Diagnostic Leukapheresis Enables Reliable Transcriptomic Profiling of Single Circulating Tumor Cells to Characterize Inter-Cellular Heterogeneity in Terms of Endocrine Resistance
Abstract
1. Introduction
2. Results
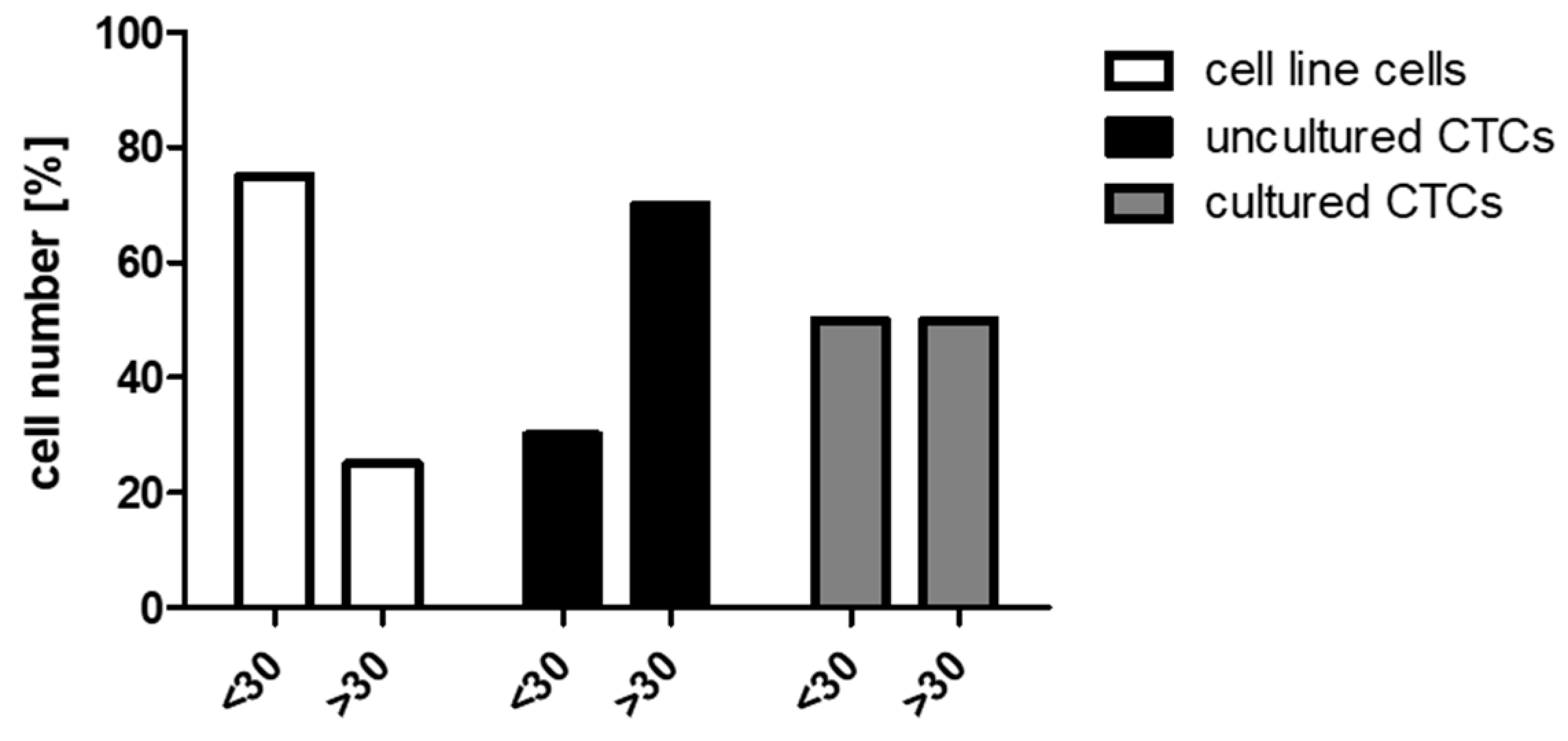
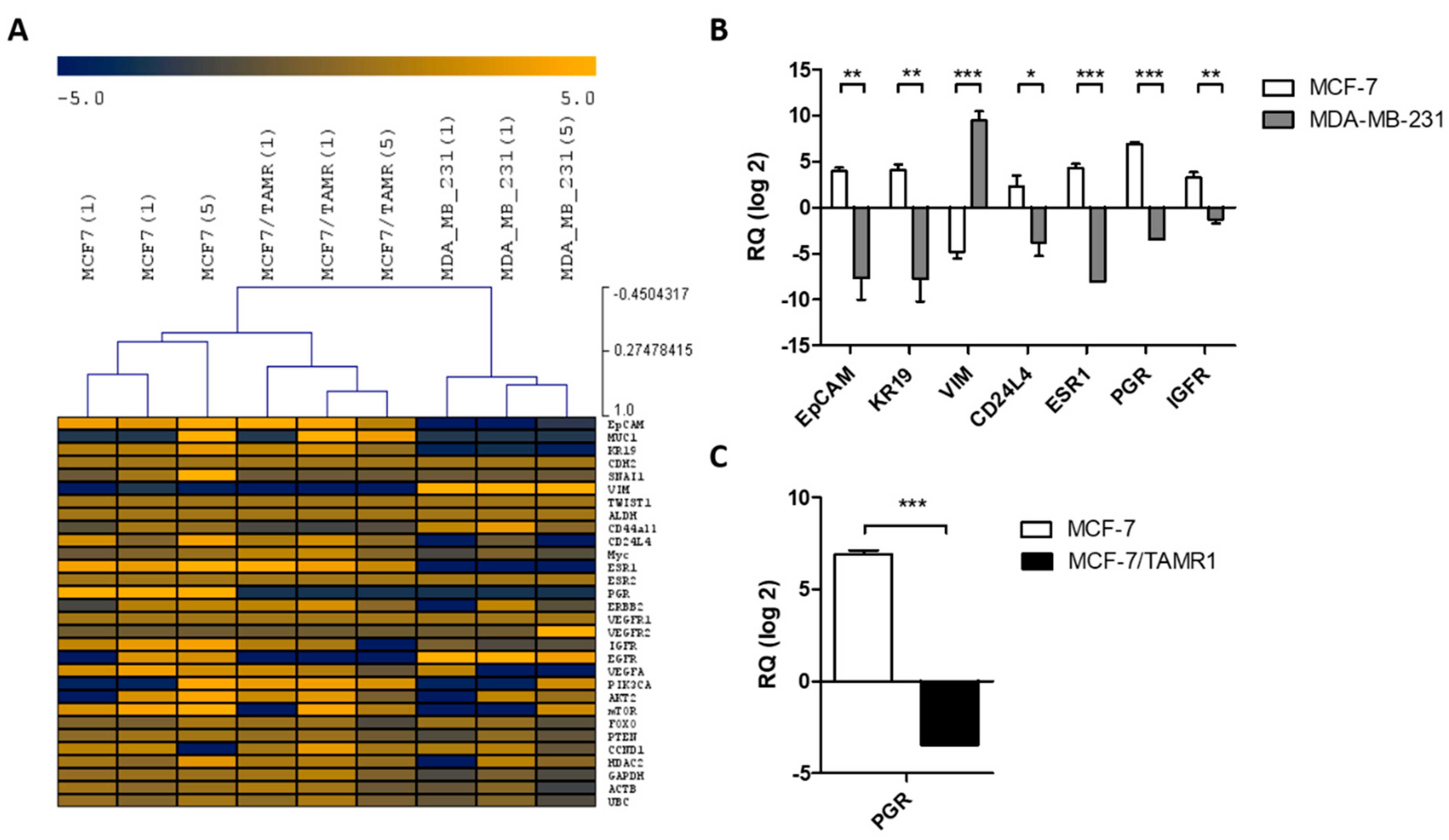
2.1. Validation of Single Cell Multi-Marker RT-qPCR Analysis
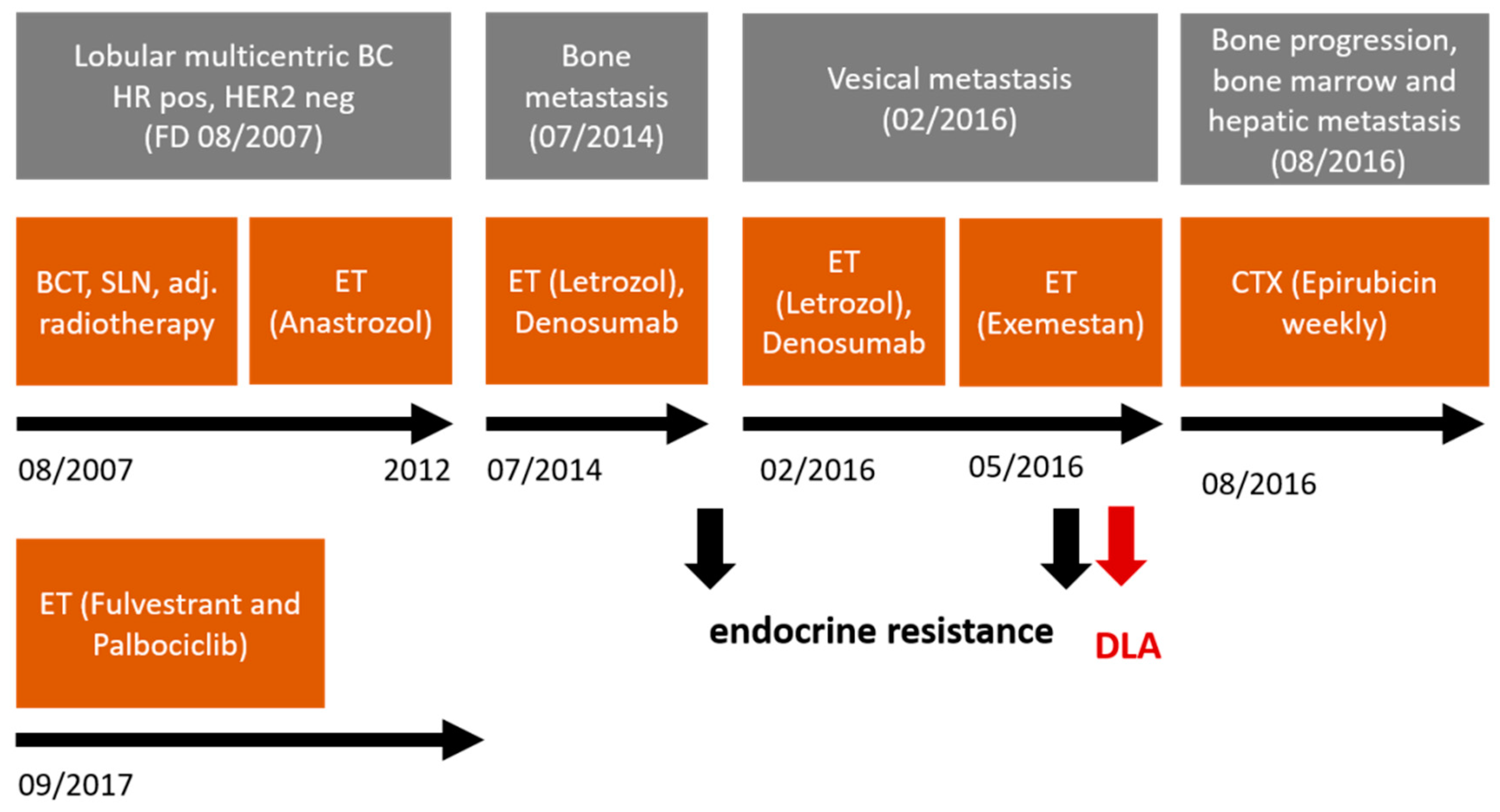
2.2. Clinical Characteristics and Therapeutic Interventions of the DLA Patient
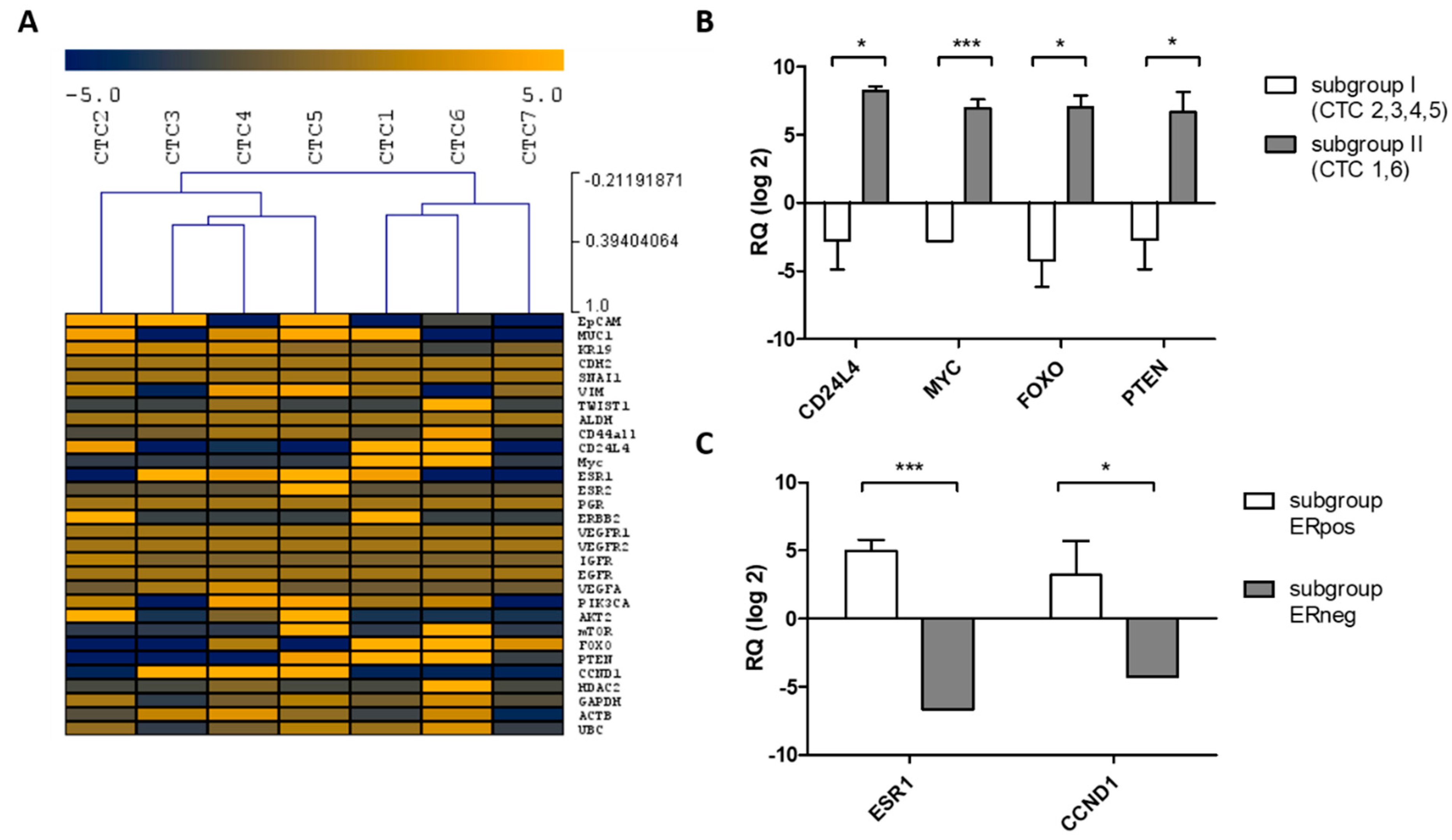
2.3. Determination of Intra-Patient Heterogeneity by Multi-Marker Gene Expression Profiling of Uncultured and Cultured CTCs
2.3.1. Uncultured CTCs
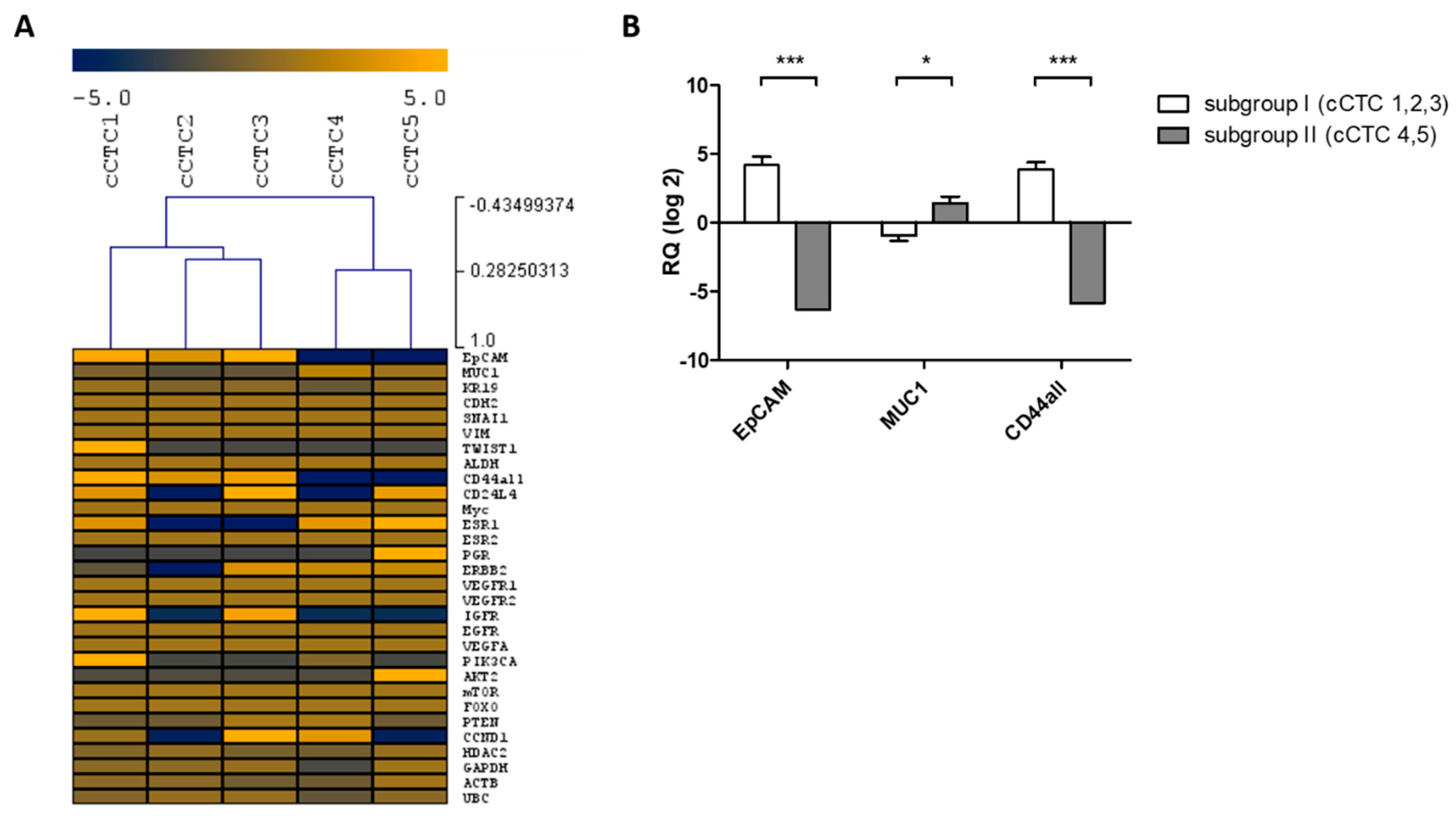
2.3.2. Cultured CTCs
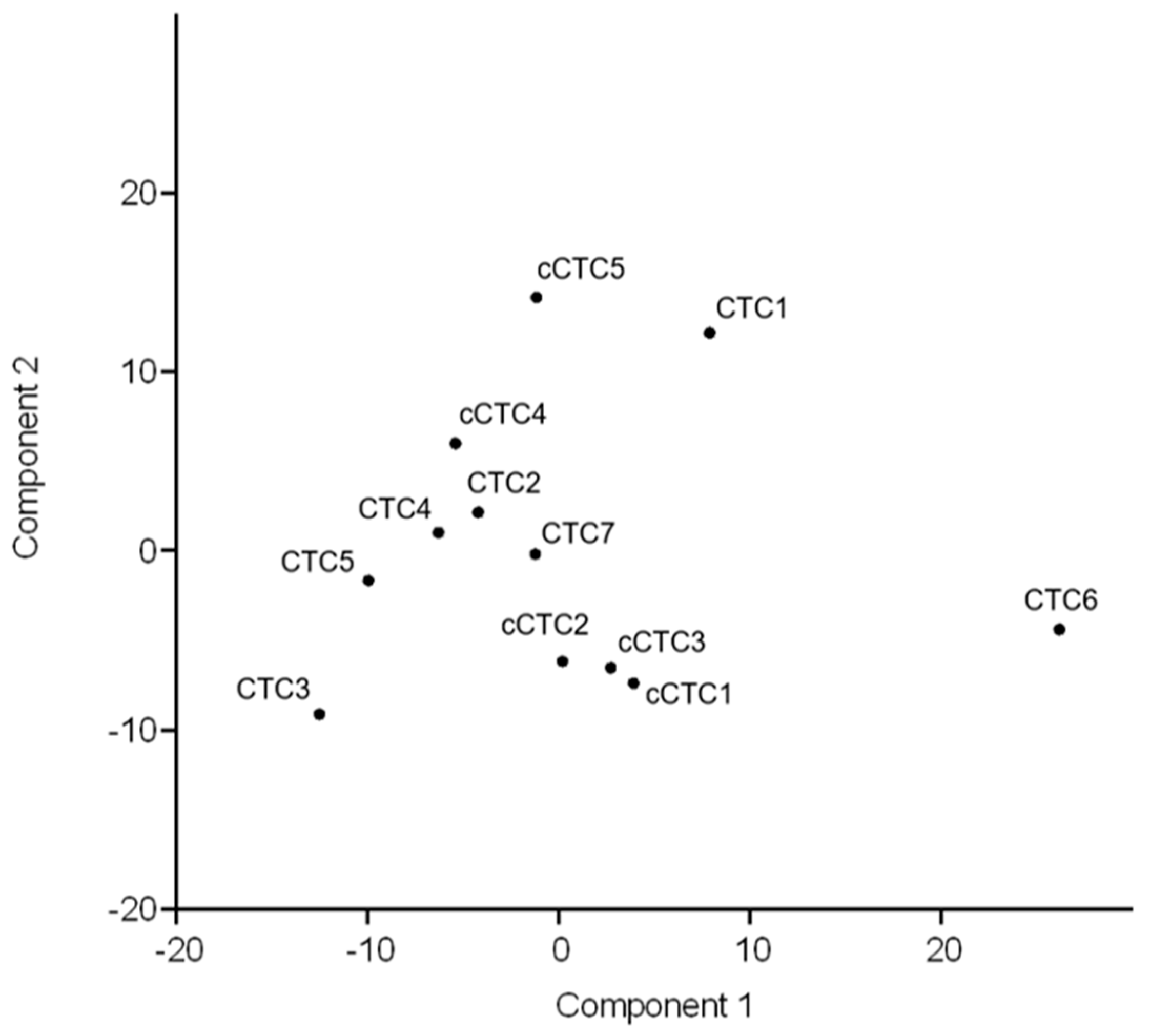
2.3.3. Concordance between Profiles of Uncultured and Cultured CTCs
3. Discussion
4. Materials and Methods
4.1. Cell Culture
4.2. Diagnostic Leukapheresis
4.3. Enrichment of Viable CTCs from Cryopreserved DLA Samples
4.4. CTC Culture
4.5. Staining and Isolation of Single Cells
4.6. RNA Isolation, cDNA, and RT-qPCR
4.7. Statistical Analysis and Single Cell Analysis
4.8. Patient Samples
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Anderson, W.F.; Chatterjee, N.; Ershler, W.B.; Brawley, O.W. Estrogen receptor breast cancer phenotypes in the surveillance, epidemiology, and end results database. Breast Cancer Res. Treat. 2002, 76, 27–36. [Google Scholar] [CrossRef] [PubMed]
- Colomer, R.; Beltran, M.; Dorcas, J.; Cortes-Funes, H.; Hornedo, J.; Valentin, V.; Vargas, C.; Mendiola, C.; Ciruelos, E. It is not time to stop progesterone receptor testing in breast cancer. J. Clin. Oncol. 2005, 23, 3868–3869. [Google Scholar] [CrossRef] [PubMed]
