The Use of Microfluidics in Cytotoxicity and Nanotoxicity Experiments
Abstract
:1. Introduction
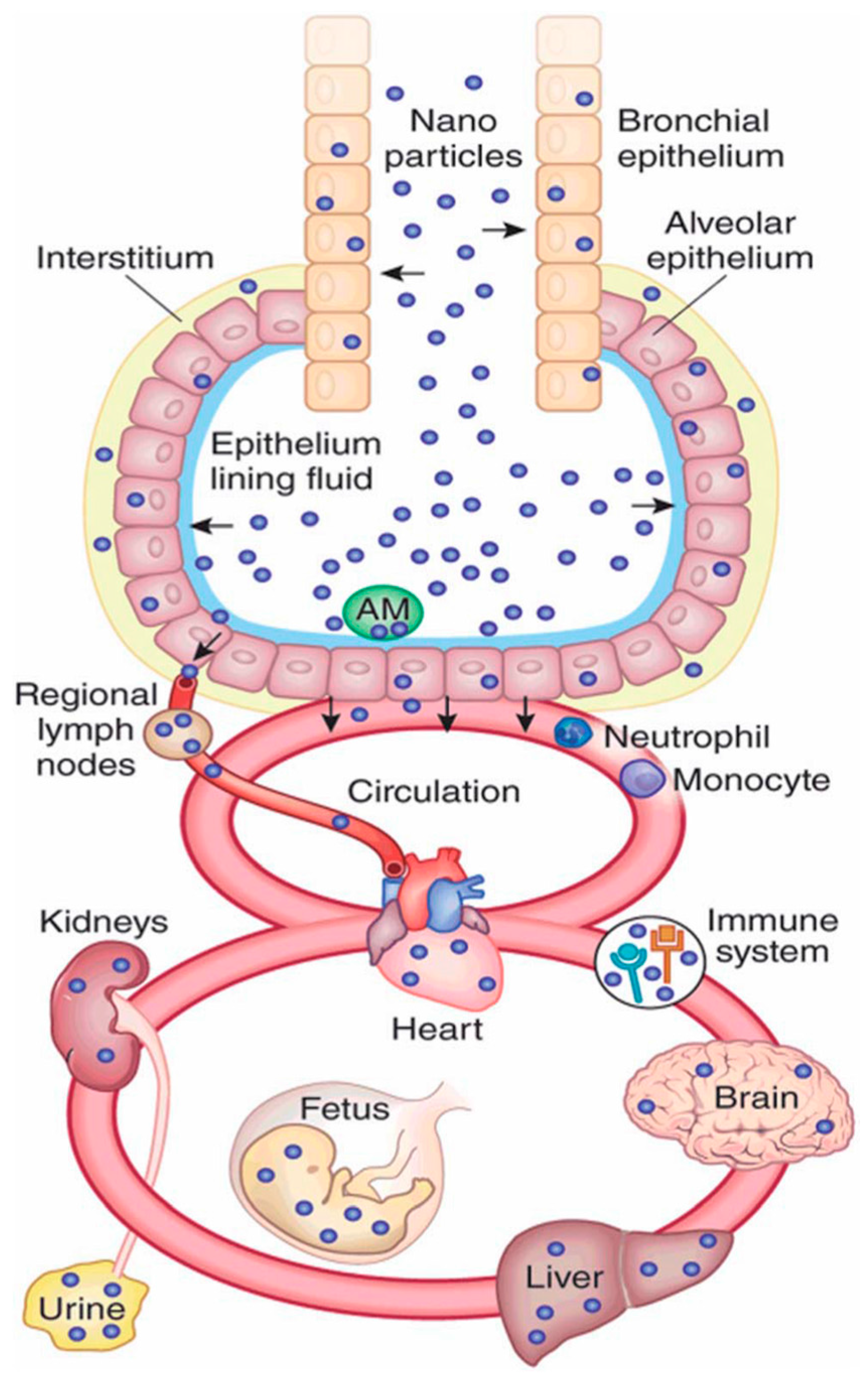
2. Nanomaterial Exposure Pathways in Biology
3. Microfluidics in Cytotoxicity Screening
3.1. Cell Capture and Immobilisation
3.2. Channel Arrays and Laminar Flow
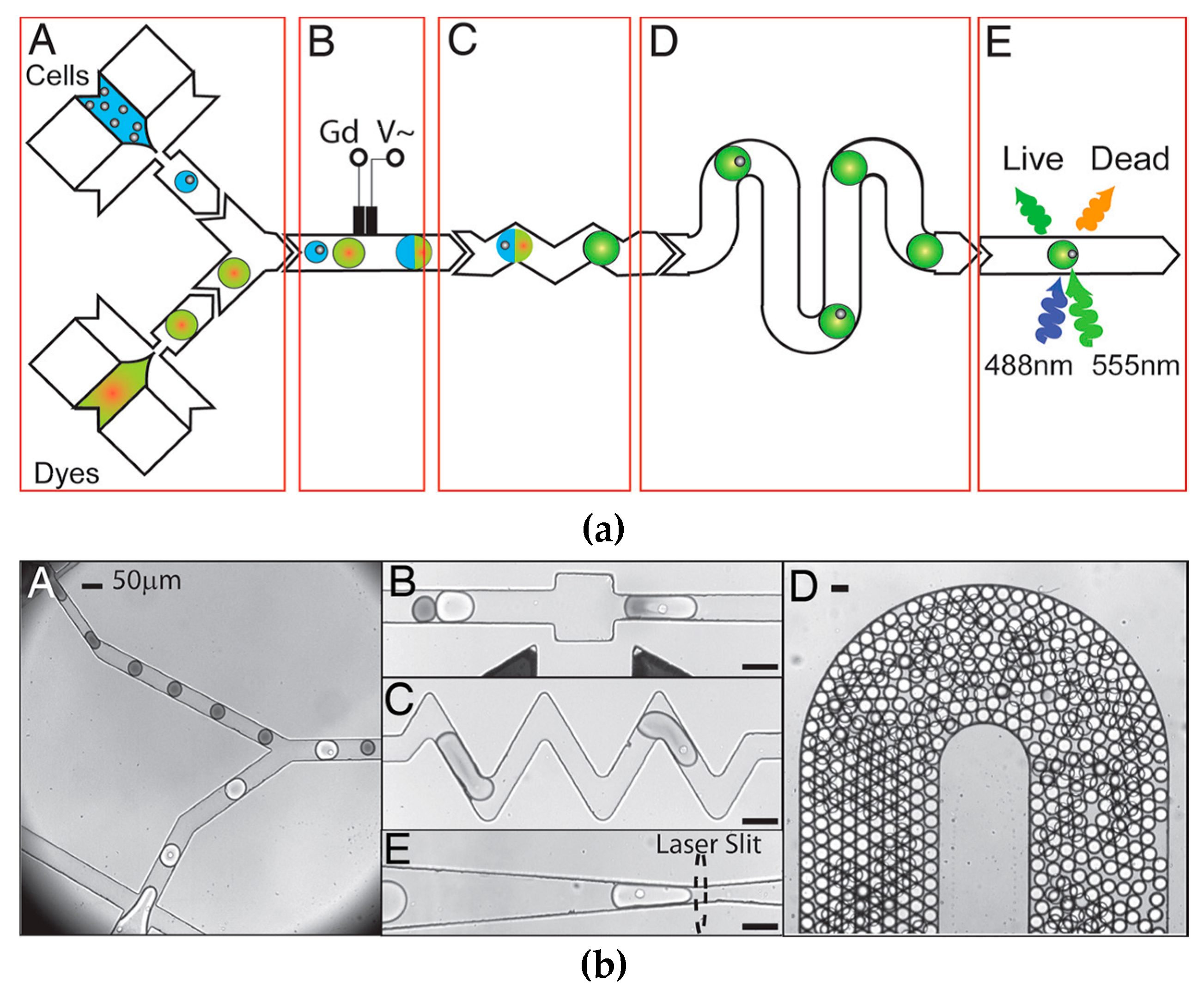
3.3. Droplet Microfluidics
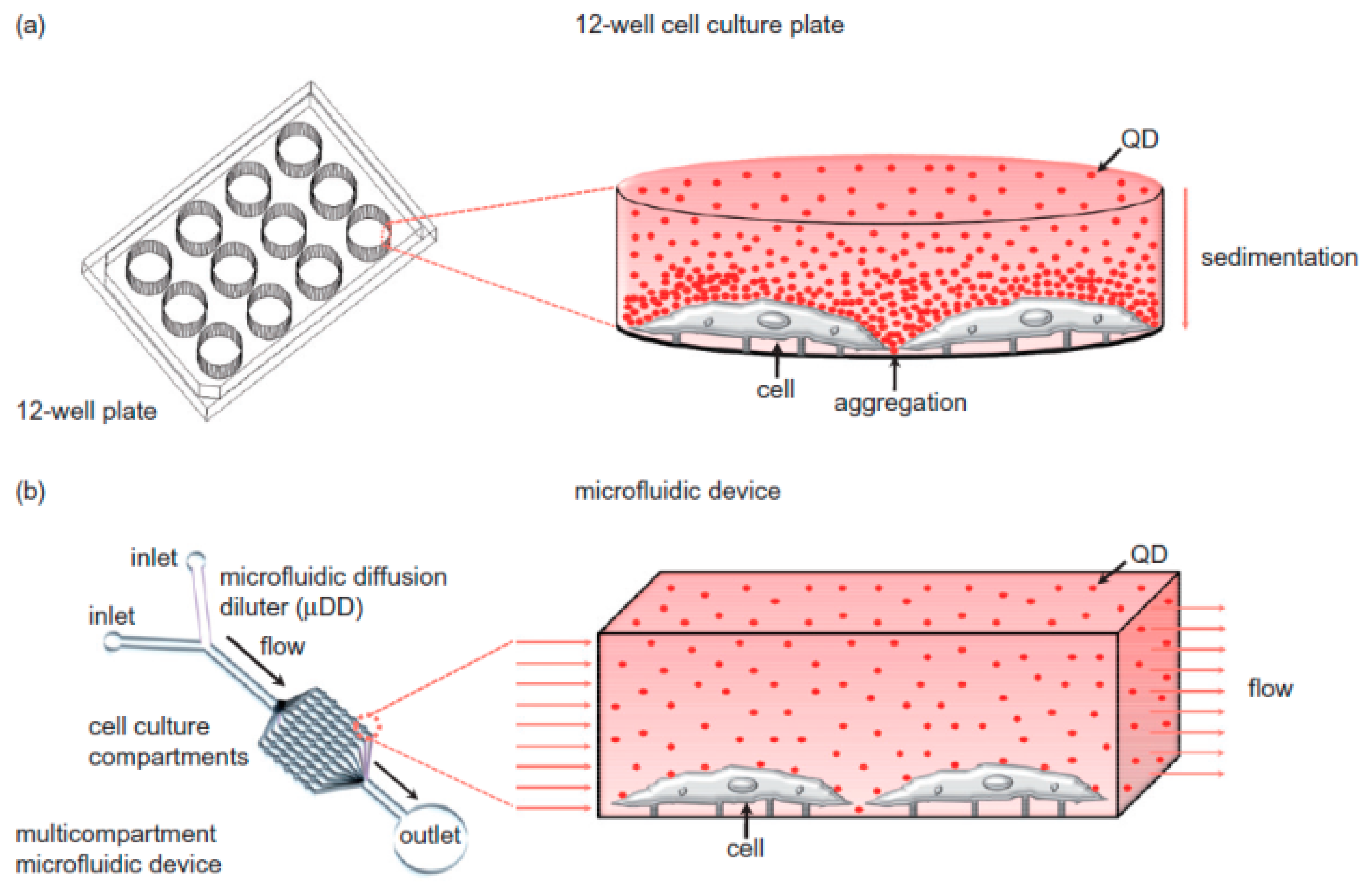
4. Microfluidics for Nanotoxicity Screening
5. Outlook
Acknowledgments
Author Contributions
Conflicts of Interest
References
- Monica, J.C.; Heintz, M.E.; Lewis, P.T. The perils of pre-emptive regulation. Nat. Nano 2007, 2, 68–70. [Google Scholar] [CrossRef] [PubMed]
- Bowman, D.M.; van Calster, G.; Friedrichs, S. Nanomaterials and regulation of cosmetics. Nat. Nano 2010, 5, 92. [Google Scholar] [CrossRef] [PubMed]
- Balas, F.; Arruebo, M.; Urrutia, J.; Santamaria, J. Reported nanosafety practices in research laboratories worldwide. Nat. Nano 2010, 5, 93–96. [Google Scholar] [CrossRef] [PubMed]
- Hunter, R. In Vitro Toxicity Testing: Technologies and Global Markets; Global Markets: A BCC Research Report: Contract No.: PHM017E; BCC Research: Wellesley, MA, USA, 2014. [Google Scholar]
- Alberts, B.; Johnson, A.; Lewis, J.; Raff, M.; Roberts, K.; Walter, P. Molecular Biology of the Cell, 4th ed.; Garland Science: New York, NY, USA, 2002. [Google Scholar]
- Bianconi, E.; Piovesan, A.; Facchin, F.; Beraudi, A.; Casadei, R.; Frabetti, F.; Vitale, L.; Pelleri, M.C.; Tassani, S.; Piva, F.; et al. An estimation of the number of cells in the human body. Ann. Hum. Biol. 2013, 40, 463–471. [Google Scholar] [CrossRef] [PubMed]
- Vickaryous, M.K.; Hall, B.K. Human cell type diversity, evolution, development, and classification with special reference to cells derived from the neural crest. Biol. Rev. Camb. Philos. Soc. 2006, 81, 425–455. [Google Scholar] [CrossRef] [PubMed]
- Jani, P.; Halbert, G.W.; Langridge, J.; Florence, A.T. Nanoparticle uptake by the rat gastrointestinal mucosa: Quantitation and particle size dependency. J. Pharm. Pharmacol. 1990, 42, 821–826. [Google Scholar] [CrossRef] [PubMed]
