The Function of Non-Coding RNAs in Lung Cancer Tumorigenesis
Abstract
1. Introduction
2. Transfer RNA
3. Circular RNAs
4. Small Nucleolar RNA (snoRNAs)
5. PIWI-Interacting RNAs
6. YRNA
7. Natural Antisense Transcripts
8. Pseudogene Transcript
9. Transcribed Ultraconserved Region (T-UCR)
10. Telomerase RNA Component
11. Conclusions and Perspectives
Author Contributions
Funding
Conflicts of Interest
Abbreviations
| NSCLC | non-small cell lung cancer |
| ncRNA | non-coding RNA |
| nt | nucleotide |
| bp | base pairs |
| TF | transcription factor |
| tRNA | transfer RNA |
| rRNA | ribosomal RNA |
| miRNA/miR | microRNA |
| siRNA | small interfering RNA |
| piRNA | piwi-interacting RNA |
| piR | piwi-interacting RNA |
| snoRNA | small nucleolar RNA |
| circRNA | circular RNA |
| tRF | tRNA-derived fragments |
| half-tRNA | tiRNA |
| ceRNA | competing endogenous RNA |
| NAT | natural antisense transcripts |
| T-UCR | transcribed ultraconserved region |
| dsRNA | double-stranded RNA |
| shRNA | short hairpin RNA |
| TERC | telomerase associated RNA |
| NAT | natural antisense transcripts |
| p-ERM | phosphorylated ERM proteins |
| NXN | nucleoredoxin |
| AHRR | Aryl-Hydrocarbon Receptor Repressor |
| CDC42EP3 | CDC42 Effector Protein 3 |
| EDC3 | Enhancer of MRNA Decapping 3 |
| HIPK2 | Homeodomain Interacting Protein Kinase 2 |
| CXCL5 | C-X-C Motif Chemokine Ligand 5 |
| SEC62 | SEC62 Homolog: Preprotein Translocation Factor |
| ZNF703 | Zinc Finger Protein 703 |
| CTNNB | Catenin Beta 1 |
| BRD | bromodomain |
| BET | extra-terminal |
| PTEN | Phosphatase and Tensin Homolog |
| TNC | Tenascin C |
| TERC | telomerase RNA component |
| BAX | BCL2 Associated X: Apoptosis Regulator |
| CCND2 | Cyclin D |
| TIAM1 | T-lymphoma invasion and metastasis-inducing protein 1 |
| NFAT5 | Nuclear factor of activated T-cells 5 |
| CO1A1 | Collagen alpha-1(I) chain |
| TRAF3 | TNF Receptor Associated Factor 3 |
| MEF2C | Myocyte Enhancer Factor 2C |
| RPCK1 | Rho-associated protein kinase 1 |
| FOXR2 | Forkhead Box R2 |
| SphK1 | Sphingosine kinase 1 |
| STAT3 | Signal transducer and activator of transcription 3 |
| CDK4 | Cyclin-dependent kinase 4 |
| NKX2-1 | NK2 homeobox 1 |
| DUXAP8 | Double Homeobox A Pseudogene 8 |
| EGR1 | Early growth response protein 1 |
| RHOB | Rho-related GTP-binding protein RhoB |
| LSD1 | Histone demethylase Lysine specific demethylase1 |
| LATS2 | Large tumour suppressor 2 |
| RRAD | Ras-related associated with diabetes |
| CHIAP2 | Chitinase: Acidic Pseudogene 2 |
| RPL13AP17 | Ribosomal Protein L13a Pseudogene 17 |
| p-ERM | phosphorylated ERM proteins |
| EMT | epithelial to mesenchymal transition |
| Ect2 | Epithelial Cell Transforming Sequence 2 |
| EV | extracellular vesicles |
References
- Bray, F.; Ferlay, J.; Soerjomataram, I.; Siegel, R.L.; Torre, L.A.; Jemal, A. Global cancer statistics 2018: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA Cancer J. Clin. 2018, 68, 394–424. [Google Scholar] [CrossRef] [PubMed]
- Alberg, A.J.; Brock, M.V.; Samet, J.M. Epidemiology of lung cancer: Looking to the future. J. Clin. Oncol. Off. J. Am. Soc. Clin. Oncol. 2005, 23, 3175–3185. [Google Scholar] [CrossRef]
- Crick, F. Central dogma of molecular biology. Nature 1970, 227, 561–563. [Google Scholar] [CrossRef] [PubMed]
- Ling, H.; Girnita, L.; Buda, O.; Calin, G.A. Non-coding RNAs: The cancer genome dark matter that matters! Clin. Chem. Lab. Med. 2017, 55, 705–714. [Google Scholar] [CrossRef] [PubMed]
- Hubé, F.; Francastel, C. Coding and Non-coding RNAs, the Frontier Has Never Been So Blurred. Front. Genet. 2018, 9, 140. [Google Scholar] [CrossRef] [PubMed]
- Brosnan, C.A.; Voinnet, O. The long and the short of noncoding RNAs. Curr. Opin. Cell Biol. 2009, 21, 416–425. [Google Scholar] [CrossRef]
- Wu, H.; Yang, L.; Chen, L.L. The Diversity of Long Noncoding RNAs and Their Generation. Trends Genet 2017, 33, 540–552. [Google Scholar] [CrossRef] [PubMed]
- Uszczynska-Ratajczak, B.; Lagarde, J.; Frankish, A.; Guigo, R.; Johnson, R. Towards a complete map of the human long non-coding RNA transcriptome. Nat. Rev. Genet. 2018, 19, 535–548. [Google Scholar] [CrossRef] [PubMed]
- Chen, B.; Huang, S. Circular RNA: An emerging non-coding RNA as a regulator and biomarker in cancer. Cancer Lett. 2018, 418, 41–50. [Google Scholar] [CrossRef]
- Groen, J.N.; Capraro, D.; Morris, K.V. The emerging role of pseudogene expressed non-coding RNAs in cellular functions. Int. J. Biochem. Cell Biol. 2014, 54, 350–355. [Google Scholar] [CrossRef]
