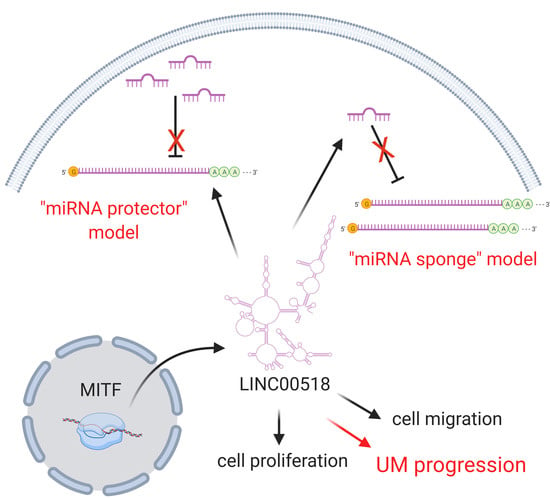
LncRNA LINC00518 Acts as an Oncogene in Uveal Melanoma by Regulating an RNA-Based Network
Abstract
Simple Summary
Abstract
1. Introduction
2. Results
2.1. Selection of lncRNAs from the UM TCGA Dataset
2.2. Differential Expression of lncRNAs in Patient Biopsies
2.3. LINC00518 May Be Involved in Metastasis-Related Processes Rather Than Cell Proliferation
2.4. LINC00518 Transient Silencing Affects the RNA-Based Network
2.5. RNA Molecular Axes of LINC00518
2.6. RNA–RNA Interaction between LINC00518 and mRNAs
2.7. Identification of Transcription Factors Potentially Regulating LINC00518 Expression
3. Discussion
4. Materials and Methods
4.1. LncRNA Selection
4.2. Patient Recruitment and Tissue, Vitreous, and Serum Sampling
4.3. RNA Isolation from FFPE Tissues, Vitreous, and Serum Samples
4.4. Subcellular Fractionation
4.5. Computational Analysis
4.6. In Vitro Functional Assays on UM Cells
- Cell cultures
- MAPK inhibition
- Cell cycle block and activation
- EMT induction
- HIF1A activation
- LINC00518 transient silencing
- Inhibition of TFs potentially regulating LINC00518 expression
4.7. Expression Analysis through Real-Time PCR
4.8. Statistical Analysis
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Souto, E.B.; Zielinska, A.; Luis, M.; Carbone, C.; Martins-Gomes, C.; Souto, S.B.; Silva, A.M. Uveal melanoma: Physiopathology and new in situ-specific therapies. Cancer Chemother. Pharmacol. 2019, 84, 15–32. [Google Scholar] [CrossRef] [PubMed]
- Berus, T.; Halon, A.; Markiewicz, A.; Orlowska-Heitzman, J.; Romanowska-Dixon, B.; Donizy, P. Clinical, Histopathological and Cytogenetic Prognosticators in Uveal Melanoma—A Comprehensive Review. Anticancer Res. 2017, 37, 6541–6549. [Google Scholar] [CrossRef] [PubMed]
- Violanti, S.S.; Bononi, I.; Gallenga, C.E.; Martini, F.; Tognon, M.; Perri, P. New Insights into Molecular Oncogenesis and Therapy of Uveal Melanoma. Cancers 2019, 11, 694. [Google Scholar] [CrossRef] [PubMed]
- Luke, J.J.; Triozzi, P.L.; McKenna, K.C.; Van Meir, E.G.; Gershenwald, J.E.; Bastian, B.C.; Gutkind, J.S.; Bowcock, A.M.; Streicher, H.Z.; Patel, P.M.; et al. Biology of advanced uveal melanoma and next steps for clinical therapeutics. Pigment Cell Melanoma Res. 2015, 28, 135–147. [Google Scholar] [CrossRef] [PubMed]
- Harbour, J.W. The genetics of uveal melanoma: An emerging framework for targeted therapy. Pigment Cell Melanoma Res. 2012, 25, 171–181. [Google Scholar] [CrossRef] [PubMed]
- Huang, T.; Alvarez, A.; Hu, B.; Cheng, S.Y. Noncoding RNAs in cancer and cancer stem cells. Chin. J. Cancer 2013, 32, 582–593. [Google Scholar] [CrossRef] [PubMed]
- Ragusa, M.; Barbagallo, C.; Brex, D.; Caponnetto, A.; Cirnigliaro, M.; Battaglia, R.; Barbagallo, D.; Di Pietro, C.; Purrello, M. Molecular Crosstalking among Noncoding RNAs: A New Network Layer of Genome Regulation in Cancer. Int. J. Genom. 2017, 2017, 4723193. [Google Scholar] [CrossRef]
