Synergistic Effects of Melittin and Plasma Treatment: A Promising Approach for Cancer Therapy
Abstract
:1. Introduction
2. Materials and Methods
2.1. Reagents and Cell Lines
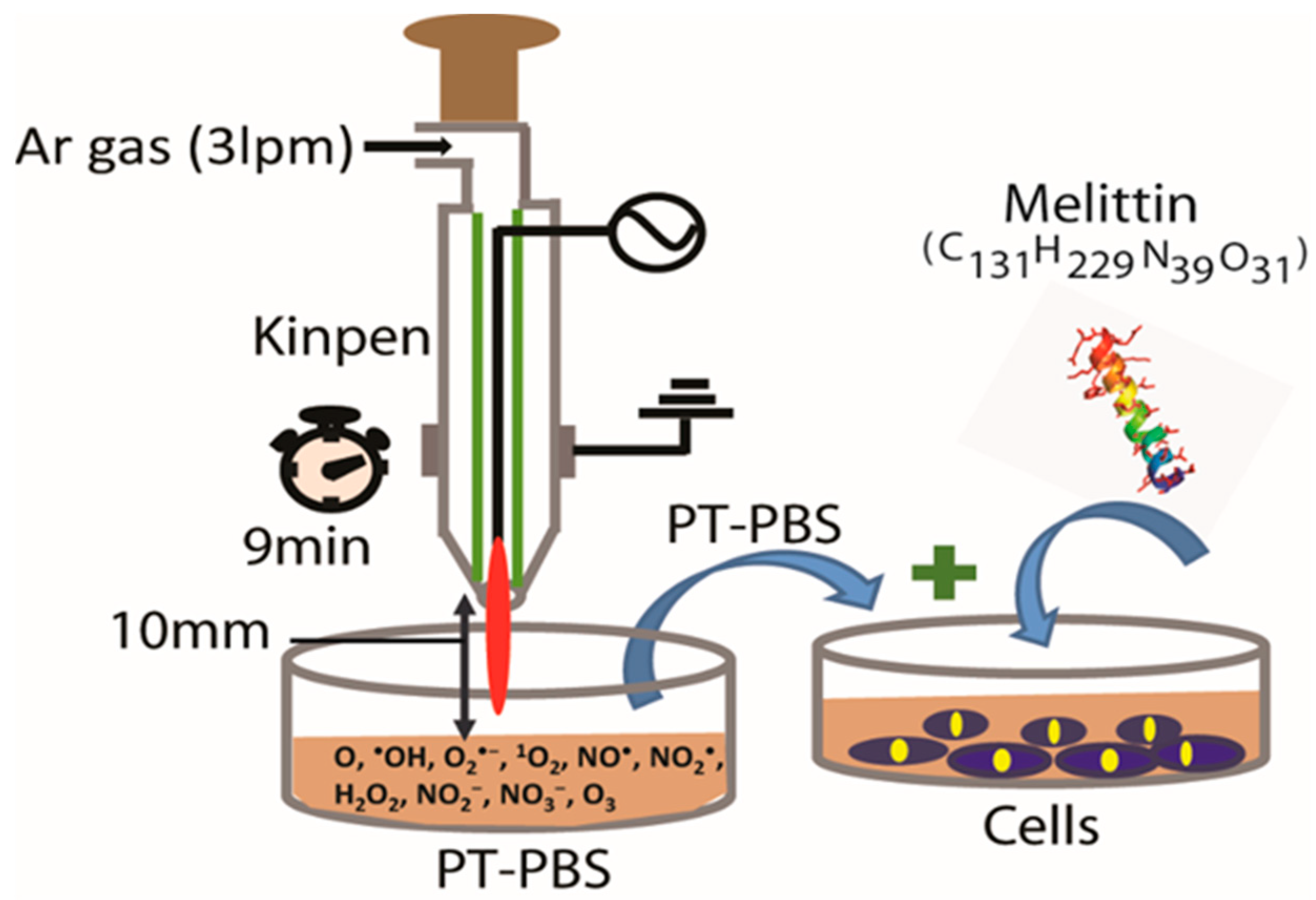
2.2. Plasma Device and Sample Preparation
2.3. Analysis of Cell Cytotoxicity
2.4. Estimation of Lipid Peroxidation
2.5. Chicken Chorioallantoic Membrane Assay (CAM Assay)
2.6. Immunohistochemical Analysis for Ki-67
2.7. Mass Spectrometry Analysis
2.8. Statistical Analysis
2.9. Computational Analysis
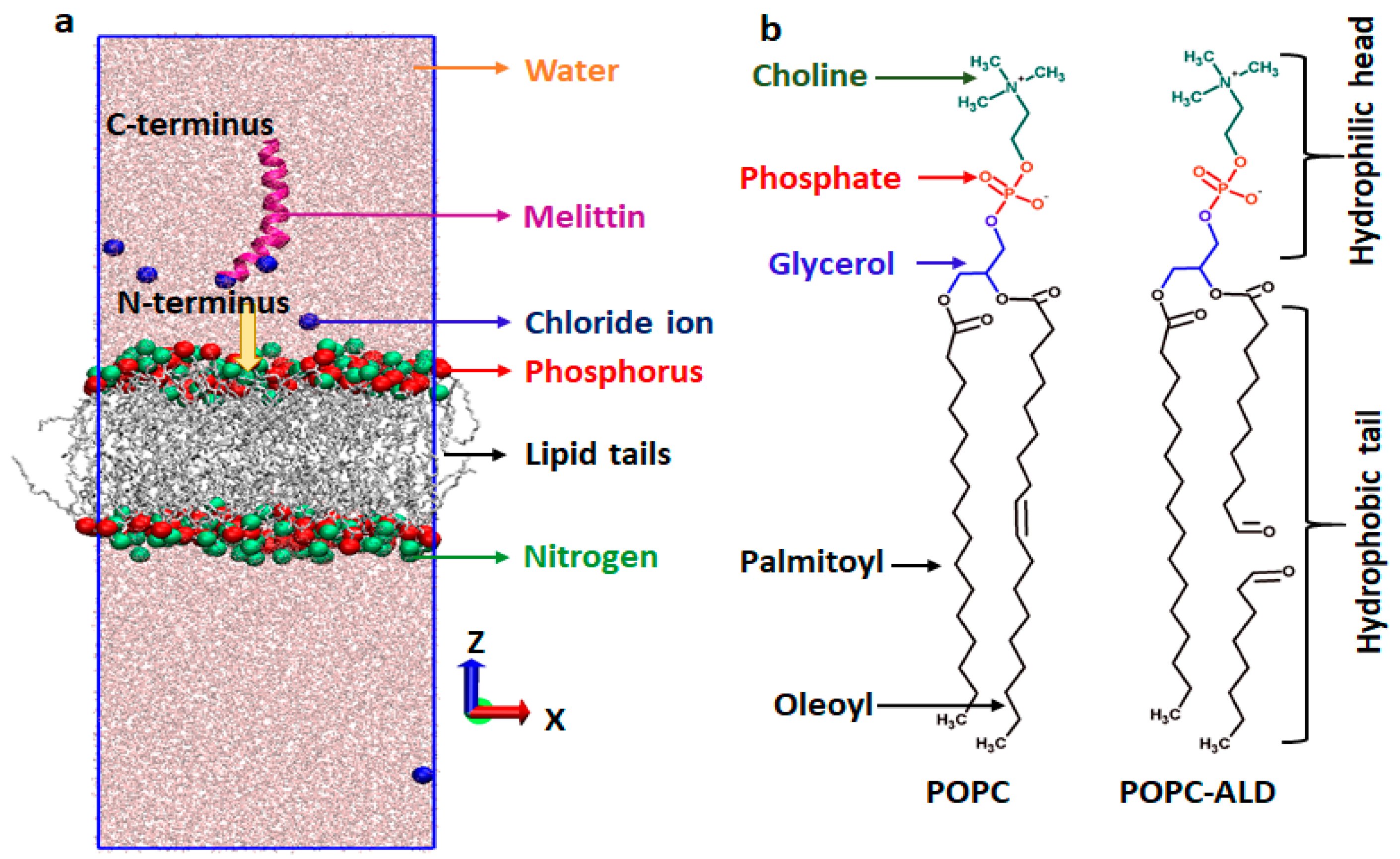
2.9.1. Simulation Setup
2.9.2. Umbrella Sampling
3. Results and Discussion
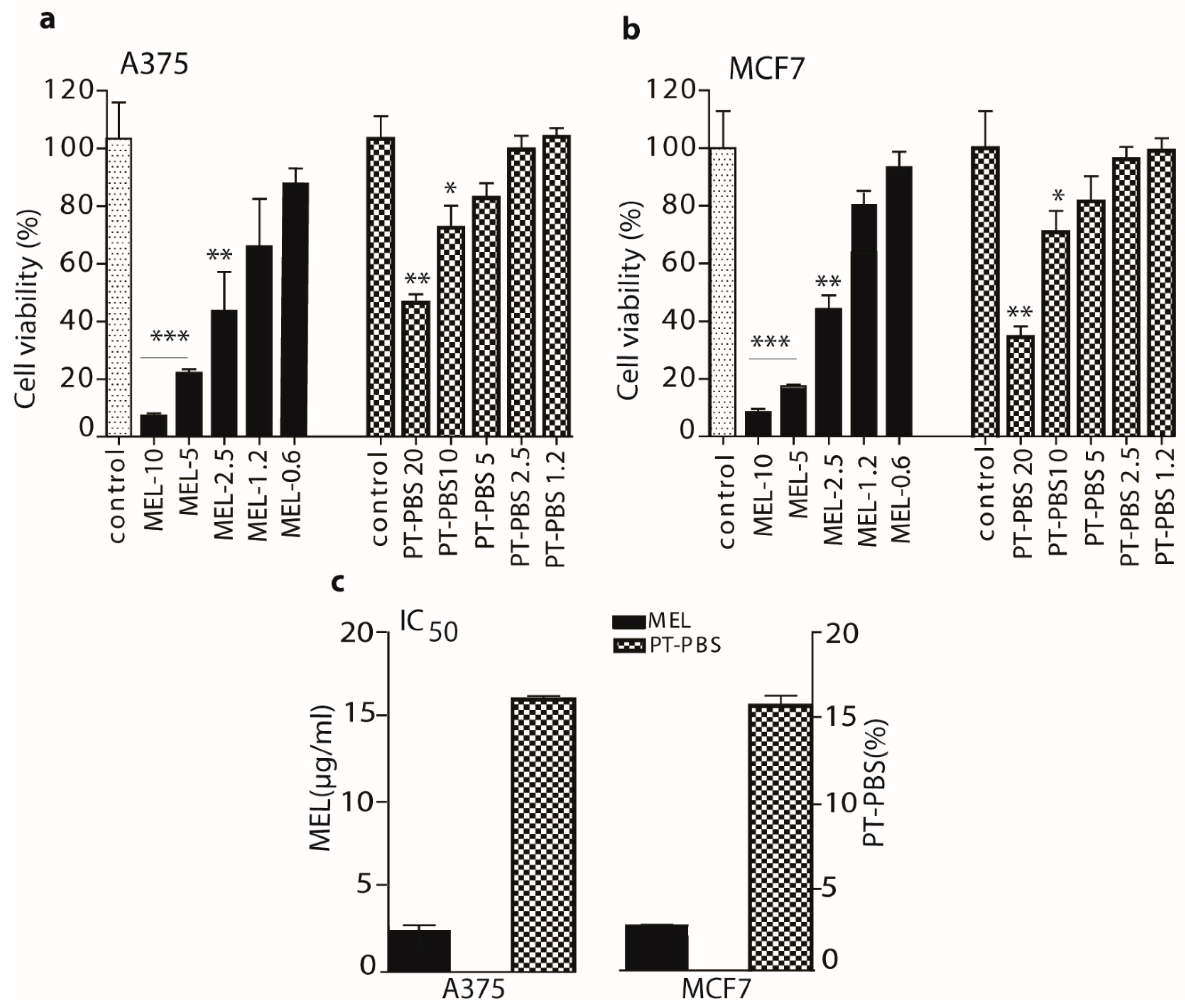
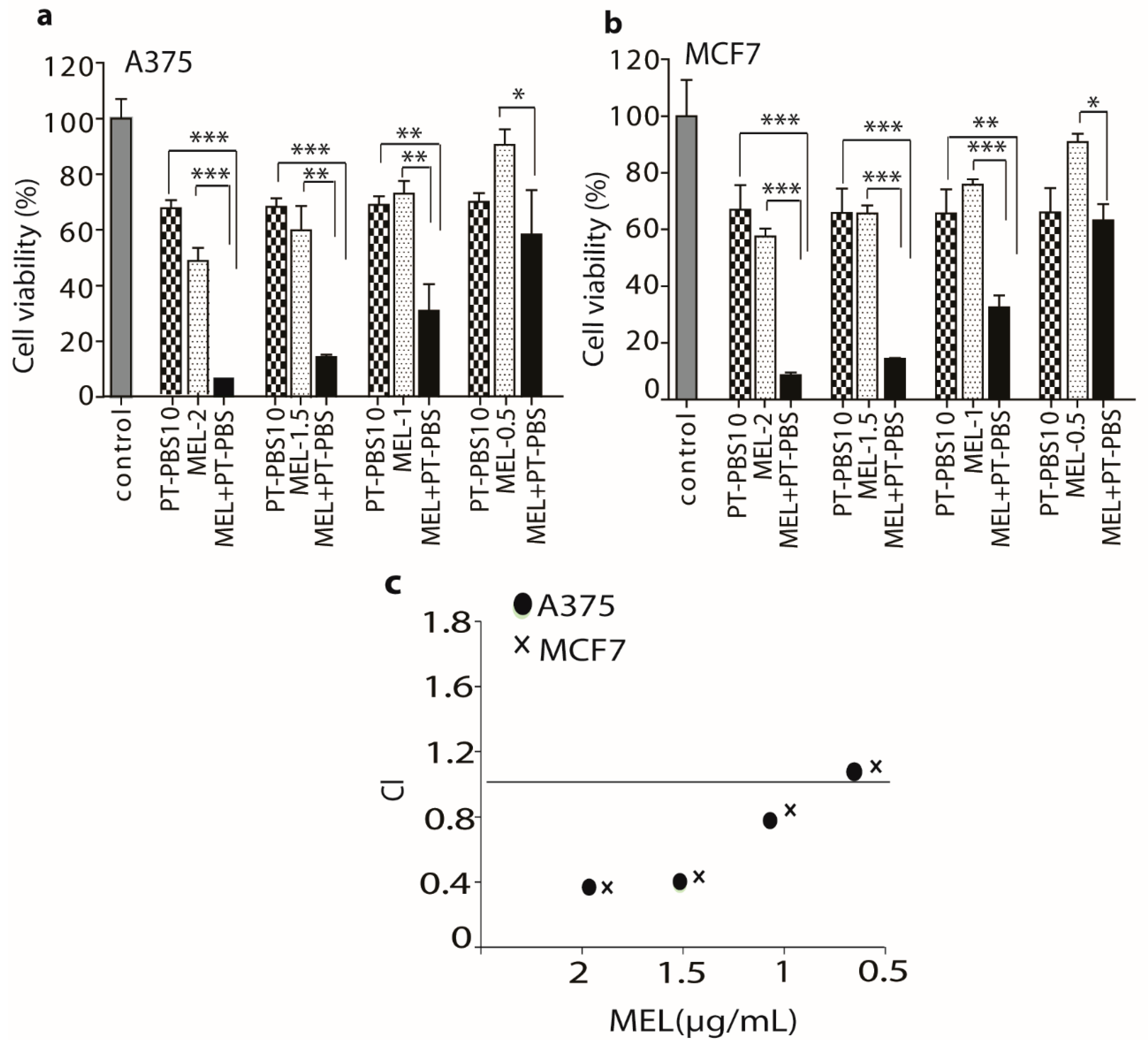
3.1. Effects of PT-PBS and MEL on Cell Viability and Dose Optimization
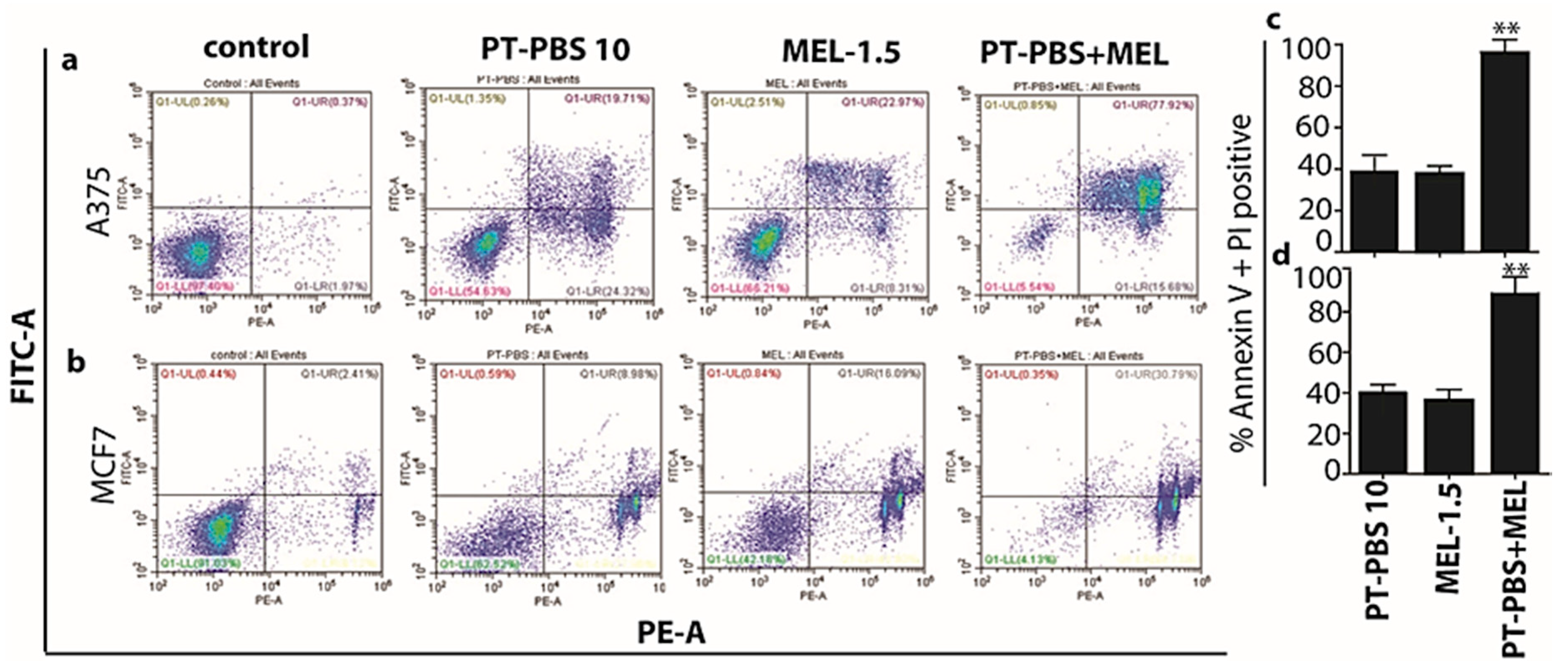
3.2. Influence of PT-PBS and MEL on Cell Death
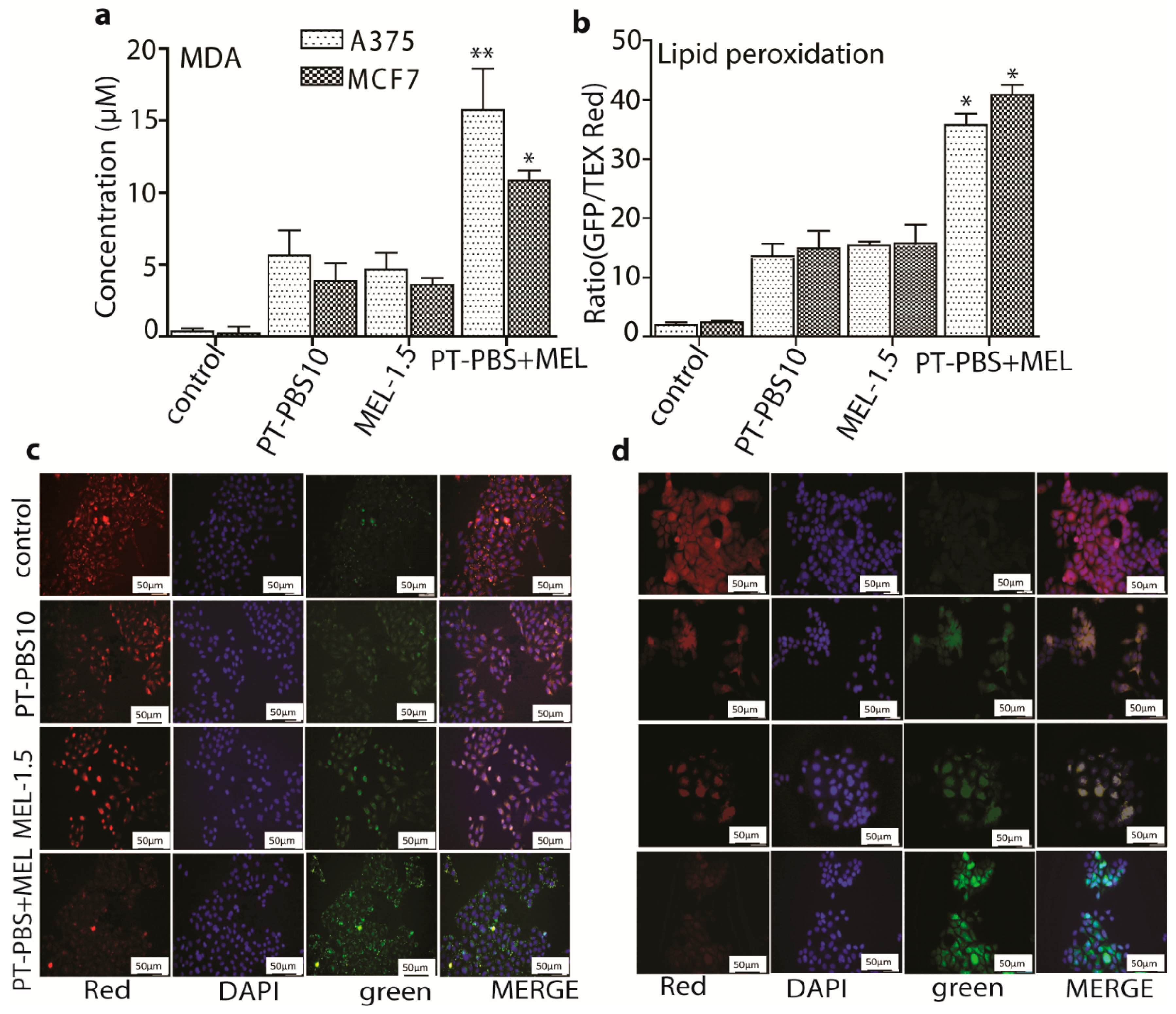
3.3. Effect of PT-PBS and MEL on Lipid Peroxidation
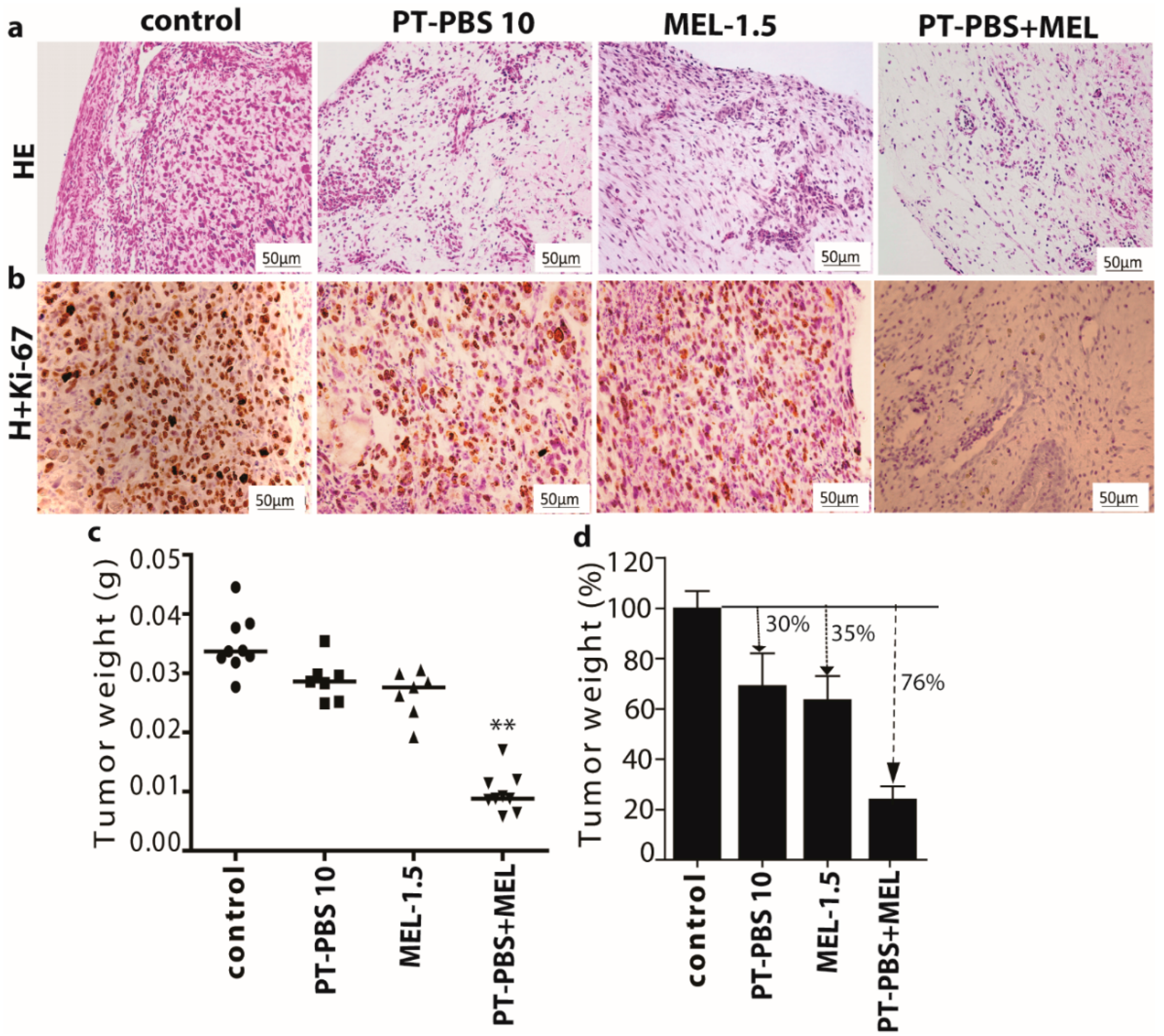
3.4. Effect of PT-PBS and MEL on Malignant Melanoma Cancer Tumors of the TUM-CAM Model
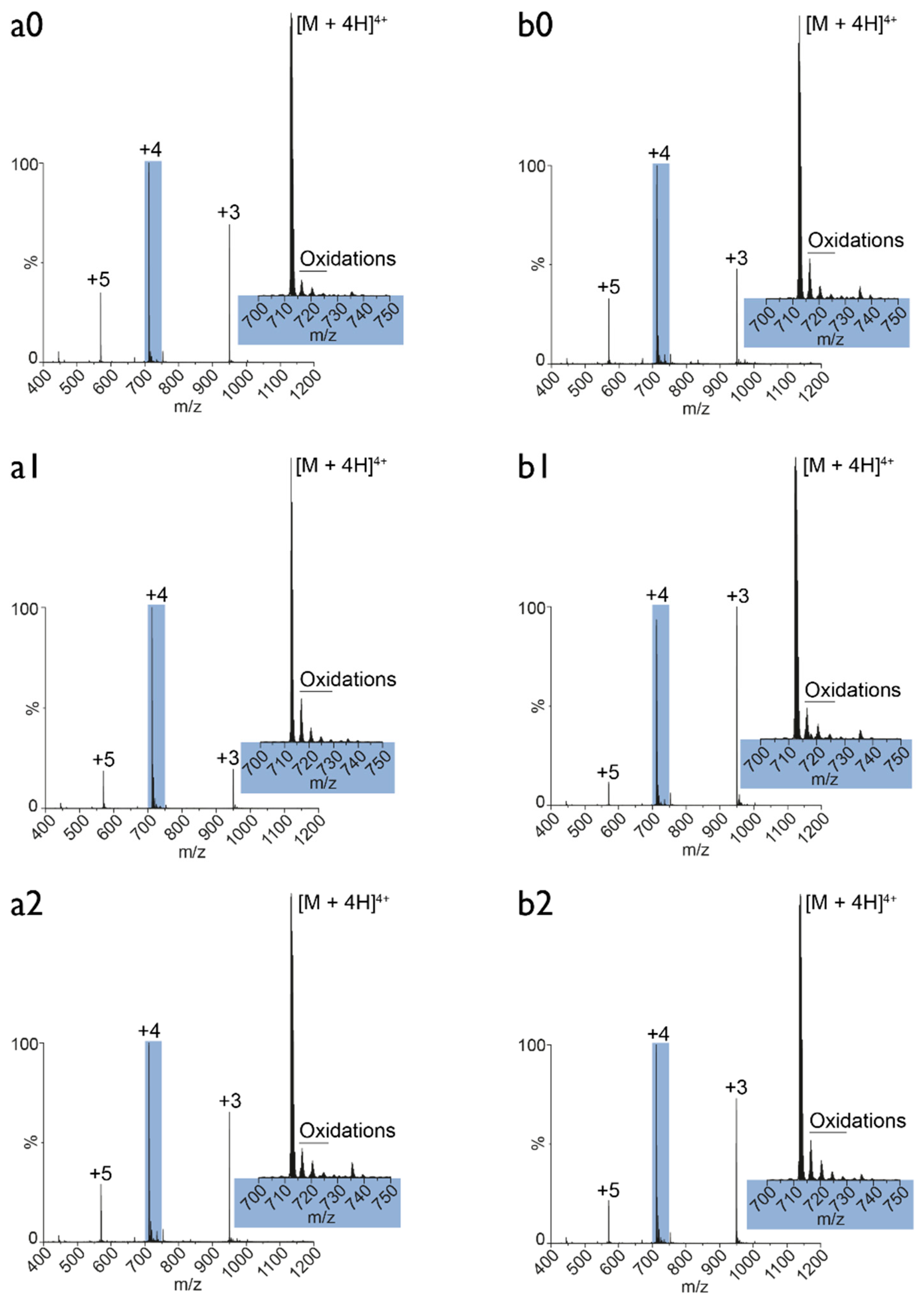
3.5. Plasma Oxidation of MEL: Mass Spectrometry Analysis
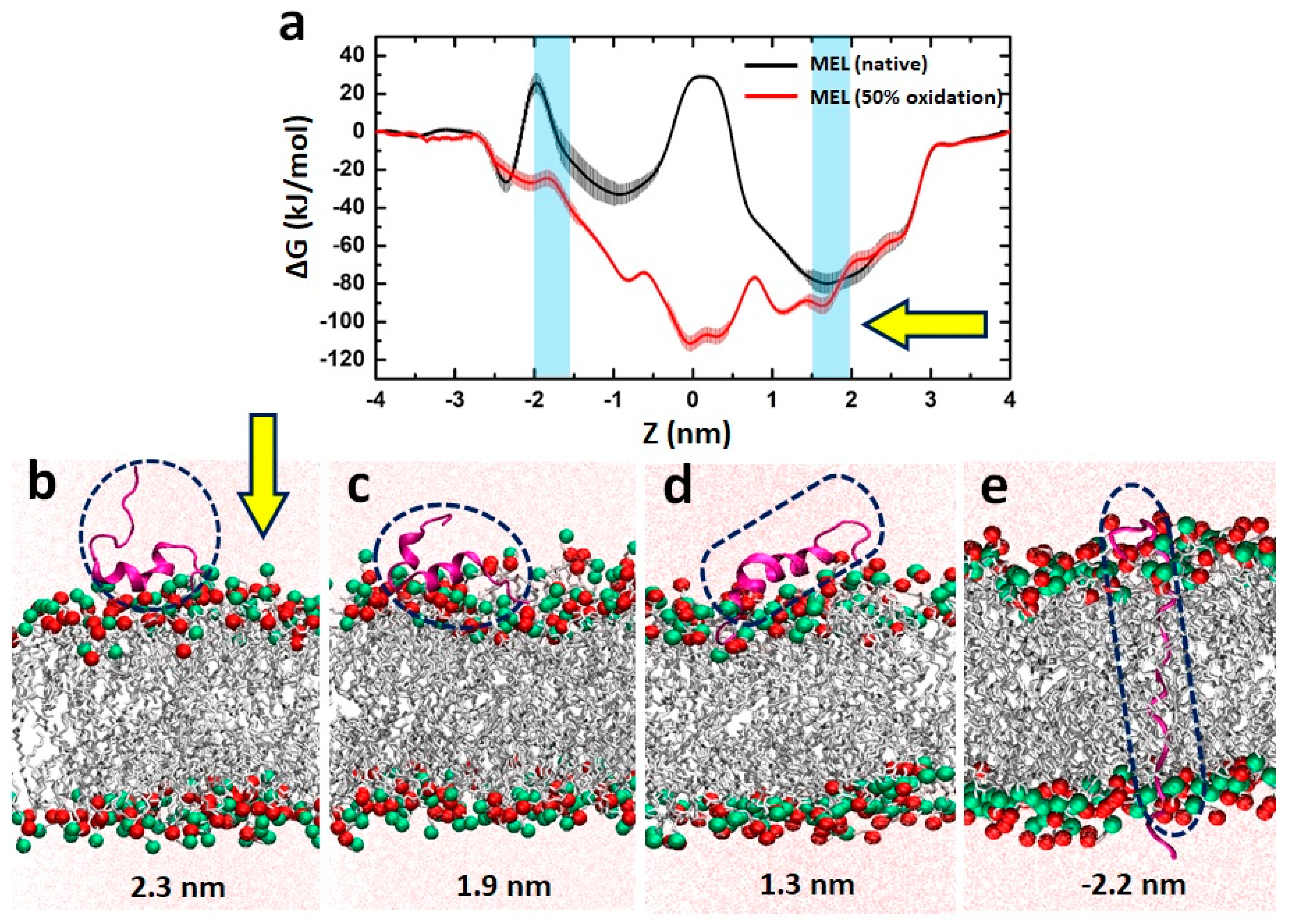
3.6. MEL Translocation across Native and Oxidized Phospholipid Membrane Revealed through MD Simulations
4. Conclusions
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Soman, N.R.; Baldwin, S.L.; Hu, G.; Marsh, J.N.; Lanza, G.M.; Heuser, J.E.; Arbeit, J.M.; Wickline, S.A.; Schlesinger, P.H. Molecularly targeted nanocarriers deliver the cytolytic peptide melittin specifically to tumor cells in mice, reducing tumor growth. J. Clin. Investig. 2009, 119, 2830–2842. [Google Scholar] [CrossRef] [PubMed]
- Leuschner, C.; Hansel, W. Membrane disrupting lytic peptides for cancer treatments. Curr. Pharm. Des. 2004, 10, 2299–2310. [Google Scholar] [CrossRef] [PubMed]
- Hansel, W.; Enright, F.; Leuschner, C. Destruction of breast cancers and their metastases by lytic peptide conjugates in vitro and in vivo. Mol. Cell. Endocrinol. 2007, 260, 183–189. [Google Scholar] [CrossRef] [PubMed]
- Kumar, C.S.; Leuschner, C.; Doomes, E.; Henry, L.; Juban, M.; Hormes, J. Efficacy of lytic peptide-bound magnetite nanoparticles in destroying breast cancer cells. J. Nanosci. Nanotechnol. 2004, 4, 245–249. [Google Scholar] [PubMed]
