Genomic Profiling of the Steroidogenic Acute Regulatory Protein in Breast Cancer: In Silico Assessments and a Mechanistic Perspective
Abstract
1. Introduction
2. Materials and Methods
2.1. TCGA Hormone Responsive Cancer Tumors and Their Correlation to Copy Number Alterations of StAR and Steroidogenic Enzyme Genes
2.2. Expression of ER, PR, and HER2 in Breast Cancer Tumors
2.3. Amplification of the StAR Gene in Breast Cancer Studies
2.4. Mutational Portraits of the StAR Gene in TCGA Hormone Responsive Cancers
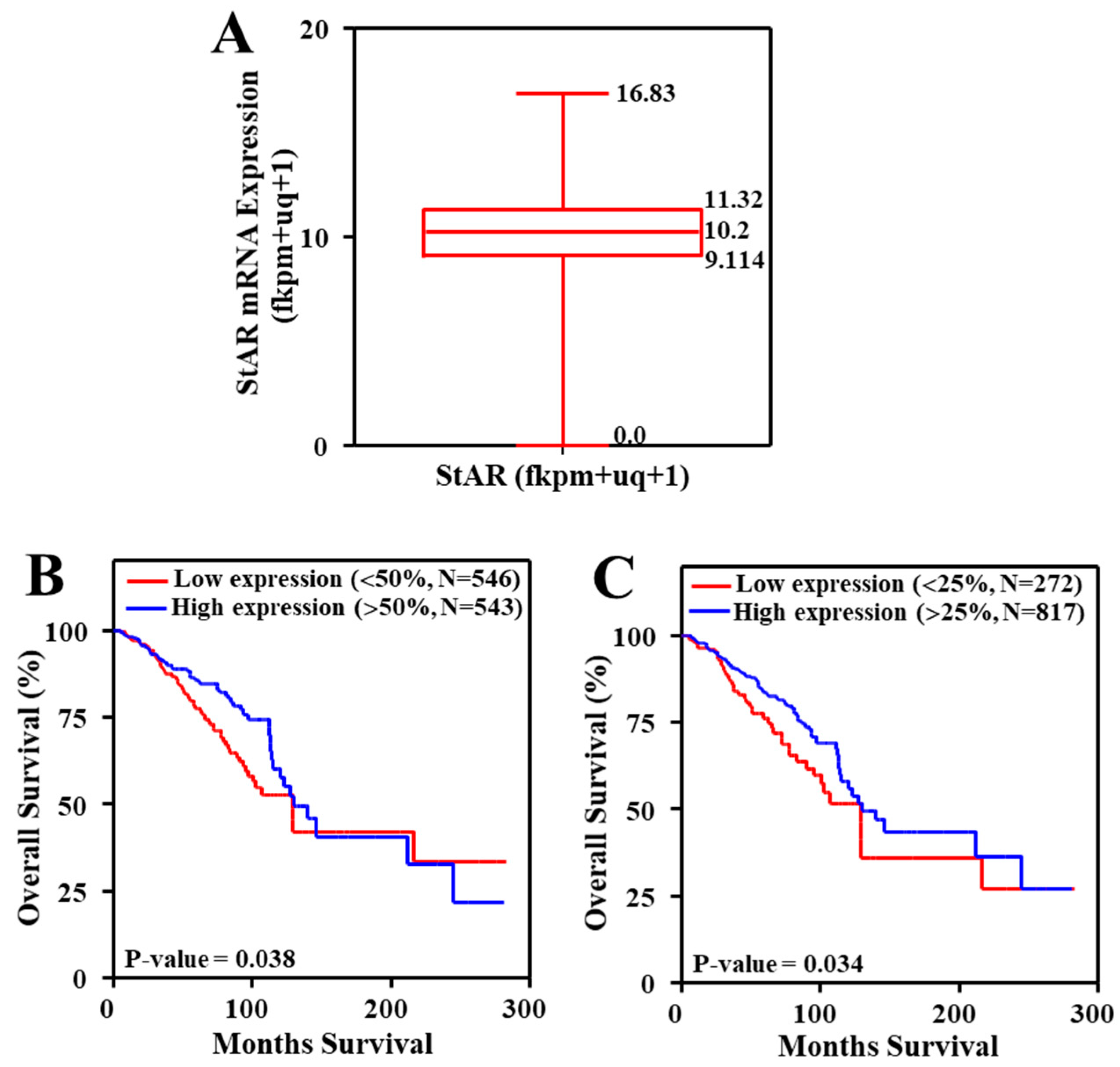
2.5. Expression of StAR mRNA in TCGA Breast Cancer Tumors and Their Correlation to TNM Stages
2.6. Generation of Kaplan-Meier Curves and Overall Survival Analyses
2.7. Statistical Analysis
3. Results
3.1. Assessment of StAR CNAs in Various Hormone Sensitive Cancers
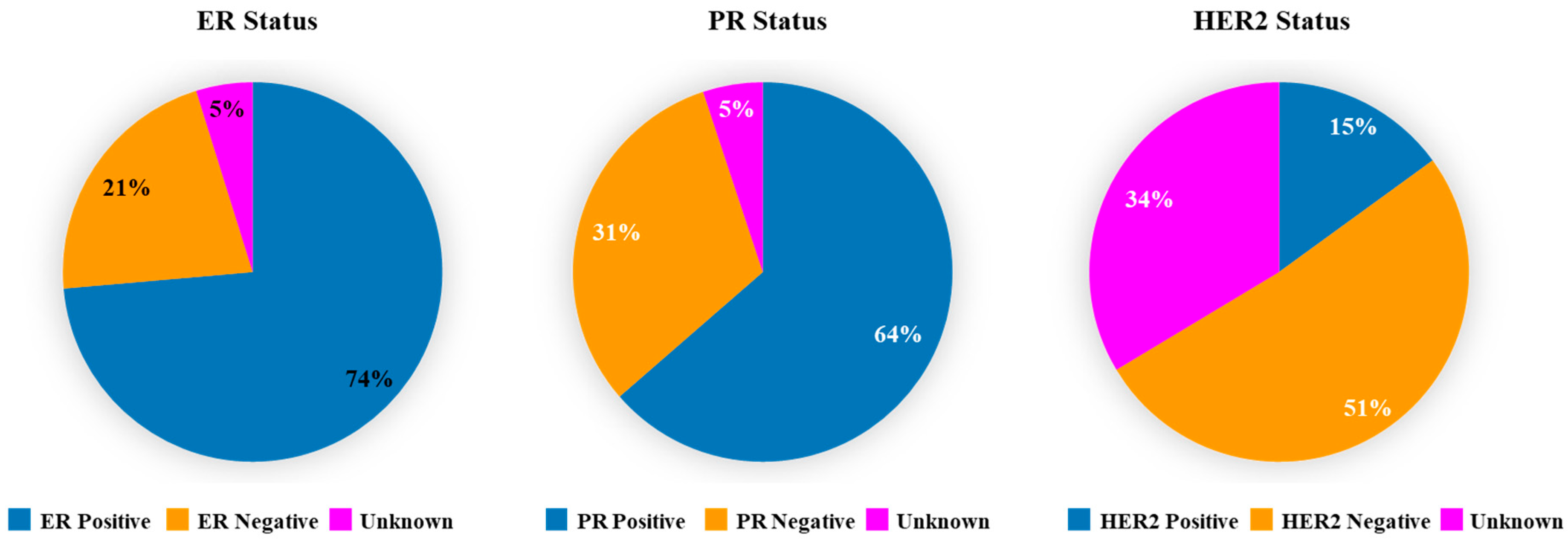
3.2. Expression of ER, PR and HER2 in TCGA Breast Cancer Tumors
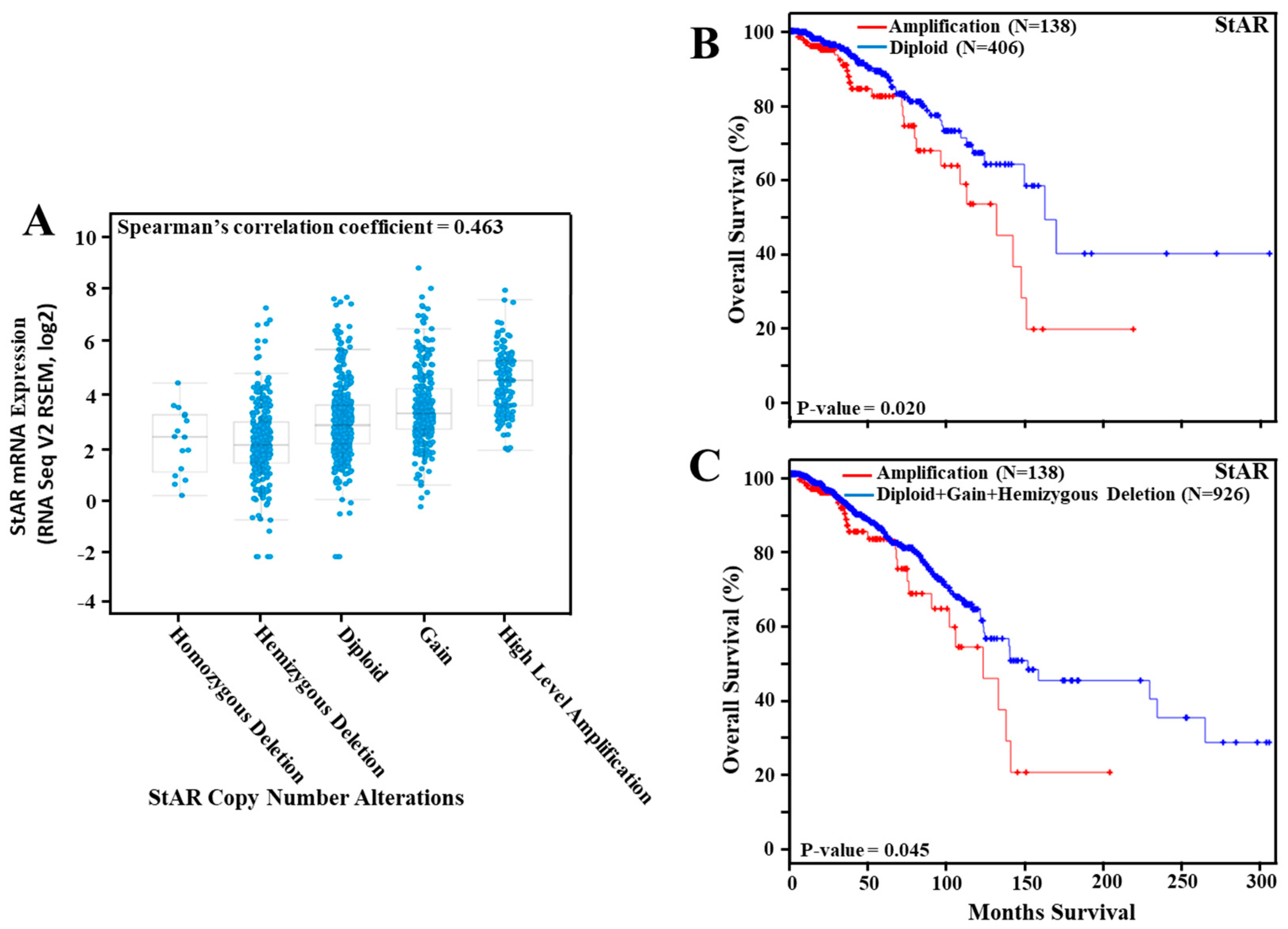
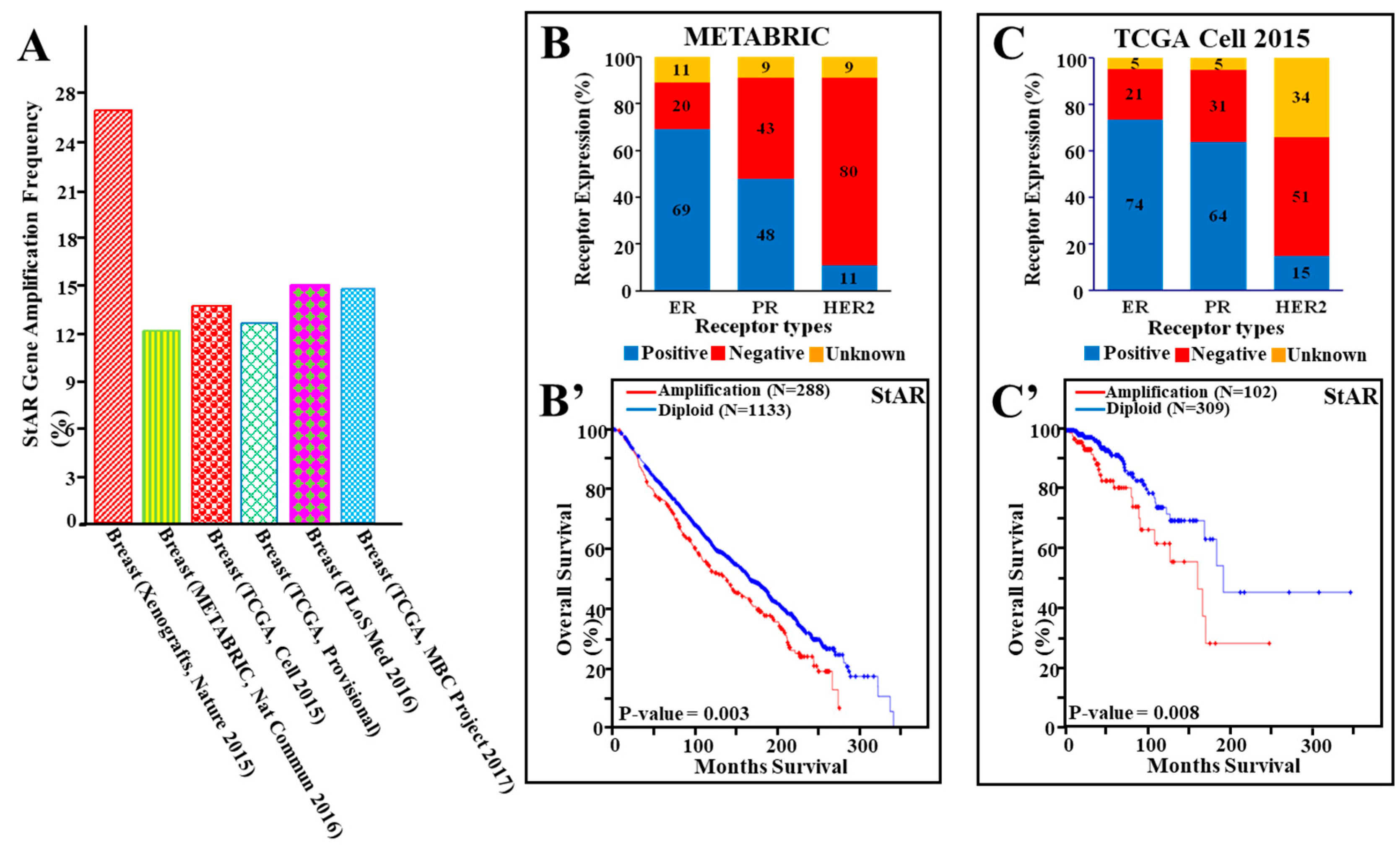
3.3. Amplification of the StAR Gene in Breast Cancer and Its Correlation to Overall Survival
3.4. StAR Gene Amplification, Hormone Receptor Expression, and Their Correlation to Cancer Survival in a Number of Breast Cancer Studies
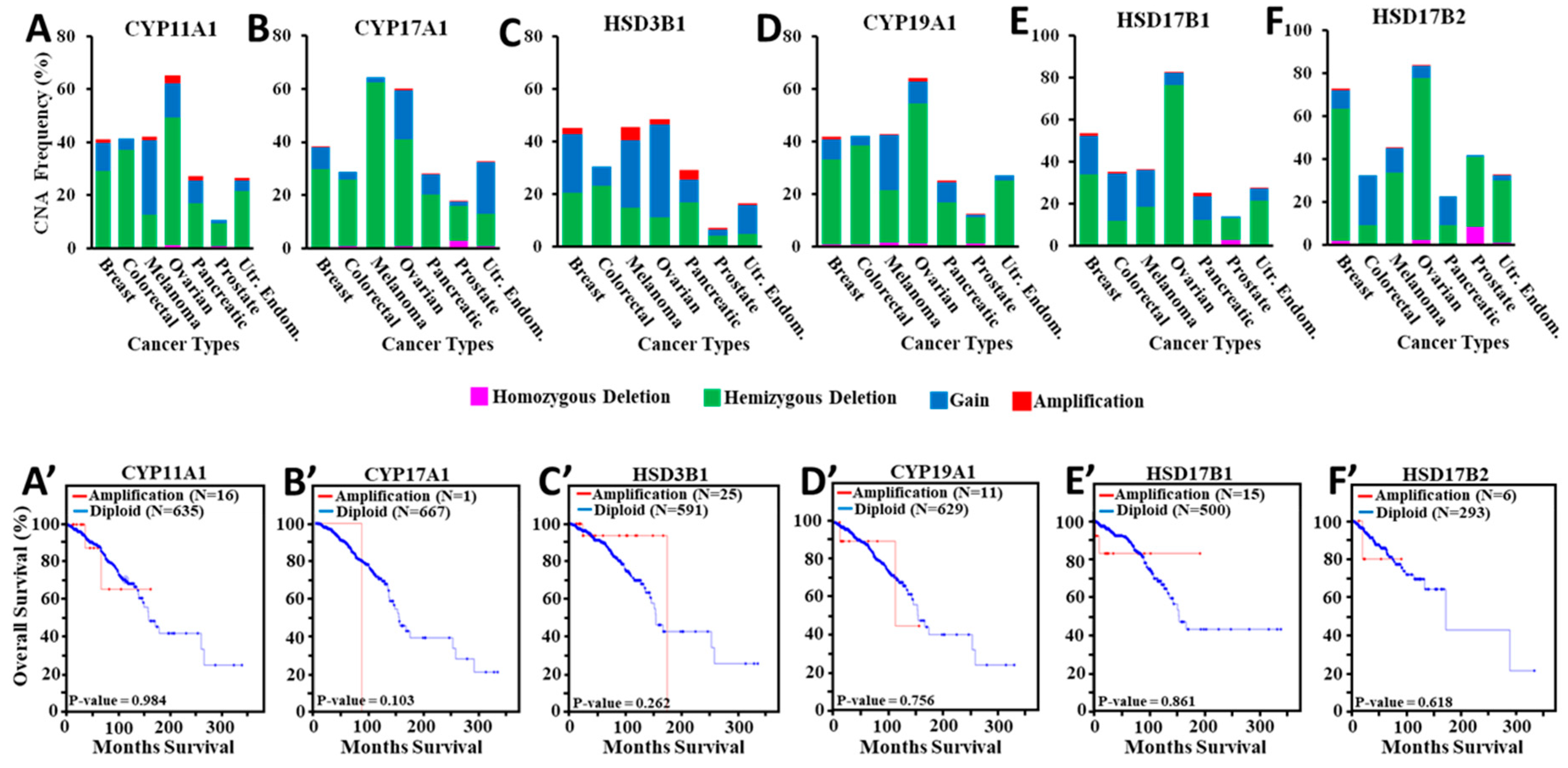
3.5. Amplification of Steroidogenic Enzyme Genes and Their Correlation to Overall Breast Cancer Survival
3.6. Assessment of StAR Gene Mutation in Hormone Sensitive Cancers
3.7. Expression of StAR mRNA in TCGA Breast Cancer Tumors and Its Association to Overall Survival
3.8. TNM Staging and Its Correlation to Breast Cancer Deaths
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Acknowledgements
Conflicts of Interest
References
- Manna, P.R.; Stocco, D.M. Regulation of the steroidogenic acute regulatory protein expression: Functional and physiological consequences. Curr. Drug Targets Immune Endocr. Metabol. Disord. 2005, 5, 93–108. [Google Scholar] [CrossRef] [PubMed]
- Manna, P.R.; Dyson, M.T.; Stocco, D.M. Regulation of the steroidogenic acute regulatory protein gene expression: Present and future perspectives. Mol. Hum. Reprod. 2009, 15, 321–333. [Google Scholar] [CrossRef]
- Papadopoulos, V.; Miller, W.L. Role of mitochondria in steroidogenesis. Best Pract. Res. Clin. Endocrinol. Metab. 2012, 26, 771–790. [Google Scholar] [PubMed]
- Manna, P.R.; Stetson, C.L.; Slominski, A.T.; Pruitt, K. Role of the steroidogenic acute regulatory protein in health and disease. Endocrine 2016, 51, 7–21. [Google Scholar] [CrossRef] [PubMed]
- Manna, P.R.; Ahmed, A.U.; Vartak, D.; Molehin, D.; Pruitt, K. Overexpression of the steroidogenic acute regulatory protein in breast cancer: Regulation by histone deacetylase inhibition. Biochem. Biophys. Res. Commun. 2019, 509, 476–482. [Google Scholar] [CrossRef] [PubMed]
