Observed Survival Interval: A Supplement to TCGA Pan-Cancer Clinical Data Resource
Abstract
1. Introduction
2. Materials and Methods
2.1. Data Retrieval and Preprocessing
2.2. Differential Expression Analysis
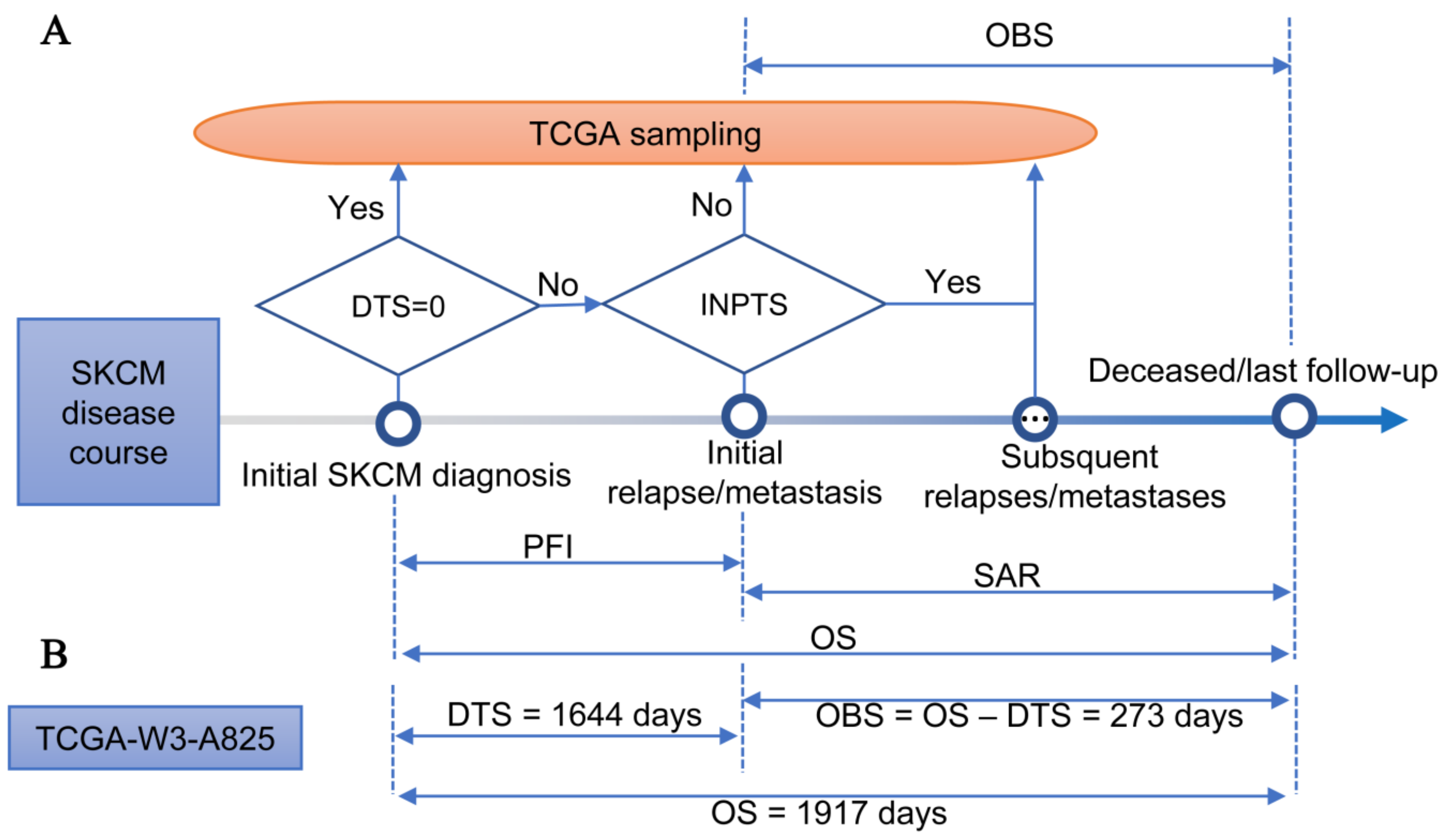
2.3. Observed Survival Interval
2.4. Comparison of OS and OBS in Association with Clinical Data
2.5. Comparison of OS and OBS in Associating miRNA Sequencing Data
2.6. Statistical Analysis
3. Results
3.1. TCGA-SKCM Dataset
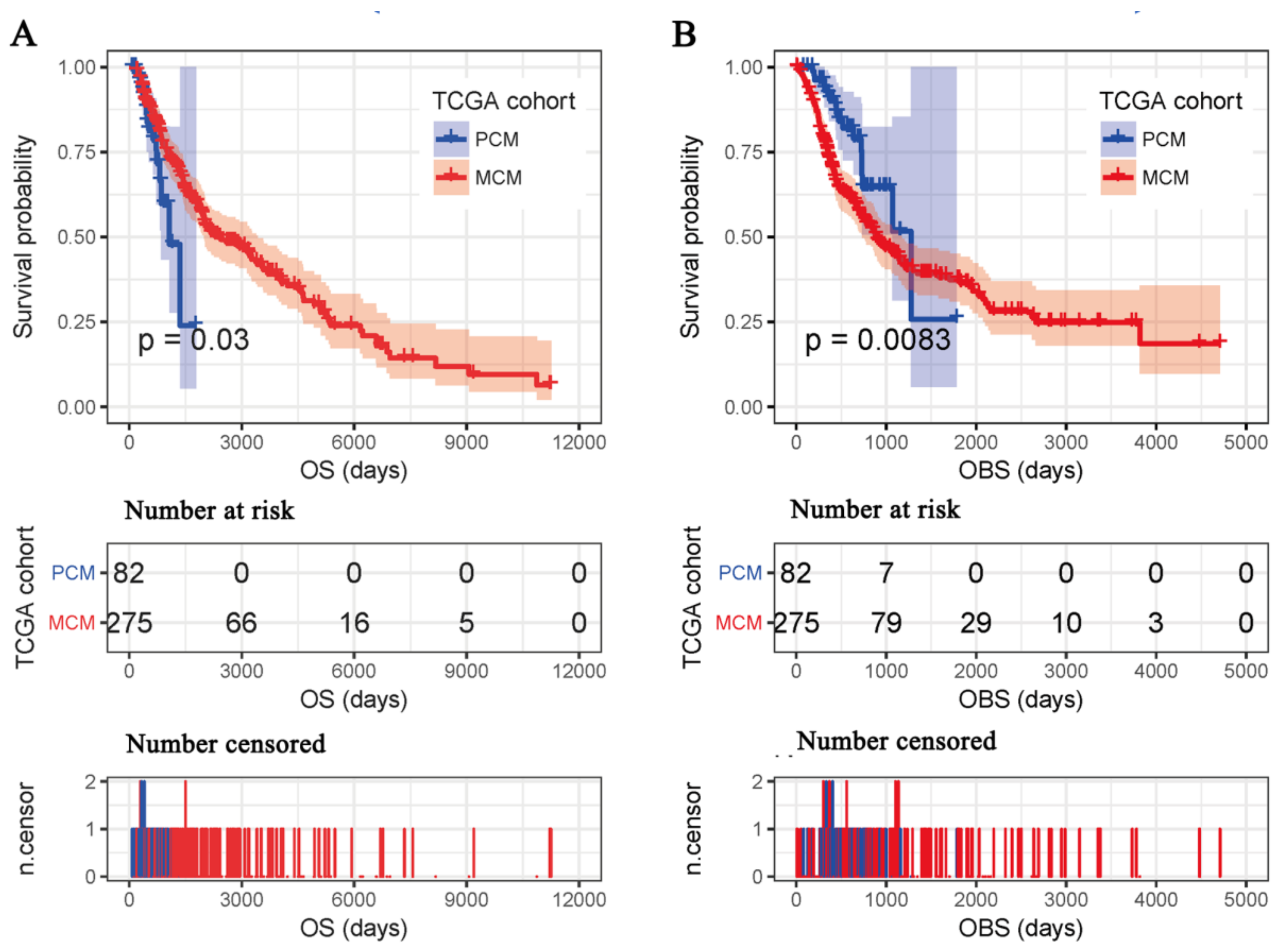
3.2. Differences between OBS and OS
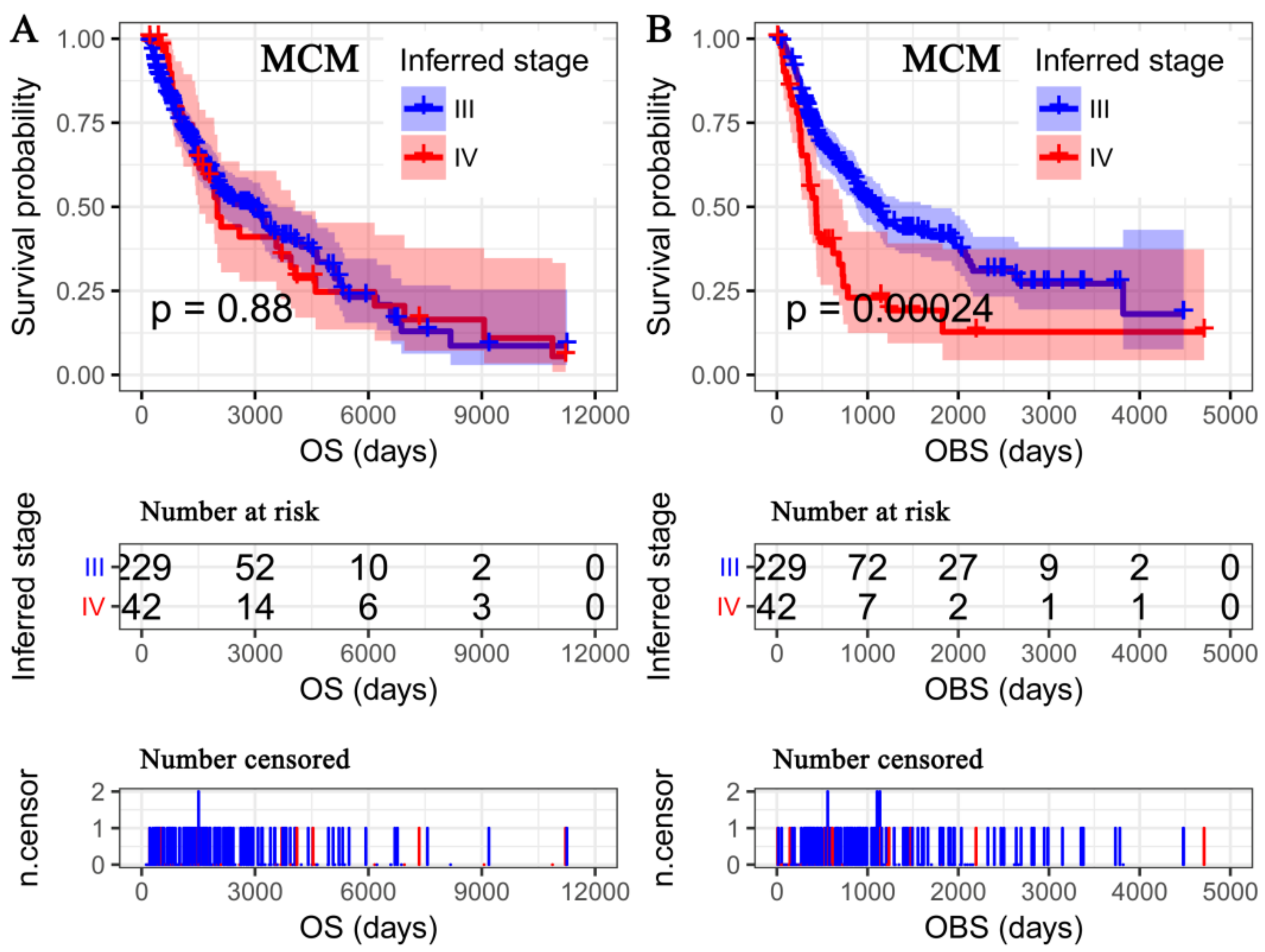
3.3. OS Deemed More Appropriate to Associate Clinicopathological Characteristics than the OBS
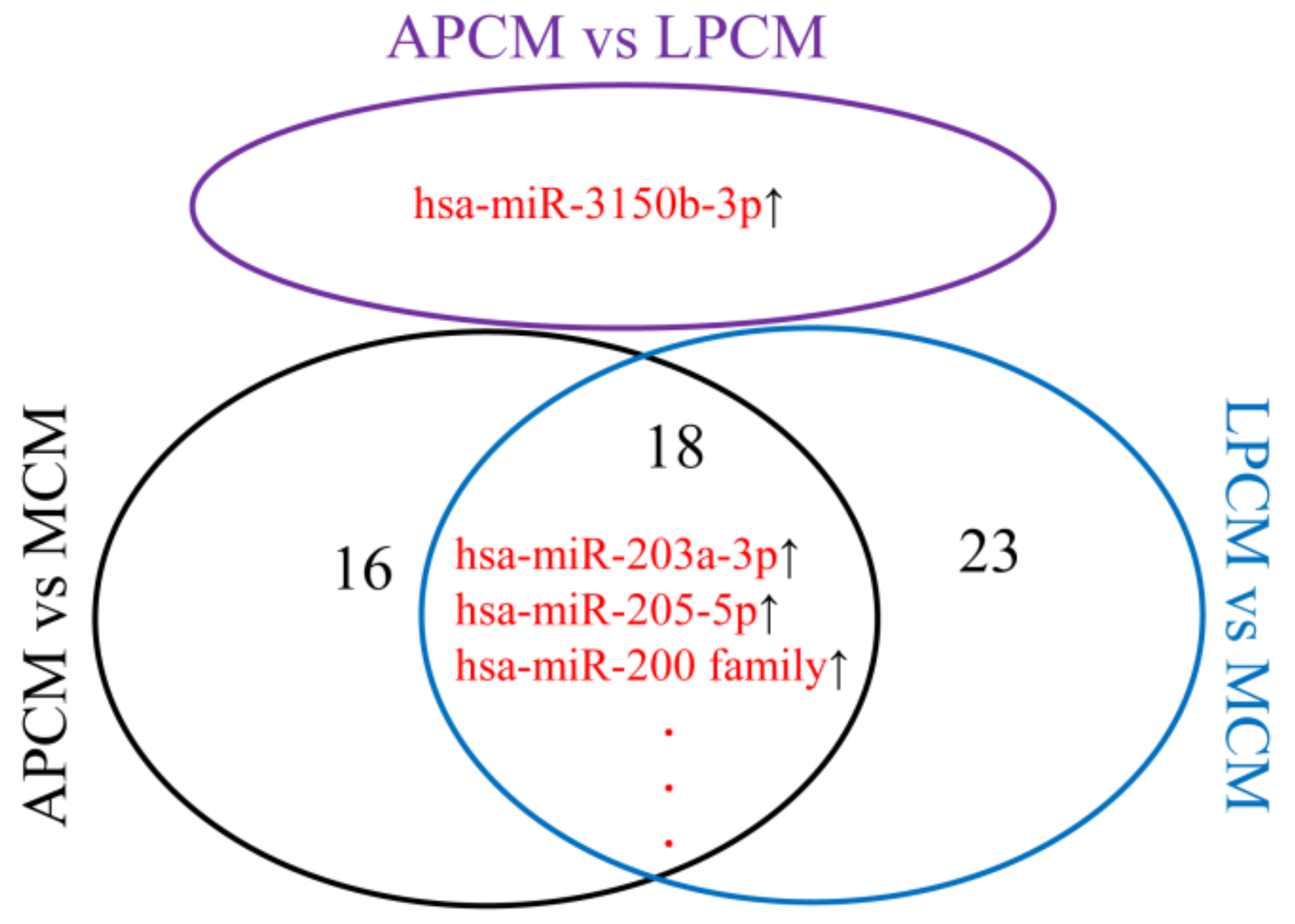
3.4. Differentially Expressed miRNAs
3.5. OBS Deemed More Appropriate to Associate miRNA-Omics Data than OS
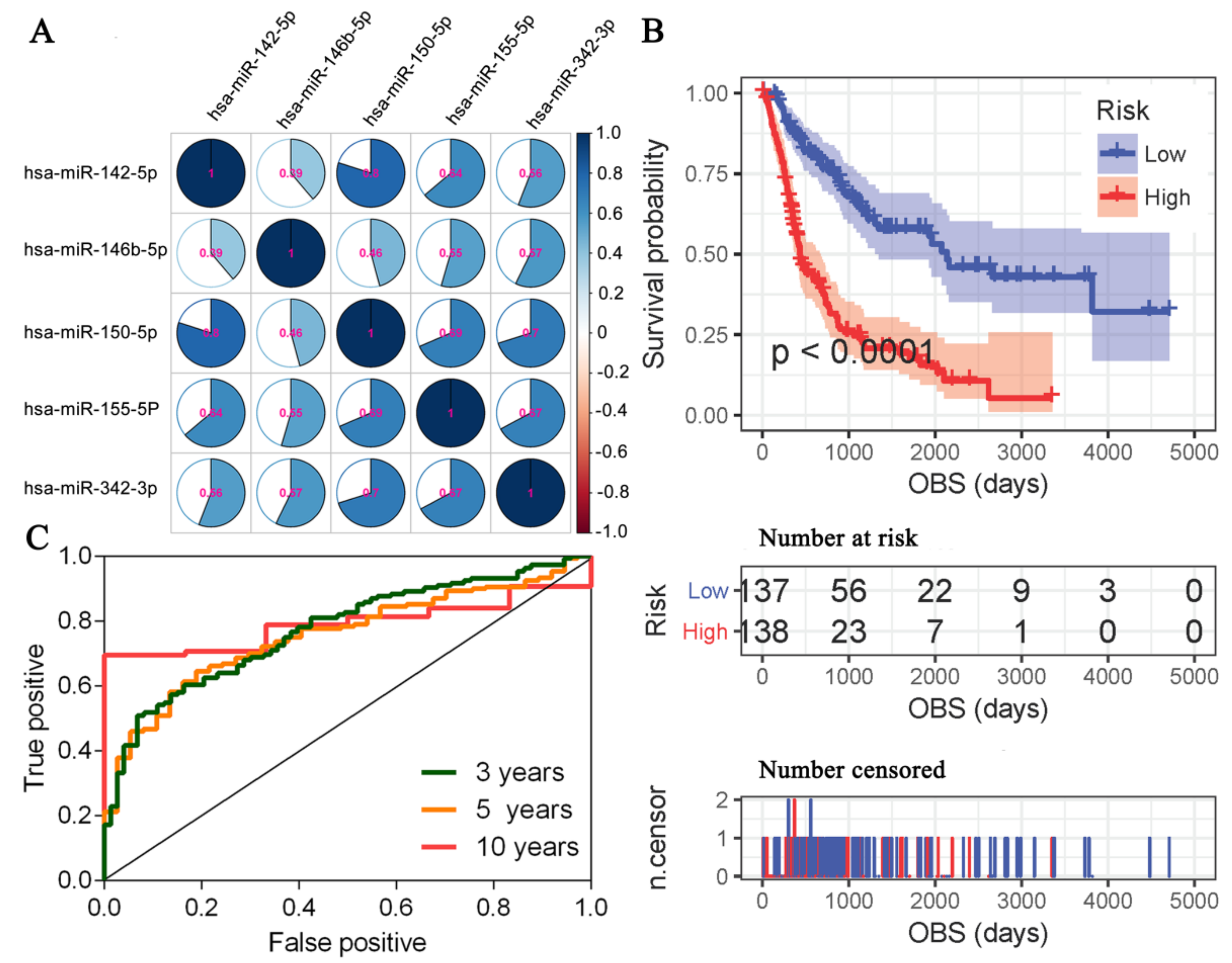
3.6. A miRNA Expression Signature for MCM Prognosis Based on the OBS
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Ali, Z.; Yousaf, N.; Larkin, J. Melanoma epidemiology, biology and prognosis. Eur. J. Cancer Suppl. 2013, 11, 81–91. [Google Scholar] [CrossRef] [PubMed]
