The Efficacy of miR-20a as a Diagnostic and Prognostic Biomarker for Colorectal Cancer: A Systematic Review and Meta-Analysis
Abstract
1. Introduction
2. Methods
2.1. Study Selection Criteria
2.2. Search Strategy
2.3. Data Extraction
2.4. Meta-Analysis
2.5. Study Quality Assessment
3. Results
3.1. Study Selection
3.2. Basic Characteristics of The Selected Studies
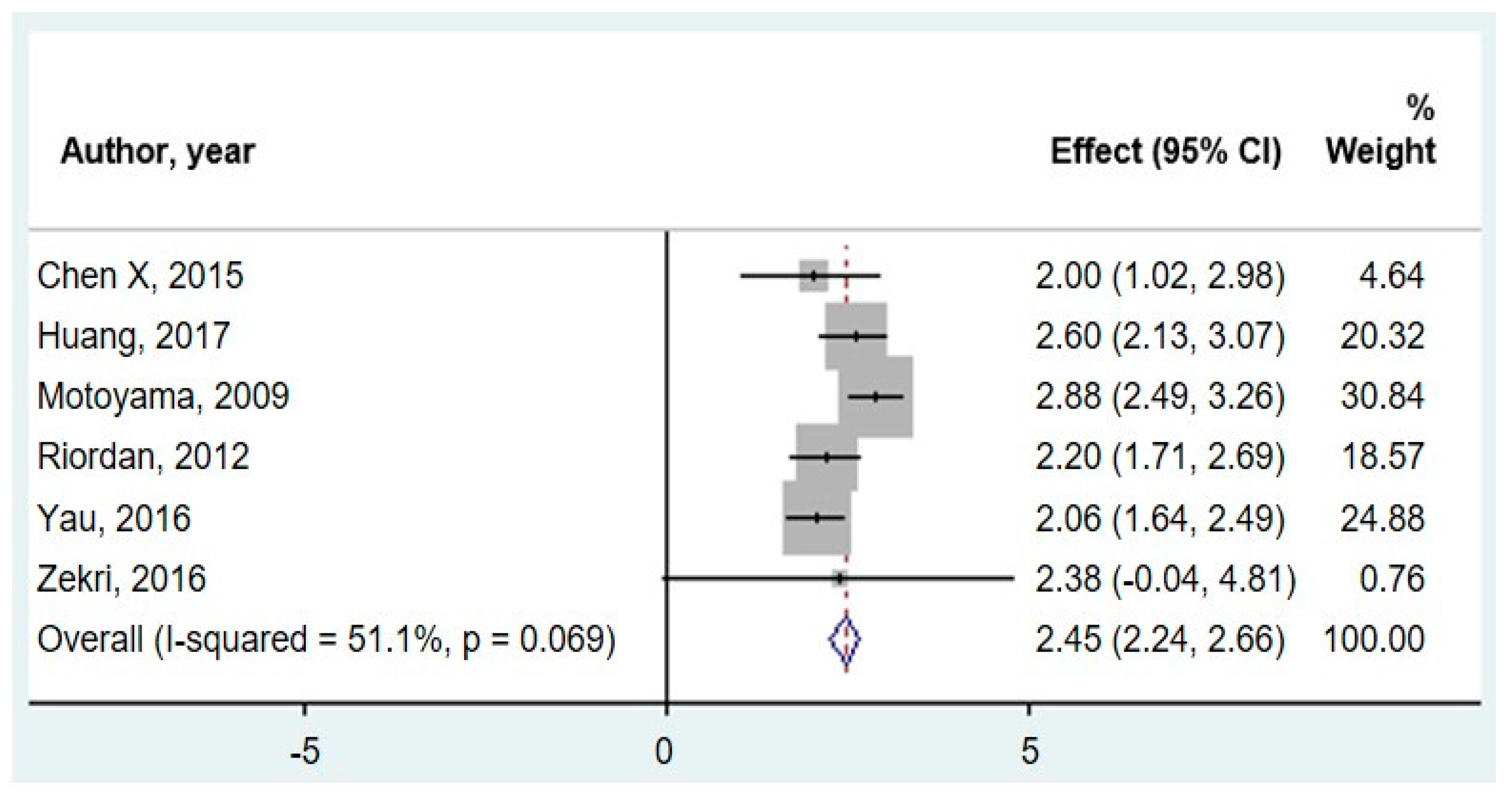
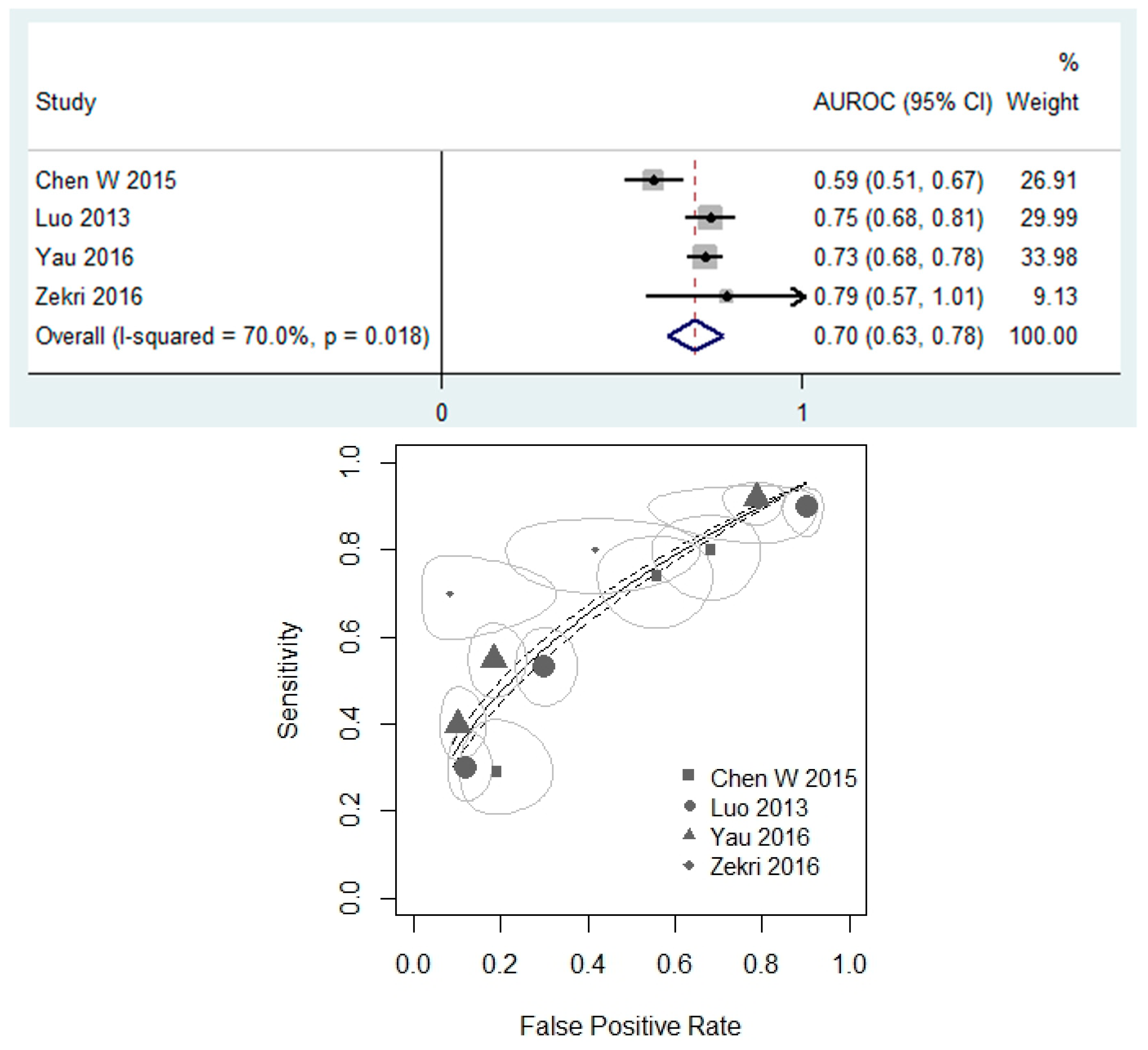
3.3. miR-20a as a Diagnostic Biomarker of CRC
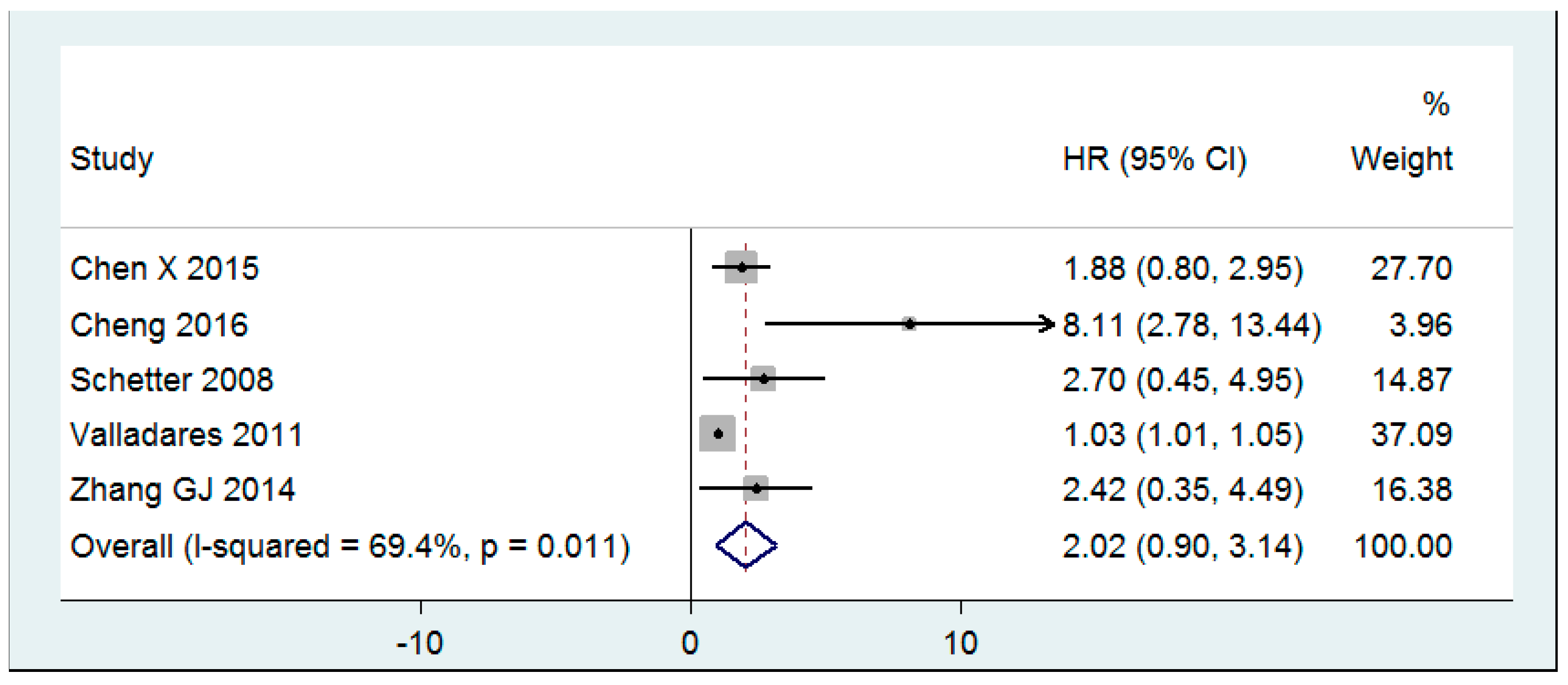
3.4. mir-20a as a Prognostic Biomarker of CRC Survival
3.5. Study Quality Assessment
4. Discussion
5. Conclusions
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Bretthauer, M. Colorectal cancer screening. J. Intern. Med. 2011, 270, 87–98. [Google Scholar] [CrossRef] [PubMed]
- Vogel, J.D.; Eskicioglu, C.; Weiser, M.R.; Feingold, D.L.; Steele, S.R. The american society of colon and rectal surgeons clinical practice guidelines for the treatment of colon cancer. Dis. Colon Rectum 2017, 60, 999–1017. [Google Scholar] [CrossRef] [PubMed]
- Habr-Gama, A.; Sao Juliao, G.P.; Perez, R.O. Nonoperative management of rectal cancer: Identifying the ideal patients. Hematol. Oncol. Clin. North Am. 2015, 29, 135–151. [Google Scholar] [CrossRef] [PubMed]
- Pozo, M.E.; Fang, S.H. Watch and wait approach to rectal cancer: A review. World J. Gastrointest. Surg. 2015, 7, 306–312. [Google Scholar] [CrossRef] [PubMed]
- Winawer, S.J.; Zauber, A.G.; Ho, M.N.; O’Brien, M.J.; Gottlieb, L.S.; Sternberg, S.S.; Waye, J.D.; Schapiro, M.; Bond, J.H.; Panish, J.F.; et al. Prevention of colorectal cancer by colonoscopic polypectomy. The national polyp study workgroup. New Engl. J. Med. 1993, 329, 1977–1981. [Google Scholar] [CrossRef] [PubMed]
- Kahi, C.J.; Imperiale, T.F.; Juliar, B.E.; Rex, D.K. Effect of screening colonoscopy on colorectal cancer incidence and mortality. Clin. Gastroenterol. Hepatol. 2009, 7, 770–775. [Google Scholar] [CrossRef] [PubMed]
- Baxter, N.N.; Warren, J.L.; Barrett, M.J.; Stukel, T.A.; Doria-Rose, V.P. Association between colonoscopy and colorectal cancer mortality in a us cohort according to site of cancer and colonoscopist specialty. J. Clin. Oncol. 2012, 30, 2664–2669. [Google Scholar] [CrossRef] [PubMed]
- Bipat, S.; van Leeuwen, M.S.; Comans, E.F.; Pijl, M.E.; Bossuyt, P.M.; Zwinderman, A.H.; Stoker, J. Colorectal liver metastases: Ct, mr imaging, and pet for diagnosis--meta-analysis. Radiology 2005, 237, 123–131. [Google Scholar] [CrossRef]
- Pema, P.J.; Bennett, W.F.; Bova, J.G.; Warman, P. Ct vs mri in diagnosis of recurrent rectosigmoid carcinoma. J. Comput. Assist. Tomogr. 1994, 18, 256–261. [Google Scholar] [CrossRef]
- Malila, N.; Oivanen, T.; Malminiemi, O.; Hakama, M. Test, episode, and programme sensitivities of screening for colorectal cancer as a public health policy in finland: Experimental design. BMJ 2008, 337, a2261. [Google Scholar] [CrossRef]
- Hardcastle, J.D.; Chamberlain, J.O.; Robinson, M.H.; Moss, S.M.; Amar, S.S.; Balfour, T.W.; James, P.D.; Mangham, C.M. Randomised controlled trial of faecal-occult-blood screening for colorectal cancer. Lancet 1996, 348, 1472–1477. [Google Scholar] [CrossRef]
- Kronborg, O.; Fenger, C.; Olsen, J.; Jorgensen, O.D.; Sondergaard, O. Randomised study of screening for colorectal cancer with faecal-occult-blood test. Lancet 1996, 348, 1467–1471. [Google Scholar] [CrossRef]