- Ragusa, M.; Barbagallo, C.; Statello, L.; Caltabiano, R.; Russo, A.; Puzzo, L.; Avitabile, T.; Longo, A.; Toro, M.D.; Barbagallo, D.; et al. miRNA profiling in vitreous humor, vitreal exosomes and serum from uveal melanoma patients: Pathological and diagnostic implications. Cancer Biol. Ther. 2015, 16, 1387–1396. [Google Scholar] [CrossRef]
- Aprile, M.; Katopodi, V.; Leucci, E.; Costa, V. LncRNAs in Cancer: From garbage to Junk. Cancers 2020, 12, 3220. [Google Scholar] [CrossRef]
- Zhang, X.Z.; Liu, H.; Chen, S.R. Mechanisms of Long Non-Coding RNAs in Cancers and Their Dynamic Regulations. Cancers 2020, 12, 1245. [Google Scholar] [CrossRef]
- Carlevaro-Fita, J.; Lanzós, A.; Feuerbach, L.; Hong, C.; Mas-Ponte, D.; Pedersen, J.S.; PCAWG Drivers and Functional Interpretation Group; Johnson, R.; PCAWG Consortium. Cancer LncRNA Census reveals evidence for deep functional conservation of long noncoding RNAs in tumorigenesis. Commun. Biol. 2020, 3, 56. [Google Scholar] [CrossRef] [PubMed]
- Jiang, M.C.; Ni, J.J.; Cui, W.Y.; Wang, B.Y.; Zhuo, W. Emerging roles of lncRNA in cancer and therapeutic opportunities. Am. J. Cancer Res. 2019, 9, 1354–1366. [Google Scholar] [PubMed]
- Wu, X.; Yuan, Y.; Ma, R.; Xu, B.; Zhang, R. lncRNA SNHG7 affects malignant tumor behaviors through downregulation of EZH2 in uveal melanoma cell lines. Oncol. Lett. 2020, 19, 1505–1515. [Google Scholar] [CrossRef] [PubMed]
- Wu, X.; Li, X.F.; Wu, Q.; Ma, R.Q.; Qian, J.; Zhang, R. LncRNA SNHG15 predicts poor prognosis in uveal melanoma and its potential pathways. Int. J. Ophthalmol. 2020, 13, 1195–1201. [Google Scholar] [CrossRef] [PubMed]
- Wu, S.; Chen, H.; Zuo, L.; Jiang, H.; Yan, H. Suppression of long noncoding RNA MALAT1 inhibits the development of uveal melanoma via microRNA-608-mediated inhibition of HOXC4. Am. J. Physiol. Cell Physiol. 2020, 318, C903–C912. [Google Scholar] [CrossRef]
- Ragusa, M.; Barbagallo, C.; Statello, L.; Condorelli, A.G.; Battaglia, R.; Tamburello, L.; Barbagallo, D.; Di Pietro, C.; Purrello, M. Non-coding landscapes of colorectal cancer. World J. Gastroenterol. 2015, 21, 11709–11739. [Google Scholar] [CrossRef]
- Barbagallo, C.; Brex, D.; Caponnetto, A.; Cirnigliaro, M.; Scalia, M.; Magnano, A.; Caltabiano, R.; Barbagallo, D.; Biondi, A.; Cappellani, A.; et al. LncRNA UCA1, Upregulated in CRC Biopsies and Downregulated in Serum Exosomes, Controls mRNA Expression by RNA-RNA Interactions. Mol. Ther. Nucleic Acids 2018, 12, 229–241. [Google Scholar] [CrossRef]
- Wang, H.B.; Wei, H.; Wang, J.S.; Li, L.; Chen, A.Y.; Li, Z.G. Down-regulated expression of LINC00518 prevents epithelial cell growth and metastasis in breast cancer through the inhibition of CDX2 methylation and the Wnt signaling pathway. Biochim. Biophys. Acta Mol. Basis Dis. 2019, 1865, 708–723. [Google Scholar] [CrossRef]
