Aptamer Therapeutics in Cancer: Current and Future
Abstract
1. Advantages of Aptamers
2. Challenges and Possible Solutions in Aptamer Therapeutics
2.1. Aptamer Stability
2.2. Renal Excretion
2.3. Safety of Aptamers
3. Aptamers as Therapeutics under Investigation in Oncology
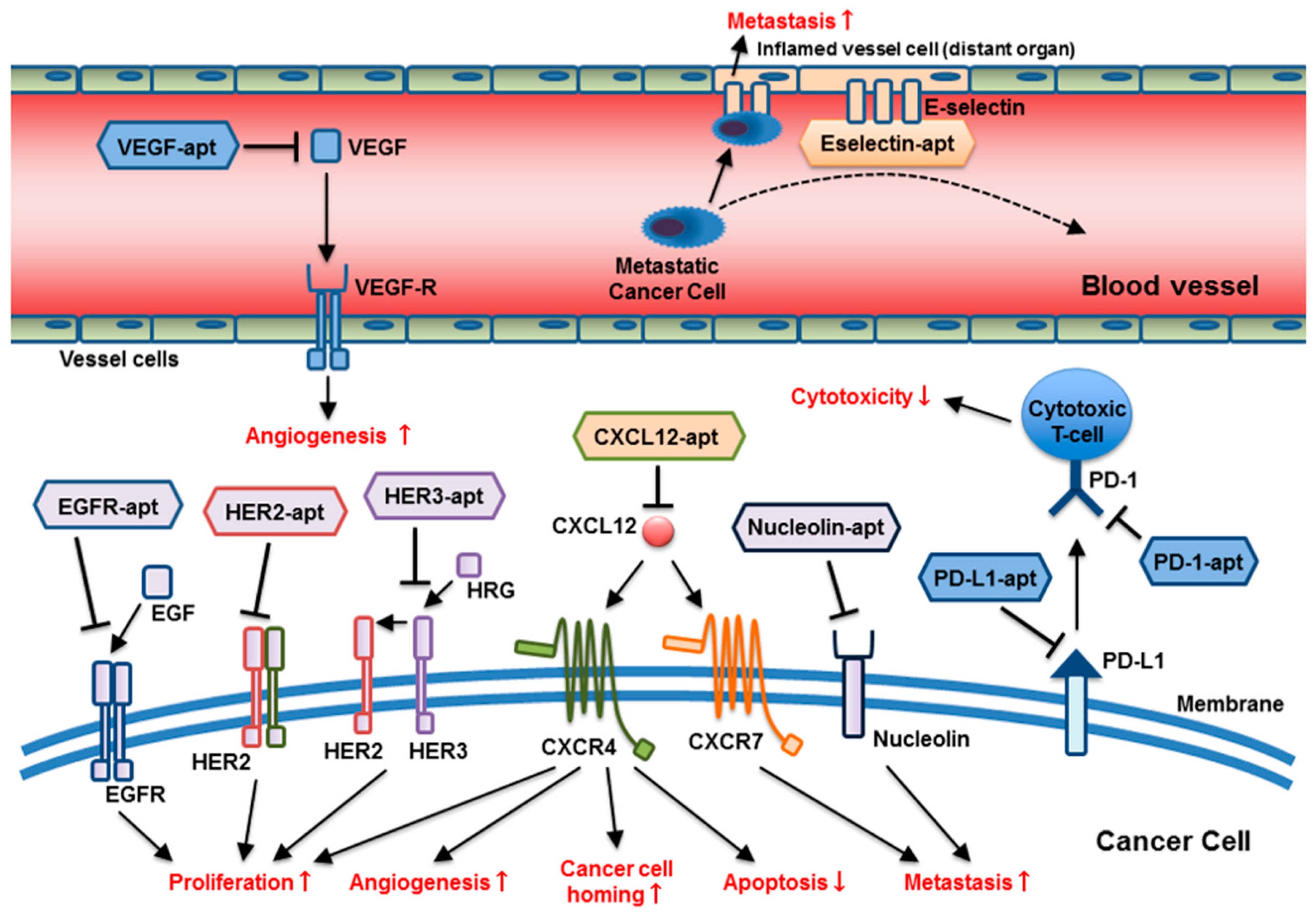
3.1. Aptamers as Tools for Cancer Therapy
3.1.1. RNA Aptamer Targeting VEGF
3.1.2. Aptamers Targeting the Epidermal Growth Factor Receptor (EGFR)
3.1.3. Aptamers Targeting HER2 and HER3
3.1.4. DNA Aptamer Targeting Nucleolin
3.1.5. DNA Aptamer Targeting PD-1/PD-L1
3.1.6. RNA Aptamer Targeting SDF-1 (CXCL12)
3.1.7. Thioaptamer Targeting E-Selectin
3.1.8. RNA Aptamer Targeting CD40
3.1.9. DNA Aptamer Targeting CTLA-4
3.1.10. RNA/DNA Aptamer Targeting C5a
3.1.11. Thioaptamer Targeting Annexin A2
3.1.12. Bispecific Aptamers
3.2. Antidote Aptamer for Controlled Therapy
4. Clinical Trials of Aptamer Application in Oncology
5. Conclusions and Future Perspectives
Acknowledgments
Conflicts of Interest
References
- Lakhin, A.V.; Tarantul, V.Z.; Gening, L.V. Aptamers: Problems, solutions and prospects. Acta Naturae 2013, 5, 34–43. [Google Scholar] [PubMed]
- Gewirtz, A.M. Oligonucleotide therapeutics: Clothing the emperor. Curr. Opin. Mol. Ther. 1999, 1, 297–306. [Google Scholar] [PubMed]
- Brody, E.N.; Gold, L. Aptamers as therapeutic and diagnostic agents. J. Biotechnol. 2000, 74, 5–13. [Google Scholar] [CrossRef]
- Yen, H.-Y.; Liu, Y.-C.; Chen, N.-Y.; Tsai, C.-F.; Wang, Y.-T.; Chen, Y.-J.; Hsu, T.-L.; Yang, P.-C.; Wong, C.-H. Effect of sialylation on EGFR phosphorylation and resistance to tyrosine kinase inhibition. Proc. Natl. Acad. Sci. USA 2015, 112, 6955–6960. [Google Scholar] [CrossRef] [PubMed]
- Yun, C.H.; Boggon, T.J.; Li, Y.; Woo, M.S.; Greulich, H.; Meyerson, M.; Eck, M.J. Structures of lung cancer-derived EGFR mutants and inhibitor complexes: Mechanism of activation and insights into differential inhibitor sensitivity. Cancer Cell 2007, 11, 217–227. [Google Scholar] [CrossRef] [PubMed]
- Li, S.; Schmitz, K.R.; Jeffrey, P.D.; Wiltzius, J.J.; Kussie, P.; Ferguson, K.M. Structural basis for inhibition of the epidermal growth factor receptor by cetuximab. Cancer Cell 2005, 7, 301–311. [Google Scholar] [CrossRef] [PubMed]
- Esposito, C.L.; Passaro, D.; Longobardo, I.; Condorelli, G.; Marotta, P.; Affuso, A.; de Franciscis, V.; Cerchia, L. A neutralizing RNA aptamer against EGFR causes selective apoptotic cell death. PLoS ONE 2011, 6, e24071. [Google Scholar] [CrossRef] [PubMed]
- Li, N.; Nguyen, H.H.; Byrom, M.; Ellington, A.D. Inhibition of cell proliferation by an anti-EGFR aptamer. PLoS ONE 2011, 6, e20299. [Google Scholar] [CrossRef] [PubMed]
- Ruckman, J.; Green, L.S.; Beeson, J.; Waugh, S.; Gillette, W.L.; Henninger, D.D.; Claesson-Welsh, L.; Janjic, N. 2′-fluoropyrimidine RNA-based aptamers to the 165-amino acid form of vascular endothelial growth factor (VEGF165). Inhibition of receptor binding and VEGF-induced vascular permeability through interactions requiring the exon 7-encoded domain. J. Biol. Chem. 1998, 273, 20556–20567. [Google Scholar] [CrossRef] [PubMed]
- Khati, M.; Schuman, M.; Ibrahim, J.; Sattentau, Q.; Gordon, S.; James, W. Neutralization of infectivity of diverse R5 clinical isolates of human immunodeficiency virus type 1 by gp120-binding 2′F-RNA aptamers. J. Virol. 2003, 77, 12692–12698. [Google Scholar] [CrossRef] [PubMed]
- Layzer, J.M.; McCaffrey, A.P.; Tanner, A.K.; Huang, Z.; Kay, M.A.; Sullenger, B.A. In vivo activity of nuclease-resistant siRNAs. RNA 2004, 10, 766–771. [Google Scholar] [CrossRef] [PubMed]
- Jellinek, D.; Green, L.S.; Bell, C.; Lynott, C.K.; Gill, N.; Vargeese, C.; Kirschenheuter, G.; McGee, D.P.; Abesinghe, P.; Pieken, W.A.; et al. Potent 2′-amino-2′-deoxypyrimidine RNA inhibitors of basic fibroblast growth factor. Biochemistry 1995, 34, 11363–11372. [Google Scholar] [CrossRef] [PubMed]
- Yan, X.; Gao, X.; Zhang, Z. Isolation and characterization of 2′-amino-modified RNA aptamers for human TNFα. Genom. Proteom. Bioinform. 2004, 2, 32–42. [Google Scholar] [CrossRef]
- Padilla, R.; Sousa, R. A Y639F/H784A T7 RNA polymerase double mutant displays superior properties for synthesizing RNAs with non-canonical NTPs. Nucleic Acids Res. 2002, 30, e138. [Google Scholar] [CrossRef] [PubMed]
- Dellafiore, M.A.; Montserrat, J.M.; Iribarren, A.M. Modified nucleoside triphosphates for in vitro selection techniques. Front. Chem. 2016, 4, 18. [Google Scholar] [CrossRef] [PubMed]
- Volk, D.E.; Lokesh, G.L.R. Development of phosphorothioate DNA and DNA thioaptamers. Biomedicines 2017, 5, 41. [Google Scholar] [CrossRef] [PubMed]
- Clercq, E.D.; Eckstein, F.; Merigan, T.C. Interferon induction increased through chemical modification of a synthetic polyribonucleotide. Science 1969, 165, 1137–1139. [Google Scholar] [CrossRef] [PubMed]
- Kruspe, S.; Giangrande, P.H. Aptamer-siRNA chimeras: Discovery, progress, and future prospects. Biomedicines 2017, 5, 45. [Google Scholar] [CrossRef] [PubMed]
- Bruno, J.G. A review of therapeutic aptamer conjugates with emphasis on new approaches. Pharmaceuticals 2013, 6, 340–357. [Google Scholar] [CrossRef] [PubMed]
- Ozalp, V.C.; Kavruk, M.; Dilek, O.; Bayrac, A.T. Aptamers: Molecular tools for medical diagnosis. Curr. Top. Med. Chem. 2015, 15, 1125–1137. [Google Scholar] [CrossRef] [PubMed]
- Seo, H.B.; Gu, M.B. Aptamer-based sandwich-type biosensors. J. Biol. Eng. 2017, 11, 11. [Google Scholar] [CrossRef] [PubMed]
- Shrivastava, S.; Jadon, N.; Jain, R. Next-generation polymer nanocomposite-based electrochemical sensors and biosensors: A review. TrAC Trends Anal. Chem. 2016, 82, 55–67. [Google Scholar] [CrossRef]
- Ma, H.; Liu, J.; Ali, M.M.; Mahmood, M.A.; Labanieh, L.; Lu, M.; Iqbal, S.M.; Zhang, Q.; Zhao, W.; Wan, Y. Nucleic acid aptamers in cancer research, diagnosis and therapy. Chem. Soc. Rev. 2015, 44, 1240–1256. [Google Scholar] [CrossRef] [PubMed]
- Mustonen, E.K.; Palomaki, T.; Pasanen, M. Oligonucleotide-based pharmaceuticals: Non-clinical and clinical safety signals and non-clinical testing strategies. Regul. Toxicol. Pharmacol. 2017, 90, 328–341. [Google Scholar] [CrossRef] [PubMed]
- Ouyang, W.; Yu, Z.; Zhao, X.; Lu, S.; Wang, Z. Aptamers in hematological malignancies and their potential therapeutic implications. Crit. Rev. Oncol. Hematol. 2016, 106, 108–117. [Google Scholar] [CrossRef] [PubMed]
- Parashar, A. Aptamers in therapeutics. J. Clin. Diagn. Res. 2016, 10, BE01-06. [Google Scholar] [CrossRef] [PubMed]
