Identifying Genetic Lesions in Ocular Adnexal Extranodal Marginal Zone Lymphomas of the MALT Subtype by Whole Genome, Whole Exome and Targeted Sequencing
Abstract
1. Introduction
2. Results
2.1. Whole Exome, Whole Genome, and Targeted Sequencing of OAML
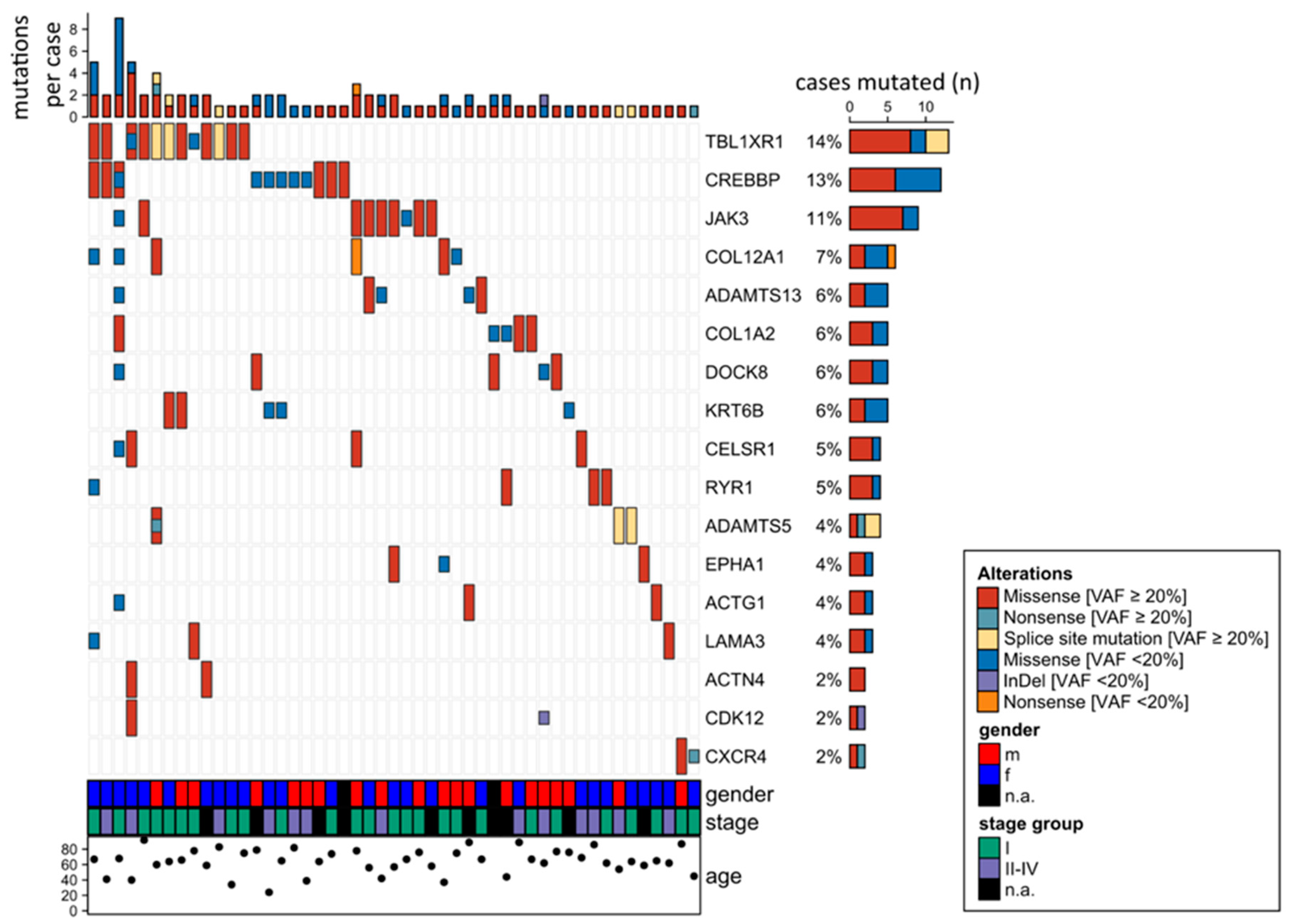
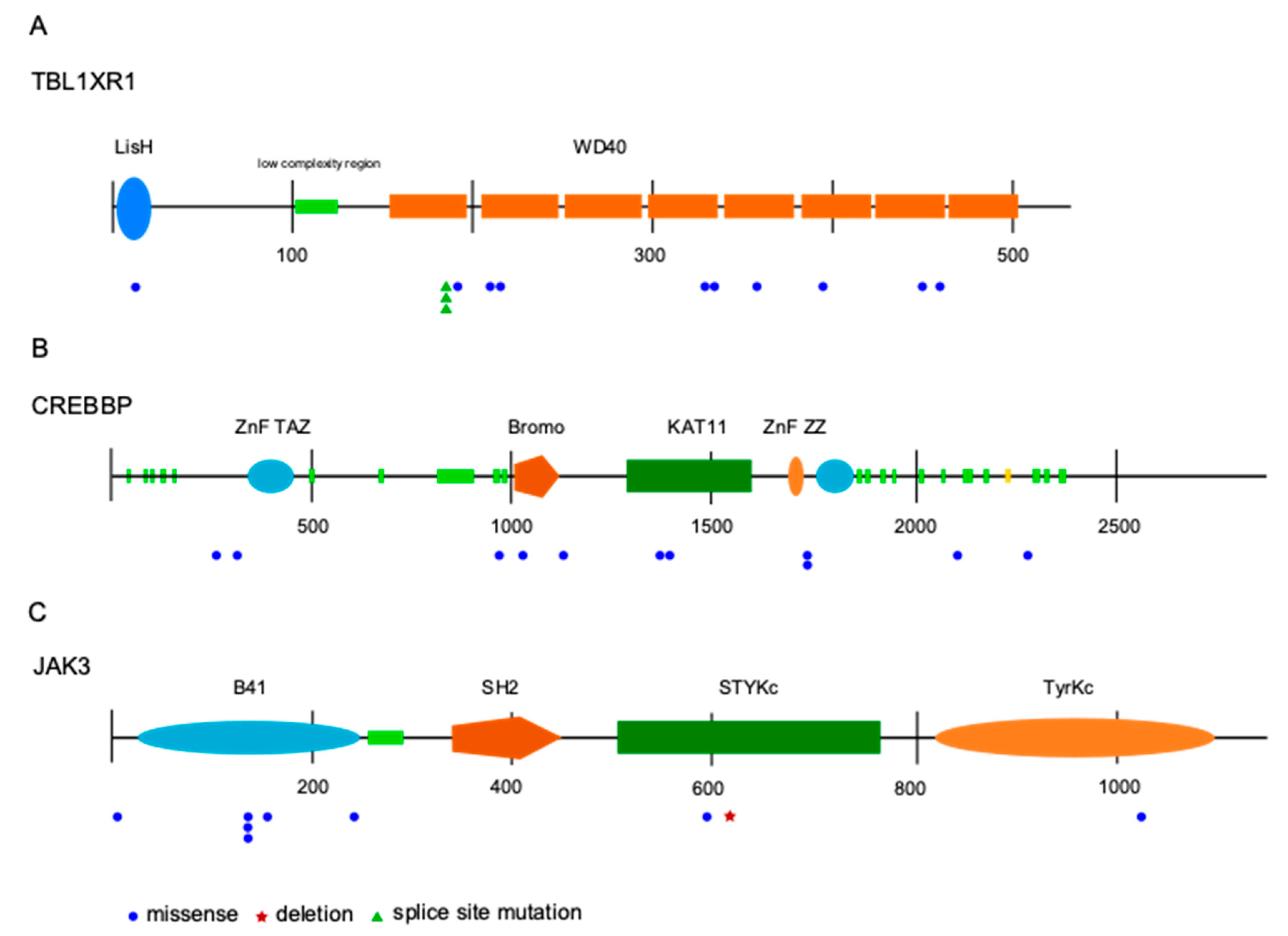
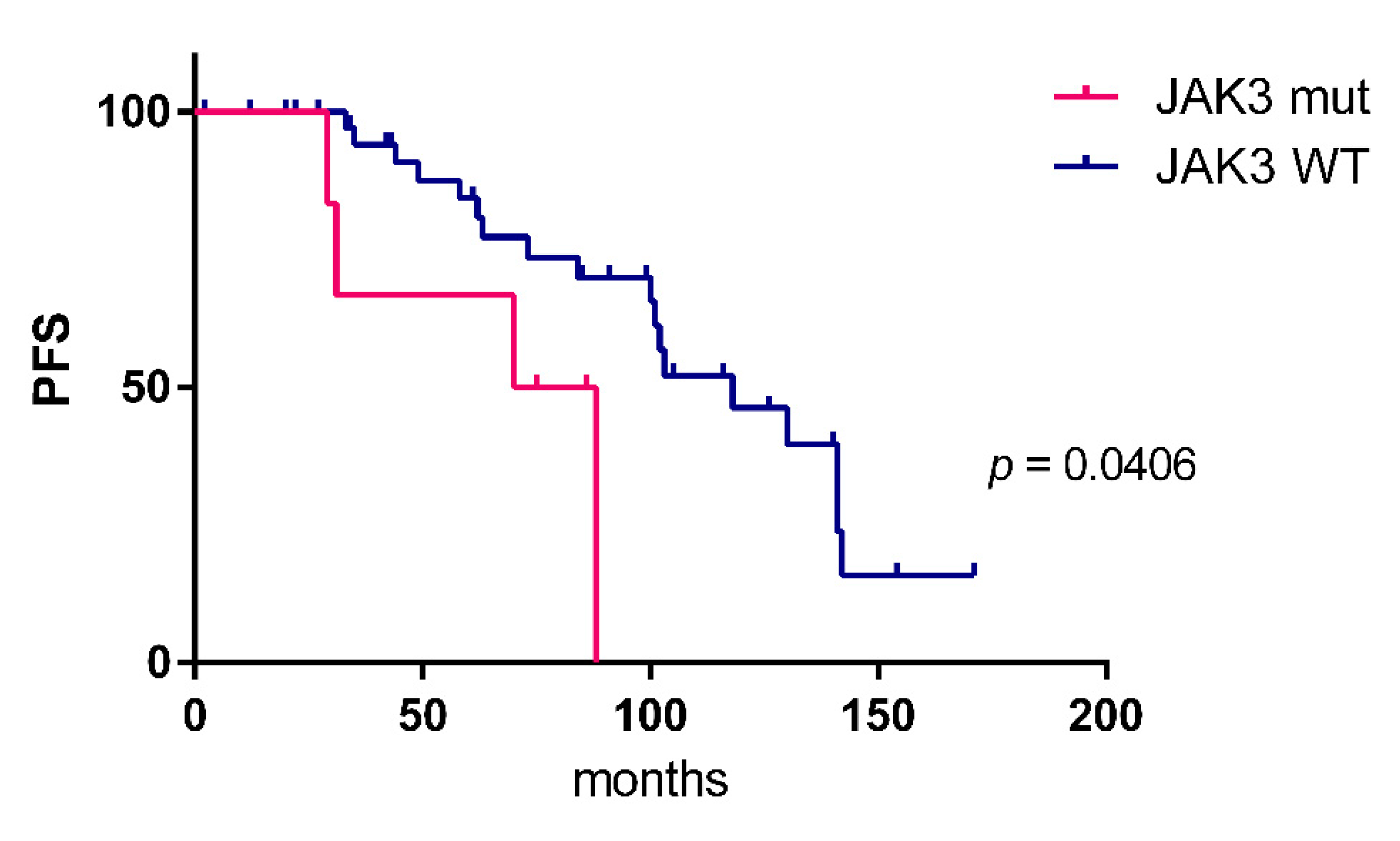
2.2. Detection of Numerous Recurrently Mutated Genes in OAML
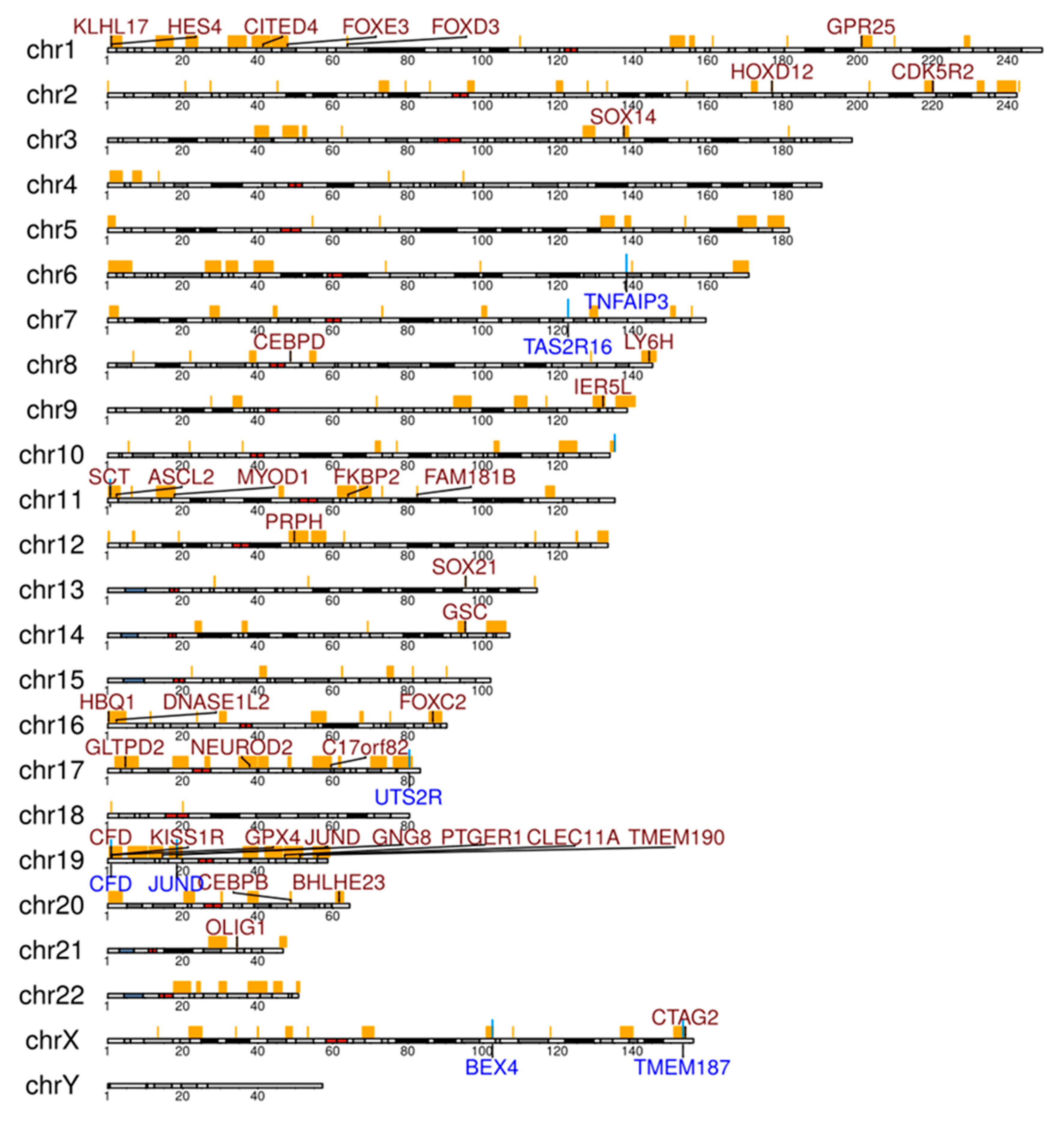
2.3. Analysis of WGS Data for CNVs, Chromosomal Translocations, and Viral Sequences
2.4. Analysis for Bacterial Sequences in the OAML Tissues
3. Discussion
4. Materials and Methods
4.1. Patients and Samples
4.2. Tumor and Non-Tumor Cell Enrichment
4.3. DNA Isolation
4.4. Whole Genome Sequencing
4.5. Gene Copy Number Analysis
4.6. Search for Viral Sequences
4.7. Translocations Analysis
4.8. Whole Exome Sequencing
4.9. Determination of the Coding Sequence of Bacterial 16S Ribosomal RNA
4.10. Targeted Re-Sequencing
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Freeman, C.; Berg, J.W.; Cutler, S.J. Occurrence and prognosis of extranodal lymphomas. Cancer 1972, 29, 252–260. [Google Scholar] [CrossRef]
- Coupland, S.E.; Krause, L.; Delecluse, H.J.; Anagnostopoulos, I.; Foss, H.D.; Hummel, M.; Bornfeld, N.; Lee, W.R.; Stein, H. Lymphoproliferative lesions of the ocular adnexa. Analysis of 112 cases. Ophthalmology 1998, 105, 1430–1441. [Google Scholar] [CrossRef]
- Ferry, J.A.; Fung, C.Y.; Zukerberg, L.; Lucarelli, M.J.; Hasserjian, R.P.; Preffer, F.I.; Harris, N.L. Lymphoma of the ocular adnexa: A study of 353 cases. Am. J. Surg. Pathol. 2007, 31, 170–184. [Google Scholar] [CrossRef]
- Knowles, D.M.; Jakobiec, F.A.; McNally, L.; Burke, J.S. Lymphoid hyperplasia and malignant lymphoma occurring in the ocular adnexa (orbit, conjunctiva, and eyelids): A prospective multiparametric analysis of 108 cases during 1977 to 1987. Hum. Pathol. 1990, 21, 959–973. [Google Scholar] [CrossRef]
- Hara, Y.; Nakamura, N.; Kuze, T.; Hashimoto, Y.; Sasaki, Y.; Shirakawa, A.; Furuta, M.; Yago, K.; Kato, K.; Abe, M. Immunoglobulin heavy chain gene analysis of ocular adnexal extranodal marginal zone B-cell lymphoma. Investig. Ophthalmol. Vis. Sci. 2001, 42, 2450–2457. [Google Scholar]
- McKelvie, P.A. Ocular adnexal lymphomas: A review. Adv. Anat. Pathol. 2010, 17, 251–261. [Google Scholar] [CrossRef] [PubMed]