- Musgrove, E.A.; Sutherland, R.L. Biological determinants of endocrine resistance in breast cancer. Nat. Rev. Cancer 2009, 9, 631–643. [Google Scholar] [CrossRef] [PubMed]
- Polzer, B.; Medoro, G.; Pasch, S.; Fontana, F.; Zorzino, L.; Pestka, A.; Andergassen, U.; Meier-Stiegen, F.; Czyz, Z.T.; Alberter, B.; et al. Molecular profiling of single circulating tumor cells with diagnostic intention. Embo. Mol. Med. 2014, 6, 1371–1386. [Google Scholar] [CrossRef] [PubMed]
- Bidard, F.C.; Proudhon, C.; Pierga, J.Y. Circulating tumor cells in breast cancer. Mol. Oncol. 2016, 10, 418–430. [Google Scholar] [CrossRef] [PubMed]
- Coumans, F.A.; Ligthart, S.T.; Uhr, J.W.; Terstappen, L.W. Challenges in the enumeration and phenotyping of ctc. Clin. Cancer Res. 2012, 18, 5711–5718. [Google Scholar] [CrossRef] [PubMed]
- Fischer, J.C.; Niederacher, D.; Topp, S.A.; Honisch, E.; Schumacher, S.; Schmitz, N.; Zacarias Föhrding, L.; Vay, C.; Hoffmann, I.; Kasprowicz, N.S.; et al. Diagnostic leukapheresis enables reliable detection of circulating tumor cells of nonmetastatic cancer patients. Proc. Natl. Acad. Sci. USA 2013, 110, 16580–16585. [Google Scholar] [CrossRef] [PubMed]
- Andree, K.C.; Mentink, A.; Zeune, L.L.; Terstappen, L.W.M.M.; Stoecklein, N.H.; Neves, R.P.; Driemel, C.; Lampignano, R.; Yang, L.; Neubauer, H.; et al. Towards a real liquid biopsy in metastatic breast and prostate cancer: Diagnostic leukapheresis increases ctc yields in a european prospective multi-center study (ctctrap). Int. J. Cancer 2018, 143, 2584–2591. [Google Scholar] [CrossRef] [PubMed]
- Fehm, T.N.; Meier-Stiegen, F.; Driemel, C.; Jäger, B.; Reinhardt, F.; Naskou, J.; Franken, A.; Neubauer, H.; Neves, R.P.L.; van Dalum, G.; et al. Diagnostic leukapheresis for ctc analysis in breast cancer patients: Ctc frequency, clinical experiences and recommendations for standardized reporting. Cytom. A. 2018, 93, 1213–1219. [Google Scholar] [CrossRef] [PubMed]