- Penney, D.P.; Ferin, J.; Oberdorster, G. Pulmonary retention of ultrafine and fine particles in rats. Am. J. Respir. Cell Mol. Biol. 1992, 6, 535–542. [Google Scholar]
- Wakefield, G.; Green, M.; Lipscomb, S.; Flutter, B. Modified titania nanomaterials for sunscreen applications—Reducing free radical generation and DNA damage. Mater. Sci. Technol. 2004, 20, 985–988. [Google Scholar] [CrossRef]
- Osmond, M.J.; McCall, M.J. Zinc oxide nanoparticles in modern sunscreens: An analysis of potential exposure and hazard. Nanotoxicology 2010, 4, 15–41. [Google Scholar] [CrossRef] [PubMed]
- Mu, L.; Sprando, R.L. Application of nanotechnology in cosmetics. Pharm. Res. 2010, 27, 1746–1749. [Google Scholar] [CrossRef] [PubMed]
- Hu, W.; Peng, C.; Luo, W.; Lv, M.; Li, X.; Li, D.; Huang, Q.; Fan, C. Graphene-Based Antibacterial Paper. ACS Nano 2010, 4, 4317–4323. [Google Scholar] [CrossRef] [PubMed]
- Li, Q.; Mahendra, S.; Lyon, D.Y.; Brunet, L.; Liga, M.V.; Li, D.; Alvarez, P.J.J. Antimicrobial nanomaterials for water disinfection and microbial control: Potential applications and implications. Water Res. 2008, 42, 4591–4602. [Google Scholar] [CrossRef] [PubMed]
- Marambio-Jones, C.; Hoek, E.M.V. A review of the antibacterial effects of silver nanomaterials and potential implications for human health and the environment. J. Nanopart. Res. 2010, 12, 1531–1551. [Google Scholar] [CrossRef]
- Betteridge, D.J. What is oxidative stress? Metabolism 2000, 49, 3–8. [Google Scholar] [CrossRef]
- Durán, N.; Guterres, S.S.; Alves, O.L. Nanotoxicology: Materials, Methodologies, and Assessments; Springer: Berlin, Germany, 2013. [Google Scholar]
- Finkel, T.; Holbrook, N.J. Oxidants, oxidative stress and the biology of ageing. Nature 2000, 408, 239–247. [Google Scholar] [CrossRef] [PubMed]
- Chen, Z.; Meng, H.; Xing, G.; Chen, C.; Zhao, Y.; Jia, G.; Wang, T.; Yuan, H.; Ye, C.; Zhao, F.; et al. Acute toxicological effects of copper nanoparticles in vivo. Toxicol. Lett. 2006, 163, 109–120. [Google Scholar] [CrossRef] [PubMed]
