3-Dimensional Plasmonic Substrates Based on Chicken Eggshell Bio-Templates for SERS-Based Bio-Sensing
Abstract
:1. Introduction
2. Methods
2.1. Preparation of Surface-Enhanced Raman Scattering (SERS) Substrates
2.2. Preparation of Bacterial Suspensions
2.3. Fabrication of 3D Flexible SERS Substrates
2.4. SERS Measurements of Benzenethiol (BT), Rhodamine 6G (R6G), and Bacteria
2.5. Morphological and Molecular Imaging
3. Results and Discussion
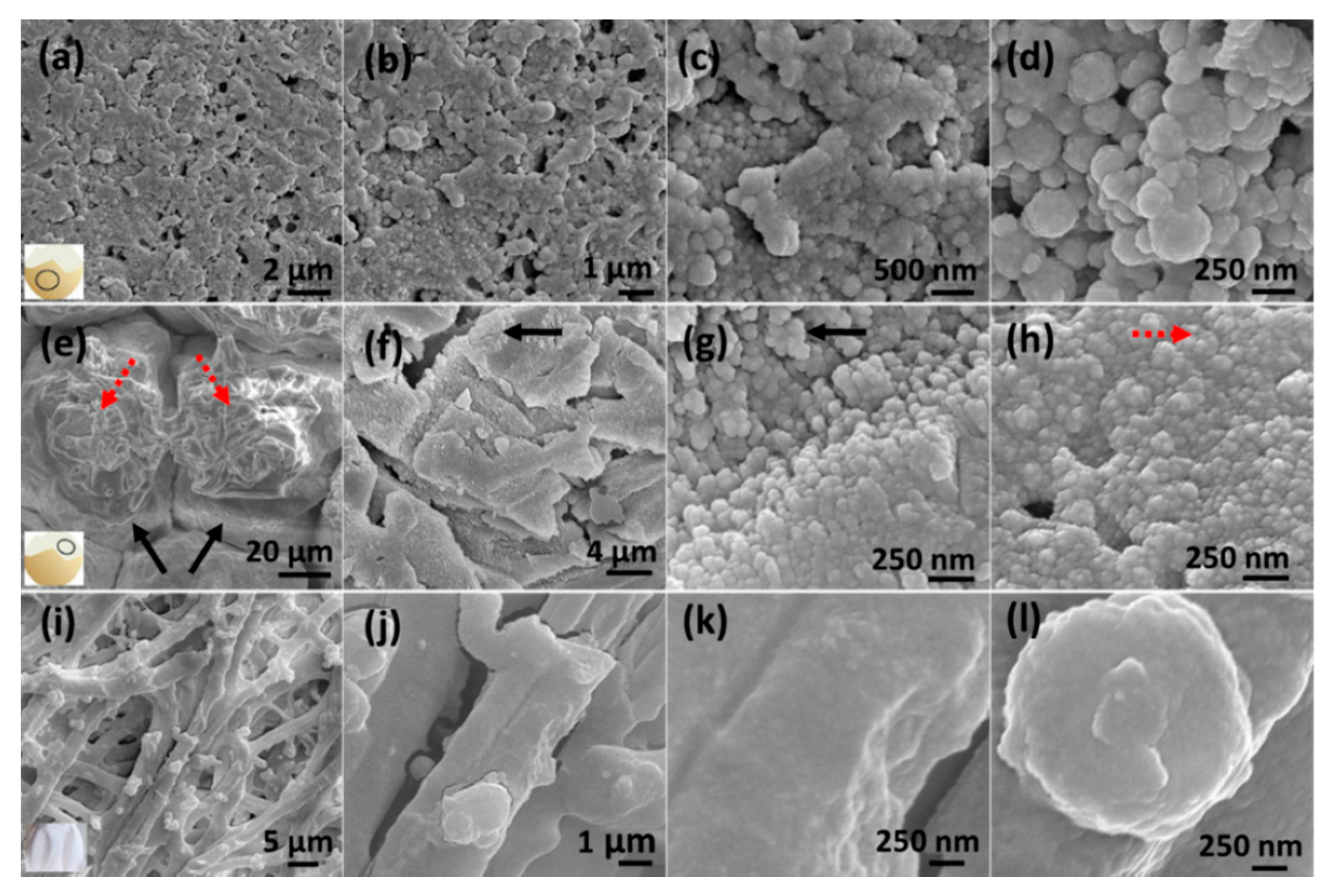
3.1. Morphological Characterization
3.2. SERS Characterization
3.3. Bio-Sensing Application
3.4. Eggshell for 3D Flexible SERS Substrate
4. Conclusions
Supplementary Materials
Acknowledgments
Author Contributions
Conflicts of Interest
References
- Albrecht, M.G.; Creighton, J.A. Anomalously intense Raman spectra of pyridine at a silver electrode. J. Am. Chem. Soc. 1977, 99, 5215–5217. [Google Scholar] [CrossRef]
- Kneipp, K.; Kneipp, H.; Itzkan, I.; Dasari, R.R.; Feld, M.S. Ultrasensitive chemical analysis by Raman spectroscopy. Chem. Rev. 1999, 99, 2957–2976. [Google Scholar] [CrossRef] [PubMed]
- Lyandres, O.; Shah, N.C.; Yonzon, C.R.; Walsh, J.T.; Glucksberg, M.R.; Van Duyne, R.P. Real-time glucose sensing by surface-enhanced Raman spectroscopy in bovine plasma facilitated by a mixed decanethiol/mercaptohexanol partition layer. Anal. Chem. 2005, 77, 6134–6139. [Google Scholar] [CrossRef] [PubMed]
- Levin, C.S.; Bishnoi, S.W.; Grady, N.K.; Halas, N.J. Determining the conformation of thiolated poly (ethylene glycol) on au nanoshells by surface-enhanced Raman scattering spectroscopic assay. Anal. Chem. 2006, 78, 3277–3281. [Google Scholar] [CrossRef] [PubMed]
- Yang, S.; Dai, X.; Stogin, B.B.; Wong, T.-S. Ultrasensitive surface-enhanced Raman scattering detection in common fluids. Proc. Natl. Acad. Sci. USA 2016, 113, 268–273. [Google Scholar] [CrossRef] [PubMed]
- Nie, S.; Emory, S.R. Probing single molecules and single nanoparticles by surface-enhanced Raman scattering. Science 1997, 275, 1102–1106. [Google Scholar] [CrossRef] [PubMed]
- Qi, J.; Zeng, J.; Zhao, F.; Lin, S.H.; Raja, B.; Strych, U.; Willson, R.C.; Shih, W.-C. Label-free, in situ SERS monitoring of individual DNA hybridization in microfluidics. Nanoscale 2014, 6, 8521–8526. [Google Scholar] [CrossRef] [PubMed]
