biol. 2014, 144, 402–409. [Google Scholar] [CrossRef]
- Chen, X.-L.; Xie, K.-X.; Yang, Z.-L.; Yuan, L.-W. Expression of FXR and HRG and their clinicopathological significance in benign and malignant pancreatic lesions. Int. J. Clin. Exp. Pathol. 2019, 12, 2111–2120. [Google Scholar]
- Koutsounas, I.; Giaginis, C.; Alexandrou, P.; Zizi-Serbetzoglou, A.; Patsouris, E.; Kouraklis, G.; Theocharis, S. Pregnane X Receptor Expression in Human Pancreatic Adenocarcinoma: Associations With Clinicopathologic Parameters, Tumor Proliferative Capacity, Patients’ Survival, and Retinoid X Receptor Expression. Pancreas 2015, 44, 1134–1140. [Google Scholar] [CrossRef]
- Kashiwagi, K.; Yanagida, M.; Matsui, D.; Tanaka, M.; Sugimoto, K.; Chen, H.; Ichikawa-Tomikawa, N.; Marubashi, S.; Suzuki, H.; Chiba, H. Expression of liver X receptors in normal and refractory carcinoma tissues of the human lung and pancreas. Histol. Histopathol. 2018, 33, 497–505. [Google Scholar]
- Saraswat, M.; Joenvaara, S.; Seppanen, H.; Mustonen, H.; Haglund, C.; Renkonen, R. Comparative proteomic profiling of the serum differentiates pancreatic cancer from chronic pancreatitis. Cancer Med. 2017, 6, 1738–1751. [Google Scholar] [CrossRef]
- Candelaria, N.R.; Addanki, S.; Zheng, J.; Nguyen-Vu, T.; Karaboga, H.; Dey, P.; Gabbi, C.; Vedin, L.-L.; Liu, K.; Wu, W.; et al. Antiproliferative effects and mechanisms of liver X receptor ligands in pancreatic ductal adenocarcinoma cells. PLoS ONE 2014, 9, e106289. [Google Scholar] [CrossRef] [PubMed]
- Massaro, C.; Thomas, J.; Phanstiel Iv, O. Investigation of Polyamine Metabolism and Homeostasis in Pancreatic Cancers. Med. Sci. 2017, 5. [Google Scholar] [CrossRef]
- Gitto, S.B.; Pandey, V.; Oyer, J.L.; Copik, A.J.; Hogan, F.C.; Phanstiel, O.T.; Altomare, D.A. Difluoromethylornithine Combined with a Polyamine Transport Inhibitor Is Effective against Gemcitabine Resistant Pancreatic Cancer. Mol. Pharm. 2018, 15, 369–376. [Google Scholar] [CrossRef] [PubMed]
- Mohammed, A.; Janakiram, N.B.; Madka, V.; Ritchie, R.L.; Brewer, M.; Biddick, L.; Patlolla, J.M.; Sadeghi, M.; Lightfoot, S.; Steele, V.E.; et al. Eflornithine (DFMO) prevents progression of pancreatic cancer by modulating ornithine decarboxylase signaling. Cancer Prev. Res. 2014, 7, 1198–1209. [Google Scholar] [CrossRef]
- Saydjari, R.; Alexander, R.W.; Barranco, S.C.; Townsend, C.M., Jr.; Thompson, J.C. The effects of 2-deoxy-D-glucose and alpha-difluoromethylornithine on the growth of pancreatic cancer in vivo. Pancreas 1989, 4, 38–43. [Google Scholar] [CrossRef]
- Chang, B.K.; Gutman, R.; Black, O., Jr. Combined effects of alpha-difluoromethylornithine and doxorubicin against pancreatic cancer cell lines in culture. Pancreas 1986, 1, 49–54. [Google Scholar] [CrossRef]
- Chang, B.K.; Bergeron, R.J.; Porter, C.W.; Liang, Y. Antitumor effects of N-alkylated polyamine analogues in human pancreatic adenocarcinoma models. Cancer Chemother. Pharmacol. 1992, 30, 179–182. [Google Scholar] [CrossRef]
- Wortham, B.W.; Patel, C.N.; Oliveira, M.A. Polyamines in bacteria: Pleiotropic effects yet specific mechanisms. Adv. Exp. Med. Biol. 2007, 603, 106–115. [Google Scholar] [CrossRef]
- Miller-Fleming, L.; Olin-Sandoval, V.; Campbell, K.; Ralser, M. Remaining Mysteries of Molecular Biology: The Role of Polyamines in the Cell. J. Mol. Biol. 2015, 427, 3389–3406. [Google Scholar] [CrossRef]
- Seiler, N. Catabolism of polyamines. Amino Acids 2004, 26, 217–233. [Google Scholar] [CrossRef]
- Lu, Y.-C.; Yeh, W.-C.; Ohashi, P.S. LPS/TLR4 signal transduction pathway. Cytokine 2008, 42, 145–151. [Google Scholar] [CrossRef] [PubMed]
