Cancer-Associated Fibroblasts: Epigenetic Regulation and Therapeutic Intervention in Breast Cancer
Simple Summary
Abstract
1. Introduction
2. Breast Cancer and CAFs
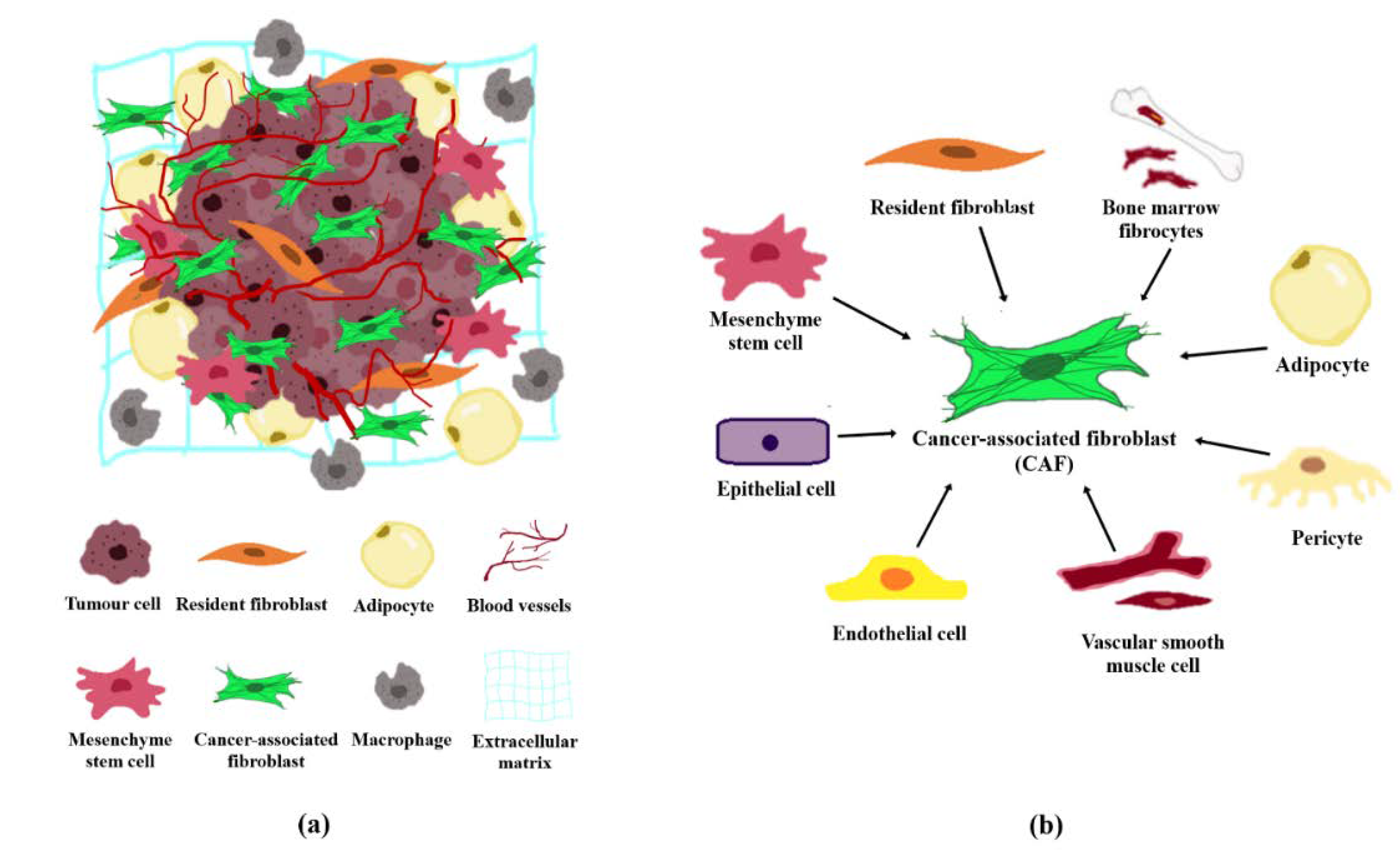
2.1. The Origin of CAFs
2.2. BCAFs Heterogeneity
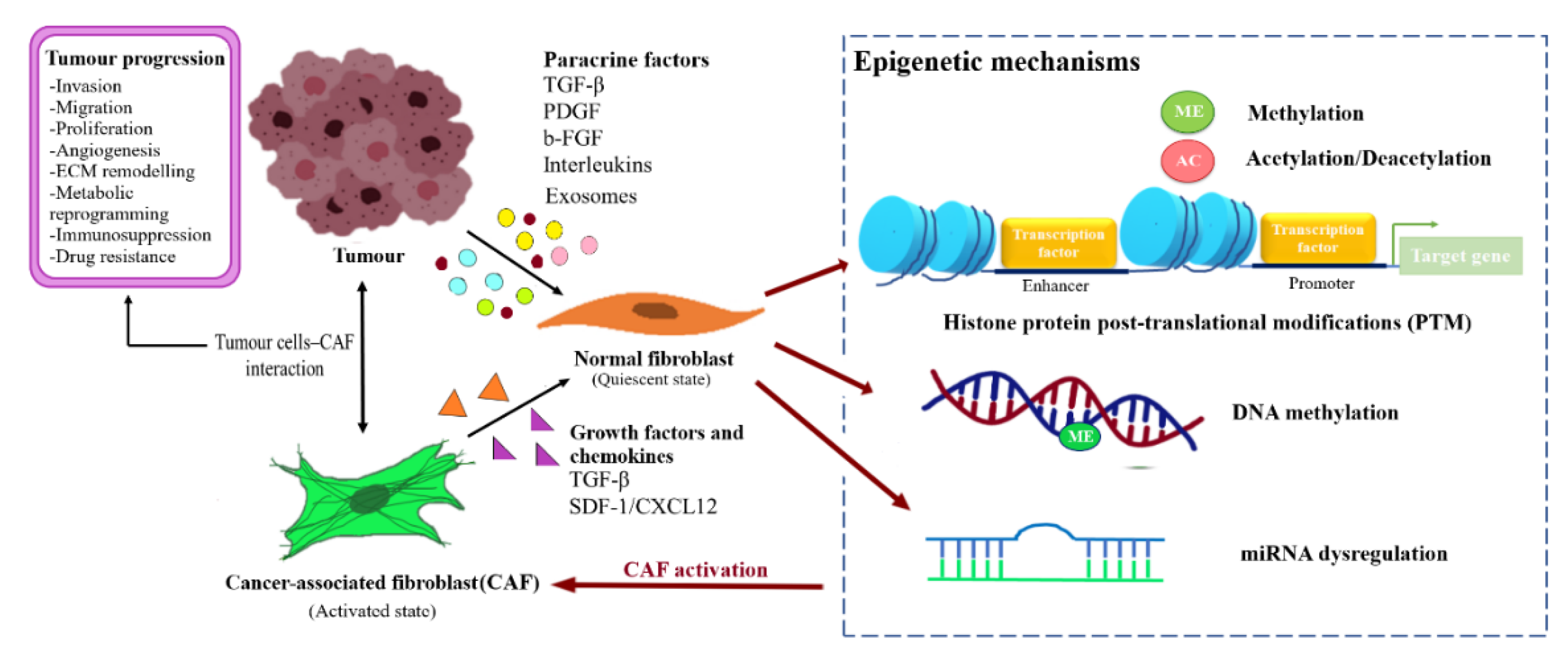
3. Reprogramming of NFs into CAFs
4. Epigenetic Regulation in BCAFs
4.1. Post-Translational Modification of Histones in BCAFs
4.2. DNA Methylation in BCAFs
4.3. miRNA Regulation in BCAFs
5. Therapeutic Intervention Targeting Epigenetics in BCAFs
6. Conclusions and Future Perspectives
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Chen, X.; Song, E. Turning foes to friends: Targeting cancer-associated fibroblasts. Nat. Rev. Drug Discov. 2018, 18, 99–115. [Google Scholar] [CrossRef]
- LeBleu, V.S.; Kalluri, R. A peek into cancer-associated fibroblasts: Origins, functions and translational impact. Dis. Models Mech. 2018, 11, dmm029447. [Google Scholar] [CrossRef] [PubMed]
- Anderberg, C.; Pietras, K. On the origin of cancer-associated fibroblasts. Cell Cycle (Georget. Tex.) 2009, 8, 1461–1465. [Google Scholar] [CrossRef] [PubMed]
- Gascard, P.; Tlsty, T.D. Carcinoma-associated fibroblasts: Orchestrating the composition of malignancy. Genes Dev. 2016, 30, 1002–1019. [Google Scholar] [CrossRef] [PubMed]
- Kalluri, R. The biology and function of fibroblasts in cancer. Nat. Rev. Cancer 2016, 16, 582–598. [Google Scholar] [CrossRef]
- Liu, T.; Han, C.; Wang, S.; Fang, P.; Ma, Z.; Xu, L.; Yin, R. Cancer-associated fibroblasts: An emerging target of anti-cancer immunotherapy. J. Hematol. Oncol. 2019, 12, e86. [Google Scholar] [CrossRef]
- Sahai, E.; Astsaturov, I.; Cukierman, E.; DeNardo, D.G.; Egeblad, M.; Evans, R.M.; Fearon, D.; Greten, F.R.; Hingorani, S.R.; Hunter, T.; et al. A framework for advancing our understanding of cancer-associated fibroblasts. Nat. Rev. Cancer 2020, 20, 174–186. [Google Scholar] [CrossRef]
- Dvorak, K.M.; Pettee, K.M.; Rubinic-Minotti, K.; Su, R.; Nestor-Kalinoski, A.; Eisenmann, K.M. Carcinoma associated fibroblasts (CAFs) promote breast cancer motility by suppressing mammalian Diaphanous-related formin-2 (mDia2). PLoS ONE 2018, 13, e0195278. [Google Scholar] [CrossRef]
- Albrengues, J.; Bertero, T.; Grasset, E.; Bonan, S.; Maiel, M.; Bourget, I.; Philippe, C.; Herraiz Serrano, C.; Benamar, S.; Croce, O.; et al. Epigenetic switch drives the conversion of fibroblasts into proinvasive cancer-associated fibroblasts. Nat. Commun. 2015, 6, 10204. [Google Scholar] [CrossRef]
- Avagliano, A.; Granato, G.; Ruocco, M.R.; Romano, V.; Belviso, I.; Carfora, A.; Montagnani, S.; Arcucci, A. Metabolic Reprogramming of Cancer Associated Fibroblasts: The Slavery of Stromal Fibroblasts. Biomed Res. Int. 2018, 2018, e6075403. [Google Scholar] [CrossRef]
- Vazquez-Villa, F.; Garcia-Ocana, M.; Galvan, J.A.; Garcia-Martinez, J.; Garcia-Pravia, C.; Menendez-Rodriguez, P.; Gonzalez-del Rey, C.; Barneo-Serra, L.; de Los Toyos, J.R. COL11A1/(pro)collagen 11A1 expression is a remarkable biomarker of human invasive carcinoma-associated stromal cells and carcinoma progression. Tumour Biol. J. Int. Soc. Oncodevelopmental Biol. Med. 2015, 36, 2213–2222. [Google Scholar] [CrossRef] [PubMed]
- Maris, P.; Blomme, A.; Palacios, A.P.; Costanza, B.; Bellahcene, A.; Bianchi, E.; Gofflot, S.; Drion, P.; Trombino, G.E.; Di Valentin, E.; et al. Asporin Is a Fibroblast-Derived TGF-beta1 Inhibitor and a Tumor Suppressor Associated with Good Prognosis in Breast Cancer. PLoS Med. 2015, 12, e1001871. [Google Scholar] [CrossRef]
- Yeung, T.L.; Leung, C.S.; Mok, S.C. CAF reprogramming inhibits ovarian cancer progression. Cell Cycle (Georget. Tex.) 2014, 13, 3783–3784. [Google Scholar] [CrossRef] [PubMed]
- Bray, F.; Ferlay, J.; Soerjomataram, I.; Siegel, R.L.; Torre, L.A.; Jemal, A. Global cancer statistics 2018: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA Cancer J. Clin. 2018, 68, 394–424. [Google Scholar] [CrossRef] [PubMed]
- Fragomeni, S.M.; Sciallis, A.; Jeruss, J.S. Molecular Subtypes and Local-Regional Control of Breast Cancer. Surg. Oncol. Clin. N. Am. 2018, 27, 95–120. [Google Scholar] [CrossRef] [PubMed]
- Louault, K.; Bonneaud, T.L.; Séveno, C.; Gomez-Bougie, P.; Nguyen, F.; Gautier, F.; Bourgeois, N.; Loussouarn, D.; Kerdraon, O.; Barillé-Nion, S.; et al. Interactions between cancer-associated fibroblasts and tumor cells promote MCL-1 dependency in estrogen receptor-positive breast cancers. Oncogene 2019, 38, 3261–3273. [Google Scholar] [CrossRef] [PubMed]
- Ao, Z.; Shah, S.H.; Machlin, L.M.; Parajuli, R.; Miller, P.C.; Rawal, S.; Williams, A.J.; Cote, R.J.; Lippman, M.E.; Datar, R.H.; et al. Identification of Cancer-Associated Fibroblasts in Circulating Blood from Patients with Metastatic Breast Cancer. Cancer Res. 2015, 75, 4681–4687. [Google Scholar] [CrossRef]
- Ruocco, M.R.; Avagliano, A.; Granato, G.; Imparato, V.; Masone, S.; Masullo, M.; Nasso, R.; Montagnani, S.; Arcucci, A. Involvement of Breast Cancer-Associated Fibroblasts in Tumor Development, Therapy Resistance and Evaluation of Potential Therapeutic Strategies. Curr. Med. Chem. 2018, 25, 3414–3434. [Google Scholar] [CrossRef]
- Shiga, K.; Hara, M.; Nagasaki, T.; Sato, T.; Takahashi, H.; Takeyama, H. Cancer-Associated Fibroblasts: Their Characteristics and Their Roles in Tumor Growth. Cancers (Basel) 2015, 7, 2443–2458. [Google Scholar] [CrossRef]
- Qiao, A.; Gu, F.; Guo, X.; Zhang, X.; Fu, L. Breast cancer-associated fibroblasts: Their roles in tumor initiation, progression and clinical applications. Front. Med. 2016, 10, 33–40. [Google Scholar] [CrossRef]
- Kadel, D.; Zhang, Y.; Sun, H.-R.; Zhao, Y.; Dong, Q.-Z.; Qin, L.-x. Current perspectives of cancer-associated fibroblast in therapeutic resistance: Potential mechanism and future strategy. Cell Biol. Toxicol. 2019. [Google Scholar] [CrossRef]
- Luo, H.; Tu, G.; Liu, Z.; Liu, M. Cancer-associated fibroblasts: A multifaceted driver of breast cancer progression. Cancer Lett. 2015, 361, 155–163. [Google Scholar] [CrossRef] [PubMed]