- Fletcher, R.H. Carcinoembryonic antigen. Ann. Intern. Med. 1986, 104, 66–73. [Google Scholar] [CrossRef] [PubMed]
- Gu, S.; Kay, M.A. How do mirnas mediate translational repression? Silence 2010, 1, 11. [Google Scholar] [CrossRef] [PubMed]
- Meng, F.; Henson, R.; Wehbe-Janek, H.; Ghoshal, K.; Jacob, S.T.; Patel, T. Microrna-21 regulates expression of the pten tumor suppressor gene in human hepatocellular cancer. Gastroenterology 2007, 133, 647–658. [Google Scholar] [CrossRef]
- Lu, Z.; Liu, M.; Stribinskis, V.; Klinge, C.M.; Ramos, K.S.; Colburn, N.H.; Li, Y. Microrna-21 promotes cell transformation by targeting the programmed cell death 4 gene. Oncogene 2008, 27, 4373–4379. [Google Scholar] [CrossRef] [PubMed]
- Sayed, D.; Rane, S.; Lypowy, J.; He, M.; Chen, I.Y.; Vashistha, H.; Yan, L.; Malhotra, A.; Vatner, D.; Abdellatif, M. Microrna-21 targets sprouty2 and promotes cellular outgrowths. Mol. Biol. Cell 2008, 19, 3272–3282. [Google Scholar] [CrossRef]
- Thum, T.; Gross, C.; Fiedler, J.; Fischer, T.; Kissler, S.; Bussen, M.; Galuppo, P.; Just, S.; Rottbauer, W.; Frantz, S.; et al. Microrna-21 contributes to myocardial disease by stimulating map kinase signalling in fibroblasts. Nature 2008, 456, 980–984. [Google Scholar] [CrossRef]
- Buscaglia, L.E.; Li, Y. Apoptosis and the target genes of microrna-21. Chin. J. Cancer 2011, 30, 371–380. [Google Scholar] [CrossRef]
- Medina, P.P.; Nolde, M.; Slack, F.J. Oncomir addiction in an in vivo model of microrna-21-induced pre-b-cell lymphoma. Nature 2010, 467, 86–90. [Google Scholar] [CrossRef]
- Yan, L.X.; Huang, X.F.; Shao, Q.; Huang, M.Y.; Deng, L.; Wu, Q.L.; Zeng, Y.X.; Shao, J.Y. Microrna mir-21 overexpression in human breast cancer is associated with advanced clinical stage, lymph node metastasis and patient poor prognosis. RNA 2008, 14, 2348–2360. [Google Scholar] [CrossRef]
- Zhou, X.; Wang, X.P.; Huang, Z.B.; Wang, J.; Zhu, W.; Shu, Y.Q.; Liu, P. Prognostic value of mir-21 in various cancers: An updating meta-analysis. PLoS ONE 2014, 9, e102413. [Google Scholar] [CrossRef] [PubMed]
- Tanzer, A.; Stadler, P.F. Molecular evolution of a microrna cluster. J. Mol. Biol. 2004, 339, 327–335. [Google Scholar] [CrossRef] [PubMed]
- Ventura, A.; Young, A.G.; Winslow, M.M.; Lintault, L.; Meissner, A.; Erkeland, S.J.; Newman, J.; Bronson, R.T.; Crowley, D.; Stone, J.R.; et al. Targeted deletion reveals essential and overlapping functions of the mir-17 through 92 family of mirna clusters. Cell 2008, 132, 875–886. [Google Scholar] [CrossRef] [PubMed]
- Liu, S.Q.; Jiang, S.; Li, C.; Zhang, B.; Li, Q.J. Mir-17-92 cluster targets phosphatase and tensin homology and ikaros family zinc finger 4 to promote th17-mediated inflammation. J. Biol. Chem. 2014, 289, 12446–12456. [Google Scholar] [CrossRef] [PubMed]
- O’Donnell, K.A.; Wentzel, E.A.; Zeller, K.I.; Dang, C.V.; Mendell, J.T. C-myc-regulated micrornas modulate e2f1 expression. Nature 2005, 435, 839–843. [Google Scholar] [CrossRef] [PubMed]
- Dews, M.; Fox, J.L.; Hultine, S.; Sundaram, P.; Wang, W.; Liu, Y.Y.; Furth, E.; Enders, G.H.; El-Deiry, W.; Schelter, J.M.; et al. The myc-mir-17~92 axis blunts tgf{beta} signaling and production of multiple tgf{beta}-dependent antiangiogenic factors. Cancer Res. 2010, 70, 8233–8246. [Google Scholar] [CrossRef] [PubMed]
