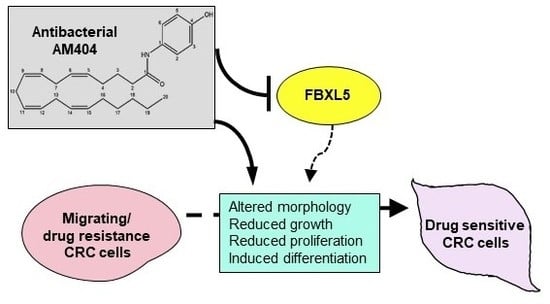
Repurposing Antibacterial AM404 As a Potential Anticancer Drug for Targeting Colorectal Cancer Stem-Like Cells
Abstract
1. Introduction
2. Results
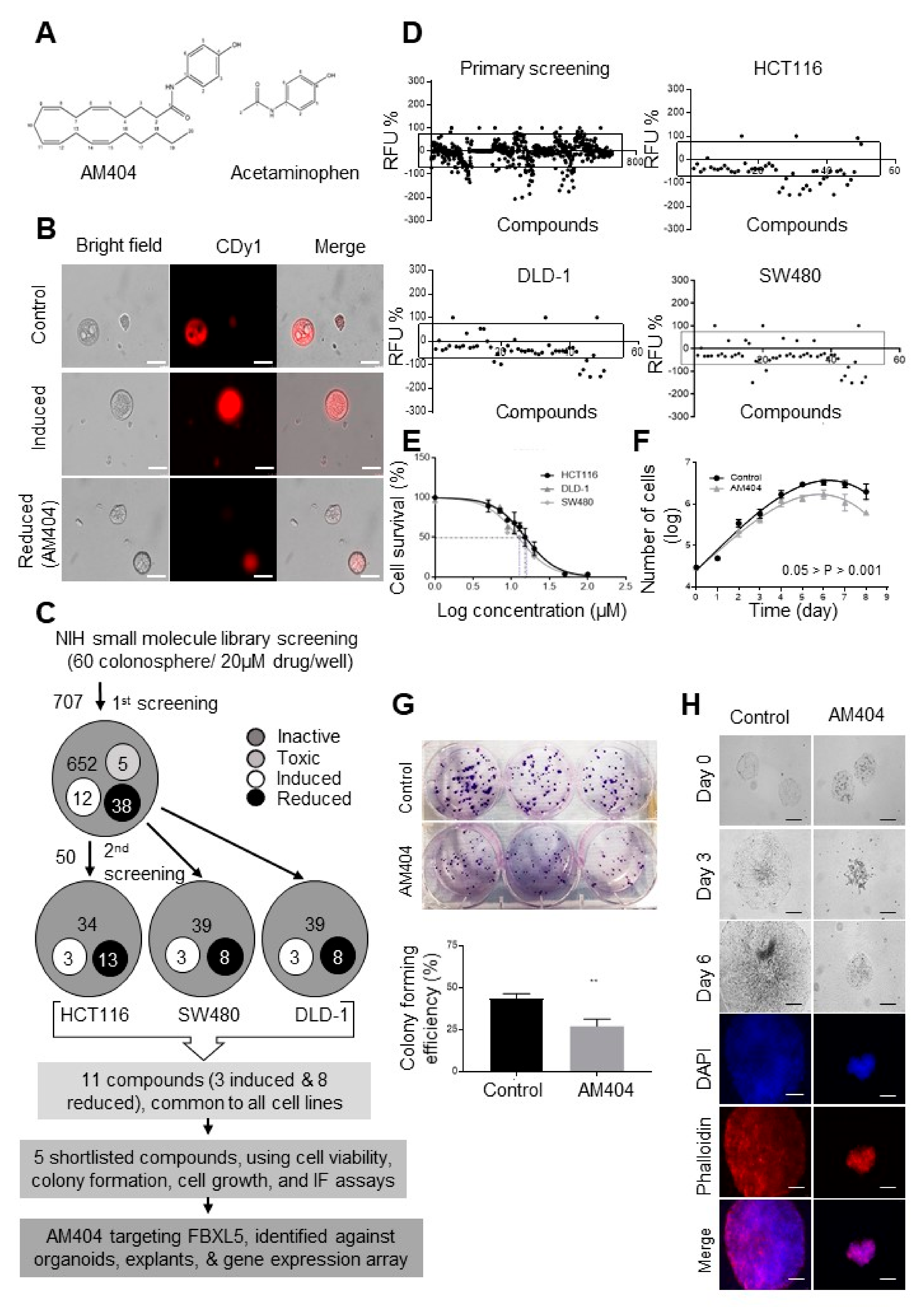
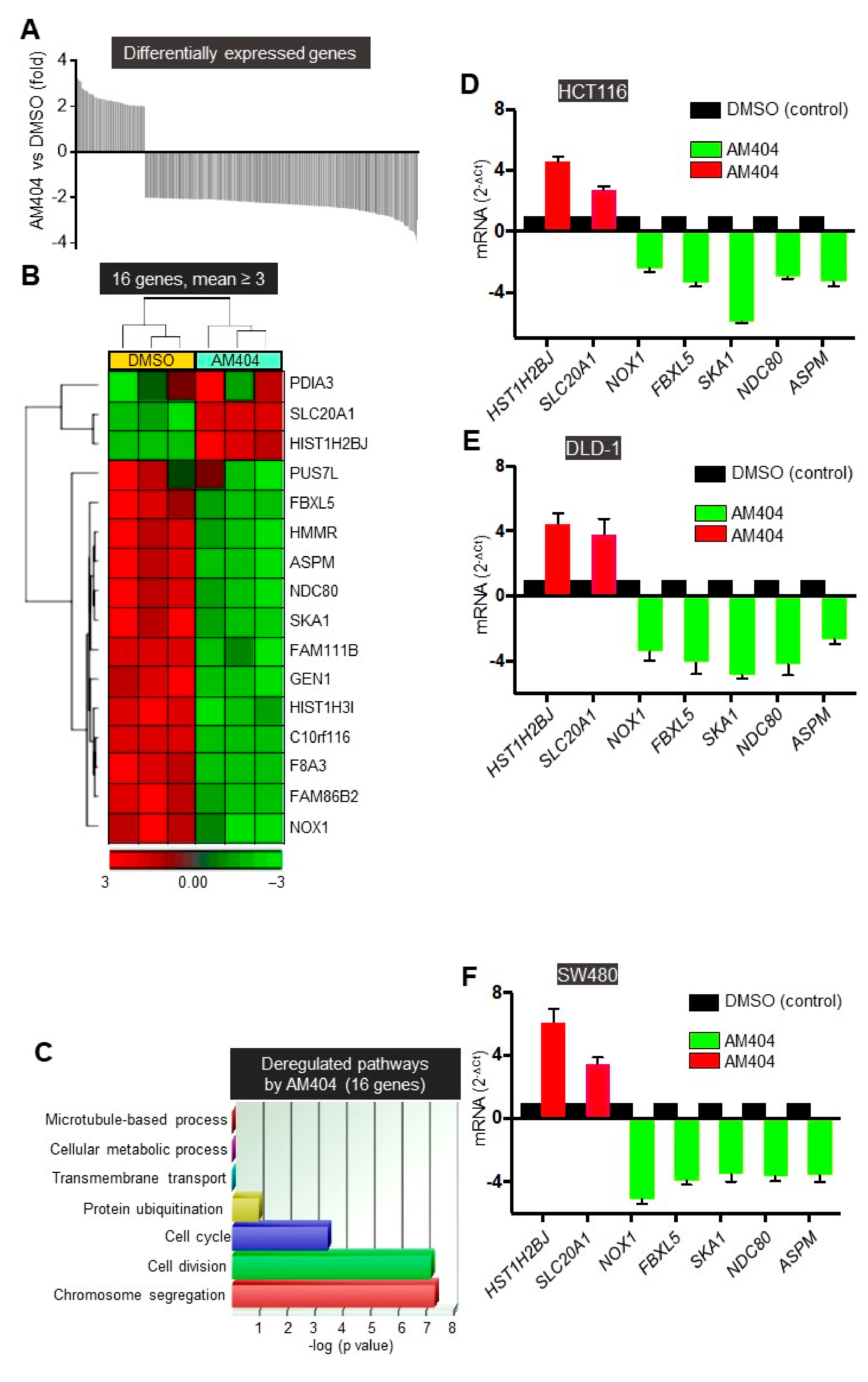
2.1. A Screen of the NIH Clinical Collection Small Molecule Library Identifies Potential Anti-Cancer Drug AM404
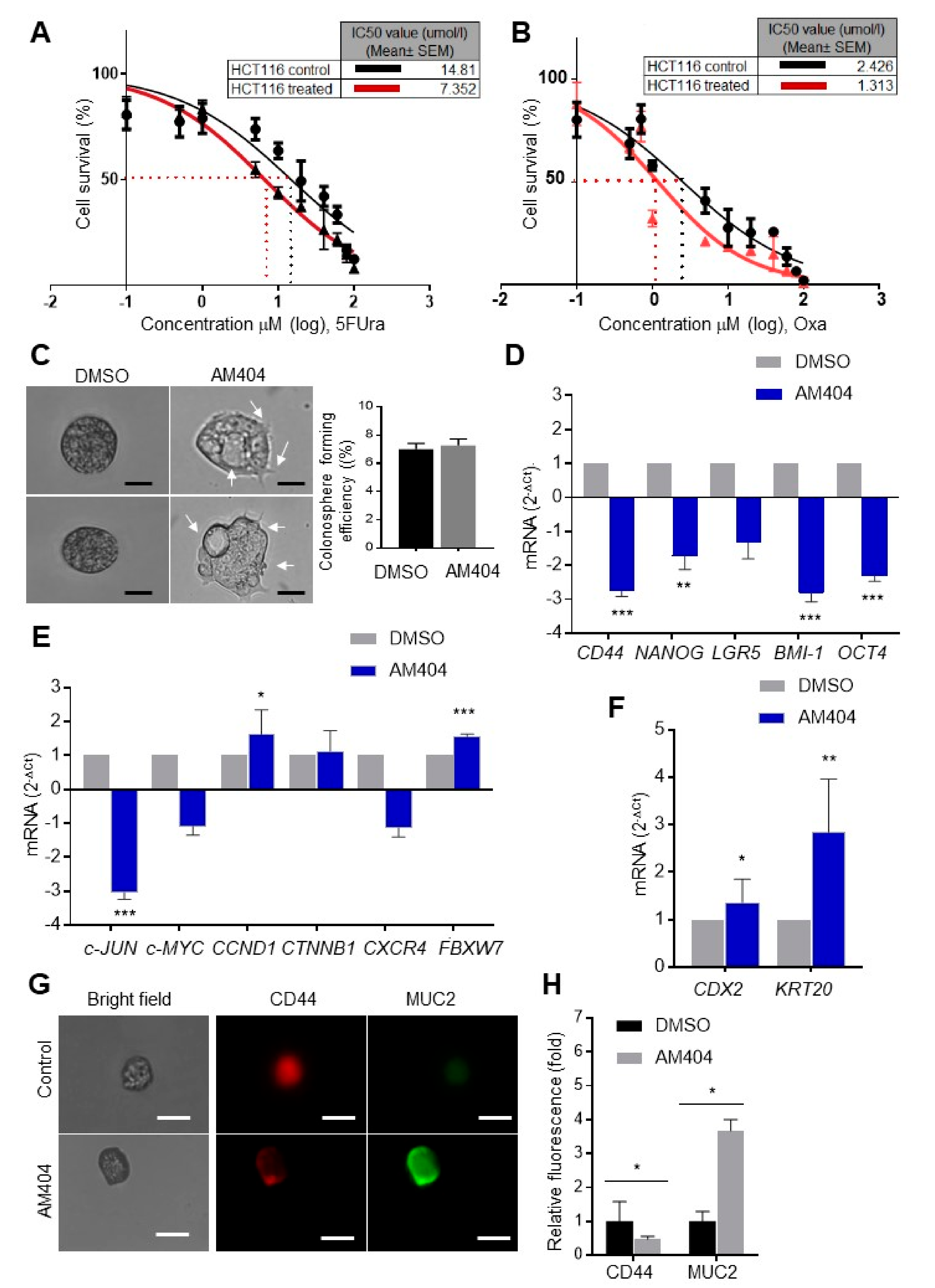
2.2. AM404 Inhibits De-Differentiation and Acquisition of Stem-Like Properties
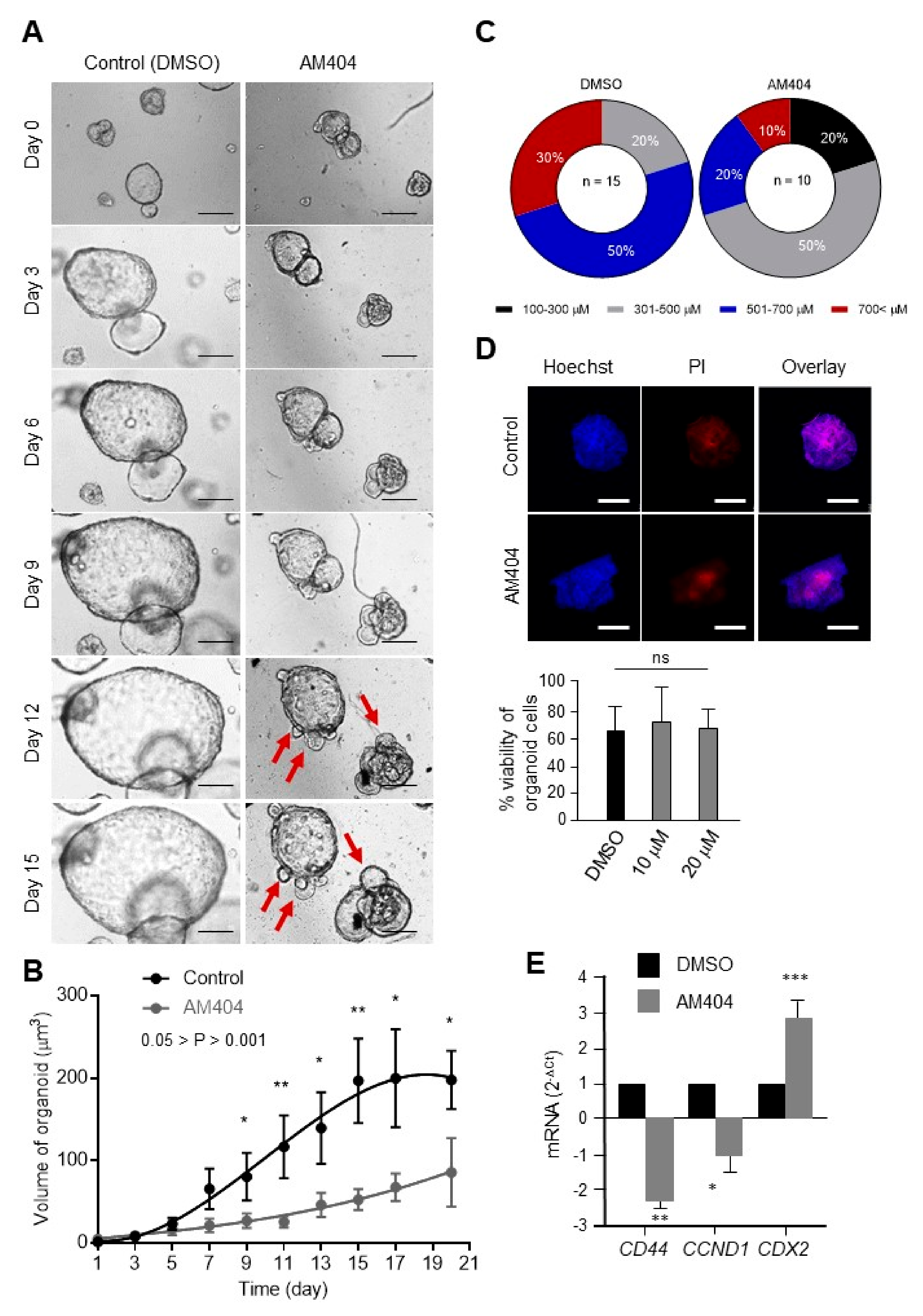
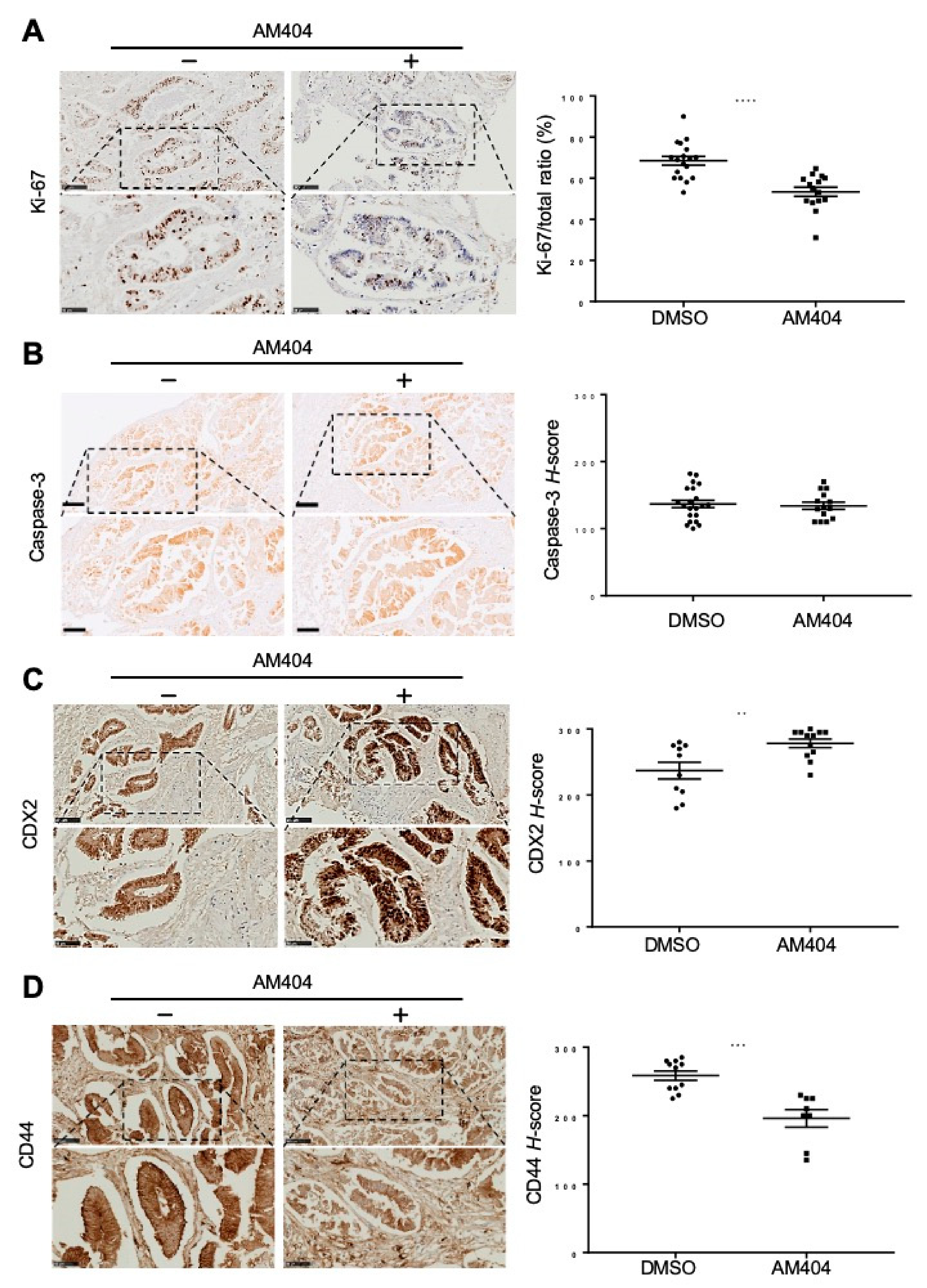
2.3. Ex Vivo Treatment of CRC Patient Biopsies Evaluates AM404 Response As an Anticancer Drug
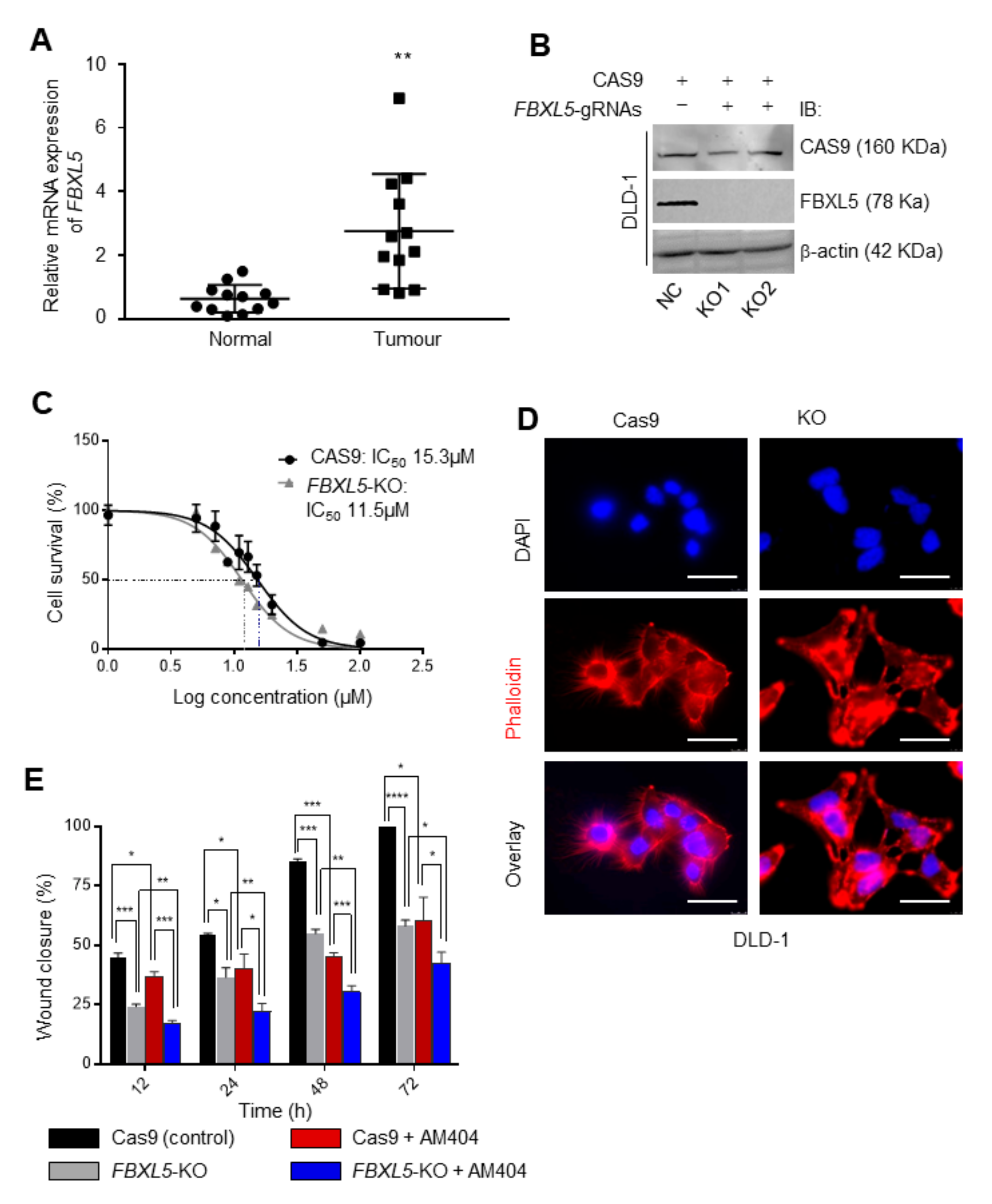
2.4. FBXL5 Attenuates AM404-Induced Anticancer Activity
3. Discussion
4. Materials and Methods
4.1. Human Tissues
4.2. Ex Vivo Explant Culture of CRC Tissues
4.3. Organoid Culture
4.4. Organoids Live/Dead Staining
4.5. Immunohistochemistry (IHC)
4.6. Cell Culture and Colonosphere Formation Assay
4.7. Screening of NIH Clinical Collection Using 3D Colonosphere
4.8. Cytotoxicity Assay
4.9. Clonogenic Assay
4.10. Cell Migration and Wound Healing Assays
4.11. Immunofluorescence and Western Blotting
4.12. Knockout of FBXL5 Using the CRISPR/Cas9 System
4.13. RNA, Transcriptomic, qRT-PCR Assay
4.14. Data Analysis and Statistics
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Arnold, M.; Sierra, M.S.; Laversanne, M.; Soerjomataram, I.; Jemal, A.; Bray, F. Global patterns and trends in colorectal cancer incidence and mortality. Gut 2017, 66, 683–691. [Google Scholar] [CrossRef] [PubMed]
- O’Brien, C.A.; Pollett, A.; Gallinger, S.; Dick, J.E. A human colon cancer cell capable of initiating tumour growth in immunodeficient mice. Nature 2007, 445, 106–110. [Google Scholar] [CrossRef] [PubMed]
- Ricci-Vitiani, L.; Lombardi, D.G.; Pilozzi, E.; Biffoni, M.; Todaro, M.; Peschle, C.; De Maria, R. Identification and expansion of human colon-cancer-initiating cells. Nature 2007, 445, 111–115. [Google Scholar] [CrossRef] [PubMed]
- Singh, A.; Settleman, J. EMT, cancer stem cells and drug resistance: An emerging axis of evil in the war on cancer. Oncogene 2010, 29, 4741–4751. [Google Scholar] [CrossRef] [PubMed]
