Automated Quantification of Extranuclear ERα Using Phosphor-Integrated Dots for Predicting Endocrine Therapy Resistance in HR+/HER2− Breast Cancer
Abstract
1. Introduction
2. Results
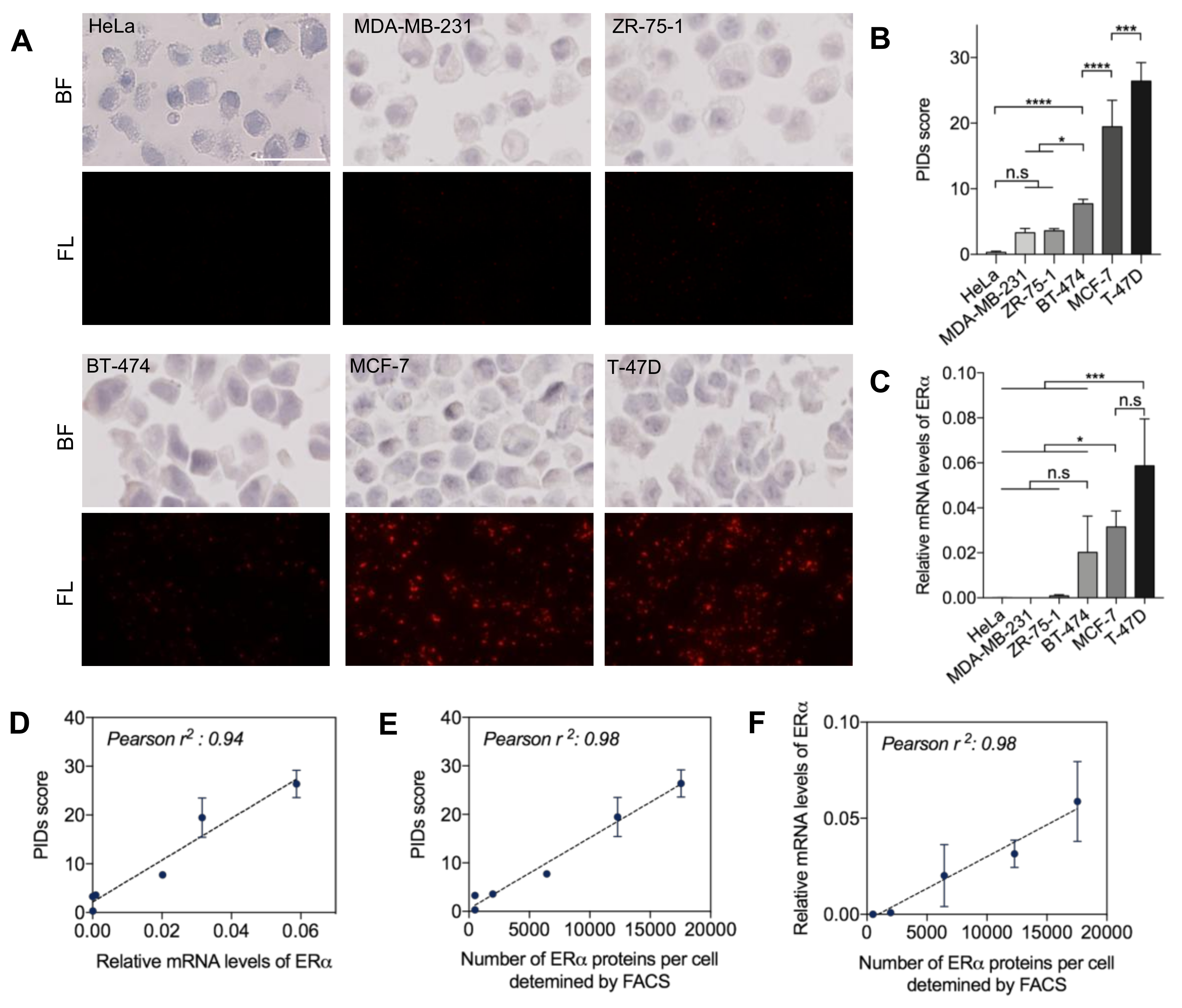
2.1. Quantitative Sensitivity of IHC-PIDs In Vitro
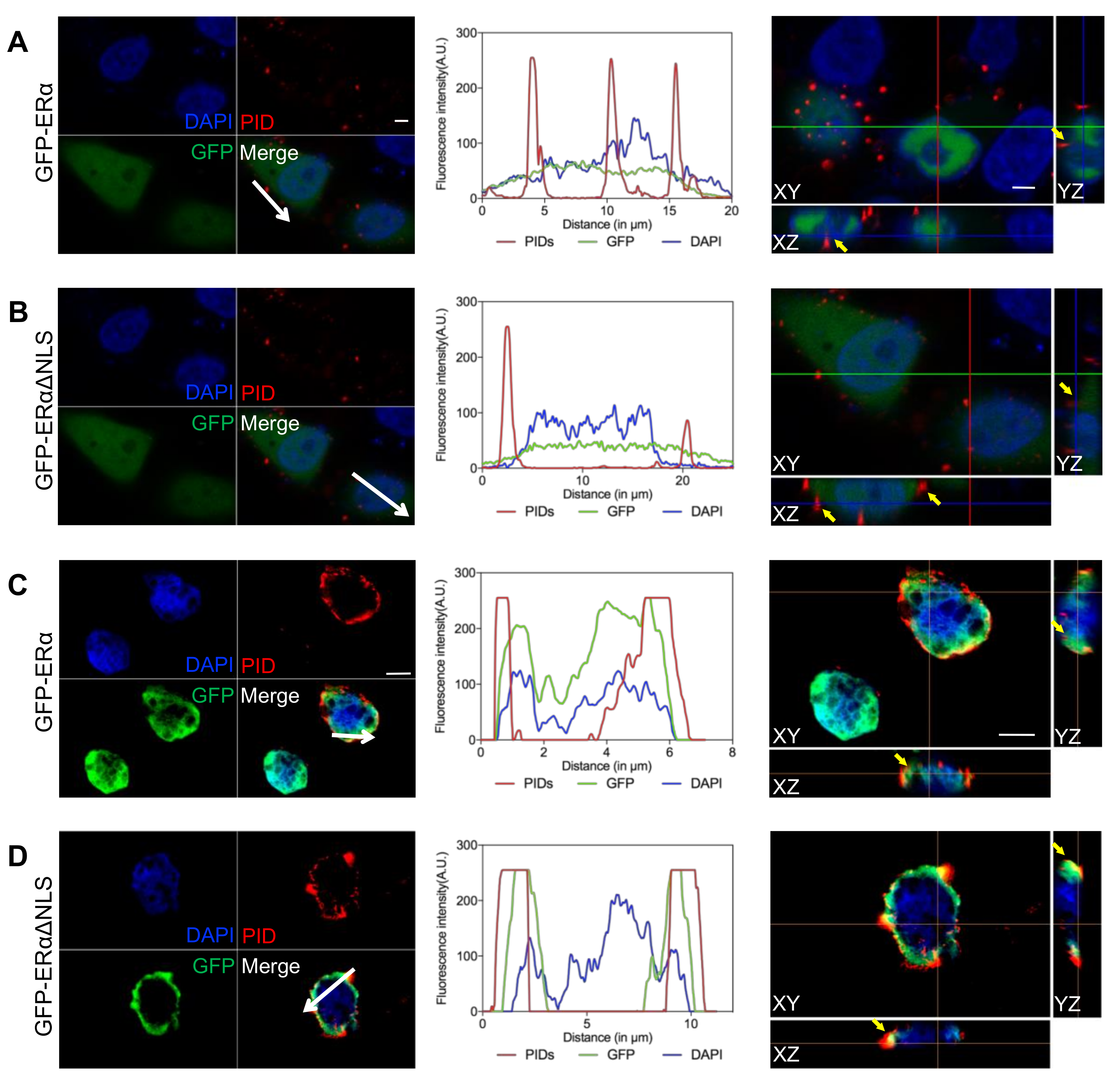
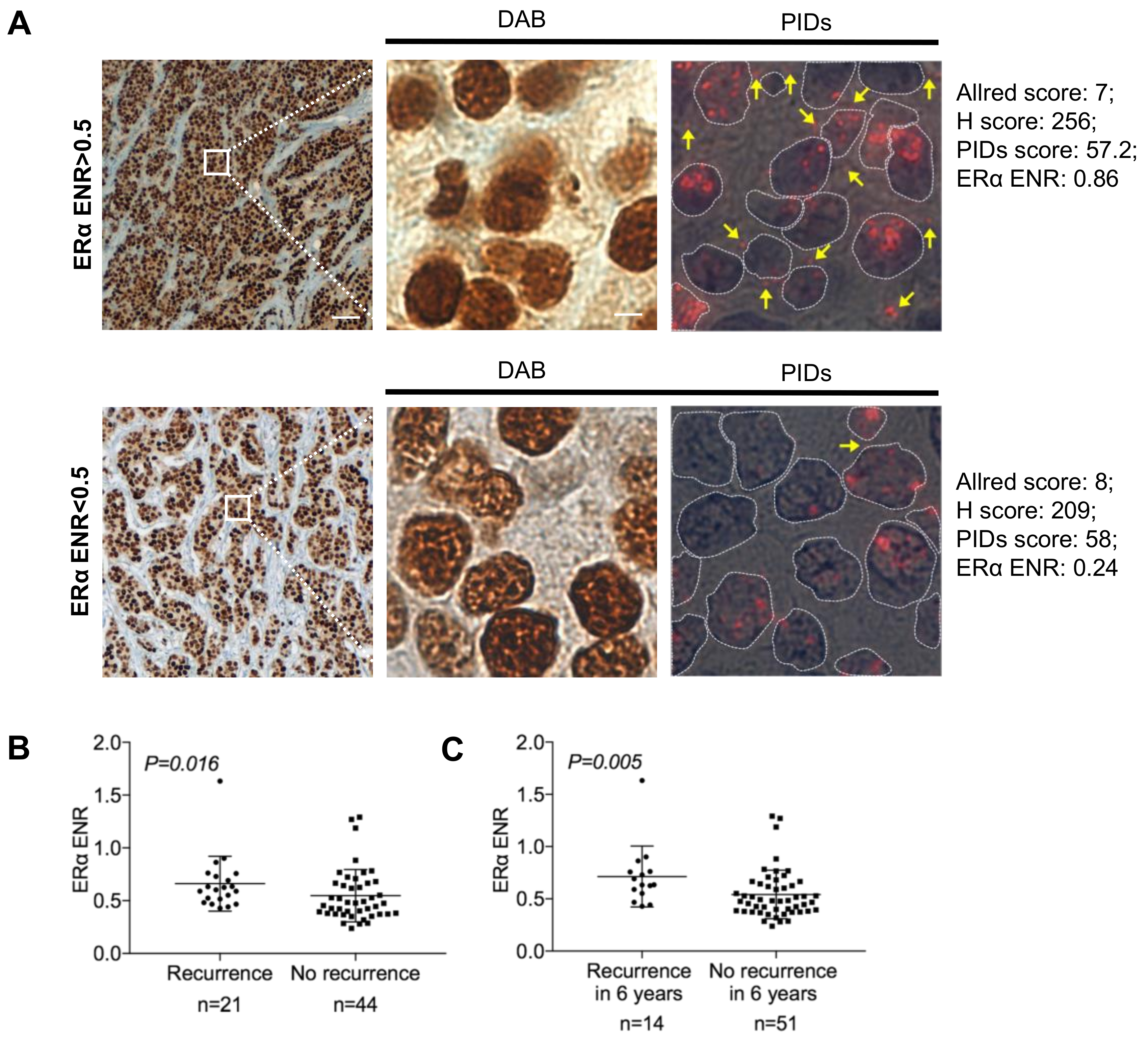
2.2. Identification of the Extranuclear ERα by IHC-PIDs
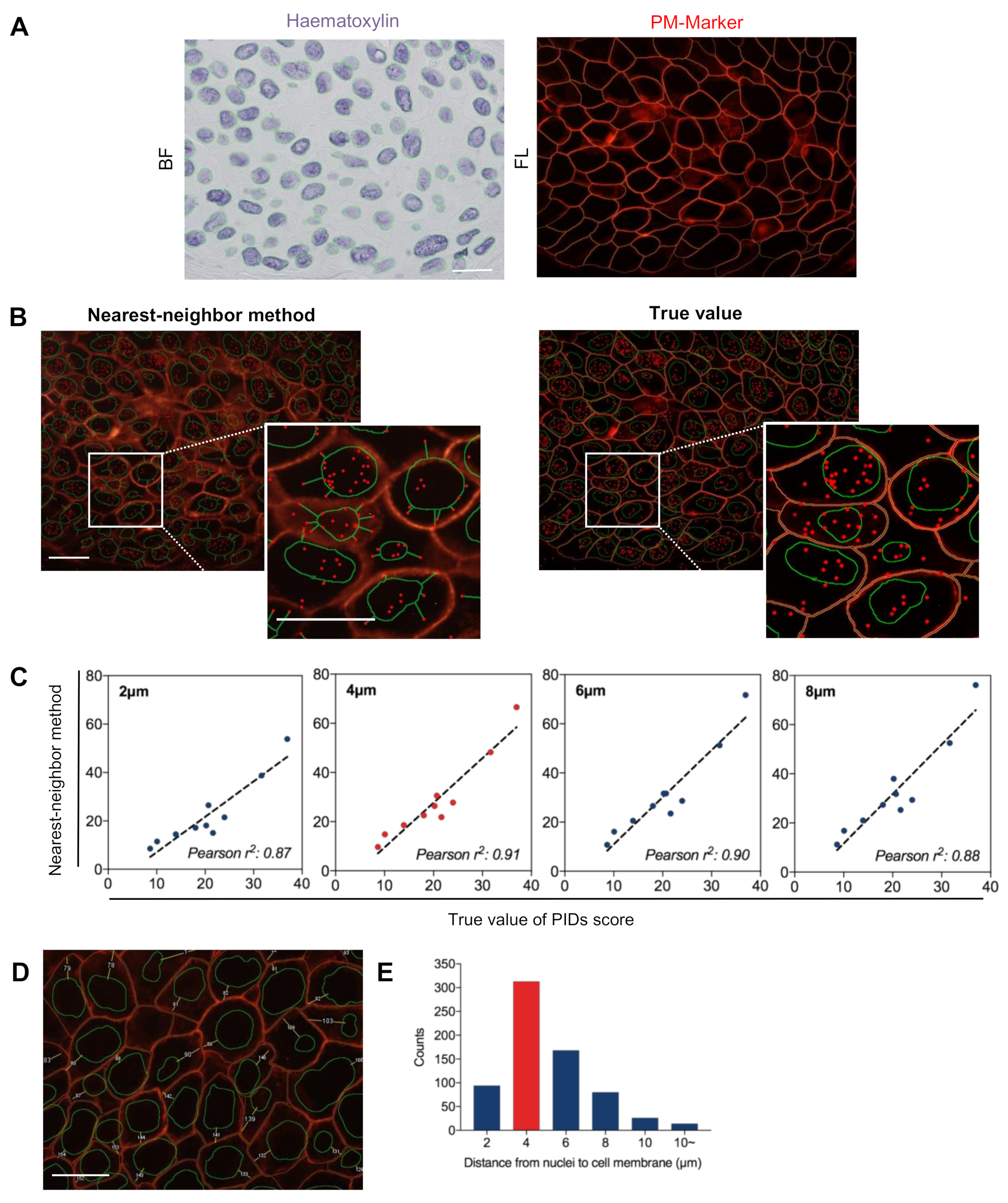
2.3. Validation of the Accuracy of the Nearest-Neighbor Method in Evaluating Extranuclear ERα
2.4. Correlation between Biological Factors and Clinical Parameters of Patients
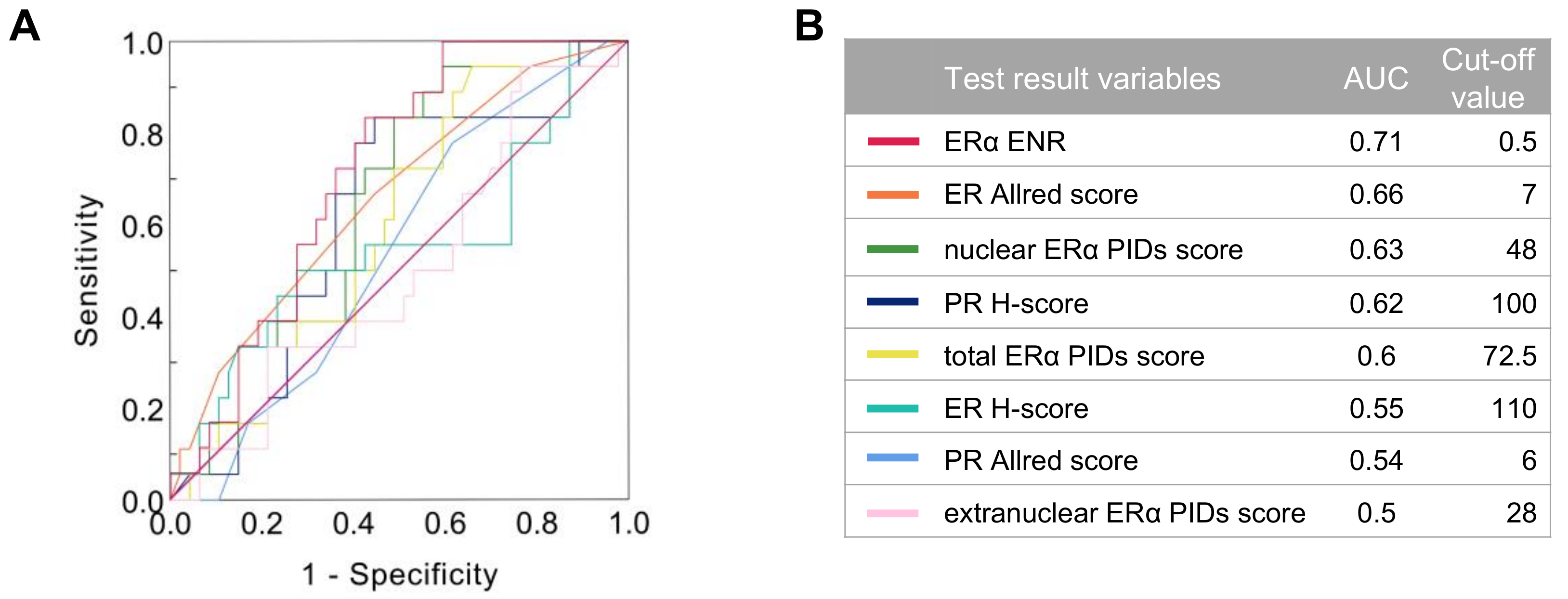
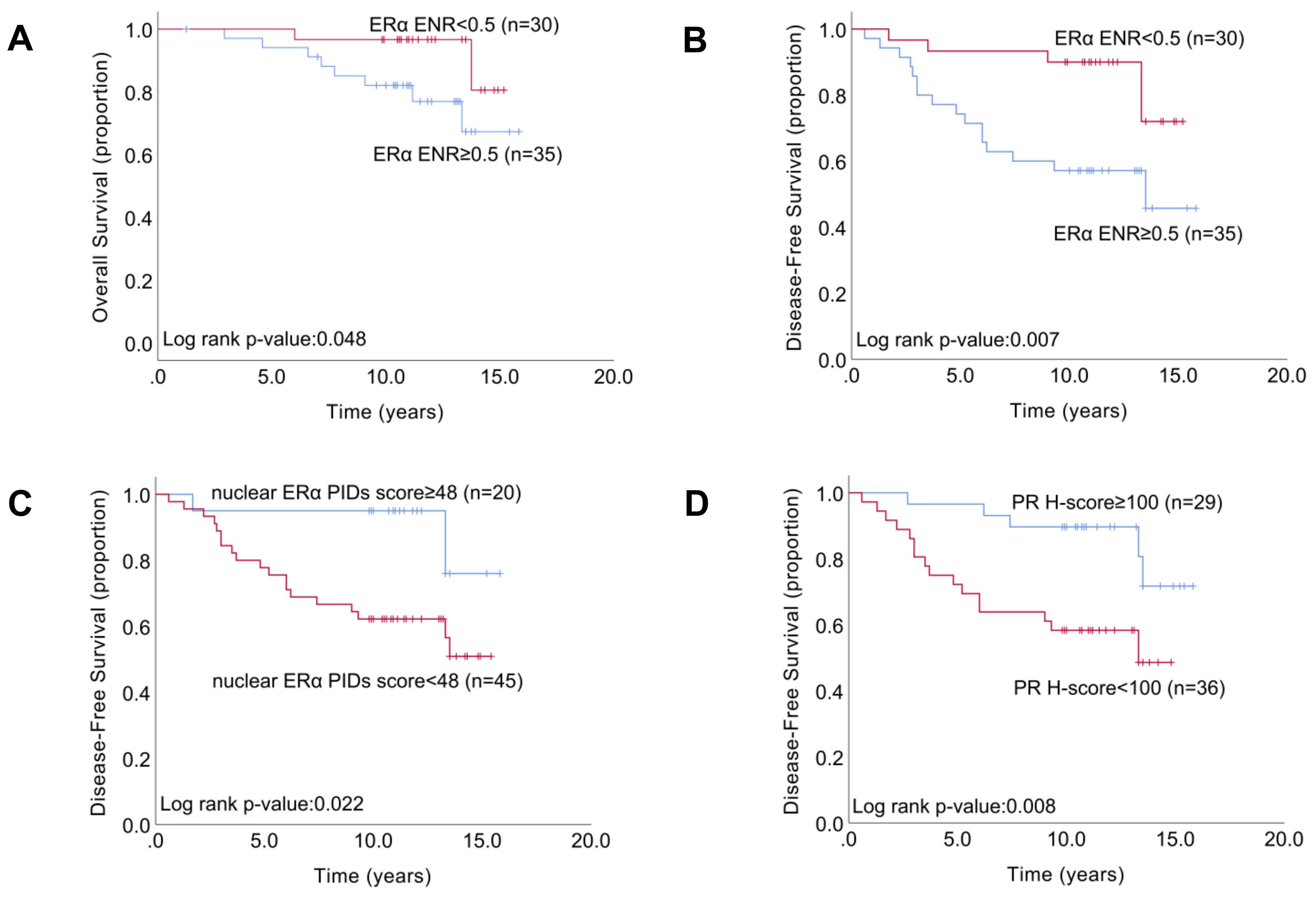
2.5. Relationship between the Indexes of ERα Expression and Patient Survival
3. Discussion
4. Materials and Methods
4.1. Patient Samples
4.2. Cell Lines and Paraffin Sections
4.3. Immunofluorescence
4.4. IHC-PIDs
4.5. Real-Time qRT-PCR
4.6. Flow Cytometry
4.7. Automatic Measurement of Nuclear and Extranuclear ERα
4.8. Statistical Analysis
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
Abbreviations
References
- Weatherman, R.V.; Fletterick, R.J.; Scanlan, T.S. Nuclear-Receptor Ligands and Ligand-Binding Domains. Annu. Rev. Biochem. 1999, 68, 559–581. [Google Scholar] [CrossRef] [PubMed]
- Clark, G.M.; Osborne, C.K.; McGuire, W.L. Correlations between estrogen receptor, progesterone receptor, and patient characteristics in human breast cancer. J. Clin. Oncol. 1984, 2, 1102–1109. [Google Scholar] [CrossRef]
- Rastelli, F.; Crispino, S. Factors predictive of response to hormone therapy in breast cancer. Tumori 2008, 94, 370–383. [Google Scholar] [CrossRef]
- Ring, A.; Dowsett, M. Mechanisms of tamoxifen resistance. Endocr. Relat. Cancer 2004, 11, 643–658. [Google Scholar] [CrossRef] [PubMed]
- Cardoso, F.; Costa, A.; Norton, L.; Senkus, E.; Aapro, M.; Andre, F.; Barrios, C.; Bergh, J.; Biganzoli, L.; Blackwell, K. ESO-ESMO 2nd international consensus guidelines for advanced breast cancer (ABC2). Ann. Oncol. 2014, 25, 1871–1888. [Google Scholar]
- Chen, Z.; Yuhanna, I.S.; Galcheva-Gargova, Z.; Karas, R.H.; Mendelsohn, M.E.; Shaul, P.W. Estrogen receptor α mediates the nongenomic activation of endothelial nitric oxide synthase by estrogen. J. Clin. Investig. 1999, 103, 401–406. [Google Scholar]