- Chanudet, E.; Zhou, Y.; Bacon, C.M.; Wotherspoon, A.C.; Muller-Hermelink, H.K.; Adam, P.; Dong, H.Y.; de Jong, D.; Li, Y.; Wei, R.; et al. Chlamydia psittaci is variably associated with ocular adnexal MALT lymphoma in different geographical regions. J. Pathol. 2006, 209, 344–351. [Google Scholar] [CrossRef] [PubMed]
- Ferreri, A.J.; Guidoboni, M.; Ponzoni, M.; De Conciliis, C.; Dell’Oro, S.; Fleischhauer, K.; Caggiari, L.; Lettini, A.A.; Dal Cin, E.; Ieri, R.; et al. Evidence for an association between Chlamydia psittaci and ocular adnexal lymphomas. J. Natl. Cancer Inst. 2004, 96, 586–594. [Google Scholar] [CrossRef]
- Du, M.Q. MALT lymphoma: Recent advances in aetiology and molecular genetics. J. Clin. Exp. Hematop. 2007, 47, 31–42. [Google Scholar] [CrossRef] [PubMed]
- Du, M.Q. MALT lymphoma: Many roads lead to nuclear factor-kappab activation. Histopathology 2011, 58, 26–38. [Google Scholar] [CrossRef] [PubMed]
- Streubel, B.; Vinatzer, U.; Lamprecht, A.; Raderer, M.; Chott, A. T(3;14)(p14.1;q32) involving IGH and FOXP1 is a novel recurrent chromosomal aberration in MALT lymphoma. Leukemia 2005, 19, 652–658. [Google Scholar] [CrossRef] [PubMed]
- Zhu, D.; Ikpatt, O.F.; Dubovy, S.R.; Lossos, C.; Natkunam, Y.; Chapman-Fredricks, J.R.; Fan, Y.S.; Lossos, I.S. Molecular and genomic aberrations in Chlamydophila psittaci negative ocular adnexal marginal zone lymphomas. Am. J. Hematol. 2013, 88, 730–735. [Google Scholar] [CrossRef] [PubMed]
- Du, M.Q. MALT lymphoma: A paradigm of NF-kappaB dysregulation. Semin. Cancer Biol. 2016, 39, 49–60. [Google Scholar] [CrossRef] [PubMed]
- Yan, Q.; Wang, M.; Moody, S.; Xue, X.; Huang, Y.; Bi, Y.; Du, M.Q. Distinct involvement of NF-kappaB regulators by somatic mutation in ocular adnexal malt lymphoma. Br. J. Haematol. 2013, 160, 851–854. [Google Scholar] [CrossRef] [PubMed]
- Zhu, J.; Wei, R.L.; Pi, Y.L.; Guo, Q. Significance of Bcl10 gene mutations in the clinical diagnosis of MALT-type ocular adnexal lymphoma in the Chinese population. Genet. Mol. Res. 2013, 12, 1194–1204. [Google Scholar] [CrossRef] [PubMed]
- Johansson, P.; Klein-Hitpass, L.; Grabellus, F.; Arnold, G.; Klapper, W.; Pfortner, R.; Duhrsen, U.; Eckstein, A.; Durig, J.; Kuppers, R. Recurrent mutations in NF-kappaB pathway components, KMT2D, and NOTCH1/2 in ocular adnexal MALT-type marginal zone lymphomas. Oncotarget 2016, 7, 62627–62639. [Google Scholar] [CrossRef]