- Wang, H.; Naghavi, M.; Allen, C.; Barber, R.M.; Bhutta, Z.A.; Carter, A.; Casey, D.C.; Charlson, F.J.; Chen, A.Z.; Coates, M.M.; et al. Global, regional, and national life expectancy, all-cause mortality, and cause-specific mortality for 249 causes of death, 1980–2015: A systematic analysis for the Global Burden of Disease Study 2015. Lancet 2016, 385, 117–171. [Google Scholar] [CrossRef]
- Siegel, R.L.; Miller, K.D.; Jemal, A. Cancer statistics, 2018. CA Cancer J. Clin. 2018, 68, 7–30. [Google Scholar] [CrossRef] [PubMed]
- Gershenwald, J.E.; Scolyer, R.A.; Hess, K.R.; Sondak, V.K.; Long, G.V.; Ross, M.I.; Lazar, A.J.; Faries, M.B.; Kirkwood, J.M.; McArthur, G.A.; et al. Melanoma staging: Evidence-based changes in the American Joint Committee on Cancer eighth edition cancer staging manual. CA Cancer J. Clin. 2017, 67, 472–492. [Google Scholar] [CrossRef] [PubMed]
- Hanniford, D.; Zhong, J.; Koetz, L.; Gaziel-Sovran, A.; Lackaye, D.J.; Shang, S.; Pavlick, A.; Shapiro, R.; Berman, R.; Darvishian, F.; et al. A miRNA-Based Signature Detected in Primary Melanoma Tissue Predicts Development of Brain Metastasis. Clin. Cancer Res. 2015, 21, 4903–4912. [Google Scholar] [CrossRef] [PubMed]
- Hu, X.; Schwarz, J.K.; Lewis, J.S.J.; Huettner, P.C.; Rader, J.S.; Deasy, J.O.; Grigsby, P.W.; Wang, X. A microRNA expression signature for cervical cancer prognosis. Cancer Res. 2010, 70, 1441–1448. [Google Scholar] [CrossRef] [PubMed]
