Immunoprofiling of Breast Cancer Antigens Using Antibodies Derived from Local Lymph Nodes
Abstract
:1. Introduction
2. Results
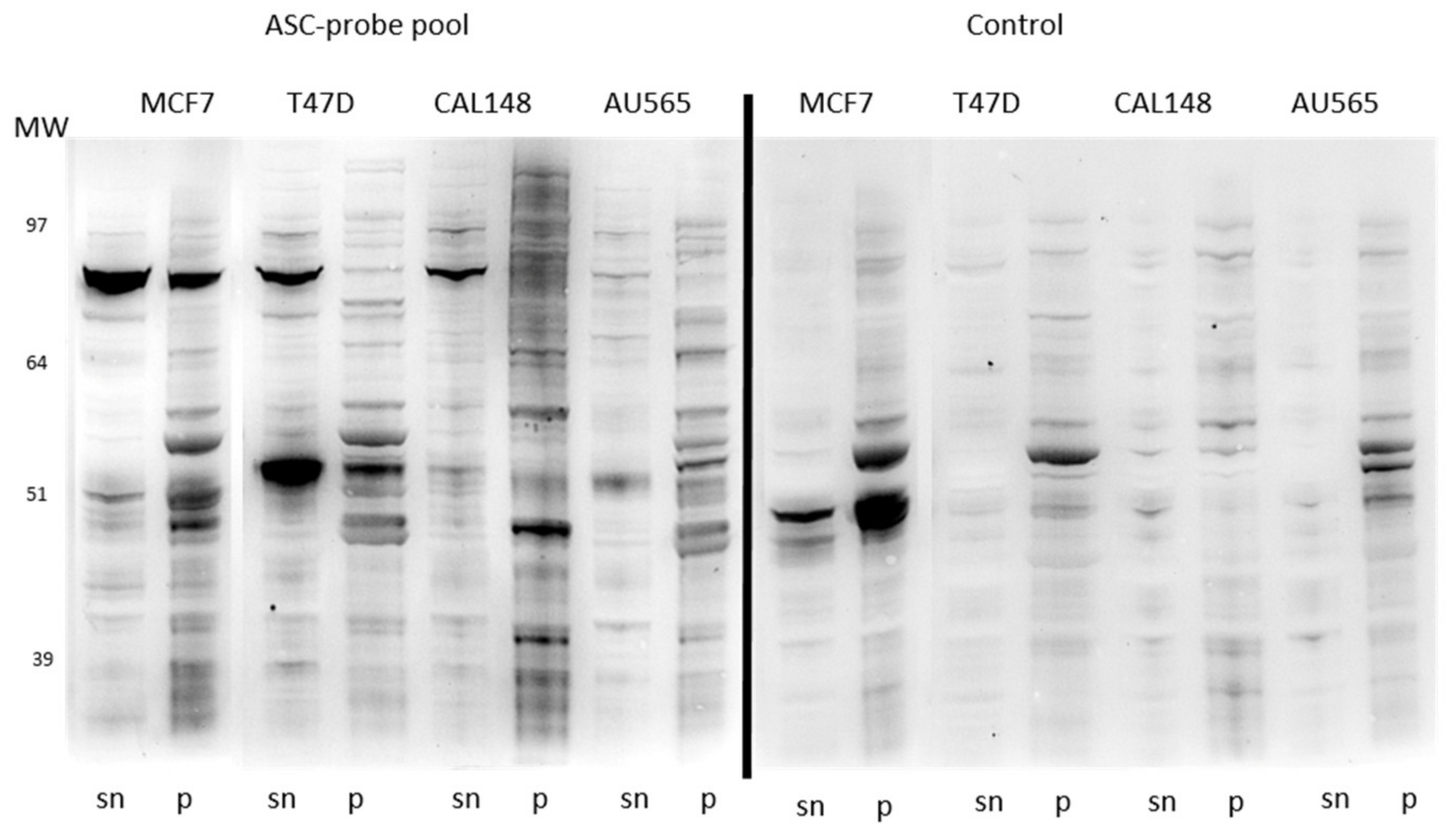
2.1. Immunoglobulin Concentration and Antibody Reactivity Profile of ASC-Probes against MCF-7 Extracts
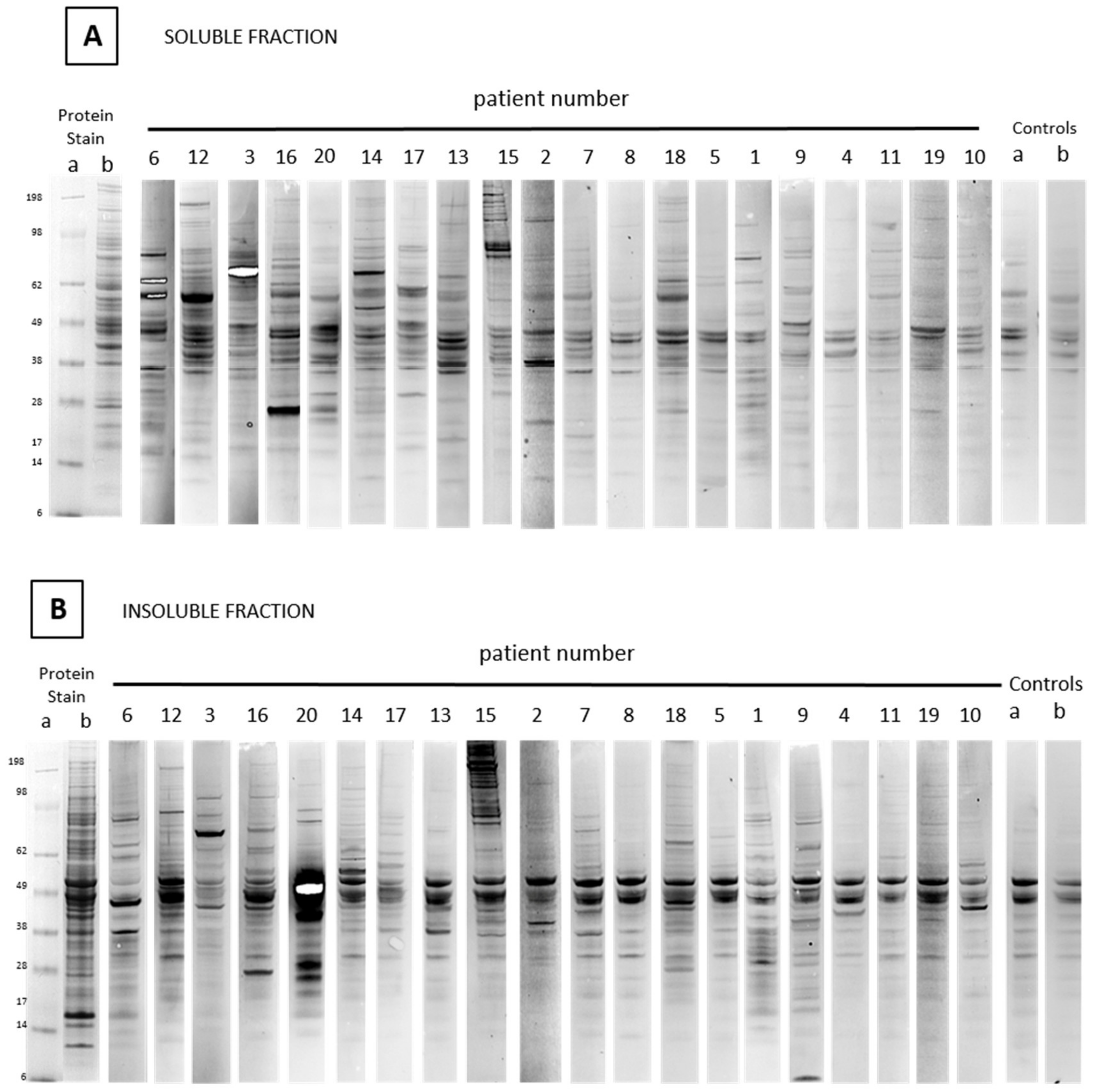
2.2. Comparative Reactivity of ASC-Probes against Different Breast Cancer Cell Lines
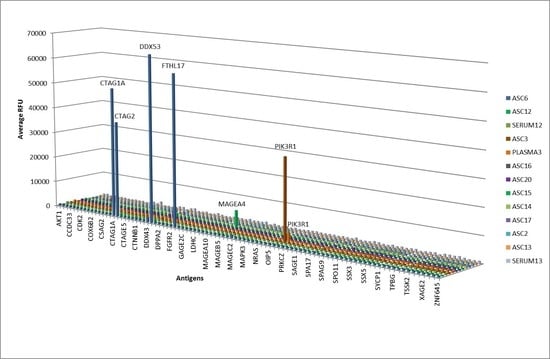
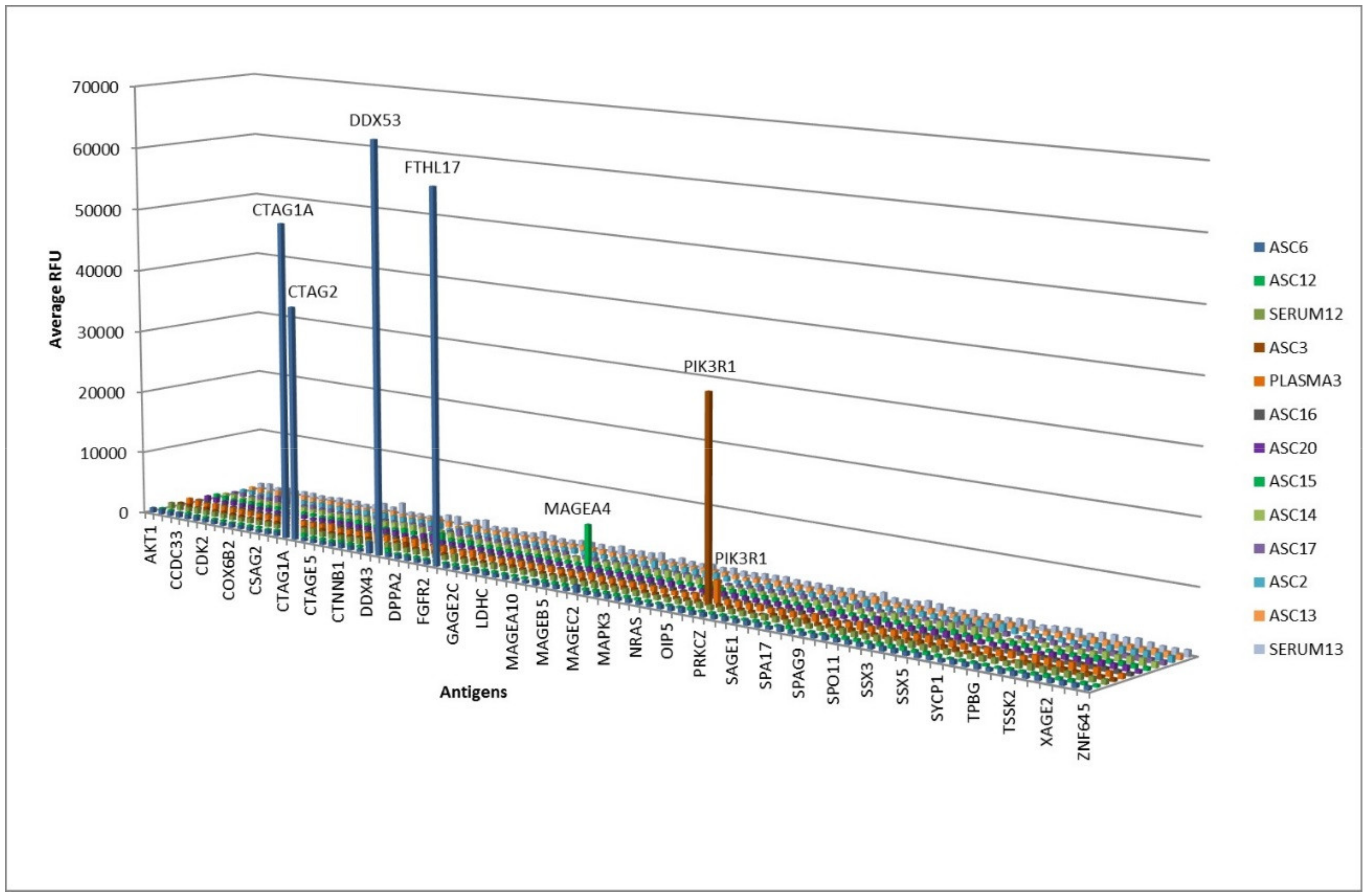
2.3. Antigen Recognition Profile of ASC-Probes on a Custom Protein Microarray
3. Discussion
4. Materials and Methods
4.1. Generation of Lymph Node ASC-Probes and Quantitation of Secreted Antibodies
4.2. Western Blotting of ASC-Probes against Soluble and Insoluble Extracts of Cancer Cell Lines
4.3. Analysis of Overall Reactivity with Image J
4.4. Antibody Profiling Using a Custom Protein Microarray
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Conflicts of Interest
References
- Gnjatic, S.; Bronte, V.; Brunet, L.R.; Butler, M.O.; Disis, M.L.; Galon, J.; Hakansson, L.G.; Hanks, B.A.; Karanikas, V.; Khleif, S.N.; et al. Identifying baseline immune-related biomarkers to predict clinical outcome of immunotherapy. J. Immunother. Cancer 2017, 5. [Google Scholar] [CrossRef] [PubMed]
- Wei, S.C.; Duffy, C.R.; Allison, J.P. Fundamental Mechanisms of Immune Checkpoint Blockade Therapy. Cancer Discov. 2018, 8, 1069–1086. [Google Scholar] [CrossRef] [PubMed]
- Desrichard, A.; Snyder, A.; Chan, T.A. Cancer Neoantigens and Applications for Immunotherapy. Clin. Cancer Res. 2016, 22, 807–812. [Google Scholar] [CrossRef] [PubMed]
- Meeusen, E.; Lim, E.; Mathivanan, S. Secreted Tumor Antigens—Immune Biomarkers for Diagnosis and Therapy. Proteomics 2017, 17. [Google Scholar] [CrossRef]