- Sachlos, E.; Risueño, R.M.; Laronde, S.; Shapovalova, Z.; Lee, J.H.; Russell, J.; Malig, M.; McNicol, J.D.; Fiebig-Comyn, A.; Graham, M.; et al. Identification of Drugs Including a Dopamine Receptor Antagonist that Selectively Target Cancer Stem Cells. Cell 2012, 149, 1284–1297. [Google Scholar] [CrossRef] [PubMed]
- Dashzeveg, N.K.; Taftaf, R.; Ramos, E.K.; Torre-Healy, L.; Chumakova, A.; Silver, D.J.; Alban, T.J.; Sinyuk, M.; Thiagarajan, P.S.; Jarrar, A.M.; et al. New Advances and Challenges of Targeting Cancer Stem Cells. Cancer Res. 2017, 77, 5222–5227. [Google Scholar] [CrossRef] [PubMed]
- Marquardt, S.; Solanki, M.; Spitschak, A.; Vera, J.; Pützer, B.M. Emerging functional markers for cancer stem cell-based therapies: Understanding signaling networks for targeting metastasis. Semin. Cancer Biol. 2018, 53, 90–109. [Google Scholar] [CrossRef]
- Batlle, E.; Clevers, H. Cancer stem cells revisited. Nat. Med. 2017, 23, 1124–1134. [Google Scholar] [CrossRef]
- Hanahan, D.; Weinberg, R. Hallmarks of cancer: The next generation. Cell 2011, 144, 646–674. [Google Scholar] [CrossRef]
- Elinav, E.; Nowarski, R.; Thaiss, C.A.; Hu, B.; Jin, C.; Flavell, R.A. Inflammation-induced cancer: Crosstalk between tumours, immune cells and microorganisms. Nat. Rev. Cancer 2013, 13, 759–771. [Google Scholar] [CrossRef]
- Ahmed, M.; Chaudhari, K.; Babaei-Jadidi, R.; Dekker, L.V.; Shams Nateri, A. Concise Review: Emerging Drugs Targeting Epithelial Cancer Stem-Like Cells. Stem Cells 2017, 35, 839–850. [Google Scholar] [CrossRef] [PubMed]
- Gupta, P.B.; Onder, T.T.; Jiang, G.; Tao, K.; Kuperwasser, C.; Weinberg, R.A.; Lander, E.S. Identification of selective inhibitors of cancer stem cells by high-throughput screening. Cell 2009, 138, 645–659. [Google Scholar] [CrossRef] [PubMed]
- Shibata, H.; Yamakoshi, H.; Sato, A.; Ohori, H.; Kakudo, Y.; Kudo, C.; Takahashi, Y.; Watanabe, M.; Takano, H.; Ishioka, C.; et al. Newly synthesized curcumin analog has improved potential to prevent colorectal carcinogenesis in vivo. Cancer Sci. 2009, 100, 956–960. [Google Scholar] [CrossRef] [PubMed]
- Naujokat, C.; Laufer, S.J.J. Targeting cancer stem cells with defined compounds and drugs. J. Cancer Res. Update 2013, 2, 36–67. [Google Scholar] [CrossRef]
- Shaheen, S.; Ahmed, M.; Lorenzi, F.; Nateri, A.S. Reports Spheroid-Formation (Colonosphere) Assay for in vitro Assessment and Expansion of Stem Cells in Colon Cancer. Stem Cell Rev. Rep. 2016, 12, 492–499. [Google Scholar] [CrossRef]