- Chang, L.; Hu, Z.; Zhou, Z.; Zhang, H. Linc00518 Contributes to Multidrug Resistance through Regulating the MiR-199a/MRP1 Axis in Breast Cancer. Cell. Physiol. Biochem. 2018, 48, 16–28. [Google Scholar] [CrossRef]
- Wang, D.W.; You, D.; Dong, J.; Liu, T.F. Knockdown of long non-coding RNA LINC00518 inhibits cervical cancer proliferation and metastasis by modulating JAK/STAT3 signaling. Eur. Rev. Med. Pharmacol. Sci. 2019, 23, 496–506. [Google Scholar] [CrossRef]
- He, J.; Sun, M.; Geng, H.; Tian, S. Long non-coding RNA Linc00518 promotes paclitaxel resistance of the human prostate cancer by sequestering miR-216b-5p. Biol. Cell 2019, 111, 39–50. [Google Scholar] [CrossRef] [PubMed]
- Luan, W.; Ding, Y.; Ma, S.; Ruan, H.; Wang, J.; Lu, F. Long noncoding RNA LINC00518 acts as a competing endogenous RNA to promote the metastasis of malignant melanoma via miR-204-5p/AP1S2 axis. Cell Death Dis. 2019, 10, 855. [Google Scholar] [CrossRef] [PubMed]
- Gerami, P.; Alsobrook, J.P., 2nd; Palmer, T.J.; Robin, H.S. Development of a novel noninvasive adhesive patch test for the evaluation of pigmented lesions of the skin. J. Am. Acad. Dermatol. 2014, 71, 237–244. [Google Scholar] [CrossRef] [PubMed]
- Gerami, P.; Yao, Z.; Polsky, D.; Jansen, B.; Busam, K.; Ho, J.; Martini, M.; Ferris, L.K. Development and validation of a noninvasive 2-gene molecular assay for cutaneous melanoma. J. Am. Acad. Dermatol. 2017, 76, 114–120.e2. [Google Scholar] [CrossRef] [PubMed]
- Ferris, L.K.; Jansen, B.; Ho, J.; Busam, K.J.; Gross, K.; Hansen, D.D.; Alsobrook, J.P., 2nd; Yao, Z.; Peck, G.L.; Gerami, P. Utility of a Noninvasive 2-Gene Molecular Assay for Cutaneous Melanoma and Effect on the Decision to Biopsy. JAMA Dermatol. 2017, 153, 675–680. [Google Scholar] [CrossRef]
- Ferris, L.K.; Gerami, P.; Skelsey, M.K.; Peck, G.; Hren, C.; Gorman, C.; Frumento, T.; Siegel, D.M. Real-world performance and utility of a noninvasive gene expression assay to evaluate melanoma risk in pigmented lesions. Melanoma Res. 2018, 28, 478–482. [Google Scholar] [CrossRef]
- Rivers, J.K.; Copley, M.R.; Svoboda, R.; Rigel, D.S. Non-Invasive Gene Expression Testing to Rule Out Melanoma. Skin Ther. Lett. 2018, 23, 1–4. [Google Scholar]
- Zhang, X.; Feng, Y.; Gao, Y.; Hu, J. Long Noncoding RNA LINC00634 Functions as an Oncogene in Esophageal Squamous Cell Carcinoma Through the miR-342-3p/Bcl2L1 Axis. Technol. Cancer Res. Treat. 2020, 19, 1533033820928508. [Google Scholar] [CrossRef]
- Zhou, J.; Zhang, C.; Pan, J.; Chen, L.; Qi, S.T. Interleukin6 induces an epithelialmesenchymal transition phenotype in human adamantinomatous craniopharyngioma cells and promotes tumor cell migration. Mol. Med. Rep. 2017, 15, 4123–4131. [Google Scholar] [CrossRef]