- Bertani, B.; Ruiz, N. Function and Biogenesis of Lipopolysaccharides. EcoSal Plus 2018, 8. [Google Scholar] [CrossRef] [PubMed]
- Yokoyama, M.T.; Carlson, J.R. Microbial metabolites of tryptophan in the intestinal tract with special reference to skatole. Am. J. Clin. Nutr. 1979, 32, 173–178. [Google Scholar] [CrossRef] [PubMed]
- Wikoff, W.R.; Anfora, A.T.; Liu, J.; Schultz, P.G.; Lesley, S.A.; Peters, E.C.; Siuzdak, G. Metabolomics analysis reveals large effects of gut microflora on mammalian blood metabolites. Proc. Natl. Acad. Sci. USA 2009, 106, 3698–3703. [Google Scholar] [CrossRef] [PubMed]
- Browne, C.A.; Clarke, G.; Dinan, T.G.; Cryan, J.F. An effective dietary method for chronic tryptophan depletion in two mouse strains illuminates a role for 5-HT in nesting behaviour. Neuropharmacology 2012, 62, 1903–1915. [Google Scholar] [CrossRef] [PubMed]
- El Aidy, S.; Kunze, W.; Bienenstock, J.; Kleerebezem, M. The microbiota and the gut-brain axis: Insights from the temporal and spatial mucosal alterations during colonisation of the germfree mouse intestine. Benef. Microbes. 2012, 3, 251–259. [Google Scholar] [CrossRef]
- Mardinoglu, A.; Shoaie, S.; Bergentall, M.; Ghaffari, P.; Zhang, C.; Larsson, E.; Backhed, F.; Nielsen, J. The gut microbiota modulates host amino acid and glutathione metabolism in mice. Mol. Syst. Biol. 2015, 11, 834. [Google Scholar] [CrossRef]
- Roager, H.M.; Licht, T.R. Microbial tryptophan catabolites in health and disease. Nat. Commun. 2018, 9, 3294. [Google Scholar] [CrossRef]
- DeMoss, R.D.; Moser, K. Tryptophanase in diverse bacterial species. J. Bacteriol. 1969, 98, 167–171. [Google Scholar] [CrossRef]
- Ma, Q.; Zhang, X.; Qu, Y. Biodegradation and Biotransformation of Indole: Advances and Perspectives. Front. Microbiol. 2018, 9, 2625. [Google Scholar] [CrossRef]
- Zelante, T.; Iannitti, R.G.; Cunha, C.; De Luca, A.; Giovannini, G.; Pieraccini, G.; Zecchi, R.; D’Angelo, C.; Massi-Benedetti, C.; Fallarino, F.; et al. Tryptophan catabolites from microbiota engage aryl hydrocarbon receptor and balance mucosal reactivity via interleukin-22. Immunity 2013, 39, 372–385. [Google Scholar] [CrossRef] [PubMed]
- Venkatesh, M.; Mukherjee, S.; Wang, H.; Li, H.; Sun, K.; Benechet, A.P.; Qiu, Z.; Maher, L.; Redinbo, M.R.; Phillips, R.S.; et al. Symbiotic bacterial metabolites regulate gastrointestinal barrier function via the xenobiotic sensor PXR and Toll-like receptor 4. Immunity 2014, 41, 296–310. [Google Scholar] [CrossRef] [PubMed]
- Lamas, B.; Richard, M.L.; Leducq, V.; Pham, H.P.; Michel, M.L.; Da Costa, G.; Bridonneau, C.; Jegou, S.; Hoffmann, T.W.; Natividad, J.M.; et al. CARD9 impacts colitis by altering gut microbiota metabolism of tryptophan into aryl hydrocarbon receptor ligands. Nat. Med. 2016, 22, 598–605. [Google Scholar] [CrossRef] [PubMed]
- Kim, C.H. Immune regulation by microbiome metabolites. Immunology 2018, 154, 220–229. [Google Scholar] [CrossRef]
- Sonner, J.K.; Keil, M.; Falk-Paulsen, M.; Mishra, N.; Rehman, A.; Kramer, M.; Deumelandt, K.; Rowe, J.; Sanghvi, K.; Wolf, L.; et al. Dietary tryptophan links encephalogenicity of autoreactive T cells with gut microbial ecology. Nat. Commun. 2019, 10, 4877. [Google Scholar] [CrossRef]
- Shi, L.Z.; Faith, N.G.; Nakayama, Y.; Suresh, M.; Steinberg, H.; Czuprynski, C.J. The aryl hydrocarbon receptor is required for optimal resistance to Listeria monocytogenes infection in mice. J. Immunol. 2007, 179, 6952–6962. [Google Scholar] [CrossRef]
- Qiu, J.; Heller, J.J.; Guo, X.; Chen, Z.-m.E.; Fish, K.; Fu, Y.-X.; Zhou, L. The aryl hydrocarbon receptor regulates gut immunity through modulation of innate lymphoid cells. Immunity 2012, 36, 92–104. [Google Scholar] [CrossRef]