- Hwang, W.L.; Jiang, J.K.; Yang, S.H.; Huang, T.S.; Lan, H.Y.; Teng, H.W.; Yang, C.Y.; Tsai, Y.P.; Lin, C.H.; Wang, H.W.; et al. MicroRNA-146a directs the symmetric division of Snail-dominant colorectal cancer stem cells. Nat. Cell Biol. 2014, 16, 268–280. [Google Scholar] [CrossRef]
- Kang, N.Y.; Yun, S.W.; Ha, H.H.; Park, S.J.; Chang, Y.T. Embryonic and induced pluripotent stem cell staining and sorting with the live-cell fluorescence imaging probe CDy1. Nat. Protoc. 2011, 6, 1044–1052. [Google Scholar] [CrossRef]
- Vendrell, M.; Park, S.J.; Chandran, Y.; Lee, C.L.; Ha, H.H.; Kang, N.Y.; Yun, S.W.; Chang, Y.T. A fluorescent screening platform for the rapid evaluation of chemicals in cellular reprogramming. Stem Cell Res. 2012, 9, 185–191. [Google Scholar] [CrossRef]
- Saliba, S.W.; Marcotegui, A.R.; Fortwängler, E.; Ditrich, J.; Perazzo, J.C.; Muñoz, E.; de Oliveira, A.C.P.; Fiebich, B.L. AM404, paracetamol metabolite, prevents prostaglandin synthesis in activated microglia by inhibiting COX activity. J. Neuroinflamm. 2017, 14, 246. [Google Scholar] [CrossRef]
- Mazaleuskaya, L.L.; Sangkuhl, K.; Thorn, C.F.; FitzGerald, G.A.; Altman, R.B.; Klein, T.E. PharmGKB summary: Pathways of acetaminophen metabolism at the therapeutic versus toxic doses. Pharm. Genom. 2015, 25, 416–426. [Google Scholar] [CrossRef]
- Sharma, C.V.; Long, J.H.; Shah, S.; Rahman, J.; Perrett, D.; Ayoub, S.S.; Mehta, V. First evidence of the conversion of paracetamol to AM404 in human cerebrospinal fluid. J. Pain Res. 2017, 10, 2703–2709. [Google Scholar] [CrossRef] [PubMed]
- Costa, B.; Siniscalco, D.; Trovato, A.E.; Comelli, F.; Sotgiu, M.L.; Colleoni, M.; Maione, S.; Rossi, F.; Giagnoni, G. AM404, an inhibitor of anandamide uptake, prevents pain behaviour and modulates cytokine and apoptotic pathways in a rat model of neuropathic pain. Br. J. Pharmacol. 2006, 148, 1022–1032. [Google Scholar] [CrossRef] [PubMed]
- Gerits, E.; Spincemaille, P.; De Cremer, K.; De Brucker, K.; Beullens, S.; Thevissen, K.; Cammue, B.P.A.; Vandamme, K.; Fauvart, M.; Verstraeten, N.; et al. Repurposing AM404 for the treatment of oral infections by Porphyromonas gingivalis. Clin. Exp. Dent. Res. 2017, 3, 69–76. [Google Scholar] [CrossRef] [PubMed]