- Mogilyansky, E.; Rigoutsos, I. The mir-17/92 cluster: A comprehensive update on its genomics, genetics, functions and increasingly important and numerous roles in health and disease. Cell Death Differ. 2013, 20, 1603–1614. [Google Scholar] [CrossRef] [PubMed]
- Peng, W.; Liu, Y.N.; Zhu, S.Q.; Li, W.Q.; Guo, F.C. The correlation of circulating pro-angiogenic mirnas’ expressions with disease risk, clinicopathological features, and survival profiles in gastric cancer. Cancer Med. 2018, 7, 3773–3791. [Google Scholar] [CrossRef]
- Zeng, X.; Xiang, J.; Wu, M.; Xiong, W.; Tang, H.; Deng, M.; Li, X.; Liao, Q.; Su, B.; Luo, Z.; et al. Circulating mir-17, mir-20a, mir-29c, and mir-223 combined as non-invasive biomarkers in nasopharyngeal carcinoma. PLoS ONE 2012, 7, e46367. [Google Scholar] [CrossRef]
- Bryant, R.J.; Pawlowski, T.; Catto, J.W.; Marsden, G.; Vessella, R.L.; Rhees, B.; Kuslich, C.; Visakorpi, T.; Hamdy, F.C. Changes in circulating microrna levels associated with prostate cancer. Br. J. Cancer 2012, 106, 768–774. [Google Scholar] [CrossRef] [PubMed]
- Cheng, D.; Zhao, S.; Tang, H.; Zhang, D.; Sun, H.; Yu, F.; Jiang, W.; Yue, B.; Wang, J.; Zhang, M.; et al. Microrna-20a-5p promotes colorectal cancer invasion and metastasis by downregulating smad4. Oncotarget 2016, 7, 45199–45213. [Google Scholar] [CrossRef] [PubMed]
- Mestdagh, P.; Bostrom, A.K.; Impens, F.; Fredlund, E.; Van Peer, G.; De Antonellis, P.; von Stedingk, K.; Ghesquiere, B.; Schulte, S.; Dews, M.; et al. The mir-17-92 microrna cluster regulates multiple components of the tgf-beta pathway in neuroblastoma. Mol. Cell 2010, 40, 762–773. [Google Scholar] [CrossRef] [PubMed]
- Zhou, J.; Guo, F.; Wang, G.; Wang, J.; Zheng, F.; Guan, X.; Chang, A.; Zhang, X.; Dai, C.; Li, S.; et al. Mir-20a regulates adipocyte differentiation by targeting lysine-specific demethylase 6b and transforming growth factor-beta signaling. Int. J. Obes. 2015, 39, 1282–1291. [Google Scholar] [CrossRef] [PubMed]
- Sokolova, V.; Fiorino, A.; Zoni, E.; Crippa, E.; Reid, J.F.; Gariboldi, M.; Pierotti, M.A. The effects of mir-20a on p21: Two mechanisms blocking growth arrest in tgf-beta-responsive colon carcinoma. J. Cell. Physiol. 2015, 230, 3105–3114. [Google Scholar] [CrossRef] [PubMed]
- Zhao, S.; Yao, D.; Chen, J.; Ding, N.; Ren, F. Mir-20a promotes cervical cancer proliferation and metastasis in vitro and in vivo. PLoS ONE 2015, 10, e0120905. [Google Scholar] [CrossRef] [PubMed]
- Littell, J.H.; Corcoran, J.; Pillai, V. Systematic Reviews and Meta-Analysis; Oxford University Press: Oxford, UK, 2008. [Google Scholar]
- Bovell, L.; Shanmugam, C.; Katkoori, V.R.; Zhang, B.; Vogtmann, E.; Grizzle, W.E.; Manne, U. Mirnas are stable in colorectal cancer archival tissue blocks. Front. Biosci. 2012, 4, 1937–1940. [Google Scholar] [CrossRef]
- Ling, H.; Spizzo, R.; Atlasi, Y.; Nicoloso, M.; Shimizu, M.; Redis, R.S.; Nishida, N.; Gafa, R.; Song, J.; Guo, Z.; et al. Ccat2, a novel noncoding rna mapping to 8q24, underlies metastatic progression and chromosomal instability in colon cancer. Genome Res. 2013, 23, 1446–1461. [Google Scholar] [CrossRef] [PubMed]
- Ahmed, F.E.; Ahmed, N.C.; Vos, P.W.; Bonnerup, C.; Atkins, J.N.; Casey, M.; Nuovo, G.J.; Naziri, W.; Wiley, J.E.; Mota, H.; et al. Diagnostic microrna markers to screen for sporadic human colon cancer in stool: I. Proof of principle. Cancer Genom. Proteom. 2013, 10, 93–113. [Google Scholar]