- Das, A.; Gorospe, M.; Panda, A.C. The coding potential of circRNAs. Aging 2018, 10, 2228–2229. [Google Scholar] [CrossRef]
- Xu, J.; Zhang, J. Are Human Translated Pseudogenes Functional? Mol. Biol. Evol. 2016, 33, 755–760. [Google Scholar] [CrossRef]
- Sonea, L.; Buse, M.; Gulei, D.; Onaciu, A.; Simon, I.; Braicu, C.; Berindan-Neagoe, I. Decoding the Emerging Patterns Exhibited in Non-coding RNAs Characteristic of Lung Cancer with Regard to their Clinical Significance. Current Genom. 2018, 19, 258–278. [Google Scholar] [CrossRef]
- Redis, R.S.; Berindan-Neagoe, I.; Pop, V.I.; Calin, G.A. Non-coding RNAs as theranostics in human cancers. J. Cellular Biochem. 2012, 113, 1451–1459. [Google Scholar]
- Starega-Roslan, J.; Krol, J.; Koscianska, E.; Kozlowski, P.; Szlachcic, W.J.; Sobczak, K.; Krzyzosiak, W.J. Structural basis of microRNA length variety. Nucleic Acids Res. 2011, 39, 257–268. [Google Scholar] [CrossRef]
- Macfarlane, L.A.; Murphy, P.R. MicroRNA: Biogenesis, Function and Role in Cancer. Curr. Genom. 2010, 11, 537–561. [Google Scholar] [CrossRef]
- Dana, H.; Chalbatani, G.M.; Mahmoodzadeh, H.; Karimloo, R.; Rezaiean, O.; Moradzadeh, A.; Mehmandoost, N.; Moazzen, F.; Mazraeh, A.; Marmari, V.; et al. Molecular Mechanisms and Biological Functions of siRNA. Int. J. Biomed. Sci. Ijbs 2017, 13, 48–57. [Google Scholar]
- Kim, V.N.; Han, J.; Siomi, M.C. Biogenesis of small RNAs in animals. Nat. Rev. Mol. Cell Biol. 2009, 10, 126–139. [Google Scholar] [CrossRef]
- Iwasaki, Y.W.; Siomi, M.C.; Siomi, H. PIWI-Interacting RNA: Its Biogenesis and Functions. Annu. Rev. Biochem. 2015, 84, 405–433. [Google Scholar] [CrossRef]
- Ozata, D.M.; Gainetdinov, I.; Zoch, A.; O’Carroll, D.; Zamore, P.D. PIWI-interacting RNAs: Small RNAs with big functions. Nat. Rev. Genet. 2019, 20, 89–108. [Google Scholar] [CrossRef]
- Lee, Y.S.; Shibata, Y.; Malhotra, A.; Dutta, A. A novel class of small RNAs: tRNA-derived RNA fragments (tRFs). Genes Dev. 2009, 23, 2639–2649. [Google Scholar] [CrossRef]
- Martinez, G.; Choudury, S.G.; Slotkin, R.K. tRNA-derived small RNAs target transposable element transcripts. Nucleic Acids Res. 2017, 45, 5142–5152. [Google Scholar] [CrossRef] [PubMed]
- Saikia, M.; Hatzoglou, M. The Many Virtues of tRNA-derived Stress-induced RNAs (tiRNAs): Discovering Novel Mechanisms of Stress Response and Effect on Human Health. J. Biol. Chem. 2015, 290, 29761–29768. [Google Scholar] [CrossRef]
- Dhahbi, J.M.; Spindler, S.R.; Atamna, H.; Boffelli, D.; Mote, P.; Martin, D.I. 5’-YRNA fragments derived by processing of transcripts from specific YRNA genes and pseudogenes are abundant in human serum and plasma. Physiol. Genom. 2013, 45, 990–998. [Google Scholar] [CrossRef] [PubMed]
- Kowalski, M.P.; Krude, T. Functional roles of non-coding Y RNAs. Int. J. Biochem. Cell Biol. 2015, 66, 20–29. [Google Scholar] [CrossRef] [PubMed]
- Stein, A.J.; Fuchs, G.; Fu, C.; Wolin, S.L.; Reinisch, K.M. Structural insights into RNA quality control: The Ro autoantigen binds misfolded RNAs via its central cavity. Cell 2005, 121, 529–539. [Google Scholar] [CrossRef]
- Scott, M.S.; Ono, M. From snoRNA to miRNA: Dual function regulatory non-coding RNAs. Biochimie 2011, 93, 1987–1992. [Google Scholar] [CrossRef]
- Dupuis-Sandoval, F.; Poirier, M.; Scott, M.S. The emerging landscape of small nucleolar RNAs in cell biology. Wiley Interdiscip. Rev. RNA 2015, 6, 381–397. [Google Scholar] [CrossRef] [PubMed]
- Rubtsova, M.P.; Vasilkova, D.P.; Naraykina, Y.V.; Dontsova, O.A. Peculiarities of Yeasts and Human Telomerase RNAs Processing. Acta Nat. 2016, 8, 14–22. [Google Scholar]
- Tseng, C.K.; Wang, H.F.; Burns, A.M.; Schroeder, M.R.; Gaspari, M.; Baumann, P. Human Telomerase RNA Processing and Quality Control. Cell Rep. 2015, 13, 2232–2243. [Google Scholar] [CrossRef]
- Rao, X.; Huang, D.; Sui, X.; Liu, G.; Song, X.; Xie, J.; Huang, D. Overexpression of WRAP53 is associated with development and progression of esophageal squamous cell carcinoma. PLoS ONE 2014, 9, e91670. [Google Scholar] [CrossRef]
- Piatek, M.J.; Henderson, V.; Zynad, H.S.; Werner, A. Natural antisense transcription from a comparative perspective. Genomics 2016, 108, 56–63. [Google Scholar] [CrossRef][Green Version]
- Wight, M.; Werner, A. The functions of natural antisense transcripts. Essays Biochem. 2013, 54, 91–101. [Google Scholar] [CrossRef]
