CTCs Expression Profiling for Advanced Breast Cancer Monitoring
Abstract
1. Introduction
2. Results
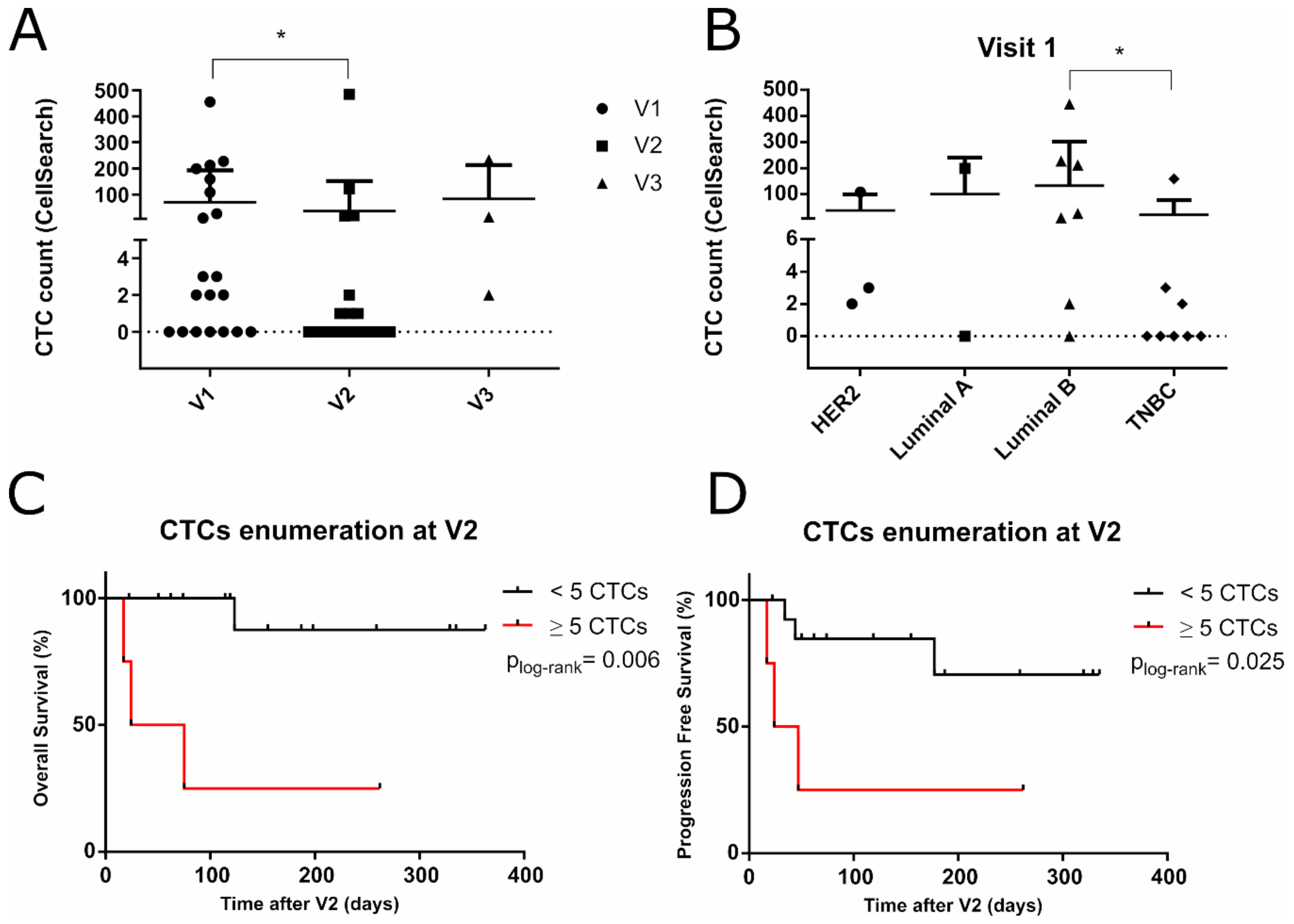
2.1. EpCAM+ CTCs Enumeration Monitoring
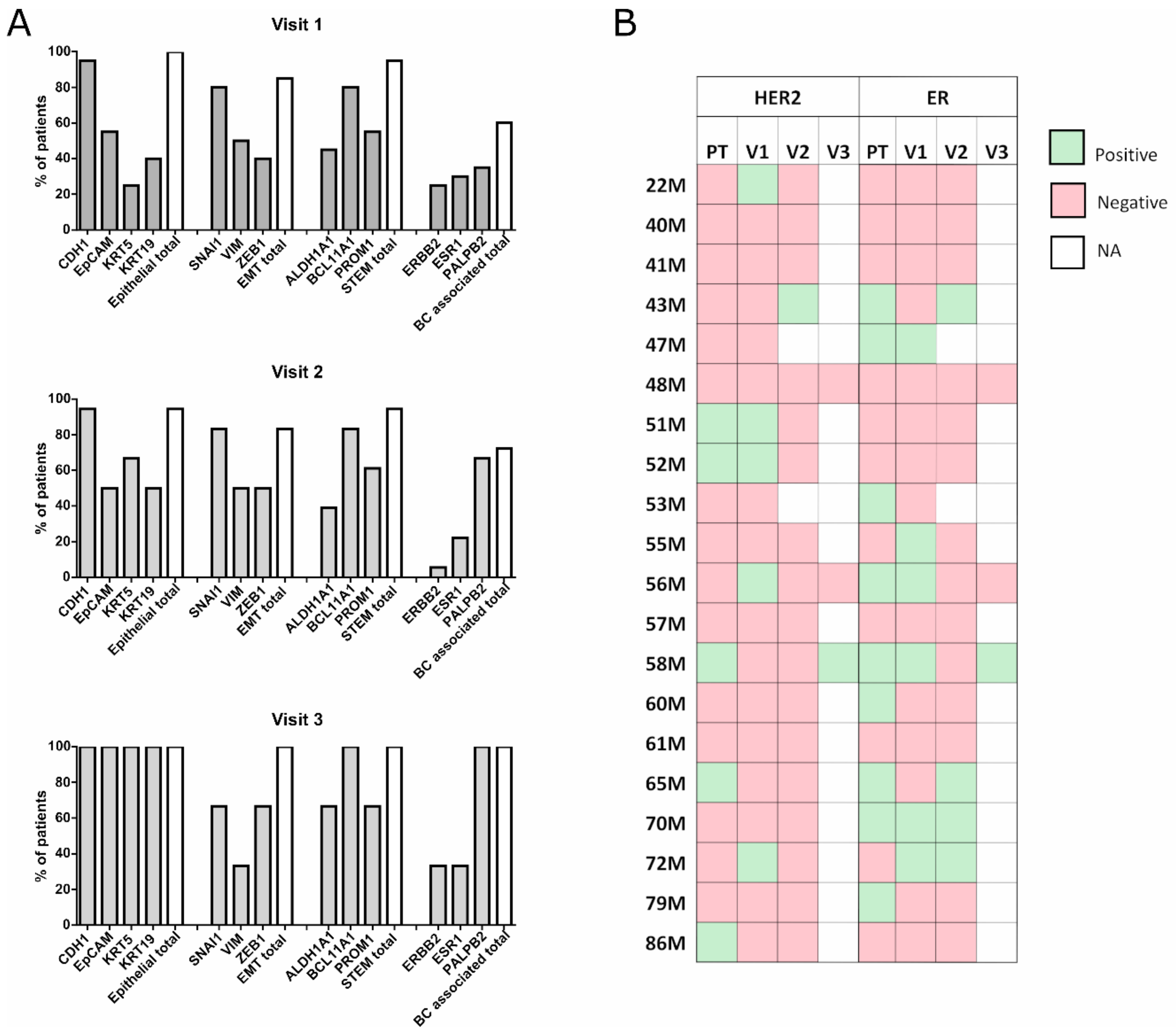
2.2. Unbiased CTC Gene Expression For Advanced BC Patients Monitoring
2.3. CTC Detection By Cell Search and Gene Expression Profiles
2.4. CTCs Expression Profile and Clinical Data
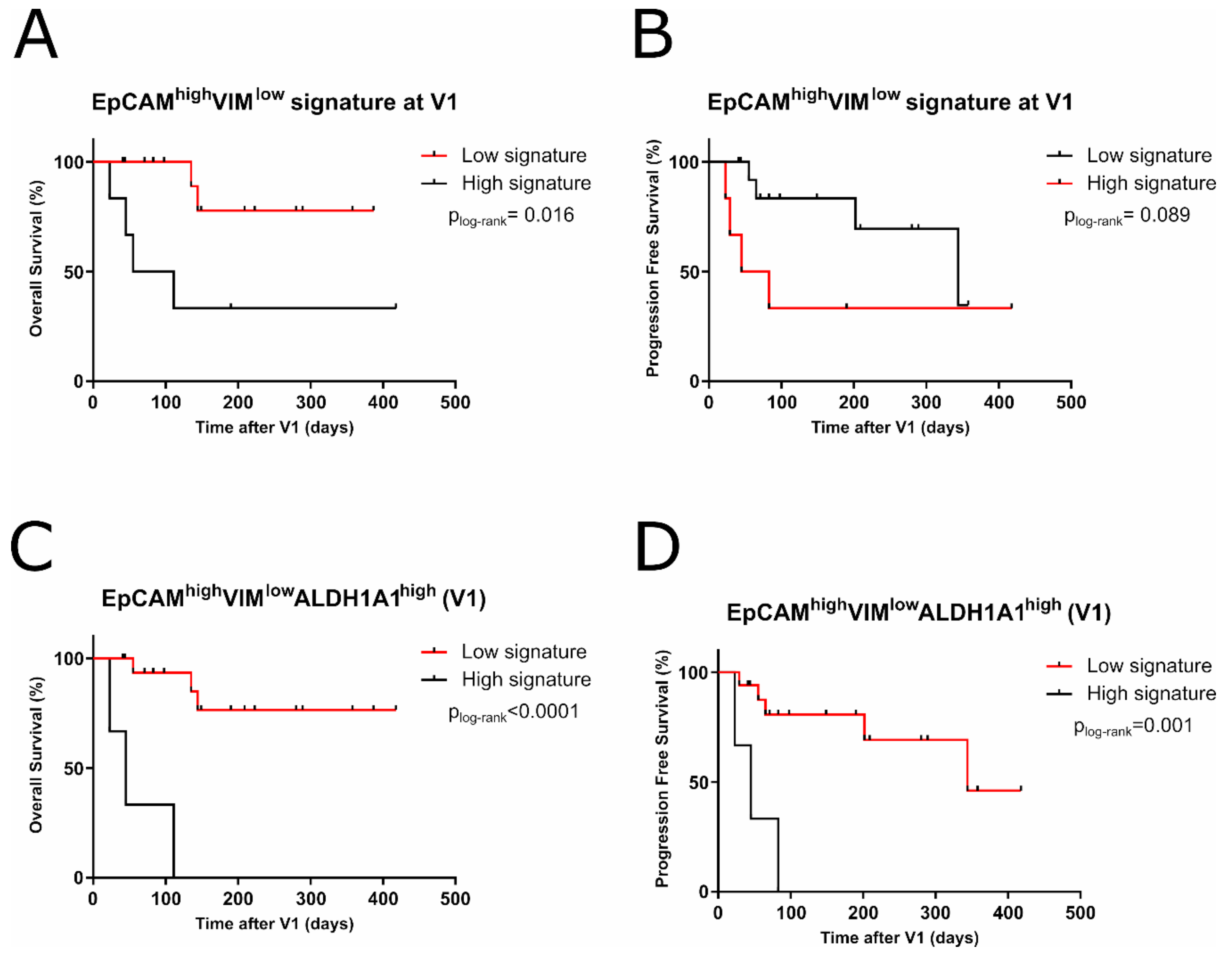
2.5. EpCAMhighVIMlowALDH1A1high CTCs Signature Predicts Shorter Outcome
3. Discussion
4. Materials and Methods
4.1. Patients
4.2. Clinical Samples
4.3. CTC Enumeration
4.4. Peripheral Blood Mononuclear Cells (PBMCs) Isolation and CTCs Enrichment
4.5. Nucleic Acid Extraction, cDNA Synthesis, Preamplification, and Quantitative Real-Time PCR (qPCR)
4.6. Statistical Analysis
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- International Agency for Research on Cancer Breast Cancer Incidence and Mortality Statistics. Available online: https://gco.iarc.fr/ (accessed on 20 October 2019).
- Scully, O.J.; Bay, B.-H.; Yip, G.; Yu, Y. Breast cancer metastasis. Cancer Genom. Proteom. 2012, 9, 311–320. [Google Scholar]
- Lu, J.; Steeg, P.S.; Price, J.E.; Krishnamurthy, S.; Mani, S.A.; Reuben, J.; Cristofanilli, M.; Dontu, G.; Bidaut, L.; Valero, V.; et al. Breast cancer metastasis: Challenges and opportunities. Proc. Cancer Res. 2009, 69, 4951–4953. [Google Scholar] [CrossRef]
- Ignatiadis, M.; Sotiriou, C. Luminal breast cancer: From biology to treatment. Nat. Rev. Clin. Oncol. 2013, 10, 494–506. [Google Scholar] [CrossRef] [PubMed]
- Cejalvo, J.M.; Martínez de Dueñas, E.; Galván, P.; García-Recio, S.; Burgués Gasión, O.; Paré, L.; Antolín, S.; Martinello, R.; Blancas, I.; Adamo, B.; et al. Intrinsic Subtypes and Gene Expression Profiles in Primary and Metastatic Breast Cancer. Cancer Res. 2017, 77, 2213–2221. [Google Scholar] [CrossRef] [PubMed]
- Cancer, T.; Atlas, G. Comprehensive molecular portraits of human breast tumours. Nature 2012, 490, 61. [Google Scholar]
- Pantel, K.; Hayes, D.F. Disseminated breast tumour cells: Biological and clinical meaning. Nat. Rev. Clin. Oncol. 2018, 15, 129–131. [Google Scholar] [CrossRef] [PubMed]
- Cristofanilli, M.; Budd, G.T.; Ellis, M.J.; Stopeck, A.; Matera, J.; Miller, M.C.; Reuben, J.M.; Doyle, G.V.; Allard, W.J.; Terstappen, L.W.M.M.; et al. Circulating tumor cells, disease progression, and survival in metastatic breast cancer. N. Engl. J. Med. 2004, 351, 781–791. [Google Scholar] [CrossRef] [PubMed]
