Proteomic Analysis Implicates Vimentin in Glioblastoma Cell Migration
Abstract
1. Introduction
2. Results
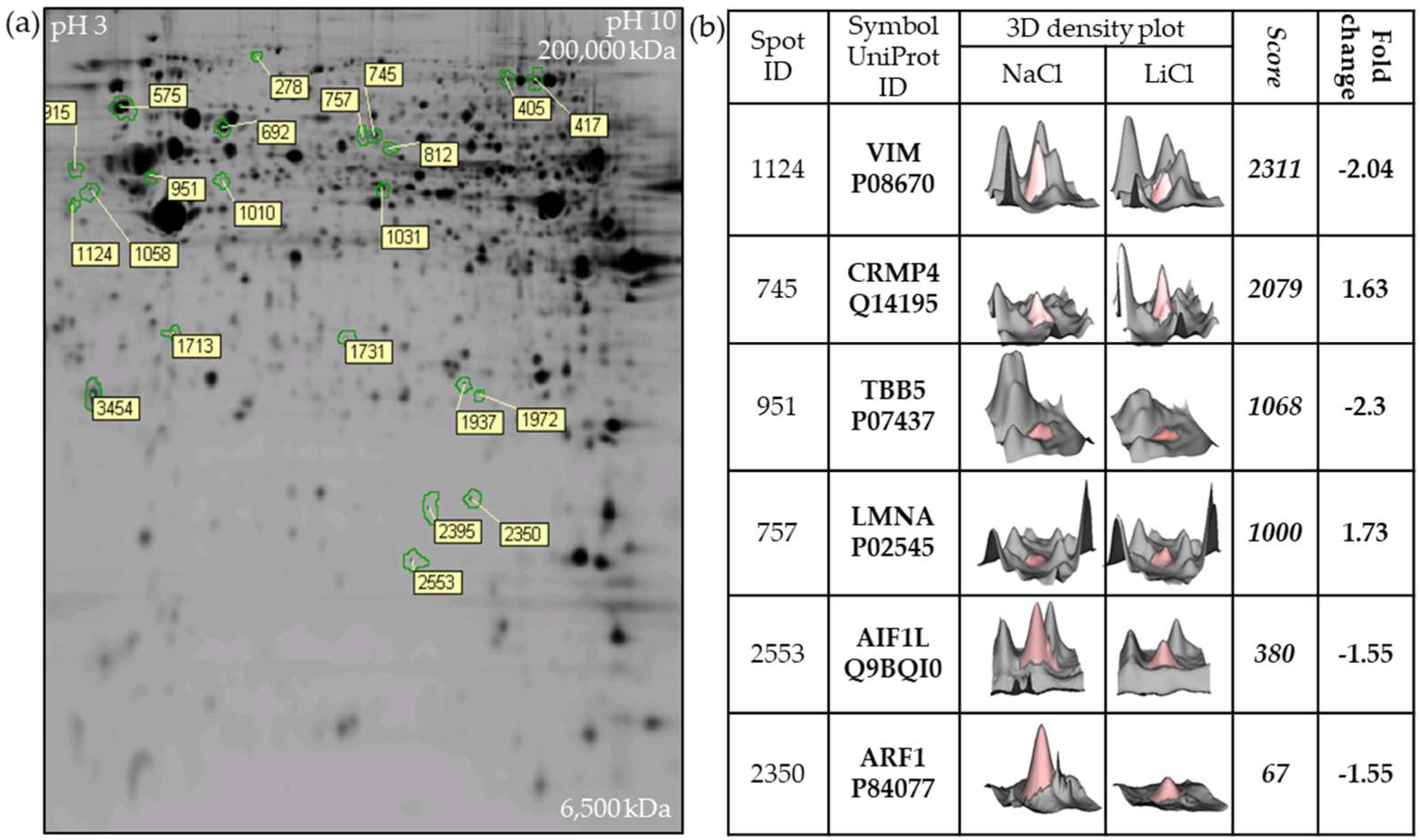
2.1. Two-Dimensional Fluorescence DIGE Proteomics Analysis Identifies Vimentin Downregulation by LiCl in GBM Cells
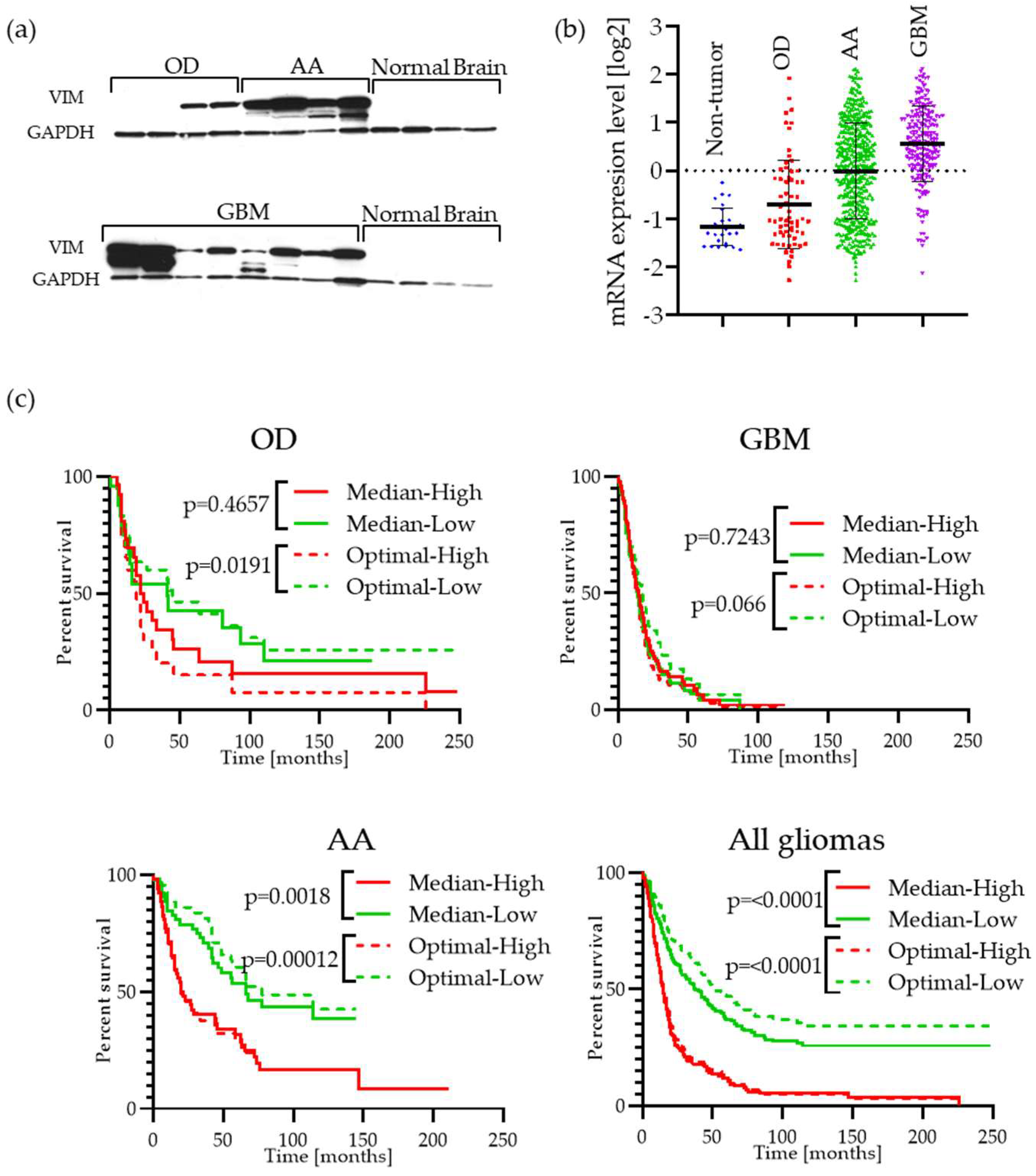
2.2. Vimentin Is Highly Expressed in GBM Patient Tumor Specimens and Is Associated with Cell Migration
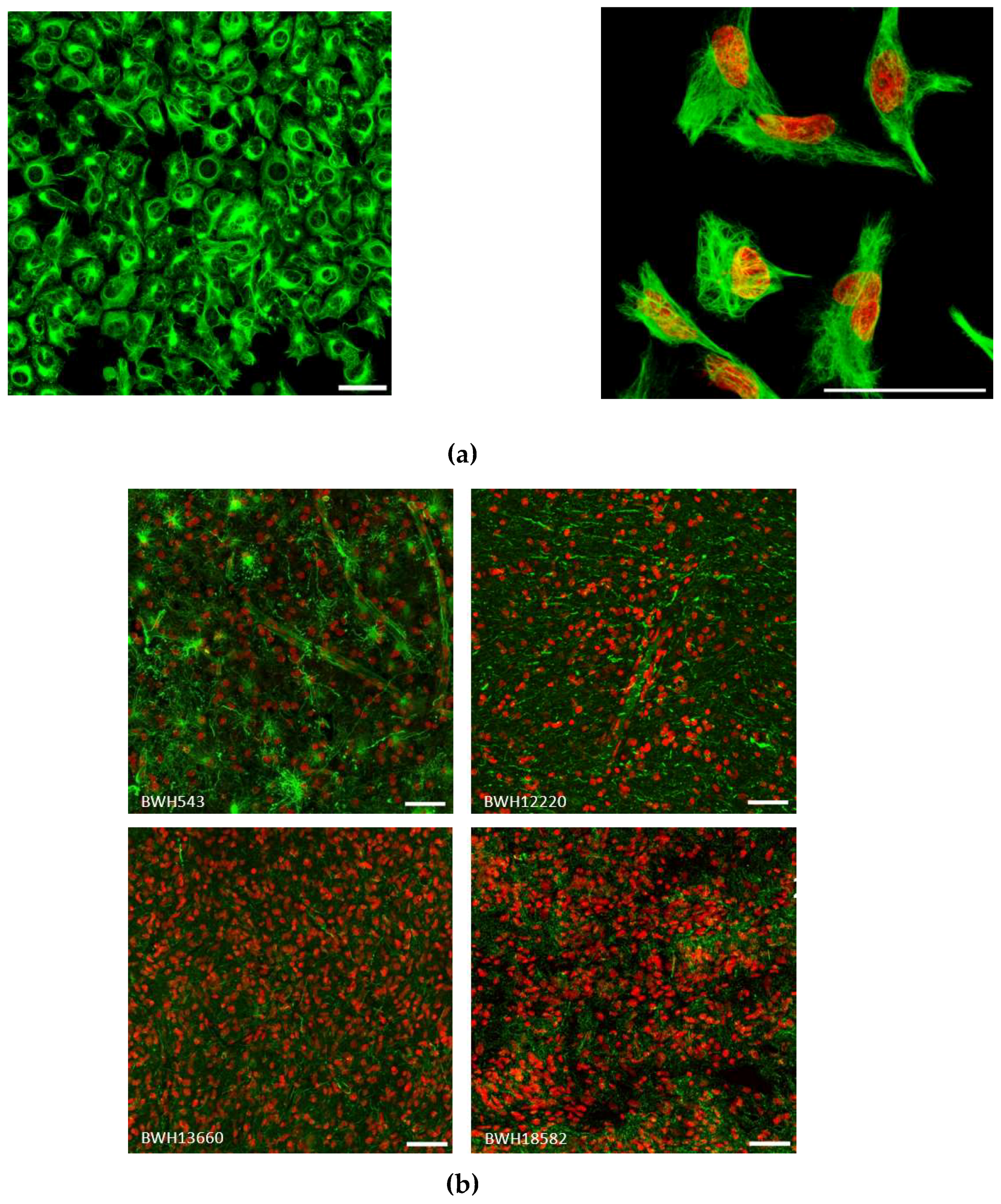
2.3. Vimentin Immunofluorescence Staining in GBM cells and in Patient Specimens
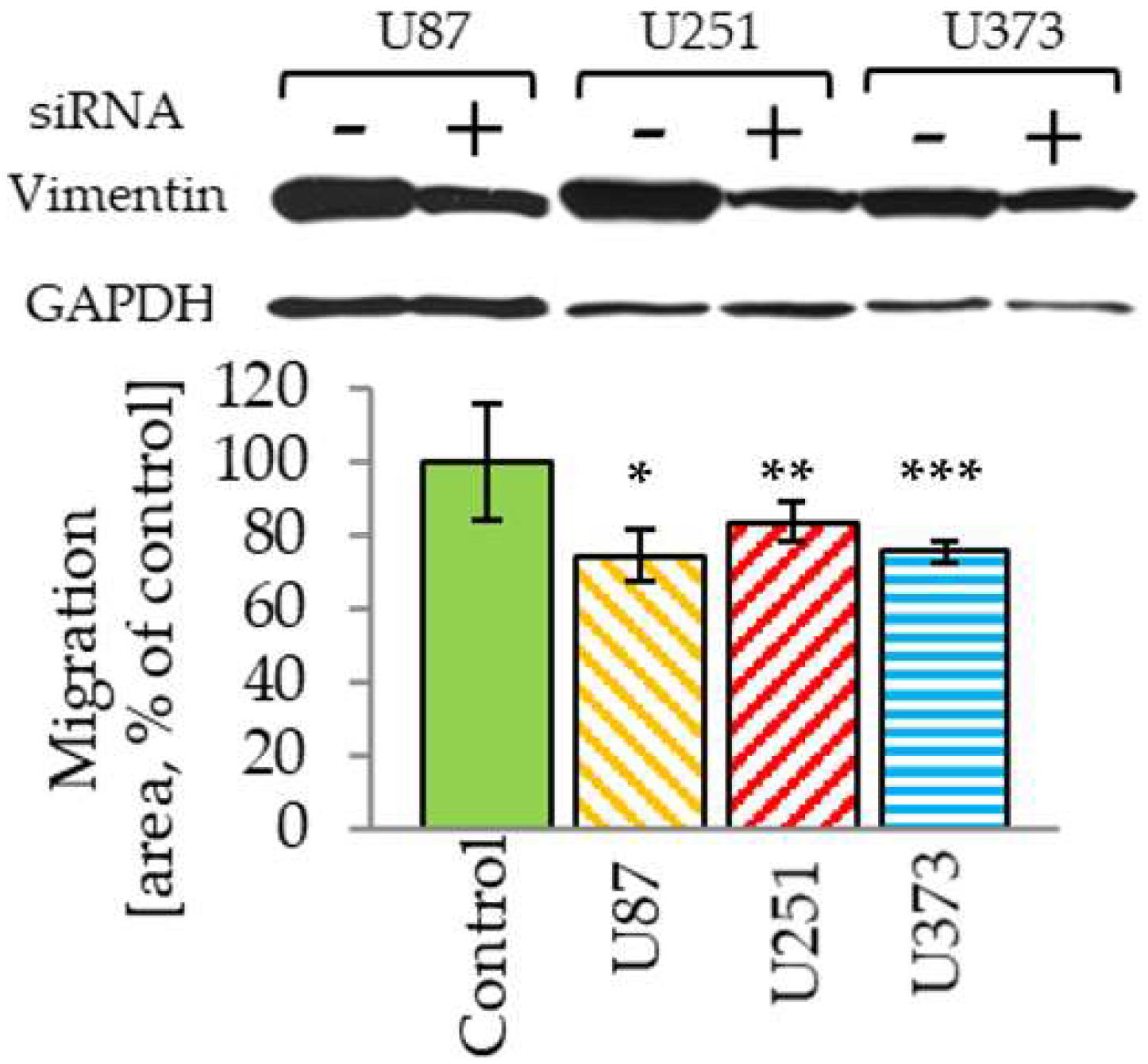
2.4. Vimentin Knockdown Reduces GBM Cell Migration
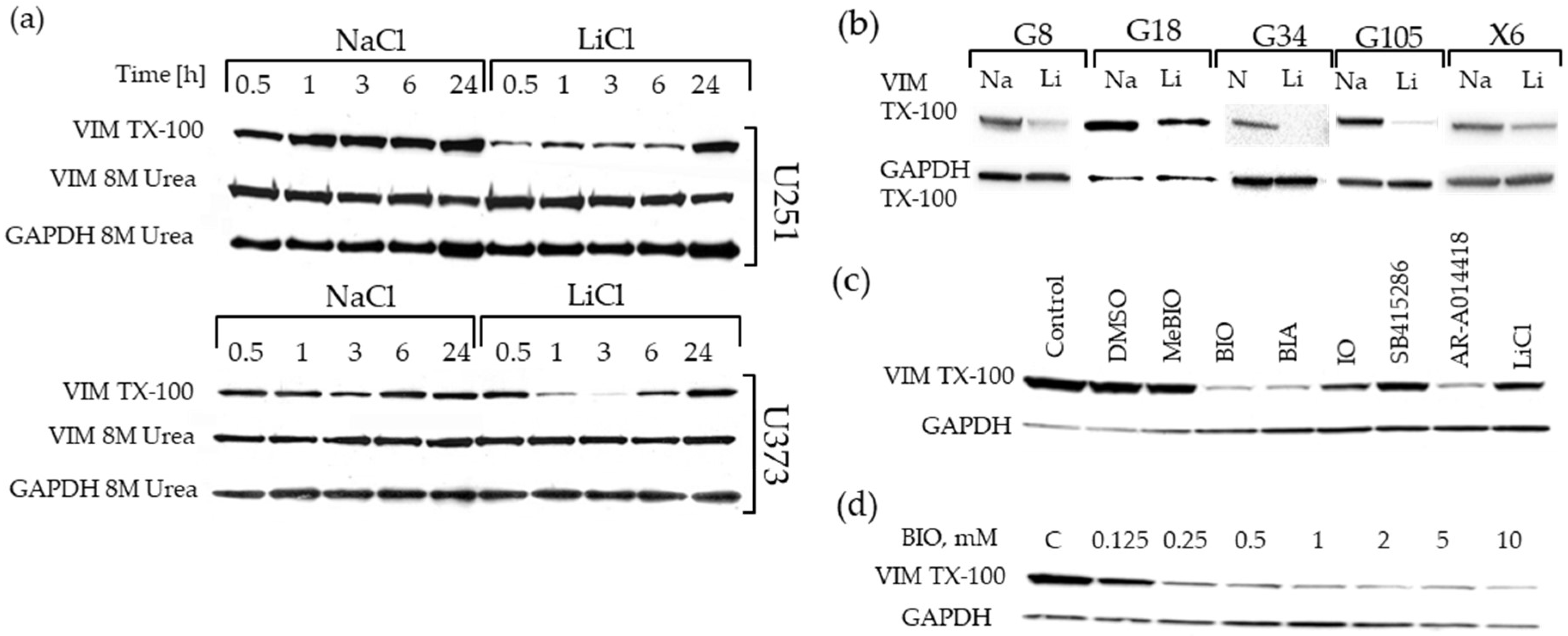
2.5. Downregulation of the Triton-x-100 Soluble Vimentin Fraction in GBM Cell Lines by GSK-3 Inhibitors
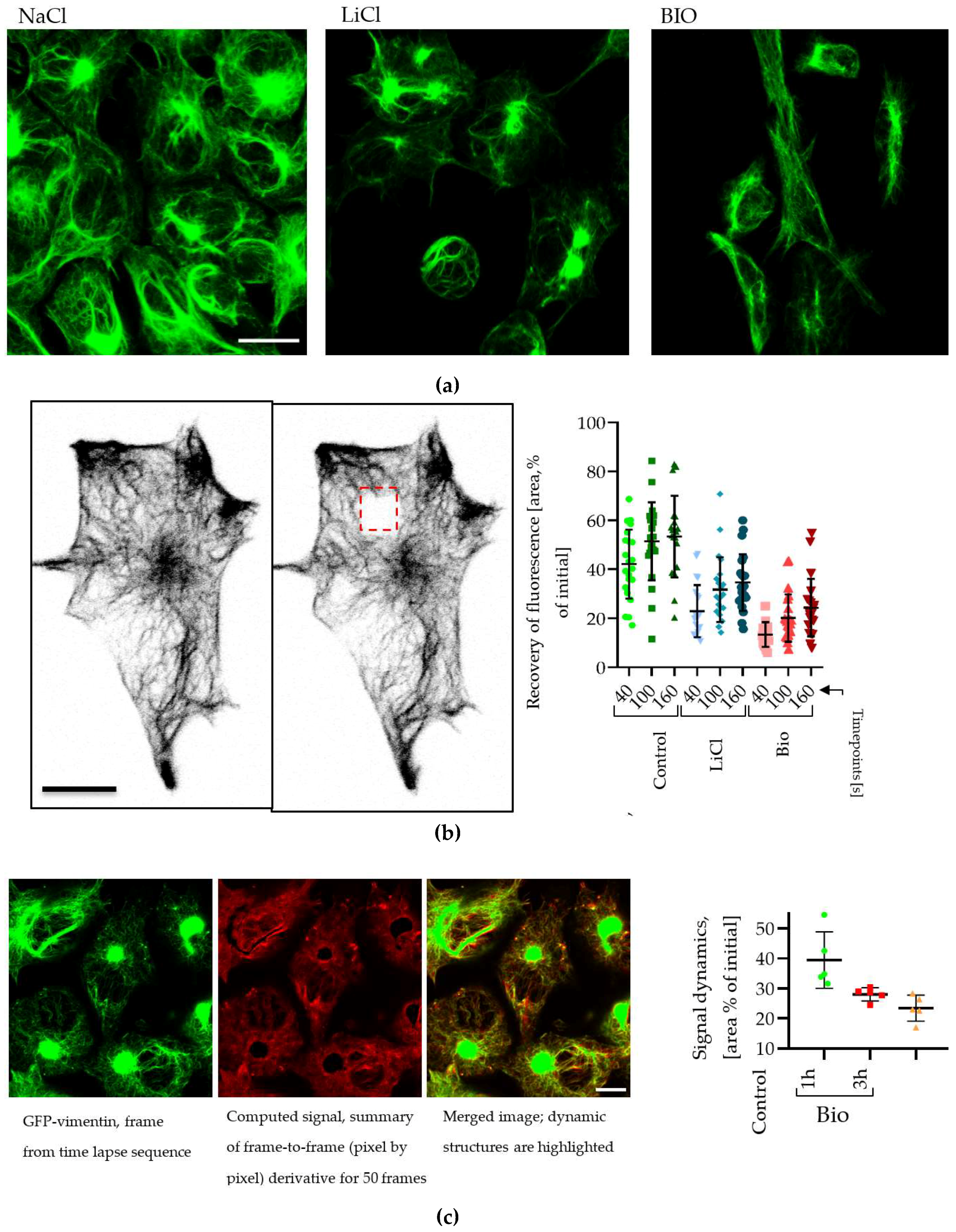
2.6. Changes in Vimentin Dynamics in LiCl and BIO Treated GBM cells
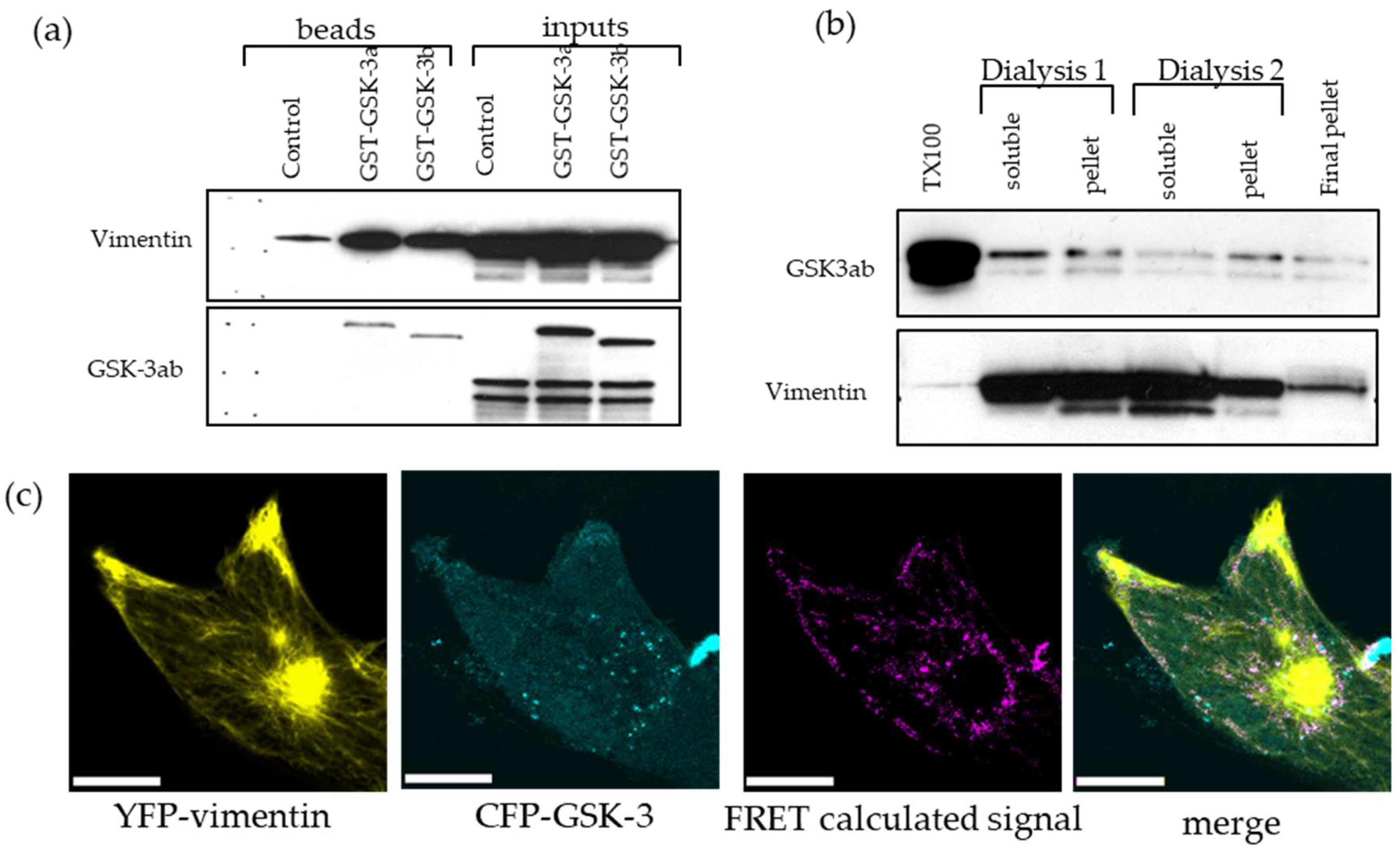
2.7. GSK-3 and Vimentin Colocalize in GBM Cells
3. Discussion
4. Materials and Methods
4.1. Cell Lines, Tissue Samples and Chemicals
4.2. Western Blotting
4.3. Cell Migration Assays
4.4. Two-Dimensional Fluorescence Difference Gel Electrophoresis