- Miller, W.L.; Bose, H.S. Early steps in steroidogenesis: Intracellular cholesterol trafficking. J. Lipid Res. 2011, 52, 2111–2135. [Google Scholar] [CrossRef]
- Manna, P.R.; Dyson, M.T.; Eubank, D.W.; Clark, B.J.; Lalli, E.; Sassone-Corsi, P.; Zeleznik, A.J.; Stocco, D.M. Regulation of steroidogenesis and the steroidogenic acute regulatory protein by a member of the cAMP response-element binding protein family. Mol. Endocrinol. 2002, 16, 184–199. [Google Scholar] [CrossRef] [PubMed]
- Miller, W.L.; Auchus, R.J. The molecular biology, biochemistry, and physiology of human steroidogenesis and its disorders. Endocr. Rev. 2011, 32, 81–151. [Google Scholar] [CrossRef]
- Manna, P.R.; Cohen-Tannoudji, J.; Counis, R.; Garner, C.W.; Huhtaniemi, I.; Kraemer, F.B.; Stocco, D.M. Mechanisms of action of hormone sensitive lipase in mouse Leydig cells: Its role in the regulation of the steroidogenic acute regulatory protein. J. Biol. Chem. 2013, 288, 8505–8518. [Google Scholar] [CrossRef] [PubMed]
- Manna, P.R.; Stetson, C.L.; Daugherty, C.; Shimizu, I.; Syapin, P.J.; Garrel, G.; Cohen-Tannoudji, J.; Huhtaniemi, I.; Slominski, A.T.; Pruitt, K.; et al. Up-regulation of steroid biosynthesis by retinoid signaling: Implications for aging. Mech. Ageing. Dev. 2015, 150, 74–82. [Google Scholar] [CrossRef] [PubMed]
- Slominski, A.T.; Manna, P.R.; Tuckey, R.C. On the role of skin in the regulation of local and systemic steroidogenic activities. Steroids 2015, 103, 72–88. [Google Scholar] [CrossRef] [PubMed]
- Slominski, A.; Semak, I.; Zjawiony, J.; Wortsman, J.; Li, W.; Szczesniewski, A.; Tuckey, R.C. The cytochrome P450scc system opens an alternate pathway of vitamin D3 metabolism. FEBS J. 2005, 272, 4080–4090. [Google Scholar] [CrossRef]
- Slominski, A.T.; Li, W.; Kim, T.K.; Semak, I.; Wang, J.; Zjawiony, J.K.; Tuckey, R.C. Novel activities of CYP11A1 and their potential physiological significance. J. Steroid Biochem. Mol. Biol. 2015, 151, 25–37. [Google Scholar] [CrossRef] [PubMed]
- Manna, P.R.; Slominski, A.T.; King, S.R.; Stetson, C.L.; Stocco, D.M. Synergistic Activation of Steroidogenic Acute Regulatory Protein Expression and Steroid Biosynthesis by Retinoids: Involvement of cAMP/PKA Signaling. Endocrinology 2014, 155, 576–591. [Google Scholar] [CrossRef]
- Slominski, A.T.; Manna, P.R.; Tuckey, R.C. Cutaneous glucocorticosteroidogenesis: Securing local homeostasis and the skin integrity. Exp. Dermatol. 2014, 23, 369–374. [Google Scholar] [CrossRef]
- Manna, P.R.; Molehin, D.; Ahmed, A.U. Dysregulation of Aromatase in Breast, Endometrial, and Ovarian Cancers: An Overview of Therapeutic Strategies. Prog. Mol. Biol. Transl. Sci. 2016, 144, 487–537. [Google Scholar] [PubMed]
- Nelson, E.R.; Wardell, S.E.; Jasper, J.S.; Park, S.; Suchindran, S.; Howe, M.K.; Carver, N.J.; Pillai, R.V.; Sullivan, P.M.; Sondhi, V.; et al. 27-Hydroxycholesterol links hypercholesterolemia and breast cancer pathophysiology. Science 2013, 342, 1094–1098. [Google Scholar] [CrossRef] [PubMed]