- Zhang, L.; Nichols, R.G.; Patterson, A.D. The aryl hydrocarbon receptor as a moderator of host-microbiota communication. Curr. Opin. Toxicol. 2017, 2, 30–35. [Google Scholar] [CrossRef]
- Jin, U.H.; Kim, S.B.; Safe, S. Omeprazole Inhibits Pancreatic Cancer Cell Invasion through a Nongenomic Aryl Hydrocarbon Receptor Pathway. Chem. Res. Toxicol. 2015, 28, 907–918. [Google Scholar] [CrossRef]
- Ansari, D.; Toren, W.; Zhou, Q.; Hu, D.; Andersson, R. Proteomic and genomic profiling of pancreatic cancer. Cell Biol. Toxicol. 2019, 35, 333–343. [Google Scholar] [CrossRef]
- Brown, J.M.; Hazen, S.L. Targeting of microbe-derived metabolites to improve human health: The next frontier for drug discovery. J. Biol. Chem. 2017, 292, 8560–8568. [Google Scholar] [CrossRef] [PubMed]
- Descamps, H.C.; Herrmann, B.; Wiredu, D.; Thaiss, C.A. The path toward using microbial metabolites as therapies. EBioMedicine 2019, 44, 747–754. [Google Scholar] [CrossRef] [PubMed]
- Szanto, M.; Dozsa, A.; Antal, D.; Szabo, K.; Kemeny, L.; Bai, P. Targeting the gut-skin axis—Probiotics as new tools for skin disorder management? Exp. Dermatol. 2019. [Google Scholar] [CrossRef] [PubMed]
- Velicer, C.M.; Heckbert, S.R.; Lampe, J.W.; Potter, J.D.; Robertson, C.A.; Taplin, S.H. Antibiotic use in relation to the risk of breast cancer. JAMA 2004, 291, 827–835. [Google Scholar] [CrossRef] [PubMed]
- Varian, B.J.; Goureshetti, S.; Poutahidis, T.; Lakritz, J.R.; Levkovich, T.; Kwok, C.; Teliousis, K.; Ibrahim, Y.M.; Mirabal, S.; Erdman, S.E. Beneficial bacteria inhibit cachexia. Oncotarget 2016, 7, 11803–11816. [Google Scholar] [CrossRef]
- Mills, S.; Stanton, C.; Lane, J.A.; Smith, G.J.; Ross, R.P. Precision Nutrition and the Microbiome, Part I: Current State of the Science. Nutrients 2019, 11, 923. [Google Scholar] [CrossRef]
| Sample Type and Sample Size | Method | Changes to Microbiome | Other Observations | Ref. |
|---|---|---|---|---|
| Changes to the oral microbiome | ||||
| 11,328 individuals in a prospective study. Dental health was monitored between 1971–1992 | Periodontitis increases the risk for pancreatic adenocarcinoma. | [81] | ||
| 10 resectable patients with pancreatic cancer and 10 matched healthy controls for oral microbiome assay, 28 resectable pancreatic cancer, 28 matched healthy controls, and 27 chronic pancreatitis samples for validation | HOMIM hybridization array | Streptococci, Veilonella, Actinobacteria, Campylobacter, and Prevotella increased in pancreatic adenocarcinoma patients. | Neisseria elongata and Streptococcus mitis were validated as biomarkers for pancreatic adenocarcinoma. | [63] |
| Pre-diagnosis blood samples from 405 pancreatic cancer cases and 416 matched controls, collected as part of the European Prospective Investigation into Cancer and Nutrition study | High serum antibodies against Porphyromonas gingivalis ATTC 53978 showed a two fold increase in risk for pancreatic adenocarcinoma. Those individuals who had high antibody titer against the commensal flora had a lower risk for pancreatic adenocarcinoma as compared to those with low titer. | [82] | ||
| 8 pancreatic adenocarcinoma patients and 22 healthy controls | 16S rDNA was amplified and sequenced | The pancreatic cancer group had higher levels of Leptotrichia, and lower levels of Porphyromonas, and Neisseria. No difference in diversity. Leptotrichia to Porphyromonas ratio was significantly higher in pancreatic adenocarcinoma patients. | [83] | |
| Among 149 orodigestive cancers 6 pancreatic adenocarcinoma cases | Treponema denticola chymotrypsin-like proteinase that can induce matrix metalloproteinases, was found in pancreatic adenocarcinoma using immunohistochemistry. | [84] | ||