- Tecalco-Cruz, A.C.; Perez-Alvarado, I.A.; Ramirez-Jarquin, J.O.; Rocha-Zavaleta, L. Nucleo-cytoplasmic transport of estrogen receptor alpha in breast cancer cells. Cell. Sig. 2017, 34, 121–132. [Google Scholar] [CrossRef] [PubMed]
- Levin, E.R. Extranuclear Steroid Receptors Are Essential for Steroid Hormone Actions. Annu. Rev. Med. 2015, 66, 271–280. [Google Scholar] [CrossRef] [PubMed]
- Nardone, A.; De Angelis, C.; Trivedi, M.V.; Osborne, C.K.; Schiff, R. The changing role of ER in endocrine resistance. Breast 2015, 24, S60–S66. [Google Scholar] [CrossRef]
- Maurer, C.; Martel, S.; Zardavas, D.; Ignatiadis, M. New agents for endocrine resistance in breast cancer. Breast 2017, 34, 1–11. [Google Scholar] [CrossRef] [PubMed]
- Santen, R.J.; Song, R.X.; Masamura, S.; Yue, W.; Fan, P.; Sogon, T.; Hayashi, S.; Nakachi, K.; Eguchi, H. Adaptation to estradiol deprivation causes up-regulation of growth factor pathways and hypersensitivity to estradiol in breast cancer cells. Adv. Exp. Med. Biol. 2008, 630, 19–34. [Google Scholar] [PubMed]
- Razandi, M.; Pedram, A.; Greene, G.L.; Levin, E.R. Cell membrane and nuclear estrogen receptors (ERs) originate from a single transcript: Studies of ERalpha and ERbeta expressed in Chinese hamster ovary cells. Mol. Endocrinol. 1999, 13, 307–319. [Google Scholar] [CrossRef] [PubMed]
- Hammond, M.E.; Hayes, D.F.; Dowsett, M.; Allred, D.C.; Hagerty, K.L.; Badve, S.; Fitzgibbons, P.L.; Francis, G.; Goldstein, N.S.; Hayes, M.; et al. American Society of Clinical Oncology/College of American Pathologists guideline recommendations for immunohistochemical testing of estrogen and progesterone receptors in breast cancer. Arch. Pathol. Lab. Med. 2010, 134, 907–922. [Google Scholar] [CrossRef] [PubMed]
- Dan, P.; Cheung, J.C.; Scriven, D.R.; Moore, E.D. Epitope-dependent localization of estrogen receptor-alpha, but not -beta, in en face arterial endothelium. Am. J. Physiol. Heart Circul. Physiol. 2003, 284, H1295–H1306. [Google Scholar] [CrossRef] [PubMed]
- Welsh, A.W.; Lannin, D.R.; Young, G.S.; Sherman, M.E.; Figueroa, J.D.; Henry, N.L.; Ryden, L.; Kim, C.; Love, R.R.; Schiff, R.; et al. Cytoplasmic estrogen receptor in breast cancer. Clin. Cancer Res. 2012, 18, 118–126. [Google Scholar] [CrossRef] [PubMed]