- Silvente-Poirot, S.; Dalenc, F.; Poirot, M. The Effects of Cholesterol-Derived Oncometabolites on Nuclear Receptor Function in Cancer. Cancer Res. 2018, 78, 4803–4808. [Google Scholar] [CrossRef]
- Brodie, A.; Njar, V.; Macedo, L.F.; Vasaitis, T.S.; Sabnis, G. The Coffey Lecture: Steroidogenic enzyme inhibitors and hormone dependent cancer. Urol. Oncol. 2009, 27, 53–63. [Google Scholar] [CrossRef]
- Richie, R.C.; Swanson, J.O. Breast cancer: A review of the literature. J. Insur. Med. 2003, 35, 85–101. [Google Scholar]
- Bulun, S.E.; Lin, Z.; Zhao, H.; Lu, M.; Amin, S.; Reierstad, S.; Chen, D. Regulation of aromatase expression in breast cancer tissue. Ann. N. Y. Acad. Sci. 2009, 1155, 121–131. [Google Scholar] [CrossRef]
- Folkerd, E.; Dowsett, M. Sex hormones and breast cancer risk and prognosis. Breast 2013, 22 (Suppl. 2), S38–S43. [Google Scholar] [CrossRef]
- Cheang, M.C.; Martin, M.; Nielsen, T.O.; Prat, A.; Voduc, D.; Rodriguez-Lescure, A.; Ruiz, A.; Chia, S.; Shepherd, L.; Ruiz-Borrego, M.; et al. Defining breast cancer intrinsic subtypes by quantitative receptor expression. Oncologist 2015, 20, 474–482. [Google Scholar] [CrossRef] [PubMed]
- Prat, A.; Pineda, E.; Adamo, B.; Galvan, P.; Fernandez, A.; Gaba, L.; Diez, M.; Viladot, M.; Arance, A.; Munoz, M. Clinical implications of the intrinsic molecular subtypes of breast cancer. Breast 2015, 24 (Suppl. 2), S26–S35. [Google Scholar] [CrossRef]
- Dai, X.; Li, T.; Bai, Z.; Yang, Y.; Liu, X.; Zhan, J.; Shi, B. Breast cancer intrinsic subtype classification, clinical use and future trends. Am. J. Cancer Res. 2015, 5, 2929–2943. [Google Scholar]
- Cancer Genome Atlas Research Network. Integrated genomic analyses of ovarian carcinoma. Nature 2011, 474, 609–615. [Google Scholar] [CrossRef]
- Cancer Genome Atlas Network. Comprehensive molecular portraits of human breast tumours. Nature 2012, 490, 61–70. [Google Scholar] [CrossRef] [PubMed]
- Cancer Genome Atlas, N. Comprehensive molecular characterization of human colon and rectal cancer. Nature 2012, 487, 330–337. [Google Scholar] [CrossRef] [PubMed]
- Kandoth, C.; Schultz, N.; Cherniack, A.D.; Akbani, R.; Liu, Y.; Shen, H.; Robertson, A.G.; Pashtan, I.; Shen, R.; Benz, C.C.; et al. Integrated genomic characterization of endometrial carcinoma. Nature 2013, 497, 67–73. [Google Scholar]
- Casper, J.; Zweig, A.S.; Villarreal, C.; Tyner, C.; Speir, M.L.; Rosenbloom, K.R.; Raney, B.J.; Lee, C.M.; Lee, B.T.; Karolchik, D.; et al. The UCSC Genome Browser database: 2018 update. Nucleic Acids Res. 2018, 46, D762–D769. [Google Scholar] [PubMed]