4.5. Fluorescence Recovery After Photobleaching (FRAP) Assay
4.6. GST Pull-Down Assay
4.7. Digital Data Resources
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Conflicts of Interest
References
- Sattiraju, A.; Sai, K.K.S.; Mintz, A. Glioblastoma Stem Cells and Their Microenvironment. Adv. Exp. Med. Biol. 2017, 1041, 119–140. [Google Scholar]
- Bellail, A.C.; Hunter, S.B.; Brat, D.J.; Tan, C.; Van Meir, E.G. Microregional extracellular matrix heterogeneity in brain modulates glioma cell invasion. Int. J. Biochem. Cell Biol. 2004, 36, 1046–1069. [Google Scholar] [CrossRef] [PubMed]
- Roomi, M.W.; Kalinovsky, T.; Rath, M.; Niedzwiecki, A. Modulation of MMP-2 and MMP-9 secretion by cytokines, inducers and inhibitors in human glioblastoma T-98G cells. Oncol. Rep. 2017, 37, 1907–1913. [Google Scholar] [CrossRef] [PubMed]
- Matias, D.; Predes, D.; Niemeyer Filho, P.; Lopes, M.C.; Abreu, J.G.; Lima, F.R.S.; Moura Neto, V. Microglia-glioblastoma interactions: New role for Wnt signaling. Biochim. Biophys. Acta Rev. Cancer 2017, 1868, 333–340. [Google Scholar] [CrossRef] [PubMed]