- Azizian, A.; Kramer, F.; Jo, P.; Wolff, H.A.; Beissbarth, T.; Skarupke, R.; Bernhardt, M.; Grade, M.; Ghadimi, B.M.; Gaedcke, J. Preoperative prediction of lymph node status by circulating mir-18b and mir-20a during chemoradiotherapy in patients with rectal cancer. World J. Surg. 2015, 39, 2329–2335. [Google Scholar] [CrossRef] [PubMed]
- Bovell, L.C.; Shanmugam, C.; Putcha, B.D.K.; Katkoori, V.R.; Zhang, B.; Bae, S.J.; Singh, K.P.; Grizzle, W.E.; Manne, U. The prognostic value of micrornas varies with patient race/ethnicity and stage of colorectal cancer. Clin. Cancer Res. 2013, 19, 3955–3965. [Google Scholar] [CrossRef]
- Brunet Vega, A.; Pericay, C.; Moya, I.; Ferrer, A.; Dotor, E.; Pisa, A.; Casalots, A.; Serra-Aracil, X.; Oliva, J.C.; Ruiz, A.; et al. Microrna expression profile in stage iii colorectal cancer: Circulating mir-18a and mir-29a as promising biomarkers. Oncol. Rep. 2013, 30, 320–326. [Google Scholar] [CrossRef]
- Caritg, O.; Navarro, A.; Moreno, I.; Martinez-Rodenas, F.; Cordeiro, A.; Munoz, C.; Ruiz-Martinez, M.; Santasusagna, S.; Castellano, J.J.; Monzo, M. Identifying high-risk stage ii colon cancer patients: A three-microrna-based score as a prognostic biomarker. Clin. Colorectal Cancer 2016, 15, e175–e182. [Google Scholar] [CrossRef]
- Chen, W.Y.; Zhao, X.J.; Yu, Z.F.; Hu, F.L.; Liu, Y.P.; Cui, B.B.; Dong, X.S.; Zhao, Y.S. The potential of plasma mirnas for diagnosis and risk estimation of colorectal cancer. Int. J. Clin. Exp. Pathol. 2015, 8, 7092–7101. [Google Scholar] [PubMed]
- Chen, X.J.; Shi, K.Q.; Wang, Y.Q.; Song, M.; Zhou, W.; Tu, H.X.; Lin, Z. Clinical value of integrated-signature mirnas in colorectal cancer: Mirna expression profiling analysis and experimental validation. Oncotarget 2015, 6, 37553–37565. [Google Scholar] [CrossRef]
- de Groen, F.L.M.; Timmer, L.M.; Menezes, R.X.; Diosdado, B.; Hooijberg, E.; Meijer, G.A.; Steenbergen, R.D.M.; Carvalho, B. Oncogenic role of mir-15a-3p in 13q amplicon-driven colorectal adenoma-to-carcinoma progression. PLoS ONE 2015, 10, 13. [Google Scholar] [CrossRef]
- Hotchi, M.; Shimada, M.; Kurita, N.; Iwata, T.; Sato, H.; Morimoto, S.; Yoshikawa, K.; Higashijima, J.; Miyatani, T. Microrna expression is able to predict response to chemoradiotherapy in rectal cancer. Mol. Clin. Oncol. 2013, 1, 137–142. [Google Scholar] [CrossRef]
- Huang, G.L.; Chen, X.J.; Cai, Y.F.; Wang, X.B.; Xing, C.G. Mir-20a-directed regulation of bid is associated with the trail sensitivity in colorectal cancer. Oncol. Rep. 2017, 37, 571–578. [Google Scholar] [CrossRef]
- Luo, X.Y.; Stock, C.; Burwinkel, B.; Brenner, H. Identification and evaluation of plasma micrornas for early detection of colorectal cancer. PLoS ONE 2013, 8, 9. [Google Scholar] [CrossRef]
- Molinari, C.; Salvi, S.; Foca, F.; Teodorani, N.; Saragoni, L.; Puccetti, M.; Passardi, A.; Tamberi, S.; Avanzolini, A.; Lucci, E.; et al. Mir-17-92a-1 cluster host gene (mir17hg) evaluation and response to neoadjuvant chemoradiotherapy in rectal cancer. Oncotargets Ther. 2016, 9, 2735–2742. [Google Scholar]
- Motoyama, K.; Inoue, H.; Takatsuno, Y.; Tanaka, F.; Mimori, K.; Uetake, H.; Sugihara, K.; Mori, M. Over- and under-expressed micrornas in human colorectal cancer. Int. J. Oncol. 2009, 34, 1069–1075. [Google Scholar]