- Cerami, E.; Gao, J.; Dogrusoz, U.; Gross, B.E.; Sumer, S.O.; Aksoy, B.A.; Jacobsen, A.; Byrne, C.J.; Heuer, M.L.; Larsson, E.; et al. The cBio cancer genomics portal: An open platform for exploring multidimensional cancer genomics data. Cancer Discov. 2012, 2, 401–404. [Google Scholar] [CrossRef] [PubMed]
- Gao, J.; Aksoy, B.A.; Dogrusoz, U.; Dresdner, G.; Gross, B.; Sumer, S.O.; Sun, Y.; Jacobsen, A.; Sinha, R.; Larsson, E.; et al. Integrative analysis of complex cancer genomics and clinical profiles using the cBioPortal. Sci. Signal. 2013, 6, pl1. [Google Scholar] [CrossRef]
- Li, B.; Dewey, C.N. RSEM: Accurate transcript quantification from RNA-Seq data with or without a reference genome. BMC Bioinform. 2011, 12, 323. [Google Scholar] [CrossRef] [PubMed]
- Pereira, B.; Chin, S.F.; Rueda, O.M.; Vollan, H.K.; Provenzano, E.; Bardwell, H.A.; Pugh, M.; Jones, L.; Russell, R.; Sammut, S.J. The somatic mutation profiles of 2,433 breast cancers refines their genomic and transcriptomic landscapes. Nat. Commun. 2016, 7, 11479. [Google Scholar] [CrossRef]
- Eirew, P.; Steif, A.; Khattra, J.; Ha, G.; Yap, D.; Farahani, H.; Gelmon, K.; Chia, S.; Mar, C.; Wan, A.; et al. Dynamics of genomic clones in breast cancer patient xenografts at single-cell resolution. Nature 2015, 518, 422–426. [Google Scholar] [CrossRef] [PubMed]
- Ciriello, G.; Gatza, M.L.; Beck, A.H.; Wilkerson, M.D.; Rhie, S.K.; Pastore, A.; Zhang, H.; McLellan, M.; Yau, C.; Kandoth, C.; et al. Comprehensive Molecular Portraits of Invasive Lobular Breast Cancer. Cell 2015, 163, 506–519. [Google Scholar] [CrossRef] [PubMed]
- Lefebvre, C.; Bachelot, T.; Filleron, T.; Pedrero, M.; Campone, M.; Soria, J.C.; Massard, C.; Levy, C.; Arnedos, M.; Lacroix-Triki, M.; et al. Mutational Profile of Metastatic Breast Cancers: A Retrospective Analysis. PLoS Med. 2016, 13, e1002201. [Google Scholar] [CrossRef]
- Bullard, J.H.; Purdom, E.; Hansen, K.D.; Dudoit, S. Evaluation of statistical methods for normalization and differential expression in mRNA-Seq experiments. BMC Bioinform. 2010, 11, 94. [Google Scholar] [CrossRef]
- O’Sullivan, B.; Brierley, J.; Byrd, D.; Bosman, F.; Kehoe, S.; Kossary, C.; Pineros, M.; Van Eycken, E.; Weir, H.K.; Gospodarowicz, M. The TNM classification of malignant tumours-towards common understanding and reasonable expectations. Lancet Oncol. 2017, 18, 849–851. [Google Scholar] [CrossRef]
- Lydiatt, W.M.; Patel, S.G.; O’Sullivan, B.; Brandwein, M.S.; Ridge, J.A.; Migliacci, J.C.; Loomis, A.M.; Shah, J.P. Head and Neck cancers-major changes in the American Joint Committee on cancer eighth edition cancer staging manual. CA Cancer J. Clin. 2017, 67, 122–137. [Google Scholar] [CrossRef]
- Sedgwick, P. How to read a Kaplan-Meier survival plot. BMJ 2014, 349, g5608. [Google Scholar] [CrossRef] [PubMed]
- Slamon, D.J.; Clark, G.M.; Wong, S.G.; Levin, W.J.; Ullrich, A.; McGuire, W.L. Human breast cancer: Correlation of relapse and survival with amplification of the HER-2/neu oncogene. Science 1987, 235, 177–182. [Google Scholar] [CrossRef]