- Akbani, R.; Akdemir, K.C.; Aksoy, B.A.; Albert, M.; Ally, A.; Amin, S.B.; Arachchi, H.; Arora, A.; Auman, J.T.; Ayala, B.; et al. Genomic Classification of Cutaneous Melanoma. Cell 2015, 161, 1681–1696. [Google Scholar] [CrossRef] [PubMed]
- Jayawardana, K.; Schramm, S.J.; Tembe, V.; Mueller, S.; Thompson, J.F.; Scolyer, R.A.; Mann, G.J.; Yang, J. Identification, Review, and Systematic Cross-Validation of microRNA Prognostic Signatures in Metastatic Melanoma. J. Investig. Dermatol. 2016, 136, 245–254. [Google Scholar] [CrossRef] [PubMed]
- Guo, J.; Yang, M.; Zhang, W.; Lu, H.; Li, J. A panel of miRNAs as prognostic indicators for clinical outcome of skin cutaneous melanoma. Int. J. Clin. Exp. Med. 2016, 9, 28–39. [Google Scholar]
- Chen, X.; Guo, W.; Xu, X.J.; Su, F.; Wang, Y.; Zhang, Y.; Wang, Q.; Zhu, L. Melanoma long non-coding RNA signature predicts prognostic survival and directs clinical risk-specific treatments. J. Dermatol. Sci. 2017, 85, 226–234. [Google Scholar] [CrossRef] [PubMed]
- Yang, S.; Xu, J.; Zeng, X. A six-long non-coding RNA signature predicts prognosis in melanoma patients. Int. J. Oncol. 2018, 52, 1178–1188. [Google Scholar] [CrossRef] [PubMed]