- Zarrinnahad, H.; Mahmoodzadeh, A.; Hamidi, M.P.; Mahdavi, M.; Moradi, A.; Bagheri, K.P.; Shahbazzadeh, D. Apoptotic effect of melittin purified from iranian honey bee venom on human cervical cancer hela cell line. Int. J. Pept. Res. Ther. 2018, 24, 563–570. [Google Scholar] [CrossRef] [PubMed]
- Jo, M.; Park, M.H.; Kollipara, P.S.; An, B.J.; Song, H.S.; Han, S.B.; Kim, J.H.; Song, M.J.; Hong, J.T. Anti-cancer effect of bee venom toxin and melittin in ovarian cancer cells through induction of death receptors and inhibition of JAK2/STAT3 pathway. Toxicol. Appl. Pharmacol. 2012, 258, 72–81. [Google Scholar] [CrossRef] [PubMed]
- Moon, D.-O.; Park, S.-Y.; Choi, Y.H.; Kim, N.D.; Lee, C.; Kim, G.-Y. Melittin induces bcl-2 and caspase-3-dependent apoptosis through downregulation of akt phosphorylation in human leukemic u937 cells. Toxicon 2008, 51, 112–120. [Google Scholar] [CrossRef] [PubMed]
- Park, M.H.; Choi, M.S.; Kwak, D.H.; Oh, K.W.; Yoon, D.Y.; Han, S.B.; Song, H.S.; Song, M.J.; Hong, J.T. Anti-cancer effect of bee venom in prostate cancer cells through activation of caspase pathway via inactivation of nf-κb. Prostate 2011, 71, 801–812. [Google Scholar] [CrossRef]
- Zhang, S.F.; Chen, Z. Melittin exerts an antitumor effect on nonsmall cell lung cancer cells. Mol. Med. Rep. 2017, 16, 3581–3586. [Google Scholar] [CrossRef]
- Clague, M.J.; Cherry, R.J. Comparison of p25 presequence peptide and melittin. Red blood cell haemolysis and band 3 aggregation. Biochem. J. 1988, 252, 791–794. [Google Scholar] [CrossRef] [Green Version]
- Voss, J.; Birmachu, W.; Hussey, D.M.; Thomas, D.D. Effects of melittin on molecular dynamics and calcium-atpase activity in sarcoplasmic reticulum membranes: Time-resolved optical anisotropy. Biochemistry 1991, 30, 7498–7506. [Google Scholar] [CrossRef] [PubMed]
- Sumikura, H.; Andersen, O.K.; Drewes, A.M.; Arendt-Nielsen, L. A comparison of hyperalgesia and neurogenic inflammation induced by melittin and capsaicin in humans. Neurosci. Lett. 2003, 337, 147–150. [Google Scholar] [CrossRef]
- Oktiansyah, R.; Juliandi, B.; Widayati, K.A.; Juniantito, V. Neuronal cell death and mouse (Mus musculus) behaviour induced by bee venom. Trop. Life Sci. Res. 2018, 29, 1–11. [Google Scholar] [CrossRef] [PubMed]
- Maher, S.; McClean, S. Melittin exhibits necrotic cytotoxicity in gastrointestinal cells which is attenuated by cholesterol. Biochem. Pharmacol. 2008, 75, 1104–1114. [Google Scholar] [CrossRef] [PubMed]