- Schneiderman, J.; London, W.B.; Brodeur, G.M.; Castleberry, R.P.; Look, A.T.; Cohn, S.L. Clinical significance of MYCN amplification and ploidy in favorable-stage neuroblastoma: A report from the Children’s Oncology Group. J. Clin. Oncol. 2008, 26, 913–918. [Google Scholar] [CrossRef] [PubMed]
- Liu, L.; Kimball, S.; Liu, H.; Holowatyj, A.; Yang, Z.Q. Genetic alterations of histone lysine methyltransferases and their significance in breast cancer. Oncotarget 2015, 6, 2466–2482. [Google Scholar] [CrossRef] [PubMed]
- Hilborn, E.; Stal, O.; Jansson, A. Estrogen and androgen-converting enzymes 17beta-hydroxysteroid dehydrogenase and their involvement in cancer: With a special focus on 17beta-hydroxysteroid dehydrogenase type 1, 2, and breast cancer. Oncotarget 2017, 8, 30552–30562. [Google Scholar] [CrossRef] [PubMed]
- Giovannetti, E.; Wang, Q.; Avan, A.; Funel, N.; Lagerweij, T.; Lee, J.H.; Caretti, V.; van der Velde, A.; Boggi, U.; Wang, Y.; et al. Role of CYB5A in pancreatic cancer prognosis and autophagy modulation. J. Natl. Cancer Inst. 2014, 106, djt346. [Google Scholar] [CrossRef] [PubMed]
- Yoshihara, K.; Shahmoradgoli, M.; Martinez, E.; Vegesna, R.; Kim, H.; Torres-Garcia, W.; Trevino, V.; Shen, H.; Laird, P.W.; Levine, D.A.; et al. Inferring tumour purity and stromal and immune cell admixture from expression data. Nat. Commun. 2013, 4, 2612. [Google Scholar] [CrossRef]
- Geng, X.; Liu, Y.; Diersch, S.; Kotzsch, M.; Grill, S.; Weichert, W.; Kiechle, M.; Magdolen, V.; Dorn, J. Clinical relevance of kallikrein-related peptidase 9, 10, 11, and 15 mRNA expression in advanced high-grade serous ovarian cancer. PLoS ONE 2017, 12, e0186847. [Google Scholar] [CrossRef]
- Slominski, A.T.; Zmijewski, M.A.; Semak, I.; Zbytek, B.; Pisarchik, A.; Li, W.; Zjawiony, J.; Tuckey, R.C. Cytochromes p450 and skin cancer: Role of local endocrine pathways. Anticancer Agents Med. Chem. 2014, 14, 77–96. [Google Scholar] [CrossRef]
- Slominski, A.; Gomez-Sanchez, C.E.; Foecking, M.F.; Wortsman, J. Metabolism of progesterone to DOC, corticosterone and 18OHDOC in cultured human melanoma cells. FEBS Lett. 1999, 455, 364–366. [Google Scholar] [CrossRef]
- Camats, N.; Pandey, A.V.; Fernandez-Cancio, M.; Fernandez, J.M.; Ortega, A.M.; Udhane, S.; Andaluz, P.; Audi, L.; Fluck, C.E. STAR splicing mutations cause the severe phenotype of lipoid congenital adrenal hyperplasia: Insights from a novel splice mutation and review of reported cases. Clin. Endocrinol. 2014, 80, 191–199. [Google Scholar] [CrossRef]
- Miller, W.L. Steroidogenic acute regulatory protein (StAR), a novel mitochondrial cholesterol transporter. Biochim. Biophys. Acta 2007, 1771, 663–676. [Google Scholar] [CrossRef]