- Goodnow, C.C.; Vinuesa, C.G.; Randall, K.L.; Mackay, F.; Brink, R. Control systems and decision making for antibody production. Nat. Immunol. 2010, 11, 681–688. [Google Scholar] [CrossRef] [PubMed]
- Ma, Y.; Hendershot, L.M. The stressful road to antibody secretion. Nat. Immunol. 2003, 4, 310–311. [Google Scholar] [CrossRef]
- Jarossay, A.; Hadzhieva, M.; Kaveri, S.V.; Lacroix-Desmazes, S.; Dimitrov, J.D. Natural and Induced Antibody Polyreactivity. Anticancer Agents Med. Chem. 2015, 15, 1230–1241. [Google Scholar] [CrossRef]
- Lacroix-Desmazes, S.; Kaveri, S.V.; Mouthon, L.; Ayouba, A.; Malanchere, E.; Coutinho, A.; Kazatchkine, M.D. Self-reactive antibodies (natural autoantibodies) in healthy individuals. J. Immunol. Methods 1998, 216, 117–137. [Google Scholar] [CrossRef]
- Duarte, J.G.; Blackburn, J.M. Advances in the development of human protein microarrays. Expert Rev. Proteom. 2017, 14, 627–641. [Google Scholar] [CrossRef] [PubMed]
- Black, M.M. Immunology of breast cancer: clinical implications. Prog. Clin. Cancer 1975, 6, 115–137. [Google Scholar] [PubMed]
- Kohrt, H.E.; Nouri, N.; Nowels, K.; Johnson, D.; Holmes, S.; Lee, P.P. Profile of immune cells in axillary lymph nodes predicts disease-free survival in breast cancer. PLoS Med. 2005, 2, e284. [Google Scholar] [CrossRef]
- Coronella-Wood, J.A.; Hersh, E.M. Naturally occurring B-cell responses to breast cancer. Cancer Immunol. Immunother. CII 2003, 52, 715–738. [Google Scholar] [CrossRef]
- Mahmoud, S.M.; Lee, A.H.; Paish, E.C.; Macmillan, R.D.; Ellis, I.O.; Green, A.R. The prognostic significance of B lymphocytes in invasive carcinoma of the breast. Breast Cancer Res. Treat. 2012, 132, 545–553. [Google Scholar] [CrossRef] [PubMed]