- Terracciano, D.; Terreri, S.; de Nigris, F.; Costa, V.; Calin, G.A.; Cimmino, A. The role of a new class of long noncoding RNAs transcribed from ultraconserved regions in cancer. Biochim. Et Biophys. Acta. Rev. Cancer 2017, 1868, 449–455. [Google Scholar] [CrossRef]
- Memczak, S.; Papavasileiou, P.; Peters, O.; Rajewsky, N. Identification and Characterization of Circular RNAs As a New Class of Putative Biomarkers in Human Blood. PLoS ONE 2015, 10, e0141214. [Google Scholar] [CrossRef]
- Zhang, Y.; Liang, W.; Zhang, P.; Chen, J.; Qian, H.; Zhang, X.; Xu, W. Circular RNAs: Emerging cancer biomarkers and targets. J. Exp. Clin. Cancer Res. Cr 2017, 36, 152. [Google Scholar] [CrossRef] [PubMed]
- Braicu, C.; Zimta, A.A.; Gulei, D.; Olariu, A.; Berindan-Neagoe, I. Comprehensive analysis of circular RNAs in pathological states: Biogenesis, cellular regulation, and therapeutic relevance. Cell. Mol. Life Sci. Cmls 2019. [Google Scholar] [CrossRef] [PubMed]
- Jingsi, T.; Mingyao, Y.; Ying, L. Functional roles of pseudogenes in cancers. Yi Chuan Hered. 2015, 37, 8–16. [Google Scholar] [CrossRef]
- Xiao-Jie, L.; Ai-Mei, G.; Li-Juan, J.; Jiang, X. Pseudogene in cancer: Real functions and promising signature. J. Med. Genet. 2015, 52, 17–24. [Google Scholar] [CrossRef]
- Huang, S.Q.; Sun, B.; Xiong, Z.P.; Shu, Y.; Zhou, H.H.; Zhang, W.; Xiong, J.; Li, Q. The dysregulation of tRNAs and tRNA derivatives in cancer. J. Exp. Clin. Cancer Res. Cr 2018, 37, 101. [Google Scholar] [CrossRef]
- Pekarsky, Y.; Balatti, V.; Palamarchuk, A.; Rizzotto, L.; Veneziano, D.; Nigita, G.; Rassenti, L.Z.; Pass, H.I.; Kipps, T.J.; Liu, C.G.; et al. Dysregulation of a family of short noncoding RNAs, tsRNAs, in human cancer. Proc. Natl. Acad. Sci. USA 2016, 113, 5071–5076. [Google Scholar] [CrossRef] [PubMed]
- Balatti, V.; Nigita, G.; Veneziano, D.; Drusco, A.; Stein, G.S.; Messier, T.L.; Farina, N.H.; Lian, J.B.; Tomasello, L.; Liu, C.G.; et al. tsRNA signatures in cancer. Proc. Natl. Acad. Sci. USA 2017, 114, 8071–8076. [Google Scholar] [CrossRef] [PubMed]
- Shao, Y.; Sun, Q.; Liu, X.; Wang, P.; Wu, R.; Ma, Z. tRF-Leu-CAG promotes cell proliferation and cell cycle in non-small cell lung cancer. Chem. Biol. Drug Des. 2017, 90, 730–738. [Google Scholar] [CrossRef]
- Liu, X.; Li, Z.; Song, Y.; Wang, R.; Han, L.; Wang, Q.; Jiang, K.; Kang, C.; Zhang, Q. AURKA induces EMT by regulating histone modification through Wnt/beta-catenin and PI3K/Akt signaling pathway in gastric cancer. Oncotarget 2016, 7, 33152–33164. [Google Scholar] [CrossRef] [PubMed]
- Han, W.; Wang, L.; Zhang, L.; Wang, Y.; Li, Y. Circular RNA circ-RAD23B promotes cell growth and invasion by miR-593-3p/CCND2 and miR-653-5p/TIAM1 pathways in non-small cell lung cancer. Biochem. Biophys. Res. Commun. 2019, 510, 462–466. [Google Scholar] [CrossRef] [PubMed]
- Chen, L.; Nan, A.; Zhang, N.; Jia, Y.; Li, X.; Ling, Y.; Dai, J.; Zhang, S.; Yang, Q.; Yi, Y.; et al. Circular RNA 100146 functions as an oncogene through direct binding to miR-361-3p and miR-615-5p in non-small cell lung cancer. Mol. Cancer 2019, 18, 13. [Google Scholar] [CrossRef]
- Qin, S.; Zhao, Y.; Lim, G.; Lin, H.; Zhang, X.; Zhang, X. Circular RNA PVT1 acts as a competing endogenous RNA for miR-497 in promoting non-small cell lung cancer progression. Biomed. Pharmacother. Biomed. Pharmacother. 2019, 111, 244–250. [Google Scholar] [CrossRef] [PubMed]
- Zhu, X.; Wang, X.; Wei, S.; Chen, Y.; Chen, Y.; Fan, X.; Han, S.; Wu, G. hsa_circ_0013958: A circular RNA and potential novel biomarker for lung adenocarcinoma. Febs J. 2017, 284, 2170–2182. [Google Scholar] [CrossRef] [PubMed]
- Li, W.; Jiang, W.; Liu, T.; Lv, J.; Guan, J. Enhanced expression of circ_0000735 forecasts clinical severity in NSCLC and promotes cell progression via sponging miR-1179 and miR-1182. Biochem. Biophys. Res. Commun. 2019, 510, 467–471. [Google Scholar] [CrossRef] [PubMed]
- Li, X.; Zhang, Z.; Jiang, H.; Li, Q.; Wang, R.; Pan, H.; Niu, Y.; Liu, F.; Gu, H.; Fan, X.; et al. Circular RNA circPVT1 Promotes Proliferation and Invasion Through Sponging miR-125b and Activating E2F2 Signaling in Non-Small Cell Lung Cancer. Cell. Physiol. Biochem. 2018, 51, 2324–2340. [Google Scholar] [CrossRef] [PubMed]
- Qiu, B.Q.; Zhang, P.F.; Xiong, D.; Xu, J.J.; Long, X.; Zhu, S.Q.; Ye, X.D.; Wu, Y.; Pei, X.; Zhang, X.M.; et al. CircRNA fibroblast growth factor receptor 3 promotes tumor progression in non-small cell lung cancer by regulating Galectin-1-AKT/ERK1/2 signaling. J. Cell. Physiol. 2018. [Google Scholar] [CrossRef] [PubMed]
