Assessment of a High Sensitivity Method for Identification of IDH1 R132x Mutations in Tumors and Plasma of Intrahepatic Cholangiocarcinoma Patients
Abstract
1. Introduction
2. Results
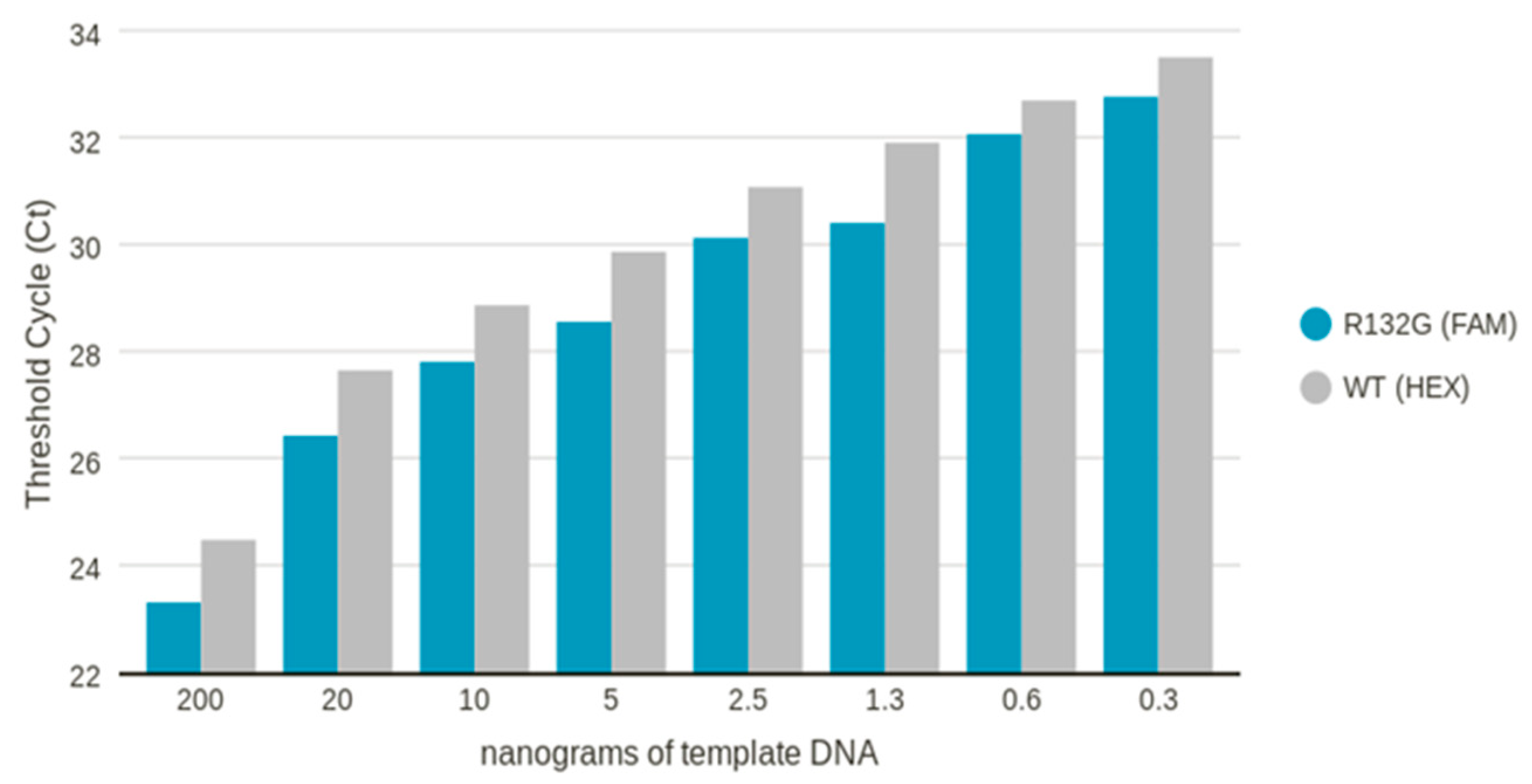
2.1. Sensitivity of IDH1 R132x qPCR
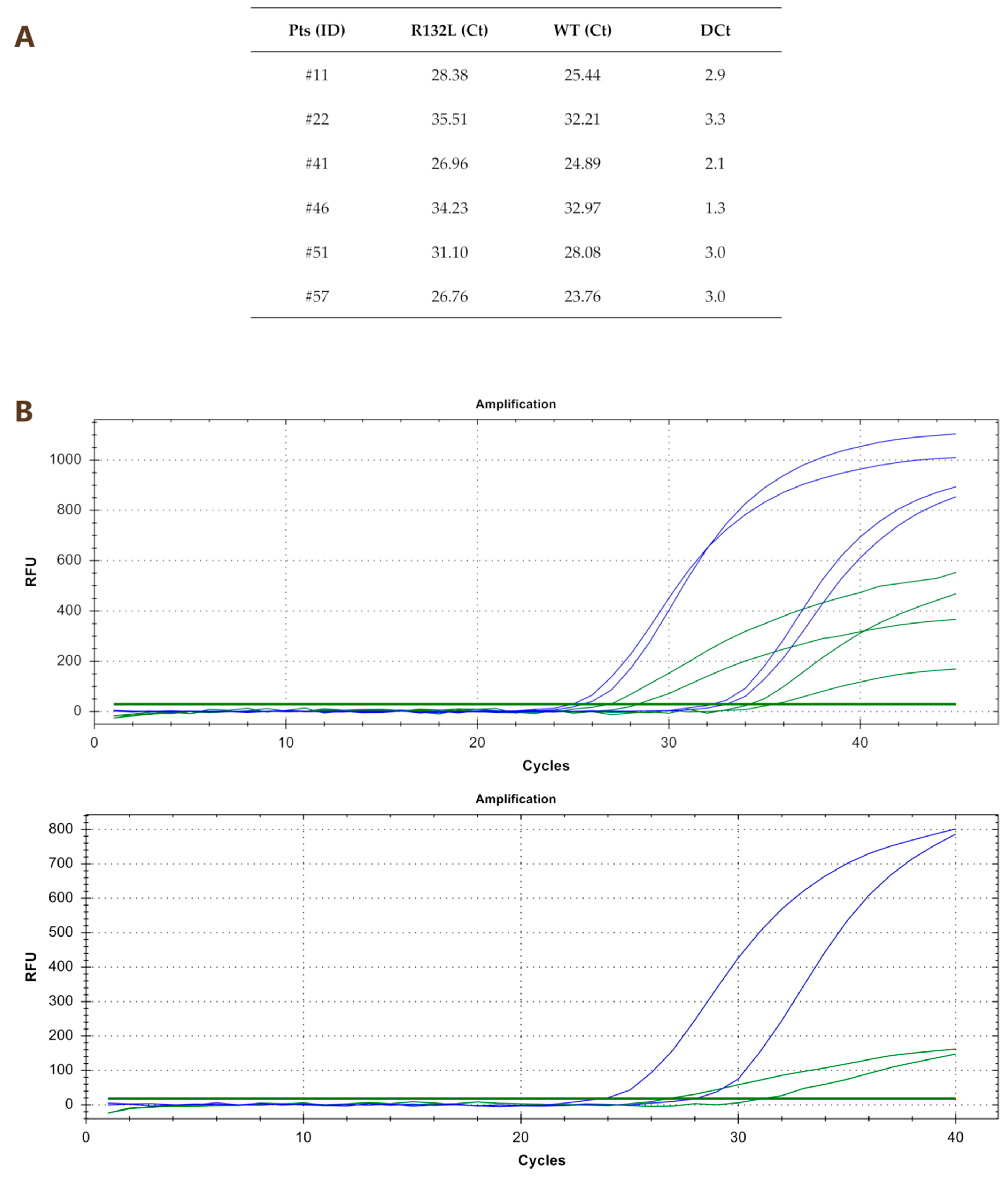
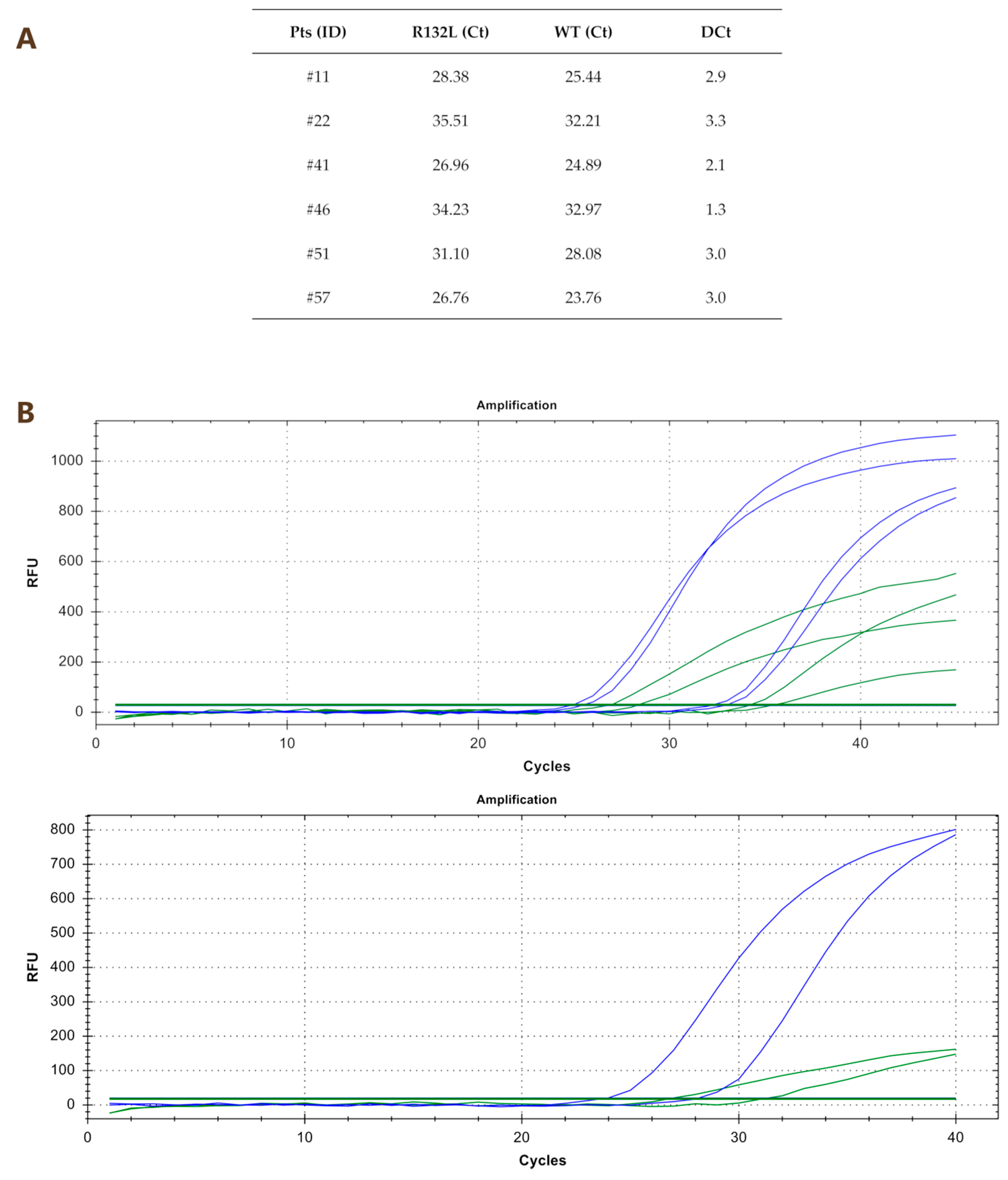
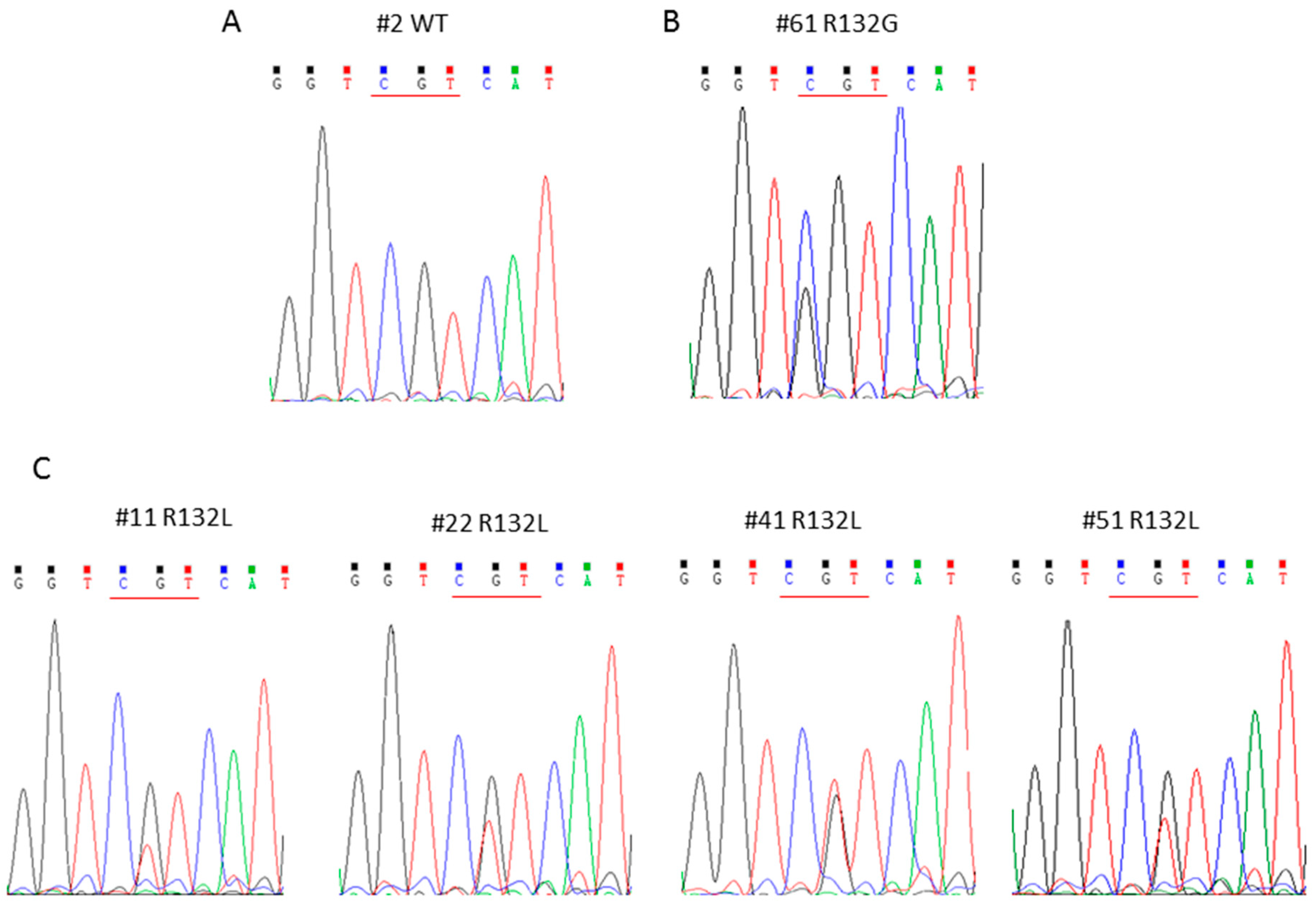
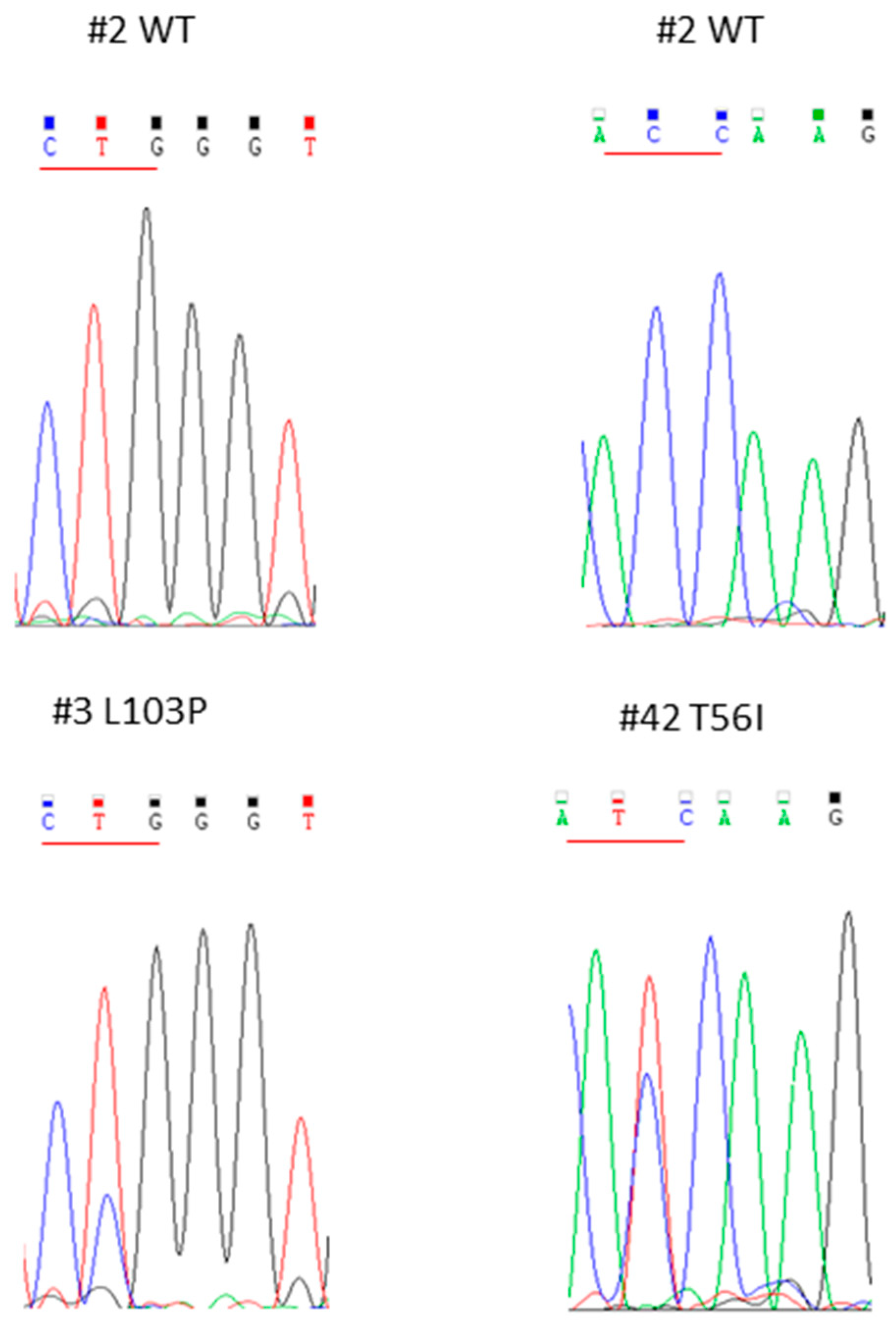
2.2. Comparison between Quantitative PCR (qPCR) and Nested PCR for the Identification of IDH1R132x Mutations
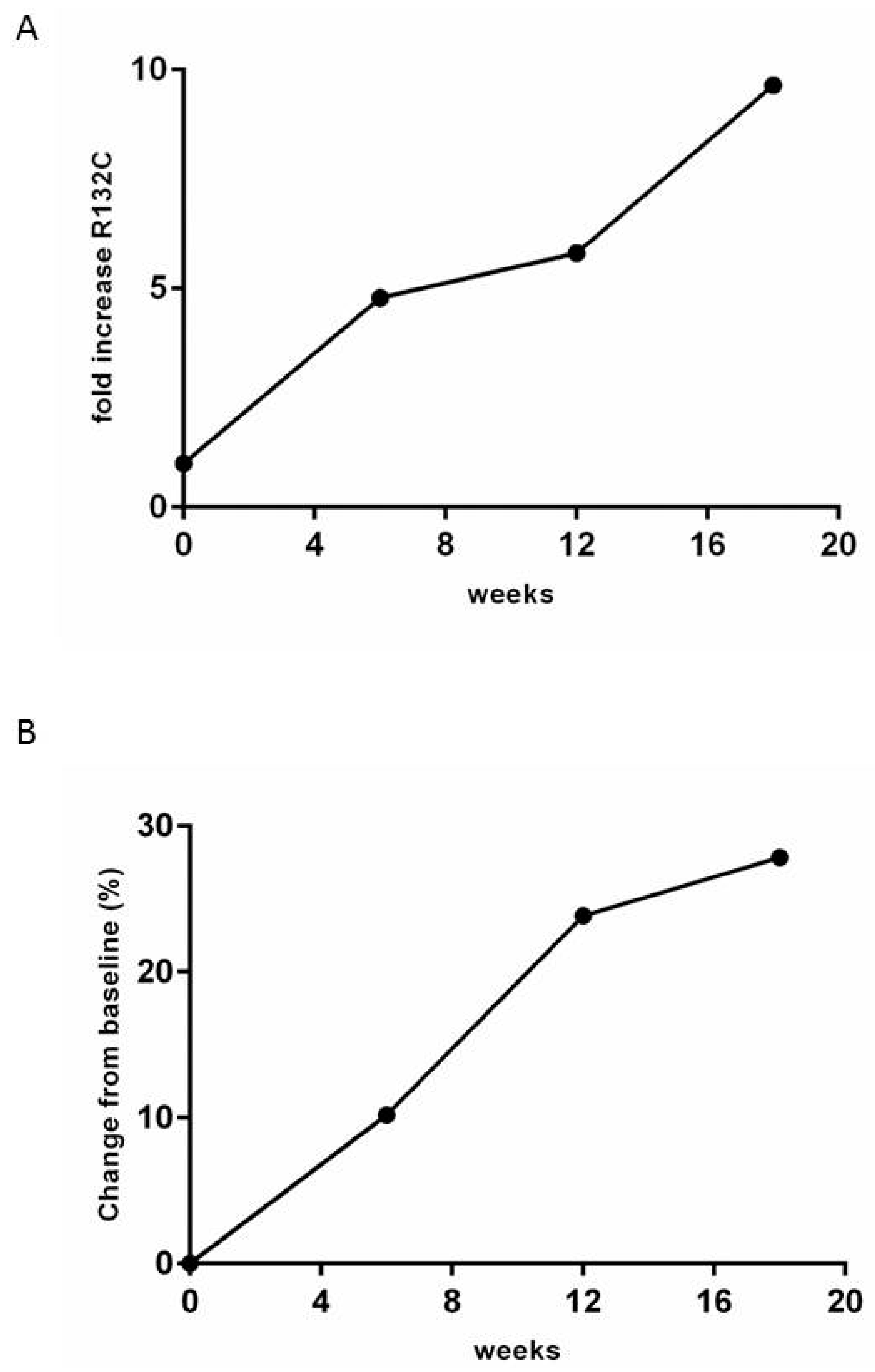
2.3. cfIDH1m R132x is a Potential Prognostic Marker in ICC in Plasma Samples
3. Discussion
4. Materials and Methods
4.1. Patients
4.2. DNA Extraction from FFPE Tissues
4.3. DNA Isolation from Plasma Samples
4.4. IDH1 Mutational Analysis
4.5. Analysis of IDH1 PCR Data
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Alvaro, D.; Crocetti, E.; Ferretti, S.; Bragazzi, M.C.; Capocaccia, R.; The AISF “Cholangiocarcinoma” Committee. Descriptive epidemiology of cholangiocarcinoma in Italy. Dig. Liver Dis. 2010, 42, 490–495. [Google Scholar] [CrossRef] [PubMed]
- Cho, K.M.; Park, H.; Oh, D.Y.; Kim, T.Y.; Lee, K.H.; Han, S.W.; Im, S.A.; Bang, Y.J. Neutrophil-to-lymphocyte ratio, platelet-to-lymphocyte ratio, and their dynamic changes during chemotherapy is useful to predict a more accurate prognosis of advanced biliary tract cancer. Oncotarget 2017, 8, 2329–2341. [Google Scholar] [CrossRef] [PubMed]