- Iglesia, M.D.; Vincent, B.G.; Parker, J.S.; Hoadley, K.A.; Carey, L.A.; Perou, C.M.; Serody, J.S. Prognostic B-cell signatures using mRNA-seq in patients with subtype-specific breast and ovarian cancer. Clin. Cancer Res. 2014, 20, 3818–3829. [Google Scholar] [CrossRef] [PubMed]
- McWilliam, H.E.; Driguez, P.; Piedrafita, D.; Maupin, K.A.; Haab, B.B.; McManus, D.P.; Meeusen, E.N. The developing schistosome worms elicit distinct immune responses in different tissue regions. Immunol. Cell Biol. 2013, 91, 477–485. [Google Scholar] [CrossRef]
- Meeusen, E.; Brandon, M. The use of antibody-secreting cell probes to reveal tissue-restricted immune responses during infection. Eur. J. Immunol. 1994, 24, 469–474. [Google Scholar] [CrossRef]
- McWilliam, H.E.; Driguez, P.; Piedrafita, D.; McManus, D.P.; Meeusen, E.N. Novel immunomic technologies for schistosome vaccine development. Parasite Immunol. 2012, 34, 276–284. [Google Scholar] [CrossRef] [PubMed]
- Rodriguez-Pinto, D.; Sparkowski, J.; Keough, M.P.; Phoenix, K.N.; Vumbaca, F.; Han, D.K.; Gundelfinger, E.D.; Beesley, P.; Claffey, K.P. Identification of novel tumor antigens with patient-derived immune-selected antibodies. Cancer Immunol. Immunother. CII 2009, 58, 221–234. [Google Scholar] [CrossRef]
- Mizukami, M.; Hanagiri, T.; Baba, T.; Fukuyama, T.; Nagata, Y.; So, T.; Ichiki, Y.; Sugaya, M.; Yasuda, M.; Takenoyama, M.; et al. Identification of tumor associated antigens recognized by IgG from tumor-infiltrating B cells of lung cancer: Correlation between Ab titer of the patient’s sera and the clinical course. Cancer Sci. 2005, 96, 882–888. [Google Scholar] [CrossRef]
- Punt, C.J.; Barbuto, J.A.; Zhang, H.; Grimes, W.J.; Hatch, K.D.; Hersh, E.M. Anti-tumor antibody produced by human tumor-infiltrating and peripheral blood B lymphocytes. Cancer Immunol. Immunother. CII 1994, 38, 225–232. [Google Scholar] [CrossRef] [PubMed]