- Papadopoulos, V.; Aghazadeh, Y.; Fan, J.; Campioli, E.; Zirkin, B.; Midzak, A. Translocator protein-mediated pharmacology of cholesterol transport and steroidogenesis. Mol. Cell. Endocrinol. 2015, 408, 90–98. [Google Scholar] [CrossRef]
- Fan, J.; Wang, K.; Zirkin, B.; Papadopoulos, V. CRISPR/Cas9Mediated Tspo Gene Mutations Lead to Reduced Mitochondrial Membrane Potential and Steroid Formation in MA-10 Mouse Tumor Leydig Cells. Endocrinology 2018, 159, 1130–1146. [Google Scholar] [CrossRef]
- Strauss, J.F., 3rd; Kishida, T.; Christenson, LK.; Fujimoto, T.; Hiroi, H. START domain proteins and the intracellular trafficking of cholesterol in steroidogenic cells. Mol. Cell. Endocrinol. 2003, 202, 59–65. [Google Scholar] [CrossRef]
- Alpy, F.; Legueux, F.; Bianchetti, L.; Tomasetto, C. START domain-containing proteins: A review of their role in lipid transport and exchange. Med. Sci. Paris 2009, 25, 181–191. [Google Scholar]
- Tomasetto, C.; Regnier, C.; Moog-Lutz, C.; Mattei, M.G.; Chenard, M.P.; Lidereau, R.; Basset, P.; Rio, M.C. Identification of four novel human genes amplified and overexpressed in breast carcinoma and localized to the q11-q21.3 region of chromosome 17. Genomics 1995, 28, 367–376. [Google Scholar] [CrossRef]
- Akiyama, N.; Sasaki, H.; Ishizuka, T.; Kishi, T.; Sakamoto, H.; Onda, M.; Hirai, H.; Yazaki, Y.; Sugimura, T.; Terada, M. Isolation of a candidate gene, CAB1, for cholesterol transport to mitochondria from the c-ERBB-2 amplicon by a modified cDNA selection method. Cancer Res. 1997, 57, 3548–3553. [Google Scholar]
- Alpy, F.; Boulay, A.; Moog-Lutz, C.; Andarawewa, K.L.; Degot, S.; Stoll, I.; Rio, M.C.; Tomasetto, C. Metastatic lymph node 64 (MLN64), a gene overexpressed in breast cancers, is regulated by Sp/KLF transcription factors. Oncogene 2003, 22, 3770–3780. [Google Scholar] [CrossRef]
- Vassilev, B.; Sihto, H.; Li, S.; Holtta-Vuori, M.; Ilola, J.; Lundin, J.; Isola, J.; Kellokumpu-Lehtinen, P.L.; Joensuu, H.; Ikonen, E. Elevated levels of StAR-related lipid transfer protein 3 alter cholesterol balance and adhesiveness of breast cancer cells: Potential mechanisms contributing to progression of HER2-positive breast cancers. Am. J. Pathol. 2015, 185, 987–1000. [Google Scholar] [CrossRef]
- Llaverias, G.; Danilo, C.; Mercier, I.; Daumer, K.; Capozza, F.; Williams, T.M.; Sotgia, F.; Lisanti, M.P.; Frank, P.G. Role of cholesterol in the development and progression of breast cancer. Am. J. Pathol. 2011, 178, 402–412. [Google Scholar] [CrossRef]
- Voisin, M.; de Medina, P.; Mallinger, A.; Dalenc, F.; Huc-Claustre, E.; Leignadier, J.; Serhan, N.; Soules, R.; Segala, G.; Mougel, A.; et al. Identification of a tumor-promoter cholesterol metabolite in human breast cancers acting through the glucocorticoid receptor. Proc. Natl. Acad. Sci. USA 2017, 114, E9346–E9355. [Google Scholar] [CrossRef] [PubMed]