| 361 incident pancreatic adenocarcinoma patients and 371 matched controls from two prospective cohort studies, the American Cancer Society Cancer Prevention Study II and the National Cancer Institute Prostate, Lung, Colorectal and Ovarian Cancer Screening Trial. | DNA was isolated from oral wash samples; 16S rRNA gene V3-V4 was amplified and sequenced using Roche 454 FLX Titanium Pyrosequencing system | Carriage of Porphyromonas gingivalis and Aggregatibacter actinomycetemcomitans were associated with a higher risk for pancreatic adenocarcinoma. Fusobacteria and its genus, Leptotrichia decreased the risk of pancreatic adenocarcinoma. The genus Alloprevotella increased the risk for pancreatic adenocarcinoma. | [85] | |
| 30 stage I pancreas head adenocarcinoma patients and 25 healthy controls | Tongue scrapes were collected and the V3-V4 16S rDNA was amplified and sequenced | Leptotrichia, Fusobacterium, Rothia, Actinomyces, Corynebacterium, Atopobium, Peptostreptococcus, Catonella, Oribacterium, Filifactor, Campylobacter, Moraxella, and Tannerella were overrepresented, while Haemophilus, Porphyromonas, and Paraprevotella were underrepresented in pancreatic adenocarcinoma patients. | Haemophilus, Porphyromonas, Leptotrichia, and Fusobacterium distinguished pancreatic adenocarcinoma patients from healthy subjects. | [86] |
| Saliva samples from 280 pancreatic adenocarcinoma cases (29 stage I, 160 stage II, 37 stage III, and 54 stage IV pancreatic tumors) of which 273 was used in the study and 285 controls | V4 region of the 16S rRNA gene was PCR amplified and sequencing was performed on the Illumina MiSeq. | Haemophilus genus showed a marginal association with pancreatic cancer risk. Enterobacteriaceae, Lachnospiraceae, Bacteroidaceae, and Staphylococcaceae showed a positive correlation with pancreatic cancer risk. | [88] | |
| Helicobacter pylori colonization (seropositivity) | ||||
| Cases with pancreatic cancer (n = 87) were matched to controls (n = 263) using age, sex and time for baseline investigation as matching variables | H.pylori serology was analyzed in stored serum samples using an enzyme-linked immunosorbent assay | H.pylori seropositivity was not associated with pancreatic cancer in the total cohort (adjusted OR 1.25 (0.75–2.09)). However, a statistically significant association was found in never smokers (OR 3.81 (1.06–13.63) adjusted for alcohol consumption) and a borderline statistically significant association was found in subjects with low alcohol consumption (OR 2.13 (0.97–4.69) adjusted for smoking). | [62] | |
| 110 patients with pancreatic cancer | A polypeptide antibody against the plasminogen-binding protein (PBP) of Helicobacter pylori and with the ubiquitin-protein ligase E3 component, n-recognin 2 (UBR2), an enzyme highly expressed in acinar cells of the pancreas | The antibody was positive in 5 of 110 patients with pancreatic cancer (5%). | [69] | |
| Venipuncture specimens were obtained from a representative sample of 761 case patients and 794 randomly selected control subjects matched by category of age and gender | Antibody seropositivity for H. pylori and its virulence protein CagA were determined by commercial enzyme-linked immunosorbent IgG assays | Compared with individuals seronegative for both H. pylori and CagA, decreased pancreas-cancer risk was seen for CagA seropositivity [adjusted OR, 0.68; 95% confidence interval (CI), 0.54–0.84], whereas some increased risk was suggested for CagA-negative H. pylori seropositivity (OR, 1.28; 95% CI, 0.76–2.13). | [70] | |
