Gene Expression Comparison between the Lymph Node-Positive and -Negative Reveals a Peculiar Immune Microenvironment Signature and a Theranostic Role for WNT Targeting in Pancreatic Ductal Adenocarcinoma: A Pilot Study
Abstract
1. Introduction
2. Results
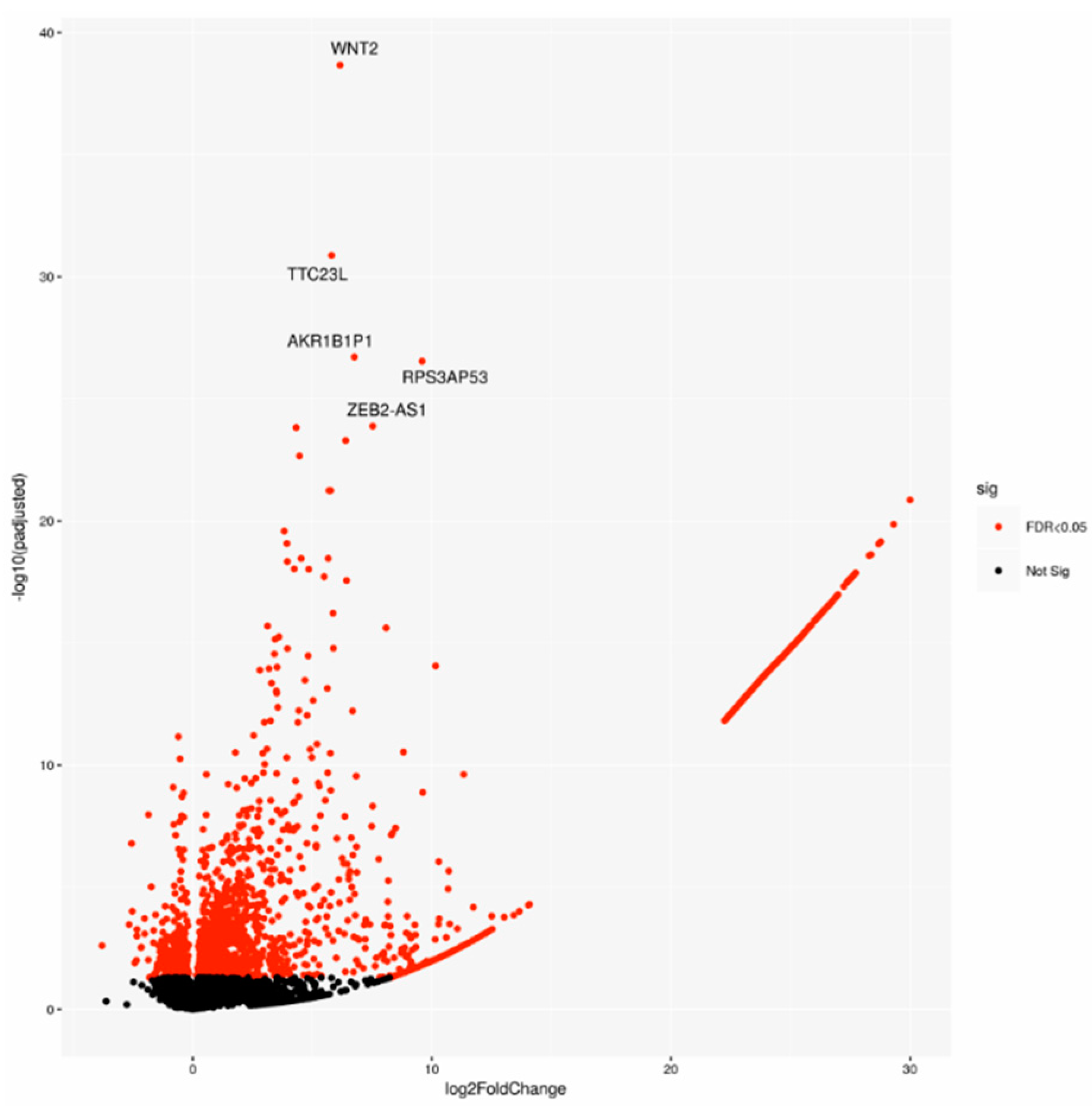
2.1. Gene Expression Analysis Comparing PDAC Patient-Derived Specimen Identified a Characteristic Biological Signature Depending on Lymph Node Status
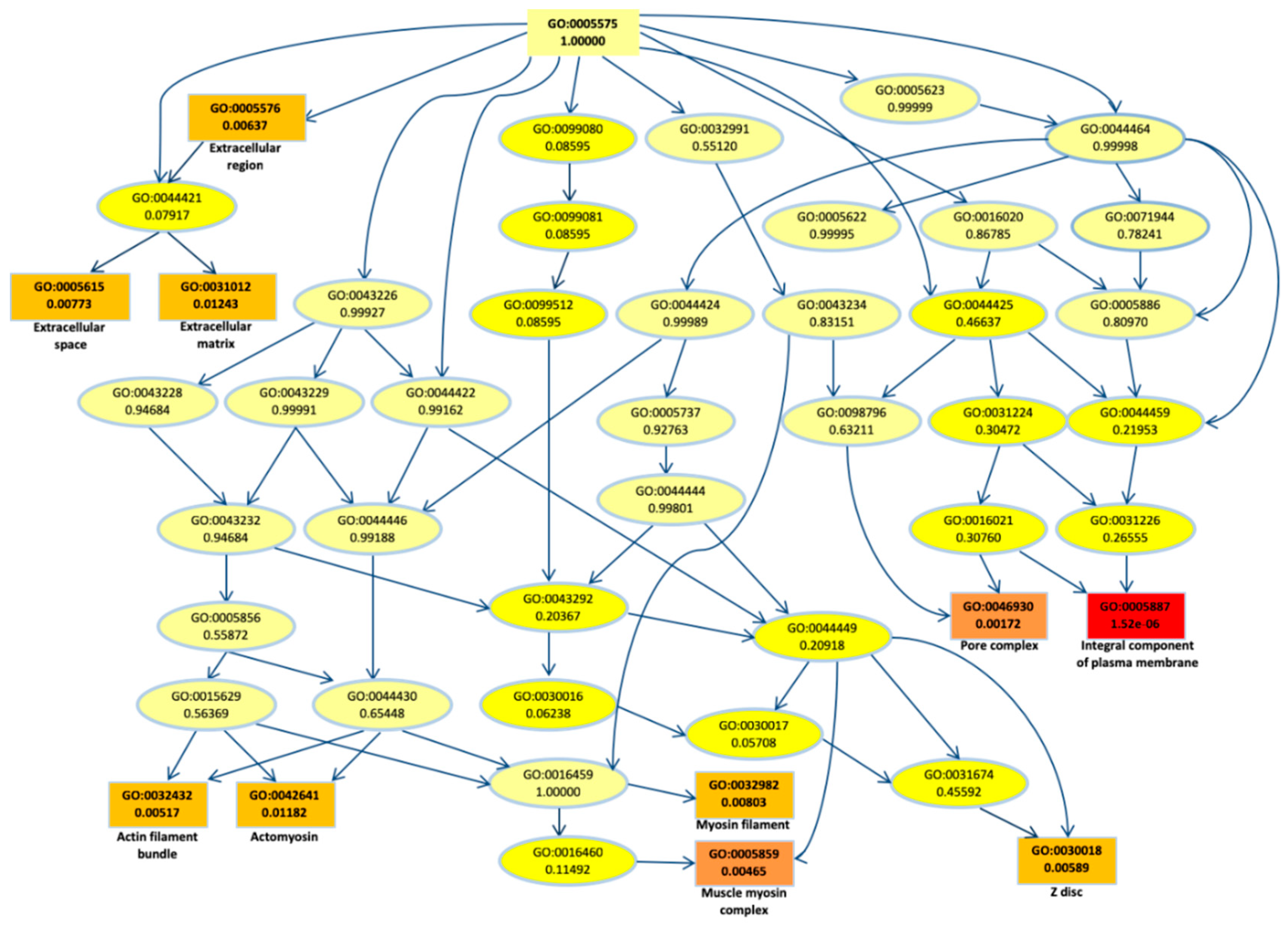
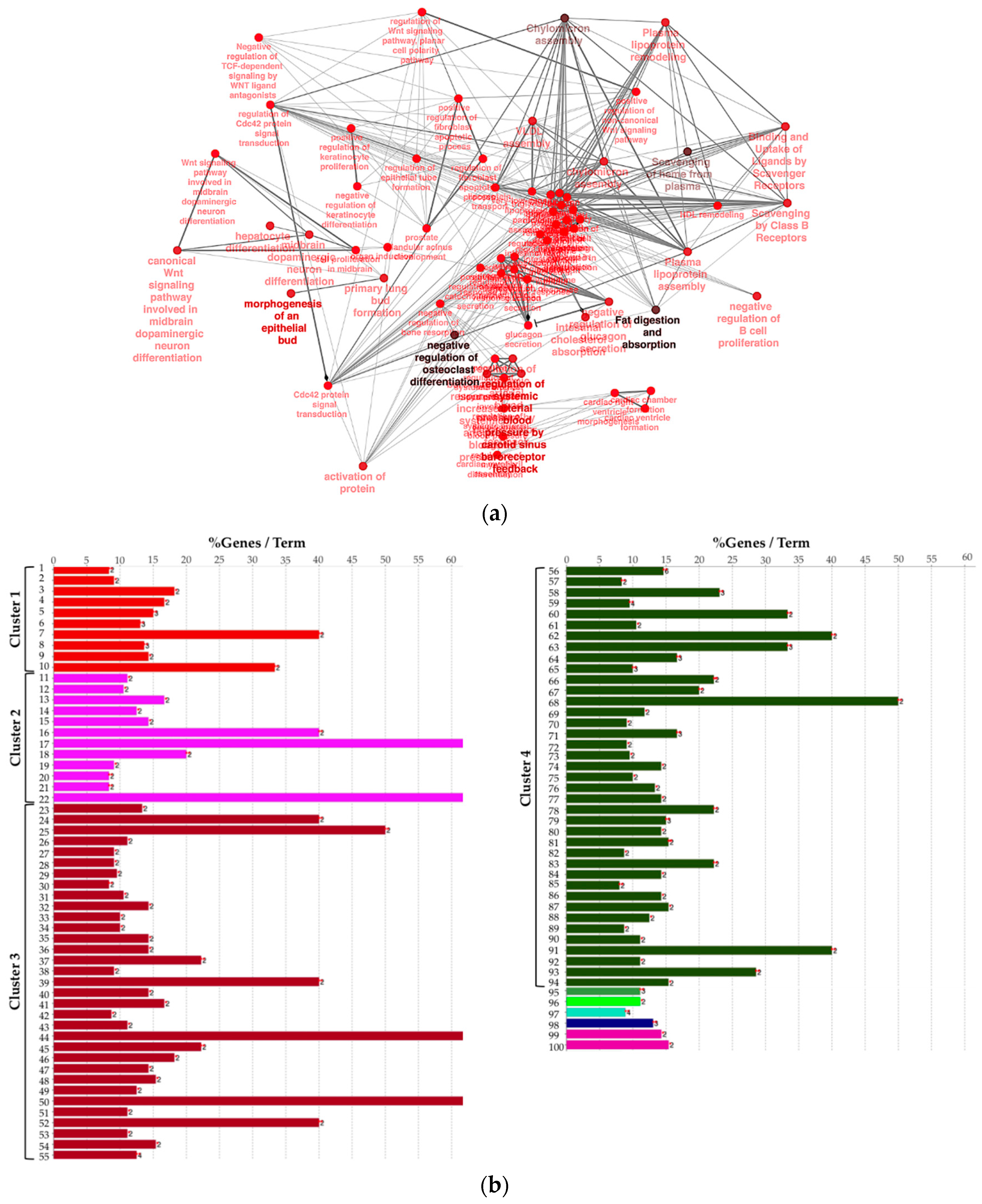
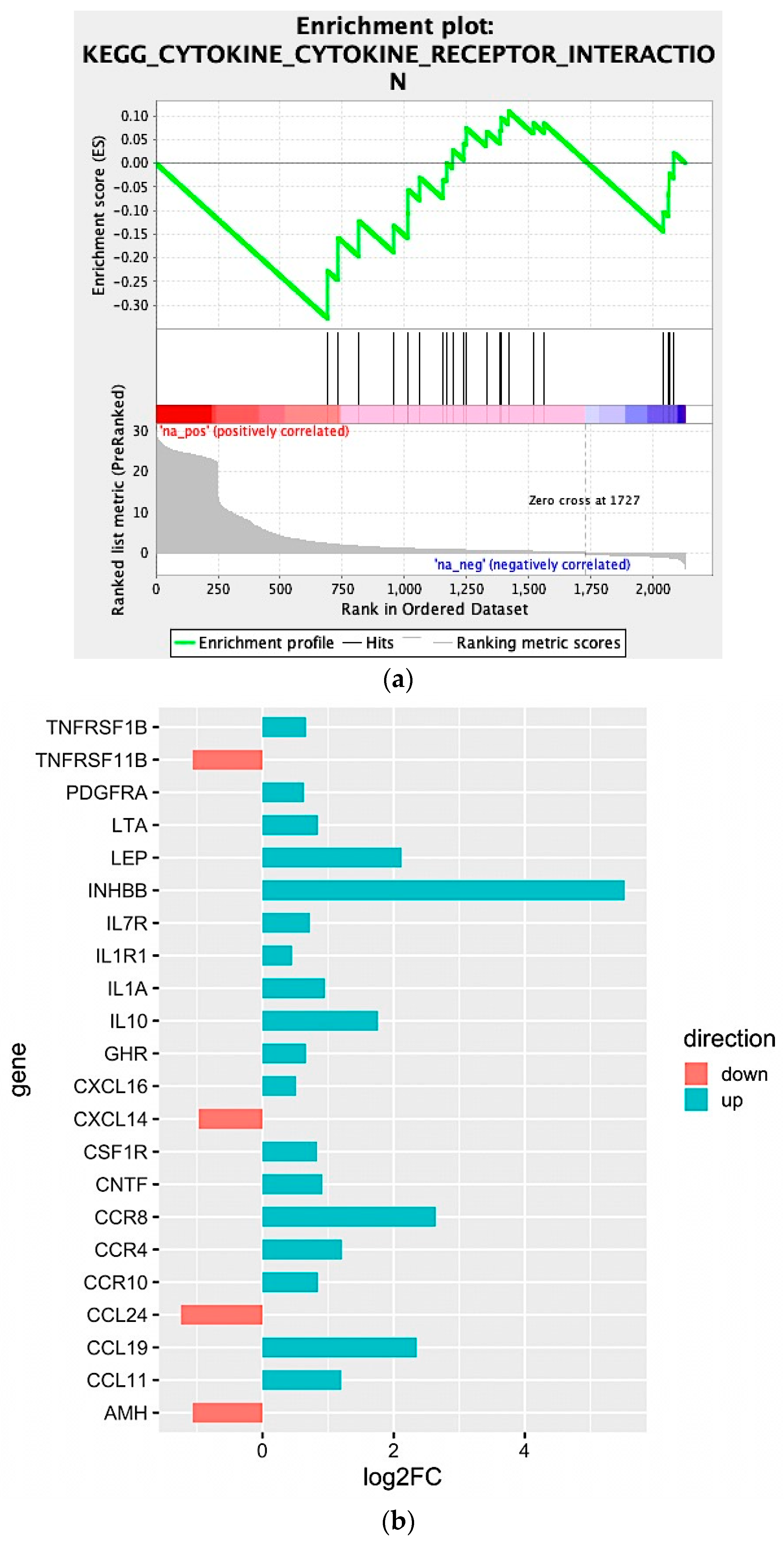
2.2. Biological Network and Gene Set Enrichment Analysis of Database of Essential Genes
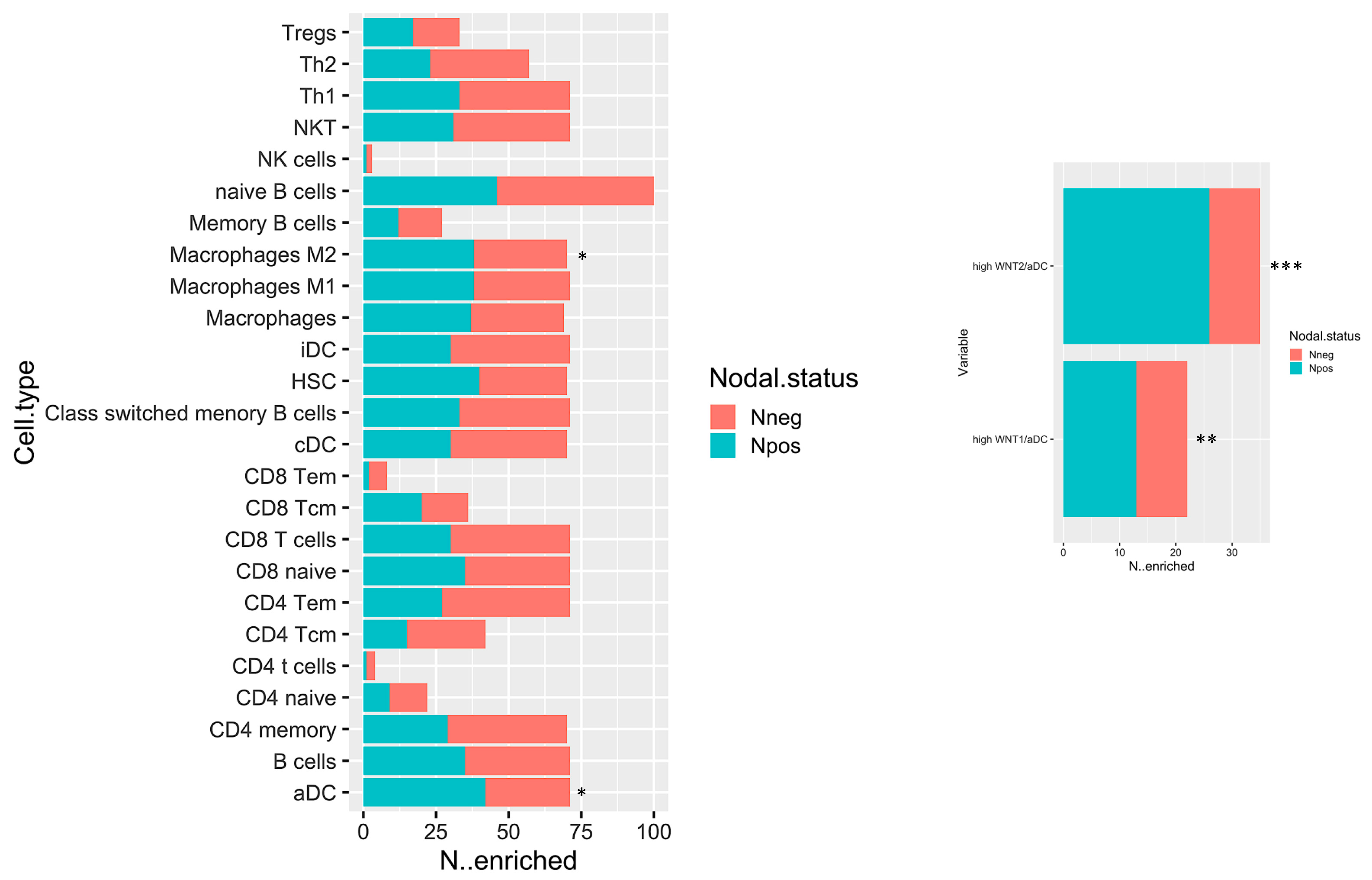
2.3. In Silico Microenvironmental Composition Estimation
2.4. WNT Inhibition Enhances Pancreatic Cancer Cell Immune-Mediated Killing In Vitro
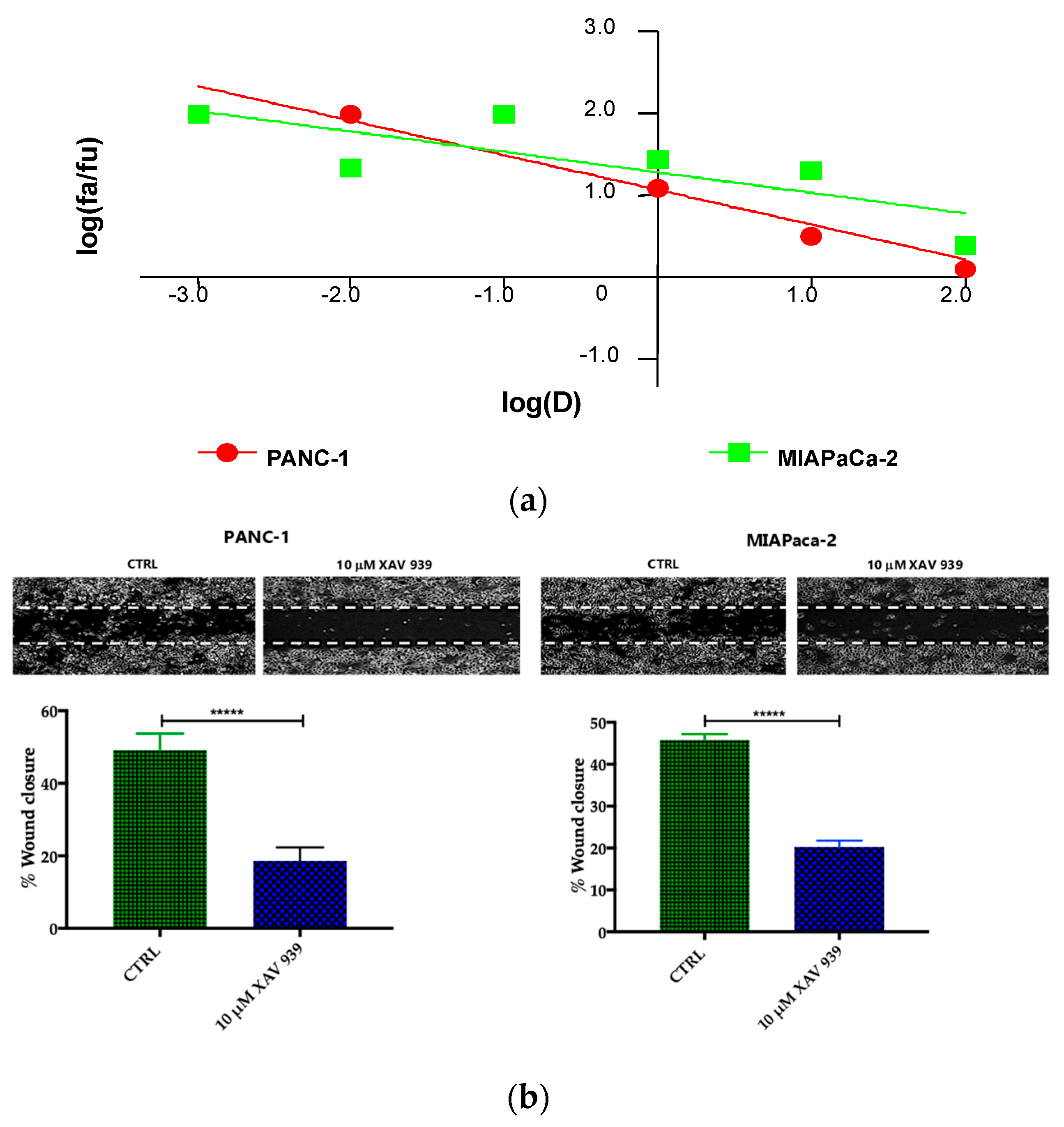
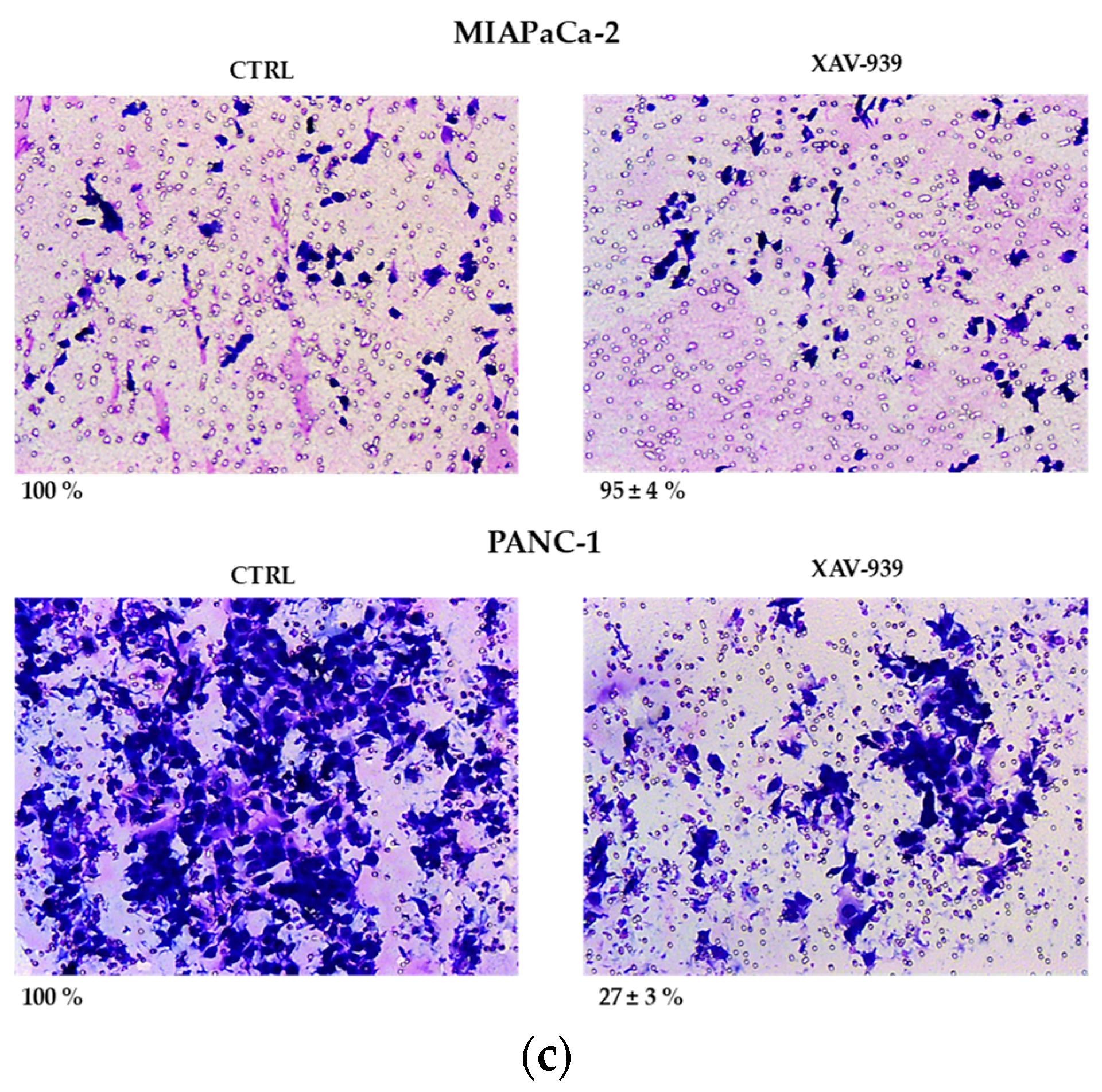
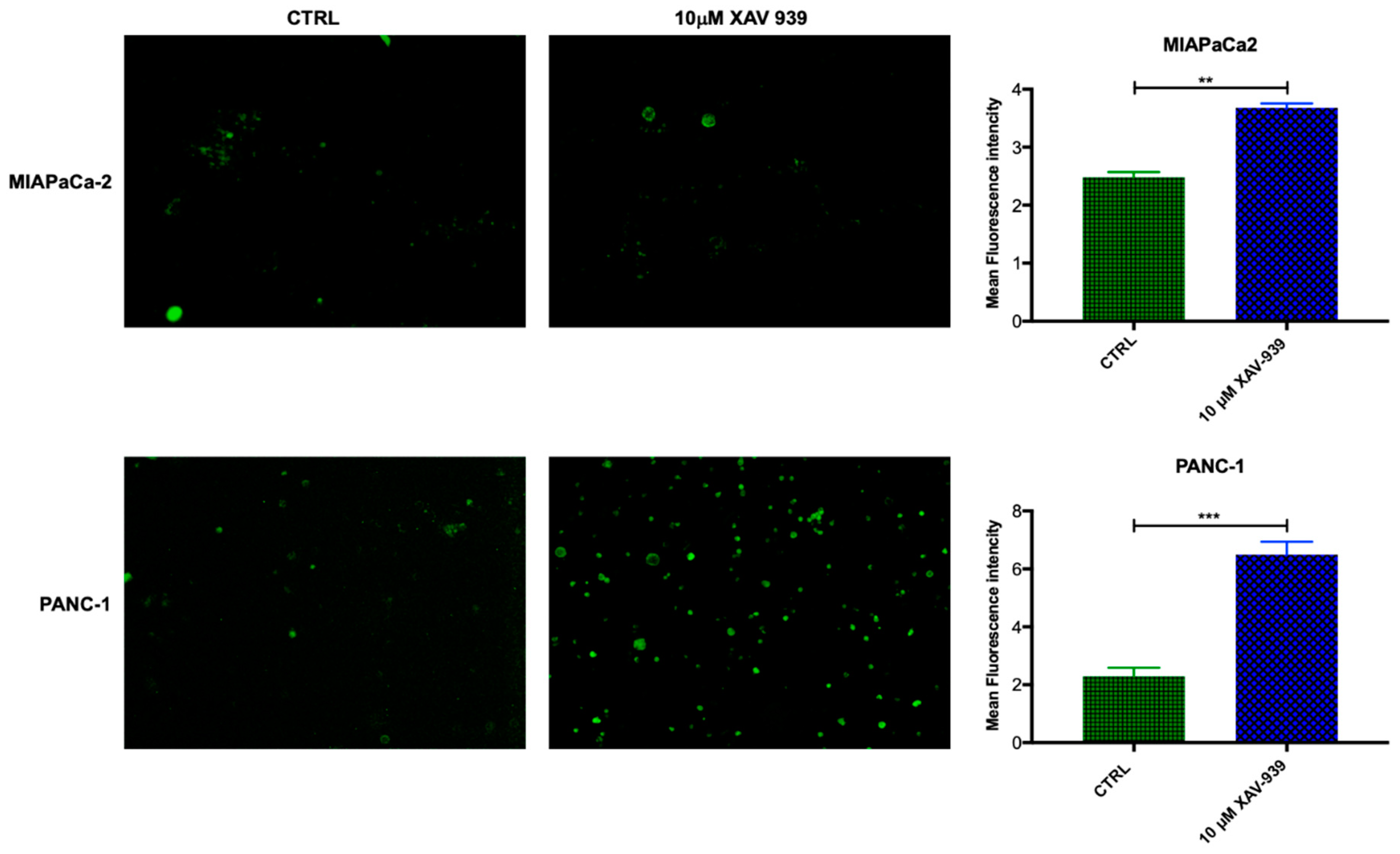
2.4.1. XAV-939 Restrains PDAC Cells Tumor Functions, Exerts Cytotoxic Effect, and Halts Cancer-Migration Potential
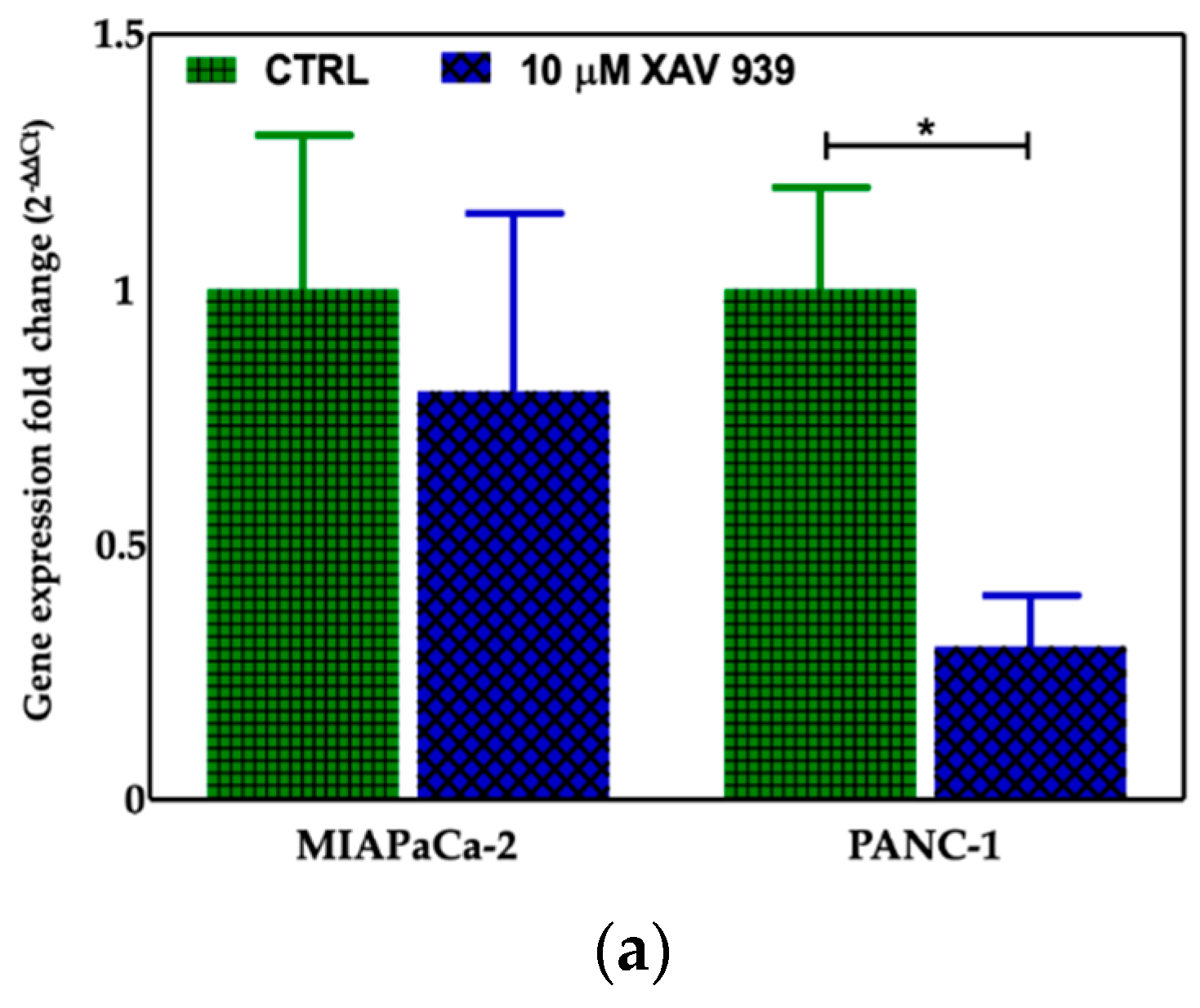
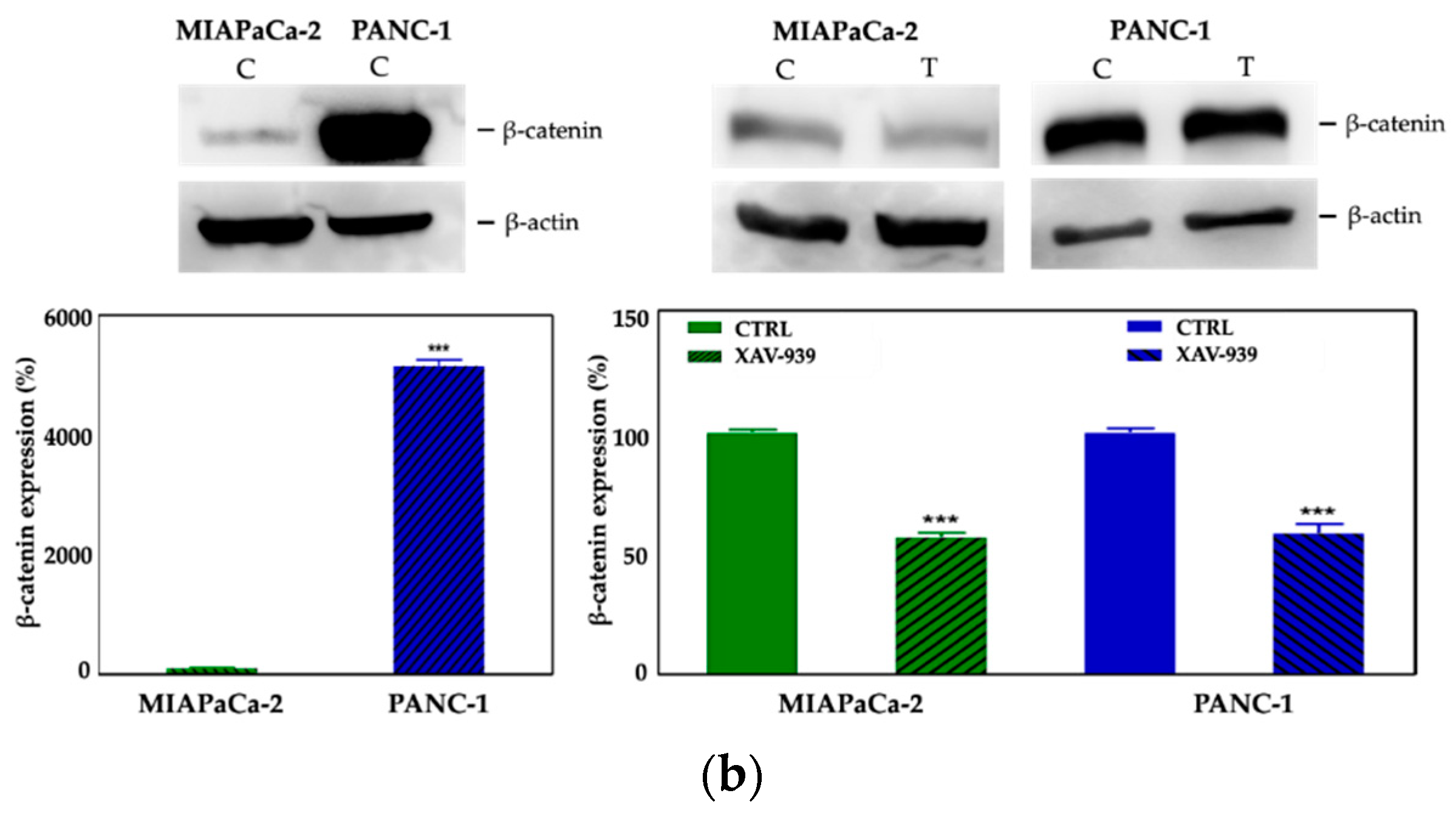
2.4.2. XAV-939 Induced Modulation of β-catenin Expression
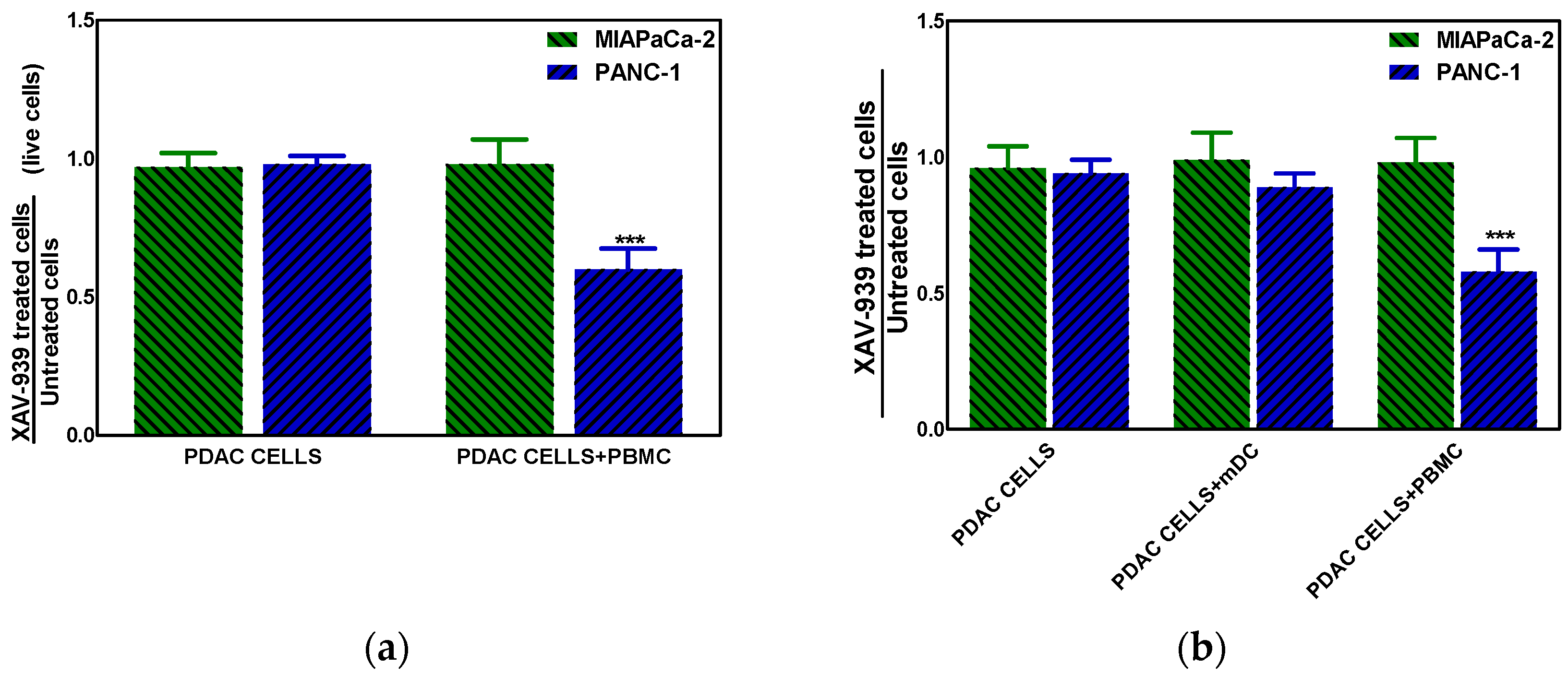
2.4.3. XAV-939 Enhances Immune Cell-Dependent PDAC Cell Killing
2.4.4. XAV-939 Modulates CD40 Expression in mDCs upon PDAC Cell Co-Culture
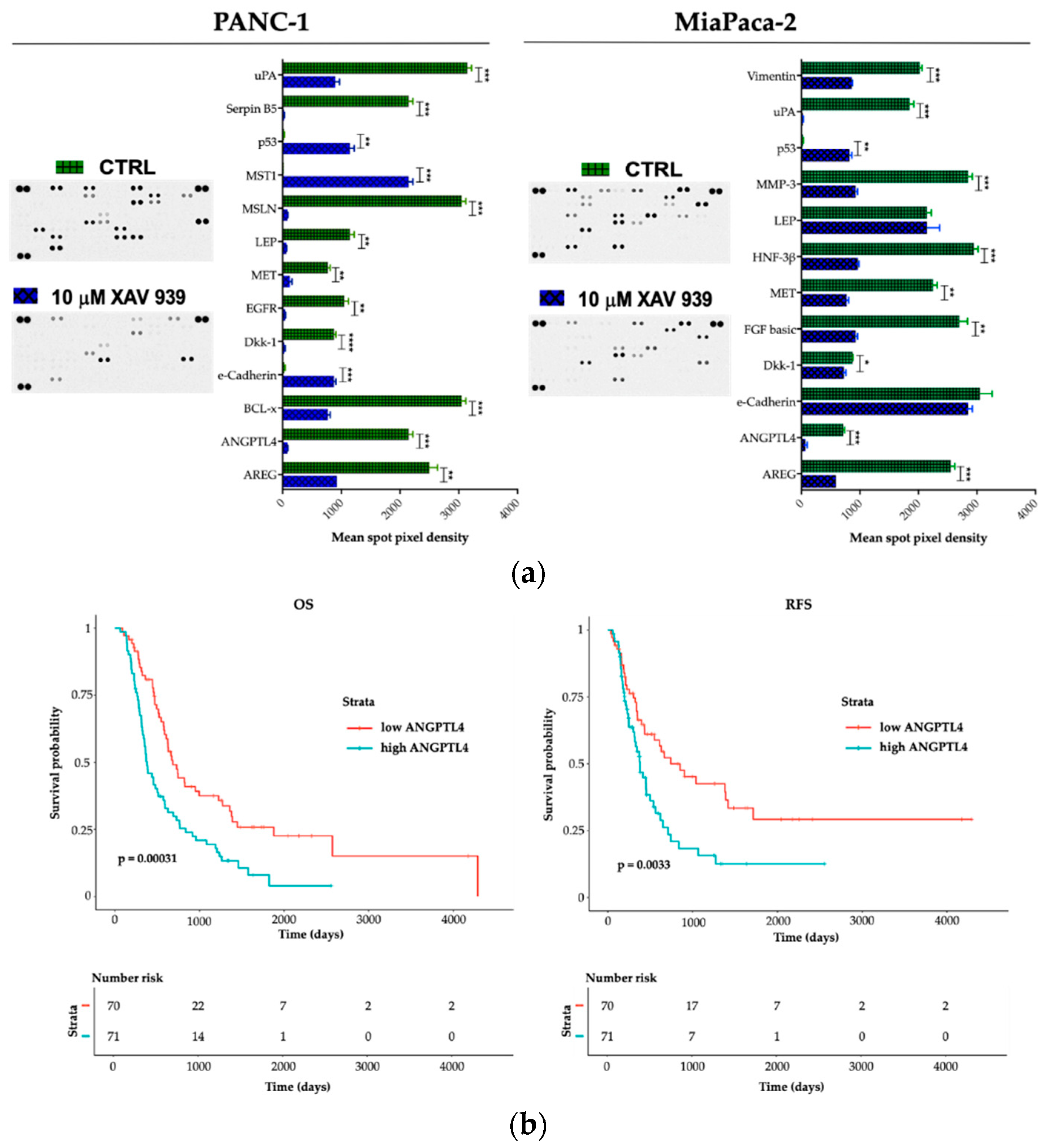
2.5. Effect of XAV-939 on Short-Term Culture of PDAC Cells Cancer-Related Protein Profile
3. Discussion
4. Material and Methods
4.1. Dataset
4.2. Differential Gene Expression Analysis, GO Enrichment, and Biological Network
4.3. In silico Estimation of Microenvironmental Cell Types
4.4. Cell Culture
4.5. β-Catenin Expression by Real-Time PCR
4.6. Cytoxicity Assay
4.7. Killing Assay
4.8. CD40 Expression in mDC
4.9. Migration Test by Wound Healing (Scratch) Assay
4.10. Invasion Assay
4.11. Western Blotting Analysis
4.12. Proteome Profiler Antibodies Array
4.13. Statistical Analyses
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- National Comprehensive Cancer Network NCCN Guidelines Version 2.2019 Pancreatic Adenocarcinoma. Available online: http://www.nccn.org/patients (accessed on 5 May 2019).