- Franken, A.; Driemel, C.; Behrens, B.; Meier-Stiegen, F.; Endris, V.; Stenzinger, A.; Niederacher, D.; Fischer, J.C.; Stoecklein, N.H.; Ruckhaeberle, E.; et al. Label-free enrichment and molecular characterization of viable circulating tumor cells from diagnostic leukapheresis products. Clin. Chem. 2019, 65, 549–558. [Google Scholar] [CrossRef] [PubMed]
- Xenidis, N.; Perraki, M.; Apostolaki, S.; Agelaki, S.; Kalbakis, K.; Vardakis, N.; Kalykaki, A.; Xyrafas, A.; Kakolyris, S.; Mavroudis, D.; et al. Differential effect of adjuvant taxane-based and taxane-free chemotherapy regimens on the ck-19 mrna-positive circulating tumour cells in patients with early breast cancer. Br. J. Cancer 2013, 108, 549–556. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Daskalaki, A.; Agelaki, S.; Perraki, M.; Apostolaki, S.; Xenidis, N.; Stathopoulos, E.; Kontopodis, E.; Hatzidaki, D.; Mavroudis, D.; Georgoulias, V. Detection of cytokeratin-19 mrna-positive cells in the peripheral blood and bone marrow of patients with operable breast cancer. Br. J. Cancer 2009, 101, 589–597. [Google Scholar] [CrossRef] [PubMed]
- Stathopoulou, A.; Ntoulia, M.; Perraki, M.; Apostolaki, S.; Mavroudis, D.; Malamos, N.; Georgoulias, V.; Lianidou, E.S. A highly specific real-time rt-pcr method for the quantitative determination of ck-19 mRNA positive cells in peripheral blood of patients with operable breast cancer. Int. J. Cancer 2006, 119, 1654–1659. [Google Scholar] [CrossRef] [PubMed]
- Gorges, T.M.; Kuske, A.; Röck, K.; Mauermann, O.; Müller, V.; Peine, S.; Verpoort, K.; Novosadova, V.; Kubista, M.; Riethdorf, S.; et al. Accession of tumor heterogeneity by multiplex transcriptome profiling of single circulating tumor cells. Clin. Chem. 2016, 62, 1504–1515. [Google Scholar] [CrossRef] [PubMed]