- Buchsbaum, R.J.; Oh, S.Y. Breast Cancer-Associated Fibroblasts: Where We Are and Where We Need to Go. Cancers (Basel) 2016, 8, 19. [Google Scholar] [CrossRef] [PubMed]
- Mathot, P.; Grandin, M.; Devailly, G.; Souaze, F.; Cahais, V.; Moran, S.; Campone, M.; Herceg, Z.; Esteller, M.; Juin, P.; et al. DNA methylation signal has a major role in the response of human breast cancer cells to the microenvironment. Oncogenesis 2017, 6, e390. [Google Scholar] [CrossRef]
- Östman, A. Cancer-associated fibroblasts: Recent developments and emerging challenges. Semin. Cancer Biol. 2014, 25, 1–2. [Google Scholar] [CrossRef]
- Eiro, N.; González, L.; Martínez-Ordoñez, A.; Fernandez-Garcia, B.; González, L.O.; Cid, S.; Dominguez, F.; Perez-Fernandez, R.; Vizoso, F.J. Cancer-associated fibroblasts affect breast cancer cell gene expression, invasion and angiogenesis. Cell. Oncol. 2018, 41, 369–378. [Google Scholar] [CrossRef]
- Zhang, X.H.-F.; Jin, X.; Malladi, S.; Zou, Y.; Wen, Y.H.; Brogi, E.; Smid, M.; Foekens, J.A.; Massagué, J. Selection of bone metastasis seeds by mesenchymal signals in the primary tumor stroma. Cell 2013, 154, 1060–1073. [Google Scholar] [CrossRef]
- Paulsson, J.; Rydén, L.; Strell, C.; Frings, O.; Tobin, N.P.; Fornander, T.; Bergh, J.; Landberg, G.; Stål, O.; Östman, A. High expression of stromal PDGFRβ is associated with reduced benefit of tamoxifen in breast cancer. J. Pathol. Clin. Res. 2016, 3, 38–43. [Google Scholar] [CrossRef]
- Kalluri, R.; Zeisberg, M. Fibroblasts in cancer. Nat. Rev. Cancer 2006, 6, 392–401. [Google Scholar] [CrossRef]
- Ronnov-Jessen, L.; Petersen, O.W.; Koteliansky, V.E.; Bissell, M.J. The origin of the myofibroblasts in breast cancer. Recapitulation of tumor environment in culture unravels diversity and implicates converted fibroblasts and recruited smooth muscle cells. J. Clin. Investig. 1995, 95, 859–873. [Google Scholar] [CrossRef]
- Kidd, S.; Spaeth, E.; Watson, K.; Burks, J.; Lu, H.; Klopp, A.; Andreeff, M.; Marini, F.C. Origins of the tumor microenvironment: Quantitative assessment of adipose-derived and bone marrow-derived stroma. PLoS ONE 2012, 7, e30563. [Google Scholar] [CrossRef] [PubMed]
- Cho, J.A.; Park, H.; Lim, E.H.; Lee, K.W. Exosomes from breast cancer cells can convert adipose tissue-derived mesenchymal stem cells into myofibroblast-like cells. Int. J. Oncol. 2012, 40, 130–138. [Google Scholar] [PubMed]
- Weber, C.E.; Kothari, A.N.; Wai, P.Y.; Li, N.Y.; Driver, J.; Zapf, M.A.; Franzen, C.A.; Gupta, G.N.; Osipo, C.; Zlobin, A. Osteopontin mediates an MZF1–TGF-β1-dependent transformation of mesenchymal stem cells into cancer-associated fibroblasts in breast cancer. Oncogene 2015, 34, e4821. [Google Scholar] [CrossRef]
- Avgustinova, A.; Iravani, M.; Robertson, D.; Fearns, A.; Gao, Q.; Klingbeil, P.; Hanby, A.M.; Speirs, V.; Sahai, E.; Calvo, F.; et al. Tumour cell-derived Wnt7a recruits and activates fibroblasts to promote tumour aggressiveness. Nat. Commun. 2016, 7, e10305. [Google Scholar] [CrossRef] [PubMed]
- Nair, N.; Calle, A.S.; Zahra, M.H.; Prieto-Vila, M.; Oo, A.K.K.; Hurley, L.; Vaidyanath, A.; Seno, A.; Masuda, J.; Iwasaki, Y. A cancer stem cell model as the point of origin of cancer-associated fibroblasts in tumor microenvironment. Sci. Rep. 2017, 7, e6838. [Google Scholar] [CrossRef]
- Heneberg, P. Paracrine tumor signaling induces transdifferentiation of surrounding fibroblasts. Crit. Rev. Oncol. Hematol. 2016, 97, 303–311. [Google Scholar] [CrossRef]
- Ohlund, D.; Handly-Santana, A.; Biffi, G.; Elyada, E.; Almeida, A.S.; Ponz-Sarvise, M.; Corbo, V.; Oni, T.E.; Hearn, S.A.; Lee, E.J.; et al. Distinct populations of inflammatory fibroblasts and myofibroblasts in pancreatic cancer. J. Exp. Med. 2017, 214, 579–596. [Google Scholar] [CrossRef]
- Costa, A.; Kieffer, Y.; Scholer-Dahirel, A.; Pelon, F.; Bourachot, B.; Cardon, M.; Sirven, P.; Magagna, I.; Fuhrmann, L.; Bernard, C. Fibroblast heterogeneity and immunosuppressive environment in human breast cancer. Cancer Cell 2018, 33, 463–479. [Google Scholar] [CrossRef]
- Du, H.; Che, G. Genetic alterations and epigenetic alterations of cancer-associated fibroblasts. Oncol. Lett. 2017, 13, 3–12. [Google Scholar] [CrossRef]
- Raz, Y.; Cohen, N.; Shani, O.; Bell, R.E.; Novitskiy, S.V.; Abramovitz, L.; Levy, C.; Milyavsky, M.; Leider-Trejo, L.; Moses, H.L.; et al. Bone marrow-derived fibroblasts are a functionally distinct stromal cell population in breast cancer. J. Exp. Med. 2018, 215, 3075–3093. [Google Scholar] [CrossRef]
- Biffi, G.; Tuveson, D.A. Diversity and biology of cancer-associated fibroblasts. Physiol. Rev. 2020. [Google Scholar] [CrossRef] [PubMed]
- O’Connell, J.T.; Sugimoto, H.; Cooke, V.G.; MacDonald, B.A.; Mehta, A.I.; LeBleu, V.S.; Dewar, R.; Rocha, R.M.; Brentani, R.R.; Resnick, M.B.; et al. VEGF-A and Tenascin-C produced by S100A4+ stromal cells are important for metastatic colonization. Proc. Natl. Acad. Sci. USA 2011, 108, 16002–16007. [Google Scholar] [CrossRef]
- Perou, C.M.; Sørlie, T.; Eisen, M.B.; Van De Rijn, M.; Jeffrey, S.S.; Rees, C.A.; Pollack, J.R.; Ross, D.T.; Johnsen, H.; Akslen, L.A. Molecular portraits of human breast tumours. Nature 2000, 406, e747. [Google Scholar] [CrossRef] [PubMed]
- Tchou, J.; Kossenkov, A.V.; Chang, L.; Satija, C.; Herlyn, M.; Showe, L.C.; Puré, E. Human breast cancer associated fibroblasts exhibit subtype specific gene expression profiles. BMC Med. Genom. 2012, 5, e39. [Google Scholar] [CrossRef] [PubMed]
- Augsten, M. Cancer-associated fibroblasts as another polarized cell type of the tumor microenvironment. Front. Oncol. 2014, 4, 62. [Google Scholar] [CrossRef] [PubMed]
- Chang, P.-H.; Hwang-Verslues, W.W.; Chang, Y.-C.; Chen, C.-C.; Hsiao, M.; Jeng, Y.-M.; Chang, K.-J.; Lee, E.Y.H.P.; Shew, J.-Y.; Lee, W.-H. Activation of Robo1 signaling of breast cancer cells by Slit2 from stromal fibroblast restrains tumorigenesis via blocking PI3K/Akt/β-catenin pathway. Cancer Res. 2012, 72, 4652–4661. [Google Scholar] [CrossRef] [PubMed]