- Poolsup, S.; Kim, C.Y. Therapeutic applications of synthetic nucleic acid aptamers. Curr. Opin. Biotechnol. 2017, 48, 180–186. [Google Scholar] [CrossRef] [PubMed]
- Yazdian-Robati, R.; Arab, A.; Ramezani, M.; Abnous, K.; Taghdisi, S.M. Application of aptamers in treatment and diagnosis of leukemia. Int. J. Pharm. 2017, 529, 44–54. [Google Scholar] [CrossRef] [PubMed]
- Zhou, G.; Wilson, G.; Hebbard, L.; Duan, W.; Liddle, C.; George, J.; Qiao, L. Aptamers: A promising chemical antibody for cancer therapy. Oncotarget 2016, 7, 13446–13463. [Google Scholar] [CrossRef] [PubMed]
- Zhu, H.; Li, J.; Zhang, X.B.; Ye, M.; Tan, W. Nucleic acid aptamer-mediated drug delivery for targeted cancer therapy. ChemMedChem 2015, 10, 39–45. [Google Scholar] [CrossRef] [PubMed]
- Robbie, G.J.; Criste, R.; Dall’Acqua, W.F.; Jensen, K.; Patel, N.K.; Losonsky, G.A.; Griffin, M.P. A novel investigational fc-modified humanized monoclonal antibody, motavizumab-yte, has an extended half-life in healthy adults. Antimicrob. Agents Chemother. 2013, 57, 6147–6153. [Google Scholar] [CrossRef] [PubMed]
- Griffin, L.C.; Tidmarsh, G.F.; Bock, L.C.; Toole, J.J.; Leung, L.L. In vivo anticoagulant properties of a novel nucleotide-based thrombin inhibitor and demonstration of regional anticoagulation in extracorporeal circuits. Blood 1993, 81, 3271–3276. [Google Scholar] [PubMed]
- Tucker, C.E.; Chen, L.S.; Judkins, M.B.; Farmer, J.A.; Gill, S.C.; Drolet, D.W. Detection and plasma pharmacokinetics of an anti-vascular endothelial growth factor oligonucleotide-aptamer (nx1838) in rhesus monkeys. J. Chromatogr. B Biomed. Sci. Appl. 1999, 732, 203–212. [Google Scholar] [CrossRef]
- Drolet, D.W.; Nelson, J.; Tucker, C.E.; Zack, P.M.; Nixon, K.; Bolin, R.; Judkins, M.B.; Farmer, J.A.; Wolf, J.L.; Gill, S.C.; et al. Pharmacokinetics and safety of an anti-vascular endothelial growth factor aptamer (nx1838) following injection into the vitreous humor of rhesus monkeys. Pharm. Res. 2000, 17, 1503–1510. [Google Scholar] [CrossRef] [PubMed]
- Siddiqui, M.A.; Keating, G.M. Pegaptanib: In exudative age-related macular degeneration. Drugs 2005, 65, 1571–1577, discussion 1578–1579. [Google Scholar] [CrossRef] [PubMed]
- Derbyshire, N.; White, S.J.; Bunka, D.H.; Song, L.; Stead, S.; Tarbin, J.; Sharman, M.; Zhou, D.; Stockley, P.G. Toggled RNA aptamers against aminoglycosides allowing facile detection of antibiotics using gold nanoparticle assays. Anal. Chem. 2012, 84, 6595–6602. [Google Scholar] [CrossRef] [PubMed]
- Kuwahara, M.; Sugimoto, N. Molecular evolution of functional nucleic acids with chemical modifications. Molecules 2010, 15, 5423–5444. [Google Scholar] [CrossRef] [PubMed]
- Lebars, I.; Richard, T.; Di Primo, C.; Toulme, J.J. LNA derivatives of a kissing aptamer targeted to the trans-activating responsive RNA element of hiv-1. Blood Cells Mol. Dis. 2007, 38, 204–209. [Google Scholar] [CrossRef] [PubMed]
- Hernandez, F.J.; Stockdale, K.R.; Huang, L.; Horswill, A.R.; Behlke, M.A.; McNamara, J.O., 2nd. Degradation of nuclease-stabilized RNA oligonucleotides in mycoplasma-contaminated cell culture media. Nucleic Acid Ther. 2012, 22, 58–68. [Google Scholar] [CrossRef] [PubMed]
- Thiviyanathan, V.; Somasunderam, A.D.; Gorenstein, D.G. Combinatorial selection and delivery of thioaptamers. Biochem. Soc. Trans. 2007, 35, 50–52. [Google Scholar] [CrossRef] [PubMed]
- Volk, D.E.; Yang, X.; Fennewald, S.M.; King, D.J.; Bassett, S.E.; Venkitachalam, S.; Herzog, N.; Luxon, B.A.; Gorenstein, D.G. Solution structure and design of dithiophosphate backbone aptamers targeting transcription factor nf-kappab. Bioorg. Chem. 2002, 30, 396–419. [Google Scholar] [CrossRef]
- Maasch, C.; Buchner, K.; Eulberg, D.; Vonhoff, S.; Klussmann, S. Physicochemical stability of NOX-E36, a 40mer L-RNA (spiegelmer) for therapeutic applications. Nucleic Acids Symp. Ser. 2008, 52, 61–62. [Google Scholar] [CrossRef] [PubMed]
- Pfander, S.; Fiammengo, R.; Kirin, S.I.; Metzler-Nolte, N.; Jaschke, A. Reversible site-specific tagging of enzymatically synthesized RNAs using aldehyde-hydrazine chemistry and protease-cleavable linkers. Nucleic Acids Res. 2007, 35, e25. [Google Scholar] [CrossRef] [PubMed]
- Nitsche, A.; Kurth, A.; Dunkhorst, A.; Panke, O.; Sielaff, H.; Junge, W.; Muth, D.; Scheller, F.; Stocklein, W.; Dahmen, C.; et al. One-step selection of vaccinia virus-binding DNA aptamers by monolex. BMC Biotechnol. 2007, 7, 48. [Google Scholar] [CrossRef] [PubMed]
- Di Giusto, D.A.; Knox, S.M.; Lai, Y.; Tyrelle, G.D.; Aung, M.T.; King, G.C. Multitasking by multivalent circular DNA aptamers. Chembiochem 2006, 7, 535–544. [Google Scholar] [CrossRef] [PubMed]
- Mori, Y.; Nakamura, Y.; Ohuchi, S. Inhibitory RNA aptamer against sp6 RNA polymerase. Biochem. Biophys. Res. Commun. 2012, 420, 440–443. [Google Scholar] [CrossRef] [PubMed]
- Guo, P. The emerging field of RNA nanotechnology. Nat. Nanotechnol. 2010, 5, 833–842. [Google Scholar] [CrossRef] [PubMed]
- Burmeister, P.E.; Lewis, S.D.; Silva, R.F.; Preiss, J.R.; Horwitz, L.R.; Pendergrast, P.S.; McCauley, T.G.; Kurz, J.C.; Epstein, D.M.; Wilson, C.; et al. Direct in vitro selection of a 2′-O-methyl aptamer to VEGF. Chem. Biol. 2005, 12, 25–33. [Google Scholar] [CrossRef] [PubMed]
- Rusconi, C.P.; Roberts, J.D.; Pitoc, G.A.; Nimjee, S.M.; White, R.R.; Quick, G., Jr.; Scardino, E.; Fay, W.P.; Sullenger, B.A. Antidote-mediated control of an anticoagulant aptamer in vivo. Nat. Biotechnol. 2004, 22, 1423–1428. [Google Scholar] [CrossRef] [PubMed]
- Lee, C.H.; Lee, S.H.; Kim, J.H.; Noh, Y.H.; Noh, G.J.; Lee, S.W. Pharmacokinetics of a cholesterol-conjugated aptamer against the hepatitis C virus (HCV) NS5B protein. Mol. Ther. Nucleic Acids 2015, 4, e254. [Google Scholar] [CrossRef] [PubMed]
- Dougan, H.; Lyster, D.M.; Vo, C.V.; Stafford, A.; Weitz, J.I.; Hobbs, J.B. Extending the lifetime of anticoagulant oligodeoxynucleotide aptamers in blood. Nucl. Med. Biol. 2000, 27, 289–297. [Google Scholar] [CrossRef]
- Heo, K.; Min, S.W.; Sung, H.J.; Kim, H.G.; Kim, H.J.; Kim, Y.H.; Choi, B.K.; Han, S.; Chung, S.; Lee, E.S.; et al. An aptamer-antibody complex (oligobody) as a novel delivery platform for targeted cancer therapies. J. Control. Release 2016, 229, 1–9. [Google Scholar] [CrossRef] [PubMed]
- Kim, Y.; Dennis, D.M.; Morey, T.; Yang, L.; Tan, W. Engineering dendritic aptamer assemblies as superior inhibitors of protein function. Chem. Asian J. 2010, 5, 56–59. [Google Scholar] [CrossRef] [PubMed]
- Musumeci, D.; Montesarchio, D. Polyvalent nucleic acid aptamers and modulation of their activity: A focus on the thrombin binding aptamer. Pharmacol. Ther. 2012, 136, 202–215. [Google Scholar] [CrossRef] [PubMed]