- Germain, C.; Gnjatic, S.; Tamzalit, F.; Knockaert, S.; Remark, R.; Goc, J.; Lepelley, A.; Becht, E.; Katsahian, S.; Bizouard, G.; et al. Presence of B cells in tertiary lymphoid structures is associated with a protective immunity in patients with lung cancer. Am. J. Respir. Crit. Care Med. 2014, 189, 832–844. [Google Scholar] [CrossRef]
- Tsou, P.; Katayama, H.; Ostrin, E.J.; Hanash, S.M. The Emerging Role of B Cells in Tumor Immunity. Cancer Res. 2016, 76, 5597–5601. [Google Scholar] [CrossRef]
- Meeusen, E.N.; Brandon, M.R. Antibody secreting cells as specific probes for antigen identification. J. Immunol. Methods 1994, 172, 71–76. [Google Scholar] [CrossRef]
- Novinger, L.J.; Ashikaga, T.; Krag, D.N. Identification of tumor-binding scFv derived from clonally related B cells in tumor and lymph node of a patient with breast cancer. Cancer Immunol. Immunother. CII 2015, 64, 29–39. [Google Scholar] [CrossRef] [PubMed]
- Criscitiello, C.; Esposito, A.; Gelao, L.; Fumagalli, L.; Locatelli, M.; Minchella, I.; Adamoli, L.; Goldhirsch, A.; Curigliano, G. Immune approaches to the treatment of breast cancer, around the corner? Breast Cancer Res. 2014, 16. [Google Scholar] [CrossRef]
- Makhoul, I.; Atiq, M.; Alwbari, A.; Kieber-Emmons, T. Breast Cancer Immunotherapy: An Update. Breast Cancer Basic Clin. Res. 2018, 12. [Google Scholar] [CrossRef] [PubMed]
- Jiang, G.; Zhang, S.; Yazdanparast, A.; Li, M.; Pawar, A.V.; Liu, Y.; Inavolu, S.M.; Cheng, L. Comprehensive comparison of molecular portraits between cell lines and tumors in breast cancer. BMC Genom. 2016, 17. [Google Scholar] [CrossRef]
- Alizadeh, A.A.; Aranda, V.; Bardelli, A.; Blanpain, C.; Bock, C.; Borowski, C.; Caldas, C.; Califano, A.; Doherty, M.; Elsner, M.; et al. Toward understanding and exploiting tumor heterogeneity. Nat. Med. 2015, 21, 846–853. [Google Scholar] [CrossRef]