- Rizvi, S.; Gores, G.J. Pathogenesis, diagnosis, and management of cholangiocarcinoma. Gastroenterology 2013, 145, 1215–1229. [Google Scholar] [CrossRef]
- de Jong, M.C.; Hong, S.M.; Augustine, M.M.; Goggins, M.G.; Wolfgang, C.L.; Hirose, K.; Schulick, R.D.; Choti, M.A.; Anders, R.A.; Pawlik, T.M. Hilar cholangiocarcinoma: Tumor depth as a predictor of outcome. Arch. Surg. 2011, 146, 697–703. [Google Scholar] [CrossRef] [PubMed]
- Ni, Q.; Shen, W.; Zhang, M.; Yang, C.; Cai, W.; Wu, M.; Yang, J. Prognostic analysis of radical resection for intrahepatic cholangiocarcinoma: A retrospective cohort study. Oncotarget 2017, 8, 75627–75637. [Google Scholar] [CrossRef] [PubMed]
- Jain, A.; Javle, M. Molecular profiling of biliary tract cancer: A target rich disease. J. Gastrointest. Oncol. 2016, 7, 797–803. [Google Scholar] [CrossRef] [PubMed]
- Lee, H.; Ross, J.S. The potential role of comprehensive genomic profiling to guide targeted therapy for patients with biliary cancer. Ther. Adv. Gastroenterol. 2017, 10, 507–520. [Google Scholar] [CrossRef] [PubMed]
- Ong, C.K.; Subimerb, C.; Pairojkul, C.; Wongkham, S.; Cutcutache, I.; Yu, W.; McPherson, J.R.; Allen, G.E.; Ng, C.C.; Wong, B.H.; et al. Exome sequencing of liver fluke-associated cholangiocarcinoma. Nat. Genet. 2012, 44, 690–693. [Google Scholar] [CrossRef] [PubMed]