- Yao, H.; Su, S.; Xia, D.; Wang, M.; Li, Z.; Chen, W.; Ren, L.; Xu, L. F-box and leucine-rich repeat protein 5 promotes colon cancer progression by modulating PTEN/PI3K/AKT signaling pathway. Biomed. Pharm. 2018, 107, 1712–1719. [Google Scholar] [CrossRef] [PubMed]
- Xiong, Y.; Sun, F.; Dong, P.; Watari, H.; Yue, J.; Yu, M.F.; Lan, C.Y.; Wang, Y.; Ma, Z.B. iASPP induces EMT and cisplatin resistance in human cervical cancer through miR-20a-FBXL5/BTG3 signaling. J. Exp. Clin. Cancer Res. 2017, 36, 48. [Google Scholar] [CrossRef]
- Machado-Oliveira, G.; Guerreiro, E.; Matias, A.C.; Facucho-Oliveira, J.; Pacheco-Leyva, I.; Bragança, J. FBXL5 modulates HIF-1alpha transcriptional activity by degradation of CITED2. Arch. Biochem. Biophys. 2015, 576, 61–72. [Google Scholar] [CrossRef]
- Dragoi, A.M.; Swiss, R.; Gao, B.; Agaisse, H. Novel strategies to enforce an epithelial phenotype in mesenchymal cells. Cancer Res. 2014, 74, 3659–3672. [Google Scholar] [CrossRef]
- Richon, V.M. Cancer biology: Mechanism of antitumour action of vorinostat (suberoylanilide hydroxamic acid), a novel histone deacetylase inhibitor. Br. J. Cancer. 2006, 95, S2. [Google Scholar] [CrossRef]
- Finnin, M.S.; Donigian, J.R.; Cohen, A.; Richon, V.M.; Rifkind, R.A.; Marks, P.A.; Breslow, R.; Pavletich, N.P. Structures of a histone deacetylase homologue bound to the TSA and SAHA inhibitors. Nature 1999, 401, 188–193. [Google Scholar] [CrossRef]
- Richon, V.M.; Emiliani, S.; Verdin, E.; Webb, Y.; Breslow, R.; Rifkind, R.A.; Marks, P.A. A class of hybrid polar inducers of transformed cell differentiation inhibits histone deacetylases. Proc. Natl. Acad. Sci. USA 1998, 95, 3003–3007. [Google Scholar] [CrossRef]
- Sancho, R.; Jandke, A.; Davis, H.; Diefenbacher, M.E.; Tomlinson, I.; Behrens, A. F-box and WD repeat domain-containing 7 regulates intestinal cell lineage commitment and is a haploinsufficient tumor suppressor. Gastroenterology 2010, 139, 929–941. [Google Scholar] [CrossRef] [PubMed]
- Welcker, M.; Clurman, B.E. FBW7 ubiquitin ligase: A tumour suppressor at the crossroads of cell division, growth and differentiation. Nature reviews. Cancer 2008, 8, 83–93. [Google Scholar] [CrossRef] [PubMed]