- Su, S.; Chen, J.; Yao, H.; Liu, J.; Yu, S.; Lao, L.; Wang, M.; Luo, M.; Xing, Y.; Chen, F. CD10+ GPR77+ cancer-associated fibroblasts promote cancer formation and chemoresistance by sustaining cancer stemness. Cell 2018, 172, 841–856.e816. [Google Scholar] [CrossRef]
- Brechbuhl, H.M.; Finlay-Schultz, J.; Yamamoto, T.M.; Gillen, A.E.; Cittelly, D.M.; Tan, A.C.; Sams, S.B.; Pillai, M.M.; Elias, A.D.; Robinson, W.A.; et al. Fibroblast Subtypes Regulate Responsiveness of Luminal Breast Cancer to Estrogen. Clin. Cancer Res. Off. J. Am. Assoc. Cancer Res. 2017, 23, 1710–1721. [Google Scholar] [CrossRef]
- Roswall, P.; Bocci, M.; Bartoschek, M.; Li, H.; Kristiansen, G.; Jansson, S.; Lehn, S.; Sjölund, J.; Reid, S.; Larsson, C.; et al. Microenvironmental control of breast cancer subtype elicited through paracrine platelet-derived growth factor-CC signaling. Nat. Med. 2018, 24, 463–473. [Google Scholar] [CrossRef]
- Martinez-Outschoorn, U.E.; Sotgia, F.; Lisanti, M.P. Caveolae and signalling in cancer. Nat. Rev. Cancer 2015, 15, 225–237. [Google Scholar] [CrossRef]
- Bartoschek, M.; Oskolkov, N.; Bocci, M.; Lovrot, J.; Larsson, C.; Sommarin, M.; Madsen, C.D.; Lindgren, D.; Pekar, G.; Karlsson, G.; et al. Spatially and functionally distinct subclasses of breast cancer-associated fibroblasts revealed by single cell RNA sequencing. Nat. Commun. 2018, 9, e5150. [Google Scholar] [CrossRef] [PubMed]
- Farmer, P.; Bonnefoi, H.; Anderle, P.; Cameron, D.; Wirapati, P.; Becette, V.; André, S.; Piccart, M.; Campone, M.; Brain, E.; et al. A stroma-related gene signature predicts resistance to neoadjuvant chemotherapy in breast cancer. Nat. Med. 2009, 15, e68. [Google Scholar] [CrossRef] [PubMed]
- Valdes-Mora, F.; Salomon, R.; Gloss, B.; Law, A.M.K.; Murphy, K.; Roden, D.L.; Castillo, L.; Colino-Sanguino, Y.; Kikhtyak, Z.; Farbehi, N.; et al. Single-cell RNAseq uncovers involution mimicry as an aberrant development pathway during breast cancer metastasis. bioRxiv 2019. [Google Scholar] [CrossRef]
- Friedman, G.; Levi-Galibov, O.; David, E.; Bornstein, C.; Giladi, A.; Dadiani, M.; Mayo, A.; Halperin, C.; Pevsner-Fischer, M.; Lavon, H.; et al. Cancer-associated fibroblast compositions change with breast cancer progression linking the ratio of S100A4+ and PDPN+ CAFs to clinical outcome. Nat. Cancer 2020, 1, 692–708. [Google Scholar] [CrossRef]
- Bauer, M.; Su, G.; Casper, C.; He, R.; Rehrauer, W.; Friedl, A. Heterogeneity of gene expression in stromal fibroblasts of human breast carcinomas and normal breast. Oncogene 2010, 29, e1732. [Google Scholar] [CrossRef]
- Lamprecht, S.; Sigal-Batikoff, I.; Shany, S.; Abu-Freha, N.; Ling, E.; Delinasios, G.; Moyal-Atias, K.; Delinasios, J.; Fich, A. Teaming up for trouble: Cancer cells, transforming growth factor-β1 signaling and the epigenetic corruption of stromal naïve fibroblasts. Cancers (Basel) 2018, 10, 61. [Google Scholar] [CrossRef]
- Singer, C.F.; Gschwantler-Kaulich, D.; Fink-Retter, A.; Haas, C.; Hudelist, G.; Czerwenka, K.; Kubista, E. Differential gene expression profile in breast cancer-derived stromal fibroblasts. Breast Cancer Res. Treat. 2008, 110, 273–281. [Google Scholar] [CrossRef]
- Martinez-Outschoorn, U.E.; Pavlides, S.; Whitaker-Menezes, D.; Daumer, K.M.; Milliman, J.N.; Chiavarina, B.; Migneco, G.; Witkiewicz, A.K.; Martinez-Cantarin, M.P.; Flomenberg, N.; et al. Tumor cells induce the cancer associated fibroblast phenotype via caveolin-1 degradation: Implications for breast cancer and DCIS therapy with autophagy inhibitors. Cell Cycle (Georget. Tex.) 2010, 9, 2423–2433. [Google Scholar] [CrossRef]
- Sadlonova, A.; Bowe, D.B.; Novak, Z.; Mukherjee, S.; Duncan, V.E.; Page, G.P.; Frost, A.R. Identification of molecular distinctions between normal breast-associated fibroblasts and breast cancer-associated fibroblasts. Cancer Microenviron 2009, 2, 9–21. [Google Scholar] [CrossRef]
- Hu, M.; Yao, J.; Cai, L.; Bachman, K.E.; Van Den Brûle, F.; Velculescu, V.; Polyak, K. Distinct epigenetic changes in the stromal cells of breast cancers. Nat. Genet. 2005, 37, e899. [Google Scholar] [CrossRef]
- Kurose, K.; Gilley, K.; Matsumoto, S.; Watson, P.H.; Zhou, X.-P.; Eng, C. Frequent somatic mutations in PTEN and TP53 are mutually exclusive in the stroma of breast carcinomas. Nat. Genet. 2002, 32, e355. [Google Scholar] [CrossRef] [PubMed]
- Patocs, A.; Zhang, L.; Xu, Y.; Weber, F.; Caldes, T.; Mutter, G.L.; Platzer, P.; Eng, C. Breast-cancer stromal cells with TP53 mutations and nodal metastases. N. Engl. J. Med. 2007, 357, 2543–2551. [Google Scholar] [CrossRef] [PubMed]
- Klymenko, Y.; Nephew, K.P. Epigenetic Crosstalk between the Tumor Microenvironment and Ovarian Cancer Cells: A Therapeutic Road Less Traveled. Cancers (Basel) 2018, 10, 295. [Google Scholar] [CrossRef]
- McCuaig, R.; Wu, F.; Dunn, J.; Rao, S.; Dahlstrom, J.E. The biological and clinical significance of stromal-epithelial interactions in breast cancer. Pathology 2017, 49, 133–140. [Google Scholar] [CrossRef] [PubMed]
- Fiori, M.E.; Di Franco, S.; Villanova, L.; Bianca, P.; Stassi, G.; De Maria, R. Cancer-associated fibroblasts as abettors of tumor progression at the crossroads of EMT and therapy resistance. Mol. Cancer 2019, 18, 70. [Google Scholar] [CrossRef] [PubMed]
- Öhlund, D.; Elyada, E.; Tuveson, D. Fibroblast heterogeneity in the cancer wound. J. Exp. Med. 2014, 211, 1503–1523. [Google Scholar] [CrossRef]
- Yang, X.; Li, Y.; Zou, L.; Zhu, Z. Role of Exosomes in Crosstalk Between Cancer-Associated Fibroblasts and Cancer Cells. Front. Oncol. 2019, 9, 356. [Google Scholar] [CrossRef] [PubMed]
- Hu, J.L.; Wang, W.; Lan, X.L.; Zeng, Z.C.; Liang, Y.S.; Yan, Y.R.; Song, F.Y.; Wang, F.F.; Zhu, X.H.; Liao, W.J.; et al. CAFs secreted exosomes promote metastasis and chemotherapy resistance by enhancing cell stemness and epithelial-mesenchymal transition in colorectal cancer. Mol. Cancer 2019, 18, e91. [Google Scholar] [CrossRef]