- Chen, C.; Sun, S.-R.; Gong, Y.-P.; Qi, C.-B.; Peng, C.-W.; Yang, X.-Q.; Liu, S.-P.; Peng, J.; Zhu, S.; Hu, M.-B.; et al. Quantum dots-based molecular classification of breast cancer by quantitative spectroanalysis of hormone receptors and HER2. Biomaterials 2011, 32, 7592–7599. [Google Scholar] [CrossRef]
- Chen, C.; Xia, H.-S.; Gong, Y.-P.; Peng, J.; Peng, C.-W.; Hu, M.-B.; Zhu, X.-B.; Pang, D.-W.; Sun, S.-R.; Li, Y. The quantitative detection of total HER2 load by quantum dots and the identification of a new subtype of breast cancer with different 5-year prognosis. Biomaterials 2010, 31, 8818–8825. [Google Scholar] [CrossRef] [PubMed]
- Gonda, K.; Miyashita, M.; Higuchi, H.; Tada, H.; Watanabe, T.M.; Watanabe, M.; Ishida, T.; Ohuchi, N. Predictive diagnosis of the risk of breast cancer recurrence after surgery by single-particle quantum dot imaging. Sci. Rep. 2015, 5, 14322. [Google Scholar] [CrossRef]
- Miyashita, M.; Gonda, K.; Tada, H.; Watanabe, M.; Kitamura, N.; Kamei, T.; Sasano, H.; Ishida, T.; Ohuchi, N. Quantitative diagnosis of HER2 protein expressing breast cancer by single-particle quantum dot imaging. Cancer Med. 2016, 5, 2813–2824. [Google Scholar] [CrossRef]
- Gonda, K.; Miyashita, M.; Watanabe, M.; Takahashi, Y.; Goda, H.; Okada, H.; Nakano, Y.; Tada, H.; Amari, M.; Ohuchi, N. Development of a quantitative diagnostic method of estrogen receptor expression levels by immunohistochemistry using organic fluorescent material-assembled nanoparticles. Biochem. Biophys. Res. Commun. 2012, 426, 409–414. [Google Scholar] [CrossRef]
- Gonda, K.; Watanabe, M.; Tada, H.; Miyashita, M.; Takahashi-Aoyama, Y.; Kamei, T.; Ishida, T.; Usami, S.; Hirakawa, H.; Kakugawa, Y.; et al. Quantitative diagnostic imaging of cancer tissues by using phosphor-integrated dots with ultra-high brightness. Sci. Rep. 2017, 7, 7509. [Google Scholar] [CrossRef]
- Yamaki, S.; Yanagimoto, H.; Tsuta, K.; Ryota, H.; Kon, M. PD-L1 expression in pancreatic ductal adenocarcinoma is a poor prognostic factor in patients with high CD8(+) tumor-infiltrating lymphocytes: Highly sensitive detection using phosphor-integrated dot staining. Int. J. Clin. Oncol. 2017, 22, 726–733. [Google Scholar] [CrossRef]