- Alix-Panabierès, C.; Pantel, K. Circulating tumor cells: Liquid biopsy of cancer. Clin. Chem. 2013, 59, 110–118. [Google Scholar] [CrossRef]
- Andree, K.C.; van Dalum, G.; Terstappen, L.W.M.M. Challenges in circulating tumor cell detection by the CellSearch system. Mol. Oncol. 2016, 10, 395–407. [Google Scholar] [CrossRef]
- Lianidou, E.S.; Markou, A.; Strati, A. Molecular characterization of circulating tumor cells in breast cancer: Challenges and promises for individualized cancer treatment. Cancer Metastasis Rev. 2012, 31, 663–671. [Google Scholar] [CrossRef]
- Boral, D.; Vishnoi, M.; Liu, H.N.; Yin, W.; Sprouse, M.L.; Scamardo, A.; Hong, D.S.; Tan, T.Z.; Thiery, J.P.; Chang, J.C.; et al. Molecular characterization of breast cancer CTCs associated with brain metastasis. Nat. Commun. 2017, 8, 196. [Google Scholar] [CrossRef] [PubMed]
- Liu, Z.; Fusi, A.; Klopocki, E.; Schmittel, A.; Tinhofer, I.; Nonnenmacher, A.; Keilholz, U. Negative enrichment by immunomagnetic nanobeads for unbiased characterization of circulating tumor cells from peripheral blood of cancer patients. J. Transl. Med. 2011, 9, 70. [Google Scholar] [CrossRef] [PubMed]
- Lianidou, E.S.; Markou, A. Circulating tumor cells as emerging tumor biomarkers in breast cancer. Clin. Chem. Lab. Med. 2011, 49, 1579–1590. [Google Scholar]
- Mego, M.; De Giorgi, U.; Dawood, S.; Wang, X.; Valero, V.; Andreopoulou, E.; Handy, B.; Ueno, N.T.; Reuben, J.M.; Cristofanilli, M. Characterization of metastatic breast cancer patients with nondetectable circulating tumor cells. Int. J. Cancer 2011, 129, 417–423. [Google Scholar] [CrossRef]
- Giordano, A.; Giuliano, M.; De Laurentiis, M.; Arpino, G.; Jackson, S.; Handy, B.C.; Ueno, N.T.; Andreopoulou, E.; Alvarez, R.H.; Valero, V.; et al. Circulating tumor cells in immunohistochemical subtypes of metastatic breast cancer: Lack of prediction in HER2-positive disease treated with targeted therapy. Ann. Oncol. Off. J. Eur. Soc. Med. Oncol. 2012, 23, 1144–1150. [Google Scholar] [CrossRef]
- Pierga, J.Y.; Bidard, F.C.; Mathiot, C.; Brain, E.; Delaloge, S.; Giachetti, S.; De Cremoux, P.; Salmon, R.; Vincent-Salomon, A.; Marty, M. Circulating tumor cell detection predicts early metastatic relapse after neoadjuvant chemotherapy in large operable and locally advanced breast cancer in a phase II randomized trial. Clin. Cancer Res. 2008, 14, 7004–7010. [Google Scholar] [CrossRef]
- Rack, B.; Schindlbeck, C.; Andergassen, U.; Lorenz, R.; Zwingers, T.; Schneeweiss, A.; Lichtenegger, W.; Beckmann, M.W.; Sommer, H.; Pantel, K.; et al. Prognostic Relevance of Circulating Tumor Cells in the Peripheral Blood of Primary Breast Cancer Patients. Cancer Res. 2010, 70, S5–S6. [Google Scholar]