- Lee, S.J.; Morrill, A.R.; Moskovits, M. Hot spots in silver nanowire bundles for surface-enhanced Raman spectroscopy. J. Am. Chem. Soc. 2006, 128, 2200–2201. [Google Scholar] [CrossRef] [PubMed]
- Chen, Y.-H.; Huang, L.; Gan, L.; Li, Z.-Y. Wavefront shaping of infrared light through a subwavelength hole. Light Sci. Appl. 2012, 1, e26. [Google Scholar] [CrossRef]
- Wang, L.; Xiong, W.; Nishijima, Y.; Yokota, Y.; Ueno, K.; Misawa, H.; Qiu, J.; Bi, G. Spectral properties of nanoengineered Ag/Au bilayer rods fabricated by electron beam lithography. Appl. Opt. 2011, 50, 5600–5605. [Google Scholar] [CrossRef] [PubMed]
- Xu, B.B.; Zhang, Y.L.; Zhang, W.Y.; Liu, X.Q.; Wang, J.N.; Zhang, X.L.; Zhang, D.D.; Jiang, H.B.; Zhang, R.; Sun, H.B. Silver-coated rose petal: Green, facile, low-cost and sustainable fabrication of a SERS substrate with unique superhydrophobicity and high efficiency. Adv. Opt. Mater. 2013, 1, 56–60. [Google Scholar] [CrossRef]
- Shih, W.-C.; Santos, G.M.; Zhao, F.; Zenasni, O.; Arnob, M.M.P. Simultaneous chemical and refractive index sensing in the 1–2.5 μm near-infrared wavelength range on nanoporous gold disks. Nano Lett. 2016, 16, 4641–4647. [Google Scholar] [CrossRef] [PubMed]
- Talley, C.E.; Jackson, J.B.; Oubre, C.; Grady, N.K.; Hollars, C.W.; Lane, S.M.; Huser, T.R.; Nordlander, P.; Halas, N.J. Surface-enhanced Raman scattering from individual Au nanoparticles and nanoparticle dimer substrates. Nano Lett. 2005, 5, 1569–1574. [Google Scholar] [CrossRef] [PubMed]
- Schwartzberg, A.M.; Grant, C.D.; Wolcott, A.; Talley, C.E.; Huser, T.R.; Bogomolni, R.; Zhang, J.Z. Unique gold nanoparticle aggregates as a highly active surface-enhanced Raman scattering substrate. J. Phys. Chem. B 2004, 108, 19191–19197. [Google Scholar] [CrossRef]
- Felidj, N.; Aubard, J.; Levi, G.; Krenn, J.; Hohenau, A.; Schider, G.; Leitner, A.; Aussenegg, F. Optimized surface-enhanced Raman scattering on gold nanoparticle arrays. Appl. Phys. Lett. 2003, 82, 3095–3097. [Google Scholar] [CrossRef]
- Haynes, C.L.; Van Duyne, R.P. Nanosphere lithography: A versatile nanofabrication tool for studies of size-dependent nanoparticle optics. J. Phys. Chem. B 2001, 105, 5599–5611. [Google Scholar] [CrossRef]
- Fan, M.; Andrade, G.F.; Brolo, A.G. A review on the fabrication of substrates for surface enhanced Raman spectroscopy and their applications in analytical chemistry. Anal. Chim. Acta 2011, 693, 7–25. [Google Scholar] [CrossRef] [PubMed]