- Ma, X.; He, Z.; Li, L.; Yang, D.; Liu, G. Expression profiles analysis of long non-coding RNAs identified novel lncRNA biomarkers with predictive value in outcome of cutaneous melanoma. Oncotarget 2017, 8, 77761–77770. [Google Scholar] [CrossRef] [PubMed]
- Jiang, Y.; Shi, X.; Zhao, Q.; Krauthammer, M.; Rothberg, B.E.; Ma, S. Integrated analysis of multidimensional omics data on cutaneous melanoma prognosis. Genomics 2016, 107, 223–230. [Google Scholar] [CrossRef] [PubMed]
- Liu, J.; Lichtenberg, T.; Hoadley, K.A.; Poisson, L.M.; Lazar, A.J.; Cherniack, A.J.; Kovatich, A.J.; Benz, C.C.; Levine, D.A.; Lee, A.V.; et al. An Integrated TCGA Pan-Cancer Clinical Data Resource to Drive High-Quality Survival Outcome Analytics. Cell 2018, 173, 400–416. [Google Scholar] [CrossRef] [PubMed]
- Xiong, J.; Guo, S.; Bing, Z.; Su, Y.; Guo, L. A Comprehensive RNA Expression Signature for Cervical Squamous Cell Carcinoma Prognosis. Front. Genet. 2018, 9, 696. [Google Scholar] [CrossRef] [PubMed]
- Reese, S.E.; Archer, K.J.; Therneau, T.M.; Atkinson, E.J.; Vachon, C.M.; de Andrade, M.; Kocher, J.P.; Eckel-Passow, J.E. A new statistic for identifying batch effects in high-throughput genomic data that uses guided principal component analysis. Bioinformatics 2013, 29, 2877–2883. [Google Scholar] [CrossRef] [PubMed]