- Soule, E.E.; Bompiani, K.M.; Woodruff, R.S.; Sullenger, B.A. Targeting two coagulation cascade proteases with a bivalent aptamer yields a potent and antidote-controllable anticoagulant. Nucleic Acid Ther. 2016, 26, 1–9. [Google Scholar] [CrossRef] [PubMed]
- Willis, M.C.; Collins, B.D.; Zhang, T.; Green, L.S.; Sebesta, D.P.; Bell, C.; Kellogg, E.; Gill, S.C.; Magallanez, A.; Knauer, S.; et al. Liposome-anchored vascular endothelial growth factor aptamers. Bioconj. Chem. 1998, 9, 573–582. [Google Scholar] [CrossRef] [PubMed]
- Zhou, J.; Soontornworajit, B.; Martin, J.; Sullenger, B.A.; Gilboa, E.; Wang, Y. A hybrid DNA aptamer-dendrimer nanomaterial for targeted cell labeling. Macromol. Biosci. 2009, 9, 831–835. [Google Scholar] [CrossRef] [PubMed]
- Chen, L.; Rashid, F.; Shah, A.; Awan, H.M.; Wu, M.; Liu, A.; Wang, J.; Zhu, T.; Luo, Z.; Shan, G. The isolation of an RNA aptamer targeting to p53 protein with single amino acid mutation. Proc. Natl. Acad. Sci. USA 2015, 112, 10002–10007. [Google Scholar] [CrossRef] [PubMed]
- Milla, P.; Dosio, F.; Cattel, L. Pegylation of proteins and liposomes: A powerful and flexible strategy to improve the drug delivery. Curr. Drug Metab. 2012, 13, 105–119. [Google Scholar] [CrossRef] [PubMed]
- Turecek, P.L.; Bossard, M.J.; Schoetens, F.; Ivens, I.A. Pegylation of biopharmaceuticals: A review of chemistry and nonclinical safety information of approved drugs. J. Pharm. Sci. 2016, 105, 460–475. [Google Scholar] [CrossRef] [PubMed]
- Pasut, G.; Veronese, F.M. State of the art in pegylation: The great versatility achieved after forty years of research. J. Control. Release 2012, 161, 461–472. [Google Scholar] [CrossRef] [PubMed]
- Morita, Y.; Kamal, M.; Kang, S.A.; Zhang, R.; Lokesh, G.L.; Thiviyanathan, V.; Hasan, N.; Woo, S.; Zhao, D.; Leslie, M.; et al. E-selectin targeting pegylated-thioaptamer prevents breast cancer metastases. Mol. Ther. Nucleic Acids 2016, 5, e399. [Google Scholar] [CrossRef] [PubMed]
- Ganson, N.J.; Povsic, T.J.; Sullenger, B.A.; Alexander, J.H.; Zelenkofske, S.L.; Sailstad, J.M.; Rusconi, C.P.; Hershfield, M.S. Pre-existing anti-polyethylene glycol antibody linked to first-exposure allergic reactions to pegnivacogin, a PEGylated RNA aptamer. J. Allergy Clin. Immunol. 2016, 137, 1610–1613. [Google Scholar] [CrossRef] [PubMed]
- Keefe, A.D.; Pai, S.; Ellington, A. Aptamers as therapeutics. Nat. Rev. Drug Discov. 2010, 9, 537–550. [Google Scholar] [CrossRef] [PubMed]
- Stessl, M.; Noe, C.R.; Winkler, J. Off-Target Effects and Safety Aspects of Phosphorothioate Oligonucleotides; Springer: Berlin/Heidelberg, Germany, 2012. [Google Scholar]
- Geary, R.S.; Yu, R.Z.; Watanabe, T.; Henry, S.P.; Hardee, G.E.; Chappell, A.; Matson, J.; Sasmor, H.; Cummins, L.; Levin, A.A. Pharmacokinetics of a tumor necrosis factor-alpha phosphorothioate 2′-O-(2-methoxyethyl) modified antisense oligonucleotide: Comparison across species. Drug Metab. Dispos. 2003, 31, 1419–1428. [Google Scholar] [CrossRef] [PubMed]
- Farman, C.A.; Kornbrust, D.J. Oligodeoxynucleotide studies in primates: Antisense and immune stimulatory indications. Toxicol. Pathol. 2003, 31 (Suppl. 1), 119–122. [Google Scholar] [PubMed]
- Goebl, N.; Berridge, B.; Wroblewski, V.J.; Brown-Augsburger, P.L. Development of a sensitive and specific in situ hybridization technique for the cellular localization of antisense oligodeoxynucleotide drugs in tissue sections. Toxicol. Pathol. 2007, 35, 541–548. [Google Scholar] [CrossRef] [PubMed]
- Sarmiento, U.M.; Perez, J.R.; Becker, J.M.; Narayanan, R. In vivo toxicological effects of rel a antisense phosphorothioates in CD-1 mice. Antisense Res. Dev. 1994, 4, 99–107. [Google Scholar] [CrossRef] [PubMed]
- Henry, S.P.; Zuckerman, J.E.; Rojko, J.; Hall, W.C.; Harman, R.J.; Kitchen, D.; Crooke, S.T. Toxicological properties of several novel oligonucleotide analogs in mice. Anticancer Drug Des. 1997, 12, 1–14. [Google Scholar] [PubMed]
- Bouchard, P.R.; Hutabarat, R.M.; Thompson, K.M. Discovery and development of therapeutic aptamers. Annu. Rev. Pharmacol. Toxicol. 2010, 50, 237–257. [Google Scholar] [CrossRef] [PubMed]
- Henry, S.P.; Taylor, J.; Midgley, L.; Levin, A.A.; Kornbrust, D.J. Evaluation of the toxicity of ISIS 2302, a phosphorothioate oligonucleotide, in a 4-week study in CD-1 mice. Antisense Nucleic Acid Drug Dev. 1997, 7, 473–481. [Google Scholar] [CrossRef] [PubMed]
- Henry, S.P.; Templin, M.V.; Gillett, N.; Rojko, J.; Levin, A.A. Correlation of toxicity and pharmacokinetic properties of a phosphorothioate oligonucleotide designed to inhibit ICAM-1. Toxicol. Pathol. 1999, 27, 95–100. [Google Scholar] [CrossRef] [PubMed]
- Kang, S.A.; Tsolmon, B.; Mann, A.P.; Zheng, W.; Zhao, L.; Zhao, Y.D.; Volk, D.E.; Lokesh, G.L.; Morris, L.; Gupta, V.; et al. Safety evaluation of intravenously administered mono-thioated aptamer against e-selectin in mice. Toxicol. Appl. Pharmacol. 2015, 287, 86–92. [Google Scholar] [CrossRef] [PubMed]
- Goel, S.; Desai, K.; Macapinlac, M.; Wadler, S.; Goldberg, G.; Fields, A.; Einstein, M.; Volterra, F.; Wong, B.; Martin, R.; et al. A phase I safety and dose escalation trial of docetaxel combined with GEM231, a second generation antisense oligonucleotide targeting protein kinase a R1α in patients with advanced solid cancers. Investig. New Drugs 2006, 24, 125–134. [Google Scholar] [CrossRef] [PubMed]
- Akdim, F.; Stroes, E.S.; Sijbrands, E.J.; Tribble, D.L.; Trip, M.D.; Jukema, J.W.; Flaim, J.D.; Su, J.; Yu, R.; Baker, B.F.; et al. Efficacy and safety of mipomersen, an antisense inhibitor of apolipoprotein B, in hypercholesterolemic subjects receiving stable statin therapy. J. Am. Coll. Cardiol. 2010, 55, 1611–1618. [Google Scholar] [CrossRef] [PubMed]
- Wong, E.; Goldberg, T. Mipomersen (kynamro): A novel antisense oligonucleotide inhibitor for the management of homozygous familial hypercholesterolemia. Pharm. Ther. 2014, 39, 119–122. [Google Scholar]
- Raal, F.J.; Santos, R.D.; Blom, D.J.; Marais, A.D.; Charng, M.J.; Cromwell, W.C.; Lachmann, R.H.; Gaudet, D.; Tan, J.L.; Chasan-Taber, S.; et al. Mipomersen, an apolipoprotein B synthesis inhibitor, for lowering of ldl cholesterol concentrations in patients with homozygous familial hypercholesterolaemia: A randomised, double-blind, placebo-controlled trial. Lancet 2010, 375, 998–1006. [Google Scholar] [CrossRef]