- Gajski, G.; Garaj-Vrhovac, V. Melittin: A lytic peptide with anticancer properties. Environ. Toxicol. Pharmacol. 2013, 36, 697–705. [Google Scholar] [CrossRef] [PubMed]
- Suttmann, H.; Retz, M.; Paulsen, F.; Harder, J.; Zwergel, U.; Kamradt, J.; Wullich, B.; Unteregger, G.; Stockle, M.; Lehmann, J. Antimicrobial peptides of the cecropin-family show potent antitumor activity against bladder cancer cells. BMC Urol. 2008, 8, 5. [Google Scholar] [CrossRef] [PubMed]
- Pan, H.; Soman, N.R.; Schlesinger, P.H.; Lanza, G.M.; Wickline, S.A. Cytolytic peptide nanoparticles (‘nanobees’) for cancer therapy. Wiley Interdiscip. Rev. Nanomed. Nanobiotechnol. 2011, 3, 318–327. [Google Scholar] [CrossRef]
- Bei, C.; Bindu, T.; Remant, K.C.; Peisheng, X. Dual secured nano-melittin for the safe and effective eradication of cancer cells. J. Mater. Chem. B 2015, 3, 25–29. [Google Scholar]
- Oršolić, N. Potentiation of bleomycin lethality in hela and v79 cells by bee venom. Arch. Ind. Hyg. Toxicol. 2009, 60, 317–326. [Google Scholar] [CrossRef]
- Alonezi, S.; Tusiimire, J.; Wallace, J.; Dufton, M.; Parkinson, J.; Young, L.; Clements, C.; Park, J.-K.; Jeon, J.-W.; Ferro, V. Metabolomic profiling of the synergistic effects of melittin in combination with cisplatin on ovarian cancer cells. Metabolites 2017, 7, 14. [Google Scholar] [CrossRef]
- Alizadehnohi, M.; Nabiuni, M.; Nazari, Z.; Safaeinejad, Z.; Irian, S. The synergistic cytotoxic effect of cisplatin and honey bee venom on human ovarian cancer cell line A2780cp. J. Venom Res. 2012, 3, 22. [Google Scholar]
- Rady, I.; Siddiqui, I.A.; Rady, M.; Mukhtar, H. Melittin, a major peptide component of bee venom, and its conjugates in cancer therapy. Cancer Lett. 2017, 402, 16–31. [Google Scholar] [CrossRef] [Green Version]
- Biswaro, L.S.; da Costa Sousa, M.G.; Rezende, T.; Dias, S.C.; Franco, O.L. Antimicrobial peptides and nanotechnology, recent advances and challenges. Front. Microbiol. 2018, 9, 855. [Google Scholar] [CrossRef]
- Kumar, N.; Attri, P.; Dewilde, S.; Bogaerts, A. Inactivation of human pancreatic ductal adenocarcinoma with atmospheric plasma treated media and water: A comparative study. J. Phys. D Appl. Phys. 2018, 51, 255401. [Google Scholar] [CrossRef]