- Cox, D.R. Regression Models and Life-Tables. J. Royal Stat. Soc. 1972, 34, 187–220. [Google Scholar] [CrossRef]
- Grambsch, P.M.; Therneau, T.M. Proportional hazards tests and diagnostics based on weighted residuals. Biometrika 1994, 81, 515–526. [Google Scholar] [CrossRef]
- Benjamini, Y.; Hochberg, Y. Controlling the false discovery rate: A practical and powerful approach to multiple testing. J. Royal Stat. Soc. Ser. B 1995, 57, 289–300. [Google Scholar] [CrossRef]
- Blanche, P.; Dartigues, J.F.; Jacqmin-Gadda, H. Estimating and comparing time-dependent areas under receiver operating characteristic curves for censored event times with competing risks. Stat. Med. 2013, 32, 5381–5397. [Google Scholar] [CrossRef] [PubMed]
- Ihaka, R.; Gentleman, R.R. A language for data analysis and graphics. J.Comput. Gr. Stat. 1996, 5, 299–314. [Google Scholar]
- Love, M.I.; Huber, W.; Anders, S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 2014, 15, 550. [Google Scholar] [CrossRef] [PubMed]
- Xu, Y.; Brenn, T.; Brown, E.R.; Doherty, V.; Melton, D.W. Differential expression of microRNAs during melanoma progression: miR-200c, miR-205 and miR-211 are downregulated in melanoma and act as tumour suppressors. Br. J. Cancer 2012, 106, 553–561. [Google Scholar] [CrossRef] [PubMed]
- Segura, M.F.; Belitskayalévy, I.; Rose, A.E.; Zarkrzewski, J.; Gaziel, A.; Hanniford, D.; Darvishian, F.; Berman, R.S.; Shapiro, R.L.; Pavlick, A.C.; et al. Melanoma microRNA signature predicts post-recurrence survival. Clin. Cancer Res. 2010, 16, 1577–1586. [Google Scholar] [CrossRef] [PubMed]
- Caramuta, S.; Egyha´zi, S.; Rodolfo, M.; Witten, D.; Hansson, J.; Larsson, C.; Lui, W.O. MicroRNA Expression Profiles Associated with Mutational Status and Survival in Malignant Melanoma. J. Investig. Dermatol. 2010, 130, 2062–2070. [Google Scholar] [CrossRef] [PubMed]
- Tembe, V.; Schramm, S.J.; Stark, M.S.; Patrick, E.; Jayaswal, V.; Tang, Y.H.; Barbour, A.; Hayward, N.K.; Thompson, J.F.; Scolyer, R.A.; et al. microRNA and mRNA expression profiling in metastatic melanoma reveal associations with BRAF mutation and patient prognosis. Pigment. Cell Melanoma Res. 2014, 28, 254–266. [Google Scholar] [CrossRef] [PubMed]
| Column Header | Deaths/Patients (%) | MS (95% CI) | ULog-rank test p | MHR (95% CI) | MWald test p |
|---|---|---|---|---|---|
| Age1 | |||||