- McGowan, M.P.; Tardif, J.C.; Ceska, R.; Burgess, L.J.; Soran, H.; Gouni-Berthold, I.; Wagener, G.; Chasan-Taber, S. Randomized, placebo-controlled trial of mipomersen in patients with severe hypercholesterolemia receiving maximally tolerated lipid-lowering therapy. PLoS ONE 2012, 7, e49006. [Google Scholar] [CrossRef] [PubMed]
- Stein, E.A.; Dufour, R.; Gagne, C.; Gaudet, D.; East, C.; Donovan, J.M.; Chin, W.; Tribble, D.L.; McGowan, M. Apolipoprotein B synthesis inhibition with mipomersen in heterozygous familial hypercholesterolemia: Results of a randomized, double-blind, placebo-controlled trial to assess efficacy and safety as add-on therapy in patients with coronary artery disease. Circulation 2012, 126, 2283–2292. [Google Scholar] [CrossRef] [PubMed]
- Sheehan, J.P.; Lan, H.C. Phosphorothioate oligonucleotides inhibit the intrinsic tenase complex. Blood 1998, 92, 1617–1625. [Google Scholar] [PubMed]
- Yacyshyn, B.R.; Bowen-Yacyshyn, M.B.; Jewell, L.; Tami, J.A.; Bennett, C.F.; Kisner, D.L.; Shanahan, W.R., Jr. A placebo-controlled trial of ICAM-1 antisense oligonucleotide in the treatment of Crohn’s disease. Gastroenterology 1998, 114, 1133–1142. [Google Scholar] [CrossRef]
- Henry, S.P.; Beattie, G.; Yeh, G.; Chappel, A.; Giclas, P.; Mortari, A.; Jagels, M.A.; Kornbrust, D.J.; Levin, A.A. Complement activation is responsible for acute toxicities in rhesus monkeys treated with a phosphorothioate oligodeoxynucleotide. Int. Immunopharmacol. 2002, 2, 1657–1666. [Google Scholar] [CrossRef]
- Henry, S.P.; Jagels, M.A.; Hugli, T.E.; Manalili, S.; Geary, R.S.; Giclas, P.C.; Levin, A.A. Mechanism of alternative complement pathway dysregulation by a phosphorothioate oligonucleotide in monkey and human serum. Nucleic Acid Ther. 2014, 24, 326–335. [Google Scholar] [CrossRef] [PubMed]
- Avci-Adali, M.; Steinle, H.; Michel, T.; Schlensak, C.; Wendel, H.P. Potential capacity of aptamers to trigger immune activation in human blood. PLoS ONE 2013, 8, e68810. [Google Scholar] [CrossRef] [PubMed]
- Weber, C.; Muller, C.; Podszuweit, A.; Montino, C.; Vollmer, J.; Forsbach, A. Toll-like receptor (TLR) 3 immune modulation by unformulated small interfering RNA or DNA and the role of CD14 (in TLR-mediated effects). Immunology 2012, 136, 64–77. [Google Scholar] [CrossRef] [PubMed]
- Akira, S.; Uematsu, S.; Takeuchi, O. Pathogen recognition and innate immunity. Cell 2006, 124, 783–801. [Google Scholar] [CrossRef] [PubMed]
- Krieg, A.M. Mechanisms and applications of immune stimulatory CpG oligodeoxynucleotides. Biochim. Biophys. Acta 1999, 1489, 107–116. [Google Scholar] [CrossRef]
- Krieg, A.M. Therapeutic potential of toll-like receptor 9 activation. Nat. Rev. Drug Discov. 2006, 5, 471–484. [Google Scholar] [CrossRef] [PubMed]
- Klinman, D.M.; Yi, A.K.; Beaucage, S.L.; Conover, J.; Krieg, A.M. CpG motifs present in bacteria DNA rapidly induce lymphocytes to secrete interleukin 6, interleukin 12, and interferon gamma. Proc. Natl. Acad. Sci. USA 1996, 93, 2879–2883. [Google Scholar] [CrossRef] [PubMed]
- Krieg, A.M.; Yi, A.K.; Matson, S.; Waldschmidt, T.J.; Bishop, G.A.; Teasdale, R.; Koretzky, G.A.; Klinman, D.M. CpG motifs in bacterial DNA trigger direct B-cell activation. Nature 1995, 374, 546–549. [Google Scholar] [CrossRef] [PubMed]
- Foy, J.W.; Rittenhouse, K.; Modi, M.; Patel, M. Local tolerance and systemic safety of pegaptanib sodium in the dog and rabbit. J. Ocul. Pharmacol. Ther. 2007, 23, 452–466. [Google Scholar] [CrossRef] [PubMed]
- Cerchia, L.; Esposito, C.L.; Camorani, S.; Catuogno, S.; Franciscis, V. Coupling aptamers to short interfering RNAs as therapeutics. Pharmaceuticals 2011, 4, 1434–1449. [Google Scholar] [CrossRef] [PubMed]
- Armstrong, J.K.; Hempel, G.; Koling, S.; Chan, L.S.; Fisher, T.; Meiselman, H.J.; Garratty, G. Antibody against poly(ethylene glycol) adversely affects peg-asparaginase therapy in acute lymphoblastic leukemia patients. Cancer 2007, 110, 103–111. [Google Scholar] [CrossRef] [PubMed]
- Hershfield, M.S.; Ganson, N.J.; Kelly, S.J.; Scarlett, E.L.; Jaggers, D.A.; Sundy, J.S. Induced and pre-existing anti-polyethylene glycol antibody in a trial of every 3-week dosing of pegloticase for refractory gout, including in organ transplant recipients. Arthritis Res. Ther. 2014, 16, R63. [Google Scholar] [CrossRef] [PubMed]
- Akiyama, H.; Kachi, S.; Silva, R.L.; Umeda, N.; Hackett, S.F.; McCauley, D.; McCauley, T.; Zoltoski, A.; Epstein, D.M.; Campochiaro, P.A. Intraocular injection of an aptamer that binds PDGF-B: A potential treatment for proliferative retinopathies. J. Cell. Physiol. 2006, 207, 407–412. [Google Scholar] [CrossRef] [PubMed]
- Sennino, B.; Falcon, B.L.; McCauley, D.; Le, T.; McCauley, T.; Kurz, J.C.; Haskell, A.; Epstein, D.M.; McDonald, D.M. Sequential loss of tumor vessel pericytes and endothelial cells after inhibition of platelet-derived growth factor B by selective aptamer AX102. Cancer Res. 2007, 67, 7358–7367. [Google Scholar] [CrossRef] [PubMed]
- Kaur, H.; Li, J.J.; Bay, B.H.; Yung, L.Y. Investigating the antiproliferative activity of high affinity DNA aptamer on cancer cells. PLoS ONE 2013, 8, e50964. [Google Scholar] [CrossRef] [PubMed]
- Gantenbein, A.R.; Sarikaya, H.; Riederer, F.; Goadsby, P.J. Postoperative hemicrania continua-like headache—A case series. J. Headache Pain 2015, 16, 526. [Google Scholar] [CrossRef] [PubMed]
- Ireson, C.R.; Kelland, L.R. Discovery and development of anticancer aptamers. Mol. Cancer Ther. 2006, 5, 2957–2962. [Google Scholar] [CrossRef] [PubMed]
- Bates, P.J.; Laber, D.A.; Miller, D.M.; Thomas, S.D.; Trent, J.O. Discovery and development of the G-rich oligonucleotide AS1411 as a novel treatment for cancer. Exp. Mol. Pathol. 2009, 86, 151–164. [Google Scholar] [CrossRef] [PubMed]
- Mongelard, F.; Bouvet, P. As-1411, a guanosine-rich oligonucleotide aptamer targeting nucleolin for the potential treatment of cancer, including acute myeloid leukemia. Curr. Opin. Mol. Ther. 2010, 12, 107–114. [Google Scholar] [PubMed]
- Fan, X.; Sun, L.; Li, K.; Yang, X.; Cai, B.; Zhang, Y.; Zhu, Y.; Ma, Y.; Guan, Z.; Wu, Y.; et al. The bioactivity of d-/l-isonucleoside- and 2′-deoxyinosine-incorporated aptamer AS1411s including DNA replication/microrna expression. Mol. Ther. Nucleic Acids 2017, 9, 218–229. [Google Scholar] [CrossRef] [PubMed]
- Vater, A.; Sahlmann, J.; Kroger, N.; Zollner, S.; Lioznov, M.; Maasch, C.; Buchner, K.; Vossmeyer, D.; Schwoebel, F.; Purschke, W.G.; et al. Hematopoietic stem and progenitor cell mobilization in mice and humans by a first-in-class mirror-image oligonucleotide inhibitor of CXCL12. Clin. Pharmacol. Ther. 2013, 94, 150–157. [Google Scholar] [CrossRef] [PubMed]