- Rack, B.; Schindlbeck, C.; Jückstock, J.; Andergassen, U.; Hepp, P.; Zwingers, T.; Friedl, T.W.P.; Lorenz, R.; Tesch, H.; Fasching, P.A.; et al. Circulating tumor cells predict survival in early average-to-high risk breast cancer patients. J. Natl. Cancer Inst. 2014, 106, dju066. [Google Scholar] [CrossRef]
- Hayes, D.F.; Cristofanilli, M.; Budd, G.T.; Ellis, M.J.; Stopeck, A.; Miller, M.C.; Matera, J.; Allard, W.J.; Doyle, G.V.; Terstappen, L.W.W.M. Circulating tumor cells at each follow-up time point during therapy of metastatic breast cancer patients predict progression-free and overall survival. Clin. Cancer Res. 2006, 12, 4218–4224. [Google Scholar] [CrossRef]
- Joosse, S.A.; Gorges, T.M.; Pantel, K. Biology, detection, and clinical implications of circulating tumor cells. EMBO Mol. Med. 2015, 7, 1–11. [Google Scholar] [CrossRef]
- Fehm, T.; Hoffmann, O.; Aktas, B.; Becker, S.; Solomayer, E.F.; Wallwiener, D.; Kimmig, R.; Kasimir-Bauer, S. Detection and characterization of circulating tumor cells in blood of primary breast cancer patients by RT-PCR and comparison to status of bone marrow disseminated cells. Breast Cancer Res. 2009, 11, 1–9. [Google Scholar] [CrossRef] [PubMed]
- Kasimir-Bauer, S.; Hoffmann, O.; Wallwiener, D.; Kimmig, R.; Fehm, T. Expression of stem cell and epithelial-mesenchymal transition markers in primary breast cancer patients with circulating tumor cells. Breast Cancer Res. 2012, 14, R15. [Google Scholar] [CrossRef] [PubMed]
- Barriere, G.; Riouallon, A.; Renaudie, J.; Tartary, M.; Rigaud, M. Mesenchymal characterization: Alternative to simple CTC detection in two clinical trials. Anticancer Res. 2012, 32, 3363–3370. [Google Scholar] [PubMed]
- Barrière, G.; Riouallon, A.; Renaudie, J.; Tartary, M.; Rigaud, M. Mesenchymal and stemness circulating tumor cells in early breast cancer diagnosis. BMC Cancer 2012, 12, 114. [Google Scholar] [CrossRef]
- Kasimir-Bauer, S.; Bittner, A.K.; König, L.; Reiter, K.; Keller, T.; Kimmig, R.; Hoffmann, O. Does primary neoadjuvant systemic therapy eradicate minimal residual disease? Analysis of disseminated and circulating tumor cells before and after therapy. Breast Cancer Res. 2016, 18, 20. [Google Scholar] [CrossRef]
- Keup, C.; Mach, P.; Aktas, B.; Tewes, M.; Kolberg, H.C.; Hauch, S.; Sprenger-Haussels, M.; Kimmig, R.; Kasimir-Bauer, S. RNA profiles of circulating tumor cells and extracellular vesicles for therapy stratification of metastatic breast cancer patients. Clin. Chem. 2018, 64, 1054–1062. [Google Scholar] [CrossRef]