- Luan, X.; Sansanaphongpricha, K.; Myers, I.; Chen, H.; Yuan, H.; Sun, D. Engineering exosomes as refined biological nanoplatforms for drug delivery. Acta Pharmacol. Sin. 2017, 38, 754–763. [Google Scholar] [CrossRef]
- Mashouri, L.; Yousefi, H.; Aref, A.R.; Ahadi, A.m.; Molaei, F.; Alahari, S.K. Exosomes: Composition, biogenesis, and mechanisms in cancer metastasis and drug resistance. Mol. Cancer 2019, 18, e75. [Google Scholar] [CrossRef]
- Donnarumma, E.; Fiore, D.; Nappa, M.; Roscigno, G.; Adamo, A.; Iaboni, M.; Russo, V.; Affinito, A.; Puoti, I.; Quintavalle, C. Cancer-associated fibroblasts release exosomal microRNAs that dictate an aggressive phenotype in breast cancer. Oncotarget 2017, 8, e19592. [Google Scholar] [CrossRef] [PubMed]
- Baroni, S.; Romero-Cordoba, S.; Plantamura, I.; Dugo, M.; D’Ippolito, E.; Cataldo, A.; Cosentino, G.; Angeloni, V.; Rossini, A.; Daidone, M. Exosome-mediated delivery of miR-9 induces cancer-associated fibroblast-like properties in human breast fibroblasts. Cell Death Dis. 2016, 7, e2312. [Google Scholar] [CrossRef] [PubMed]
- Ringuette Goulet, C.; Bernard, G.; Tremblay, S.; Chabaud, S.; Bolduc, S.; Pouliot, F. Exosomes Induce Fibroblast Differentiation into Cancer-Associated Fibroblasts through TGFbeta Signaling. Mol. Cancer Res. MCR 2018, 16, 1196–1204. [Google Scholar] [CrossRef]
- Toullec, A.; Gerald, D.; Despouy, G.; Bourachot, B.; Cardon, M.; Lefort, S.; Richardson, M.; Rigaill, G.; Parrini, M.C.; Lucchesi, C.; et al. Oxidative stress promotes myofibroblast differentiation and tumour spreading. EMBO Mol. Med. 2010, 2, 211–230. [Google Scholar] [CrossRef] [PubMed]
- Laitala, A.; Erler, J.T. Hypoxic Signalling in Tumour Stroma. Front. Oncol. 2018, 8, 189. [Google Scholar] [CrossRef] [PubMed]
- Luo, H.; Yang, G.; Yu, T.; Luo, S.; Wu, C.; Sun, Y.; Liu, M.; Tu, G. GPER-mediated proliferation and estradiol production in breast cancer-associated fibroblasts. Endocr. Relat. Cancer 2014, 21, 355–369. [Google Scholar] [CrossRef] [PubMed]
- Ren, J.; Guo, H.; Wu, H.; Tian, T.; Dong, D.; Zhang, Y.; Sui, Y.; Zhang, Y.; Zhao, D.; Wang, S.; et al. GPER in CAFs regulates hypoxia-driven breast cancer invasion in a CTGF-dependent manner. Oncol. Rep. 2015, 33, 1929–1937. [Google Scholar] [CrossRef]
- Sylvestre, M.; Tarte, K.; Roulois, D. Epigenetic mechanisms driving tumor supportive microenvironment differentiation and function: A role in cancer therapy? Epigenomics 2020, 12, 157–169. [Google Scholar] [CrossRef]
- Ren, Y.; Jia, H.-h.; Xu, Y.-q.; Zhou, X.; Zhao, X.-h.; Wang, Y.-f.; Song, X.; Zhu, Z.-y.; Sun, T.; Dou, Y.; et al. Paracrine and epigenetic control of CAF-induced metastasis: The role of HOTAIR stimulated by TGF-ß1 secretion. Mol. Cancer 2018, 17, e5. [Google Scholar] [CrossRef]
- Bai, J.; Xi, Q. Crosstalk between TGF-β signaling and epigenome. Acta Biochim. Biophys. Sin. 2017, 50, 60–67. [Google Scholar] [CrossRef]
- Yu, Y.; Xiao, C.H.; Tan, L.D.; Wang, Q.S.; Li, X.Q.; Feng, Y.M. Cancer-associated fibroblasts induce epithelial-mesenchymal transition of breast cancer cells through paracrine TGF-β signalling. Br. J. Cancer 2014, 110, 724–732. [Google Scholar] [CrossRef] [PubMed]
- Suriyamurthy, S.; Baker, D.; ten Dijke, P.; Iyengar, P.V. Epigenetic Reprogramming of TGF-β Signaling in Breast Cancer. Cancers (Basel) 2019, 11, 726. [Google Scholar] [CrossRef] [PubMed]
- Liu, S.; Chen, S.; Zeng, J. TGF-β signaling: A complex role in tumorigenesis (Review). Mol. Med. Rep. 2018, 17, 699–704. [Google Scholar] [CrossRef] [PubMed]
- Cardenas, H.; Vieth, E.; Lee, J.; Segar, M.; Liu, Y.; Nephew, K.P.; Matei, D. TGF-β induces global changes in DNA methylation during the epithelial-to-mesenchymal transition in ovarian cancer cells. Epigenetics 2014, 9, 1461–1472. [Google Scholar] [CrossRef] [PubMed]
- Barcellos-Hoff, M.H.; Akhurst, R.J. Transforming growth factor-beta in breast cancer: Too much, too late. Breast Cancer Res. 2009, 11, 202. [Google Scholar] [CrossRef] [PubMed]
- Marks, D.L.; Olson, R.L.; Fernandez-Zapico, M.E. Epigenetic control of the tumor microenvironment. Epigenomics 2016, 8, 1671–1687. [Google Scholar] [CrossRef] [PubMed]
- Elsheikh, S.E.; Green, A.R.; Rakha, E.A.; Powe, D.G.; Ahmed, R.A.; Collins, H.M.; Soria, D.; Garibaldi, J.M.; Paish, C.E.; Ammar, A.A.; et al. Global Histone Modifications in Breast Cancer Correlate with Tumor Phenotypes, Prognostic Factors, and Patient Outcome. Cancer Res. 2009, 69, e3802. [Google Scholar] [CrossRef]
- Tyan, S.W.; Hsu, C.H.; Peng, K.L.; Chen, C.C.; Kuo, W.H.; Lee, E.Y.; Shew, J.Y.; Chang, K.J.; Juan, L.J.; Lee, W.H. Breast cancer cells induce stromal fibroblasts to secrete ADAMTS1 for cancer invasion through an epigenetic change. PLoS ONE 2012, 7, e35128. [Google Scholar] [CrossRef]
- Saldanha, S.N.; Tollefsbol, T.O. Chapter 7—Epigenetic Approaches to Cancer Therapy. In Epigenetics in Human Disease, 2nd ed.; Tollefsbol, T.O., Ed.; Academic Press: London, UK, 2018; Volume 6, pp. 219–247. [Google Scholar]
- Zeisberg, E.M.; Zeisberg, M. The role of promoter hypermethylation in fibroblast activation and fibrogenesis. J. Pathol. 2013, 229, 264–273. [Google Scholar] [CrossRef]
- Nicholson, T.B.; Veland, N.; Chen, T. Chapter 3—Writers, Readers, and Erasers of Epigenetic Marks. In Epigenetic Cancer Therapy; Gray, S.G., Ed.; Academic Press: Boston, MA, USA, 2015; pp. 31–66. [Google Scholar] [CrossRef]
- Dey, P. Epigenetic changes in tumor microenvironment. Indian J. Cancer 2011, 48, e507. [Google Scholar] [CrossRef]
- Fiegl, H.; Millinger, S.; Goebel, G.; Müller-Holzner, E.; Marth, C.; Laird, P.W.; Widschwendter, M. Breast Cancer DNA Methylation Profiles in Cancer Cells and Tumor Stroma: Association with HER-2/neu Status in Primary Breast Cancer. Cancer Res. 2006, 66, e29. [Google Scholar] [CrossRef] [PubMed]
- Dai, J.L.; Wang, L.; Sahin, A.A.; Broemeling, L.D.; Schutte, M.; Pan, Y. NHERF (Na+/H+ Exchanger Regulatory Factor) gene mutations in human breast cancer. Oncogene 2004, 23, 8681–8687. [Google Scholar] [CrossRef] [PubMed]