- Rocha, R.; Nunes, C.; Rocha, G.; Oliveira, F.; Sanches, F.; Gobbi, H. Rabbit monoclonal antibodies show higher sensitivity than mouse monoclonals for estrogen and progesterone receptor evaluation in breast cancer by immunohistochemistry. Pathol. Res. Pract. 2008, 204, 655–662. [Google Scholar] [CrossRef] [PubMed]
- Madeira, K.P.; Daltoe, R.D.; Sirtoli, G.M.; Rezende, L.C.; Carvalho, A.A.; Guimaraes Idos, S.; Silva, I.V.; Rangel, L.B. Comparison of immunohistochemical analysis with estrogen receptor SP1 and 1D5 monoclonal antibodies in breast cancer. Pathol. Res. Pract. 2012, 208, 657–661. [Google Scholar] [CrossRef] [PubMed]
- Sobin, L.H.; Gospodarowicz, M.K.; Wittekind, C. TNM Classification of Malignant Tumours; John Wiley & Sons: Hoboken, NJ, USA, 2011. [Google Scholar]
- Allred, D.C.; Harvey, J.M.; Berardo, M.; Clark, G.M. Prognostic and predictive factors in breast cancer by immunohistochemical analysis. Mod. Pathol. 1998, 11, 155–168. [Google Scholar]
- Arias-Pulido, H.; Smith, H.O.; Joste, N.E.; Bocklage, T.; Qualls, C.R.; Chavez, A.; Prossnitz, E.R.; Verschraegen, C.F. Estrogen and progesterone receptor status and outcome in epithelial ovarian cancers and low malignant potential tumors. Gynecol. Oncol. 2009, 114, 480–485. [Google Scholar] [CrossRef]
- Sato-Tadano, A.; Suzuki, T.; Amari, M.; Takagi, K.; Miki, Y.; Tamaki, K.; Watanabe, M.; Ishida, T.; Sasano, H.; Ohuchi, N. Hexokinase II in breast carcinoma: A potent prognostic factor associated with hypoxia-inducible factor-1alpha and Ki-67. Cancer Sci. 2013, 104, 1380–1388. [Google Scholar] [CrossRef]
- Garcia-Becerra, R.; Santos, N.; Diaz, L.; Camacho, J. Mechanisms of resistance to endocrine therapy in breast cancer: Focus on signaling pathways, miRNAs and genetically based resistance. Int. J. Mol. Sci. 2012, 14, 108–145. [Google Scholar] [CrossRef] [PubMed]
- Fan, P.; Wang, J.; Santen, R.J.; Yue, W. Long-term treatment with tamoxifen facilitates translocation of estrogen receptor α out of the nucleus and enhances its interaction with EGFR in MCF-7 breast cancer cells. Cancer Res. 2007, 67, 1352–1360. [Google Scholar] [CrossRef]
- Kulkoyluoglu, E.; Madak-Erdogan, Z. Nuclear and extranuclear-initiated estrogen receptor signaling crosstalk and endocrine resistance in breast cancer. Steroids 2016, 114, 41–47. [Google Scholar] [CrossRef]