- Zhou, Y.; Zheng, X.; Xu, B.; Chen, L.; Wang, Q.; Deng, H.; Jiang, J. Circular RNA hsa_circ_0004015 regulates the proliferation, invasion, and TKI drug resistance of non-small cell lung cancer by miR-1183/PDPK1 signaling pathway. Biochem. Biophys. Res. Commun. 2019, 508, 527–535. [Google Scholar] [CrossRef] [PubMed]
- Chen, J.; Xu, S.; Chen, S.; Zong, Z.; Han, X.; Zhao, Y.; Shang, H. CircPUM1 promotes the malignant behavior of lung adenocarcinoma by regulating miR-326. Biochem. Biophys. Res. Commun. 2019, 508, 844–849. [Google Scholar] [CrossRef] [PubMed]
- Li, L.; Li, W.; Chen, N.; Zhao, H.; Xu, G.; Zhao, Y.; Pan, X.; Zhang, X.; Zhou, L.; Yu, D.; et al. FLI1 Exonic Circular RNAs as a Novel Oncogenic Driver to Promote Tumor Metastasis in Small Cell Lung Cancer. Clin. Cancer Res. Off. J. Am. Assoc. Cancer Res. 2019, 25, 1302–1317. [Google Scholar] [CrossRef]
- Tian, X.; Zhang, L.; Jiao, Y.; Chen, J.; Shan, Y.; Yang, W. CircABCB10 promotes nonsmall cell lung cancer cell proliferation and migration by regulating the miR-1252/FOXR2 axis. J. Cell. Biochem. 2019, 120, 3765–3772. [Google Scholar] [CrossRef]
- Yu, H.; Chen, Y.; Jiang, P. Circular RNA HIPK3 exerts oncogenic properties through suppression of miR-124 in lung cancer. Biochem. Biophys. Res. Commun. 2018, 506, 455–462. [Google Scholar] [CrossRef]
- Wan, L.; Zhang, L.; Fan, K.; Cheng, Z.X.; Sun, Q.C.; Wang, J.J. Circular RNA-ITCH Suppresses Lung Cancer Proliferation via Inhibiting the Wnt/beta-Catenin Pathway. Biomed Res. Int. 2016, 2016, 1579490. [Google Scholar] [CrossRef] [PubMed]
- Nan, A.; Chen, L.; Zhang, N.; Jia, Y.; Li, X.; Zhou, H.; Ling, Y.; Wang, Z.; Yang, C.; Liu, S.; et al. Circular RNA circNOL10 Inhibits Lung Cancer Development by Promoting SCLM1-Mediated Transcriptional Regulation of the Humanin Polypeptide Family. Adv. Sci. (Weinh. Baden-Wurtt. Ger.) 2019, 6, 1800654. [Google Scholar] [CrossRef]
- Russell, A.G.; Watanabe, Y.; Charette, J.M.; Gray, M.W. Unusual features of fibrillarin cDNA and gene structure in Euglena gracilis: Evolutionary conservation of core proteins and structural predictions for methylation-guide box C/D snoRNPs throughout the domain Eucarya. Nucleic Acids Res. 2005, 33, 2781–2791. [Google Scholar] [CrossRef]
- Khanna, M.; Wu, H.; Johansson, C.; Caizergues-Ferrer, M.; Feigon, J. Structural study of the H/ACA snoRNP components Nop10p and the 3’ hairpin of U65 snoRNA. RNA 2006, 12, 40–52. [Google Scholar] [CrossRef]
- Newman, D.R.; Kuhn, J.F.; Shanab, G.M.; Maxwell, E.S. Box C/D snoRNA-associated proteins: Two pairs of evolutionarily ancient proteins and possible links to replication and transcription. RNA 2000, 6, 861–879. [Google Scholar] [CrossRef]
- Taft, R.J.; Glazov, E.A.; Lassmann, T.; Hayashizaki, Y.; Carninci, P.; Mattick, J.S. Small RNAs derived from snoRNAs. RNA 2009, 15, 1233–1240. [Google Scholar] [CrossRef]
- Bai, B.; Yegnasubramanian, S.; Wheelan, S.J.; Laiho, M. RNA-Seq of the nucleolus reveals abundant SNORD44-derived small RNAs. PLoS ONE 2014, 9, e107519. [Google Scholar] [CrossRef]
- Martens-Uzunova, E.S.; Olvedy, M.; Jenster, G. Beyond microRNA--novel RNAs derived from small non-coding RNA and their implication in cancer. Cancer Lett. 2013, 340, 201–211. [Google Scholar] [CrossRef]
- Zhong, F.; Zhou, N.; Wu, K.; Guo, Y.; Tan, W.; Zhang, H.; Zhang, X.; Geng, G.; Pan, T.; Luo, H.; et al. A SnoRNA-derived piRNA interacts with human interleukin-4 pre-mRNA and induces its decay in nuclear exosomes. Nucleic Acids Res. 2015, 43, 10474–10491. [Google Scholar] [CrossRef]
- Ono, M.; Yamada, K.; Avolio, F.; Scott, M.S.; van Koningsbruggen, S.; Barton, G.J.; Lamond, A.I. Analysis of human small nucleolar RNAs (snoRNA) and the development of snoRNA modulator of gene expression vectors. Mol. Biol. Cell 2010, 21, 1569–1584. [Google Scholar] [CrossRef]
- Mei, Y.P.; Liao, J.P.; Shen, J.; Yu, L.; Liu, B.L.; Liu, L.; Li, R.Y.; Ji, L.; Dorsey, S.G.; Jiang, Z.R.; et al. Small nucleolar RNA 42 acts as an oncogene in lung tumorigenesis. Oncogene 2012, 31, 2794–2804. [Google Scholar] [CrossRef]
- Nogueira Jorge, N.A.; Wajnberg, G.; Ferreira, C.G.; de Sa Carvalho, B.; Passetti, F. snoRNA and piRNA expression levels modified by tobacco use in women with lung adenocarcinoma. PLoS ONE 2017, 12, e0183410. [Google Scholar] [CrossRef]