- Meng, H.; Chen, Z.; Xing, G.; Yuan, H.; Chen, C.; Zhao, F.; Zhang, C.; Zhao, Y. Ultrahigh reactivity provokes nanotoxicity: Explanation of oral toxicity of nano-copper particles. Toxicol. Lett. 2007, 175, 102–110. [Google Scholar] [CrossRef] [PubMed]
- Ye, Y.; Liu, J.; Xu, J.; Sun, L.; Chen, M.; Lan, M. Nano-SiO2 induces apoptosis via activation of p53 and Bax mediated by oxidative stress in human hepatic cell line. Toxicol. In Vitro 2010, 24, 751–758. [Google Scholar] [CrossRef] [PubMed]
- Ahamed, M. Silica nanoparticles-induced cytotoxicity, oxidative stress and apoptosis in cultured A431 and A549 cells. Hum. Exp. Toxicol. 2013, 32, 186–195. [Google Scholar] [CrossRef] [PubMed]
- Park, E.-J.; Yi, J.; Chung, K.-H.; Ryu, D.-Y.; Choi, J.; Park, K. Oxidative stress and apoptosis induced by titanium dioxide nanoparticles in cultured BEAS-2B cells. Toxicol. Lett. 2008, 180, 222–229. [Google Scholar] [CrossRef] [PubMed]
- Ko, K.S.; Kong, I.C. Toxic effects of nanoparticles on bioluminescence activity, seed germination, and gene mutation. Appl. Microbiol. Biotechnol. 2014, 98, 3295–3303. [Google Scholar] [CrossRef] [PubMed]
- Moran, J.H.; Schnellmann, R.G. A rapid beta-NADH-linked fluorescence assay for lactate dehydrogenase in cellular death. J. Pharmacol. Toxicol. Methods 1996, 36, 41–44. [Google Scholar] [CrossRef]
- Zhao, J.; Castranova, V. Toxicology of nanomaterials used in nanomedicine. J. Toxicol. Environ. Health Part B Crit. Rev. 2011, 14, 593–632. [Google Scholar] [CrossRef] [PubMed]
- Wang, T.; Bai, J.; Jiang, X.; Nienhaus, G.U. Cellular uptake of nanoparticles by membrane penetration: A study combining confocal microscopy with FTIR spectroelectrochemistry. ASC Nano 2012, 6, 1251–1519. [Google Scholar] [CrossRef] [PubMed]
- Kreyling, W.G.; Hirn, S.; Schleh, C. Nanoparticles in the lung. Nat. Biotechnol. 2010, 28, 1275. [Google Scholar] [CrossRef] [PubMed]
- Guo, Y.; Terazzi, E.; Seemann, R.; Fleury, J.B.; Baulin, V.A. Direct proof of spontaneous translocation of lipid-covered hydrophobic nanoparticles through a phospholipid bilayer. Sci. Adv. 2016, 2, e1600261. [Google Scholar] [CrossRef] [PubMed]