- Kumar, N.; Park, J.H.; Jeon, S.N.; Park, B.S.; Choi, E.H.; Attri, P. The action of microsecond-pulsed plasma-activated media on the inactivation of human lung cancer cells. J. Phys. D Appl. Phys. 2016, 49, 115401. [Google Scholar] [CrossRef]
- Siu, A.; Volotskova, O.; Cheng, X.; Khalsa, S.S.; Bian, K.; Murad, F.; Keidar, M.; Sherman, J.H. Differential effects of cold atmospheric plasma in the treatment of malignant glioma. PLoS ONE 2015, 10, e0126313. [Google Scholar] [CrossRef]
- Lunov, O.; Zablotskii, V.; Churpita, O.; Lunova, M.; Jirsa, M.; Dejneka, A.; Kubinova, S. Chemically different non-thermal plasmas target distinct cell death pathways. Sci. Rep. 2017, 7, 600. [Google Scholar] [CrossRef]
- Kumar, N.; Attri, P.; Choi, E.H.; Sup Uhm, H. Influence of water vapour with non-thermal plasma jet on the apoptosis of sk-br-3 breast cancer cells. RSC Adv. 2015, 5, 14670–14677. [Google Scholar] [CrossRef]
- Shaw, P.; Kumar, N.; Kwak, H.S.; Park, J.H.; Uhm, H.S.; Bogaerts, A.; Choi, E.H.; Attri, P. Bacterial inactivation by plasma treated water enhanced by reactive nitrogen species. Sci. Rep. 2018, 8, 11268. [Google Scholar] [CrossRef]
- Furuta, R.; Kurake, N.; Ishikawa, K.; Takeda, K.; Hashizume, H.; Tanaka, H.; Kondo, H.; Sekine, M.; Hori, M. Intracellular responses to reactive oxygen and nitrogen species, and lipid peroxidation in apoptotic cells cultivated in plasma-activated medium. Plasma Process. Polym. 2017, 14, 1700123. [Google Scholar] [CrossRef]
- Kumar, N.; Attri, P.; Yadav, D.K.; Choi, J.; Choi, E.H.; Uhm, H.S. Induced apoptosis in melanocytes cancer cell and oxidation in biomolecules through deuterium oxide generated from atmospheric pressure non-thermal plasma jet. Sci. Rep. 2014, 4, 7589. [Google Scholar] [CrossRef] [Green Version]
- Irani, S.; Shahmirani, Z.; Atyabi, S.M.; Mirpoor, S. Induction of growth arrest in colorectal cancer cells by cold plasma and gold nanoparticles. Arch. Med. Sci. AMS 2015, 11, 1286–1295. [Google Scholar] [CrossRef]
- Cheng, X.; Murphy, W.; Recek, N.; Yan, D.; Cvelbar, U.; Vesel, A.; Mozetič, M.; Canady, J.; Keidar, M.; Sherman, J.H. Synergistic effect of gold nanoparticles and cold plasma on glioblastoma cancer therapy. J. Phys. D Appl. Phys. 2014, 47, 335402. [Google Scholar] [CrossRef]
- Soliman, W.; Bhattacharjee, S.; Kaur, K. Interaction of an antimicrobial peptide with a model lipid bilayer using molecular dynamics simulation. Langmuir 2009, 25, 6591–6595. [Google Scholar] [CrossRef]