- Li, N.; Babaei-Jadidi, R.; Lorenzi, F.; Spencer-Dene, B.; Clarke, P.; Domingo, E.; Tulchinsky, E.; Vries, R.G.J.; Kerr, D.; Pan, Y.; et al. An FBXW7-ZEB2 axis links EMT and tumour microenvironment to promote colorectal cancer stem cells and chemoresistance. Oncogenesis 2019, 8, 13. [Google Scholar] [CrossRef] [PubMed]
- Babaei-Jadidi, R.; Li, N.; Saadeddin, A.; Spencer-Dene, B.; Jandke, A.; Muhammad, B.; Ibrahim, E.E.; Muraleedharan, R.; Abuzinadah, M.; Davis, H.; et al. FBXW7 influences murine intestinal homeostasis and cancer, targeting Notch, Jun, and DEK for degradation. J. Exp. Med. 2011, 208, 295–312. [Google Scholar] [CrossRef]
- Yeung, T.M.; Gandhi, S.C.; Wilding, J.L.; Muschel, R.; Bodmer, W.F. Cancer stem cells from colorectal cancer-derived cell lines. Proc. Natl. Acad. Sci. USA 2010, 107, 3722–3727. [Google Scholar] [CrossRef]
- Buczacki, S.J.A.; Popova, S.; Biggs, E.; Koukorava, C.; Buzzelli, J.; Vermeulen, L.; Hazelwood, L.; Francies, H.; Garnett, M.J.; Winton, D.J. Itraconazole targets cell cycle heterogeneity in colorectal cancer. J. Exp. Med. 2018, 215, 1891–1912. [Google Scholar] [CrossRef]
- Tarpgaard, L.S.; Qvortrup, C.; Nygård, S.B.; Nielsen, S.L.; Andersen, D.R.; Jensen, N.F.; Stenvang, J.; Detlefsen, S.; Brünner, N.; Pfeiffer, P.; et al. A phase II study of Epirubicin in oxaliplatin-resistant patients with metastatic colorectal cancer and TOP2A gene amplification. BMC Cancer 2016, 16, 91. [Google Scholar] [CrossRef]
- Longley, D.B.; Harkin, D.P.; Johnston, P.G. 5-fluorouracil: Mechanisms of action and clinical strategies. Nat. Rev. Cancer 2003, 3, 330–338. [Google Scholar] [CrossRef]
- Lorenzi, F.; Babaei-Jadidi, R.; Sheard, J.; Spencer-Dene, B.; Nateri, A.S. Fbxw7-associated drug resistance is reversed by induction of terminal differentiation in murine intestinal organoid culture. Mol. Ther. Methods Clin. Dev. 2016, 3, 16024. [Google Scholar] [CrossRef][Green Version]
- Chou, T.C.; Talalay, P. Quantitative analysis of dose-effect relationships: The combined effects of multiple drugs or enzyme inhibitors. Adv. Enzym. Regul. 1984, 22, 27–55. [Google Scholar] [CrossRef]
- Foucquier, J.; Guedj, M. Analysis of drug combinations: Current methodological landscape. Pharmacol. Res. Perspect. 2015, 3, e00149. [Google Scholar] [CrossRef]