- Ozcan, O.; Kara, M.; Yumrutas, O.; Bozgeyik, E.; Bozgeyik, I.; Celik, O.I. Mtus1 and its targeting mirnas in colorectal carcinoma: Significant associations. Tumor Biol. 2016, 37, 6637–6645. [Google Scholar] [CrossRef]
- Pellatt, D.F.; Stevens, J.R.; Wolff, R.K.; Mullany, L.E.; Herrick, J.S.; Samowitz, W.; Slattery, M.L. Expression profiles of mirna subsets distinguish human colorectal carcinoma and normal colonic mucosa. Clin. Transl. Gastroenterol. 2016, 7, 10. [Google Scholar] [CrossRef]
- Riordan, A.M.; Thomas, M.K.; Ronnekleiv-Kelly, S.; Warner, T.; Geiger, P.G.; Kennedy, G.D. Utility of micro-ribonucleic acid profile for predicting recurrence of rectal cancer. J. Surg. Res. 2012, 177, 87–92. [Google Scholar] [CrossRef]
- Ristau, J.; Staffa, J.; Schrotz-King, P.; Gigic, B.; Makar, K.W.; Hoffmeister, M.; Brenner, H.; Ulrich, A.; Schneider, M.; Ulrich, C.M.; et al. Suitability of circulating mirnas as potential prognostic markers in colorectal cancer. Cancer Epidemiol. Biomark. Prev. 2014, 23, 2632–2637. [Google Scholar] [CrossRef]
- Rotelli, M.T.; Di Lena, M.; Cavallini, A.; Lippolis, C.; Bonfrate, L.; Chetta, N.; Portincasa, P.; Altomare, D.F. Fecal microrna profile in patients with colorectal carcinoma before and after curative surgery. Int. J. Colorectal Dis. 2015, 30, 891–898. [Google Scholar] [CrossRef]
- Schetter, A.J.; Leung, S.Y.; Sohn, J.J.; Zanetti, K.A.; Bowman, E.D.; Yanaihara, N.; Yuen, S.T.; Chan, T.L.; Kwong, D.L.W.; Au, G.K.H.; et al. Microrna expression profiles associated with prognosis and therapeutic outcome in colon adenocarcinoma. JAMA J. Am. Med. Assoc. 2008, 299, 425–436. [Google Scholar] [CrossRef]
- Tsuchida, A.; Ohno, S.; Wu, W.H.; Borjigin, N.; Fujita, K.; Aoki, T.; Ueda, S.; Takanashi, M.; Kuroda, M. Mir-92 is a key oncogenic component of the mir-17-92 cluster in colon cancer. Cancer Sci. 2011, 102, 2264–2271. [Google Scholar] [CrossRef]
- Valladares-Ayerbes, M.; Blanco, M.; Haz, M.; Medina, V.; Iglesias-Diaz, P.; Lorenzo-Patino, M.J.; Reboredo, M.; Santamarina, I.; Figueroa, A.; Anton-Aparicio, L.M.; et al. Prognostic impact of disseminated tumor cells and microrna-17-92 cluster deregulation in gastrointestinal cancer. Int. J. Oncol. 2011, 39, 1253–1264. [Google Scholar] [CrossRef]
- Volinia, S.; Calin, G.A.; Liu, C.G.; Ambs, S.; Cimmino, A.; Petrocca, F.; Visone, R.; Iorio, M.; Roldo, C.; Ferracin, M.; et al. A microrna expression signature of human solid tumors defines cancer gene targets. Proc. Natl. Acad. Sci. USA 2006, 103, 2257–2261. [Google Scholar] [CrossRef]
- Wang, Y.X.; Zhang, X.Y.; Zhang, B.F.; Yang, C.Q.; Chen, X.M.; Gao, H.J. Initial study of microrna expression profiles of colonic cancer without lymph node metastasis. J. Dig. Dis. 2010, 11, 50–54. [Google Scholar] [CrossRef]
- Xu, T.; Jing, C.Q.; Shi, Y.L.; Miao, R.Z.; Peng, L.P.; Kong, S.; Ma, Y.; Li, L.P. Microrna-20a enhances the epithelial-to-mesenchymal transition of colorectal cancer cells by modulating matrix metalloproteinases. Exp. Ther. Med. 2015, 10, 683–688. [Google Scholar] [CrossRef]
- Yamada, A.; Horimatsu, T.; Okugawa, Y.; Nishida, N.; Honjo, H.; Ida, H.; Kou, T.; Kusaka, T.; Sasaki, Y.; Yagi, M.; et al. Serum mir-21, mir-29a, and mir-125b are promising biomarkers for the early detection of colorectal neoplasia. Clin. Cancer Res. 2015, 21, 4234–4242. [Google Scholar] [CrossRef]