- Sia, D.; Tovar, V.; Moeini, A.; Llovet, J.M. Intrahepatic cholangiocarcinoma: Pathogenesis and rationale for molecular therapies. Oncogene 2013, 32, 4861–4870. [Google Scholar] [CrossRef] [PubMed]
- Peraldo Neia, C.; Cavalloni, G.; Balsamo, A.; Venesio, T.; Napoli, F.; Sassi, F.; Martin, V.; Frattini, M.; Aglietta, M.; Leone, F. Screening for the FIG-ROS1 fusion in biliary tract carcinomas by nested PCR. Genes Chromosomes Cancer 2014, 53, 1033–1040. [Google Scholar] [CrossRef] [PubMed]
- Lombardi, P.; Marino, D.; Fenocchio, E.; Chilà, G.; Aglietta, M.; Leone, F. Emerging molecular target antagonists for the treatment of biliary tract cancer. Expert. Opin. Emerg. Drugs 2018, 23, 63–75. [Google Scholar] [CrossRef]
- Farshidfar, F.; Zheng, S.; Gingras, M.C.; Newton, Y.; Shih, J.; Robertson, A.G.; Hinoue, T.; Hoadley, K.A.; Gibb, E.A.; Roszik, J.; et al. Integrative Genomic Analysis of Cholangiocarcinoma Identifies Distinct IDH-Mutant Molecular Profiles. Cell Rep. 2017, 19, 2878–2880. [Google Scholar] [CrossRef] [PubMed]
- Raineri, S.; Mellor, J. Linking Metabolism and Epigenetics. Front. Genet. 2018, 9, 493. [Google Scholar] [CrossRef]
- Grassian, A.R.; Pagliarini, R.; Chiang, D.Y. Mutations of isocitrate dehydrogenase 1 and 2 in intrahepatic cholangiocarcinoma. Curr. Opin. Gastroenterol. 2014, 30, 295–302. [Google Scholar] [CrossRef]