- Scanlan, M.J.; Gure, A.O.; Jungbluth, A.A.; Old, L.J.; Chen, Y.T. Cancer/testis antigens: An expanding family of targets for cancer immunotherapy. Immunol. Rev. 2002, 188, 22–32. [Google Scholar] [CrossRef]
- Schmid, P.; Adams, S.; Rugo, H.S.; Schneeweiss, A.; Barrios, C.H.; Iwata, H.; Dieras, V.; Hegg, R.; Im, S.A.; Shaw Wright, G.; et al. Atezolizumab and Nab-Paclitaxel in Advanced Triple-Negative Breast Cancer. N. Engl. J. Med. 2018. [Google Scholar] [CrossRef]
- Yan, L.X.; Liu, Y.H.; Xiang, J.W.; Wu, Q.N.; Xu, L.B.; Luo, X.L.; Zhu, X.L.; Liu, C.; Xu, F.P.; Luo, D.L.; et al. PIK3R1 targeting by miR-21 suppresses tumor cell migration and invasion by reducing PI3K/AKT signaling and reversing EMT, and predicts clinical outcome of breast cancer. Int. J. Oncol. 2016, 48, 471–484. [Google Scholar] [CrossRef] [PubMed]
- Network, C.G.A. Comprehensive molecular portraits of human breast tumours. Nature 2012, 490, 61–70. [Google Scholar] [CrossRef]
- McDaniel, J.R.; Pero, S.C.; Voss, W.N.; Shukla, G.S.; Sun, Y.; Schaetzle, S.; Lee, C.H.; Horton, A.P.; Harlow, S.; Gollihar, J.; et al. Identification of tumor-reactive B cells and systemic IgG in breast cancer based on clonal frequency in the sentinel lymph node. Cancer Immunol. Immunother. CII 2018, 67, 729–738. [Google Scholar] [CrossRef] [PubMed]
- Petibois, C.; Cazorla, G.; Cassaigne, A.; Deleris, G. Plasma protein contents determined by Fourier-transform infrared spectrometry. Clin. Chem. 2001, 47, 730–738. [Google Scholar] [PubMed]
- Da Gama Duarte, J.; Goosen, R.W.; Lawry, P.J.; Blackburn, J.M. PMA: Protein Microarray Analyser, a user-friendly tool for data processing and normalization. BMC Res. Notes 2018, 11, 156. [Google Scholar] [CrossRef] [PubMed]
| Patient No. | Age | Tumor Grade | ER | PR | HER2 (ISH) | TNM Classification | Tumor Size (mm) | Pathological Features |