| Changes to the duodenal microbiome | ||||
| 14 patients with pancreatic head cancer and 14 healthy controls. Endoscopic duodenal mucosal biopsies | 16S rRNA gene pyrosequencing after the PCR amplification of the V3-V4 region. The rarefaction curves did not approach a plateau. | Acinetobacter, Aquabacterium, Oceanobacillus, Rahnella, Massilia, Delftia, Deinococcus, and Sphingobium were more abundant in the duodenal mucosa of pancreatic cancer patients, whereas the duodenal microbiomes of healthy controls were enriched with Porphyromonas, Paenibacillus, Enhydrobacter, Escherichia, Shigella, and Pseudomonas. Alpha and beta diversity were not different between the two groups. | Pancreatic adenocarcinoma patients have a higher incidence of H. pylori colonization. | [64] |
| Changes to the pancreatic microbiome | ||||
| 283 patients with pancreatic ductal adenocarcinoma (PDAC) | Genomic DNA extracted from FFPE tissue specimens assessed using TaqMan primer/probe sets to detect Fusobacterium species | 8.8% detection rate of Fusobacterium species in pancreatic cancers; however, tumor Fusobacterium status was not associated with any clinical and molecular features. In multivariate Cox regression analysis, compared with the Fusobacterium species-negative group, higher cancer-specific mortality rates were observed in the Fusobacterium positive group. | [89] | |
| Human FFPE pancreatic adenocarcinoma samples (n = 27) | Illumina sequencing of V1-V3 hypervariable regions of 16S RNA gene. | Differential presence of Acinetobacter, Afipia, Corynebacterium, Deltia, Enterobacter, Enterococcus, Escherichia, Klebsiella, Propionibacterium, Pseudomonas, Rastoria, Sphingomonas, Staphylococcus, and Streptococcus between healthy, pancreatitis, and pancreatic adenocarcinoma tissues. | In the pancreas, the microbiome could not discriminate between healthy, pancreatitis, and pancreatic adenocarcinoma states. Culturable bacteria are present in the human pancreas with a mean of ~1 × 105 (aerobic) and ~1 × 105 (anaerobic) cfu/g of tissue after 48 h of culture. | [65] |
| Pancreatic juice from pancreatic cancer (n = 20) and duodenal cancer/bile duct cancer (n = 16) patients | PCR identification of bacterial species by 16S ribosomal RNA gene. | Enterococcus faecalis may be involved in pancreas carcinogenesis. | E. faecalis is present in pancreas tissue in cancer patients. Antibodies against E. faecalis capsular polysaccharide is elevated in chronic pancreatitis patients. | [90] |
| Patients with pancreatic adenocarcinoma (n = 32), matched healthy individuals (n = 31). stool and pancreas tissue were assessed | Sequencing of the V3-V4 hypervariable region of the 16S RNA gene after PCR amplification | Proteobacteria are more abundant in patients with pancreatic adenocarcinoma as compared to gut cancer patients. | [41] | |
| 105 subjects were enrolled of which 27 had pancreatic adenocarcinoma, 57 had intraductal papillary mucinous neoplasms and 21 had benign lesions | Pancreas cyst fluid was collected, total bacterial 16S copy number was assessed, and 16S DNA was sequenced | Fusobacterium nucleatum and Granulicatella adiacens were associated with high-grade dysplasia. In network analysis, the network’s nodes were Actinobacteria (Cutibacterium acnes), Bacteroidetes, Firmicutes (Streptococcus anginous, Granulicatella adiacens), and Proteobacteria (Klebsiella aerogenes). | The number of 16S reads increases in precancerous and cancer cases. | [87] |