- Sato, T.; Vries, R.G.; Snippert, H.J.; van de Wetering, M.; Barker, N.; Stange, D.E.; van Es, J.H.; Abo, A.; Kujala, P.; Peters, P.J.; et al. Single Lgr5 stem cells build crypt-villus structures in vitro without a mesenchymal niche. Nature 2009, 459, 262–265. [Google Scholar] [CrossRef]
- Kashfi, S.M.H.; Almozyan, S.; Jinks, N.; Koo, B.K.; Nateri, A.S. Morphological alterations of cultured human colorectal matched tumour and healthy organoids. Oncotarget 2018, 9, 10572–10584. [Google Scholar] [CrossRef]
- Xu, H.; Jiao, Y.; Qin, S.; Zhao, W.; Chu, Q.; Wu, K. Organoid technology in disease modelling, drug development, personalized treatment and regeneration medicine. Exp. Hematol. Oncol. 2018, 7, 30. [Google Scholar] [CrossRef] [PubMed]
- Karekla, E.; Liao, W.J.; Sharp, B.; Pugh, J.; Reid, H.; Quesne, J.L.; Moore, D.; Pritchard, C.; MacFarlane, M.; Pringle, J.H. Ex Vivo Explant Cultures of Non-Small Cell Lung Carcinoma Enable Evaluation of Primary Tumor Responses to Anticancer Therapy. Cancer Res. 2017, 77, 2029–2039. [Google Scholar] [CrossRef] [PubMed]
- Majumder, B.; Baraneedharan, U.; Thiyagarajan, S.; Radhakrishnan, P.; Narasimhan, H.; Dhandapani, M.; Brijwani, N.; Pinto, D.D.; Prasath, A.; Shanthappa, B.U.; et al. Predicting clinical response to anticancer drugs using an ex vivo platform that captures tumour heterogeneity. Nat. Commun 2015, 6, 6169. [Google Scholar] [CrossRef] [PubMed]
- Qualtrough, D.; Hinoi, T.; Fearon, E.; Paraskeva, C. Expression of CDX2 in normal and neoplastic human colon tissue and during differentiation of an in vitro model system. Gut 2002, 51, 184–190. [Google Scholar] [CrossRef]
- Merlos-Suárez, A.; Barriga, F.M.; Jung, P.; Iglesias, M.; Céspedes, M.V.; Rossell, D.; Sevillano, M.; Hernando-Momblona, X.; da Silva-Diz, V.; Muñoz, P.; et al. The Intestinal Stem Cell Signature Identifies Colorectal Cancer Stem Cells and Predicts Disease Relapse. Cell Stem Cell 2011, 8, 511–524. [Google Scholar] [CrossRef]
- Todaro, M.; Gaggianesi, M.; Catalano, V.; Benfante, A.; Iovino, F.; Biffoni, M.; Apuzzo, T.; Sperduti, I.; Volpe, S.; Cocorullo, G.; et al. CD44v6 is a marker of constitutive and reprogrammed cancer stem cells driving colon cancer metastasis. Cell Stem Cell 2014, 14, 342–356. [Google Scholar] [CrossRef]
- Nateri, A.S.; Riera-Sans, L.; Da Costa, C.; Behrens, A. The ubiquitin ligase SCFFbw7 antagonizes apoptotic JNK signaling. Science 2004, 303, 1374–1378. [Google Scholar] [CrossRef]