- Turaga, S.M.; Lathia, J.D. Adhering towards tumorigenicity: Altered adhesion mechanisms in glioblastoma cancer stem cells. CNS Oncol. 2016, 5, 251–259. [Google Scholar] [CrossRef] [PubMed]
- Kłopocka, W.; Korczyński, J.; Pomorski, P. Cytoskeleton and nucleotide signaling in glioma C6 cells. Adv. Exp. Med. Biol. 2013, 986, 103–119. [Google Scholar]
- Schoumacher, M.; Goldman, R.D.; Louvard, D.; Vignjevic, D.M. Actin, microtubules, and vimentin intermediate filaments cooperate for elongation of invadopodia. J. Cell Biol. 2010, 189, 541–556. [Google Scholar] [CrossRef]
- Nowicki, M.O.; Dmitrieva, N.; Stein, A.M.; Cutter, J.L.; Godlewski, J.; Saeki, Y.; Nita, M.; Berens, M.E.; Sander, L.M.; Newton, H.B.; et al. Lithium inhibits invasion of glioma cells; possible involvement of glycogen synthase kinase-3. Neuro-Oncology 2008, 10, 690–699. [Google Scholar] [CrossRef]
- Mendez, M.G.; Kojima, S.-I.; Goldman, R.D. Vimentin induces changes in cell shape, motility, and adhesion during the epithelial to mesenchymal transition. FASEB J. 2010, 24, 1838–1851. [Google Scholar] [CrossRef]
- Lowery, J.; Kuczmarski, E.R.; Herrmann, H.; Goldman, R.D. Intermediate Filaments Play a Pivotal Role in Regulating Cell Architecture and Function. J. Biol. Chem. 2015, 290, 17145–17153. [Google Scholar] [CrossRef]