- Yantiss, R.K.; Goodarzi, M.; Zhou, X.K.; Rennert, H.; Pirog, E.C.; Banner, B.F.; Chen, Y.T. Clinical, pathologic, and molecular features of early-onset colorectal carcinoma. Am. J. Surg. Pathol. 2009, 33, 572–582. [Google Scholar] [CrossRef]
- Yau, T.O.; Wu, C.W.; Tang, C.M.; Chen, Y.X.; Fang, J.Y.; Dong, Y.J.; Liang, Q.Y.; Ng, S.S.M.; Chan, F.K.L.; Sung, J.J.Y.; et al. Microrna-20a in human faeces as a non-invasive biomarker for colorectal cancer. Oncotarget 2016, 7, 1559–1568. [Google Scholar] [CrossRef]
- Zekri, A.R.N.; Youssef, A.S.E.; Lotfy, M.M.; Gabr, R.; Ahmed, O.S.; Nassar, A.; Hussein, N.; Omran, D.; Medhat, E.; Eid, S.; et al. Circulating serum mirnas as diagnostic markers for colorectal cancer. PLoS ONE 2016, 11, 14. [Google Scholar] [CrossRef]
- Zhang, J.; Zhang, K.J.; Bi, M.S.; Jiao, X.L.; Zhang, D.L.; Dong, Q. Circulating microrna expressions in colorectal cancer as predictors of response to chemotherapy. Anti-Cancer Drugs 2014, 25, 346–352. [Google Scholar] [CrossRef]
- Zhang, J.X.; Song, W.; Chen, Z.H.; Wei, J.H.; Liao, Y.J.; Lei, J.; Hu, M.; Chen, G.Z.; Liao, B.; Lu, J.; et al. Prognostic and predictive value of a microrna signature in stage ii colon cancer: A microrna expression analysis. Lancet Oncol. 2013, 14, 1295–1306. [Google Scholar] [CrossRef]
- Pickering, M.T.; Stadler, B.M.; Kowalik, T.F. Mir-17 and mir-20a temper an e2f1-induced g1 checkpoint to regulate cell cycle progression. Oncogene 2009, 28, 140–145. [Google Scholar] [CrossRef]
- Tan, E.; Gouvas, N.; Nicholls, R.J.; Ziprin, P.; Xynos, E.; Tekkis, P.P. Diagnostic precision of carcinoembryonic antigen in the detection of recurrence of colorectal cancer. Surg. Oncol. 2009, 18, 15–24. [Google Scholar] [CrossRef]
- Zhang, S.Y.; Lin, M.; Zhang, H.B. Diagnostic value of carcinoembryonic antigen and carcinoma antigen 19-9 for colorectal carcinoma. Int. J. Clin. Exp. Pathol. 2015, 8, 9404–9409. [Google Scholar]
- Wang, Y.R.; Yan, J.X.; Wang, L.N. The diagnostic value of serum carcino-embryonic antigen, alpha fetoprotein and carbohydrate antigen 19-9 for colorectal cancer. J. Cancer Res. Ther. 2014, 10, 307–309. [Google Scholar]
- Xia, X.; Yang, B.; Zhai, X.; Liu, X.; Shen, K.; Wu, Z.; Cai, J. Prognostic role of microrna-21 in colorectal cancer: A meta-analysis. PLoS ONE 2013, 8, e80426. [Google Scholar] [CrossRef]
- Chen, Z.; Liu, H.; Jin, W.; Ding, Z.; Zheng, S.; Yu, Y. Tissue microrna-21 expression predicted recurrence and poor survival in patients with colorectal cancer—a meta-analysis. Onco Targets Ther. 2016, 9, 2615–2624. [Google Scholar]
- Zhao, Y.; Dai, C.; Wang, M.; Kang, H.; Lin, S.; Yang, P.; Liu, X.; Liu, K.; Xu, P.; Zheng, Y.; et al. Clinicopathological and prognostic significance of metastasis-associated in colon cancer-1 (macc1) overexpression in colorectal cancer: A meta-analysis. Oncotarget 2016, 7, 62966–62975. [Google Scholar] [CrossRef]
- Jing, C.; Jin, Y.H.; You, Z.; Qiong, Q.; Jun, Z. Prognostic value of amphiregulin and epiregulin mrna expression in metastatic colorectal cancer patients. Oncotarget 2016, 7, 55890–55899. [Google Scholar] [CrossRef]
- Lee, Y.C.; Lee, Y.L.; Chuang, J.P.; Lee, J.C. Differences in survival between colon and rectal cancer from seer data. PLoS ONE 2013, 8, e78709. [Google Scholar] [CrossRef]