| Human fecal samples and specimens of pancreatic tissue were collected under sterile conditions from healthy volunteers and patients undergoing surgery for PDA or pancreatic endocrine tumors (benign disease) | PCR amplification and sequencing of the ITS1 region of the 18S rRNA gene using Illumina sequencing | Pancreatic adenocarcinoma tumors with fungal infiltration were enriched for Malassezia spp. | Ligation of mannose-binding lectin (MBL), which binds to glycans of the fungal wall to activate the complement cascade, was required for oncogenic progression. | [68] |
| Long-term surviving (n = 22) and short-term surviving (n = 21) pancreatic adenocarcinoma patients. | From the tumor and feces 16S rDNA V4 region was amplified by PCR and sequenced in the MiSeq platform (Illumina). | Intratumoral microbiome signature (Pseudoxanthomonas-Streptomyces-Saccharopolyspora-Bacillus clausii) is highly predictive of long-term survivorship. | The microbiome that provides long-term survival can be transplanted. | [91] |
| 50 patients with pancreatic adenocarcinoma were enrolled. In cases where a biliary stent was inserted prior to surgery, the stent was removed and cultured. In other cases, swabs of bile or pancreatic fluid and tissue from the bile duct or pancreas were obtained and cultured. | Classical culture | 96% of the specimens demonstrated the presence of microbes, 90% of all cases were polymicrobial. The most frequent species found were Enterobacteriaceae, Enterococcus species, Candida species, and Streptococcus milleri | [92] | |
| 152 Italian patients of which 72 had pancreas head adenocarcinoma patients were present | Classical culture | The most common bacteria among pancreas head adenocarcinoma patients were E. coli, K. pneumoniae, and P. aeruginosa, and less frequently, Alcaligenes spp., Serratia spp., and Enterococcus spp. | Although pancreas head carcinoma patients were not assessed separately, only such patients were present in the shortest survival cohort enabling the assessment of that patient population. E. coli, K. pneumoniae, and P. aeruginosa showed a high percentage of resistance to third-generation cephalosporins (3GCs), aminoglycosides class, and quinolone group, especially to levofloxacin, but the same bacteria were sensitive to carbapenems. | [93] |
| 50 patients with pancreatic adenocarcinoma, 34 other organs (i.e., controls). In total, 189 tissue samples (pancreatic duct, duodenum, pancreas), 57 swabs (bile duct, jejunum, stomach), and 12 stool samples. | The 16S rRNA V3–V4 hypervariable regions were amplified using Illumina MiSeq | Lactobacillus ssp. was significantly higher in noncancer subjects compared with cancer subjects and the relative abundance of Fusobacterium spp was higher in cancer subjects compared with noncancer subjects. | [94] | |
| Changes to stool microbiome | ||||
| Prospective study, 85 pancreatic cancer (PC) and 57 matched healthy controls (HC) | MiSeq sequencing | Phylum Bacteroidetes was significantly increased, while Firmicutes and Proteobacteria were decreased in PC patients versus healthy controls. | Gut microbial diversity decreased in pancreatic adenocarcinoma. Alpha diversity decreased. The abundance of certain pathogens and lipopolysaccharides-producing bacteria increased. Probiotics and butyrate-producing bacteria decreased. Changes to the microbiome can be used as markers to detect pancreatic adenocarcinoma and the obstructive and non-obstructive forms. | [43] |
| Patients with pancreatic adenocarcinoma (n = 32), and matched healthy individuals (n = 31). stool and pancreatic tissue were assessed | Sequencing of the V3-V4 hypervariable region of the 16S RNA gene after PCR amplification | Proteobacteria are more abundant in patients with pancreatic adenocarcinoma as compared to healthy controls. | [41] | |