- Moody, S.; Thompson, J.S.; Chuang, S.S.; Liu, H.; Raderer, M.; Vassiliou, G.; Wlodarska, I.; Wu, F.; Cogliatti, S.; Robson, A.; et al. Novel GPR34 and CCR6 mutation and distinct genetic profiles in MALT lymphomas of different sites. Haematologica 2018, 103, 1329–1336. [Google Scholar] [CrossRef]
- Cascione, L.; Rinaldi, A.; Bruscaggin, A.; Tarantelli, C.; Arribas, A.J.; Kwee, I.; Pecciarini, L.; Mensah, A.A.; Spina, V.; Chung, E.Y.L.; et al. Novel insights into the genetics and epigenetics of MALT lymphoma unveiled by next generation sequencing analyses. Haematologica 2019, 104, e558–e561. [Google Scholar] [CrossRef]
- Jung, H.; Yoo, H.Y.; Lee, S.H.; Shin, S.; Kim, S.C.; Lee, S.; Joung, J.G.; Nam, J.Y.; Ryu, D.; Yun, J.W.; et al. The mutational landscape of ocular marginal zone lymphoma identifies frequent alterations in TNFAIP3 followed by mutations in TBL1XR1 and CREBBP. Oncotarget 2017, 8, 17038–17049. [Google Scholar] [CrossRef]
- Yoon, H.G.; Chan, D.W.; Huang, Z.Q.; Li, J.; Fondell, J.D.; Qin, J.; Wong, J. Purification and functional characterization of the human N-CoR complex: The roles of HDAC3, TBL1 and TBLR1. EMBO J. 2003, 22, 1336–1346. [Google Scholar] [CrossRef]
- De Decker, M.; Lavaert, M.; Roels, J.; Tilleman, L.; Vandekerckhove, B.; Leclercq, G.; Van Nieuwerburgh, F.; Van Vlierberghe, P.; Taghon, T. HES1 and HES4 have non-redundant roles downstream of Notch during early human T cell development. Haematologica 2020. [Google Scholar] [CrossRef] [PubMed]
- Heckman, C.A.; Wheeler, M.A.; Boxer, L.M. Regulation of Bcl-2 expression by C/EBP in t(14;18) lymphoma cells. Oncogene 2003, 22, 7891–7899. [Google Scholar] [CrossRef] [PubMed]
- Pal, R.; Janz, M.; Galson, D.L.; Gries, M.; Li, S.; Johrens, K.; Anagnostopoulos, I.; Dorken, B.; Mapara, M.Y.; Borghesi, L.; et al. C/EBPbeta regulates transcription factors critical for proliferation and survival of multiple myeloma cells. Blood 2009, 114, 3890–3898. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Vegesna, V.; Takeuchi, S.; Hofmann, W.K.; Ikezoe, T.; Tavor, S.; Krug, U.; Fermin, A.C.; Heaney, A.; Miller, C.W.; Koeffler, H.P. C/EBP-beta, C/EBP-delta, PU.1, AML1 genes: Mutational analysis in 381 samples of hematopoietic and solid malignancies. Leuk. Res. 2002, 26, 451–457. [Google Scholar] [CrossRef]
- Bi, Y.; Zeng, N.; Chanudet, E.; Huang, Y.; Hamoudi, R.A.; Liu, H.; Dong, G.; Watkins, A.J.; Ley, S.C.; Zou, L.; et al. A20 inactivation in ocular adnexal MALT lymphoma. Haematologica 2012, 97, 926–930. [Google Scholar] [CrossRef]