- Shi, L.; Cembran, A.; Gao, J.; Veglia, G. Tilt and azimuthal angles of a transmembrane peptide: A comparison between molecular dynamics calculations and solid-state nmr data of sarcolipin in lipid membranes. Biophys. J. 2009, 96, 3648–3662. [Google Scholar] [CrossRef]
- Dorairaj, S.; Allen, T.W. On the thermodynamic stability of a charged arginine side chain in a transmembrane helix. Proc. Natl. Acad. Sci. USA 2007, 104, 4943–4948. [Google Scholar] [CrossRef] [Green Version]
- Manna, M.; Mukhopadhyay, C. Molecular dynamics simulations of the interactions of kinin peptides with an anionic popg bilayer. Langmuir 2011, 27, 3713–3722. [Google Scholar] [CrossRef]
- Andersson, M.; Ulmschneider, J.P.; Ulmschneider, M.B.; White, S.H. Conformational states of melittin at a bilayer interface. Biophys. J. 2013, 104, L12–L14. [Google Scholar] [CrossRef]
- Irudayam, S.J.; Pobandt, T.; Berkowitz, M.L. Free energy barrier for melittin reorientation from a membrane-bound state to a transmembrane state. J. Phys. Chem. B 2013, 117, 13457–13463. [Google Scholar] [CrossRef]
- Irudayam, S.J.; Berkowitz, M.L. Binding and reorientation of melittin in a popc bilayer: Computer simulations. Biochim. Biophys. Acta Biomembr. 2012, 1818, 2975–2981. [Google Scholar] [CrossRef]
- Bekeschus, S.; Schmidt, A.; Weltmann, K.-D.; von Woedtke, T. The plasma jet kinpen—A powerful tool for wound healing. Clin. Plasma Med. 2016, 4, 19–28. [Google Scholar] [CrossRef]
- Van Boxem, W.; Van der Paal, J.; Gorbanev, Y.; Vanuytsel, S.; Smits, E.; Dewilde, S.; Bogaerts, A. Anti-cancer capacity of plasma-treated pbs: Effect of chemical composition on cancer cell cytotoxicity. Sci. Rep. 2017, 7, 16478. [Google Scholar] [CrossRef]
- Reis, A.; Domingues, M.R.; Amado, F.M.; Ferrer-Correia, A.J.; Domingues, P. Separation of peroxidation products of diacyl-phosphatidylcholines by reversed-phase liquid chromatography-mass spectrometry. Biomed. Chromatogr. BMC 2005, 19, 129–137. [Google Scholar] [CrossRef]
- Van Der Spoel, D.; Lindahl, E.; Hess, B.; Groenhof, G.; Mark, A.E.; Berendsen, H.J. Gromacs: Fast, flexible, and free. J. Comput. Chem. 2005, 26, 1701–1718. [Google Scholar] [CrossRef]
- Chiu, S.-W.; Pandit, S.A.; Scott, H.L.; Jakobsson, E. An improved united atom force field for simulation of mixed lipid bilayers. J. Phys. Chem. B 2009, 113, 2748–2763. [Google Scholar] [CrossRef]