| Long-term surviving (n = 22) and short-term surviving (n = 21) patients. Sequencing of intratumor and stool microbiomes. | 16S rDNA V4 region was amplified by PCR and sequenced in the MiSeq platform (Illumina). | Intra-tumoral microbiome signature occurs in pancreatic adenocarcinoma patients (Pseudoxanthomonas-Streptomyces-Saccharopolyspora-Bacillus clausii) that is highly predictive of long-term survivorship. | The microbiome that provides long-term survival can be transplanted. | [91] |
| 30 patients with pancreatic adenocarcinoma, 6 patients with pre-cancerous lesions, 13 healthy subjects, and 16 with non-alcoholic fatty liver disease | 16S RNA was PCR amplified and was sequenced using the Illumina MiSeq platform and LEfSe linear discriminant analysis (LDA) was performed | Patterns of the microbiome can separate pancreatic adenocarcinoma patients from healthy subjects and patients with comorbidities (NAFLD, etc.) and can discriminate between the etiology of pancreatic adenocarcinoma. | [100] | |
| Parent Metabolite | Bioactive Metabolite | Genus | Species | Reference |
|---|---|---|---|---|
| complex carbohydrates simple carbohydrates mucin | acetate | Lactobacillus | [108] | |
| Bifidobacterium | [108] | |||
| Bacteroides | [108] | |||
| Prevotella | [108] | |||
| Ruminococcus | [108] | |||
| Streptococcus | [108] | |||
| propionate | Lachnospiraceae | [119] | ||
| Bacteroidetes | [119] | |||
| Negativicutes | [119] | |||
| Phascolarctobacterium | [120,121] | |||
| Dialister | [120,121] | |||
| Veillonella | [120,121] | |||
| Salmonella | [120,121] | |||
| Megasphaera elsdenii | [120,121] | |||
| Coprococcus catus | [120,121] | |||
| Akkermansia muciniphila | [118] | |||
| Ruminococcus obeum | [119] | |||
| Roseburia inulinivorans | [119] | |||
| butyrate | Odoribacter | [120,121,122,123] | ||
| Anaeotruncus | [120,121,122,123] | |||
| Faecalibacterium prausnitzii | [120,121,122,123] | |||
| Eubacterium rectale | [120,121,122,123] | |||
| Roseburia faecis | [120,121,122,123] | |||
| Clostridium leptum | [120,121,122,123] | |||
| Coprococcus eutactus | [120,121,122,123] | |||
| Faecalibacterium prausnitzii | [120,121,122,123] | |||
| Eubacterium rectale | [120,121,122,123] | |||
| Anaerostipes caccae | [120,121,122,123] | |||
| Eubacterium hallii | [120,121,122,123] | |||
| unnamed cultured species SS2/1 | [120,121,122,123] | |||
| primary bile acids | secondary bile acids | Clostridium | [124,125,126,127,128,129] | |
| Lactobacillus | [124,125,126,127,128,129] | |||
| Bifidobacterium | [124,125,126,127,128,129] | |||
| Eubacterium | [124,125,126,128] | |||
| Ruminococcus | [124,125,126,128] | |||
| Escherichia | [124,125,126,128] | |||
| Bacteroides fragilis | [124,125,126,127,128,129] | |||
| Bacteroides vulgatus | [124,125,126,127,128,129] | |||
| Listeria monocytogenes | [124,125,126,127,128,129] | |||
| polyamines | E. coli | [108,130,131] | ||
| Enterococcus faecalis | [108,130,131] | |||
| Staphylococcus aureus | [108,130,131] | |||
| Haemophilus influenzae | [108,130,131] | |||
| Neisseria flava | [108,130,131] | |||
| Pseudomonas aeruginosa | [108,130,131] | |||
| Campylobacter jejuni | [108,130,131] | |||
| Yersinia pestis | [108,130,131] | |||
| Vibrio cholerae | [108,130,131] | |||
| Bacteroides dorei | [108,130,131] | |||
| Bacteroides thetaiotaomicron | [108,130,131] | |||
| Bacteroides fragilis | [108,130,131] | |||
| Bacillus subtilis | [108,130,131] | |||
| Proteus mirabilis | [108,130,131] | |||
| lysine | cadaverine | Streptococcus | [132] | |
| Shigella flexneri | [132] | |||
| Shigella sonnei | [132] | |||