- Chanudet, E.; Ye, H.; Ferry, J.; Bacon, C.M.; Adam, P.; Muller-Hermelink, H.K.; Radford, J.; Pileri, S.A.; Ichimura, K.; Collins, V.P.; et al. A20 deletion is associated with copy number gain at the TNFA/B/C locus and occurs preferentially in translocation-negative MALT lymphoma of the ocular adnexa and salivary glands. J. Pathol. 2009, 217, 420–430. [Google Scholar] [CrossRef]
- Honma, K.; Tsuzuki, S.; Nakagawa, M.; Karnan, S.; Aizawa, Y.; Kim, W.S.; Kim, Y.D.; Ko, Y.H.; Seto, M. TNFAIP3 is the target gene of chromosome band 6q23.3-q24.1 loss in ocular adnexal marginal zone B cell lymphoma. Genes Chromosomes Cancer 2008, 47, 1–7. [Google Scholar] [CrossRef]
- Choung, H.; Kim, Y.A.; Kim, N.; Lee, M.J.; Khwarg, S.I. Clinicopathologic Study of Chromosomal Aberrations in Ocular Adnexal Lymphomas of Korean Patients. Korean J. Ophthalmol. 2015, 29, 285–293. [Google Scholar] [CrossRef][Green Version]
- Ferreri, A.J.; Dolcetti, R.; Du, M.Q.; Doglioni, C.; Resti, A.G.; Politi, L.S.; De Conciliis, C.; Radford, J.; Bertoni, F.; Zucca, E.; et al. Ocular adnexal MALT lymphoma: An intriguing model for antigen-driven lymphomagenesis and microbial-targeted therapy. Ann. Oncol. 2008, 19, 835–846. [Google Scholar] [CrossRef]
- Kim, W.S.; Honma, K.; Karnan, S.; Tagawa, H.; Kim, Y.D.; Oh, Y.L.; Seto, M.; Ko, Y.H. Genome-wide array-based comparative genomic hybridization of ocular marginal zone B cell lymphoma: Comparison with pulmonary and nodal marginal zone B cell lymphoma. Genes Chromosomes Cancer 2007, 46, 776–783. [Google Scholar] [CrossRef]
- Liu, F.; He, Y.; Cao, Q.; Liu, N.; Zhang, W. TBL1XR1 Is Highly Expressed in Gastric Cancer and Predicts Poor Prognosis. Dis. Markers 2016, 2016, 2436518. [Google Scholar] [CrossRef] [PubMed]
- Zhang, T.; Liu, C.; Yu, Y.; Geng, J.; Meng, Q.; Xu, S.; Zhou, F.; Chen, Y.; Jin, S.; Shen, J.; et al. TBL1XR1 is involved in c-Met-mediated tumorigenesis of human nonsmall cell lung cancer. Cancer Gene Ther. 2019. [Google Scholar] [CrossRef]
- Pasqualucci, L.; Dominguez-Sola, D.; Chiarenza, A.; Fabbri, G.; Grunn, A.; Trifonov, V.; Kasper, L.H.; Lerach, S.; Tang, H.; Ma, J.; et al. Inactivating mutations of acetyltransferase genes in B-cell lymphoma. Nature 2011, 471, 189–195. [Google Scholar] [CrossRef]