- Köster, S.; Weitz, D.A.; Goldman, R.D.; Aebi, U.; Herrmann, H. Intermediate filament mechanics in vitro and in the cell: From coiled coils to filaments, fibers and networks. Curr. Opin. Cell Biol. 2015, 32, 82–91. [Google Scholar] [CrossRef] [PubMed]
- Cole, A.R.; Knebel, A.; Morrice, N.A.; Robertson, L.A.; Irving, A.J.; Connolly, C.N.; Sutherland, C. GSK-3 phosphorylation of the Alzheimer epitope within collapsin response mediator proteins regulates axon elongation in primary neurons. J. Biol. Chem. 2004, 279, 50176–50180. [Google Scholar] [CrossRef] [PubMed]
- Nagini, S.; Sophia, J.; Mishra, R. Glycogen synthase kinases: Moonlighting proteins with theranostic potential in cancer. In Seminars in Cancer Biology; Academic Press: Cambridge, MA, USA, 2018. [Google Scholar]
- Patel, P.; Woodgett, J.R. Glycogen Synthase Kinase 3: A Kinase for All Pathways? Curr. Top. Dev. Biol. 2017, 123, 277–302. [Google Scholar] [PubMed]
- Tejeda-Muñoz, N.; Robles-Flores, M. Glycogen synthase kinase 3 in Wnt signaling pathway and cancer. IUBMB Life 2015, 67, 914–922. [Google Scholar] [CrossRef]