- Kettiger, H.; Schipanski, A.; Wick, P.; Huwyler, J. Engineered nanomaterial uptake and tissue distribution: From cell to organism. Int. J. Nanomed. 2013, 8, 3255–3269. [Google Scholar]
- Yang, P.-H.; Sun, X.; Chiu, J.-F.; Sun, H.; He, Q.-Y. Transferrin-mediated gold nanoparticle cellular uptake. Bioconjug. Chem. 2005, 16, 494–496. [Google Scholar] [CrossRef] [PubMed]
- Oh, N.; Park, J.-H. Endocytosis and exocytosis of nanoparticles in mammalian cells. Int. J. Nanomed. 2014, 9, 51–62. [Google Scholar]
- Fischer, H.C.; Chan, W.C.W. Nanotoxicity: The growing need for in vivo study. Curr. Opin.Biotechnol. 2007, 18, 565–571. [Google Scholar] [CrossRef] [PubMed]
- Chiappini, C.; Almeida, C. 8-Silicon nanoneedles for drug delivery. In Semiconducting Silicon Nanowires for Biomedical Applications; Imperial College Press: London, UK, 2014; pp. 144–167. [Google Scholar]
- Kolhar, P.; Doshi, N.; Mitragotri, S. Polymer nanoneedle-mediated intracellular drug delivery. Small 2011, 7, 2094–2100. [Google Scholar] [CrossRef] [PubMed]
- Carmona, H.; Valadez, H.; Yun, Y.; Sankar, J.; Estala, L.; Gomez, F.A. Development of microfluidic-based assays to estimate the binding between osteocalcin (BGLAP) and fluorescent antibodies. Talanta 2015, 132, 676–679. [Google Scholar] [CrossRef] [PubMed]
- Hosokawa, M.; Hayashi, T.; Mori, T.; Yoshino, T.; Nakasono, S.; Matsunaga, T. Microfluidic device with chemical gradient for single-cell cytotoxicity assays. Anal. Chem. 2011, 83, 3648–3654. [Google Scholar] [CrossRef] [PubMed]
- Ng, E.; Hoshino, K.; Zhang, X. Microfluidic immunodetection of cancer cells via site-specific microcontact printing of antibodies on nanoporous surface. Methods 2013, 63, 266–275. [Google Scholar] [CrossRef] [PubMed]
- Pasirayi, G.; Scott, S.M.; Islam, M.; O’Hare, L.; Bateson, S.; Ali, Z. Low cost microfluidic cell culture array using normally closed valves for cytotoxicity assay. Talanta 2014, 129, 491–498. [Google Scholar] [CrossRef] [PubMed]
- Cunha-Matos, C.A.; Millington, O.R.; Wark, A.W.; Zagnoni, M. Real-time assessment of nanoparticle-mediated antigen delivery and cell response. Lab Chip 2016, 16, 3374–3381. [Google Scholar] [CrossRef] [PubMed]