- Liu, S.C.; Alomran, R.; Chernikova, S.B.; Lartey, F.; Stafford, J.; Jang, T.; Merchant, M.; Zboralski, D.; Zollner, S.; Kruschinski, A.; et al. Blockade of SDF-1 after irradiation inhibits tumor recurrences of autochthonous brain tumors in rats. Neuro Oncol. 2014, 16, 21–28. [Google Scholar] [CrossRef] [PubMed]
- Hoellenriegel, J.; Zboralski, D.; Maasch, C.; Rosin, N.Y.; Wierda, W.G.; Keating, M.J.; Kruschinski, A.; Burger, J.A. The spiegelmer NOX-A12, a novel CXCL12 inhibitor, interferes with chronic lymphocytic leukemia cell motility and causes chemosensitization. Blood 2014, 123, 1032–1039. [Google Scholar] [CrossRef] [PubMed]
- Zboralski, D.; Hoehlig, K.; Eulberg, D.; Fromming, A.; Vater, A. Increasing tumor-infiltrating T cells through inhibition of CXCL12 with NOX-A12 synergizes with PD-1 blockade. Cancer Immunol. Res. 2017, 5, 950–956. [Google Scholar] [CrossRef] [PubMed]
- Wan, Y.; Tamuly, D.; Allen, P.B.; Kim, Y.T.; Bachoo, R.; Ellington, A.D.; Iqbal, S.M. Proliferation and migration of tumor cells in tapered channels. Biomed. Microdevices 2013, 15, 635–643. [Google Scholar] [CrossRef] [PubMed]
- Buerger, C.; Nagel-Wolfrum, K.; Kunz, C.; Wittig, I.; Butz, K.; Hoppe-Seyler, F.; Groner, B. Sequence-specific peptide aptamers, interacting with the intracellular domain of the epidermal growth factor receptor, interfere with STAT3 activation and inhibit the growth of tumor cells. J. Biol. Chem. 2003, 278, 37610–37621. [Google Scholar] [CrossRef] [PubMed]
- Wang, D.L.; Song, Y.L.; Zhu, Z.; Li, X.L.; Zou, Y.; Yang, H.T.; Wang, J.J.; Yao, P.S.; Pan, R.J.; Yang, C.J.; et al. Selection of DNA aptamers against epidermal growth factor receptor with high affinity and specificity. Biochem. Biophys. Res. Commun. 2014, 453, 681–685. [Google Scholar] [CrossRef] [PubMed]
- Kim, M.Y.; Jeong, S. In vitro selection of RNA aptamer and specific targeting of ErbB2 in breast cancer cells. Nucleic Acid Ther. 2011, 21, 173–178. [Google Scholar] [CrossRef] [PubMed]
- Mahlknecht, G.; Maron, R.; Mancini, M.; Schechter, B.; Sela, M.; Yarden, Y. Aptamer to ErbB-2/HER2 enhances degradation of the target and inhibits tumorigenic growth. Proc. Natl. Acad. Sci. USA 2013, 110, 8170–8175. [Google Scholar] [CrossRef] [PubMed]
- Lee, Y.J.; Kim, I.S.; Park, S.A.; Kim, Y.; Lee, J.E.; Noh, D.Y.; Kim, K.T.; Ryu, S.H.; Suh, P.G. Periostin-binding DNA aptamer inhibits breast cancer growth and metastasis. Mol. Ther. 2013, 21, 1004–1013. [Google Scholar] [CrossRef] [PubMed]
- Hicke, B.J.; Stephens, A.W.; Gould, T.; Chang, Y.F.; Lynott, C.K.; Heil, J.; Borkowski, S.; Hilger, C.S.; Cook, G.; Warren, S.; et al. Tumor targeting by an aptamer. J. Nucl. Med. 2006, 47, 668–678. [Google Scholar] [PubMed]
- Daniels, D.A.; Chen, H.; Hicke, B.J.; Swiderek, K.M.; Gold, L. A tenascin-c aptamer identified by tumor cell selex: Systematic evolution of ligands by exponential enrichment. Proc. Natl. Acad. Sci. USA 2003, 100, 15416–15421. [Google Scholar] [CrossRef] [PubMed]
- Hicke, B.J.; Marion, C.; Chang, Y.F.; Gould, T.; Lynott, C.K.; Parma, D.; Schmidt, P.G.; Warren, S. Tenascin-C aptamers are generated using tumor cells and purified protein. J. Biol. Chem. 2001, 276, 48644–48654. [Google Scholar] [CrossRef] [PubMed]
- Zamay, T.N.; Kolovskaya, O.S.; Glazyrin, Y.E.; Zamay, G.S.; Kuznetsova, S.A.; Spivak, E.A.; Wehbe, M.; Savitskaya, A.G.; Zubkova, O.A.; Kadkina, A.; et al. DNA-aptamer targeting vimentin for tumor therapy in vivo. Nucleic Acid Ther. 2014, 24, 160–170. [Google Scholar] [CrossRef] [PubMed]
- Lee, Y.J.; Han, S.R.; Kim, N.Y.; Lee, S.H.; Jeong, J.S.; Lee, S.W. An RNA aptamer that binds carcinoembryonic antigen inhibits hepatic metastasis of colon cancer cells in mice. Gastroenterology 2012, 143, 155–165. [Google Scholar] [CrossRef] [PubMed]
- Ojima, A.; Matsui, T.; Maeda, S.; Takeuchi, M.; Inoue, H.; Higashimoto, Y.; Yamagishi, S. DNA aptamer raised against advanced glycation end products inhibits melanoma growth in nude mice. Lab. Investig. 2014, 94, 422–429. [Google Scholar] [CrossRef] [PubMed]
- Mi, J.; Zhang, X.; Rabbani, Z.N.; Liu, Y.; Reddy, S.K.; Su, Z.; Salahuddin, F.K.; Viles, K.; Giangrande, P.H.; Dewhirst, M.W.; et al. RNA aptamer-targeted inhibition of NF-κB suppresses non-small cell lung cancer resistance to doxorubicin. Mol. Ther. 2008, 16, 66–73. [Google Scholar] [CrossRef] [PubMed]
- Cerchia, L.; Esposito, C.L.; Camorani, S.; Rienzo, A.; Stasio, L.; Insabato, L.; Affuso, A.; de Franciscis, V. Targeting AXL with an high-affinity inhibitory aptamer. Mol. Ther. 2012, 20, 2291–2303. [Google Scholar] [CrossRef] [PubMed]
- Mi, Z.; Guo, H.; Russell, M.B.; Liu, Y.; Sullenger, B.A.; Kuo, P.C. RNA aptamer blockade of osteopontin inhibits growth and metastasis of mda-mb231 breast cancer cells. Mol. Ther. 2009, 17, 153–161. [Google Scholar] [CrossRef] [PubMed]
- Talbot, L.J.; Mi, Z.; Bhattacharya, S.D.; Kim, V.; Guo, H.; Kuo, P.C. Pharmacokinetic characterization of an RNA aptamer against osteopontin and demonstration of in vivo efficacy in reversing growth of human breast cancer cells. Surgery 2011, 150, 224–230. [Google Scholar] [CrossRef] [PubMed]
- Zhang, X.; Zhang, J.; Ma, Y.; Pei, X.; Liu, Q.; Lu, B.; Jin, L.; Wang, J.; Liu, J. A cell-based single-stranded DNA aptamer specifically targets gastric cancer. Int. J. Biochem. Cell Biol. 2014, 46, 1–8. [Google Scholar] [CrossRef] [PubMed]
- Cao, H.Y.; Yuan, A.H.; Chen, W.; Shi, X.S.; Miao, Y. A DNA aptamer with high affinity and specificity for molecular recognition and targeting therapy of gastric cancer. BMC Cancer 2014, 14, 699. [Google Scholar] [CrossRef] [PubMed]
- Li, S.; Wang, W.; Ding, H.; Xu, H.; Zhao, Q.; Li, J.; Li, H.; Xia, W.; Su, X.; Chen, Y.; et al. Aptamer BC15 against heterogeneous nuclear ribonucleoprotein A1 has potential value in diagnosis and therapy of hepatocarcinoma. Nucleic Acid Ther. 2012, 22, 391–398. [Google Scholar] [PubMed]
- Dassie, J.P.; Hernandez, L.I.; Thomas, G.S.; Long, M.E.; Rockey, W.M.; Howell, C.A.; Chen, Y.; Hernandez, F.J.; Liu, X.Y.; Wilson, M.E.; et al. Targeted inhibition of prostate cancer metastases with an RNA aptamer to prostate-specific membrane antigen. Mol. Ther. 2014, 22, 1910–1922. [Google Scholar] [CrossRef] [PubMed]
- Mann, A.P.; Somasunderam, A.; Nieves-Alicea, R.; Li, X.; Hu, A.; Sood, A.K.; Ferrari, M.; Gorenstein, D.G.; Tanaka, T. Identification of thioaptamer ligand against E-selectin: Potential application for inflamed vasculature targeting. PLoS ONE 2010, 5, e13050. [Google Scholar] [CrossRef] [PubMed]