- Schiff, R.; Osborne, C.K. Endocrinology and hormone therapy in breast cancer: New insight into estrogen receptor-alpha function and its implication for endocrine therapy resistance in breast cancer. BCR 2005, 7, 205–211. [Google Scholar] [CrossRef]
- La Rosa, P.; Pesiri, V.; Leclercq, G.; Marino, M.; Acconcia, F. Palmitoylation regulates 17beta-estradiol-induced estrogen receptor-alpha degradation and transcriptional activity. Mol. Endocrinol. 2012, 26, 762–774. [Google Scholar] [CrossRef] [PubMed]
- Finn, R.S.; Crown, J.P.; Lang, I.; Boer, K.; Bondarenko, I.M.; Kulyk, S.O.; Ettl, J.; Patel, R.; Pinter, T.; Schmidt, M. The cyclin-dependent kinase 4/6 inhibitor palbociclib in combination with letrozole versus letrozole alone as first-line treatment of oestrogen receptor-positive, HER2-negative, advanced breast cancer (PALOMA-1/TRIO-18): A randomised phase 2 study. Lancet Oncol. 2015, 16, 25–35. [Google Scholar] [CrossRef]
- Bachelot, T.; Bourgier, C.; Cropet, C.; Ray-Coquard, I.; Ferrero, J.-M.; Freyer, G.; Abadie-Lacourtoisie, S.; Eymard, J.-C.; Debled, M.; Spaëth, D. Randomized phase II trial of everolimus in combination with tamoxifen in patients with hormone receptor–positive, human epidermal growth factor receptor 2–negative metastatic breast cancer with prior exposure to aromatase inhibitors: A GINECO study. J. Clin. Oncol. 2012, 30, 2718–2724. [Google Scholar] [CrossRef]
- Cortes, J.; Im, S.A.; Holgado, E.; Perez-Garcia, J.M.; Schmid, P.; Chavez-MacGregor, M. The next era of treatment for hormone receptor-positive, HER2-negative advanced breast cancer: Triplet combination-based endocrine therapies. Cancer Treat. Rev. 2017, 61, 53–60. [Google Scholar] [CrossRef]
- Dowsett, M.; Sestak, I.; Lopez-Knowles, E.; Sidhu, K.; Dunbier, A.K.; Cowens, J.W.; Ferree, S.; Storhoff, J.; Schaper, C.; Cuzick, J. Comparison of PAM50 risk of recurrence score with oncotype DX and IHC4 for predicting risk of distant recurrence after endocrine therapy. J. Clin. Oncol. 2013, 31, 2783–2790. [Google Scholar] [CrossRef]
- Orucevic, A.; Bell, J.L.; McNabb, A.P.; Heidel, R.E. Oncotype DX breast cancer recurrence score can be predicted with a novel nomogram using clinicopathologic data. Breast Cancer Res. Treat. 2017, 163, 51–61. [Google Scholar] [CrossRef]