- Diaz, V.M.; de Herreros, A.G. F-box proteins: Keeping the epithelial-to-mesenchymal transition (EMT) in check. Semin. Cancer Biol. 2016, 36, 71–79. [Google Scholar] [CrossRef] [PubMed]
- Muto, Y.; Nishiyama, M.; Nita, A.; Moroishi, T.; Nakayama, K.I. Essential role of FBXL5-mediated cellular iron homeostasis in maintenance of hematopoietic stem cells. Nat. Commun. 2017, 8, 16114. [Google Scholar] [CrossRef] [PubMed]
- Liang, C.C.; Park, A.Y.; Guan, J.L. In vitro scratch assay: A convenient and inexpensive method for analysis of cell migration in vitro. Nat. Protoc. 2007, 2, 329. [Google Scholar] [CrossRef] [PubMed]
- Diakos, C.I.; Charles, K.A.; McMillan, D.C.; Clarke, S.J. Cancer-related inflammation and treatment effectiveness. Lancet Oncol. 2014, 15, e493–e503. [Google Scholar] [CrossRef]
- Lasry, A.; Zinger, A.; Ben-Neriah, Y. Inflammatory networks underlying colorectal cancer. Nat. Immunol. 2016, 17, 230. [Google Scholar] [CrossRef]
- Caballero, F.J.; Soler-Torronteras, R.; Lara-Chica, M.; García, V.; Fiebich, B.L.; Muñoz, E.; Calzado, M.A. AM404 inhibits NFAT and NF-κB signaling pathways and impairs migration and invasiveness of neuroblastoma cells. Eur. J. Pharmacol. 2015, 746, 221–232. [Google Scholar] [CrossRef]
- De Lago, E.; Gustafsson, S.B.; Fernández-Ruiz, J.; Nilsson, J.; Jacobsson, S.O.; Fowler, C.J. Acyl-based anandamide uptake inhibitors cause rapid toxicity to C6 glioma cells at pharmacologically relevant concentrations. J. Neurochem. 2006, 99, 677–688. [Google Scholar] [CrossRef]
- Kleppe, M.; Levine, R.L. Tumor heterogeneity confounds and illuminates: Assessing the implications. Nat. Med. 2014, 20, 342–344. [Google Scholar] [CrossRef]
- Li, Y.; Rogoff, H.A.; Keates, S.; Gao, Y.; Murikipudi, S.; Mikule, K.; Leggett, D.; Li, W.; Pardee, A.B.; Li, C.J. Suppression of cancer relapse and metastasis by inhibiting cancer stemness. Proc. Natl. Acad. Sci. USA 2015, 112, 1839–1844. [Google Scholar] [CrossRef]
- Chen, D.; Pamu, S.; Cui, Q.; Chan, T.H.; Dou, Q.P. Novel epigallocatechin gallate (EGCG) analogs activate AMP-activated protein kinase pathway and target cancer stem cells. Bioorg. Med. Chem. 2012, 20, 3031–3037. [Google Scholar] [CrossRef]