- Stiles, P.L.; Dieringer, J.A.; Shah, N.C.; Van Duyne, R.P. Surface-enhanced Raman spectroscopy. Ann. Rev. Anal. Chem. 2008, 1, 601–626. [Google Scholar] [CrossRef] [PubMed]
- Stoddart, P.; Cadusch, P.; Boyce, T.; Erasmus, R.; Comins, J. Optical properties of chitin: Surface-enhanced Raman scattering substrates based on antireflection structures on cicada wings. Nanotechnology 2006, 17, 680. [Google Scholar] [CrossRef]
- Garrett, N.L.; Vukusic, P.; Ogrin, F.; Sirotkin, E.; Winlove, C.P.; Moger, J. Spectroscopy on the wing: Naturally inspired SERS substrates for biochemical analysis. J. Biophotonics 2009, 2, 157. [Google Scholar] [CrossRef] [PubMed]
- Zheng, B.; Qian, L.; Yuan, H.; Xiao, D.; Yang, X.; Paau, M.C.; Choi, M.M. Preparation of gold nanoparticles on eggshell membrane and their biosensing application. Talanta 2010, 82, 177–183. [Google Scholar] [CrossRef] [PubMed]
- Kumar, G.P. Gold nanoparticle-coated biomaterial as SERS micro-probes. Bull. Mater. Sci. 2011, 34, 417–422. [Google Scholar] [CrossRef]
- Tan, Y.; Zang, X.; Gu, J.; Liu, D.; Zhu, S.; Su, H.; Feng, C.; Liu, Q.; Lau, W.M.; Moon, W.-J.; et al. Morphological effects on surface-enhanced Raman scattering from silver butterfly wing scales synthesized via photoreduction. Langmuir 2011, 27, 11742–11746. [Google Scholar] [CrossRef] [PubMed]
- Lin, P.Y.; Hsieh, C.W.; Tsai, P.C.; Hsieh, S. Porosity-controlled eggshell membrane as 3D SERS-active substrate. ChemPhysChem 2014, 15, 1577–1580. [Google Scholar] [CrossRef] [PubMed]
- Tanahashi, I.; Harada, Y. Silver nanoparticles deposited on TiO2-coated cicada and butterfly wings as naturally inspired SERS substrates. J. Mater. Chem. C 2015, 3, 5721–5726. [Google Scholar] [CrossRef]
- Wang, M.; Meng, G.; Huang, Q.; Tang, H.; Li, Z.; Zhang, Z. CNTs-anchored eggshell membrane decorated with Ag-NPs as cheap but effective SERS substrates. Sci. China Mater. 2015, 58, 198–203. [Google Scholar] [CrossRef]
- Zeng, J.; Zhao, F.; Li, M.; Li, C.-H.; Lee, T.R.; Shih, W.-C. Morphological control and plasmonic tuning of nanoporous gold disks by surface modifications. J. Mater. Chem. C 2015, 3, 247–252. [Google Scholar] [CrossRef]
- Jeong, J.W.; Arnob, M.M.P.; Baek, K.M.; Lee, S.Y.; Shih, W.C.; Jung, Y.S. 3D cross-point plasmonic nanoarchitectures containing dense and regular hot spots for surface-enhanced Raman spectroscopy analysis. Adv. Mater. 2016, 28, 8695–8704. [Google Scholar] [CrossRef] [PubMed]