- Ma, C.X.; Sanchez, C.G.; Ellis, M.J. Predicting endocrine therapy responsiveness in breast cancer. Oncology 2009, 23, 133–142. [Google Scholar] [PubMed]
- Xu, L.; Mao, X.; Imrali, A.; Syed, F.; Mutsvangwa, K.; Berney, D.; Cathcart, P.; Hines, J.; Shamash, J.; Lu, Y.J. Optimization and evaluation of a novel size based circulating tumor cell isolation system. PLoS ONE 2015, 10, e0138032. [Google Scholar] [CrossRef] [PubMed]
- Ståhlberg, A.; Rusnakova, V.; Forootan, A.; Anderova, M.; Kubista, M. Rt-qpcr work-flow for single-cell data analysis. Methods 2013, 59, 80–88. [Google Scholar] [CrossRef]
- Kumar, N.; Singh, A.; Kulkarni, R.V. Transcriptional bursting in gene expression: Analytical results for general stochastic models. PLoS Comput. Biol. 2015, 11, e1004292. [Google Scholar] [CrossRef]
- Powell, A.A.; Talasaz, A.H.; Zhang, H.; Coram, M.A.; Reddy, A.; Deng, G.; Telli, M.L.; Advani, R.H.; Carlson, R.W.; Mollick, J.A.; et al. Single cell profiling of circulating tumor cells: Transcriptional heterogeneity and diversity from breast cancer cell lines. PLoS ONE 2012, 7, e33788. [Google Scholar] [CrossRef]
- de Cremoux, P.; Tran-Perennou, C.; Brockdorff, B.L.; Boudou, E.; Brünner, N.; Magdelénat, H.; Lykkesfeldt, A.E. Validation of real-time rt-pcr for analysis of human breast cancer cell lines resistant or sensitive to treatment with antiestrogens. Endocr. Relat. Cancer 2003, 10, 409–418. [Google Scholar] [CrossRef]
- Elowitz, M.B.; Levine, A.J.; Siggia, E.D.; Swain, P.S. Stochastic gene expression in a single cell. Science 2002, 297, 1183–1186. [Google Scholar] [CrossRef] [PubMed]
- Raj, A.; Peskin, C.S.; Tranchina, D.; Vargas, D.Y.; Tyagi, S. Stochastic mrna synthesis in mammalian cells. PLoS Biol. 2006, 4, e309. [Google Scholar] [CrossRef] [PubMed]