- Simpson, E.; Santen, R.J. Celebrating 75 years of oestradiol. J. Mol. Endocrinol. 2015, 55, T1–T20. [Google Scholar] [CrossRef]
- Zhao, H.; Zhou, L.; Shangguan, A.J.; Bulun, S.E. Aromatase expression and regulation in breast and endometrial cancer. J. Mol. Endocrinol. 2016, 57, 19–33. [Google Scholar] [CrossRef]
- Manna, P.R.; Chandrala, S.P.; King, S.R.; Jo, Y.; Counis, R.; Huhtaniemi, I.T.; Stocco, D.M. Molecular mechanisms of insulin-like growth factor-I mediated regulation of the steroidogenic acute regulatory protein in mouse leydig cells. Mol. Endocrinol. 2006, 20, 362–378. [Google Scholar] [CrossRef]
- Manna, P.R.; Soh, J.W.; Stocco, D.M. The involvement of specific PKC isoenzymes in phorbol ester-mediated regulation of steroidogenic acute regulatory protein expression and steroid synthesis in mouse Leydig cells. Endocrinology 2011, 152, 313–325. [Google Scholar] [CrossRef]
- Arakane, F.; King, S.R.; Du, Y.; Kallen, C.B.; Walsh, L.P.; Watari, H.; Stocco, D.M.; Strauss, J.F., 3rd. Phosphorylation of steroidogenic acute regulatory protein (StAR) modulates its steroidogenic activity. J. Biol. Chem. 1997, 272, 32656–32662. [Google Scholar] [CrossRef]
- Clark, B.J.; Ranganathan, V.; Combs, R. Steroidogenic acute regulatory protein expression is dependent upon post-translational effects of cAMP-dependent protein kinase A. Mol. Cell. Endocrinol. 2001, 173, 183–192. [Google Scholar] [CrossRef]
- Castro-Piedras, I.; Sharma, M.; den Bakker, M.; Molehin, D.; Martinez, E.G.; Vartak, D.; Pruitt, W.M.; Deitrick, J.; Almodovar, S.; Pruitt, K. DVL1 and DVL3 differentially localize to CYP19A1 promoters and regulate aromatase mRNA in breast cancer cells. Oncotarget 2018, 9, 35639–35654. [Google Scholar] [CrossRef]
- Simpson, E.R.; Misso, M.; Hewitt, K.N.; Hill, R.A.; Boon, W.C.; Jones, M.E.; Kovacic, A.; Zhou, J.; Clyne, C.D. Estrogen--the good, the bad, and the unexpected. Endocr. Rev. 2005, 26, 322–330. [Google Scholar] [CrossRef] [PubMed]
- Molehin, D.; Castro-Piedras, I.; Sharma, M.; Sennoune, S.R.; Arena, D.; Manna, P.R.; Pruitt, K. Aromatase Acetylation Patterns and Altered Activity in Response to Sirtuin Inhibition. Mol. Cancer Res. 2018, 16, 1530–1542. [Google Scholar] [CrossRef] [PubMed]
- Bulun, S.E.; Lin, Z.; Imir, G.; Amin, S.; Demura, M.; Yilmaz, B.; Martin, R.; Utsunomiya, H.; Thung, S.; Gurates, B.; et al. Regulation of aromatase expression in estrogen-responsive breast and uterine disease: From bench to treatment. Pharmacol. Rev. 2005, 57, 359–383. [Google Scholar] [CrossRef] [PubMed]
| CNAs | Breast N (%) | Colorectal N (%) | Melanoma N (%) | Ovarian N (%) | Pancreatic N (%) | Prostate N (%) | Uterine Endometrial N (%) |
|---|---|---|---|---|---|---|---|
| Homozygous Deletion | 16 (1.48) | 8 (1.30) | 3 (0.82) | 7 (1.21) | 0 (0.00) | 32 (6.50) | 7 (1.30) |
| Hemizygous Deletion | 252 (23.33) | 140 (22.73) | 69 (18.80) | 215 (37.13) | 42 (22.83) | 133 (27.03) | 66 (12.24) |