- Aktas, B.; Tewes, M.; Fehm, T.; Hauch, S.; Kimmig, R.; Kasimir-Bauer, S. Stem cell and epithelial-mesenchymal transition markers are frequently overexpressed in circulating tumor cells of metastatic breast cancer patients. Breast Cancer Res. 2009, 11, R46. [Google Scholar] [CrossRef]
- Xenidis, N.; Vlachonikolis, I.; Mavroudis, D.; Perraki, M.; Stathopoulou, A.; Malamos, N.; Kouroussis, C.; Kakolyris, S.; Apostolaki, S.; Vardakis, N.; et al. Peripheral blood circulating cytokeratin-19 mRNA-positive cells after the completion of adjuvant chemotherapy in patients with operable breast cancer. Ann. Oncol. Off. J. Eur. Soc. Med. Oncol. 2003, 14, 849–855. [Google Scholar] [CrossRef]
- Tewes, M.; Aktas, B.; Welt, A.; Mueller, S.; Hauch, S.; Kimmig, R.; Kasimir-Bauer, S. Molecular profiling and predictive value of circulating tumor cells in patients with metastatic breast cancer: An option for monitoring response to breast cancer related therapies. Breast Cancer Res. Treat. 2009, 115, 581–590. [Google Scholar] [CrossRef]
- Bredemeier, M.; Edimiris, P.; Mach, P.; Kubista, M.; Sjöback, R.; Rohlova, E.; Kolostova, K.; Hauch, S.; Aktas, B.; Tewes, M.; et al. Gene expression signatures in circulating tumor cells correlate with response to therapy in metastatic breast cancer. Clin. Chem. 2017, 63, 1585–1593. [Google Scholar] [CrossRef]
- Markiewicz, A.; Topa, J.; Nagel, A.; Skokowski, J.; Seroczynska, B.; Stokowy, T.; Welnicka-Jaskiewicz, M.; Zaczek, A. Spectrum of Epithelial-Mesenchymal Transition Phenotypes in Circulating Tumour Cells from Early Breast Cancer Patients. Cancers 2019, 11, 59. [Google Scholar] [CrossRef] [PubMed]
- Bredemeier, M.; Edimiris, P.; Tewes, M.; Mach, P.; Aktas, B. Establishment of a multimarker qPCR panel for the molecular characterization of circulating tumor cells in blood samples of metastatic breast cancer patients during the course of palliative treatment. Oncotarget 2016, 7, 41677–41690. [Google Scholar] [CrossRef] [PubMed]
- Aaltonen, K.E.; Novosadova, V.; Bendahl, P.-O.; Graffman, C.; Larsson, A.-M.; Ryden, L. Molecular characterization of circulating tumor cells from patients with metastatic breast cancer reflects evolutionary changes in gene expression under the pressure of systemic therapy. Oncotarget 2017, 8, 45544–45565. [Google Scholar] [CrossRef] [PubMed]