- Zheng, D.; Zhang, J.; Ni, J.; Luo, J.; Wang, J.; Tang, L.; Zhang, L.; Wang, L.; Xu, J.; Su, B.; et al. Small nucleolar RNA 78 promotes the tumorigenesis in non-small cell lung cancer. J. Exp. Clin. Cancer Res. Cr 2015, 34, 49. [Google Scholar] [CrossRef]
- Mannoor, K.; Shen, J.; Liao, J.; Liu, Z.; Jiang, F. Small nucleolar RNA signatures of lung tumor-initiating cells. Mol. Cancer 2014, 13, 104. [Google Scholar] [CrossRef]
- Gong, J.; Li, Y.; Liu, C.J.; Xiang, Y.; Li, C.; Ye, Y.; Zhang, Z.; Hawke, D.H.; Park, P.K.; Diao, L.; et al. A Pan-cancer Analysis of the Expression and Clinical Relevance of Small Nucleolar RNAs in Human Cancer. Cell Rep. 2017, 21, 1968–1981. [Google Scholar] [CrossRef]
- Gao, L.; Ma, J.; Mannoor, K.; Guarnera, M.A.; Shetty, A.; Zhan, M.; Xing, L.; Stass, S.A.; Jiang, F. Genome-wide small nucleolar RNA expression analysis of lung cancer by next-generation deep sequencing. Int. J. Cancer 2015, 136, E623–E629. [Google Scholar] [CrossRef]
- Su, Y.; Guarnera, M.A.; Fang, H.; Jiang, F. Small non-coding RNA biomarkers in sputum for lung cancer diagnosis. Mol. Cancer 2016, 15, 36. [Google Scholar] [CrossRef]
- Han, Y.N.; Li, Y.; Xia, S.Q.; Zhang, Y.Y.; Zheng, J.H.; Li, W. PIWI Proteins and PIWI-Interacting RNA: Emerging Roles in Cancer. Cell. Physiol. Biochem. Int. J. Exp. Cell. Physiol. Biochem. Pharmacol. 2017, 44, 1–20. [Google Scholar] [CrossRef]
- Dorner, S.; Eulalio, A.; Huntzinger, E.; Izaurralde, E. Delving into the diversity of silencing pathways. Symp. Micrornas Sirnas: Biol. Funct. Mech. 2007, 8, 723–729. [Google Scholar] [CrossRef]
- Luo, S.; Lu, J. Silencing of Transposable Elements by piRNAs in Drosophila: An Evolutionary Perspective. Genom. Proteom. Bioinformat. 2017, 15, 164–176. [Google Scholar] [CrossRef] [PubMed]
- Siomi, M.C.; Sato, K.; Pezic, D.; Aravin, A.A. PIWI-interacting small RNAs: The vanguard of genome defence. Nat. Rev. Mol. Cell Biol. 2011, 12, 246. [Google Scholar] [CrossRef]
- Reeves, M.E.; Firek, M.; Jliedi, A.; Amaar, Y.G. Identification and characterization of RASSF1C piRNA target genes in lung cancer cells. Oncotarget 2017, 8, 34268–34282. [Google Scholar] [CrossRef]
- Mei, Y.; Wang, Y.; Kumari, P.; Shetty, A.C.; Clark, D.; Gable, T.; MacKerell, A.D.; Ma, M.Z.; Weber, D.J.; Yang, A.J.; et al. A piRNA-like small RNA interacts with and modulates p-ERM proteins in human somatic cells. Nat. Commun. 2015, 6, 7316. [Google Scholar] [CrossRef]
- Fehon, R.G.; McClatchey, A.I.; Bretscher, A. Organizing the cell cortex: The role of ERM proteins. Nat. Rev. Mol. Cell Biol. 2010, 11, 276–287. [Google Scholar] [CrossRef]
- Zhang, S.J.; Yao, J.; Shen, B.Z.; Li, G.B.; Kong, S.S.; Bi, D.D.; Pan, S.H.; Cheng, B.L. Role of piwi-interacting RNA-651 in the carcinogenesis of non-small cell lung cancer. Oncol. Lett. 2018, 15, 940–946. [Google Scholar] [CrossRef]
- Li, C.; Qin, F.; Hu, F.; Xu, H.; Sun, G.; Han, G.; Wang, T.; Guo, M. Characterization and selective incorporation of small non-coding RNAs in non-small cell lung cancer extracellular vesicles. Cell Biosci. 2018, 8, 2. [Google Scholar] [CrossRef]
- Liao, J.; Yu, L.; Mei, Y.; Guarnera, M.; Shen, J.; Li, R.; Liu, Z.; Jiang, F. Small nucleolar RNA signatures as biomarkers for non-small-cell lung cancer. Mol. Cancer 2010, 9, 198. [Google Scholar] [CrossRef] [PubMed]
- Balbin, O.A.; Malik, R.; Dhanasekaran, S.M.; Prensner, J.R.; Cao, X.; Wu, Y.M.; Robinson, D.; Wang, R.; Chen, G.; Beer, D.G.; et al. The landscape of antisense gene expression in human cancers. Genome Res. 2015, 25, 1068–1079. [Google Scholar] [CrossRef]
- Yuan, X.S.; Cao, L.X.; Hu, Y.J.; Bao, F.C.; Wang, Z.T.; Cao, J.L.; Yuan, P.; Lv, W.; Hu, J. Clinical, cellular, and bioinformatic analyses reveal involvement of WRAP53 overexpression in carcinogenesis of lung adenocarcinoma. Tumour Biol. J. Int. Soc. Oncodevelopmental Biol. Med. 2017, 39. [Google Scholar] [CrossRef] [PubMed]
- Shi, R.; Jiao, Z.; Yu, A.; Wang, T. Long noncoding antisense RNA FAM83A-AS1 promotes lung cancer cell progression by increasing FAM83A. J. Cell. Biochem. 2019. [Google Scholar] [CrossRef] [PubMed]
- He, J.; Wu, K.; Guo, C.; Zhou, J.K.; Pu, W.; Deng, Y.; Zuo, Y.; Zhao, Y.; Liu, L.; Wei, Y.Q.; et al. Long non-coding RNA AFAP1-AS1 plays an oncogenic role in promoting cell migration in non-small cell lung cancer. Cell. Mol. Life Sci. Cmls 2018, 75, 4667–4681. [Google Scholar] [CrossRef] [PubMed]