- Degryse, S.; de Bock, C.E.; Cox, L.; Demeyer, S.; Gielen, O.; Mentens, N.; Jacobs, K.; Geerdens, E.; Gianfelici, V.; Hulselmans, G.; et al. JAK3 mutants transform hematopoietic cells through JAK1 activation, causing T-cell acute lymphoblastic leukemia in a mouse model. Blood 2014, 124, 3092–3100. [Google Scholar] [CrossRef] [PubMed]
- Elliott, N.E.; Cleveland, S.M.; Grann, V.; Janik, J.; Waldmann, T.A.; Dave, U.P. FERM domain mutations induce gain of function in JAK3 in adult T-cell leukemia/lymphoma. Blood 2011, 118, 3911–3921. [Google Scholar] [CrossRef] [PubMed]
- Lupardus, P.J.; Ultsch, M.; Wallweber, H.; Bir Kohli, P.; Johnson, A.R.; Eigenbrot, C. Structure of the pseudokinase-kinase domains from protein kinase TYK2 reveals a mechanism for Janus kinase (JAK) autoinhibition. Proc. Natl. Acad. Sci. USA 2014, 111, 8025–8030. [Google Scholar] [CrossRef] [PubMed]
- Hasegawa, M.; Kojima, M.; Shioya, M.; Tamaki, Y.; Saitoh, J.; Sakurai, H.; Kitamoto, Y.; Suzuki, Y.; Niibe, H.; Nakano, T. Treatment results of radiotherapy for malignant lymphoma of the orbit and histopathologic review according to the WHO classification. Int. J. Radiat. Oncol. Biol. Phys. 2003, 57, 172–176. [Google Scholar] [CrossRef]
- Wenzel, C.; Fiebiger, W.; Dieckmann, K.; Formanek, M.; Chott, A.; Raderer, M. Extranodal marginal zone B-cell lymphoma of mucosa-associated lymphoid tissue of the head and neck area: High rate of disease recurrence following local therapy. Cancer 2003, 97, 2236–2241. [Google Scholar] [CrossRef]
- Derenzini, E.; Younes, A. Targeting the JAK-STAT pathway in lymphoma: A focus on pacritinib. Expert Opin. Investig. Drugs 2013, 22, 775–785. [Google Scholar] [CrossRef]
- Liu, J.; Li, H.; Shen, S.; Sun, L.; Yuan, Y.; Xing, C. Alternative splicing events implicated in carcinogenesis and prognosis of colorectal cancer. J. Cancer 2018, 9, 1754–1764. [Google Scholar] [CrossRef]
- Qiu, Z.; Lin, A.; Li, K.; Lin, W.; Wang, Q.; Wei, T.; Zhu, W.; Luo, P.; Zhang, J. A novel mutation panel for predicting etoposide resistance in small-cell lung cancer. Drug Des. Devel. Ther. 2019, 13, 2021–2041. [Google Scholar] [CrossRef] [PubMed]
- Liu, K.; Ren, Y.; Pang, L.; Qi, Y.; Jia, W.; Tao, L.; Hu, Z.; Zhao, J.; Zhang, H.; Li, L.; et al. Papillary renal cell carcinoma: A clinicopathological and whole-genome exon sequencing study. Int. J. Clin. Exp. Pathol. 2015, 8, 8311–8335. [Google Scholar] [PubMed]