- Kang, S.A.; Hasan, N.; Mann, A.P.; Zheng, W.; Zhao, L.; Morris, L.; Zhu, W.; Zhao, Y.D.; Suh, K.S.; Dooley, W.C.; et al. Blocking the adhesion cascade at the premetastatic niche for prevention of breast cancer metastasis. Mol. Ther. 2015, 23, 1044–1054. [Google Scholar] [CrossRef] [PubMed]
- Kang, S.A.; Blache, C.A.; Bajana, S.; Hasan, N.; Kamal, M.; Morita, Y.; Gupta, V.; Tsolmon, B.; Suh, K.S.; Gorenstein, D.G.; et al. The effect of soluble E-selectin on tumor progression and metastasis. BMC Cancer 2016, 16, 331. [Google Scholar]
- McNamara, J.O.; Kolonias, D.; Pastor, F.; Mittler, R.S.; Chen, L.; Giangrande, P.H.; Sullenger, B.; Gilboa, E. Multivalent 4-1BB binding aptamers costimulate CD8+ T cells and inhibit tumor growth in mice. J. Clin. Investig. 2008, 118, 376–386. [Google Scholar] [CrossRef] [PubMed]
- Dollins, C.M.; Nair, S.; Boczkowski, D.; Lee, J.; Layzer, J.M.; Gilboa, E.; Sullenger, B.A. Assembling OX40 aptamers on a molecular scaffold to create a receptor-activating aptamer. Chem. Biol. 2008, 15, 675–682. [Google Scholar] [CrossRef] [PubMed]
- Pratico, E.D.; Sullenger, B.A.; Nair, S.K. Identification and characterization of an agonistic aptamer against the T cell costimulatory receptor, OX40. Nucleic Acid Ther. 2013, 23, 35–43. [Google Scholar] [CrossRef] [PubMed]
- Pastor, F.; Soldevilla, M.M.; Villanueva, H.; Kolonias, D.; Inoges, S.; de Cerio, A.L.; Kandzia, R.; Klimyuk, V.; Gleba, Y.; Gilboa, E.; et al. CD28 aptamers as powerful immune response modulators. Mol. Ther. Nucleic Acids 2013, 2, e98. [Google Scholar] [CrossRef] [PubMed]
- Santulli-Marotto, S.; Nair, S.K.; Rusconi, C.; Sullenger, B.; Gilboa, E. Multivalent RNA aptamers that inhibit CTLA-4 and enhance tumor immunity. Cancer Res. 2003, 63, 7483–7489. [Google Scholar] [PubMed]
- Pastor, F.; Kolonias, D.; McNamara, J.O., 2nd; Gilboa, E. Targeting 4-1BB costimulation to disseminated tumor lesions with bi-specific oligonucleotide aptamers. Mol. Ther. 2011, 19, 1878–1886. [Google Scholar] [CrossRef] [PubMed]
- Eder, J.P.; Vande Woude, G.F.; Boerner, S.A.; LoRusso, P.M. Novel therapeutic inhibitors of the c-Met signaling pathway in cancer. Clin. Cancer Res. 2009, 15, 2207–2214. [Google Scholar] [CrossRef] [PubMed]
- Schrand, B.; Berezhnoy, A.; Brenneman, R.; Williams, A.; Levay, A.; Kong, L.Y.; Rao, G.; Zhou, S.; Heimberger, A.B.; Gilboa, E. Targeting 4-1BB costimulation to the tumor stroma with bispecific aptamer conjugates enhances the therapeutic index of tumor immunotherapy. Cancer Immunol. Res. 2014, 2, 867–877. [Google Scholar] [CrossRef] [PubMed]
- Prodeus, A.; Abdul-Wahid, A.; Fischer, N.W.; Huang, E.H.; Cydzik, M.; Gariepy, J. Targeting the PD-1/PD-L1 immune evasion axis with DNA aptamers as a novel therapeutic strategy for the treatment of disseminated cancers. Mol. Ther. Nucleic Acids 2015, 4, e237. [Google Scholar] [CrossRef] [PubMed]
- Lai, W.Y.; Huang, B.T.; Wang, J.W.; Lin, P.Y.; Yang, P.C. A novel PD-L1-targeting antagonistic DNA aptamer with antitumor effects. Mol. Ther. Nucleic Acids 2016, 5, e397. [Google Scholar] [CrossRef] [PubMed]
- Berezhnoy, A.; Stewart, C.A.; McNamara, J.O., 2nd; Thiel, W.; Giangrande, P.; Trinchieri, G.; Gilboa, E. Isolation and optimization of murine IL-10 receptor blocking oligonucleotide aptamers using high-throughput sequencing. Mol. Ther. 2012, 20, 1242–1250. [Google Scholar] [CrossRef] [PubMed]
- Roth, F.; De La Fuente, A.C.; Vella, J.L.; Zoso, A.; Inverardi, L.; Serafini, P. Aptamer-mediated blockade of IL4α triggers apoptosis of mdscs and limits tumor progression. Cancer Res. 2012, 72, 1373–1383. [Google Scholar] [CrossRef] [PubMed]
- Zheng, J.; Zhao, S.; Yu, X.; Huang, S.; Liu, H.Y. Simultaneous targeting of CD44 and epcam with a bispecific aptamer effectively inhibits intraperitoneal ovarian cancer growth. Theranostics 2017, 7, 1373–1388. [Google Scholar] [CrossRef] [PubMed]
- Hervas-Stubbs, S.; Soldevilla, M.M.; Villanueva, H.; Mancheno, U.; Bendandi, M.; Pastor, F. Identification of TIM3 2′-fluoro oligonucleotide aptamer by HT-SELEX for cancer immunotherapy. Oncotarget 2016, 7, 4522–4530. [Google Scholar] [CrossRef] [PubMed]
- Soldevilla, M.M.; Villanueva, H.; Bendandi, M.; Inoges, S.; Lopez-Diaz de Cerio, A.; Pastor, F. 2-fluoro-RNA oligonucleotide CD40 targeted aptamers for the control of B lymphoma and bone-marrow aplasia. Biomaterials 2015, 67, 274–285. [Google Scholar] [CrossRef] [PubMed]
- Huang, B.T.; Lai, W.Y.; Chang, Y.C.; Wang, J.W.; Yeh, S.D.; Lin, E.P.; Yang, P.C. A CTLA-4 antagonizing DNA aptamer with antitumor effect. Mol. Ther. Nucleic Acids 2017, 8, 520–528. [Google Scholar] [CrossRef] [PubMed]
- Ajona, D.; Ortiz-Espinosa, S.; Moreno, H.; Lozano, T.; Pajares, M.J.; Agorreta, J.; Bertolo, C.; Lasarte, J.J.; Vicent, S.; Hoehlig, K.; et al. A combined PD-1/C5a blockade synergistically protects against lung cancer growth and metastasis. Cancer Discov. 2017, 7, 694–703. [Google Scholar] [CrossRef] [PubMed]
- Drolet, D.W.; Green, L.S.; Gold, L.; Janjic, N. Fit for the eye: Aptamers in ocular disorders. Nucleic Acid Ther. 2016, 26, 127–146. [Google Scholar] [CrossRef] [PubMed]
- Eyetech Study, G. Preclinical and phase 1A clinical evaluation of an anti-VEGF pegylated aptamer (EYE001) for the treatment of exudative age-related macular degeneration. Retina 2002, 22, 143–152. [Google Scholar] [CrossRef]
- Zhang, H.; Berezov, A.; Wang, Q.; Zhang, G.; Drebin, J.; Murali, R.; Greene, M.I. ErbB receptors: From oncogenes to targeted cancer therapies. J. Clin. Investig. 2007, 117, 2051–2058. [Google Scholar] [CrossRef] [PubMed]
- Yarden, Y.; Sliwkowski, M.X. Untangling the ErbB signalling network. Nat. Rev. Mol. Cell Biol. 2001, 2, 127–137. [Google Scholar] [CrossRef] [PubMed]
- Gala, K.; Chandarlapaty, S. Molecular pathways: HER3 targeted therapy. Clin. Cancer Res. 2014, 20, 1410–1416. [Google Scholar] [CrossRef] [PubMed]
- Chen, C.H.; Chernis, G.A.; Hoang, V.Q.; Landgraf, R. Inhibition of heregulin signaling by an aptamer that preferentially binds to the oligomeric form of human epidermal growth factor receptor-3. Proc. Natl. Acad. Sci. USA 2003, 100, 9226–9231. [Google Scholar] [CrossRef] [PubMed]
- Berger, C.M.; Gaume, X.; Bouvet, P. The roles of nucleolin subcellular localization in cancer. Biochimie 2015, 113, 78–85. [Google Scholar] [CrossRef] [PubMed]
- Rosenberg, J.E.; Bambury, R.M.; Van Allen, E.M.; Drabkin, H.A.; Lara, P.N.; Harzstark, A.L.; Wagle, N.; Figlin, R.A.; Smith, G.W.; Garraway, L.A.; et al. A phase II trial of AS1411 (a novel nucleolin-targeted DNA aptamer) in metastatic renal cell carcinoma. Investig. New Drugs 2014, 32, 178–187. [Google Scholar] [CrossRef] [PubMed]