- Liu, F.; Song, S.; Yi, Z.; Zhang, M.; Li, J.; Yang, F.; Yin, H.; Yu, X.; Guan, C.; Liu, Y.; et al. HGF induces EMT in non-small-cell lung cancer through the hBVR pathway. Eur. J. Pharmacol. 2017, 811, 180–190. [Google Scholar] [CrossRef]
- Xie, G.; Yao, Q.; Liu, Y.; Du, S.; Liu, A.; Guo, Z.; Sun, A.; Ruan, J.; Chen, L.; Ye, C.; et al. IL-6-induced epithelial-mesenchymal transition promotes the generation of breast cancer stem-like cells analogous to mammosphere cultures. Int. J. Oncol. 2012, 40, 1171–1179. [Google Scholar] [CrossRef] [PubMed]
- Miao, J.W.; Liu, L.J.; Huang, J. Interleukin-6-induced epithelial-mesenchymal transition through signal transducer and activator of transcription 3 in human cervical carcinoma. Int. J. Oncol. 2014, 45, 165–176. [Google Scholar] [CrossRef] [PubMed]
- Jiao, D.; Wang, J.; Lu, W.; Tang, X.; Chen, J.; Mou, H.; Chen, Q.Y. Curcumin inhibited HGF-induced EMT and angiogenesis through regulating c-Met dependent PI3K/Akt/mTOR signaling pathways in lung cancer. Mol. Ther. Oncolytics 2016, 3, 16018. [Google Scholar] [CrossRef] [PubMed]
- Nagai, T.; Arao, T.; Furuta, K.; Sakai, K.; Kudo, K.; Kaneda, H.; Tamura, D.; Aomatsu, K.; Kimura, H.; Fujita, Y.; et al. Sorafenib inhibits the hepatocyte growth factor-mediated epithelial mesenchymal transition in hepatocellular carcinoma. Mol. Cancer Ther. 2011, 10, 169–177. [Google Scholar] [CrossRef] [PubMed]
- Gong, C.; Shen, J.; Fang, Z.; Qiao, L.; Feng, R.; Lin, X.; Li, S. Abnormally expressed JunB transactivated by IL-6/STAT3 signaling promotes uveal melanoma aggressiveness via epithelial-mesenchymal transition. Biosci. Rep. 2018, 38, BSR20180532. [Google Scholar] [CrossRef] [PubMed]
- Ye, M.; Hu, D.; Tu, L.; Zhou, X.; Lu, F.; Wen, B.; Wu, W.; Lin, Y.; Zhou, Z.; Qu, J. Involvement of PI3K/Akt signaling pathway in hepatocyte growth factor-induced migration of uveal melanoma cells. Investig. Ophthalmol. Vis. Sci. 2008, 49, 497–504. [Google Scholar] [CrossRef]
- Li, Q.; Ma, R.; Zhang, M. CoCl2 increases the expression of hypoxic markers HIF-1alpha, VEGF and CXCR4 in breast cancer MCF-7 cells. Oncol. Lett. 2018, 15, 1119–1124. [Google Scholar] [CrossRef]
- Liu, Z.J.; Semenza, G.L.; Zhang, H.F. Hypoxia-inducible factor 1 and breast cancer metastasis. J. Zhejiang Univ. Sci. B 2015, 16, 32–43. [Google Scholar] [CrossRef]
- Woodman, S.E. Metastatic uveal melanoma: Biology and emerging treatments. Cancer J. 2012, 18, 148–152. [Google Scholar] [CrossRef]
- Noh, J.H.; Kim, K.M.; McClusky, W.G.; Abdelmohsen, K.; Gorospe, M. Cytoplasmic functions of long noncoding RNAs. Wiley Interdiscip. Rev. RNA 2018, 9, e1471. [Google Scholar] [CrossRef]
- Jo, J.H.; Park, S.B.; Park, S.; Lee, H.S.; Kim, C.; Jung, D.E.; Song, S.Y. Novel Gastric Cancer Stem Cell-Related Marker LINGO2 Is Associated with Cancer Cell Phenotype and Patient Outcome. Int. J. Mol. Sci. 2019, 20, 555. [Google Scholar] [CrossRef] [PubMed]