- Reinhardt, F.; Franken, A.; Fehm, T.; Neubauer, H. Navigation through inter- and intratumoral heterogeneity of endocrine resistance mechanisms in breast cancer: A potential role for liquid biopsies? Tumour Biol. 2017, 39, 1010428317731511. [Google Scholar] [CrossRef] [PubMed]
- Riethdorf, S.; Müller, V.; Zhang, L.; Rau, T.; Loibl, S.; Komor, M.; Roller, M.; Huober, J.; Fehm, T.; Schrader, I.; et al. Detection and her2 expression of circulating tumor cells: Prospective monitoring in breast cancer patients treated in the neoadjuvant geparquattro trial. Clin. Cancer Res. 2010, 16, 2634–2645. [Google Scholar] [CrossRef] [PubMed]
- Meng, S.; Tripathy, D.; Shete, S.; Ashfaq, R.; Haley, B.; Perkins, S.; Beitsch, P.; Khan, A.; Euhus, D.; Osborne, C.; et al. Her-2 gene amplification can be acquired as breast cancer progresses. Proc. Natl. Acad. Sci. USA 2004, 101, 9393–9398. [Google Scholar] [CrossRef] [PubMed]
- Aktas, B.; Kasimir-Bauer, S.; Müller, V.; Janni, W.; Fehm, T.; Wallwiener, D.; Pantel, K.; Tewes, M.; Group, D.S. Comparison of the her2, estrogen and progesterone receptor expression profile of primary tumor, metastases and circulating tumor cells in metastatic breast cancer patients. Cancer 2016, 16, 522. [Google Scholar] [PubMed]
- Montenegro, M.F.; Collado-González, M.e.M.; Fernández-Pérez, M.P.; Hammouda, M.B.; Tolordava, L.; Gamkrelidze, M.; Rodríguez-López, J.N. Promoting e2f1-mediated apoptosis in oestrogen receptor-α-negative breast cancer cells. Cancer 2014, 14, 539. [Google Scholar] [CrossRef] [PubMed]
- Shoman, N.; Klassen, S.; McFadden, A.; Bickis, M.G.; Torlakovic, E.; Chibbar, R. Reduced pten expression predicts relapse in patients with breast carcinoma treated by tamoxifen. Mod. Pathol. 2005, 18, 250–259. [Google Scholar] [CrossRef] [PubMed]