| ≤50 years | 58/113 (51.33) | 4062 (2022–5370) | |||
| >50 years | 115/244 (47.13) | 1927 (1524–2927) | 0.011 | 1.27 (0.82–1.96) | 0.282 |
| Gender | |||||
| Male | 60/138 (43.48) | 2004 (1640–4507) | |||
| Female | 111/219 (50.68) | 2402 (1960–3424) | 0.736 | ||
| Breslow depth1 | |||||
| ≤2 mm | 55/114 (48.25) | 3943 (3139–5318) | |||
| >2 mm | 80/169 (47.33) | 1424 (1103–2004) | <0.001 | 1.44 (0.93–2.22) | 0.098 |
| Pathological stage2 | |||||
| I–II | 81/179 (45.25) | 3266 (2402–4601) | |||
| III–IV | 75/153 (49.02) | 1490 (988–2071) | <0.001 | 1.82 (1.22–2.72) | 0.004 |
| Ulceration1 | |||||
| No | 57/111 (51.35) | 2402 (1927–4222) | |||
| Yes | 58/133 (43.61) | 1354 (1059–2028) | <0.001 | 1.53 (1.00–2.35) | 0.052 |
| Primary tumor site | |||||
| Extremities | 76/158 (48.1) | 2071 (1910–4000) | |||
| Head and neck | 11/23 (47.83) | 2192 (787–NA) | |||
| Trunk | 59/128 (46.09) | 3139 (1691–5107) | 0.787 | ||
| Radiation therapy | |||||
| No | 165/355 (49.25) | 2192 (1917–3266) | |||
| Yes | 7/14 (50.00) | 1341–NA | 0.892 | ||
| Chemotherapy | |||||
| No | 121/251 (48.21) | 2173 (1832–3564) | |||
| Yes | 40/75 (53.33) | 2184 (1917–3683) | 0.813 | ||
| Column Header | MHR (95% CI) | MWald Test P | Type |