- Fane, M.; Harris, L.; Smith, A.G.; Piper, M. Nuclear factor one transcription factors as epigenetic regulators in cancer. Int. J. Cancer 2017, 140, 2634–2641. [Google Scholar] [CrossRef] [PubMed]
- Lee, J.S.; Xiao, J.; Patel, P.; Schade, J.; Wang, J.; Deneen, B.; Erdreich-Epstein, A.; Song, H.R. A novel tumor-promoting role for nuclear factor IA in glioblastomas is mediated through negative regulation of p53, p21, and PAI1. Neuro-Oncology 2014, 16, 191–203. [Google Scholar] [CrossRef] [PubMed]
- Song, H.R.; Gonzalez-Gomez, I.; Suh, G.S.; Commins, D.L.; Sposto, R.; Gilles, F.H.; Deneen, B.; Erdreich-Epstein, A. Nuclear factor IA is expressed in astrocytomas and is associated with improved survival. Neuro-Oncology 2010, 12, 122–132. [Google Scholar] [CrossRef] [PubMed]
- Lin, D.D.; Shen, Y.; Qiao, S.; Liu, W.W.; Zheng, L.; Wang, Y.N.; Cui, N.; Wang, Y.F.; Zhao, S.; Shi, J.H. Upregulation of OTUD7B (Cezanne) Promotes Tumor Progression via AKT/VEGF Pathway in Lung Squamous Carcinoma and Adenocarcinoma. Front. Oncol. 2019, 9, 862. [Google Scholar] [CrossRef] [PubMed]
- Lei, S.; He, Z.; Chen, T.; Guo, X.; Zeng, Z.; Shen, Y.; Jiang, J. Long noncoding RNA 00976 promotes pancreatic cancer progression through OTUD7B by sponging miR-137 involving EGFR/MAPK pathway. J. Exp. Clin. Cancer Res. 2019, 38, 470. [Google Scholar] [CrossRef]
- Wright, P.K. Targeting vesicle trafficking: An important approach to cancer chemotherapy. Recent Pat. Anticancer Drug Discov. 2008, 3, 137–147. [Google Scholar] [CrossRef]
- Sun, W.; Tian, B.X.; Wang, S.H.; Liu, P.J.; Wang, Y.C. The function of SEC22B and its role in human diseases. Cytoskeleton 2020, 77, 303–312. [Google Scholar] [CrossRef]
- Luftman, K.; Hasan, N.; Day, P.; Hardee, D.; Hu, C. Silencing of VAMP3 inhibits cell migration and integrin-mediated adhesion. Biochem. Biophys. Res. Commun. 2009, 380, 65–70. [Google Scholar] [CrossRef]
- Clancy, J.W.; Sedgwick, A.; Rosse, C.; Muralidharan-Chari, V.; Raposo, G.; Method, M.; Chavrier, P.; D’Souza-Schorey, C. Regulated delivery of molecular cargo to invasive tumour-derived microvesicles. Nat. Commun. 2015, 6, 6919. [Google Scholar] [CrossRef]
- D’Mello, S.A.; Finlay, G.J.; Baguley, B.C.; Askarian-Amiri, M.E. Signaling Pathways in Melanogenesis. Int. J. Mol. Sci. 2016, 17, 1144. [Google Scholar] [CrossRef] [PubMed]
- Chen, X.; Wang, J.; Shen, H.; Lu, J.; Li, C.; Hu, D.N.; Dong, X.D.; Yan, D.; Tu, L. Epigenetics, microRNAs, and carcinogenesis: Functional role of microRNA-137 in uveal melanoma. Investig. Ophthalmol. Vis. Sci. 2011, 52, 1193–1199. [Google Scholar] [CrossRef] [PubMed]
- Fagerberg, L.; Hallstrom, B.M.; Oksvold, P.; Kampf, C.; Djureinovic, D.; Odeberg, J.; Habuka, M.; Tahmasebpoor, S.; Danielsson, A.; Edlund, K.; et al. Analysis of the human tissue-specific expression by genome-wide integration of transcriptomics and antibody-based proteomics. Mol. Cell. Proteom. 2014, 13, 397–406. [Google Scholar] [CrossRef] [PubMed]