- Felts, J.L.; Zhu, J.; Han, B.; Smith, S.J.; Truica, C.I. An Analysis of Oncotype DX Recurrence Scores and Clinicopathologic Characteristics in Invasive Lobular Breast Cancer. Breast J. 2017, 23, 677–686. [Google Scholar] [CrossRef] [PubMed]
- Gygi, S.P.; Rochon, Y.; Franza, B.R.; Aebersold, R. Correlation between protein and mRNA abundance in yeast. Mol. Cell. Biol. 1999, 19, 1720–1730. [Google Scholar] [CrossRef]
- Chen, G.; Gharib, T.G.; Huang, C.-C.; Taylor, J.M.; Misek, D.E.; Kardia, S.L.; Giordano, T.J.; Iannettoni, M.D.; Orringer, M.B.; Hanash, S.M. Discordant protein and mRNA expression in lung adenocarcinomas. Mol. Cell. Proteom. 2002, 1, 304–313. [Google Scholar] [CrossRef]
- Pentheroudakis, G.; Kalogeras, K.T.; Wirtz, R.M.; Grimani, I.; Zografos, G.; Gogas, H.; Stropp, U.; Pectasides, D.; Skarlos, D.; Hennig, G.; et al. Gene expression of estrogen receptor, progesterone receptor and microtubule-associated protein Tau in high-risk early breast cancer: A quest for molecular predictors of treatment benefit in the context of a Hellenic Cooperative Oncology Group trial. Breast Cancer Res. Treat. 2009, 116, 131–143. [Google Scholar] [CrossRef]
- Prat, A.; Cheang, M.C.U.; Martín, M.; Parker, J.S.; Carrasco, E.; Caballero, R.; Tyldesley, S.; Gelmon, K.; Bernard, P.S.; Nielsen, T.O.; et al. Prognostic Significance of Progesterone Receptor–Positive Tumor Cells Within Immunohistochemically Defined Luminal A Breast Cancer. J. Clin. Oncol. 2013, 31, 203–209. [Google Scholar] [CrossRef] [PubMed]
- Shi, L.; Dong, B.; Li, Z.; Lu, Y.; Ouyang, T.; Li, J.; Wang, T.; Fan, Z.; Fan, T.; Lin, B. Expression of ER-α36, a novel variant of estrogen receptor α, and resistance to tamoxifen treatment in breast cancer. J. Clin. Oncol. 2009, 27, 3423. [Google Scholar] [CrossRef] [PubMed]
- Gu, Y.; Chen, T.; Lopez, E.; Wu, W.; Wang, X.; Cao, J.; Teng, L. The therapeutic target of estrogen receptor-alpha36 in estrogen-dependent tumors. J. Trans. Med. 2014, 12, 16. [Google Scholar] [CrossRef] [PubMed]
- Chantalat, E.; Boudou, F.; Laurell, H.; Palierne, G.; Houtman, R.; Melchers, D.; Rochaix, P.; Filleron, T.; Stella, A.; Burlet-Schiltz, O. The AF-1-deficient estrogen receptor ERα46 isoform is frequently expressed in human breast tumors. Breast Cancer Res. 2016, 18, 123. [Google Scholar] [CrossRef]