- Zhou, J.; Wang, S.; Nie, F.; Feng, L.; Zhu, G.; Jiang, L. Elaborate architecture of the hierarchical hen’s eggshell. Nano Res. 2011, 4, 171–179. [Google Scholar] [CrossRef]
- Patel, I.; Premasiri, W.; Moir, D.; Ziegler, L. Barcoding bacterial cells: A SERS-based methodology for pathogen identification. J. Raman Spectrosc. 2008, 39, 1660–1672. [Google Scholar] [CrossRef] [PubMed]
- Qi, J.; Shih, W.-C. Parallel Raman microspectroscopy using programmable multipoint illumination. Opt. Lett. 2012, 37, 1289–1291. [Google Scholar] [CrossRef] [PubMed]
- Narendran, S.; Ji, Q.; Eric, D.Y.; Alexander, J.L.; Dina, C.L.; Raphael, E.P.; Kirill, V.L.; Wei-Chuan, S. Line-scan Raman microscopy complements optical coherence tomography for tumor boundary detection. Laser Phys. Lett. 2014, 11, 105602. [Google Scholar]
- Qi, J.; Bechtel, K.L.; Shih, W.-C. Automated image curvature assessment and correction for high-throughput Raman spectroscopy and microscopy. Biomed. Spectrosc. Imaging 2014, 3, 359–368. [Google Scholar]
- Tan, Y.; Gu, J.; Xu, L.; Zang, X.; Liu, D.; Zhang, W.; Liu, Q.; Zhu, S.; Su, H.; Feng, C. High-density hotspots engineered by naturally piled-up subwavelength structures in three-dimensional copper butterfly wing scales for surface-enhanced Raman scattering detection. Adv. Funct. Mater. 2012, 22, 1578–1585. [Google Scholar] [CrossRef]
- Jiwei, Q.; Yudong, L.; Ming, Y.; Qiang, W.; Zongqiang, C.; Wudeng, W.; Wenqiang, L.; Xuanyi, Y.; Jingjun, X.; Qian, S. Large-area high-performance SERS substrates with deep controllable sub-10-nm gap structure fabricated by depositing Au film on the cicada wing. Nanoscale Res. Lett. 2013, 8, 1. [Google Scholar] [CrossRef] [PubMed]
- Zhang, B.; Wang, H.; Lu, L.; Ai, K.; Zhang, G.; Cheng, X. Large-area silver-coated silicon nanowire arrays for molecular sensing using surface-enhanced Raman spectroscopy. Adv. Funct. Mater. 2008, 18, 2348–2355. [Google Scholar] [CrossRef]
- Tan, Y.; Gu, J.; Zang, X.; Xu, W.; Shi, K.; Xu, L.; Zhang, D. Versatile fabrication of intact three-dimensional metallic butterfly wing scales with hierarchical sub-micrometer structures. Angew. Chem. Int. Ed. 2011, 50, 8307–8311. [Google Scholar] [CrossRef] [PubMed]
- Chen, J.; Su, H.; You, X.; Gao, J.; Lau, W.M.; Zhang, D. 3D TiO2 submicrostructures decorated by silver nanoparticles as SERS substrate for organic pollutants detection and degradation. Mater. Res. Bull. 2014, 49, 560–565. [Google Scholar] [CrossRef]