- Calvo, F.; Ranftl, R.; Hooper, S.; Farrugia, A.J.; Moeendarbary, E.; Bruckbauer, A.; Batista, F.; Charras, G.; Sahai, E. Cdc42EP3/BORG2 and Septin Network Enables Mechano-transduction and the Emergence of Cancer-Associated Fibroblasts. Cell Rep. 2015, 13, 2699–2714. [Google Scholar] [CrossRef] [PubMed]
- Farrugia, A.J.; Calvo, F. The Borg family of Cdc42 effector proteins Cdc42EP1–5. Biochem. Soc. Trans. 2016, 44, e1709. [Google Scholar] [CrossRef] [PubMed]
- Onuchic, V.; Hartmaier, R.J.; Boone, D.N.; Samuels, M.L.; Patel, R.Y.; White, W.M.; Garovic, V.D.; Oesterreich, S.; Roth, M.E.; Lee, A.V. Epigenomic deconvolution of breast tumors reveals metabolic coupling between constituent cell types. Cell Rep. 2016, 17, 2075–2086. [Google Scholar] [CrossRef] [PubMed]
- Kim, S.H.; Lee, H.Y.; Jung, S.P.; Kim, S.; Lee, J.E.; Nam, S.J.; Bae, J.W. Role of secreted type I collagen derived from stromal cells in two breast cancer cell lines. Oncol. Lett. 2014, 8, 507–512. [Google Scholar] [CrossRef]
- Zhao, L.; Sun, Y.; Hou, Y.; Peng, Q.; Wang, L.; Luo, H.; Tang, X.; Zeng, Z.; Liu, M. MiRNA expression analysis of cancer-associated fibroblasts and normal fibroblasts in breast cancer. Int. J. Biochem. Cell Biol. 2012, 44, 2051–2059. [Google Scholar] [CrossRef]
- Lee, Y.T.; Tan, Y.J.; Oon, C.E. Molecular targeted therapy: Treating cancer with specificity. Eur. J. Pharmacol. 2018, 834, 188–196. [Google Scholar] [CrossRef]
- Aprelikova, O.; Yu, X.; Palla, J.; Wei, B.-R.; John, S.; Yi, M.; Stephens, R.; Simpson, R.M.; Risinger, J.I.; Jazaeri, A. The role of miR-31 and its target gene SATB2 in cancer-associated fibroblasts. Cell Cycle (Georget. Tex.) 2010, 9, 4387–4398. [Google Scholar] [CrossRef]
- Mitra, A.K.; Zillhardt, M.; Hua, Y.; Tiwari, P.; Murmann, A.E.; Peter, M.E.; Lengyel, E. MicroRNAs reprogram normal fibroblasts into cancer-associated fibroblasts in ovarian cancer. Cancer Discov. 2012, 2, 1100–1108. [Google Scholar] [CrossRef]
- Nouraee, N.; Van Roosbroeck, K.; Vasei, M.; Semnani, S.; Samaei, N.M.; Naghshvar, F.; Omidi, A.A.; Calin, G.A.; Mowla, S.J. Expression, tissue distribution and function of miR-21 in esophageal squamous cell carcinoma. PLoS ONE 2013, 8, e73009. [Google Scholar]
- Josson, S.; Gururajan, M.; Sung, S.; Hu, P.; Shao, C.; Zhau, H.; Liu, C.; Lichterman, J.; Duan, P.; Li, Q. Stromal fibroblast-derived miR-409 promotes epithelial-to-mesenchymal transition and prostate tumorigenesis. Oncogene 2015, 34, e2690. [Google Scholar] [CrossRef] [PubMed]
- Shen, H.; Yu, X.; Yang, F.; Zhang, Z.; Shen, J.; Sun, J.; Choksi, S.; Jitkaew, S.; Shu, Y. Reprogramming of normal fibroblasts into cancer-associated fibroblasts by miRNAs-mediated CCL2/VEGFA signaling. PLoS Genet. 2016, 12, e1006244. [Google Scholar] [CrossRef] [PubMed]
- Tang, X.; Hou, Y.; Yang, G.; Wang, X.; Tang, S.; Du, Y.; Yang, L.; Yu, T.; Zhang, H.; Zhou, M. Stromal miR-200s contribute to breast cancer cell invasion through CAF activation and ECM remodeling. Cell Death Differ. 2016, 23, e132. [Google Scholar] [CrossRef] [PubMed]
- Al-Ansari, M.M.; Aboussekhra, A. miR-146b-5p mediates p16-dependent repression of IL-6 and suppresses paracrine procarcinogenic effects of breast stromal fibroblasts. Oncotarget 2015, 6, 30006–30016. [Google Scholar] [CrossRef] [PubMed]
- Bronisz, A.; Godlewski, J.; Wallace, J.; Merchant, A.S.; Nowicki, M.O.; Mathsyaraja, H.; Srinivasan, R.; Trimboli, A.J.; Martin, C.K.; Li, F. Reprogramming of the tumour microenvironment by stromal PTEN-regulated miR-320. Nat. Cell Biol. 2012, 14, e159. [Google Scholar] [CrossRef] [PubMed]
- Rask, L.; Balslev, E.; Jørgensen, S.; Eriksen, J.; Flyger, H.; Møller, S.; Høgdall, E.; Litman, T.; Schnack Nielsen, B. High expression of miR-21 in tumor stroma correlates with increased cancer cell proliferation in human breast cancer. Apmis 2011, 119, 663–673. [Google Scholar] [CrossRef]
- Verghese, E.T.; Drury, R.; Green, C.A.; Holliday, D.L.; Lu, X.; Nash, C.; Speirs, V.; Thorne, J.L.; Thygesen, H.H.; Zougman, A. MiR-26b is down-regulated in carcinoma-associated fibroblasts from ER-positive breast cancers leading to enhanced cell migration and invasion. J. Pathol. 2013, 231, 388–399. [Google Scholar] [CrossRef]
- Shah, S.H.; Miller, P.; Garcia-Contreras, M.; Ao, Z.; Machlin, L.; Issa, E.; El-Ashry, D. Hierarchical paracrine interaction of breast cancer associated fibroblasts with cancer cells via hMAPK-microRNAs to drive ER-negative breast cancer phenotype. Cancer Biol. Ther. 2015, 16, 1671–1681. [Google Scholar] [CrossRef]
- Santolla, M.F.; Lappano, R.; Cirillo, F.; Rigiracciolo, D.C.; Sebastiani, A.; Abonante, S.; Tassone, P.; Tagliaferri, P.; Di Martino, M.T.; Maggiolini, M. miR-221 stimulates breast cancer cells and cancer-associated fibroblasts (CAFs) through selective interference with the A20/c-Rel/CTGF signaling. J. Exp. Clin. Cancer Res. 2018, 37, e94. [Google Scholar] [CrossRef]
- DeFilippis, R.A.; Chang, H.; Dumont, N.; Rabban, J.T.; Chen, Y.-Y.; Fontenay, G.V.; Berman, H.K.; Gauthier, M.L.; Zhao, J.; Hu, D. CD36 repression activates a multicellular stromal program shared by high mammographic density and tumor tissues. Cancer Discov. 2012, 2, 826–839. [Google Scholar] [CrossRef] [PubMed]
- Smith, B.; Agarwal, P.; Bhowmick, N.A. MicroRNA applications for prostate, ovarian and breast cancer in the era of precision medicine. Endocr. Relat. Cancer 2017, 24, R157–R172. [Google Scholar] [CrossRef]
- Chiappinelli, K.B.; Zahnow, C.A.; Ahuja, N.; Baylin, S.B. Combining Epigenetic and Immunotherapy to Combat Cancer. Cancer Res. 2016, 76, 1683–1689. [Google Scholar] [CrossRef]
- Falahi, F.; van Kruchten, M.; Martinet, N.; Hospers, G.A.P.; Rots, M.G. Current and upcoming approaches to exploit the reversibility of epigenetic mutations in breast cancer. Breast Cancer Res. 2014, 16, e412. [Google Scholar] [CrossRef] [PubMed]
- Strauss, J.; Figg, W.D. Using Epigenetic Therapy to Overcome Chemotherapy Resistance. Anticancer Res. 2016, 36, 1–4. [Google Scholar] [PubMed]
- Biffi, G.; Tuveson, D.A. Deciphering cancer fibroblasts. J. Exp. Med. 2018, 215, 2967–2968. [Google Scholar] [CrossRef] [PubMed]