- Wong-Ekkabut, J.; Xu, Z.; Triampo, W.; Tang, I.M.; Tieleman, D.P.; Monticelli, L. Effect of lipid peroxidation on the properties of lipid bilayers: A molecular dynamics study. Biophys. J. 2007, 93, 4225–4236. [Google Scholar] [CrossRef]
- Martinez, L.; Andrade, R.; Birgin, E.G.; Martinez, J.M. Packmol: A package for building initial configurations for molecular dynamics simulations. J. Comput. Chem. 2009, 30, 2157–2164. [Google Scholar] [CrossRef]
- Van der Paal, J.; Neyts, E.C.; Verlackt, C.C.W.; Bogaerts, A. Effect of lipid peroxidation on membrane permeability of cancer and normal cells subjected to oxidative stress. Chem. Sci. 2016, 7, 489–498. [Google Scholar] [CrossRef]
- Parrinello, M. Polymorphic transitions in single crystals: A new molecular dynamics method. J. Appl. Phys. 1981, 52, 7182. [Google Scholar] [CrossRef]
- Hoover, W.G. Canonical dynamics: Equilibrium phase-space distributions. Phys. Rev. A 1985, 31, 1695–1697. [Google Scholar] [CrossRef] [Green Version]
- Essmann, U. A smooth particle mesh ewald potential. J. Chem. Phys. 1995, 103, 8577–8592. [Google Scholar] [CrossRef]
- Yang, L.; Harroun, T.A.; Weiss, T.M.; Ding, L.; Huang, H.W. Barrel-stave model or toroidal model? A case study on melittin pores. Biophys. J. 2001, 81, 1475–1485. [Google Scholar] [CrossRef]
- Kastner, J. Umbrella sampling. Wiley Interdiscip. Rev. Comput. Mol. Sci. 2011, 1, 932–942. [Google Scholar] [CrossRef]
- Hernandez, J.L.; Padilla, L.; Dakhel, S.; Coll, T.; Hervas, R.; Adan, J.; Masa, M.; Mitjans, F.; Martinez, J.M.; Coma, S.; et al. Therapeutic targeting of tumor growth and angiogenesis with a novel anti-s100a4 monoclonal antibody. PLoS ONE 2013, 8, e72480. [Google Scholar] [CrossRef]
- Reynolds, C.P.; Maurer, B.J. Evaluating response to antineoplastic drug combinations in tissue culture models. Methods Mol. Med. 2005, 110, 173–183. [Google Scholar]
- Tu, W.C.; Wu, C.C.; Hsieh, H.L.; Chen, C.Y.; Hsu, S.L. Honeybee venom induces calcium-dependent but caspase-independent apoptotic cell death in human melanoma A2058 cells. Toxicon 2008, 52, 318–329. [Google Scholar] [CrossRef]
- Vermeylen, S.; De Waele, J.; Vanuytsel, S.; De Backer, J.; Van der Paal, J.; Ramakers, M.; Leyssens, K.; Marcq, E.; Van Audenaerde, J.; Smits, E.L.J.; et al. Cold atmospheric plasma treatment of melanoma and glioblastoma cancer cells. Plasma Process. Polym. 2016, 13, 1195–1205. [Google Scholar] [CrossRef]