- Zhu, Q.-S.; Rosenblatt, K.; Huang, K.-L.; Lahat, G.; Brobey, R.; Bolshakov, S.; Nguyen, T.; Ding, Z.; Belousov, R.; Bill, K.; et al. Vimentin is a novel AKT1 target mediating motility and invasion. Oncogene 2011, 30, 457–470. [Google Scholar] [CrossRef]
- Williams, S.P.; Nowicki, M.O.; Liu, F.; Press, R.; Godlewski, J.; Abdel-Rasoul, M.; Kaur, B.; Fernandez, S.A.; Chiocca, E.A.; Lawler, S.E. Indirubins decrease glioma invasion by blocking migratory phenotypes in both the tumor and stromal endothelial cell compartments. Cancer Res. 2011, 71, 5374–5380. [Google Scholar] [CrossRef] [PubMed]
- Majewska, E.; Szeliga, M. AKT/GSK3β Signaling in Glioblastoma. Neurochem. Res. 2017, 42, 918–924. [Google Scholar] [CrossRef]
- Cheng, F.; Shen, Y.; Mohanasundaram, P.; Lindström, M.; Ivaska, J.; Ny, T.; Eriksson, J.E. Vimentin coordinates fibroblast proliferation and keratinocyte differentiation in wound healing via TGF-β-Slug signaling. Proc. Natl. Acad. Sci. USA 2016, 113, E4320–E4327. [Google Scholar] [CrossRef]
- Menko, A.S.; Bleaken, B.M.; Libowitz, A.A.; Zhang, L.; Stepp, M.A.; Walker, J.L. A central role for vimentin in regulating repair function during healing of the lens epithelium. Mol. Biol. Cell 2014, 25, 776–790. [Google Scholar] [CrossRef] [PubMed]