| Diploid | 406 (37.59) | 271 (43.99) | 184 (50.14) | 220 (38.00) | 109 (59.24) | 258 (52.44) | 345 (64.01) |
| Gain | 268 (24.81) | 182 (29.55) | 110 (29.97) | 117 (20.21) | 25 (13.59) | 55 (11.18) | 111 (20.59) |
| High Level Amplification | 138 (12.78) | 15 (2.44) | 1 (0.27) | 20 (3.45) | 8 (4.35) | 14 (2.85) | 10 (1.86) |
| Total Number of Tumors | 1080 | 616 | 367 | 579 | 184 | 492 | 539 |
| TNM Staging | Low Expression (<50%) | High Expression (>50%) | Low Expression (<25%) | High Expression (>25%) | ||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Tumor Nos. | Death Nos. | % Deaths | Tumor Nos. | Death Nos. | % Deaths | Tumor Nos. | Death Nos. | % Deaths | Tumor Nos. | Death Nos. | % Deaths | |
| T1 | 120 | 17 | 14.2 | 158 | 16 | 10.1 | 54 | 8 | 14.8 | 224 | 25 | 11.2 |
| T2 | 336 | 44 | 13.1 | 294 | 33 | 11.2 | 169 | 25 | 14.8 | 461 | 52 | 11.3 |
| T3 | 63 | 11 | 17.5 | 75 | 14 | 18.7 | 32 | 9 | 28.1 | 106 | 16 | 15.1 |
| T4 | 26 | 8 | 30.8 | 14 | 7 | 50 | 16 | 5 | 31.3 | 24 | 10 | 41.7 |
| N0 | 257 | 22 | 8.6 | 257 | 22 | 8.6 | 120 | 15 | 12.5 | 394 | 29 | 7.4 |
| N1 | 169 | 31 | 18.3 | 190 | 28 | 14.7 | 91 | 19 | 20.1 | 268 | 40 | 14.9 |
| N2 | 75 | 16 | 21.3 | 45 | 6 | 13.3 | 38 | 6 | 15.8 | 82 | 16 | 19.5 |
| N3 | 33 | 5 | 15.2 | 45 | 12 | 26.7 | 16 | 4 | 25.0 | 60 | 11 | 18.3 |
| M0 | 459 | 62 | 13.5 | 621 | 32 | 5.2 | 224 | 26 | 11.6 | 672 | 83 | 12.4 |
| M1 | 13 | 10 | 76.9 | 8 | 5 | 62.5 | 4 | 2 | 50 | 12 | 9 | 75.0 |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Manna, P.R.; Ahmed, A.U.; Yang, S.; Narasimhan, M.; Cohen-Tannoudji, J.; Slominski, A.T.; Pruitt, K. Genomic Profiling of the Steroidogenic Acute Regulatory Protein in Breast Cancer: In Silico Assessments and a Mechanistic Perspective. Cancers 2019, 11, 623. https://doi.org/10.3390/cancers11050623
Manna PR, Ahmed AU, Yang S, Narasimhan M, Cohen-Tannoudji J, Slominski AT, Pruitt K. Genomic Profiling of the Steroidogenic Acute Regulatory Protein in Breast Cancer: In Silico Assessments and a Mechanistic Perspective. Cancers. 2019; 11(5):623. https://doi.org/10.3390/cancers11050623
Chicago/Turabian StyleManna, Pulak R., Ahsen U. Ahmed, Shengping Yang, Madhusudhanan Narasimhan, Joëlle Cohen-Tannoudji, Andrzej T. Slominski, and Kevin Pruitt. 2019. "Genomic Profiling of the Steroidogenic Acute Regulatory Protein in Breast Cancer: In Silico Assessments and a Mechanistic Perspective" Cancers 11, no. 5: 623. https://doi.org/10.3390/cancers11050623
APA StyleManna, P. R., Ahmed, A. U., Yang, S., Narasimhan, M., Cohen-Tannoudji, J., Slominski, A. T., & Pruitt, K. (2019). Genomic Profiling of the Steroidogenic Acute Regulatory Protein in Breast Cancer: In Silico Assessments and a Mechanistic Perspective. Cancers, 11(5), 623. https://doi.org/10.3390/cancers11050623