- Paschke, S.; Jafarov, S.; Staib, L.; Kreuser, E.D.; Maulbecker-Armstrong, C.; Roitman, M.; Holm, T.; Harris, C.C.; Link, K.H.; Kornmann, M. Are colon and rectal cancer two different tumor entities? A proposal to abandon the term colorectal cancer. Int. J. Mol. Sci. 2018, 19, 2577. [Google Scholar] [CrossRef]
- Sanz-Pamplona, R.; Cordero, D.; Berenguer, A.; Lejbkowicz, F.; Rennert, H.; Salazar, R.; Biondo, S.; Sanjuan, X.; Pujana, M.A.; Rozek, L.; et al. Gene expression differences between colon and rectum tumors. Clin. Cancer Res. 2011, 17, 7303–7312. [Google Scholar] [CrossRef]

| Author/Year | Study Design | Cancer Pathology | TNM Stage | Biological Specimen | Sample Size (Cancer, Control) | % Female (Cancer, Control) | Measurement Method | Reported Value(s) |
|---|---|---|---|---|---|---|---|---|
| Ahmed 2013 | CC | C | I–IV | F | 40, 20 | 42.5, 50 | Microarray, qPCR | E |
| Azizian 2015 | COH | R | NR | S | 42, 0 | 40.5, NA | qPCR | -- |
| Bovell 2013 | CC | CR | I–IV | T | 381, 381 | 50.7, 50.7 | qPCR | E |
| Brunet 2013 | CC | CR | III–IV | S, T | 42, 26 | 35.7, 50 | Microarray | E |
| Caritg 2016 | COH | C | III–IV | T | 69, 0 | 37.7, NA | qPCR | -- |
| Chen X 2015 | CC | CR | II–III | T | 11, 11 | 18.2, 18.2 | qPCR | E, OS |
| Chen W 2015 | CC | CR | I–IV | S | 100, 79 | 40, 44.9 | qPCR | AUROC, E |
| Cheng 2016 | COH | CR | I–IV | T | 544, 0 | 44.9, NA | qPCR | DFS, OS |
| de Groen 2015 | CC | CR | NR | T | 52, 48 | NR | qPCR | E |
| Hotchi 2013 | COH | R | III–IV | T | 43, 0 | 27.9, NA | qPCR | -- |
| Huang 2017 | CC | CR | NR | S, T | 70, 70 | NR | qPCR | E |
| Luo 2013 | CC | CR | I–IV | S | 180, 244 | 43.3, 57 | qPCR | AUROC, E |
| Molinari 2016 | COH | R | II–IV | T | 108, 0 | 30.6, NA | qPCR | -- |
| Motoyama 2009 | CC | CR | I–III | T | 67, 67 | NR | qPCR | E |
| Ozcan 2016 | CC | CR | II–IV | T | 54, 42 | 37, NR | qPCR | E |
| Pellatt 2016 | CC | CR | NR | T | 1141, 812 | 45.2, 45.7 | qPCR | E |
| Riordan 2012 | CC | R | I–III | T | 17, 17 | NR | qPCR | E |
| Ristau 2014 | COH | CR | I–IV | T | 35, 0 | 40, NA | qPCR | -- |
| Rotelli 2015 | CC | CR | I–III | F, T | 20, 20 | 35, NR | qPCR | E |
| Schetter 2008 | COH | C | I–IV | T | 197, 197 | 38.1, 38.1 | Microarray, qPCR | E, OS |
| Tsuchida 2011 | CC | C | I–IV | T | 13, 9 | NR | qPCR | E |
| Valladares 2011 | COH | CR | I–IV | T | 38, 0 | 34.2, NA | qPCR | DFS, OS |
| Volinia 2006 | CC | C | NR | T | 46, 8 | NR | Microarray | E |
| Wang 2010 | CC | CR | NR | T | 3, 3 | NR | Microarray | E |
| Xu 2015 | CC | CR | NR | T | 30, 30 | 36.7, 36.7 | qPCR | E |
| Yamada 2015 | CC | CR | NR | S | 160, 77 | 33.8, 45.5 | qPCR | E |
| Yantiss 2009 | CC | CR | I–IV | T | 69, 0 | 44.9, NA | qPCR | -- |
| Yau 2016 | CC | CR | I–IV | F, T | 198, 198 | 41.1, 57.6 | qPCR | AUROC, E |
| Zekri 2016 | CC | CR | NR | S | 130, 24 | 38.5, 37.5 | Microarray | AUROC, E |
| Zhang GJ 2014 | COH | CR | I–IV | T | 86, 86 | 38.4, 38.4 | qPCR | E, OS |
| Zhang J 2014 | COH | CR | III–IV | S | 253, 0 | 44.7, 0 | Microarray | -- |
| Zhang JX 2013 | CC | C | II–IV | T | 735, 735 | 41.3, 41.3 | Microarray | -- |
| Author/Year | miR-20a (Expression in CRC vs. Control) | Fold Change (CRC vs. Control) | p-Value | CRC Patients |
|---|---|---|---|---|
| Feces | ||||
| Ahmed 2013 | up | NR | NR | 40 |
| Rotelli 2015 | up | 1.1461 | p < 0.0001 | 20 |