- Banerjee, S.; Modi, S.; McGinn, O.; Zhao, X.; Dudeja, V.; Ramakrishnan, S.; Saluja, A.K. Impaired Synthesis of Stromal Components in Response to Minnelide Improves Vascular Function, Drug Delivery, and Survival in Pancreatic Cancer. Clin. Cancer Res. Off. J. Am. Assoc. Cancer Res. 2016, 22, 415–425. [Google Scholar] [CrossRef]
- Sherman, M.H.; Yu, R.T.; Engle, D.D.; Ding, N.; Atkins, A.R.; Tiriac, H.; Collisson, E.A.; Connor, F.; Van Dyke, T.; Kozlov, S.; et al. Vitamin D Receptor-Mediated Stromal Reprogramming Suppresses Pancreatitis and Enhances Pancreatic Cancer Therapy. Cell 2014, 159, 80–93. [Google Scholar] [CrossRef]
- Froeling, F.E.; Feig, C.; Chelala, C.; Dobson, R.; Mein, C.E.; Tuveson, D.A.; Clevers, H.; Hart, I.R.; Kocher, H.M. Retinoic acid-induced pancreatic stellate cell quiescence reduces paracrine Wnt-β-catenin signaling to slow tumor progression. Gastroenterology 2011, 141, 1486–1497. [Google Scholar] [CrossRef]
- Stahl, M.; Kohrman, N.; Gore, S.D.; Kim, T.K.; Zeidan, A.M.; Prebet, T. Epigenetics in cancer: A hematological perspective. PLoS Genet. 2016, 12, e1006193. [Google Scholar]
- Stresemann, C.; Brueckner, B.; Musch, T.; Stopper, H.; Lyko, F. Functional diversity of DNA methyltransferase inhibitors in human cancer cell lines. Cancer Res. 2006, 66, 2794–2800. [Google Scholar] [CrossRef]
- Sheikh, S.; Bekheet, M.; Olzscha, H.; La Thangue, N.B. Predicting and monitoring responses to epigenetic drugs. In Drug Discovery in Cancer Epigenetics; Elsevier: Oxford, UK, 2016; pp. 373–406. [Google Scholar]
- Cheng, Y.; He, C.; Wang, M.; Ma, X.; Mo, F.; Yang, S.; Han, J.; Wei, X. Targeting epigenetic regulators for cancer therapy: Mechanisms and advances in clinical trials. Signal Transduct. Target. Ther. 2019, 4, e62. [Google Scholar] [CrossRef]
- An, Y.; Kiang, A.; Lopez, J.P.; Kuo, S.Z.; Yu, M.A.; Abhold, E.L.; Chen, J.S.; Wang-Rodriguez, J.; Ongkeko, W.M. Cigarette smoke promotes drug resistance and expansion of cancer stem cell-like side population. PLoS ONE 2012, 7, e47919. [Google Scholar] [CrossRef] [PubMed]
- Liang, G.; Gonzales, F.A.; Jones, P.A.; Orntoft, T.F.; Thykjaer, T. Analysis of gene induction in human fibroblasts and bladder cancer cells exposed to the methylation inhibitor 5-aza-2′-deoxycytidine. Cancer Res. 2002, 62, 961–966. [Google Scholar] [PubMed]
- Jiang, L.; Gonda, T.A.; Gamble, M.V.; Salas, M.; Seshan, V.; Tu, S.; Twaddell, W.S.; Hegyi, P.; Lazar, G.; Steele, I.; et al. Global hypomethylation of genomic DNA in cancer-associated myofibroblasts. Cancer Res. 2008, 68, 9900–9908. [Google Scholar] [CrossRef]
- Carr, B.I.; Reilly, J.G.; Smith, S.S.; Winberg, C.; Riggs, A. The tumorigenicity of 5-azacytidine in the male Fischer rat. Carcinogenesis 1984, 5, 1583–1590. [Google Scholar] [CrossRef]
- Chik, F.; Szyf, M. Effects of specific DNMT gene depletion on cancer cell transformation and breast cancer cell invasion; toward selective DNMT inhibitors. Carcinogenesis 2011, 32, 224–232. [Google Scholar] [CrossRef] [PubMed]
- Jackson-Grusby, L.; Beard, C.; Possemato, R.; Tudor, M.; Fambrough, D.; Csankovszki, G.; Dausman, J.; Lee, P.; Wilson, C.; Lander, E.; et al. Loss of genomic methylation causes p53-dependent apoptosis and epigenetic deregulation. Nat. Genet. 2001, 27, 31–39. [Google Scholar] [CrossRef] [PubMed]
- Yoshida, G.J.; Azuma, A.; Miura, Y.; Orimo, A. Activated Fibroblast Program Orchestrates Tumor Initiation and Progression; Molecular Mechanisms and the Associated Therapeutic Strategies. Int. J. Mol. Sci. 2019, 20, 2256. [Google Scholar] [CrossRef] [PubMed]
- Pazolli, E.; Alspach, E.; Milczarek, A.; Prior, J.; Piwnica-Worms, D.; Stewart, S.A. Chromatin remodeling underlies the senescence-associated secretory phenotype of tumor stromal fibroblasts that supports cancer progression. Cancer Res. 2012, 72, 2251–2261. [Google Scholar] [CrossRef] [PubMed]
- Nguyen, A.H.; Elliott, I.A.; Wu, N.; Matsumura, C.; Vogelauer, M.; Attar, N.; Dann, A.; Ghukasyan, R.; Toste, P.A.; Patel, S.G.; et al. Histone deacetylase inhibitors provoke a tumor supportive phenotype in pancreatic cancer associated fibroblasts. Oncotarget 2017, 8, 19074–19088. [Google Scholar] [CrossRef] [PubMed]
- Kim, D.J.; Dunleavey, J.M.; Xiao, L.; Ollila, D.W.; Troester, M.A.; Otey, C.A.; Li, W.; Barker, T.H.; Dudley, A.C. Suppression of TGFβ-mediated conversion of endothelial cells and fibroblasts into cancer associated (myo)fibroblasts via HDAC inhibition. Br. J. Cancer 2018, 118, 1359–1368. [Google Scholar] [CrossRef] [PubMed]
- Li, A.; Chen, P.; Leng, Y.; Kang, J. Histone deacetylase 6 regulates the immunosuppressive properties of cancer-associated fibroblasts in breast cancer through the STAT3-COX2-dependent pathway. Oncogene 2018, 37, 5952–5966. [Google Scholar] [CrossRef] [PubMed]
- Knutson, S.K.; Kawano, S.; Minoshima, Y.; Warholic, N.M.; Huang, K.-C.; Xiao, Y.; Kadowaki, T.; Uesugi, M.; Kuznetsov, G.; Kumar, N. Selective inhibition of EZH2 by EPZ-6438 leads to potent antitumor activity in EZH2-mutant non-Hodgkin lymphoma. Mol. Cancer Ther. 2014, 13, 842–854. [Google Scholar] [CrossRef]
- White, J.C.; Pucci, P.; Crea, F. The role of histone lysine demethylases in cancer cells’ resistance to tyrosine kinase inhibitors. Cancer Drug Resist. 2019, 2, 326–334. [Google Scholar] [CrossRef]
- Munster, P.N.; Thurn, K.T.; Thomas, S.; Raha, P.; Lacevic, M.; Miller, A.; Melisko, M.; Ismail-Khan, R.; Rugo, H.; Moasser, M.; et al. A phase II study of the histone deacetylase inhibitor vorinostat combined with tamoxifen for the treatment of patients with hormone therapy-resistant breast cancer. Br. J. Cancer 2011, 104, 1828–1835. [Google Scholar] [CrossRef]
- Yardley, D.A.; Ismail-Khan, R.R.; Melichar, B.; Lichinitser, M.; Munster, P.N.; Klein, P.M.; Cruickshank, S.; Miller, K.D.; Lee, M.J.; Trepel, J.B. Randomized phase II, double-blind, placebo-controlled study of exemestane with or without entinostat in postmenopausal women with locally recurrent or metastatic estrogen receptor-positive breast cancer progressing on treatment with a nonsteroidal aromatase inhibitor. J. Clin. Oncol. 2013, 31, 2128–2135. [Google Scholar] [CrossRef]