- Su, L.; Zhang, P.; Zheng, D.-W.; Wang, Y.-J.-Q.; Zhong, R.-G. Rapid detection of Escherichia coli and salmonella typhimurium by surface-enhanced Raman scattering. Optoelectron. Lett. 2015, 11, 157–160. [Google Scholar] [CrossRef]
- Zhou, H.; Yang, D.; Ivleva, N.P.; Mircescu, N.E.; Niessner, R.; Haisch, C. SERS detection of bacteria in water by in situ coating with Ag nanoparticles. Anal. Chem. 2014, 86, 1525–1533. [Google Scholar] [CrossRef] [PubMed]
- Fan, C.; Hu, Z.; Mustapha, A.; Lin, M. Rapid detection of food-and waterborne bacteria using surface-enhanced Raman spectroscopy coupled with silver nanosubstrates. Appl. Microbiol. Biotechnol. 2011, 92, 1053–1061. [Google Scholar] [CrossRef] [PubMed]
- Liu, Y.; Chen, Y.-R.; Nou, X.; Chao, K. Potential of surface-enhanced Raman spectroscopy for the rapid identification of Escherichia coli and Listeria monocytogenes cultures on silver colloidal nanoparticles. Appl. Spectrosc. 2007, 61, 824–831. [Google Scholar] [CrossRef] [PubMed]
- Jarvis, R.M.; Brooker, A.; Goodacre, R. Surface-enhanced Raman spectroscopy for bacterial discrimination utilizing a scanning electron microscope with a Raman spectroscopy interface. Anal. Chem. 2004, 76, 5198–5202. [Google Scholar] [CrossRef] [PubMed]
- Jarvis, R.M.; Goodacre, R. Characterisation and identification of bacteria using SERS. Chem. Soc. Rev. 2008, 37, 931–936. [Google Scholar] [CrossRef] [PubMed]
- Premasiri, W.; Moir, D.; Klempner, M.; Krieger, N.; Jones, G.; Ziegler, L. Characterization of the surface enhanced Raman scattering (SERS) of bacteria. J. Phys. Chem. B 2005, 109, 312–320. [Google Scholar] [CrossRef] [PubMed]
- Jarvis, R.M.; Goodacre, R. Discrimination of bacteria using surface-enhanced Raman spectroscopy. Anal. Chem. 2004, 76, 40–47. [Google Scholar] [CrossRef] [PubMed]
- Yang, D.; Zhou, H.; Haisch, C.; Niessner, R.; Ying, Y. Reproducible E. coli detection based on label-free SERS and mapping. Talanta 2016, 146, 457–463. [Google Scholar] [CrossRef] [PubMed]
- Li, J.; Du, Y.; Qi, J.; Sneha, R.; Chang, A.; Mohan, C.; Shih, W.C. Raman spectroscopy as a diagnostic tool for monitoring acute nephritis. J. Biophotonics 2015, 9, 260–269. [Google Scholar] [CrossRef] [PubMed]