- Sung, C.O.; Kim, S.C.; Karnan, S.; Karube, K.; Shin, H.J.; Nam, D.H.; Suh, Y.L.; Kim, S.H.; Kim, J.Y.; Kim, S.J.; et al. Genomic profiling combined with gene expression profiling in primary central nervous system lymphoma. Blood 2011, 117, 1291–1300. [Google Scholar] [CrossRef] [PubMed]
- Anderton, J.A.; Lindsey, J.C.; Lusher, M.E.; Gilbertson, R.J.; Bailey, S.; Ellison, D.W.; Clifford, S.C. Global analysis of the medulloblastoma epigenome identifies disease-subgroup-specific inactivation of COL1A2. Neuro Oncol. 2008, 10, 981–994. [Google Scholar] [CrossRef]
- Kalmar, A.; Peterfia, B.; Hollosi, P.; Galamb, O.; Spisak, S.; Wichmann, B.; Bodor, A.; Toth, K.; Patai, A.V.; Valcz, G.; et al. DNA hypermethylation and decreased mRNA expression of MAL, PRIMA1, PTGDR and SFRP1 in colorectal adenoma and cancer. BMC Cancer 2015, 15, 736. [Google Scholar] [CrossRef]
- Randall, K.L.; Lambe, T.; Johnson, A.L.; Treanor, B.; Kucharska, E.; Domaschenz, H.; Whittle, B.; Tze, L.E.; Enders, A.; Crockford, T.L.; et al. Dock8 mutations cripple B cell immunological synapses, germinal centers and long-lived antibody production. Nat. Immunol. 2009, 10, 1283–1291. [Google Scholar] [CrossRef]
- Schramm, A.; Koster, J.; Assenov, Y.; Althoff, K.; Peifer, M.; Mahlow, E.; Odersky, A.; Beisser, D.; Ernst, C.; Henssen, A.G.; et al. Mutational dynamics between primary and relapse neuroblastomas. Nat. Genet. 2015, 47, 872–877. [Google Scholar] [CrossRef]
- Swerdlow, S.H.; Campo, E.; Pileri, S.A.; Harris, N.L.; Stein, H.; Siebert, R.; Advani, R.; Ghielmini, M.; Salles, G.A.; Zelenetz, A.D.; et al. The 2016 revision of the World Health Organization classification of lymphoid neoplasms. Blood 2016, 127, 2375–2390. [Google Scholar] [CrossRef]
- Li, H.; Durbin, R. Fast and accurate short read alignment with Burrows-Wheeler transform. Bioinformatics 2009, 25, 1754–1760. [Google Scholar] [CrossRef]
- Rimmer, A.; Phan, H.; Mathieson, I.; Iqbal, Z.; Twigg, S.R.; Consortium, W.G.S.; Wilkie, A.O.; McVean, G.; Lunter, G. Integrating mapping-, assembly- and haplotype-based approaches for calling variants in clinical sequencing applications. Nat. Genet. 2014, 46, 912–918. [Google Scholar] [CrossRef]
- Cingolani, P.; Platts, A.; Wang le, L.; Coon, M.; Nguyen, T.; Wang, L.; Land, S.J.; Lu, X.; Ruden, D.M. A program for annotating and predicting the effects of single nucleotide polymorphisms, SnpEff: SNPs in the genome of Drosophila melanogaster strain w1118; iso-2; iso-3. Fly 2012, 6, 80–92. [Google Scholar] [CrossRef] [PubMed]