- Ramaswamy, B.; Fiskus, W.; Cohen, B.; Pellegrino, C.; Hershman, D.L.; Chuang, E.; Luu, T.; Somlo, G.; Goetz, M.; Swaby, R.; et al. Phase I-II study of vorinostat plus paclitaxel and bevacizumab in metastatic breast cancer: Evidence for vorinostat-induced tubulin acetylation and Hsp90 inhibition in vivo. Breast Cancer Res. Treat. 2012, 132, 1063–1072. [Google Scholar] [CrossRef]
- Stearns, V.; Jacobs, L.K.; Fackler, M.; Tsangaris, T.N.; Rudek, M.A.; Higgins, M.; Lange, J.; Cheng, Z.; Slater, S.A.; Jeter, S.C.; et al. Biomarker modulation following short-term vorinostat in women with newly diagnosed primary breast cancer. Clin. Cancer Res. Off. J. Am. Assoc. Cancer Res. 2013, 19, 4008–4016. [Google Scholar] [CrossRef]
- Walters, R.S.; Theriault, R.L.; Holmes, F.A.; Hortobagyi, G.N.; Esparza, L. Phase II trial of fazarabine (ARA-AC, arabinosyl-5-azacytosine) in metastatic breast cancer. Investig. New Drugs 1992, 10, 43–44. [Google Scholar] [CrossRef]
- Fridrichova, I.; Zmetakova, I. MicroRNAs Contribute to Breast Cancer Invasiveness. Cells 2019, 8, 1361. [Google Scholar] [CrossRef] [PubMed]
- Al-Harbi, B.; Hendrayani, S.F.; Silva, G.; Aboussekhra, A. Let-7b inhibits cancer-promoting effects of breast cancer-associated fibroblasts through IL-8 repression. Oncotarget 2018, 9, 17825–17838. [Google Scholar] [CrossRef] [PubMed]
- Hendrayani, S.F.; Al-Khalaf, H.H.; Aboussekhra, A. Curcumin triggers p16-dependent senescence in active breast cancer-associated fibroblasts and suppresses their paracrine procarcinogenic effects. Neoplasia (N. Y.) 2013, 15, 631–640. [Google Scholar] [CrossRef] [PubMed]
- Yang, F.; Ning, Z.; Ma, L.; Liu, W.; Shao, C.; Shu, Y.; Shen, H. Exosomal miRNAs and miRNA dysregulation in cancer-associated fibroblasts. Mol. Cancer 2017, 16, e148. [Google Scholar] [CrossRef]
- Barata, P.; Sood, A.K.; Hong, D.S. RNA-targeted therapeutics in cancer clinical trials: Current status and future directions. Cancer Treat. Rev. 2016, 50, 35–47. [Google Scholar] [CrossRef]
- Kuninty, P.R.; Schnittert, J.; Storm, G.; Prakash, J. MicroRNA Targeting to Modulate Tumor Microenvironment. Front. Oncol. 2016, 6, e3. [Google Scholar] [CrossRef] [PubMed]
- Mueller, M.M.; Fusenig, N.E. Friends or foes—Bipolar effects of the tumour stroma in cancer. Nat. Rev. Cancer 2004, 4, e839. [Google Scholar] [CrossRef]
- Lonning, S.; Mannick, J.; M McPherson, J. Antibody Targeting of TGF-β in Cancer Patients. Curr. Pharm. Biotechnol. 2011, 12, 2176–2189. [Google Scholar] [CrossRef]
- Holmgaard, R.B.; Schaer, D.A.; Li, Y.; Castaneda, S.P.; Murphy, M.Y.; Xu, X.; Inigo, I.; Dobkin, J.; Manro, J.R.; Iversen, P.W.; et al. Targeting the TGFβ pathway with galunisertib, a TGFβRI small molecule inhibitor, promotes anti-tumor immunity leading to durable, complete responses, as monotherapy and in combination with checkpoint blockade. J. Immunother. Cancer 2018, 6, 47. [Google Scholar] [CrossRef]
- Rai, A.; Greening, D.W.; Chen, M.; Xu, R.; Ji, H.; Simpson, R.J. Exosomes Derived from Human Primary and Metastatic Colorectal Cancer Cells Contribute to Functional Heterogeneity of Activated Fibroblasts by Reprogramming Their Proteome. Proteomics 2019, 19, e1800148. [Google Scholar] [CrossRef]
- Rhim, A.D.; Oberstein, P.E.; Thomas, D.H.; Mirek, E.T.; Palermo, C.F.; Sastra, S.A.; Dekleva, E.N.; Saunders, T.; Becerra, C.P.; Tattersall, I.W.; et al. Stromal elements act to restrain, rather than support, pancreatic ductal adenocarcinoma. Cancer Cell 2014, 25, 735–747. [Google Scholar] [CrossRef] [PubMed]
- Flaberg, E.; Markasz, L.; Petranyi, G.; Stuber, G.; Dicső, F.; Alchihabi, N.; Oláh, È.; Csízy, I.; Józsa, T.; Andrén, O.; et al. High-throughput live-cell imaging reveals differential inhibition of tumor cell proliferation by human fibroblasts. Int. J. Cancer 2011, 128, 2793–2802. [Google Scholar] [CrossRef] [PubMed]
- Zhou, L.; Yang, K.; Randall Wickett, R.; Zhang, Y. Dermal fibroblasts induce cell cycle arrest and block epithelial-mesenchymal transition to inhibit the early stage melanoma development. Cancer Med. 2016, 5, 1566–1579. [Google Scholar] [CrossRef] [PubMed]
- Bozóky, B.; Savchenko, A.; Csermely, P.; Korcsmáros, T.; Dúl, Z.; Pontén, F.; Székely, L.; Klein, G. Novel signatures of cancer-associated fibroblasts. Int. J. Cancer 2013, 133, 286–293. [Google Scholar] [CrossRef] [PubMed]
- Gieniec, K.A.; Butler, L.M.; Worthley, D.L.; Woods, S.L. Cancer-associated fibroblasts-heroes or villains? Br. J. Cancer 2019, 121, 293–302. [Google Scholar] [CrossRef] [PubMed]
- Choi, S.-Y.; Sung, R.; Lee, S.-J.; Lee, T.-G.; Kim, N.; Yoon, S.M.; Lee, E.J.; Chae, H.B.; Youn, S.J.; Park, S.M. Podoplanin, α-smooth muscle actin or S100A4 expressing cancer-associated fibroblasts are associated with different prognosis in colorectal cancers. J. Korean Med. Sci. 2013, 28, 1293–1301. [Google Scholar] [CrossRef]
- Habib, N.; Avraham-Davidi, I.; Basu, A.; Burks, T.; Shekhar, K.; Hofree, M.; Choudhury, S.R.; Aguet, F.; Gelfand, E.; Ardlie, K.; et al. Massively parallel single-nucleus RNA-seq with DroNc-seq. Nat. Methods 2017, 14, 955–958. [Google Scholar] [CrossRef]
| Subpopulation (s) | Functionality | Marker (s) | Ref. |
|---|---|---|---|
| CAF-S1 | Immunosuppression | FAP, αSMA, CXCL12, 1L6 | [38,41] |
| CAF-S2 | N/A | Low to negative expression of all biomarkers | |
| CAF-S3 | N/A | PDGFRβ | |
| CAF-S4 | Contractile signature, oxidative metabolism | αSMA, CD29 | |
| vCAFs | Vascular development and angiogenesis | NID2, CD31, αSMA, PDGFRβ | [41,51] |
| mCAFs | Provider of extracellular matrix and EMT | PDGFRα, FBLN1 | |
| cCAFs | Engage in cell cycle and cell division, angiogenesis | Ki-67, PDGFRβ | |
| dCAFs | Cell differentiation and development | SCRG1, SOX9 | |
| ECM-CAFs | EMT, ECM interaction and tissue remodelling | TNC | [41,53] |
| Inflammatory CAFs | Immunomodulation, monocyte recruitment | Ly6C, C3, CXCL12, PDGFRα | |