- Nakamura, K.; Terai, Y.; Tanabe, A.; Ono, Y.J.; Hayashi, M.; Maeda, K.; Fujiwara, S.; Ashihara, K.; Nakamura, M.; Tanaka, Y.; et al. Cd24 expression is a marker for predicting clinical outcome and regulates the epithelial-mesenchymal transition in ovarian cancer via both the akt and erk pathways. Oncol. Rep. 2017, 37, 3189–3200. [Google Scholar] [CrossRef]
- Zhang, Y.; Wei, J.; Wang, H.; Xue, X.; An, Y.; Tang, D.; Yuan, Z.; Wang, F.; Wu, J.; Zhang, J.; et al. Epithelial mesenchymal transition correlates with cd24+cd44+ and cd133+ cells in pancreatic cancer. Oncol. Rep. 2012, 27, 1599–1605. [Google Scholar]
- Thangavel, C.; Dean, J.L.; Ertel, A.; Knudsen, K.E.; Aldaz, C.M.; Witkiewicz, A.K.; Clarke, R.; Knudsen, E.S. Therapeutically activating rb: Reestablishing cell cycle control in endocrine therapy-resistant breast cancer. Endocr. Relat. Cancer 2011, 18, 333–345. [Google Scholar] [CrossRef] [PubMed]
- Turner, N.C.; Ro, J.; André, F.; Loi, S.; Verma, S.; Iwata, H.; Harbeck, N.; Loibl, S.; Huang Bartlett, C.; Zhang, K.; et al. Palbociclib in hormone-receptor-positive advanced breast cancer. N. Engl. J. Med. 2015, 373, 209–219. [Google Scholar] [CrossRef] [PubMed]
- Finn, R.S.; Martin, M.; Rugo, H.S.; Jones, S.; Im, S.A.; Gelmon, K.; Harbeck, N.; Lipatov, O.N.; Walshe, J.M.; Moulder, S.; et al. Palbociclib and letrozole in advanced breast cancer. N. Engl. J. Med. 2016, 375, 1925–1936. [Google Scholar] [CrossRef] [PubMed]
- Hortobagyi, G.N.; Stemmer, S.M.; Burris, H.A.; Yap, Y.S.; Sonke, G.S.; Paluch-Shimon, S.; Campone, M.; Blackwell, K.L.; André, F.; Winer, E.P.; et al. Ribociclib as first-line therapy for hr-positive, advanced breast cancer. N. Engl. J. Med. 2016, 375, 1738–1748. [Google Scholar] [CrossRef] [PubMed]