- Polioudaki, H.; Agelaki, S.; Chiotaki, R.; Politaki, E.; Mavroudis, D.; Matikas, A.; Georgoulias, V.; Theodoropoulos, P.A. Variable expression levels of keratin and vimentin reveal differential EMT status of circulating tumor cells and correlation with clinical characteristics and outcome of patients with metastatic breast cancer. BMC Cancer 2015, 15, 399. [Google Scholar] [CrossRef] [PubMed]
- Fujisue, M.; Nishimura, R.; Okumura, Y.; Tashima, R.; Nishiyama, Y.; Osako, T.; Toyozumi, Y.; Arima, N. Clinical Significance of CK19 Negative Breast Cancer. Cancers 2012, 5, 1–11. [Google Scholar] [CrossRef]
- Han, E.K.; Sgambato, A.; Jiang, W.; Zhang, Y.J.; Santella, R.M.; Doki, Y.; Cacace, A.M.; Schieren, I.; Weinstein, I.B. Stable overexpression of cyclin D1 in a human mammary epithelial cell line prolongs the S-phase and inhibits growth. Oncogene 1995, 10, 953–961. [Google Scholar]
- Fettig, L.M.; McGinn, O.; Finlay-Schultz, J.; LaBarbera, D.V.; Nordeen, S.K.; Sartorius, C.A. Cross talk between progesterone receptors and retinoic acid receptors in regulation of cytokeratin 5-positive breast cancer cells. Oncogene 2017, 36, 6074–6084. [Google Scholar] [CrossRef][Green Version]
- Joosse, S.A.; Hannemann, J.; Spötter, J.; Bauche, A.; Andreas, A.; Müller, V.; Pantel, K. Changes in keratin expression during metastatic progression of breast cancer: Impact on the detection of circulating tumor cells. Clin. Cancer Res. 2012, 18, 993–1003. [Google Scholar] [CrossRef]
- Alieva, M.; van Rheenen, J.; Broekman, M.L.D. Potential impact of invasive surgical procedures on primary tumor growth and metastasis. Clin. Exp. Metastasis 2018, 35, 319–331. [Google Scholar] [CrossRef]
- Mosly, D.; Turnbull, A.; Sims, A.; Ward, C.; Langdon, S. Predictive markers of endocrine response in breast cancer. World J. Exp. Med. 2018, 8, 1. [Google Scholar] [CrossRef]
- Khaled, W.T.; Choon Lee, S.; Stingl, J.; Chen, X.; Raza Ali, H.; Rueda, O.M.; Hadi, F.; Wang, J.; Yu, Y.; Chin, S.-F.; et al. BCL11A is a triple-negative breast cancer gene with critical functions in stem and progenitor cells. Nat. Commun. 2015, 6, 5987. [Google Scholar] [CrossRef] [PubMed]
- Dairkee, S.H.; Ljung, B.M.; Smith, H.; Hackett, A. Immunolocalization of a human basal epithelium specific keratin in benign and malignant breast disease. Breast Cancer Res. Treat. 1987, 10, 11–20. [Google Scholar] [CrossRef] [PubMed]
- Lehn, S.; Fernö, M.; Jirström, K.; Rydén, L.; Landberg, G. A non-functional retinoblastoma tumor suppressor (RB) pathway in premenopausal breast cancer is associated with resistance to tamoxifen. Cell Cycle 2011, 10, 956–962. [Google Scholar] [CrossRef] [PubMed]
- Caligiuri, I.; Toffoli, G.; Giordano, A.; Rizzolio, F. pRb controls estrogen receptor alpha protein stability and activity. Oncotarget 2013, 4, 875–883. [Google Scholar] [CrossRef]