- Peraldo-Neia, C.; Ostano, P.; Cavalloni, G.; Pignochino, Y.; Sangiolo, D.; De Cecco, L.; Marchesi, E.; Ribero, D.; Scarpa, A.; De Rose, A.M.; et al. Transcriptomic analysis and mutational status of IDH1 in paired primary-recurrent intrahepatic cholangiocarcinoma. BMC Genom. 2018, 19, 440. [Google Scholar] [CrossRef]
- Cercek, A.; Kemeny, N.E.; Boerner, T.; Tan, B.R.; Chou, J.F.; Gonen, M.; Boucher, T.M.; Hauser, H.; Do Richard, K.G.; Lowery, M.A.; et al. A bi-institutional phase II study of hepatic arterial infusion (HAI) with floxuridine (FUDR) and dexamethasone (Dex) combined with systemic gemcitabine and oxaliplatin (GemOx) for unresectable intrahepatic cholangiocarcinoma (ICC). J. Clin. Oncol. 2018, 36, 4092. [Google Scholar] [CrossRef]
- Wang, P.; Dong, Q.; Zhang, C.; Kuan, P.F.; Liu, Y.; Jeck, W.R.; Andersen, J.B.; Jiang, W.; Savich, G.L.; Tan, T.X.; et al. Mutations in isocitrate dehydrogenase 1 and 2 occur frequently in intrahepatic cholangiocarcinomas and share hypermethylation targets with glioblastomas. Oncogene 2013, 32, 3091–3100. [Google Scholar] [CrossRef]
- Jusakul, A.; Cutcutache, I.; Yong, C.H.; Lim, J.Q.; Huang, M.N.; Padmanabhan, N.; Nellore, V.; Kongpetch, S.; Ng, A.W.T.; Ng, L.M.; et al. Whole-Genome and Epigenomic Landscapes of Etiologically Distinct Subtypes of Cholangiocarcinoma. Cancer Discov. 2017, 7, 1116–1135. [Google Scholar] [CrossRef] [PubMed]