- Chen, C.; Zhao, S.; Karnad, A.; Freeman, J.W. The biology and role of cd44 in cancer progression: Therapeutic implications. J. Hematol. Oncol. 2018, 11, 64. [Google Scholar] [CrossRef] [PubMed]
- Nath, S.; Mukherjee, P. Muc1: A multifaceted oncoprotein with a key role in cancer progression. Trends Mol. Med. 2014, 20, 332–342. [Google Scholar] [CrossRef] [PubMed]
- Bitler, B.G.; Goverdhan, A.; Schroeder, J.A. Muc1 regulates nuclear localization and function of the epidermal growth factor receptor. J. Cell Sci. 2010, 123, 1716–1723. [Google Scholar] [CrossRef] [PubMed]
- Fribbens, C.; Garcia Murillas, I.; Beaney, M.; Hrebien, S.; O’Leary, B.; Kilburn, L.; Howarth, K.; Epstein, M.; Green, E.; Rosenfeld, N.; et al. Tracking evolution of aromatase inhibitor resistance with circulating tumour DNA analysis in metastatic breast cancer. Ann. Oncol. 2018, 29, 145–153. [Google Scholar] [CrossRef] [PubMed]
- Ma, C.X.; Reinert, T.; Chmielewska, I.; Ellis, M.J. Mechanisms of aromatase inhibitor resistance. Nat. Rev. Cancer 2015, 15, 261–275. [Google Scholar] [CrossRef]
- Garrido-Castro, A.C.; Goel, S. Cdk4/6 inhibition in breast cancer: Mechanisms of response and treatment failure. Curr. Breast Cancer Rep. 2017, 9, 26–33. [Google Scholar] [CrossRef]
- Yu, M.; Bardia, A.; Aceto, N.; Bersani, F.; Madden, M.W.; Donaldson, M.C.; Desai, R.; Zhu, H.; Comaills, V.; Zheng, Z.; et al. Cancer therapy. Ex vivo culture of circulating breast tumor cells for individualized testing of drug susceptibility. Science 2014, 345, 216–220. [Google Scholar] [CrossRef] [PubMed]
- Neumann, M.H.; Schneck, H.; Decker, Y.; Schömer, S.; Franken, A.; Endris, V.; Pfarr, N.; Weichert, W.; Niederacher, D.; Fehm, T.; et al. Isolation and characterization of circulating tumor cells using a novel workflow combining the cellsearch® system and the cellcelector™. Biotechnol. Prog. 2017, 33, 125–132. [Google Scholar] [CrossRef] [PubMed]