- Zhang, H.; Xiong, Y.; Xia, R.; Wei, C.; Shi, X.; Nie, F. The pseudogene-derived long noncoding RNA SFTA1P is down-regulated and suppresses cell migration and invasion in lung adenocarcinoma. Tumour Biol. J. Int. Soc. Oncodevelopmental Biol. Med. 2017, 39. [Google Scholar] [CrossRef]
- Sun, M.; Nie, F.Q.; Zang, C.; Wang, Y.; Hou, J.; Wei, C.; Li, W.; He, X.; Lu, K.H. The Pseudogene DUXAP8 Promotes Non-small-cell Lung Cancer Cell Proliferation and Invasion by Epigenetically Silencing EGR1 and RHOB. Mol. Ther. J. Am. Soc. Gene Ther. 2017, 25, 739–751. [Google Scholar] [CrossRef]
- Wei, C.C.; Nie, F.Q.; Jiang, L.L.; Chen, Q.N.; Chen, Z.Y.; Chen, X.; Pan, X.; Liu, Z.L.; Lu, B.B.; Wang, Z.X. The pseudogene DUXAP10 promotes an aggressive phenotype through binding with LSD1 and repressing LATS2 and RRAD in non small cell lung cancer. Oncotarget 2017, 8, 5233–5246. [Google Scholar] [CrossRef]
- Gao, X.; Gao, X.; Li, C.; Zhang, Y.; Gao, L. Knockdown of Long Noncoding RNA uc.338 by siRNA Inhibits Cellular Migration and Invasion in Human Lung Cancer Cells. Oncol. Res. 2016, 24, 337–343. [Google Scholar] [CrossRef]
- Vannini, I.; Wise, P.M.; Challagundla, K.B.; Plousiou, M.; Raffini, M.; Bandini, E.; Fanini, F.; Paliaga, G.; Crawford, M.; Ferracin, M.; et al. Transcribed ultraconserved region 339 promotes carcinogenesis by modulating tumor suppressor microRNAs. Nat. Commun. 2017, 8, 1801. [Google Scholar] [CrossRef]
- Zhou, J.; Wang, C.; Gong, W.; Wu, Y.; Xue, H.; Jiang, Z.; Shi, M. uc.454 Inhibited Growth by Targeting Heat Shock Protein Family A Member 12B in Non-Small-Cell Lung Cancer. Mol. Ther. Nucleic Acids 2018, 12, 174–183. [Google Scholar] [CrossRef]
- Langley, A.R.; Chambers, H.; Christov, C.P.; Krude, T. Ribonucleoprotein Particles Containing Non-Coding Y RNAs, Ro60, La and Nucleolin Are Not Required for Y RNA Function in DNA Replication. PLoS ONE 2010, 5, e13673. [Google Scholar] [CrossRef]
- Driedonks, T.A.P.; Nolte-’t Hoen, E.N.M. Circulating Y-RNAs in Extracellular Vesicles and Ribonucleoprotein Complexes; Implications for the Immune System. Front. Immunol. 2019, 9, 3164. [Google Scholar] [CrossRef]
- Gulei, D.; Irimie, A.I.; Cojocneanu-Petric, R.; Schultze, J.L.; Berindan-Neagoe, I. Exosomes-Small Players, Big Sound. Bioconjugate Chem. 2018, 29, 635–648. [Google Scholar] [CrossRef]
- Braicu, C.; Tomuleasa, C.; Monroig, P.; Cucuianu, A.; Berindan-Neagoe, I.; Calin, G.A. Exosomes as divine messengers: Are they the Hermes of modern molecular oncology? Cell Death Differ. 2015, 22, 34–45. [Google Scholar] [CrossRef]
- Werner, A.; Swan, D. What are natural antisense transcripts good for? Biochem. Soc. Trans. 2010, 38, 1144–1149. [Google Scholar] [CrossRef]
- Sun, Y.; Li, D.; Zhang, R.; Peng, S.; Zhang, G.; Yang, T.; Qian, A. Strategies to identify natural antisense transcripts. Biochimie 2017, 132, 131–151. [Google Scholar] [CrossRef]
- Zinad, H.S.; Natasya, I.; Werner, A. Natural Antisense Transcripts at the Interface between Host Genome and Mobile Genetic Elements. Front. Microbiol. 2017, 8, 2292. [Google Scholar] [CrossRef]
- Nishizawa, M.; Okumura, T.; Ikeya, Y.; Kimura, T. Regulation of Inducible Gene Expression by Natural Antisense Transcripts. Front. Biosci. 2012, 17, 938–958. [Google Scholar] [CrossRef][Green Version]
- Wanowska, E.; Kubiak, M.R.; Rosikiewicz, W.; Makałowska, I.; Szcześniak, M.W. Natural antisense transcripts in diseases: From modes of action to targeted therapies. Wiley Interdiscip. Rev. RNA 2018, 9, e1461. [Google Scholar] [CrossRef]
- Kim, D.S.; Lee, W.K.; Park, J.Y. Promoter methylation of Wrap53alpha, an antisense transcript of p53, is associated with the poor prognosis of patients with non-small cell lung cancer. Oncol. Lett. 2018, 16, 5823–5828. [Google Scholar] [CrossRef]
- Tutar, Y. Pseudogenes. Comp. Funct. Genom. 2012, 2012, 424526. [Google Scholar] [CrossRef]
- Pink, R.C.; Wicks, K.; Caley, D.P.; Punch, E.K.; Jacobs, L.; Carter, D.R.F. Pseudogenes: Pseudo-functional or key regulators in health and disease? RNA 2011, 17, 792–798. [Google Scholar] [CrossRef]
- Cooke, S.L.; Shlien, A.; Marshall, J.; Pipinikas, C.P.; Martincorena, I.; Tubio, J.M.; Li, Y.; Menzies, A.; Mudie, L.; Ramakrishna, M.; et al. Processed pseudogenes acquired somatically during cancer development. Nat. Commun. 2014, 5, 3644. [Google Scholar] [CrossRef]
- Shang, J.; Wang, Z.; Chen, W.; Yang, Z.; Zheng, L.; Wang, S.; Li, S. Pseudogene CHIAP2 inhibits proliferation and invasion of lung adenocarcinoma cells by means of the WNT pathway. J. Cell. Physiol. 2019. [Google Scholar] [CrossRef]