| Yau 2016 | up | 2.063 | p = 0.0065 | 198 |
| Feces weighted fold change ± SD: 1.99 ± 0.65 | ||||
| Serum | ||||
| Brunet 2013 | up | 1.378 | p = 0.593 | 42 |
| Chen W 2015 | up | 1.64 | p = 0.038 | 100 |
| Huang 2017 | up | 2.6 | p < 0.05 | 70 |
| Luo 2013 | up | NR | p = 0.001 | 180 |
| Yamada 2015 | down | 0.98 | p = 0.834 | 160 |
| Zekri 2016 | up | 2.385 | p = 0.017565 | 130 |
| Serum weighted fold change ± SD: 1.73 ± 0.68 | ||||
| Tumor | ||||
| Bovell 2013 | up | 2.05 | NR | 345 |
| Chen X 2015 | up | 2 | p = 0.004 | 11 |
| de Groen 2015 | up | NR | p < 0.01 | 52 |
| Motoyama 2009 | up | 2.875 | p < 0.05 | 67 |
| Ozcan 2016 | up | 8.9509 | p = 0.002625 | 54 |
| Pellatt 2016 | up | 3.765 | NR | 1141 |
| Riordan 2012 | up | 2.2 | p < 0.00001 | 17 |
| Schetter 2008 | up | 2.3 | p < 0.001 | 197 |
| Tsuchida 2011 | up | NR | p < 0.05 | 13 |
| Volinia 2006 | up | NR | p = 0.0025 | 46 |
| Wang 2010 | up | 2.41 | NR | 3 |
| Xu 2015 | up | NR | p < 0.001 | 30 |
| Zhang GJ 2014 | up | NR | p < 0.001 | 86 |
| Tumor weighted fold change ± SD: 2.86 ± 2.61 | ||||
| Overall weighted fold change ± SD: 2.35 ± 1.89 | ||||
| Criterion | Mean | SD |
|---|---|---|
| 1. Was the research question clearly stated? | 1.92 | 0.27 |
| 2. Were the inclusion and exclusion criteria clearly stated? | 1.44 | 0.71 |
| 3. Were the subjects in the study representative of the pathological population? | 1.68 | 0.59 |
| 4. Were the main findings of the study clearly described? | 1.77 | 0.46 |
| 5. Was a control group included? Did it consist of the non-tumor specimen from healthy age- and gender-matched subjects? | 1.78 | 0.49 |
| 6. Were diagnostic or prognostic measures clearly defined (e.g., TNM stage, five-year survival rate, etc.) | 1.27 | 0.70 |
| 7. Were samples collected from a relevant source (tumor tissue, blood, or feces) in a manner to prevent degradation and contamination? | 1.81 | 0.39 |
| 8. Was miRNA expression measured with a conventional, well-validated technique (e.g., miRNA-seq, microarray, qPCR, etc.)? | 2.00 | 0.00 |
| 9. Was a sample size justification via power analysis provided? | 0.13 | 0.45 |
| 10. Were potential confounders properly controlled in the analysis? | 1.27 | 0.45 |
| Total Score | 15.06 | 1.87 |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Moody, L.; Dvoretskiy, S.; An, R.; Mantha, S.; Pan, Y.-X. The Efficacy of miR-20a as a Diagnostic and Prognostic Biomarker for Colorectal Cancer: A Systematic Review and Meta-Analysis. Cancers 2019, 11, 1111. https://doi.org/10.3390/cancers11081111
Moody L, Dvoretskiy S, An R, Mantha S, Pan Y-X. The Efficacy of miR-20a as a Diagnostic and Prognostic Biomarker for Colorectal Cancer: A Systematic Review and Meta-Analysis. Cancers. 2019; 11(8):1111. https://doi.org/10.3390/cancers11081111
Chicago/Turabian StyleMoody, Laura, Svyatoslav Dvoretskiy, Ruopeng An, Suparna Mantha, and Yuan-Xiang Pan. 2019. "The Efficacy of miR-20a as a Diagnostic and Prognostic Biomarker for Colorectal Cancer: A Systematic Review and Meta-Analysis" Cancers 11, no. 8: 1111. https://doi.org/10.3390/cancers11081111
APA StyleMoody, L., Dvoretskiy, S., An, R., Mantha, S., & Pan, Y.-X. (2019). The Efficacy of miR-20a as a Diagnostic and Prognostic Biomarker for Colorectal Cancer: A Systematic Review and Meta-Analysis. Cancers, 11(8), 1111. https://doi.org/10.3390/cancers11081111