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Reinhardt, F.; Franken, A.; Meier-Stiegen, F.; Driemel, C.; Stoecklein, N.H.; Fischer, J.C.; Niederacher, D.; Ruckhaeberle, E.; Fehm, T.; Neubauer, H. Diagnostic Leukapheresis Enables Reliable Transcriptomic Profiling of Single Circulating Tumor Cells to Characterize Inter-Cellular Heterogeneity in Terms of Endocrine Resistance. Cancers 2019, 11, 903. https://doi.org/10.3390/cancers11070903
Reinhardt F, Franken A, Meier-Stiegen F, Driemel C, Stoecklein NH, Fischer JC, Niederacher D, Ruckhaeberle E, Fehm T, Neubauer H. Diagnostic Leukapheresis Enables Reliable Transcriptomic Profiling of Single Circulating Tumor Cells to Characterize Inter-Cellular Heterogeneity in Terms of Endocrine Resistance. Cancers. 2019; 11(7):903. https://doi.org/10.3390/cancers11070903
Chicago/Turabian StyleReinhardt, Florian, André Franken, Franziska Meier-Stiegen, Christiane Driemel, Nikolas H. Stoecklein, Johannes C. Fischer, Dieter Niederacher, Eugen Ruckhaeberle, Tanja Fehm, and Hans Neubauer. 2019. "Diagnostic Leukapheresis Enables Reliable Transcriptomic Profiling of Single Circulating Tumor Cells to Characterize Inter-Cellular Heterogeneity in Terms of Endocrine Resistance" Cancers 11, no. 7: 903. https://doi.org/10.3390/cancers11070903
APA StyleReinhardt, F., Franken, A., Meier-Stiegen, F., Driemel, C., Stoecklein, N. H., Fischer, J. C., Niederacher, D., Ruckhaeberle, E., Fehm, T., & Neubauer, H. (2019). Diagnostic Leukapheresis Enables Reliable Transcriptomic Profiling of Single Circulating Tumor Cells to Characterize Inter-Cellular Heterogeneity in Terms of Endocrine Resistance. Cancers, 11(7), 903. https://doi.org/10.3390/cancers11070903