- Robertson, A.G.; Shih, J.; Yau, C.; Gibb, E.A.; Oba, J.; Mungall, K.L.; Hess, J.M.; Uzunangelov, V.; Walter, V.; Danilova, L.; et al. Integrative Analysis Identifies Four Molecular and Clinical Subsets in Uveal Melanoma. Cancer Cell 2018, 33, 151. [Google Scholar] [CrossRef] [PubMed]
- Bakalian, S.; Marshall, J.C.; Logan, P.; Faingold, D.; Maloney, S.; Di Cesare, S.; Martins, C.; Fernandes, B.F.; Burnier, M.N., Jr. Molecular pathways mediating liver metastasis in patients with uveal melanoma. Clin. Cancer Res. 2008, 14, 951–956. [Google Scholar] [CrossRef]
- Mann, M.; Wright, P.R.; Backofen, R. IntaRNA 2.0: Enhanced and customizable prediction of RNA-RNA interactions. Nucleic Acids Res. 2017, 45, W435–W439. [Google Scholar] [CrossRef]
- Wang, R.; He, Y.; Robinson, V.; Yang, Z.; Hessler, P.; Lasko, L.M.; Lu, X.; Bhathena, A.; Lai, A.; Uziel, T.; et al. Targeting Lineage-specific MITF Pathway in Human Melanoma Cell Lines by A-485, the Selective Small-molecule Inhibitor of p300/CBP. Mol. Cancer Ther. 2018, 17, 2543–2550. [Google Scholar] [CrossRef]
- Ding, J.; Zhang, Z.M.; Xia, Y.; Liao, G.Q.; Pan, Y.; Liu, S.; Zhang, Y.; Yan, Z.S. LSD1-mediated epigenetic modification contributes to proliferation and metastasis of colon cancer. Br. J. Cancer 2013, 109, 994–1003. [Google Scholar] [CrossRef]
- Kashyap, V.; Ahmad, S.; Nilsson, E.M.; Helczynski, L.; Kenna, S.; Persson, J.L.; Gudas, L.J.; Mongan, N.P. The lysine specific demethylase-1 (LSD1/KDM1A) regulates VEGF-A expression in prostate cancer. Mol. Oncol. 2013, 7, 555–566. [Google Scholar] [CrossRef]











| 92.1 | MEL270 | ||||||
|---|---|---|---|---|---|---|---|
| miRNAs Regulating mRNA Expression | miRNA/mRNA | 24 h | 48 h | Correlation vs. LINC00518 | 24 h | 48 h | Correlation vs. LINC00518 |
| miR-143-3p | −1.08 (0.55) | 1.71 (0.61) | −0.2 (0.53) | 1.58 (0.51) | −1.63 (0.12) | −0.35 (0.26) | |
| miR-145-5p | −1.19 (0.47) | 1.38 (0.46) | −0.49 (0.1) | 1.98 (0.65) | 1.08 (0.87) | 0.54 (0.06) | |
| miR-199a-5p | 1.23 (0.99) | 1.46 (0.08) | −0.5 (0.09) | 1.13 (0.53) | 1.07 (0.87) | −0.16 (0.61) | |
| miR-212-5p | 1 (0.63) | 1.33 (0.86) | −0.35 (0.26) | 1.6 (0.99) | −1.09 (0.51) | 0.47 (0.12) | |
| miR-3191-3p | 1.94 (0.96) | 2.5 (0.08) | 0.11 (0.73) | 1.84 (0.33) | 1.21 (0.68) | 0.35 (0.26) | |
| miR-497-5p | −1.12 (0.48) | 1.65 (0.48) | −0.48 (0.11) | 1.22 (0.86) | 1.19 (0.72) | 0.44 (0.15) | |
| miR-212-5p, miR-3191-3p | AHCYL2 | −1.22 (0.18) | 1.82 (0.16) | −0.38 (0.22) | 1 (0.98) | 1.19 (0.21) | 0.65 (0.02) |
| miR-199a-5p | CRTAP | −1.23 (0.11) | 1.11 (0.37) | 0.51 (0.09) | −1.05 (0.75) | 1.27 (0.13) | 0.64 (0.02) |
| miR-3191-3p | ENTPD1 | 1.04 (0.96) | 1.59 (0.18) | 0.52 (0.08) | −1.02 (0.81) | 1.03 (0.82) | 0.74 (0.005) |
| miR-145-5p | F11R | 1.33 (0.24) | 1.08 (0.12) | −0.7 (0.01) | 1.04 (0.67) | 1.07 (0.54) | 0.59 (0.04) |