- Yang, W.S.; Stockwell, B.R. Ferroptosis: Death by lipid peroxidation. Trends Cell Biol. 2016, 26, 165–176. [Google Scholar] [CrossRef]
- Uchida, K.; Shiraishi, M.; Naito, Y.; Torii, Y.; Nakamura, Y.; Osawa, T. Activation of stress signaling pathways by the end product of lipid peroxidation. 4-hydroxy-2-nonenal is a potential inducer of intracellular peroxide production. J. Biol. Chem. 1999, 274, 2234–2242. [Google Scholar] [CrossRef]
- Boys, B.L.; Kuprowski, M.C.; Noel, J.J.; Konermann, L. Protein oxidative modifications during electrospray ionization: Solution phase electrochemistry or corona discharge-induced radical attack? Anal. Chem. 2009, 81, 4027–4034. [Google Scholar] [CrossRef]
- Wong, J.W.; Maleknia, S.D.; Downard, K.M. Hydroxyl radical probe of the calmodulin-melittin complex interface by electrospray ionization mass spectrometry. J. Am. Soc. Mass Spectrom. 2005, 16, 225–233. [Google Scholar] [CrossRef] [Green Version]
- Lyu, Y.; Xiang, N.; Zhu, X.; Narsimhan, G. Potential of mean force for insertion of antimicrobial peptide melittin into a pore in mixed dopc/dopg lipid bilayer by molecular dynamics simulation. J. Chem. Phys. 2017, 146, 155101. [Google Scholar] [CrossRef]
- Lin, L.; Wang, L.L.; Liu, Y.D.; Xu, C.; Tu, Y.; Zhou, J.Y. Non-thermal plasma inhibits tumor growth and proliferation and enhances the sensitivity to radiation in vitro and in vivo. Oncol. Rep. 2018, 40, 3405–3415. [Google Scholar] [CrossRef]
- Zucker, S.N.; Zirnheld, J.; Bagati, A.; DiSanto, T.M.; Des Soye, B.; Wawrzyniak, J.A.; Etemadi, K.; Nikiforov, M.; Berezney, R. Preferential induction of apoptotic cell death in melanoma cells as compared with normal keratinocytes using a non-thermal plasma torch. Cancer Biol. Ther. 2012, 13, 1299–1306. [Google Scholar] [CrossRef]
- Utsumi, F.; Kajiyama, H.; Nakamura, K.; Tanaka, H.; Hori, M.; Kikkawa, F. Selective cytotoxicity of indirect nonequilibrium atmospheric pressure plasma against ovarian clear-cell carcinoma. SpringerPlus 2014, 3, 398. [Google Scholar] [CrossRef]
- Kim, S.J.; Chung, T.H. Cold atmospheric plasma jet-generated rons and their selective effects on normal and carcinoma cells. Sci. Rep. 2016, 6, 20332. [Google Scholar] [CrossRef]
- Bogaerts, A.; Yusupov, M.; Razzokov, J.; Van der Paal, J. Plasma for cancer treatment: How can rons penetrate through the cell membrane? Answers from computer modeling. Front. Chem. Sci. Eng. 2019, 13, 253–263. [Google Scholar] [CrossRef]