- Goyal, L.; Govindan, A.; Sheth, R.A.; Nardi, V.; Blaszkowsky, L.S.; Faris, J.E.; Clark, J.W.; Ryan, D.P.; Kwak, E.L.; Allen, J.N.; et al. Prognosis and Clinicopathologic Features of Patients With Advanced Stage Isocitrate Dehydrogenase (IDH) Mutant and IDH Wild-Type Intrahepatic Cholangiocarcinoma. Oncologist 2015, 20, 1019–1027. [Google Scholar] [CrossRef]
- Jiao, Y.; Pawlik, T.M.; Anders, R.A.; Selaru, F.M.; Streppel, M.M.; Lucas, D.J.; Niknafs, N.; Guthrie, V.B.; Maitra, A.; Argani, P.; et al. Exome sequencing identifies frequent inactivating mutations in BAP1, ARID1A and PBRM1 in intrahepatic cholangiocarcinomas. Nat. Genet. 2013, 45, 1470–1473. [Google Scholar] [CrossRef]
- Gao, J.; Wu, H.; Shi, X.; Huo, Z.; Zhang, J.; Liang, Z. Comparison of Next-Generation Sequencing, Quantitative PCR, and Sanger Sequencing for Mutation Profiling of EGFR, KRAS, PIK3CA and BRAF in Clinical Lung Tumors. Clin. Lab. 2016, 62, 689–696. [Google Scholar] [CrossRef] [PubMed]
- Moeini, A.; Sia, D.; Bardeesy, N.; Mazzaferro, V.; Llovet, J.M. Molecular Pathogenesis and Targeted Therapies for Intrahepatic Cholangiocarcinoma. Clin. Cancer Res. 2016, 22, 291–300. [Google Scholar] [CrossRef] [PubMed]
- Kipp, B.R.; Voss, J.S.; Kerr, S.E.; Barr Fritcher, E.G.; Graham, R.P.; Zhang, L.; Highsmith, W.E.; Zhang, J.; Roberts, L.R.; Gores, G.J.; et al. Isocitrate dehydrogenase 1 and 2 mutations in cholangiocarcinoma. Hum. Pathol. 2012, 43, 1552–1558. [Google Scholar] [CrossRef]