- Koutsioumpa, M.; Papadimitriou, E. Cell surface nucleolin as a target for anti-cancer therapies. Recent Pat. Anticancer Drug Discov. 2014, 9, 137–152. [Google Scholar] [CrossRef] [PubMed]
- Girvan, A.C.; Teng, Y.; Casson, L.K.; Thomas, S.D.; Jüliger, S.; Ball, M.W.; Klein, J.B.; Pierce, W.M.; Barve, S.S.; Bates, P.J. AGRO100 inhibits activation of nuclear factor-κB (NF-κB) by forming a complex with NF-κB essential modulator (NEMO) and nucleolin. Mol. Cancer Ther. 2006, 5, 1790–1799. [Google Scholar] [CrossRef] [PubMed]
- Soundararajan, S.; Chen, W.; Spicer, E.K.; Courtenay-Luck, N.; Fernandes, D.J. The nucleolin targeting aptamer AS1411 destabilizes Bcl-2 messenger RNA in human breast cancer cells. Cancer Res. 2008, 68, 2358–2365. [Google Scholar] [CrossRef] [PubMed]
- Jain, N.; Zhu, H.; Khashab, T.; Ye, Q.; George, B.; Mathur, R.; Singh, R.K.; Berkova, Z.; Wise, J.F.; Braun, F.K.; et al. Targeting nucleolin for better survival in diffuse large B-cell lymphoma. Leukemia 2017, 32, 663–674. [Google Scholar] [CrossRef] [PubMed]
- Gros, A.; Robbins, P.F.; Yao, X.; Li, Y.F.; Turcotte, S.; Tran, E.; Wunderlich, J.R.; Mixon, A.; Farid, S.; Dudley, M.E.; et al. PD-1 identifies the patient-specific CD8(+) tumor-reactive repertoire infiltrating human tumors. J. Clin. Investig. 2014, 124, 2246–2259. [Google Scholar] [CrossRef] [PubMed]
- Sayyed, S.G.; Hägele, H.; Kulkarni, O.P.; Endlich, K.; Segerer, S.; Eulberg, D.; Klussmann, S.; Anders, H.-J. Podocytes produce homeostatic chemokine stromal cell-derived factor-1/CXCL12, which contributes to glomerulosclerosis, podocyte loss and albuminuria in a mouse model of type 2 diabetes. Diabetologia 2009, 52, 2445–2454. [Google Scholar] [CrossRef] [PubMed]
- Marasca, R.; Maffei, R. NOX-A12: Mobilizing CLL away from home. Blood 2014, 123, 952–953. [Google Scholar] [CrossRef] [PubMed]
- Guo, F.; Wang, Y.; Liu, J.; Mok, S.C.; Xue, F.; Zhang, W. CXCL12/CXCR4: A symbiotic bridge linking cancer cells and their stromal neighbors in oncogenic communication networks. Oncogene 2016, 35, 816–826. [Google Scholar] [CrossRef] [PubMed]
- Feig, C.; Jones, J.O.; Kraman, M.; Wells, R.J.; Deonarine, A.; Chan, D.S.; Connell, C.M.; Roberts, E.W.; Zhao, Q.; Caballero, O.L.; et al. Targeting CXCL12 from FAP-expressing carcinoma-associated fibroblasts synergizes with anti-PD-L1 immunotherapy in pancreatic cancer. Proc. Natl. Acad. Sci. USA 2013, 110, 20212–20217. [Google Scholar] [CrossRef] [PubMed]
- Mann, A.P.; Bhavane, R.C.; Somasunderam, A.; Liz Montalvo-Ortiz, B.; Ghaghada, K.B.; Volk, D.; Nieves-Alicea, R.; Suh, K.S.; Ferrari, M.; Annapragada, A.; et al. Thioaptamer conjugated liposomes for tumor vasculature targeting. Oncotarget 2011, 2, 298–304. [Google Scholar] [CrossRef] [PubMed]
- Mann, A.P.; Tanaka, T.; Somasunderam, A.; Liu, X.; Gorenstein, D.G.; Ferrari, M. E-selectin-targeted porous silicon particle for nanoparticle delivery to the bone marrow. Adv. Mater. 2011, 23, H278–H282. [Google Scholar] [CrossRef] [PubMed]
- Mai, J.; Huang, Y.; Mu, C.; Zhang, G.; Xu, R.; Guo, X.; Xia, X.; Volk, D.E.; Lokesh, G.L.; Thiviyanathan, V.; et al. Bone marrow endothelium-targeted therapeutics for metastatic breast cancer. J. Control. Release 2014, 187, 22–29. [Google Scholar] [CrossRef] [PubMed]
- Nishino, M.; Ramaiya, N.H.; Hatabu, H.; Hodi, F.S. Monitoring immune-checkpoint blockade: Response evaluation and biomarker development. Nat. Rev. Clin. Oncol. 2017, 14, 655–668. [Google Scholar] [CrossRef] [PubMed]
- Mangala, L.S.; Wang, H.; Jiang, D.; Wu, S.Y.; Somasunderam, A.; Volk, D.E.; Lokesh, G.L.; Li, X.; Pradeep, S.; Yang, X.; et al. Improving vascular maturation using noncoding RNAs increases antitumor effect of chemotherapy. JCI Insight 2016, 1, e87754. [Google Scholar] [CrossRef] [PubMed]
- Melero, I.; Hervas-Stubbs, S.; Glennie, M.; Pardoll, D.M.; Chen, L. Immunostimulatory monoclonal antibodies for cancer therapy. Nat. Rev. Cancer 2007, 7, 95–106. [Google Scholar] [CrossRef] [PubMed]
- Attarwala, H. TGN1412: From discovery to disaster. J. Young Pharm. 2010, 2, 332–336. [Google Scholar] [CrossRef] [PubMed]
- Bartkowiak, T.; Curran, M.A. 4-1BB agonists: Multi-potent potentiators of tumor immunity. Front. Oncol. 2015, 5, 117. [Google Scholar] [CrossRef] [PubMed]
- Schrand, B.; Berezhnoy, A.; Brenneman, R.; Williams, A.; Levay, A.; Gilboa, E. Reducing toxicity of 4-1BB costimulation: Targeting 4-1BB ligands to the tumor stroma with BI-specific aptamer conjugates. Oncoimmunology 2015, 4, e970918. [Google Scholar] [CrossRef] [PubMed]
- Soldevilla, M.M.; Villanueva, H.; Casares, N.; Lasarte, J.J.; Bendandi, M.; Inoges, S.; Lopez-Diaz de Cerio, A.; Pastor, F. MRP1-CD28 bi-specific oligonucleotide aptamers: Target costimulation to drug-resistant melanoma cancer stem cells. Oncotarget 2016, 7, 23182–23196. [Google Scholar] [CrossRef] [PubMed]
- Rusconi, C.P.; Scardino, E.; Layzer, J.; Pitoc, G.A.; Ortel, T.L.; Monroe, D.; Sullenger, B.A. RNA aptamers as reversible antagonists of coagulation factor IXa. Nature 2002, 419, 90–94. [Google Scholar] [CrossRef] [PubMed]
- Bompiani, K.M.; Monroe, D.M.; Church, F.C.; Sullenger, B.A. A high affinity, antidote-controllable prothrombin and thrombin-binding RNA aptamer inhibits thrombin generation and thrombin activity. J. Thromb. Haemost. 2012, 10, 870–880. [Google Scholar] [CrossRef] [PubMed]
- Ng, E.W.M.; Shima, D.T.; Calias, P.; Cunningham, E.T., Jr.; Guyer, D.R.; Adamis, A.P. Pegaptanib, a targeted anti-VEGF aptamer for ocular vascular disease. Nat. Rev. Drug Dis. 2006, 5, 123–132. [Google Scholar] [CrossRef] [PubMed]
- Jellinek, D.; Green, L.S.; Bell, C.; Janjic, N. Inhibition of receptor binding by high-affinity RNA ligands to vascular endothelial growth factor. Biochemistry 1994, 33, 10450–10456. [Google Scholar] [CrossRef] [PubMed]
- Green, L.S.; Jellinek, D.; Bell, C.; Beebe, L.A.; Feistner, B.D.; Gill, S.C.; Jucker, F.M.; Janjić, N. Nuclease-resistant nucleic acid ligands to vascular permeability factor/vascular endothelial growth factor. Chem. Biol. 1995, 2, 683–695. [Google Scholar] [CrossRef]
- Study of AS1411 in Advanced Solid Tumours. Available online: https://ClinicalTrials.gov/show/NCT00881244 (accessed on 12 December 2017).
- A phase II Study of AS1411 in Renal Cell Carcinoma. Available online: https://ClinicalTrials.gov/show/NCT00740441 (accessed on 12 December 2017).
- Phase II Study of AS1411 Combined with Cytarabine to Treat Acute Myeloid Leukemia. Available online: https: //ClinicalTrials.gov/show/NCT00512083 (accessed on 12 December 2017).