- Khorana, A.A.; Mangu, P.B.; Berlin, J.; Engebretson, A.; Hong, T.S.; Maitra, A.; Mohile, S.G.; Mumber, M.; Schulick, R.; Shapiro, M.; et al. Potentially Curable Pancreatic Cancer: American Society of Clinical Oncology Clinical Practice Guideline Update. J. Clin. Oncol. 2017, 35, 2324–2328. [Google Scholar] [CrossRef] [PubMed]
- Isaji, S.; Mizuno, S.; Windsor, J.A.; Bassi, C.; Fernández-del Castillo, C.; Hackert, T.; Hayasaky, A.; Katz, M.H.G.; Kim, S.W.; Kishiwada, M.; et al. International consensus on definition and criteria of borderline resecTable pancreatic ductal adenocarcinoma 2017. Pancreatology 2018, 18, 2–11. [Google Scholar] [CrossRef] [PubMed]
- Kaissis, G.A.; Lohöfer, F.K.; Ziegelmayer, S.; Danner, J.; Jäger, C.; Schirren, R.; Ankerst, D.; Ceyhan, G.O.; Friess, H.; Rummeny, E.J.; et al. Borderline-resectable pancreatic adenocarcinoma: Contour irregularity of the venous confluence in pre-operative computed tomography predicts histopathological infiltration. PLoS ONE 2019, 14, e0208717. [Google Scholar] [CrossRef] [PubMed]
- Silvestris, N.; Brunetti, O.; Vasile, E.; Cellini, F.; Cataldo, I.; Pusceddu, V.; Cattaneo, M.; Partelli, S.; Scartozzi, M.; Aprile, G.; et al. Multimodal treatment of resectable pancreatic ductal adenocarcinoma. Crit. Rev. Oncol. Hematol. 2017, 111, 152–165. [Google Scholar] [CrossRef] [PubMed]
- Bailey, P.; Chang, D.K.; Nones, K.; Johns, A.L.; Patch, A.M.; Gingras, M.C.; Miller, D.K.; Christ, A.N.; Bruxner, T.J.C.; Quinn, M.C.; et al. Genomic analyses identify molecular subtypes of pancreatic cancer. Nature 2016, 531, 47–52. [Google Scholar] [CrossRef] [PubMed]
- Raphael, B.J.; Hruban, R.H.; Aguirre, A.J.; Moffitt, R.A.; Yeh, J.J.; Stewart, C.; Robertson, A.G.; Cherniack, A.D.; Gupta, M.; Getz, G.; et al. The Cancer Genome Atlas Research Network. Integrated genomic characterization of pancreatic ductal adenocarcinoma. Cancer Cell 2017, 32, 185–203. [Google Scholar] [CrossRef] [PubMed]
- Puleo, F.; Nicolle, R.; Blum, Y.; Cros, J.; Marisa, L.; Demetter, P.; Quertinmont, E.; Svrcek, M.; Elarouci, N.; Iovanna, J.; et al. Stratification of pancreatic ductal adenocarcinomas based on tumor and microenvironment features. Gastroenterology 2018, 155, 1999–2013. [Google Scholar] [CrossRef]
- Xiao, Z.; Luo, G.; Liu, C.; Wu, C.; Liu, L.; Liu, Z.; Ni, Q.; Long, J.; Yu, X. Molecular mechanism underlying lymphatic metastasis in pancreatic cancer. BioMed Res. Int. 2014, 2014, 15. [Google Scholar] [CrossRef]