- Xu, T.; Li, D.; He, Y.; Zhang, F.; Qiao, M.; Chen, Y. The expression level of CSDAP1 in lung cancer and its clinical significance. Oncol. Lett. 2018, 16, 4361–4366. [Google Scholar] [CrossRef]
- Peng, J.C.; Shen, J.; Ran, Z.H. Transcribed ultraconserved region in human cancers. RNA Biol. 2013, 10, 1771–1777. [Google Scholar] [CrossRef]
- Leao, R.; Apolonio, J.D.; Lee, D.; Figueiredo, A.; Tabori, U.; Castelo-Branco, P. Mechanisms of human telomerase reverse transcriptase (hTERT) regulation: Clinical impacts in cancer. J. Biomed. Sci. 2018, 25, 22. [Google Scholar] [CrossRef]
- Webb, C.J.; Zakian, V.A. Telomerase RNA is more than a DNA template. RNA Biol. 2016, 13, 683–689. [Google Scholar] [CrossRef] [PubMed]
- Eldholm, V.; Haugen, A.; Zienolddiny, S. CTCF mediates the TERT enhancer-promoter interactions in lung cancer cells: Identification of a novel enhancer region involved in the regulation of TERT gene. Int. J. Cancer 2014, 134, 2305–2313. [Google Scholar] [CrossRef] [PubMed]
- Ye, G.; Tan, N.; Meng, C.; Li, J.; Jing, L.; Yan, M.; Jin, T.; Chen, F. Genetic variations in TERC and TERT genes are associated with lung cancer risk in a Chinese Han population. Oncotarget 2017, 8, 110145–110152. [Google Scholar] [CrossRef] [PubMed]
- Li, X.; Xu, X.; Fang, J.; Wang, L.; Mu, Y.; Zhang, P.; Yao, Z.; Ma, Z.; Liu, Z. Rs2853677 modulates Snail1 binding to the TERT enhancer and affects lung adenocarcinoma susceptibility. Oncotarget 2016, 7, 37825–37838. [Google Scholar] [CrossRef] [PubMed]
- Arinaga, M.; Shimizu, S.; Gotoh, K.; Haruki, N.; Takahashi, T.; Takahashi, T.; Mitsudomi, T. Expression of human telomerase subunit genes in primary lung cancer and its clinical significance. Ann. Thorac. Surg. 2000, 70, 401–405. [Google Scholar] [CrossRef]
| Type of RNAs | Length | Region of the DNA | Localization | Interaction | Molecular Role | Ref. |
|---|---|---|---|---|---|---|
| Short ncRNAs | ||||||
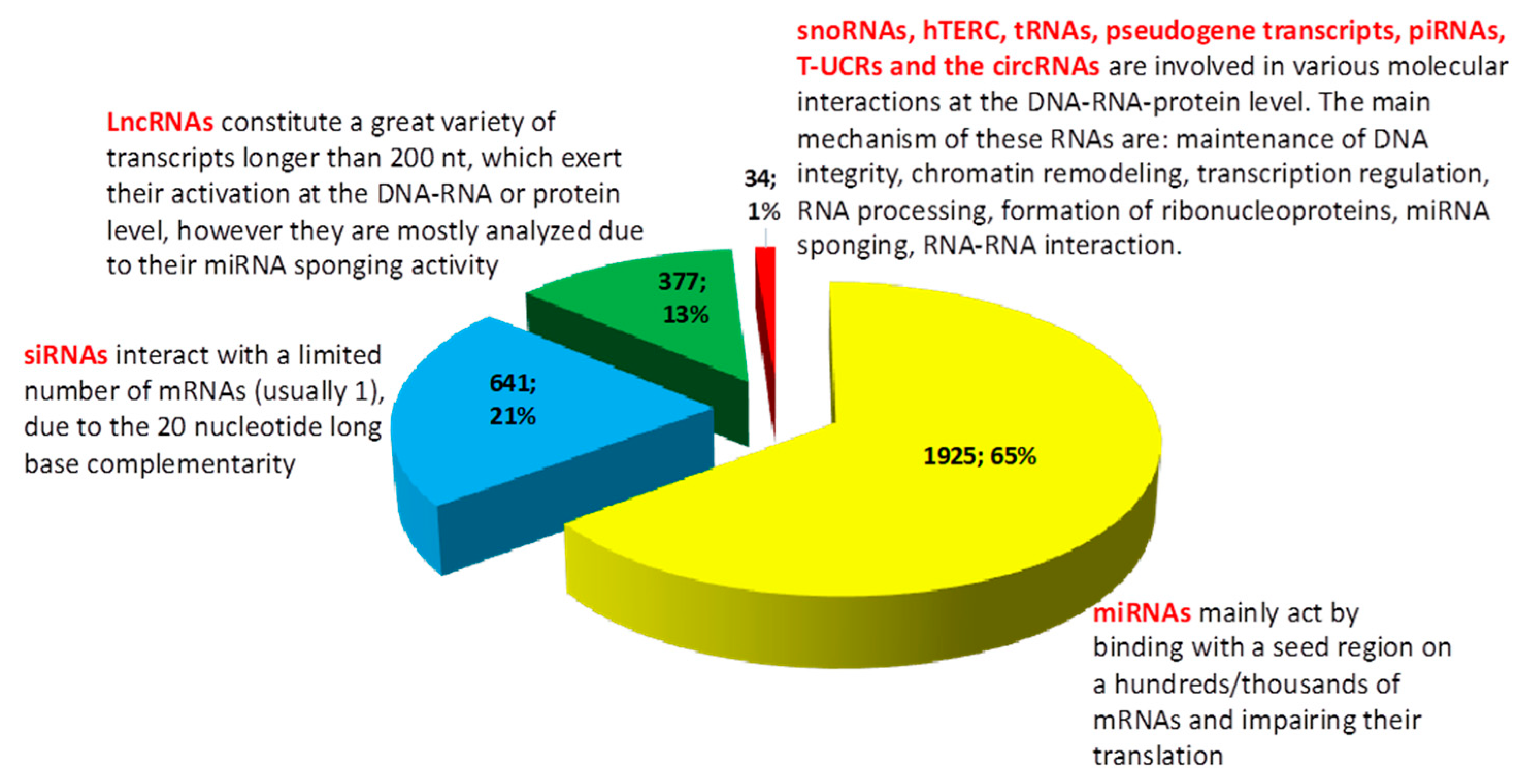
| miRNA | 22–24 nt | Intergenic regions and introns of protein-coding genes. | Nucleus, Cytoplasm | mRNA, circRNA, NAT, pseudogene transcript, T-UCR | Translation suppression | [15,16] |
| siRNA | 20–30 nt | Pseudogenes, intergenic repetitive sequence, endo-siRNA gene clusters | Cytoplasm | mRNA | Translation repression | [17,18] |
| piRNA | 24–31 nt | Flamenco locus, containing fragmented transposons | Cytoplasm | mRNA | Translation repression, modulation of transposons | [19,20] |
| tRF | 20 nt | tRNA coding transcripts | Nucleus | mRNA, transportable elements | Cell proliferation, translation repression, target transportable element | [21,22] |
| tiRNA | 30–40 nt | tRNA coding transcripts | Cytoplasm | mRNA, tRF | Translation repression, signaling molecule | [23] |
| YRNA | 84–112 nt | yDNA | Nucleus | DNA, primary RNA transcripts | Misfolded RNA degradation, DNA replication | [24,25,26] |
| snoRNA | 60–200 nt C/D snoRNAs, 120–250 nt for H/ACA snoRNAs | introns, promoter region of Pol II | Nucleus, Cytoplasm | mRNA, DNA, primary transcripts | Transcription regulation by binding to TATA box, rRNA processing, splicing, miRNA-like functions, chromatin remodeling, DNA replication, generation of piRNA/miRNA | [27,28] |
| Long non-coding RNA | ||||||
| TERC | 451 nt | promoter region of pol II | Nucleus | DNA, telomerase | Telomere length maintenance | [29,30] |
| NAT | >200 nt Depending on the antisense gene length | antisense strand of protein-coding transcript | Nucleus, Cytoplasm | DNA, miRNA, mRNA | Inhibition of the mRNA, epigenetic gene silencing, masking miRNA binding site on the mRNA, entrapment of splicing machinery | [31,32,33] |
| T-UCR | >200 nt | ultraconserved regions of the DNA | Cytoplasm | miRNA | miRNA sponge | [34] |
| Long non-coding RNAs with coding potential | ||||||
| circRNA | 200–800 nt | circular RNA | Nucleus, Cytoplasm | miRNA, mRNA, rRNA | Epigenetic silencing of genes, miRNA sponge, Translation repression, protein-coding function, protein scaffold | [35,36,37] |
| Pseudogene transcripts | >200 nt depending on the pseudogene length | Pseudotranscripts/pseudogene transcripts | Nucleus, Cytoplasm | miRNA, siRNA, | Translation repression, miRNA sponge, generation of miRNA/siRNA | [38,39] |
| Name | Targeted miRNA | Indirect Effect Over mRNA | Biological Effect | Reference |
|---|---|---|---|---|
| circ-RAD23B | miR-593-3p | CCND2 | Promotes invasion | [45] |