- Soundararajan, S.; Wang, L.; Sridharan, V.; Chen, W.; Courtenay-Luck, N.; Jones, D.; Spicer, E.K.; Fernandes, D.J. Plasma membrane nucleolin is a receptor for the anticancer aptamer AS1411 in MV4-11 leukemia cells. Mol. Pharmacol. 2009, 76, 984–991. [Google Scholar] [CrossRef] [PubMed]
- Chen, W.; Sridharan, V.; Soundararajan, S.; Otake, Y.; Stuart, R.; Jones, D.; Fernandes, D. Activity and mechanism of action of AS1411 in acute myeloid leukemia cells. Blood 2007, 110, 1604. [Google Scholar]
- Stuart, R.K.; Stockerl-Goldstein, K.; Cooper, M.; Devetten, M.; Herzig, R.; Medeiros, B.; Schiller, G.; Wei, A.; Acton, G.; Rizzieri, D. Randomized phase ii trial of the nucleolin targeting aptamer AS1411 combined with high-dose cytarabine in relapsed/refractory acute myeloid leukemia (AML). J. Clin. Oncol. 2009, 27, 7019. [Google Scholar]
- Olaptesed (NOX-A12) Alone and in Combination with Pembrolizumab in Colorectal and Pancreatic Cancer. Available online: https://ClinicalTrials.gov/show/NCT03168139 (accessed on 12 December 2017).
- NOX-A12 First-in-Human (FIH) Study. Available online: https://ClinicalTrials.gov/show/NCT00976378 (accessed on 12 December 2017).
- NOX-A12 Multiple Ascending Dose Study in Healthy Volunteers. Available online: https://ClinicalTrials.gov/show/NCT01194934 (accessed on 12 December 2017).
- Waldschmidt, J.M.; Simon, A.; Wider, D.; Muller, S.J.; Follo, M.; Ihorst, G.; Decker, S.; Lorenz, J.; Chatterjee, M.; Azab, A.K.; et al. Cxcl12 and Cxcr7 are relevant targets to reverse cell adhesion-mediated drug resistance in multiple myeloma. Br. J. Haematol. 2017, 179, 36–49. [Google Scholar] [CrossRef] [PubMed]
- Roccaro, A.M.; Sacco, A.; Purschke, W.G.; Moschetta, M.; Buchner, K.; Maasch, C.; Zboralski, D.; Zollner, S.; Vonhoff, S.; Mishima, Y.; et al. SDF-1 inhibition targets the bone marrow niche for cancer therapy. Cell Rep. 2014, 9, 118–128. [Google Scholar] [CrossRef] [PubMed]
- NOX-A12 in Combination with Bortezomib and Dexamethasone in Relapsed Multiple Myeloma. Available online: https://ClinicalTrials.gov/show/NCT01521533 (accessed on 12 December 2017).
- Ludwig, H.; Weisel, K.; Petrucci, M.T.; Leleu, X.; Cafro, A.M.; Garderet, L.; Leitgeb, C.; Foa, R.; Greil, R.; Yakoub-Agha, I.; et al. Olaptesed pegol, an anti-CXCL12/SDF-1 spiegelmer, alone and with bortezomib-dexamethasone in relapsed/refractory multiple myeloma: A phase IIa study. Leukemia 2017, 31, 997–1000. [Google Scholar] [CrossRef] [PubMed]
- NOX-A12 in Combination with Bendamustine and Rituximab in Relapsed Chronic Lymphocytic Leukemia (CLL). Available online: https://ClinicalTrials.gov/show/NCT01486797 (accessed on 12 December 2017).
- Steurer, M.; Montillo, M.; Scarfò, L.; Mauro, F.R.; Andel, J.; Wildner, S.; Trentin, L.; Janssens, A.; Burgstaller, S.; Kruschinski, A.; et al. Results from a phase IIa study of the anti-CXCL12 spiegelmer olaptesed pegol (NOX-A12) in combination with bendamustine/rituximab in patients with chronic lymphocytic leukemia. Blood 2014, 124, 1996. [Google Scholar]
- Buffery, D. The 2015 oncology drug pipeline: Innovation drives the race to cure cancer. Am. Health Drug Benefits 2015, 8, 216–222. [Google Scholar] [PubMed]
| Aptamer Name | Targets | References |
|---|---|---|
| ARC126 (RNA) AX102 (RNA) | PDGF-B | Akiyama, Kachi et al., 2006 [96], Sennino, Falcon et al., 2007 [97] |
| SL (2)-B (DNA) RNV66 (DNA) | VEGF-165 | Kaur, Li et al., 2013 [98], Gantenbein, Sarikaya et al., 2015 [99] |
| AS1411 (DNA) | Nucleolin | Ireson and Kelland 2006 [100], Bates, Laber et al., 2009 [101], Mongelard and Bouvet 2010 [102] |
| FCL-II (DNA, modified form AS1411) | Fan, Sun et al., 2017 [103] | |
| NOX-A12 (RNA) | CXCL12 | Vater, Sahlmann et al., 2013 [104], Liu, Alomran et al., 2014 [105], Hoellenriegel, Zboralski et al., 2014 [106], Zboralski, Hoehlig et al., 2017 [107] |
| E0727 (RNA) CL428 (RNA) KDI130 (RNA) TuTu2231 (RNA) | EGFR | Li, Nguyen et al., 2011 [8], Esposito, Passaro et al., 2011 [7], Wan, Tamuly et al., 2013 [108], Buerger, Nagel-Wolfrum et al., 2003 [109], Wang, Song et al., 2014 [110] |
| Trimeric apt (DNA) | HER2 | Kim and Jeong 2011 [111], Mahlknecht, Maron et al., 2013 [112] |
| PNDA-3 (DNA) | Periostin | Lee, Kim et al., 2013 [113] |
| TTA140,41 (DNA) GBI-1042 (DNA) | TN-C | Hicke, Stephens et al., 2006 [114], Daniels, Chen et al., 2003 [115], Hicke, Marion et al., 2001 [116] |
| NAS-24 (DNA) | Vimentin | Zamay, Kolovskaya et al., 2014 [117] |
| YJ-1 (RNA) | CEA | Lee, Han et al., 2012 [118] |
| AGE-apt (DNA) | AGE | Ojima, Matsui et al., 2014 [119] |
| A-P50 (RNA) | NF-κB | Mi, Zhang et al., 2008 [120] |
| GL21.T (RNA) | Axl | Cerchia, Esposito et al., 2012 [121] |
| OPN-R3 (RNA) | OPN | Mi, Guo et al., 2009 [122], Talbot, Mi et al., 2011 [123] |
| AGC03 (DNA) cy-apt (DNA) | HGC-27 | Zhang, Zhang et al., 2014 [124], Cao, Yuan et al., 2014 [125] |
| BC15 (DNA) | hnRNP A1 | Li, Wang et al., 2012 [126] |
| A9g (RNA) | PSMA | Dassie, Hernandez et al., 2014 [127] |
| ESTA (DNA) | E-selectin | Mann, Somasunderam et al., 2010 [128], Kang, Hasan et al., 2015 [129], Kang, Blache et al., 2016 [130], Morita, Kamal et al., 2016 [62] |
| M12-23 (RNA) | 4-1 BB | McNamara, Kolonias et al., 2008 [131] |
| OX40-apt (RNA) | OX40 | Dollins, Nair et al., 2008 [132], Pratico, Sullenger et al., 2013 [133] |
| CD28-apt (RNA) | CD28 | Pastor, Soldevilla et al., 2013 [134] |
| Del60 (RNA) | CTLA-4 | Santulli-Marotto, Nair et al., 2003 [135] |
| PSMA-4-1BB-apt (RNA) | PSMA/4-1BB | Pastor, Kolonias et al., 2011 [136] |
| CD16α/c-Met-apt (RNA) | CD16α/c-Met | Eder, Vande Woude et al., 2009 [137] |
| VEGF-4-1BB apt (DNA) | VEGF/4-1BB | Schrand, Berezhnoy et al., 2014 [138] |
| MP7 (DNA) | PD-1 | Prodeus, Abdul-Wahid et al., 2015 [139] |
| aptPD-L1 (DNA) | PD-L1 | Lai, Huang et al., 2016 [140] |
| R5A1 (RNA) | IL10R | Berezhnoy, Stewart et al., 2012 [141] |
| CL-42 (RNA) | IL4Rα | Roth, De La Fuente et al., 2012 [142] |
| CD44-EpCAM aptamer (RNA) | CD44/EpCAM | Zheng, Zhao et al., 2017 [143] |
| TIM3Apt (RNA) | TIM3 | Hervas-Stubbs, Soldevilla et al., 2016 [144] |
| CD40apt (RNA) | CD40 | Soldevilla, Villanueva et al., 2015 [145] |
| AptCTLA-4 (DNA) | CTLA-4 | Huang, Lai et al., 2017 [146] |
| AON-D21 l-Aptamer (RNA/DNA) | C5a | Ajona, Ortiz-Espinosa et al., 2017 [147] |
© 2018 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Morita, Y.; Leslie, M.; Kameyama, H.; Volk, D.E.; Tanaka, T. Aptamer Therapeutics in Cancer: Current and Future. Cancers 2018, 10, 80. https://doi.org/10.3390/cancers10030080
Morita Y, Leslie M, Kameyama H, Volk DE, Tanaka T. Aptamer Therapeutics in Cancer: Current and Future. Cancers. 2018; 10(3):80. https://doi.org/10.3390/cancers10030080
Chicago/Turabian StyleMorita, Yoshihiro, Macall Leslie, Hiroyasu Kameyama, David E. Volk, and Takemi Tanaka. 2018. "Aptamer Therapeutics in Cancer: Current and Future" Cancers 10, no. 3: 80. https://doi.org/10.3390/cancers10030080
APA StyleMorita, Y., Leslie, M., Kameyama, H., Volk, D. E., & Tanaka, T. (2018). Aptamer Therapeutics in Cancer: Current and Future. Cancers, 10(3), 80. https://doi.org/10.3390/cancers10030080