- Chen, J.; Qin, G.; Wang, J.; Yu, J.; Shen, B.; Li, S.; Ren, Y.; Zuo, L.; Shen, W.; Das, B. One-step fabrication of sub-10-nm plasmonic nanogaps for reliable SERS sensing of microorganisms. Biosens. Bioelectron. 2013, 44, 191–197. [Google Scholar] [CrossRef] [PubMed]
- Schuster, K.C.; Reese, I.; Urlaub, E.; Gapes, J.R.; Lendl, B. Multidimensional information on the chemical composition of single bacterial cells by confocal Raman microspectroscopy. Anal. Chem. 2000, 72, 5529–5534. [Google Scholar] [CrossRef] [PubMed]
- KahRaman, M.; Zamaleeva, A.I.; Fakhrullin, R.F.; Culha, M. Layer-by-layer coating of bacteria with noble metal nanoparticles for surface-enhanced Raman scattering. Anal. Bioanal. Chem. 2009, 395, 2559. [Google Scholar] [CrossRef] [PubMed]
- Chu, H.; Huang, Y.; Zhao, Y. Silver nanorod arrays as a surface-enhanced Raman scattering substrate for foodborne pathogenic bacteria detection. Appl. Spectrosc. 2008, 62, 922–931. [Google Scholar] [CrossRef] [PubMed]
- Yang, L.; Yan, B.; Premasiri, W.R.; Ziegler, L.D.; Negro, L.D.; Reinhard, B.M. Engineering nanoparticle cluster arrays for bacterial biosensing: The role of the building block in multiscale SERS substrates. Adv. Funct. Mater. 2010, 20, 2619–2628. [Google Scholar] [CrossRef]
- Shiohara, A.; Langer, J.; Polavarapu, L.; Liz-Marzán, L.M. Solution processed polydimethylsiloxane/gold nanostar flexible substrates for plasmonic sensing. Nanoscale 2014, 6, 9817–9823. [Google Scholar] [CrossRef] [PubMed]
- Wu, W.; Liu, L.; Dai, Z.; Liu, J.; Yang, S.; Zhou, L.; Xiao, X.; Jiang, C.; Roy, V.A. Low-cost, disposable, flexible and highly reproducible screen printed SERS substrates for the detection of various chemicals. Sci. Rep. 2015, 5, 10208. [Google Scholar] [CrossRef] [PubMed]
- Polavarapu, L.; Liz-Marzán, L.M. Towards low-cost flexible substrates for nanoplasmonic sensing. Phys. Chem. Chem. Phys. 2013, 15, 5288–5300. [Google Scholar] [CrossRef] [PubMed]
- Chung, A.J.; Huh, Y.S.; Erickson, D. Large area flexible SERS active substrates using engineered nanostructures. Nanoscale 2011, 3, 2903–2908. [Google Scholar] [CrossRef] [PubMed]
- Singh, J.; Chu, H.; Abell, J.; Tripp, R.A.; Zhao, Y. Flexible and mechanical strain resistant large area SERS active substrates. Nanoscale 2012, 4, 3410–3414. [Google Scholar] [CrossRef] [PubMed]
- Lu, G.; Li, H.; Zhang, H. Gold-nanoparticle-embedded polydimethylsiloxane elastomers for highly sensitive Raman detection. Small 2012, 8, 1336–1340. [Google Scholar] [CrossRef] [PubMed]