|---|---|---|---|---|---|---|---|---|
| 1 | 67 | 2 | + | + | neg | T2N1M0 | 33 | IDC |
| 2 | 49 | 1 | + | + | neg | T2N2M0 | 26 | Invasive micropapillary carcinoma |
| 3 | 42 | 3 | + | + | 2+ Non-amplified | T2N2M0 | 28 | IDC |
| 4 | 44 | 3 | neg | neg | 3+ Amplified | T3N2M0 | 65 | IDC |
| 5 | 66 | 3 | neg | neg | 3+ Amplified | T1N1M0 | 15 and 4.5 | IDC, multifocal |
| 6 | 58 | 3 | neg | neg | 2+ Amplified | TXN2M0 | NA | Occult Primary Breast Cancer |
| 7 | 73 | 2 | + | + | neg | T3N2M1 | 110 | IDC |
| 8 | 72 | 3 | + | + | neg | T2N1M0 | 22 | IDC |
| 9 | 78 | 2 | + | + | neg | TXN3M0 | NA | ILC |
| 10 | 44 | 2 | + | + | neg | T4N3M0 | 70 | IDC, multifocal |
| 11 | 58 | 1 | + | + | neg | T2N1M0 | 25 | IDC |
| 12 | 60 | 2 | + | + | neg | T3N2M0 | 60 | IDC |
| 13 | 63 | 3 | + | + | 2+ Amplified | T1N1M0 | 13 | IDC |
| 14 | 71 | 3 | + | + | neg | T4N2M0 | 60 | ILC |
| 15 | 64 | 3 | + | + | neg | T2N2M0 | 65 and 13 | IDC, multifocal |
| 16 | 45 | 3 | + | + | neg | T2N2M0 | 25 and 15 | IDC, multifocal |
| 17 | 39 | 3 | neg | neg | 3+ Amplified | NA | 4 (DCIS) | Residual DCIS post preoperative chemotherapy. Pathological complete response. |
| 18 | 60 | 3 | + | neg | 3+ Amplified | T2N1M0 | 40 | IDC |
| 19 | 39 | 3 | NA | NA | NA | T2N1M0 | 32 | IDC |
| 20 | 75 | 2 | + | + | neg | T2N2M0 | 21 + 15 | IDC, multifocal |
| Patient Number | µg IgG/mL | Relative Reactivity Ratio ASC-Probe/Control 1 | ||
|---|---|---|---|---|
| Soluble (S) | Insoluble (I) | Total (Ratio S/I) | ||
| Strong | ||||
| 6 | 5.882 | 12.69 | 11.54 | 24.23 (1.2) |
| 12 | 0.902 | 15.82 | 3.34 | 19.16 (4.7) |
| 3 | 3.6 | 12.33 | 6.58 | 18.92 (1.9) |
| 16 | 0.453 | 5.52 | 3.20 | 8.72 (1.7) |
| 20 | 1.150 | 2.29 | 5.04 | 7.34 (0.5) |
| 15 | 0.396 | 1.44 | 3.21 | 4.66 (0.4) |
| 14 | 0.341 | 3.90 | 2.04 | 5.94 (1.9) |