- Weibull, E.; Matsui, S.; Sakai, M.; Andersson Svahn, H.; Ohashi, T. Microfluidic device for generating a stepwise concentration gradient on a microwell slide for cell analysis. Biomicrofluidics 2013, 7, 064115. [Google Scholar] [CrossRef] [PubMed]
- Tong, Z.; Ivask, A.; Guo, K.; McCormick, S.; Lombi, E.; Priest, C.; Voelcker, N.H. Crossed flow microfluidics for high throughput screening of bioactive chemical-cell interactions. Lab Chip 2017, 17, 501–510. [Google Scholar] [CrossRef] [PubMed]
- Roth, E.A.; Xu, T.; Das, M.; Gregory, C.; Hickman, J.J.; Boland, T. Inkjet printing for high-throughput cell patterning. Biomaterials 2004, 25, 3707–3715. [Google Scholar] [CrossRef] [PubMed]
- Melamed, S.; Elad, T.; Belkin, S. Microbial sensor cell arrays. Curr. Opin. Biotechnol. 2012, 23, 2–8. [Google Scholar] [CrossRef] [PubMed]
- Leclerc, E.; El Kirat, K.; Griscom, L. In situ micropatterning technique by cell crushing for co-cultures inside microfluidic biochips. Biomed. Microdevices 2008, 10, 169–177. [Google Scholar] [CrossRef] [PubMed]
- Wu, J.; Chen, Q.; Liu, W.; He, Z.; Lin, J.-M. Recent advances in microfluidic 3D cellular scaffolds for drug assays. TrAC Trends Anal. Chem. 2017, 87, 19–31. [Google Scholar] [CrossRef]
- Toh, Y.-C.; Lim, T.C.; Tai, D.; Xiao, G.; van Noort, D.; Yu, H. A microfluidic 3D hepatocyte chip for drug toxicity testing. Lab Chip 2009, 9, 2026–2035. [Google Scholar] [CrossRef] [PubMed]
- Yoo, S.K.; Lee, J.H.; Yun, S.-S.; Gu, M.B.; Lee, J.H. Fabrication of a bio-MEMS based cell-chip for toxicity monitoring. Biosens. Bioelectron. 2007, 22, 1586–1592. [Google Scholar] [CrossRef] [PubMed]
- Wagner, I.; Materne, E.M.; Brincker, S.; Süßbier, U.; Frädrich, C.; Busek, M.; Sonntag, F.; Sakharov, D.A.; Trushkin, E.V.; Tonevitsky, A.G.; et al. A dynamic multi-organ-chip for long-term cultivation and substance testing proven by 3D human liver and skin tissue co-culture. Lab Chip 2013, 13, 3538–3547. [Google Scholar] [CrossRef] [PubMed]
- SynVivo. SynTox 3D Toxicology Model—Organ Specific Physiological Responses: SynVivo. Available online: http://www.synvivobio.com/syntox/ (accessed on 4 April 2017).