- Luke, J.J.; Bao, R.; Sweis, R.F.; Spranger, S.; Gajewski, T.F. WNT/β-catenin pathway activation correlates with immune exclusion across human cancers. Clin. Cancer Res. 2019, 25, 3074–3083. [Google Scholar] [CrossRef]
- Wang, B.; Tian, T.; Kalland, K.H.; Ke, X.; Qu, Y. Targeting Wnt/β-catenin signaling for cancer immunotherapy. Trends Pharmacol. Sci. 2018, 39, 648–658. [Google Scholar] [CrossRef]
- Dangaj, D.; Barras, D.; Coukos, G. Tumor Landscapes: β-Catenin Drives Immune Desertification. Clin. Cancer Res. 2019, 25, 3074. [Google Scholar] [CrossRef] [PubMed]
- Deer, E.L.; González-Hernández, J.; Coursen, J.D.; Shea, J.E.; Ngatia, J.; Scaife, C.L.; Firpo, M.A.; Mulvihill, S.J. Phenotype and genotype of pancreatic cancer cell lines. Pancreas 2010, 39, 425–435. [Google Scholar] [CrossRef] [PubMed]
- Hanke, N.; Alizadeh, D.; Katsanis, E.; Larmonier, N. Dendritic cell tumor killing activity and its potential applications in cancer immunotherapy. Crit. Rev. Immunol. 2013, 33, 1–21. [Google Scholar] [CrossRef] [PubMed]
- Afifi, M.M.; Austin, L.A.; MacKey, M.A.; El-Sayed, M.A. XAV939: From a small inhibitor to a potent drug bioconjugate when delivered by gold nanoparticles. Bioconjug. Chem. 2014, 25, 207–215. [Google Scholar] [CrossRef] [PubMed]
- Zhan, T.; Rindtorff, N.; Boutros, M. Wnt signaling in cancer. Oncogene 2017, 36, 1461–1473. [Google Scholar] [CrossRef] [PubMed]
- Qi, J.; Yu, Y.; Öztürk, Ö.A.; Holland, J.D.; Besser, D.; Fritzmann, J.; Wulf-Goldenberg, A.; Eckert, K.; Fichtner, I.; Birchmeier, W. New Wnt/β-catenin target genes promote experimental metastasis and migration of colorectal cancer cells through different signals. Gut 2016, 65, 1690–1701. [Google Scholar] [CrossRef] [PubMed]
- Bao, R.; Christova, T.; Song, S.; Angers, S.; Yan, X.; Attisano, L. Inhibition of Tankyrases Induces Axin Stabilization and Blocks Wnt Signalling in Breast Cancer Cells. PLoS ONE 2012, 7, e48670. [Google Scholar] [CrossRef] [PubMed]
- Swafford, D.; Manicassamy, S. Wnt signaling in dendritic cells: Its role in regulation of immunity and tolerance. Discov. Med. 2015, 19, 303–310. [Google Scholar]
- Hellman, P.; Eriksson, H. Early Activation Markers of Human Peripheral Dendritic Cells. Hum. Immunol. 2007, 68, 324–333. [Google Scholar] [CrossRef]
- Strobel, O.; Neoptolemos, J.; Jaeger, D.; Buechler, M.W. Optimizing the outcomes of pancreatic cancer surgery. Nat. Rev. Clin. Oncol. 2019, 16, 11–26. [Google Scholar] [CrossRef]