- Lin, L.; Wang, G.; Ming, J.; Meng, X.; Han, B.; Sun, B.; Cai, J.; Jiang, C. Analysis of expression and prognostic significance of vimentin and the response to temozolomide in glioma patients. Tumour Biol. J. Int. Soc. Oncodev. Biol. Med. 2016, 37, 15333–15339. [Google Scholar] [CrossRef]
- Hothorn, T.; Lausen, B. On the exact distribution of maximally selected rank statistics. Comput. Stat. Data Anal. 2003, 43, 121–137. [Google Scholar] [CrossRef]
- Ádám, C.; Fekete, A.; Bőgel, G.; Németh, Z.; Tőkési, N.; Ovádi, J.; Liliom, K.; Pesti, S.; Geiszt, M.; Buday, L. Accumulation of the PX domain mutant Frank-ter Haar syndrome protein Tks4 in aggresomes. Cell Commun. Signal. 2015, 13, 33. [Google Scholar] [CrossRef] [PubMed]
- Uhlén, M.; Fagerberg, L.; Hallström, B.M.; Lindskog, C.; Oksvold, P.; Mardinoglu, A.; Sivertsson, Å.; Kampf, C.; Sjöstedt, E.; Asplund, A.; et al. Proteomics. Tissue-based map of the human proteome. Science 2015, 347, 1260419. [Google Scholar] [CrossRef] [PubMed]
- Jope, R.S. Lithium and GSK-3: One inhibitor, two inhibitory actions, multiple outcomes. Trends Pharmacol. Sci. 2003, 24, 441–443. [Google Scholar] [CrossRef]