| Escherichia coli | [132] | |||
| tryptophan | tryptamine | Clostridium sporogenes | [133] | |
| Ruminococcus gnavus | [133] | |||
| indole acetic acid | Lactobacillus | [133] | ||
| Clostridium | [133] | |||
| Bacteroides | [133] |
| Study | Direction of Regulation in Patients | Biological Process | Biochemical Process |
|---|---|---|---|
| [67] | upregulated | Nucleotide biosynthesis | |
| Lipid biosynthesis | |||
| Polyamine biosynthesis | |||
| Hexitol fermentation | |||
| Carbohydrate metabolism | |||
| Vitamin biosynthesis and metabolism | |||
| [43] | downregulated | transport systems | Phosphate transport system (M00222) |
| Cobalt transport system (M00245) | |||
| Mannopine transport system (M00301) | |||
| Glutamate transport system (M00233) | |||
| Trehalose-maltose transport system (M00204) | |||
| Spermidine-putrescine transport system (M00299) | |||
| amino acid metabolism | Histidine biosynthesis (M00026) | ||
| Glutamate transport system (M00233) | |||
| metabolic pathways | Complex I/NADH dehydrogenase (M00144) | ||
| Pentose phosphate pathway/non-oxidative phase (M00007) | |||
| V type ATPase (M00159) | |||
| Pyrimidine deoxyribonucleotide biosynthesis (M00053) | |||
| Pyruvate ferredoxin oxidoreductase (M00310) | |||
| upregulated | amino acid metabolism | Leucine biosynthesis (M00019) | |
| Twin arginine translocation/Tat system (M00336) | |||
| Histidine degradation (M00045) | |||
| Methionine salvage pathway (M00034) | |||
| Lysine arginine ornithine transport system (M00225) | |||
| Dipeptide transport system (M00324) | |||
| Arginine transport system (M00229) | |||
| Histidine transport system (M00226) | |||
| carbohydrate metabolism | Oligogalacturonide transport system (M00202) | ||
| Entner Doudoroff pathway (M00008) | |||
| transport systems | Putative spermidine putrescine transport system (M00193) | ||
| Microcin C transport system (M00349) | |||
| Putrescine transport system (M00300) | |||
| Sec secretion system (M00335) | |||
| Histidine transport system (M00226) | |||
| metabolic pathways | Pyridoxal biosynthesis (M00124), Citrate cycle (M00011) | ||
| Complex II/succinate dehydrogenase (M00150) | |||
| Glyoxylate cycle (M00012) | |||
| C5 isoprenoid biosynthesis/non-mevalonate pathway (M00096) | |||
| Ubiquinone biosynthesis (M00117) | |||
| Prokaryotic GABA biosynthesis (M00136) | |||
| Lipopolysaccharide biosynthesis (M00060) | |||
| Bacterial DNA polymerase III complex (M00260) | |||
| polyamine biosynthesis and transport | Polyamine biosynthesis (M00133) | ||
| Spermidine putrescine transport system (M00299) |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Kiss, B.; Mikó, E.; Sebő, É.; Toth, J.; Ujlaki, G.; Szabó, J.; Uray, K.; Bai, P.; Árkosy, P. Oncobiosis and Microbial Metabolite Signaling in Pancreatic Adenocarcinoma. Cancers 2020, 12, 1068. https://doi.org/10.3390/cancers12051068
Kiss B, Mikó E, Sebő É, Toth J, Ujlaki G, Szabó J, Uray K, Bai P, Árkosy P. Oncobiosis and Microbial Metabolite Signaling in Pancreatic Adenocarcinoma. Cancers. 2020; 12(5):1068. https://doi.org/10.3390/cancers12051068
Chicago/Turabian StyleKiss, Borbála, Edit Mikó, Éva Sebő, Judit Toth, Gyula Ujlaki, Judit Szabó, Karen Uray, Péter Bai, and Péter Árkosy. 2020. "Oncobiosis and Microbial Metabolite Signaling in Pancreatic Adenocarcinoma" Cancers 12, no. 5: 1068. https://doi.org/10.3390/cancers12051068
APA StyleKiss, B., Mikó, E., Sebő, É., Toth, J., Ujlaki, G., Szabó, J., Uray, K., Bai, P., & Árkosy, P. (2020). Oncobiosis and Microbial Metabolite Signaling in Pancreatic Adenocarcinoma. Cancers, 12(5), 1068. https://doi.org/10.3390/cancers12051068