- Honselmann, K.C.; Pergolini, I.; Castillo, C.F.; Deshpande, V.; Ting, D.; Taylor, M.S.; Bolm, L.; Qadan, M.; Wellner, U.; Sandini, M.; et al. Timing But Not Patterns of Recurrence Is Different Between Node-negative and Node-positive Resected Pancreatic Cancer. Ann. Surg. 2019, 42, 93–99. [Google Scholar] [CrossRef] [PubMed]
- Valastyan, S.; Weinberg, R.A. Tumor metastasis: Molecular insights and evolving paradigms. Cell 2011, 147, 275–292. [Google Scholar] [CrossRef] [PubMed]
- Chen, D.S.; Mellman, I. Elements of cancer immunity and the cancer—immune set point. Nature 2017, 541, 321–330. [Google Scholar] [CrossRef] [PubMed]
- Jiang, H.; Li, Q.; He, C.; Li, F.; Sheng, H.; Shen, X.; Zhang, X.; Zhu, S.; Chen, H.; Chen, X.; et al. Activation of the Wnt pathway through Wnt2 promotes metastasis in pancreatic cancer. Am. J. Cancer Res. 2014, 4, 537–544. [Google Scholar] [PubMed]
- Fu, L.; Zhang, C.; Zhang, L.Y.; Dong, S.S.; Lu, L.H.; Chen, J.; Dai, Y.; Li, Y.; Kong, K.L.; Kwong, D.L.; et al. Wnt2 secreted by tumour fibroblasts promotes tumour progression in oesophageal cancer by activation of the Wnt/β-catenin signalling pathway. Gut 2011, 60, 1635–1643. [Google Scholar] [CrossRef] [PubMed]
- You, L.; He, B.; Xu, Z.; Uematsu, K.; Mazieres, J.; Fujii, N.; Mikami, I.; Reguart, N.; McIntosh, J.K.; Kashani-Sabet, M.; et al. An anti-Wnt-2 monoclonal antibody induces apoptosis in malignant melanoma cells and inhibits tumor growth. Cancer Res. 2004, 64, 5385–5389. [Google Scholar] [CrossRef]
- Tabatabai, R.; Linhares, Y.; Bolos, D.; Mita, M.; Mita, A. Targeting the Wnt pathway in cancer: A review of novel therapeutics. Target. Oncol. 2017, 12, 623–641. [Google Scholar] [CrossRef] [PubMed]
- Binnewies, M.; Roberts, E.W.; Kersten, K.; Chan, V.; Fearon, D.F.; Merad, M.; Coussens, L.M.; Gabrilovich, D.I.; Ostrand-Rosenberg, S.; Hedrick, C.C.; et al. Understanding the tumor immune microenvironment (TIME) for effective therapy. Nat. Med. 2018, 24, 541–550. [Google Scholar] [CrossRef]
- Yang, Y.; Ye, Y.C.; Chen, Y.; Zhao, J.L.; Gao, C.C.; Han, H.; Liu, W.C.; Qin, H.Y. Crosstalk between hepatic tumor cells and macrophages via Wnt/β-catenin signaling promotes M2-like macrophage polarization and reinforces tumor malignant behaviors. Cell Death Dis. 2018, 9, 793. [Google Scholar] [CrossRef]