| 17 | 0.790 | 3.59 | 1.60 | 5.20 (2.2) |
| 2 | 0.468 | 2.47 | 1.71 | 4.18 (1.4) |
| 13 | 0.423 | 3.26 | 1.77 | 5.03 (1.8) |
| Weak/negative | ||||
| 7 | 0.549 | 1.85 | 1.44 | 3.29 (1.3) |
| 8 | 0.421 | 2.10 | 1.35 | 3.45 (1.6) |
| 18 | 0.760 | 1.53 | 1.04 | 2.57 (1.5) |
| 1 | 0.174 | 0.79 | 1.20 | 1.99 (0.7) |
| 10 | 0.207 | 0.62 | 0.66 | 1.28 (0.9) |
| 5 | 0.173 | 0.8 | 1.3 | 2.2 (0.6) |
| 19 | 1.060 | 0.75 | 0.75 | 1.50 (1.0) |
| 4 | 0.188 | 0.9 | 0.7 | 1.6 (1.3) |
| 9 | 0.539 | 0.91 | 0.89 | 1.80 (1.0) |
| 11 | 0.912 | 0.87 | 0.68 | 1.55 (1.3) |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Young, A.R.; Duarte, J.D.G.; Coulson, R.; O’Brien, M.; Deb, S.; Lopata, A.; Behren, A.; Mathivanan, S.; Lim, E.; Meeusen, E. Immunoprofiling of Breast Cancer Antigens Using Antibodies Derived from Local Lymph Nodes. Cancers 2019, 11, 682. https://doi.org/10.3390/cancers11050682
Young AR, Duarte JDG, Coulson R, O’Brien M, Deb S, Lopata A, Behren A, Mathivanan S, Lim E, Meeusen E. Immunoprofiling of Breast Cancer Antigens Using Antibodies Derived from Local Lymph Nodes. Cancers. 2019; 11(5):682. https://doi.org/10.3390/cancers11050682
Chicago/Turabian StyleYoung, Anna Rachel, Jessica Da Gama Duarte, Rhiannon Coulson, Megan O’Brien, Siddhartha Deb, Alex Lopata, Andreas Behren, Suresh Mathivanan, Elgene Lim, and Els Meeusen. 2019. "Immunoprofiling of Breast Cancer Antigens Using Antibodies Derived from Local Lymph Nodes" Cancers 11, no. 5: 682. https://doi.org/10.3390/cancers11050682
APA StyleYoung, A. R., Duarte, J. D. G., Coulson, R., O’Brien, M., Deb, S., Lopata, A., Behren, A., Mathivanan, S., Lim, E., & Meeusen, E. (2019). Immunoprofiling of Breast Cancer Antigens Using Antibodies Derived from Local Lymph Nodes. Cancers, 11(5), 682. https://doi.org/10.3390/cancers11050682