- Ma, B.; Zhang, G.; Qin, J.; Lin, B. Characterization of drug metabolites and cytotoxicity assay simultaneously using an integrated microfluidic device. Lab Chip 2009, 9, 232–238. [Google Scholar] [CrossRef] [PubMed]
- Cooksey, G.A.; Elliott, J.T.; Plant, A.L. Reproducibility and Robustness of a Real-Time Microfluidic Cell Toxicity Assay. Anal. Chem. 2011, 83, 3890–3896. [Google Scholar] [CrossRef] [PubMed]
- Eddings, M.A.; Eckman, J.W.; Arana, C.A.; Papalia, G.A.; Connolly, J.E.; Gale, B.K.; Myszka, D.G. “Spot and hop”: Internal referencing for surface plasmon resonance imaging using a three-dimensional microfluidic flow cell array. Anal. Biochem. 2009, 385, 309–313. [Google Scholar] [CrossRef] [PubMed]
- Wada, K.-I.; Taniguchi, A.; Kobayashi, J.; Yamato, M.; Okano, T. Live Cells-Based Cytotoxic Sensorchip Fabricated in a Microfluidic System. Biotechnol. Bioeng. 2007, 99, 1513–1517. [Google Scholar] [CrossRef] [PubMed]
- Gori, M.; Simonelli, M.C.; Giannitelli, S.M.; Businaro, L.; Trombetta, M.; Rainer, A. Investigating Nonalcoholic Fatty Liver Disease in a Liver-on-a-Chip Microfluidic Device. PLoS ONE 2016, 11, e0159729. [Google Scholar] [CrossRef] [PubMed]
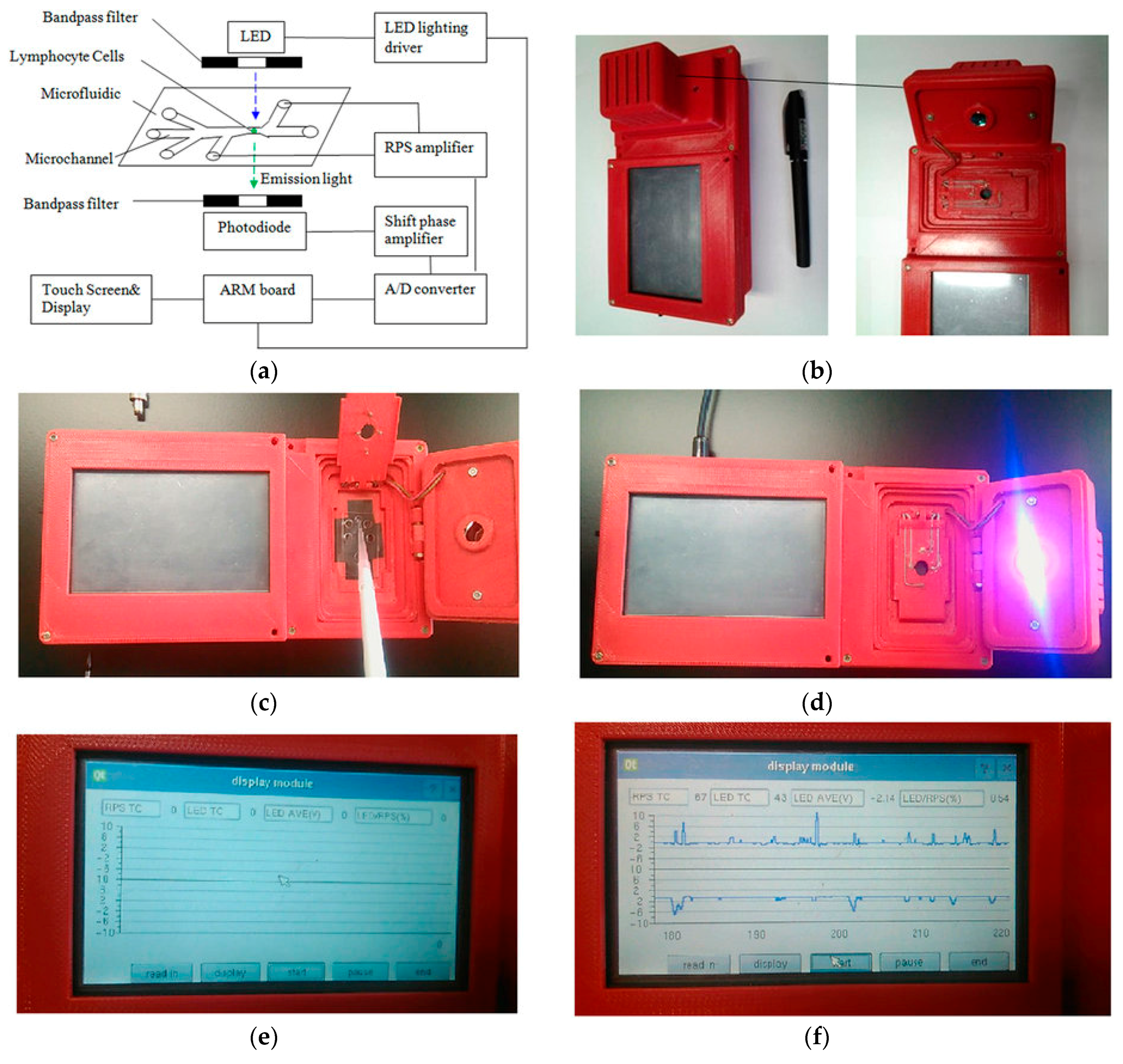
- Wang, J.; Fan, Z.; Zhao, Y.; Song, Y.; Chu, H.; Song, W.; Song, Y.; Pan, X.; Sun, Y.; Li, D. A new hand-held microfluidic cytometer for evaluating irradiation damage by analysis of the damaged cells distribution. Sci. Rep. 2016, 6, 23165. [Google Scholar] [CrossRef] [PubMed]
- Morgan, A.J.L.; Hidalgo San Jose, L.; Jamieson, W.D.; Wymant, J.M.; Song, B.; Stephens, P.; Barrow, D.A.; Castell, O.K. Simple and versatile 3D printed microfluidics using fused filament fabrication. PLoS ONE 2016, 11, e0152023. [Google Scholar] [CrossRef] [PubMed]
- Brouzes, E.; Medkova, M.; Savenelli, N.; Marran, D.; Twardowski, M.; Hutchinson, J.B.; Rothberg, J.M.; Link, D.R.; Perrimon, N.; Samuels, M.L. Droplet microfluidic technology for single-cell high-throughput screening. Proc. Natl. Acad. Sci. 2009, 106, 14195–14200. [Google Scholar] [CrossRef] [PubMed]
- Konry, T.; Sarkar, S.; Sabhachandani, P.; Stroopinsky, D.; Palmer, K.; Cohen, N.; Rosenblatt, J.; Avigan, D.; Konry, T. Dynamic analysis of immune and cancer cell interactions at single cell level in microfluidic droplets. Biomicrofluidics 2016, 10, 054115. [Google Scholar]