- Wolfer, A.; Wittner, B.S.; Irimia, D.; Flavin, R.J.; Lupien, M.; Gunawardane, R.N.; Meyer, C.A.; Lightcap, E.S.; Tamayo, P.; Mesirov, J.P.; et al. MYC regulation of a “poor-prognosis” metastatic cancer cell state. Proc. Natl. Acad. Sci. USA 2010, 107, 3698–3703. [Google Scholar] [CrossRef]
- Yang, A.; Qin, S.; Schulte, B.A.; Ethier, S.P.; Tew, K.D.; Wang, G.Y. MYC Inhibition Depletes Cancer Stem-like Cells in Triple-Negative Breast Cancer. Cancer Res. 2017, 77, 6641–6650. [Google Scholar] [CrossRef]
- McNeil, C.M.; Sergio, C.M.; Anderson, L.R.; Inman, C.K.; Eggleton, S.A.; Murphy, N.C.; Millar, E.K.A.; Crea, P.; Kench, J.G.; Alles, M.C.; et al. c-Myc overexpression and endocrine resistance in breast cancer. J. Steroid Biochem. Mol. Biol. 2006, 102, 147–155. [Google Scholar] [CrossRef]
- Reinhardt, F.; Franken, A.; Meier-Stiegen, F.; Driemel, C.; Stoecklein, N.H.; Fischer, J.C.; Niederacher, D.; Ruckhaeberle, E.; Fehm, T.; Neubauer, H. Diagnostic leukapheresis enables reliable transcriptomic profiling of single circulating tumor cells to characterize inter-cellular heterogeneity in terms of endocrine resistance. Cancers 2019, 11, 903. [Google Scholar] [CrossRef]
- Sheikh, A.; Hussain, S.A.; Ghori, Q.; Naeem, N.; Fazil, A.; Giri, S.; Sathian, B.; Mainali, P.; Al Tamimi, D.M. The spectrum of genetic mutations in breast cancer. Asian Pac. J. Cancer Prev. 2015, 16, 2177–2185. [Google Scholar] [CrossRef]
- Park, J.Y.; Zhang, F.; Andreassen, P.R. PALB2: The hub of a network of tumor suppressors involved in DNA damage responses. Biochim. Biophys. Acta Rev. Cancer 2014, 1846, 263–275. [Google Scholar] [CrossRef]
- Li, J.; Li, M.; Chen, P.; Ba, Q. High expression of PALB2 predicts poor prognosis in patients with advanced breast cancer. FEBS Open Bio. 2017, 8, 56–63. [Google Scholar] [CrossRef] [PubMed]
- Nepomuceno, T.C.; De Gregoriis, G.; de Oliveira, F.M.B.; Suarez-Kurtz, G.; Monteiro, A.N.; Carvalho, M.A. The role of PALB2 in the DNA damage response and cancer predisposition. Int. J. Mol. Sci. 2017, 18, 1886. [Google Scholar] [CrossRef] [PubMed]
- Yu, M.; Bardia, A.; Wittner, B.S.; Stott, S.L.; Smas, M.E.; Ting, D.T.; Isakoff, S.J.; Ciciliano, J.C.; Wells, M.N.; Shah, A.M.; et al. Circulating Breast Tumor Cells Exhibit Dynamic Changes in Epithelial and Mesenchymal Composition. Science 2013, 339, 580–584. [Google Scholar] [CrossRef] [PubMed]
- Pinto, C.A.; Widodo, E.; Waltham, M.; Thompson, E.W. Breast cancer stem cells and epithelial mesenchymal plasticity-Implications for chemoresistance. Cancer Lett. 2013, 341, 56–62. [Google Scholar] [CrossRef]