| Contractile myofibroblasts | Actin cytoskeleton organisation | αSMA, MYLK | |
| sCAFs | protein-folding and metabolic regulation | S100A4 | [54] |
| pCAFs | immune regulation, cell migration, wound healing | PDPN |
| Protein/Genes | Regulation | Ref. |
|---|---|---|
| Growth Factors and Cytokines | Up | [57] |
| Granulocyte-macrophage colony-stimulating factor receptor (GMCSFR), Pigment epithelium-differentiation factor (PEDF), Kirsten rat sarcoma viral oncogene homolog protein (K-RAS), Effector cell protease receptor 1 (EPR-1), Hepatoma transmembrane kinase ligand (HTK ligand), Alpha-2-macroglobulin receptor-associated protein (RAP) | ||
| Cell-Cell Interaction, Communication and Cell Structure Maintenance | Up | |
| Osteopontin (OPN), Transmembrane 4 superfamily protein (SAS), Tapasin (NGS-17), Troponin I fast-twitch isoform (TNNI2) | ||
| Proteolytic Enzymes Regulation | Up | |
| Dihydrodiol dehydrogenase (DHDH), Protein tyrosine phosphatase (PTP), Hematopoietic consensus tyrosine-lacking kinase (HYL) Pro-cathepsin L (zymogen of proteolytic enzyme cathepsin L), Beta-D-galactosidase (β-gal), Tryptophanyl tRNA synthetase (IFNWRS), Rolipram-sensitive 30, 50-cyclic AMP phosphodiesterase | ||
| Cell Cycle and Metabolism | Up | |
| Guanine nucleotide-binding protein G(q) subunit alpha (GNAQ), Circadian locomoter output cycles protein kaput (CLOCK), 63-kDa protein kinase related to rat ERK3; also known as mitogen-activated protein kinase 4 (MAPK4), Phospholipase A2 (RASF-A PLA2), Pescadillo homolog protein (PES1), Cell division cycle protein 2 (CDC2), Glutamine-dependent asparagine synthetase (Ts11), Calnexin (CNX) | ||
| Cell Paracrine and Intracellular Signalling | Up | [55] |
| Solute carrier family 24 member 3 (SLC24A3), Insulin like growth factor binding protein 2 (IGFBP2), Tumor necrosis factor ligand superfamily member 4 (TNFSF4), Ras protein specific guanine nucleotide releasing factor 2 (RASGRF2), Polypeptide N-acetylgalactosaminyltransferase 3 (GALNT3), Gap junction protein alpha 5 (GJA5), WNT1 inducible signalling pathway protein 1 (WISP1), V-Ets erythroblastosis virus E26 oncogene homolog (ERG), V-Yes-1 Yamaguchi sarcoma viral related oncogene homolog (LYN) | ||
| Cell Adhesion and Migration | Up | |
| Transforming growth factor beta 2 (TGF-β2), ST6 N-acetylgalactosaminide alpha-2,6-sialyltransferase 5 (ST6GALNAC5), Collagen type X alpha 1 chain (COL10A1), Synaptopodin (SYNPO), Heparin binding EGF like growth factor (HBEGF) | ||
| ECM Remodelling | Up | [59] |
| Plasminogen activator inhibitor 2 (PAI2), Tissue plasminogen activator (PLAT), Matrix metallopeptidase 1 (MMP1), Dickkopf WNT signalling pathway inhibitor 1 (DKK1), Neuregulin 1 (NRG1), Tissue factor pathway inhibitor 2 (TFP12) | ||
| Steroid Hormone Metabolism | Down | [55,57] |
| Aldo-keto reductase family 1 member C1 (AKR1C1), Aldo-keto reductase family 1 member C2 (AKR1C2), Phosphatidic acid phosphatase type 2B (PPAP2B), Growth hormone regulated TBC protein 1 (GRTP1) | ||
| Cell Adhesion and Migration | Down | [55,57] |
| Slit homolog 3 protein (SLIT3), Osteoblast-specific factor 2 (OSF2), OB-cadherin-1 (CDH11), Cytokine-inducible nuclear protein | ||
| Tumour Suppressor Genes | Down | [20,55,58] |
| Tumour protein P53 (TP53), Cyclin-dependent kinase inhibitor 1A (CDKN1A or P21), Caveolin 1 (CAV1), Phosphatase and tensin homolog (PTEN), Epithelial membrane protein 1 (EMP1) |
| microRNA(s) | Regulation Status in BCAFs | Effects on Breast Tumorigenesis | Ref. |
|---|---|---|---|
| miR-320 | Down | Increased ETS2 gene expression resulting in enhanced BC cell invasion and angiogenesis in vitro | [108] |
| miR-26b, miR-101, miR-141, miR-200b, miR-200c, miR-205, miR-342-3p, let-7g | Down | Enhanced TGF-β signalling in clinically derived BCAFs leading to EMT and BC cell invasion | [99] |
| miR-31-3p, miR-221-3p, miR-221-5p | Up | Enhanced TGF-β signalling in clinically derived BCAFs leading to EMT and BC cell invasion | [99] |
| miR-31 | Down | Increased SATB2 gene expression resulting in BCAFs invasion and BC cell migration | [39,101] |
| miR-9 | Up | Enhanced BC cell migration, invasion and motility in vitro | [72] |
| miR-21 | Up | Increased Ki-67 protein expression and BC cell proliferation in clinical tumour samples. | [109] |
| miR-26b | Down | Upregulated IL-6 signalling pathway leadingto enhanced BC cell migration | [110] |
| miR-221, miR-222 | Up | Repressed ER expression which led to lower recurrence-free period and overall survival in BC patients Reduced A20 (aka. TNFAIP3) gene expression subsequently inhibits NF-κB activity resulting in promotion BCAFs and BC cell growth and migration | [111,112] |
| miR-155 | Up | Repressed CD36 in stroma cells leading to the promotion of proliferation, migration and angiogenesis in BC cells | [113] |
| miR-133b | Up | Promoted BC cell growth through inhibition of apoptosis | [114] |
| miR-148a | Down | Promoted BC cell migration in vitro | [39] |
| miR-21, miR-143, miR-378e | Up | Promoted stemness, EMT, anchorage-independent cell growth, and invasive capacity of BC cells in vitro | [71] |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Lee, Y.T.; Tan, Y.J.; Falasca, M.; Oon, C.E. Cancer-Associated Fibroblasts: Epigenetic Regulation and Therapeutic Intervention in Breast Cancer. Cancers 2020, 12, 2949. https://doi.org/10.3390/cancers12102949
Lee YT, Tan YJ, Falasca M, Oon CE. Cancer-Associated Fibroblasts: Epigenetic Regulation and Therapeutic Intervention in Breast Cancer. Cancers. 2020; 12(10):2949. https://doi.org/10.3390/cancers12102949
Chicago/Turabian StyleLee, Yeuan Ting, Yi Jer Tan, Marco Falasca, and Chern Ein Oon. 2020. "Cancer-Associated Fibroblasts: Epigenetic Regulation and Therapeutic Intervention in Breast Cancer" Cancers 12, no. 10: 2949. https://doi.org/10.3390/cancers12102949
APA StyleLee, Y. T., Tan, Y. J., Falasca, M., & Oon, C. E. (2020). Cancer-Associated Fibroblasts: Epigenetic Regulation and Therapeutic Intervention in Breast Cancer. Cancers, 12(10), 2949. https://doi.org/10.3390/cancers12102949