- Mahto, S.K.; Yoon, T.H.; Rhee, S.W. A new perspective on in vitro assessment method for evaluating quantum dot toxicity by using microfluidics technology. Biomicrofluidics 2010, 4, 034111. [Google Scholar] [CrossRef] [PubMed]
- Mahto, S.K.; Charwat, V.; Ertl, P.; Rothen-Rutishauser, B.; Rhee, S.W.; Sznitman, J. Microfluidic platforms for advanced risk assessments of nanomaterials. Nanotoxicology 2015, 9, 381–395. [Google Scholar] [CrossRef] [PubMed]
- Geiser, M.; Rothen-Rutishauser, B.; Kapp, N.; Schürch, S.; Kreyling, W.; Schulz, H.; Semmler, M.; Hof, V.I.; Heyder, J.; Gehr, P. Ultrafine particles cross cellular membranes by nonphagocytic mechanisms in lungs and in cultured cells. Environ. Health Perspect. 2005, 113, 1555–1560. [Google Scholar] [CrossRef] [PubMed]
- Rothen-Rutishauser, B.M.; Schurch, S.; Haenni, B.; Kapp, N.; Gehr, P. Interaction of fine particles and nanoparticles with red blood cells visualized with advanced microscopic techniques. Environ. Sci. Technol. 2006, 40, 4353–4359. [Google Scholar] [CrossRef] [PubMed]
- Balbus, J.M.; Maynard, A.D.; Colvin, V.L.; Castranova, V.; Daston, G.P.; Denison, R.A.; Dreher, K.L.; Goering, P.L.; Goldberg, A.M.; Kulinowski, K.M.; et al. Meeting report: Hazard assessment for nanoparticles--report from an interdisciplinary workshop. Environ. Health Perspect. 2007, 115, 1654–1659. [Google Scholar] [CrossRef] [PubMed]
- Nel, A.E.; Madler, L.; Velegol, D.; Xia, T.; Hoek, E.M.V.; Somasundaran, P.; Klaessig, F.; Castranova, V.; Thompson, M. Understanding biophysicochemical interactions at the nano-bio interface. Nat. Mater. 2009, 8, 543–557. [Google Scholar] [CrossRef] [PubMed]
- Richter, L.; Charwat, V.; Jungreuthmayer, C.; Bellutti, F.; Brueckl, H.; Ertl, P. Monitoring cellular stress responses to nanoparticles using a lab-on-a-chip. Lab Chip 2011, 11, 2551–2560. [Google Scholar] [CrossRef] [PubMed]
- Cunha-Matos, C.A.; Millington, O.M.; Wark, A.W.; Zagnoni, M. (Eds.) Real-Time Multimodal Imaging of Nanoparticle-Cell Interactions in High-Throughput Microfluidics. In Proceedings of the 18th International Conference on Miniaturized Systems for Chemistry and Life Sciences, San Antonio, TX, USA, 26–30 October 2014. [Google Scholar]
- Velve-Casquillas, G.; Le Berre, M.; Piel, M.; Tran, P.T. Microfluidic tools for cell biological research. Nano Today 2010, 5, 28–47. [Google Scholar] [CrossRef] [PubMed]