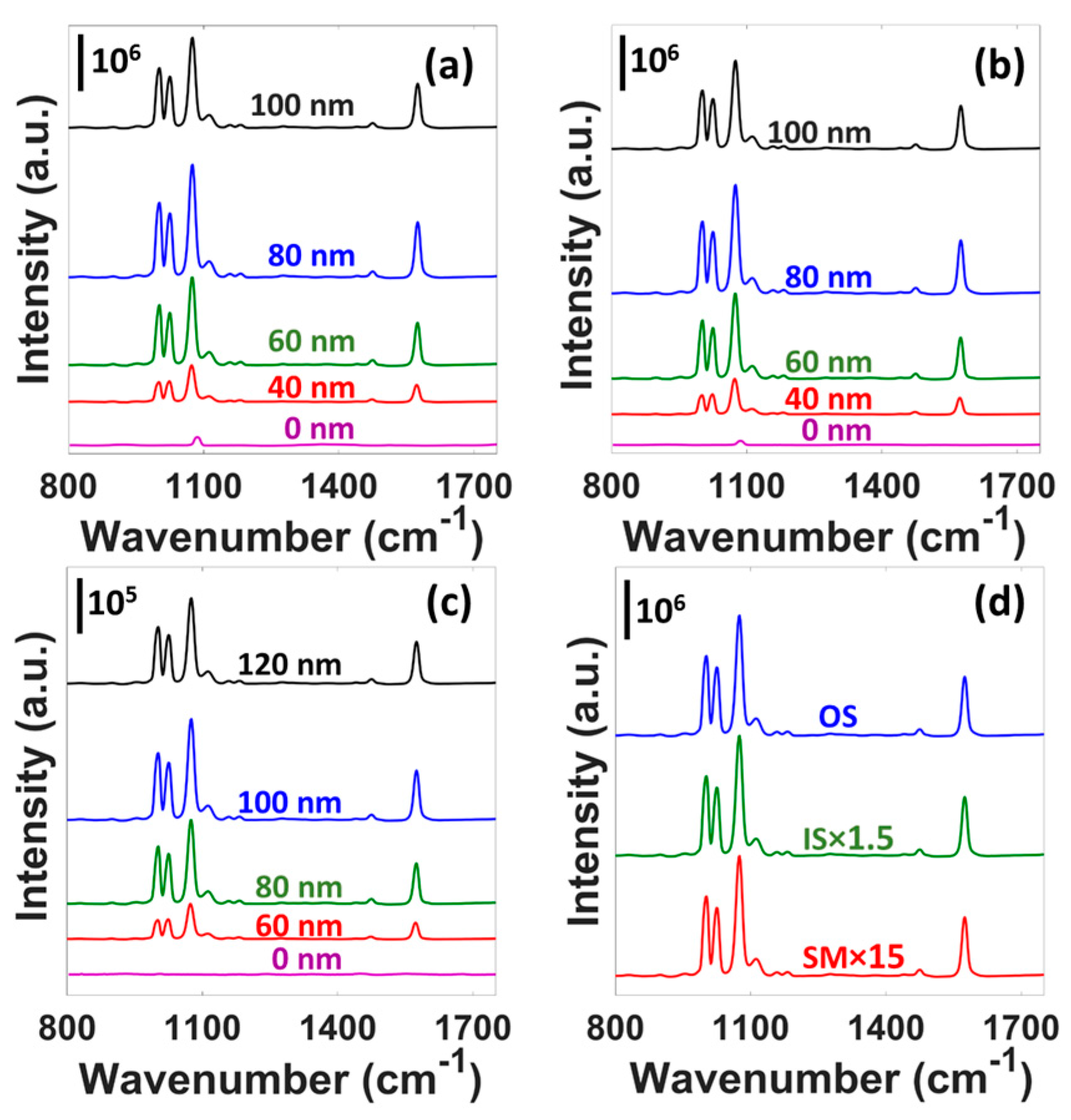





| Peak Position (cm−1) | 1003 | 1025 | 1076 | 1575 |
|---|---|---|---|---|
| %RSD (OS) | 4.98 | 5.02 | 6.07 | 4.09 |
| %RSD (IS) | 13.23 | 12.74 | 14.5 | 12.32 |
| %RSD (SM) | 5.87 | 5.64 | 8.83 | 5.67 |
| Raman Shift (cm−1) | Assignment |
|---|---|
| 550 | Proteins, S–S stretch |
| 600 | Monosubstituted benzenes, CCC ring deformation |
| 632 | Proteins (tyrosine), C–S stretching and C–C twisting |
| 666 | Guanine, (C–S) |
| 707 | Adenine (nucleic acids) |
| 732 | Adenine, trans conformation of (C–S) or tryptophan |
| 794 | Cytosine, uracil (ring, stretch) (nucleic acids) |
| 830 | O–P–O stretching or tyrosine |
| 890 | Carbohydrates, COC stretch |
| 930 | Carbohydrates, C–COO stretch |
| 963 | C=C stretch or tyrosine |
| 1000 | Phenylalanine (the symmetric ring breathing mode) (proteins) |
| 1037 | Phenylalanine (the in-plane C–H bending mode) (proteins) |
| 1079 | Phenylalanine |
| 1107 | CC skeletal and COC stretch, glycosidic link (carbohydrates) |
| 1138 | Carbohydrates, C–N and C–C stretch |
| 1163 | 2-Methyltetradecanoic acid or 15-Methylpalmitic acid |
| 1264 | Cytosine, adenine(DNA), amide III random coil |
| 1343 | (C–H2) deformation, or tryptophan |
| 1369 | Not specified |
| 1449 | Lipids, C–H deformation |
| 1485 | (C–H2) (protein) |
| 1529 | Adenine, cytosine, guanine |
| 1560 | Amide II, NH deformation and CN stretching |
© 2017 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Arnob, M.M.P.; Shih, W.-C. 3-Dimensional Plasmonic Substrates Based on Chicken Eggshell Bio-Templates for SERS-Based Bio-Sensing. Micromachines 2017, 8, 196. https://doi.org/10.3390/mi8060196
Arnob MMP, Shih W-C. 3-Dimensional Plasmonic Substrates Based on Chicken Eggshell Bio-Templates for SERS-Based Bio-Sensing. Micromachines. 2017; 8(6):196. https://doi.org/10.3390/mi8060196
Chicago/Turabian StyleArnob, Md Masud Parvez, and Wei-Chuan Shih. 2017. "3-Dimensional Plasmonic Substrates Based on Chicken Eggshell Bio-Templates for SERS-Based Bio-Sensing" Micromachines 8, no. 6: 196. https://doi.org/10.3390/mi8060196
APA StyleArnob, M. M. P., & Shih, W.-C. (2017). 3-Dimensional Plasmonic Substrates Based on Chicken Eggshell Bio-Templates for SERS-Based Bio-Sensing. Micromachines, 8(6), 196. https://doi.org/10.3390/mi8060196