- Lowery, M.A.; Ptashkin, R.; Jordan, E.; Berger, M.F.; Zehir, A.; Capanu, M.; Kemeny, N.E.; O’Reilly, E.M.; El-Dika, I.; Jarnagin, W.R.; et al. Comprehensive Molecular Profiling of Intrahepatic and Extrahepatic Cholangiocarcinomas: Potential Targets for Intervention. Clin. Cancer Res. 2018, 24, 4154–4161. [Google Scholar] [CrossRef] [PubMed]
- Borger, D.R.; Tanabe, K.K.; Fan, K.C.; Lopez, H.U.; Fantin, V.R.; Straley, K.S.; Schenkein, D.P.; Hezel, A.F.; Ancukiewicz, M.; Liebman, H.M.; et al. Frequent mutation of isocitrate dehydrogenase (IDH)1 and IDH2 in cholangiocarcinoma identified through broad-based tumor genotyping. Oncologist 2012, 17, 72–79. [Google Scholar] [CrossRef]
- Saha, S.K.; Parachoniak, C.A.; Bardeesy, N. IDH mutations in liver cell plasticity and biliary cancer. Cell Cycle 2014, 13, 3176–3182. [Google Scholar] [CrossRef] [PubMed]
- Brennan, C.W.; Verhaak, R.G.; McKenna, A.; Campos, B.; Noushmehr, H.; Salama, S.R.; Zheng, S.; Chakravarty, D.; Sanborn, J.Z.; Berman, S.H.; et al. The somatic genomic landscape of glioblastoma. Cell 2013, 155, 462–477. [Google Scholar] [CrossRef]
| IDH1 Status | 3000 Copies (Ct) | 300 Copies (Ct) | 30 Copies (Ct) | 3 Copies (Ct) | E | R2 | Slope |
|---|---|---|---|---|---|---|---|
| WT | 22.3 | 25.8 | 30.0 | 34.4 | 75.8 | 0.99 | −4.1 |
| R132C | 27.3 | 30.6 | 34.0 | 37.1 | 98.9 | 0.99 | −3.4 |