- Shah, P.; Kaushik, A.; Zhu, X.; Zhang, C.; Li, C.-Z. Chip based single cell analysis for nanotoxicity assessment. Analyst 2014, 139, 2088–2098. [Google Scholar] [CrossRef] [PubMed]
- Rothbauer, M.; Praisler, I.; Docter, D.; Stauber, R.H.; Ertl, P. Microfluidic impedimetric cell regeneration assay to monitor the enhanced cytotoxic effect of nanomaterial perfusion. Biosensors 2015, 5, 736–749. [Google Scholar] [CrossRef] [PubMed]
- Kim, D.; Lin, Y.-S.; Haynes, C.L. On-chip evaluation of shear stress effect on cytotoxicity of mesoporous silica nanoparticles. Anal. Chem. 2012, 83, 8377–8382. [Google Scholar] [CrossRef] [PubMed]
- Liu, Y.; Wang, S.; Wang, Y. Patterned fibers embedded microfluidic chips based on PLA and PDMS for Ag nanoparticle safety testing. Polymers 2016, 8, 402. [Google Scholar] [CrossRef]
- Park, J.Y.; Yoon, T.H. Microfluidic image cytometry (μFIC) assessments of silver nanoparticle cytotoxicity. Bull. Korean Chem. Soc. 2012, 33, 4023–4027. [Google Scholar] [CrossRef]
- Shah, P. Development of a Lab-on-a-Chip Device for Rapid Nanotoxicity Assessment In Vitro. Ph.D. Thesis, Florida International University, Miami, FL, USA, 2014. [Google Scholar]
- Huh, D.; Matthews, B.D.; Mammoto, A.; Montoya-Zavala, M.; Hsin, H.Y.; Ingber, D.E. Reconstituting organ-level lung functions on a chip. Science 2010, 328, 1662–1668. [Google Scholar] [CrossRef] [PubMed]

© 2017 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
McCormick, S.C.; Kriel, F.H.; Ivask, A.; Tong, Z.; Lombi, E.; Voelcker, N.H.; Priest, C. The Use of Microfluidics in Cytotoxicity and Nanotoxicity Experiments. Micromachines 2017, 8, 124. https://doi.org/10.3390/mi8040124
McCormick SC, Kriel FH, Ivask A, Tong Z, Lombi E, Voelcker NH, Priest C. The Use of Microfluidics in Cytotoxicity and Nanotoxicity Experiments. Micromachines. 2017; 8(4):124. https://doi.org/10.3390/mi8040124
Chicago/Turabian StyleMcCormick, Scott C., Frederik H. Kriel, Angela Ivask, Ziqiu Tong, Enzo Lombi, Nicolas H. Voelcker, and Craig Priest. 2017. "The Use of Microfluidics in Cytotoxicity and Nanotoxicity Experiments" Micromachines 8, no. 4: 124. https://doi.org/10.3390/mi8040124
APA StyleMcCormick, S. C., Kriel, F. H., Ivask, A., Tong, Z., Lombi, E., Voelcker, N. H., & Priest, C. (2017). The Use of Microfluidics in Cytotoxicity and Nanotoxicity Experiments. Micromachines, 8(4), 124. https://doi.org/10.3390/mi8040124