- Kamburov, A.; Stelzl, U.; Lehrach, H.; Herwig, R. The ConsensusPathDB interaction database: 2013 update. Nucleic Acids Res. 2013, 41, D793–D800. [Google Scholar] [CrossRef] [PubMed]
- McKenna, A.; Hanna, M.; Banks, E.; Sivachenko, A.; Cibulskis, K.; Kernytsky, A.; Garimella, K.; Altshuler, D.; Gabriel, S.; Daly, M.; et al. The Genome Analysis Toolkit: A MapReduce framework for analyzing next-generation DNA sequencing data. Genome Res. 2010, 20, 1297–1303. [Google Scholar] [CrossRef] [PubMed]
- Bankevich, A.; Nurk, S.; Antipov, D.; Gurevich, A.A.; Dvorkin, M.; Kulikov, A.S.; Lesin, V.M.; Nikolenko, S.I.; Pham, S.; Prjibelski, A.D.; et al. SPAdes: A new genome assembly algorithm and its applications to single-cell sequencing. J. Comput. Biol. 2012, 19, 455–477. [Google Scholar] [CrossRef] [PubMed]
- Altschul, S.F.; Gish, W.; Miller, W.; Myers, E.W.; Lipman, D.J. Basic local alignment search tool. J. Mol. Biol. 1990, 215, 403–410. [Google Scholar] [CrossRef]
- Bhaduri, A.; Qu, K.; Lee, C.S.; Ungewickell, A.; Khavari, P.A. Rapid identification of non-human sequences in high-throughput sequencing datasets. Bioinformatics 2012, 28, 1174–1175. [Google Scholar] [CrossRef]
- Chen, K.; Wallis, J.W.; McLellan, M.D.; Larson, D.E.; Kalicki, J.M.; Pohl, C.S.; McGrath, S.D.; Wendl, M.C.; Zhang, Q.; Locke, D.P.; et al. BreakDancer: An algorithm for high-resolution mapping of genomic structural variation. Nat. Methods 2009, 6, 677–681. [Google Scholar] [CrossRef]
- Bolger, A.M.; Lohse, M.; Usadel, B. Trimmomatic: A flexible trimmer for Illumina sequence data. Bioinformatics 2014, 30, 2114–2120. [Google Scholar] [CrossRef]
- Li, H.; Handsaker, B.; Wysoker, A.; Fennell, T.; Ruan, J.; Homer, N.; Marth, G.; Abecasis, G.; Durbin, R.; 1000 Genome Project Data Processing Subgroup. The Sequence Alignment/Map format and SAMtools. Bioinformatics 2009, 25, 2078–2079. [Google Scholar] [CrossRef]
- Klindworth, A.; Pruesse, E.; Schweer, T.; Peplies, J.; Quast, C.; Horn, M.; Glockner, F.O. Evaluation of general 16S ribosomal RNA gene PCR primers for classical and next-generation sequencing-based diversity studies. Nucleic Acids Res. 2013, 41, e1. [Google Scholar] [CrossRef]
- Stollberg, S.; Kammerer, D.; Neubauer, E.; Schulz, S.; Simonitsch-Klupp, I.; Kiesewetter, B.; Raderer, M.; Lupp, A. Differential somatostatin and CXCR4 chemokine receptor expression in MALT-type lymphoma of gastric and extragastric origin. J. Cancer Res. Clin. Oncol. 2016, 142, 2239–2247. [Google Scholar] [CrossRef] [PubMed]