- Valilou, S.F.; Keshavarz-Fathi, M.; Silvestris, N.; Argentiero, A.; Rezaei, N. The role of inflammatory cytokines and tumor associated macrophages (TAMs) in microenvironment of pancreatic cancer. Cytokine Growth Factor Rev. 2018, 39, 46–61. [Google Scholar] [CrossRef]
- Amedei, A.; Niccolai, E.; Prisco, D. Pancreatic cancer: Role of the immune system in cancer progression and vaccine-based immunotherapy. Hum. Vaccin. Immunother. 2014, 10, 3354–3368. [Google Scholar] [CrossRef] [PubMed]
- Gnoni, A.; Licchetta, A.; Scarpa, A.; Azzariti, A.; Brunetti, A.; Simone, G.; Nardulli, P.; Santini, D.; Aieta, M.; Delcuratolo, S.; et al. Carcinogenesis of pancreatic adenocarcinoma: Precursor lesions. Int. J. Mol. Sci. 2013, 14, 19731–19762. [Google Scholar] [CrossRef] [PubMed]
- Zhao, F.; Xiao, C.; Evans, K.S.; Theivanthiran, T.; DeVito, N.; Holtzhausen, A.; Liu, J.; Liu, X.; Boczkowski, D.; Nair, S.; et al. Paracrine wnt5a-β-catenin signaling triggers a metabolic program that drives dendritic cell tolerization. Immunity 2018, 48, 147–160. [Google Scholar] [CrossRef] [PubMed]
- Suryawanshi, A.; Tadagavadi, R.K.; Swafford, D.; Manicassamy, S. Modulation of inflammatory responses by Wnt/β-catenin signaling in dendritic cells: A novel immunotherapy target for autoimmunity and cancer. Front. Immunol. 2016, 7, 460. [Google Scholar] [CrossRef] [PubMed]
- Barilla, R.M.; Diskin, B.; Caso, R.C.; Lee, K.B.; Mohan, N.; Buttar, C.; Adam, S.; Sekendiz, Z.; Wang, J.; Salas, R.D.; et al. Specialized dendritic cells induce tumor-promoting IL-10+ IL-17+ FoxP3 neg regulatory CD4+ T cells in pancreatic carcinoma. Nat. Commun. 2019, 10, 1424. [Google Scholar] [CrossRef] [PubMed]
- Porcelli, L.; Iacobazzi, R.M.; Di Fonte, R.; Serratì, S.; Intini, A.; Solimando, A.G.; Brunetti, O.; Calabrese, A.; Leonetti, F.; Azzariti, A.; et al. CAFs and TGF-β Signaling Activation by Mast Cells Contribute to Resistance to Gemcitabine/Nabpaclitaxel in Pancreatic Cancer. Cancers 2019, 11, 330. [Google Scholar] [CrossRef] [PubMed]
- Gradiz, R.; Silva, H.C.; Carvalho, L.; Botelho, M.F.; Mota-Pinto, A. MIA PaCa-2 and PANC-1–pancreas ductal adenocarcinoma cell lines with neuroendocrine differentiation and somatostatin receptors. Sci. Rep. 2016, 6, 21648. [Google Scholar] [CrossRef] [PubMed]
- Dregalla, R.C.; Zhou, J.; Idate, R.R.; Battaglia, C.L.; Liber, H.L.; Bailey, S.M. Regulatory roles of tankyrase 1 at telomeres and in DNA repair: Suppression of T-SCE and stabilization of DNA-PKcs. Aging (Albany NY) 2010, 2, 691. [Google Scholar] [CrossRef]