| miR-145-5p, miR-212-5p | IFFO2 | 1.01 (0.99) | 1.42 (0.26) | 0.13 (0.68) | 1.01 (0.87) | 1.2 (0.29) | 0.73 (0.007) |
| miR-145-5p, miR-3191-3p | IP6K1 | 1.13 (0.62) | 2.41 (0.25) | −0.22 (0.49) | 1.15 (0.61) | 1.05 (0.71) | 0.74 (0.005) |
| miR-3191-3p | KCTD15 | 1.02 (0.87) | −1.15 (0.1) | 0.22 (0.49) | 1.05 (0.72) | 1.05 (0.93) | 0.18 (0.57) |
| miR-143-3p | KLF8 | −1.68 (0.07) | −1.12 (0.26) | 0.48 (0.11) | −1.39 (0.04) | 1.28 (0.46) | 0.15 (0.64) |
| miR-497-5p | LINGO2 | −1.68 (0.09) | −1.57 (0.15) | 0.59 (0.04) | −1.47 (0.04) | −1.14 (0.18) | 0.42 (0.17) |
| miR-143-3p | NFIA | −1.47 (0.13) | 1.56 (0.25) | 0.58 (0.04) | −1.48 (0.04) | 1.12 (0.48) | 0.34 (0.27) |
| miR-145-5p, miR-497-5p | OTUD7B | −1.4 (0.03) | 1.26 (0.54) | 0.45 (0.14) | −1.23 (0.04) | 1 (0.59) | 0.75 (0.004) |
| miR-3191-3p | RAB43 | −1.52 (0.11) | 1.27 (0.43) | 0.43 (0.16) | −1.06 (0.65) | −1.11 (0.75) | 0.72 (0.008) |
| miR-212-5p | RABGAP1L | 1 (0.76) | 1.55 (0.06) | −0.29 (0.36) | 1.24 (0.12) | 1.13 (0.25) | 0.31 (0.32) |
| miR-199a-5p, miR-212-5p | SEC22C | −1.31 (0.001) | 1.36 (0.16) | −0.11 (0.73) | −1.11 (0.03) | −1.07 (0.01) | 0.71 (0.009) |
| miR-497-5p | SEMA6A | 1.38 (0.17) | 1.38 (0.34) | −0.58 (0.04) | −1.03 (0.99) | −1.06 (0.04) | 0.41 (0.18) |
| miR-3191-3p | VAMP3 | −1.37 (0.02) | 1.31 (0.25) | 0.58 (0.04) | −1.05 (0.26) | 1.36 (0.18) | 0.58 (0.04) |
| miR-199a-5p | ZBTB20 | −1.03 (0.75) | −1.1 (0.17) | −0.31 (0.32) | 1.22 (0.19) | 1.21 (0.35) | 0.04 (0.9) |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Barbagallo, C.; Caltabiano, R.; Broggi, G.; Russo, A.; Puzzo, L.; Avitabile, T.; Longo, A.; Reibaldi, M.; Barbagallo, D.; Di Pietro, C.; et al. LncRNA LINC00518 Acts as an Oncogene in Uveal Melanoma by Regulating an RNA-Based Network. Cancers 2020, 12, 3867. https://doi.org/10.3390/cancers12123867
Barbagallo C, Caltabiano R, Broggi G, Russo A, Puzzo L, Avitabile T, Longo A, Reibaldi M, Barbagallo D, Di Pietro C, et al. LncRNA LINC00518 Acts as an Oncogene in Uveal Melanoma by Regulating an RNA-Based Network. Cancers. 2020; 12(12):3867. https://doi.org/10.3390/cancers12123867
Chicago/Turabian StyleBarbagallo, Cristina, Rosario Caltabiano, Giuseppe Broggi, Andrea Russo, Lidia Puzzo, Teresio Avitabile, Antonio Longo, Michele Reibaldi, Davide Barbagallo, Cinzia Di Pietro, and et al. 2020. "LncRNA LINC00518 Acts as an Oncogene in Uveal Melanoma by Regulating an RNA-Based Network" Cancers 12, no. 12: 3867. https://doi.org/10.3390/cancers12123867
APA StyleBarbagallo, C., Caltabiano, R., Broggi, G., Russo, A., Puzzo, L., Avitabile, T., Longo, A., Reibaldi, M., Barbagallo, D., Di Pietro, C., Purrello, M., & Ragusa, M. (2020). LncRNA LINC00518 Acts as an Oncogene in Uveal Melanoma by Regulating an RNA-Based Network. Cancers, 12(12), 3867. https://doi.org/10.3390/cancers12123867