- Meijer, L.; Skaltsounis, A.-L.; Magiatis, P.; Polychronopoulos, P.; Knockaert, M.; Leost, M.; Ryan, X.P.; Vonica, C.A.; Brivanlou, A.; Dajani, R.; et al. GSK-3-selective inhibitors derived from Tyrian purple indirubins. Chem. Biol. 2003, 10, 1255–1266. [Google Scholar] [CrossRef] [PubMed]
- Gnad, F.; Ren, S.; Cox, J.; Olsen, J.V.; Macek, B.; Oroshi, M.; Mann, M. PHOSIDA (phosphorylation site database): Management, structural and evolutionary investigation, and prediction of phosphosites. Genome Biol. 2007, 8, R250. [Google Scholar] [CrossRef] [PubMed]
- Boyden, S. The chemotactic effect of mixtures of antibody and antigen on polymorphonuclear leucocytes. J. Exp. Med. 1962, 115, 453–466. [Google Scholar] [CrossRef]
- Chen, W.; Wong, C.; Vosburgh, E.; Levine, A.J.; Foran, D.J.; Xu, E.Y. High-throughput image analysis of tumor spheroids: A user-friendly software application to measure the size of spheroids automatically and accurately. J. Vis. Exp. 2014, 89, 51639. [Google Scholar] [CrossRef] [PubMed]