- Karicheva, O.; Rodriguez-Vargas, J.M.; Wadier, N.; Martin-Hernandez, K.; Vauchelles, R.; Magroun, N.; Tissier, A.; Schreiber, V.; Dantzer, F. PARP3 controls TGFβ and ROS driven epithelial-to-mesenchymal transition and stemness by stimulating a TG2-Snail-E-cadherin axis. Oncotarget 2016, 7, 64109. [Google Scholar] [CrossRef]
- Cui, J.; Zhou, Z.; Yang, H.; Jiao, F.; Li, N.; Gao, Y.; Wang, L.; Chen, J.; Quan, M. MST1 Suppresses Pancreatic Cancer Progression via ROS-Induced Pyroptosis. Mol. Cancer Res. 2019, 17. [Google Scholar] [CrossRef]
- Tsai, B.P.; Hoverter, N.P.; Waterman, M.L. Blending hippo and WNT: Sharing messengers and regulation. Cell 2012, 151, 1401–1403. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Taha, Z.; Janse van Rensburg, H.; Yang, X. The Hippo pathway: Immunity and cancer. Cancers 2018, 10, 94. [Google Scholar] [CrossRef] [PubMed]
- Zhao, T.; Jiang, W.; Wang, X.; Wang, H.; Zheng, C.; Li, Y.; Sun, Y.; Huang, C.; Han, Z.B.; Yang, S.; et al. ESE3 inhibits pancreatic cancer metastasis by upregulating E-cadherin. Cancer Res. 2017, 77, 874–885. [Google Scholar] [CrossRef] [PubMed]
- Uhlen, M.; Zhang, C.; Lee, S.; Sjöstedt, E.; Fagerberg, L.; Bidkhori, G.; Benfeitas, R.; Arif, M.; Liu, Z.; Edfors, F.; et al. A pathology atlas of the human cancer transcriptome. Science 2017, 357, eaan2507. [Google Scholar] [CrossRef] [PubMed]
- Huang, R.L.; Teo, Z.; Chong, H.C.; Zhu, P.; Tan, M.J.; Tan, C.K.; Lam, C.R.I.; Sng, M.K.; Leong, D.T.W.; Tan, S.M.; et al. ANGPTL4 modulates vascular junction integrity by integrin signaling and disruption of intercellular VE-cadherin and claudin-5 clusters. Blood 2011, 118, 3990–4002. [Google Scholar] [CrossRef]
- Tan, M.J.; Teo, Z.; Sng, M.K.; Zhu, P.; Tan, N.S. Emerging roles of angiopoietin-like 4 in human cancer. Mol. Cancer Res. 2012, 10, 677–688. [Google Scholar] [CrossRef]
- Phua, T.; Sng, M.K.; Tan, E.H.P.; Chee, D.S.L.; Li, Y.; Wee, J.W.K.; Teo, Z.; Chan, J.S.K.; Lim, M.M.K.; Tan, C.K.; et al. Angiopoietin-like 4 mediates colonic inflammation by regulating chemokine transcript stability via tristetraprolin. Sci. Rep. 2017, 7, 44351. [Google Scholar] [CrossRef]
- Yang, X.; Cheng, Y.; Su, G. A review of the multifunctionality of angiopoietin-like 4 in eye disease. Biosci. Rep. 2018, 38, BSR20180557. [Google Scholar] [CrossRef]