| R132G | 24.6 | 28.1 | 31.4 | 34.7 | 95.0 | 0.99 | −3.5 |
| R132H | 28.2 | 30.8 | 33.1 | 35.8 | 133.6 | 0.99 | −2.7 |
| R132L | 23.3 | 27.1 | 30.2 | 32.7 | 103.2 | 0.98 | −3.3 |
| R132S | 22.6 | 26.4 | 29.7 | 32.6 | 102.9 | 0.99 | −3.3 |
| IDH1 Mutation | 8% Mutated Plasmid | 4% Mutated Plasmid | 2% Mutated Plasmid | |||
|---|---|---|---|---|---|---|
| M-FAM Ct | WT-HEX Ct | M-FAM Ct | WT-HEX Ct | M-FAM Ct | WT-HEX Ct | |
| R132C | 26.0 | 28.9 | 26.9 | 28.6 | 27.9 | 28.5 |
| R132G | 27.0 | 28.9 | 27.8 | 28.5 | 28.9 | 28.5 |
| R132H | ND | 28.9 | ND | 28.6 | ND | 28.3 |
| R132L | 28.5 | 28.3 | 33.1 | 28.2 | 37.0 | 28.7 |
| R132S | 27.0 | 28.9 | 27.3 | 28.5 | 28.4 | 28.4 |
| IDH1 Mutated Sample | 50% | 10% | |||
|---|---|---|---|---|---|
| ID | Mutation | M-FAM (Ct) | WT-HEX (Ct) | M-FAM (Ct) | WT-HEX (Ct) |
| #11 | R132L | 29.2 | 29.1 | 36.7 | 28.8 |
| #41 | R132L | 27.6 | 27.7 | ND | 28.4 |
| #61 | R132G | 27.8 | 29.1 | 31.3 | 29.0 |
| Patient ID | Mutation Detected with Sanger | Mutation Detected with qPCR |
|---|---|---|
| #11 | R132L | R132L |
| #15 | R132C | |
| #21 | R312C | |
| #22 | R132L | R132L |
| #41 | R132L | R132L |
| #45 | R312C | |
| #46 | R132L | |
| #51 | R132L | R132L |
| #57 | R132L | |