- Blundon, M.; Ganesan, V.; Redler, B.; Van, P.T.; Minden, J.S. Two-Dimensional Difference Gel Electrophoresis. Methods Mol. Biol. 2019, 1855, 229–247. [Google Scholar]
- Yoon, M.; Moir, R.D.; Prahlad, V.; Goldman, R.D. Motile properties of vimentin intermediate filament networks in living cells. J. Cell Biol. 1998, 143, 147–157. [Google Scholar] [CrossRef] [PubMed]

© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Nowicki, M.O.; Hayes, J.L.; Chiocca, E.A.; Lawler, S.E. Proteomic Analysis Implicates Vimentin in Glioblastoma Cell Migration. Cancers 2019, 11, 466. https://doi.org/10.3390/cancers11040466
Nowicki MO, Hayes JL, Chiocca EA, Lawler SE. Proteomic Analysis Implicates Vimentin in Glioblastoma Cell Migration. Cancers. 2019; 11(4):466. https://doi.org/10.3390/cancers11040466
Chicago/Turabian StyleNowicki, Michal O., Josie L. Hayes, E. Antonio Chiocca, and Sean E. Lawler. 2019. "Proteomic Analysis Implicates Vimentin in Glioblastoma Cell Migration" Cancers 11, no. 4: 466. https://doi.org/10.3390/cancers11040466
APA StyleNowicki, M. O., Hayes, J. L., Chiocca, E. A., & Lawler, S. E. (2019). Proteomic Analysis Implicates Vimentin in Glioblastoma Cell Migration. Cancers, 11(4), 466. https://doi.org/10.3390/cancers11040466