- Waddell, N.; Pajic, M.; Patch, A.M.; Chang, D.K.; Kassahn, K.S.; Bailey, P.; Johns, A.L.; Miller, D.; Nones, K.; Quek, K.; et al. Whole genomes redefine the mutational landscape of pancreatic cancer. Nature 2015, 518, 495. [Google Scholar] [CrossRef]
- Love, M.I.; Huber, W.; Anders, S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 2014, 15, 550. [Google Scholar] [CrossRef]
- Alexa, A.; Rahnenfuhrer, J. TopGo: Enrichment Analysis for Gene Ontology. R package version 2.34.0. 2018. Available online: https://bioconductor.org/packages/release/bioc/html/topGO.html (accessed on 25 June 2019).
- Bindea, G.; Mlecnik, B.; Hackl, H.; Charoentong, P.; Tosolini, M.; Kirilovsky, A.; Fridman, W.; Pagès, F.; Trajanoski, Z.; Galon, J. ClueGO: A Cytoscape plug-in to decipher functionally grouped gene ontology and pathway annotation networks. Bioinformatics 2009, 25, 1091–1093. [Google Scholar] [CrossRef] [PubMed]
- Subramanian, A.; Tamayo, P.; Mootha, V.K.; Mukherjee, S.; Ebert, B.L.; Gillette, M.A.; Paulovich, A.; Pomeroy, S.L.; Golub, T.R.; Lander, E.S.; et al. Gene set enrichment analysis: A knowledge-based approach for interpreting genome-wide expression profiles. Proc. Natl. Acad. Sci. USA 2005, 102, 15545–15550. [Google Scholar] [CrossRef] [PubMed]
- Aran, D.; Hu, Z.; Butte, A.J. xCell: Digitally portraying the tissue cellular heterogeneity landscape. Genome Biol. 2017, 18, 220. [Google Scholar] [CrossRef] [PubMed]
- Porcelli, L.; Stolfa, D.; Stefanachi, A.; Di Fonte, R.; Garofoli, M.; Iacobazzi, R.M.; Silvestris, N.; Guarini, A.; Cellamare, S.; Azzariti, A. Synthesis and biological evaluation of N-biphenyl-nicotinic based moiety compounds: A new class of antimitotic agents for the treatment of Hodgkin Lymphoma. Cancer Lett. 2019, 445, 1–10. [Google Scholar] [CrossRef] [PubMed]
- Solimando, A.G.; Brandl, A.; Mattenheimer, K.; Graf, C.; Ritz, M.; Ruckdeschel, A.; Stühmer, T.; Mokhtari, Z.; Rudelius, M.; Dotterweich, J.; et al. JAM-A as a prognostic factor and new therapeutic target in multiple myeloma. Leukemia 2018, 32, 736–743. [Google Scholar] [CrossRef] [PubMed]
- Rao, L.; De Veirman, K.; Giannico, D.; Saltarella, I.; Desantis, V.; Frassanito, M.A.; Solimando, A.G.; Ribatti, D.; Prete, M.; Harstrick, A.; et al. Targeting angiogenesis in multiple myeloma by the VEGF and HGF blocking DARPin® protein MP0250: A preclinical study. Oncotarget 2018, 9, 13366–13381. [Google Scholar] [CrossRef]