- Van der Paal, J.; Verheyen, C.; Neyts, E.C.; Bogaerts, A. Hampering effect of cholesterol on the permeation of reactive oxygen species through phospholipids bilayer: Possible explanation for plasma cancer selectivity. Sci. Rep. 2017, 7, 39526. [Google Scholar] [CrossRef]
- Cairns, R.A.; Harris, I.S.; Mak, T.W. Regulation of cancer cell metabolism. Nat. Rev. Cancer 2011, 11, 85–95. [Google Scholar] [CrossRef] [Green Version]
- Liedtke, K.R.; Diedrich, S.; Pati, O.; Freund, E.; Flieger, R.; Heidecke, C.D.; Partecke, L.I.; Bekeschus, S. Cold physical plasma selectively elicits apoptosis in murine pancreatic cancer cells in vitro and in ovo. Anticancer Res. 2018, 38, 5655–5663. [Google Scholar] [CrossRef]
- Privat-Maldonado, A.; Gorbanev, Y.; Dewilde, S.; Smits, E.; Bogaerts, A. Reduction of human glioblastoma spheroids using cold atmospheric plasma: The combined effect of short- and long-lived reactive species. Cancers 2018, 10, 394. [Google Scholar] [CrossRef]
- Radi, R.; Beckman, J.S.; Bush, K.M.; Freeman, B.A. Peroxynitrite-induced membrane lipid peroxidation: The cytotoxic potential of superoxide and nitric oxide. Arch. Biochem. Biophys. 1991, 288, 481–487. [Google Scholar] [CrossRef]
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Shaw, P.; Kumar, N.; Hammerschmid, D.; Privat-Maldonado, A.; Dewilde, S.; Bogaerts, A. Synergistic Effects of Melittin and Plasma Treatment: A Promising Approach for Cancer Therapy. Cancers 2019, 11, 1109. https://doi.org/10.3390/cancers11081109
Shaw P, Kumar N, Hammerschmid D, Privat-Maldonado A, Dewilde S, Bogaerts A. Synergistic Effects of Melittin and Plasma Treatment: A Promising Approach for Cancer Therapy. Cancers. 2019; 11(8):1109. https://doi.org/10.3390/cancers11081109
Chicago/Turabian StyleShaw, Priyanka, Naresh Kumar, Dietmar Hammerschmid, Angela Privat-Maldonado, Sylvia Dewilde, and Annemie Bogaerts. 2019. "Synergistic Effects of Melittin and Plasma Treatment: A Promising Approach for Cancer Therapy" Cancers 11, no. 8: 1109. https://doi.org/10.3390/cancers11081109
APA StyleShaw, P., Kumar, N., Hammerschmid, D., Privat-Maldonado, A., Dewilde, S., & Bogaerts, A. (2019). Synergistic Effects of Melittin and Plasma Treatment: A Promising Approach for Cancer Therapy. Cancers, 11(8), 1109. https://doi.org/10.3390/cancers11081109