| #61 | R132G | R132G |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Peraldo-Neia, C.; Scatolini, M.; Grosso, E.; Lombardi, P.; Filippi, R.; Raggi, C.; Marchiò, C.; Cavalloni, G.; Aglietta, M.; Leone, F. Assessment of a High Sensitivity Method for Identification of IDH1 R132x Mutations in Tumors and Plasma of Intrahepatic Cholangiocarcinoma Patients. Cancers 2019, 11, 454. https://doi.org/10.3390/cancers11040454
Peraldo-Neia C, Scatolini M, Grosso E, Lombardi P, Filippi R, Raggi C, Marchiò C, Cavalloni G, Aglietta M, Leone F. Assessment of a High Sensitivity Method for Identification of IDH1 R132x Mutations in Tumors and Plasma of Intrahepatic Cholangiocarcinoma Patients. Cancers. 2019; 11(4):454. https://doi.org/10.3390/cancers11040454
Chicago/Turabian StylePeraldo-Neia, Caterina, Maria Scatolini, Enrico Grosso, Pasquale Lombardi, Roberto Filippi, Chiara Raggi, Caterina Marchiò, Giuliana Cavalloni, Massimo Aglietta, and Francesco Leone. 2019. "Assessment of a High Sensitivity Method for Identification of IDH1 R132x Mutations in Tumors and Plasma of Intrahepatic Cholangiocarcinoma Patients" Cancers 11, no. 4: 454. https://doi.org/10.3390/cancers11040454
APA StylePeraldo-Neia, C., Scatolini, M., Grosso, E., Lombardi, P., Filippi, R., Raggi, C., Marchiò, C., Cavalloni, G., Aglietta, M., & Leone, F. (2019). Assessment of a High Sensitivity Method for Identification of IDH1 R132x Mutations in Tumors and Plasma of Intrahepatic Cholangiocarcinoma Patients. Cancers, 11(4), 454. https://doi.org/10.3390/cancers11040454