- Bulfoni, M.; Gerratana, L.; Del Ben, F.; Marzinotto, S.; Sorrentino, M.; Turetta, M.; Scoles, G.; Toffoletto, B.; Isola, M.; Beltrami, C.A.; et al. In patients with metastatic breast cancer the identification of circulating tumor cells in epithelial-to-mesenchymal transition is associated with a poor prognosis. Breast Cancer Res. 2016, 18, 30. [Google Scholar] [CrossRef]
- Markiewicz, A.; Zaczek, A.J. The Landscape of Circulating Tumor Cell Research in the Context of Epithelial-Mesenchymal Transition. Pathobiology 2017, 84, 264–283. [Google Scholar] [CrossRef]
- Lou, Y.; Preobrazhenska, O.; auf dem Keller, U.; Sutcliffe, M.; Barclay, L.; McDonald, P.C.; Roskelley, C.; Overall, C.M.; Dedhar, S. Epithelial-mesenchymal transition (EMT) is not sufficient for spontaneous murine breast cancer metastasis. Dev. Dyn. 2008, 237, 2755–2768. [Google Scholar] [CrossRef]
- Fischer, K.R.; Durrans, A.; Lee, S.; Sheng, J.; Li, F.; Wong, S.T.C.; Choi, H.; El Rayes, T.; Ryu, S.; Troeger, J.; et al. Epithelial-to-mesenchymal transition is not required for lung metastasis but contributes to chemoresistance. Nature 2015, 527, 472–476. [Google Scholar] [CrossRef]
- Jolly, M.K.; Ware, K.E.; Gilja, S.; Somarelli, J.A.; Levine, H. EMT and MET: Necessary or permissive for metastasis? Mol. Oncol. 2017, 11, 755–769. [Google Scholar] [CrossRef]
- Padmanaban, V.; Krol, I.; Suhail, Y.; Szczerba, B.M.; Aceto, N.; Bader, J.S.; Ewald, A.J. E-cadherin is required for metastasis in multiple models of breast cancer. Nature 2019, 573, 439–444. [Google Scholar] [CrossRef]
- De Wit, S.; Manicone, M.; Rossi, E.; Lampignano, R.; Yang, L.; Zill, B.; Rengel-Puertas, A.; Ouhlen, M.; Crespo, M.; Berghuis, A.M.S.; et al. EpCAMhigh and EpCAMlow circulating tumour cells in metastatic prostate and breast cancer patients. Oncotarget 2018, 9, 35705–35716. [Google Scholar] [CrossRef] [PubMed]
- Charafe-Jauffret, E.; Ginestier, C.; Iovino, F.; Tarpin, C.; Diebel, M.; Esterni, B.; Houvenaeghel, G.; Extra, J.-M.; Bertucci, F.; Jacquemier, J.; et al. Aldehyde dehydrogenase 1-positive cancer stem cells mediate metastasis and poor clinical outcome in inflammatory breast cancer. Clin. Cancer Res. 2010, 16, 45–55. [Google Scholar] [CrossRef] [PubMed]
- Ginestier, C.; Hur, M.H.; Charafe-Jauffret, E.; Monville, F.; Dutcher, J.; Brown, M.; Jacquemier, J.; Viens, P.; Kleer, C.G.; Liu, S.; et al. ALDH1 is a marker of normal and malignant human mammary stem cells and a predictor of poor clinical outcome. Cell Stem Cell 2007, 1, 555–567. [Google Scholar] [CrossRef] [PubMed]