| miR-653-5p | TIAM1 | |||
| circRNA 100146 | miR-361-3p | NFAT5, COL1A1, TRAF3 | Promotes invasion and cell proliferation | [46] |
| miR-615-5p | MEF2C | |||
| circPVT1 | miR-497 | BCL-2 | Promotes apoptosis, impairs proliferation | [47] |
| miR-125b | E2F1 | Stimulated in vivo tumorigenesis | [50] | |
| circFGFR3 | miR-22-3p | Gal-1, p-AKT, and p-ERK1/2 | Promotes invasion | [51] |
| circ_0004015 | miR-1183 | PDPK1 | Decreases survival rate, promotes viability, proliferation, invasion and maybe gefitinib resistance | [52] |
| circPUM1 | miR-326 | CCND1 and BCL-2 | Promotes proliferation, invasion and migration | [53] |
| circFLI1 | miR-584-3p | ROCK1 | Promotes metastasis | [54] |
| circABCB10 | miR-1252 | FOXR2 | Promotes proliferation and migration | [55] |
| circHIPK3 | miR-124 | SphK1, STAT3 and CDK4 | Promotes proliferation and impairs apoptosis | [56] |
| Type of RNAs | Transcript | Up/Down | Biological Role | Reference |
|---|---|---|---|---|
| piRNA | piR-34871, piR-52200 | UP | It stimulates cell proliferation and apoptosis via RASSF1C gene | [78] |
| piR-35127, piR-46545 | DOWN | There was no experimental modulation of its expression. | [78] | |
| piR-L-163 | DOWN | It binds to p-ERM proteins that interact with transmembrane and cytoskeleton proteins thus impairing cell migration, cell cycle progression; | [79] | |
| tRNA fragments | ts-46, ts-47, ts-101 and ts-53 | DOWN | It impairs cell proliferation | [42] |
| tRF-Leu-CAG1 tRF-Leu-CAG2 | UP | It stimulates cell cycle progression and cell proliferation; by targeting AURKA protein | [43] | |
| YRNA | hY4 RNA | UP | It was found in the extracellular vesicle from lung tumor cells, it stimulates cell proliferation | [82] |
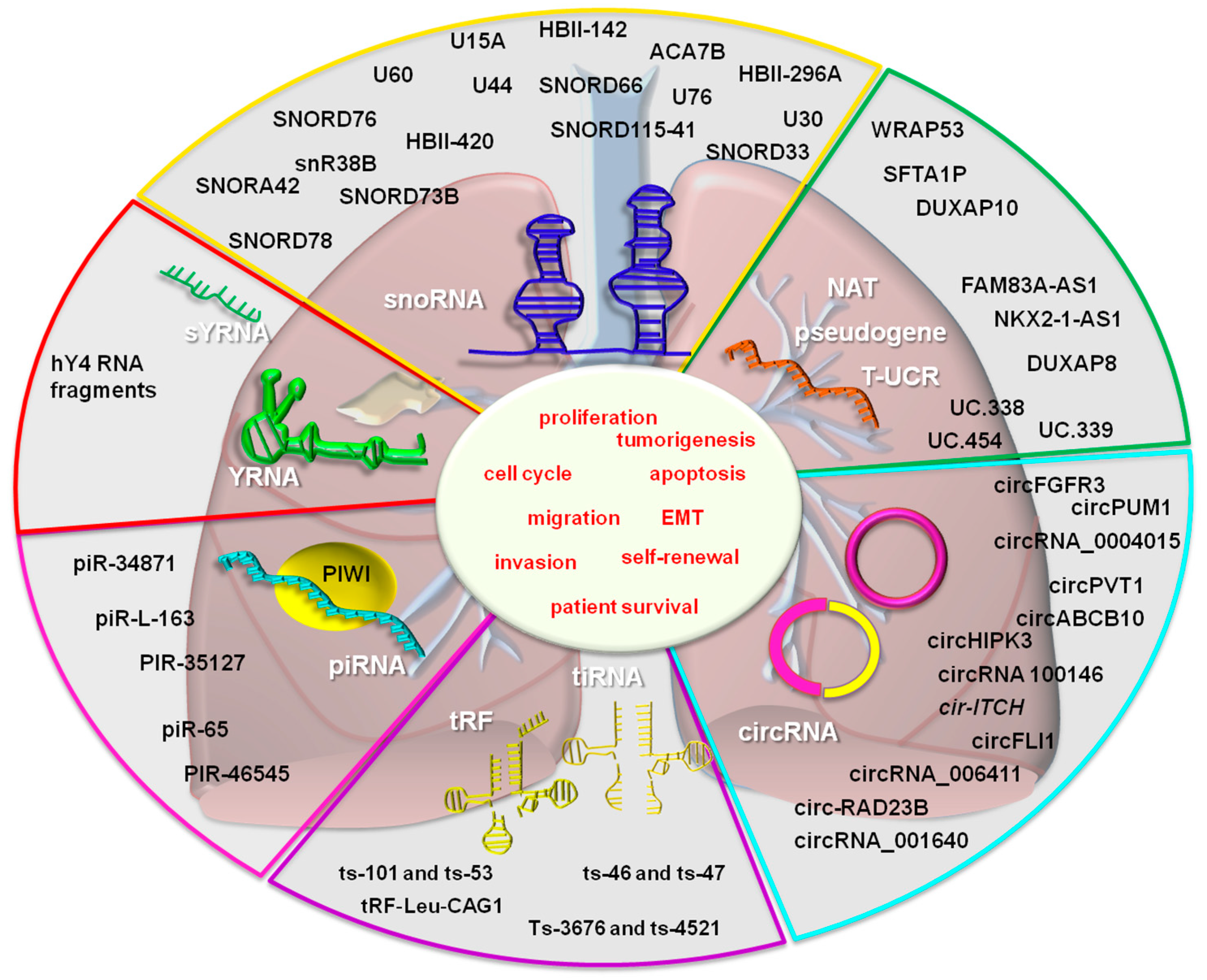
| SNORD | SNORA42 | UP | It maintains the tumor initiating phenotype of cancer cells | [70] |
| U60, U63, U28, U51, U104, HBII-419, U59B, HBII-142, HBI-100, U30 | UP | There was no experimental modulation of its expression. | [68] | |
| HBII-420 | DOWN | |||
| SNORD78 | UP | It increases the in vivo tumorigenesis, and in vitro lung malignant cell proliferation, cell cycle progression, invasion, self-renewal capacity, and maintenance of cell steamness | [69] | |
| SNORA47, SNORA68, SNORA78, SNORA21, SNORD28 SNORD66 | UP | There was no experimental modulation of its expression. | [72] | |
| SNORD33, SNORD66 SNORD76 | UP | There was no experimental modulation of its expression. | [83] | |
| NAT | NKX2-1-AS1 | UP | It increases lung malignant cell proliferation rate | [84] |
| WRAP53 | It increases lung malignant cell proliferation rate | [85] | ||
| FAM83A | It causes in vivo pronounced tumor progression | [86] | ||
| AFAP1-AS1 | It causes increased invasion and metastasis capacity of lung malignant cells | [87] | ||
| Pseudogene transcripts | SFTA1P | DOWN | It impairs cell migration and invasion | [88] |
| DUXAP8 | UP | It increases malignant cell survival and proliferation, it stimulates in vivo tumorigenesis | [89] | |
| DUXAP10 | It increases malignant cell survival, proliferation, and migration, it stimulates in vivo tumorigenesis | [90] | ||
| T-UCR | Uc.338 | UP | It increases malignant cell cycle progression, invasion, and migration | [91] |
| Uc.339 | UP | It increases malignant cell cycle progression, and migration | [92] | |
| Uc.454 | DOWN | It decreases malignant cell cycle progression, invasion, and migration | [93] |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Braicu, C.; Zimta, A.-A.; Harangus, A.; Iurca, I.; Irimie, A.; Coza, O.; Berindan-Neagoe, I. The Function of Non-Coding RNAs in Lung Cancer Tumorigenesis. Cancers 2019, 11, 605. https://doi.org/10.3390/cancers11050605
Braicu C, Zimta A-A, Harangus A, Iurca I, Irimie A, Coza O, Berindan-Neagoe I. The Function of Non-Coding RNAs in Lung Cancer Tumorigenesis. Cancers. 2019; 11(5):605. https://doi.org/10.3390/cancers11050605
Chicago/Turabian StyleBraicu, Cornelia, Alina-Andreea Zimta, Antonia Harangus, Ioana Iurca, Alexandru Irimie, Ovidiu Coza, and Ioana Berindan-Neagoe. 2019. "The Function of Non-Coding RNAs in Lung Cancer Tumorigenesis" Cancers 11, no. 5: 605. https://doi.org/10.3390/cancers11050605
APA StyleBraicu, C., Zimta, A.-A., Harangus, A., Iurca, I., Irimie, A., Coza, O., & Berindan-Neagoe, I. (2019). The Function of Non-Coding RNAs in Lung Cancer Tumorigenesis. Cancers, 11(5), 605. https://doi.org/10.3390/cancers11050605