- Subik, K.; Lee, J.F.; Baxter, L.; Strzepek, T.; Costello, D.; Crowley, P.; Xing, L.; Hung, M.C.; Bonfiglio, T.; Hicks, D.G.; et al. The Expression Patterns of ER, PR, HER2, CK5/6, EGFR, Ki-67 and AR by Immunohistochemical Analysis in Breast Cancer Cell Lines. Breast Cancer 2010, 4, 35–41. [Google Scholar] [CrossRef]
- Hayashi, S.; Hajiro-Nakanishi, K.; Makino, Y.; Eguchi, H.; Yodoi, J.; Tanaka, H. Functional modulation of estrogen receptor by redox state with reference to thioredoxin as a mediator. Nucleic Acids Res. 1997, 25, 4035–4040. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Tokuda, E.; Seino, Y.; Arakawa, A.; Saito, M.; Kasumi, F.; Hayashi, S.; Yamaguchi, Y. Estrogen receptor-alpha directly regulates sensitivity to paclitaxel in neoadjuvant chemotherapy for breast cancer. Breast Cancer Res. Treat. 2012, 133, 427–436. [Google Scholar] [CrossRef]
| Characteristics | Patients N (%) | Characteristics | Patients N (%) |
|---|---|---|---|
| Age | Ki67 LI (%) | ||
| <50 | 25(38) | <14 | 44 (68) |
| ≥50 | 40(62) | ≥14 | 21 (32) |
| Stage1 | Histological grade | ||
| I | 30 (46) | 1 | 20 (31) |
| II | 29 (45) | 2 | 34 (52) |
| III | 6 (9) | 3 | 11 (17) |
| Lymph node metastasis | Adjuvant chemotherapy | ||
| Positive | 27 (42) | + | 19 (29) |
| Negative | 38 (58) | - | 46 (71) |
| Characteristics | No. (%) of Patients | p Value 1 | |
|---|---|---|---|
| No Recurrence | Relapse | ||
| (n = 44) | (n = 21) | ||
| ER Allred score | |||
| <6 | 21 (32) | 12 (18) | |
| ≥6 | 21 (35) | 9 (14) | 0.477 |
| PR Allred score | |||
| <6 | 27 (42) | 16 (25) | |
| ≥6 | 17 (26) | 5 (8) | 0.237 |
| Ki67 LI (%) | |||
| <14 | 32 (49) | 13 (20) | |
| ≥14 | 12 (18) | 8 (12) | 0.377 |
| ER H-score | |||
| <110 | 13 (20) | 9 (14) | |
| ≥110 | 31 (48) | 12 (18) | 0.289 |
| PR H-score | |||
| <100 | 20 (31) | 16 (25) | |
| ≥100 | 24 (37) | 5 (8) | 0.02 |
| total ERα PIDs score | |||
| <72.5 | 29 (45) | 29 (29) | |
| ≥72.5 | 15 (23) | 2 (3) | 0.035 |
| nuclear ERα PIDs score | |||
| <48 | 26 (40) | 19 (29) | |
| ≥48 | 18 (28) | 2 (3) | 0.01 |
| extranuclear ERα PIDs score | |||
| <28 | 33 (51) | 20 (31) | |
| ≥28 | 11 (17) | 1 (2) | 0.049 |
| ERα ENR | |||
| <0.5 | 25 (38) | 5 (8) | |
| ≥0.5 | 19 (29) | 16 (25) | 0.013 |
| Characteristics | Univariate | Multivariate | ||||
|---|---|---|---|---|---|---|
| Hazard Ratio | 95% CI | p Value | Hazard Ratio | 95% CI | p Value | |
| Lymph node metastasis | ||||||