- Ma, F.; Li, H.; Li, Y.; Ding, X.; Wang, H.; Fan, Y.; Lin, C.; Qian, H.; Xu, B. Aldehyde dehydrogenase 1 (ALDH1) expression is an independent prognostic factor in triple negative breast cancer (TNBC). Medicine 2017, 96, e6561. [Google Scholar] [CrossRef]
- Yao, J.; Jin, Q.; Wang, X.-D.; Zhu, H.-J.; Ni, Q.-C. Aldehyde dehydrogenase 1 expression is correlated with poor prognosis in breast cancer. Medicine 2017, 96, e7171. [Google Scholar] [CrossRef]
- Raimondi, C.; Gradilone, A.; Naso, G.; Vincenzi, B. Epithelial-mesenchymal transition and stemness features in circulating tumor cells from breast cancer patients. Breast Cancer Res. Treat. 2011, 130, 449–455. [Google Scholar] [CrossRef]
- Strati, A.; Nikolaou, M.; Georgoulias, V.; Lianidou, E.S. Prognostic significance of TWIST1, CD24, CD44, and ALDH1 transcript quantification in EpCAM-positive circulating tumor cells from early stage breast cancer patients. Cells 2019, 8, 652. [Google Scholar] [CrossRef]

| CTCs | Gene Expression (ERRB2, EpCAM, ESR1, GDF15) | p-Value | |
| High | Low | ||
| 0 CTCs | 1 | 6 | 0.019 |
| >1 CTCs | 9 | 4 | |
| Gene Expression (ERRB2, EpCAM, KRT19) | p-Value | ||
| High | Low | ||
| <5 CTCs | 3 | 9 | 0.006 |
| ≥5 CTCs | 7 | 1 | |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Pereira-Veiga, T.; Martínez-Fernández, M.; Abuin, C.; Piñeiro, R.; Cebey, V.; Cueva, J.; Palacios, P.; Blanco, C.; Muinelo-Romay, L.; Abalo, A.; et al. CTCs Expression Profiling for Advanced Breast Cancer Monitoring. Cancers 2019, 11, 1941. https://doi.org/10.3390/cancers11121941
Pereira-Veiga T, Martínez-Fernández M, Abuin C, Piñeiro R, Cebey V, Cueva J, Palacios P, Blanco C, Muinelo-Romay L, Abalo A, et al. CTCs Expression Profiling for Advanced Breast Cancer Monitoring. Cancers. 2019; 11(12):1941. https://doi.org/10.3390/cancers11121941
Chicago/Turabian StylePereira-Veiga, Thais, Mónica Martínez-Fernández, Carmen Abuin, Roberto Piñeiro, Victor Cebey, Juan Cueva, Patricia Palacios, Cristina Blanco, Laura Muinelo-Romay, Alicia Abalo, and et al. 2019. "CTCs Expression Profiling for Advanced Breast Cancer Monitoring" Cancers 11, no. 12: 1941. https://doi.org/10.3390/cancers11121941
APA StylePereira-Veiga, T., Martínez-Fernández, M., Abuin, C., Piñeiro, R., Cebey, V., Cueva, J., Palacios, P., Blanco, C., Muinelo-Romay, L., Abalo, A., Costa, C., & López-López, R. (2019). CTCs Expression Profiling for Advanced Breast Cancer Monitoring. Cancers, 11(12), 1941. https://doi.org/10.3390/cancers11121941