- Jung, J.W.; Park, S.B.; Lee, S.J.; Seo, M.S.; Trosko, J.E.; Kang, K.S. Metformin represses self-renewal of the human breast carcinoma stem cells via inhibition of estrogen receptor-mediated OCT4 expression. PLoS ONE 2011, 6, e28068. [Google Scholar] [CrossRef] [PubMed]
- Singh, B.N.; Fu, J.; Srivastava, R.K.; Shankar, S. Hedgehog signaling antagonist GDC-0449 (Vismodegib) inhibits pancreatic cancer stem cell characteristics: Molecular mechanisms. PLoS ONE 2011, 6, e27306. [Google Scholar] [CrossRef] [PubMed]
- Detre, S.; Saclani Jotti, G.; Dowsett, M. A “quickscore” method for immunohistochemical semiquantitation: Validation for oestrogen receptor in breast carcinomas. J. Clin. Pathol. 1995, 48, 876–878. [Google Scholar] [CrossRef] [PubMed]
- Muhammad, B.A.; Almozyan, S.; Babaei-Jadidi, R.; Onyido, E.K.; Saadeddin, A.; Kashfi, S.H.; Spencer-Dene, B.; Ilyas, M.; Lourdusamy, A.; Behrens, A.; et al. FLYWCH1, a Novel Suppressor of Nuclear Beta-Catenin, Regulates Migration and Morphology in Colorectal Cancer. Mol. Cancer Res. 2018, 16, 1977–1990. [Google Scholar] [CrossRef]
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Ahmed, M.; Jinks, N.; Babaei-Jadidi, R.; Kashfi, H.; Castellanos-Uribe, M.; T. May, S.; Mukherjee, A.; Nateri, A.S. Repurposing Antibacterial AM404 As a Potential Anticancer Drug for Targeting Colorectal Cancer Stem-Like Cells. Cancers 2020, 12, 106. https://doi.org/10.3390/cancers12010106
Ahmed M, Jinks N, Babaei-Jadidi R, Kashfi H, Castellanos-Uribe M, T. May S, Mukherjee A, Nateri AS. Repurposing Antibacterial AM404 As a Potential Anticancer Drug for Targeting Colorectal Cancer Stem-Like Cells. Cancers. 2020; 12(1):106. https://doi.org/10.3390/cancers12010106
Chicago/Turabian StyleAhmed, Mehreen, Nicholas Jinks, Roya Babaei-Jadidi, Hossein Kashfi, Marcos Castellanos-Uribe, Sean T. May, Abhik Mukherjee, and Abdolrahman S. Nateri. 2020. "Repurposing Antibacterial AM404 As a Potential Anticancer Drug for Targeting Colorectal Cancer Stem-Like Cells" Cancers 12, no. 1: 106. https://doi.org/10.3390/cancers12010106
APA StyleAhmed, M., Jinks, N., Babaei-Jadidi, R., Kashfi, H., Castellanos-Uribe, M., T. May, S., Mukherjee, A., & Nateri, A. S. (2020). Repurposing Antibacterial AM404 As a Potential Anticancer Drug for Targeting Colorectal Cancer Stem-Like Cells. Cancers, 12(1), 106. https://doi.org/10.3390/cancers12010106