- Aksenova, V.; Turoverova, L.; Khotin, M.; Magnusson, K.E.; Tulchinsky, E.; Melino, G.; Pinaev, G.P.; Barlev, N.; Tentler, D. Actin-binding protein alpha-actinin 4 (ACTN4) is a transcriptional co-activator of RelA/p65 sub-unit of NF-kB. Oncotarget 2013, 4, 362–372. [Google Scholar] [CrossRef] [PubMed]
| Characteristics | No. of Patients (n = 84) | Percentage of Patients |
|---|---|---|
| Age at diagnosis (years) | ||
| Median | 66 | |
| Range | 24–92 | |
| Sex | ||
| Male | 40/79 | 51 |
| Female | 39/79 | 49 |
| Ann-Arbor stage ot diagnosis | ||
| IEA | 48/66 | 73 |
| II–IV | 18/66 | 27 |
| Localisation at diagnosis | ||
| Orbita | 29/72 | 40 |
| Conjunctiva | 18/72 | 25 |
| Lid | 18/72 | 25 |
| Lacrimal gland | 7/72 | 10 |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Johansson, P.; Klein-Hitpass, L.; Budeus, B.; Kuhn, M.; Lauber, C.; Seifert, M.; Roeder, I.; Pförtner, R.; Stuschke, M.; Dührsen, U.; et al. Identifying Genetic Lesions in Ocular Adnexal Extranodal Marginal Zone Lymphomas of the MALT Subtype by Whole Genome, Whole Exome and Targeted Sequencing. Cancers 2020, 12, 986. https://doi.org/10.3390/cancers12040986
Johansson P, Klein-Hitpass L, Budeus B, Kuhn M, Lauber C, Seifert M, Roeder I, Pförtner R, Stuschke M, Dührsen U, et al. Identifying Genetic Lesions in Ocular Adnexal Extranodal Marginal Zone Lymphomas of the MALT Subtype by Whole Genome, Whole Exome and Targeted Sequencing. Cancers. 2020; 12(4):986. https://doi.org/10.3390/cancers12040986
Chicago/Turabian StyleJohansson, Patricia, Ludger Klein-Hitpass, Bettina Budeus, Matthias Kuhn, Chris Lauber, Michael Seifert, Ingo Roeder, Roman Pförtner, Martin Stuschke, Ulrich Dührsen, and et al. 2020. "Identifying Genetic Lesions in Ocular Adnexal Extranodal Marginal Zone Lymphomas of the MALT Subtype by Whole Genome, Whole Exome and Targeted Sequencing" Cancers 12, no. 4: 986. https://doi.org/10.3390/cancers12040986
APA StyleJohansson, P., Klein-Hitpass, L., Budeus, B., Kuhn, M., Lauber, C., Seifert, M., Roeder, I., Pförtner, R., Stuschke, M., Dührsen, U., Eckstein, A., Dürig, J., & Küppers, R. (2020). Identifying Genetic Lesions in Ocular Adnexal Extranodal Marginal Zone Lymphomas of the MALT Subtype by Whole Genome, Whole Exome and Targeted Sequencing. Cancers, 12(4), 986. https://doi.org/10.3390/cancers12040986