| N ≥ 1/N0 | 3.5 | 1.5–9.2 | 0.005 | 3.2 | 1.3–8.7 | 0.009 |
| ER status (A score) | ||||||
| <6/≥6 | 1.4 | 0.6–3.5 | 0.407 | |||
| PR status (A score) | ||||||
| <6/≥6 | 2.2 | 0.8–6.7 | 0.115 | |||
| Ki67 IL (%) | ||||||
| ≥14/<14 | 1.5 | 0.6–3.5 | 0.4 | |||
| ER H-score | ||||||
| <110/≥110 | 1.6 | 0.6–3.8 | 0.307 | |||
| PR H-score | ||||||
| <100/≥100 | 3.5 | 1.4–10.8 | 0.008 | 3.9 | 1.5–12.5 | 0.006 |
| total ERα PIDs score | ||||||
| <72.5/≥72.5 | 3.8 | 1.1–24.1 | 0.031 | |||
| nuclear ERα PIDs score | ||||||
| <48/≥48 | 4.8 | 1.4–30 | 0.01 | |||
| extra-nuclear ERα PIDs score | ||||||
| <28/≥28 | 4.9 | 1–87.3 | 0.048 | |||
| ERα ENR | ||||||
| ≥0.5/<0.5 | 3.6 | 1.4–11 | 0.007 | 3.8 | 1.4–11.8 | 0.006 |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Guo, Z.; Tada, H.; Kitamura, N.; Hamada, Y.; Miyashita, M.; Harada-Shoji, N.; Sato, A.; Hamanaka, Y.; Tsuboi, K.; Harada, N.; et al. Automated Quantification of Extranuclear ERα Using Phosphor-Integrated Dots for Predicting Endocrine Therapy Resistance in HR+/HER2− Breast Cancer. Cancers 2019, 11, 526. https://doi.org/10.3390/cancers11040526
Guo Z, Tada H, Kitamura N, Hamada Y, Miyashita M, Harada-Shoji N, Sato A, Hamanaka Y, Tsuboi K, Harada N, et al. Automated Quantification of Extranuclear ERα Using Phosphor-Integrated Dots for Predicting Endocrine Therapy Resistance in HR+/HER2− Breast Cancer. Cancers. 2019; 11(4):526. https://doi.org/10.3390/cancers11040526
Chicago/Turabian StyleGuo, Zhaorong, Hiroshi Tada, Narufumi Kitamura, Yoh Hamada, Minoru Miyashita, Narumi Harada-Shoji, Akiko Sato, Yohei Hamanaka, Kouki Tsuboi, Nobuhisa Harada, and et al. 2019. "Automated Quantification of Extranuclear ERα Using Phosphor-Integrated Dots for Predicting Endocrine Therapy Resistance in HR+/HER2− Breast Cancer" Cancers 11, no. 4: 526. https://doi.org/10.3390/cancers11040526
APA StyleGuo, Z., Tada, H., Kitamura, N., Hamada, Y., Miyashita, M., Harada-Shoji, N., Sato, A., Hamanaka, Y., Tsuboi, K., Harada, N., Takano-Kasuya, M., Okada, H., Nakano, Y., Ohuchi, N., Hayashi, S.-i., Ishida, T., & Gonda, K. (2019). Automated Quantification of Extranuclear ERα Using Phosphor-Integrated Dots for Predicting Endocrine Therapy Resistance in HR+/HER2− Breast Cancer. Cancers, 11(4), 526. https://doi.org/10.3390/cancers11040526