- Uhlén, M.; Fagerberg, L.; Hallström, B.M.; Lindskog, C.; Oksvold, P.; Mardinoglu, A.; Sivertsson, Å.; Kampf, C.; Sjöstedt, E.; Asplund, A.; et al. Tissue-based map of the human proteome. Science 2015, 347, 1260419. [Google Scholar] [CrossRef]
- Thul, P.J.; Åkesson, L.; Wiking, M.; Mahdessian, D.; Geladaki, A.; Blal, H.A.; Alm, T.; Asplund, A.; Björk, L.; Breckels, L.M.; et al. A subcellular map of the human proteome. Science 2017, 356, eaal3321. [Google Scholar] [CrossRef]
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Argentiero, A.; De Summa, S.; Di Fonte, R.; Iacobazzi, R.M.; Porcelli, L.; Da Vià, M.; Brunetti, O.; Azzariti, A.; Silvestris, N.; Solimando, A.G. Gene Expression Comparison between the Lymph Node-Positive and -Negative Reveals a Peculiar Immune Microenvironment Signature and a Theranostic Role for WNT Targeting in Pancreatic Ductal Adenocarcinoma: A Pilot Study. Cancers 2019, 11, 942. https://doi.org/10.3390/cancers11070942
Argentiero A, De Summa S, Di Fonte R, Iacobazzi RM, Porcelli L, Da Vià M, Brunetti O, Azzariti A, Silvestris N, Solimando AG. Gene Expression Comparison between the Lymph Node-Positive and -Negative Reveals a Peculiar Immune Microenvironment Signature and a Theranostic Role for WNT Targeting in Pancreatic Ductal Adenocarcinoma: A Pilot Study. Cancers. 2019; 11(7):942. https://doi.org/10.3390/cancers11070942
Chicago/Turabian StyleArgentiero, Antonella, Simona De Summa, Roberta Di Fonte, Rosa Maria Iacobazzi, Letizia Porcelli, Matteo Da Vià, Oronzo Brunetti, Amalia Azzariti, Nicola Silvestris, and Antonio Giovanni Solimando. 2019. "Gene Expression Comparison between the Lymph Node-Positive and -Negative Reveals a Peculiar Immune Microenvironment Signature and a Theranostic Role for WNT Targeting in Pancreatic Ductal Adenocarcinoma: A Pilot Study" Cancers 11, no. 7: 942. https://doi.org/10.3390/cancers11070942
APA StyleArgentiero, A., De Summa, S., Di Fonte, R., Iacobazzi, R. M., Porcelli, L., Da Vià, M., Brunetti, O., Azzariti, A., Silvestris, N., & Solimando, A. G. (2019). Gene Expression Comparison between the Lymph Node-Positive and -Negative Reveals a Peculiar Immune Microenvironment Signature and a Theranostic Role for WNT Targeting in Pancreatic Ductal Adenocarcinoma: A Pilot Study. Cancers, 11(7), 942. https://doi.org/10.3390/cancers11070942