|---|---|---|---|
| hsa-miR-155-5p | 0.73 (0.63–0.85) | 3.15×10−5 | Protective1 |
| hsa-miR-4461 | 1.29 (1.13–1.46) | 1.07×10−4 | Risky2 |
| hsa-miR-504-5p | 0.80 (0.71–0.92) | 1.17×10−3 | Protective |
| hsa-miR-625-5p | 0.67 (0.53–0.86) | 1.35×10−3 | Protective |
| hsa-miR-664b-5p | 0.69 (0.58–0.83) | 4.39×10−5 | Protective |
| SRS | 2.28 (1.89–2.74) | <2.00×10−16 | Risky |
| Inferred stage | 1.32 (0.88–1.98) | 0.18 | Risky |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Xiong, J.; Bing, Z.; Guo, S. Observed Survival Interval: A Supplement to TCGA Pan-Cancer Clinical Data Resource. Cancers 2019, 11, 280. https://doi.org/10.3390/cancers11030280
Xiong J, Bing Z, Guo S. Observed Survival Interval: A Supplement to TCGA Pan-Cancer Clinical Data Resource. Cancers. 2019; 11(3):280. https://doi.org/10.3390/cancers11030280
Chicago/Turabian StyleXiong, Jie, Zhitong Bing, and Shengyu Guo. 2019. "Observed Survival Interval: A Supplement to TCGA Pan-Cancer Clinical Data Resource" Cancers 11, no. 3: 280. https://doi.org/10.3390/cancers11030280
APA StyleXiong, J., Bing, Z., & Guo, S. (2019). Observed Survival Interval: A Supplement to TCGA Pan-Cancer Clinical Data Resource. Cancers, 11(3), 280. https://doi.org/10.3390/cancers11030280




