SMILE Platform: An Innovative Microfluidic Approach for On-Chip Sample Manipulation and Analysis in Oral Cancer Diagnosis
Abstract
:1. Introduction
2. Materials and Methods
2.1. Materials
2.2. Computational Modeling
2.3. Design, Fabrication, and Sealing of PMMA Substrates
2.4. Microchannel Passivation and Functionalization
2.5. Experimental Tests for Particles and Coloured Liquids
2.6. Experimental Tests for Cells
3. Results and Discussion
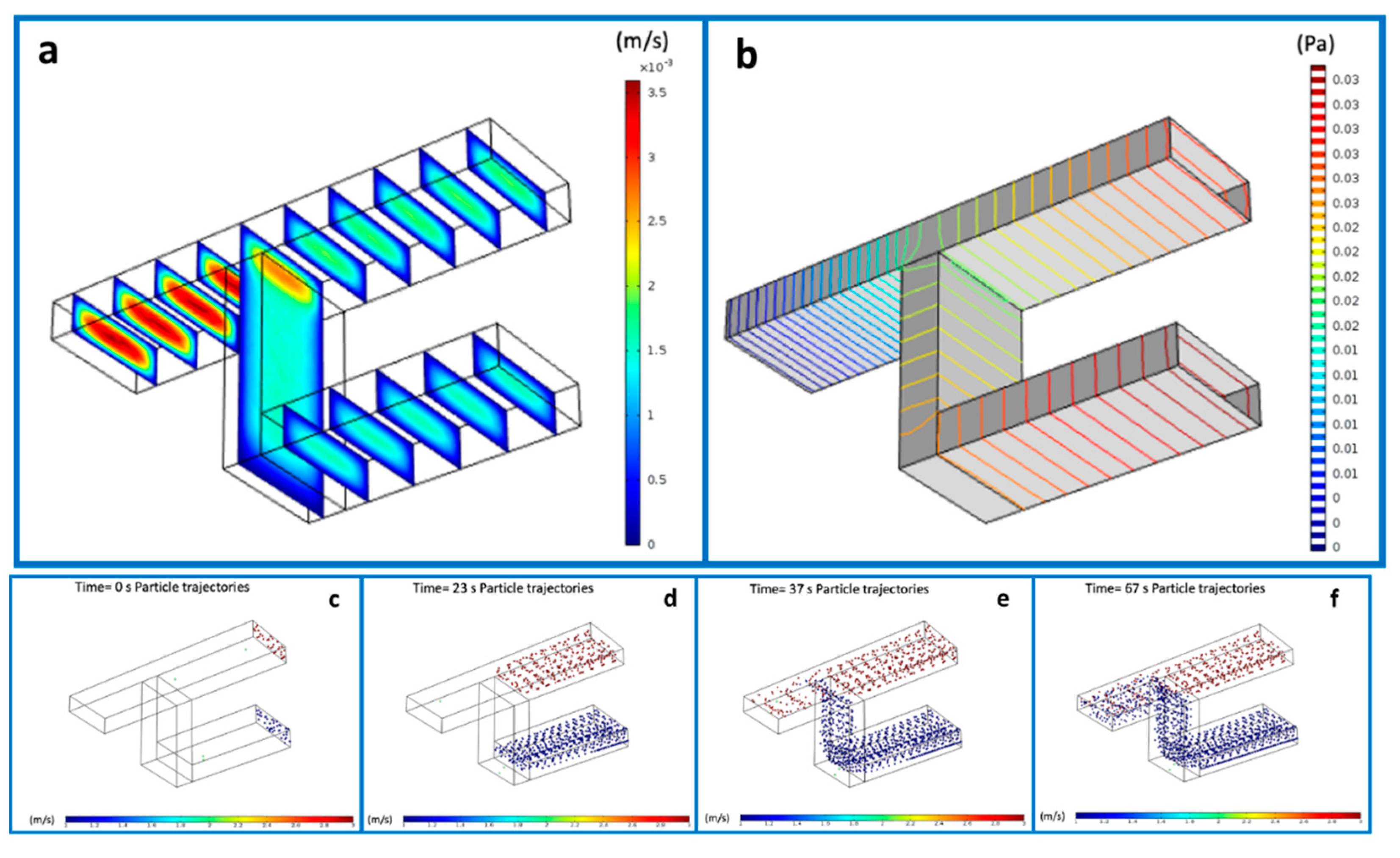
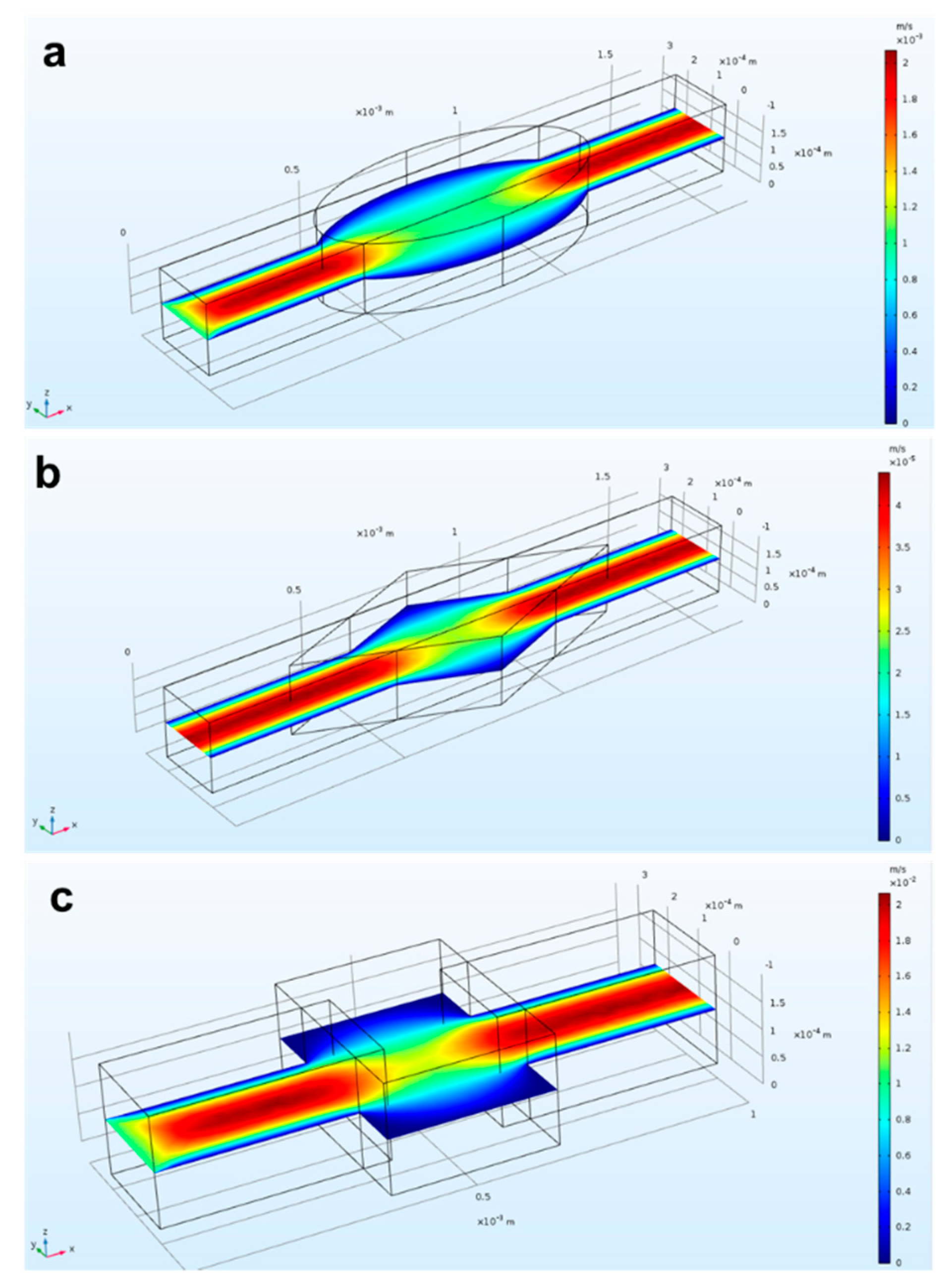
3.1. Verification of the Numerical Model
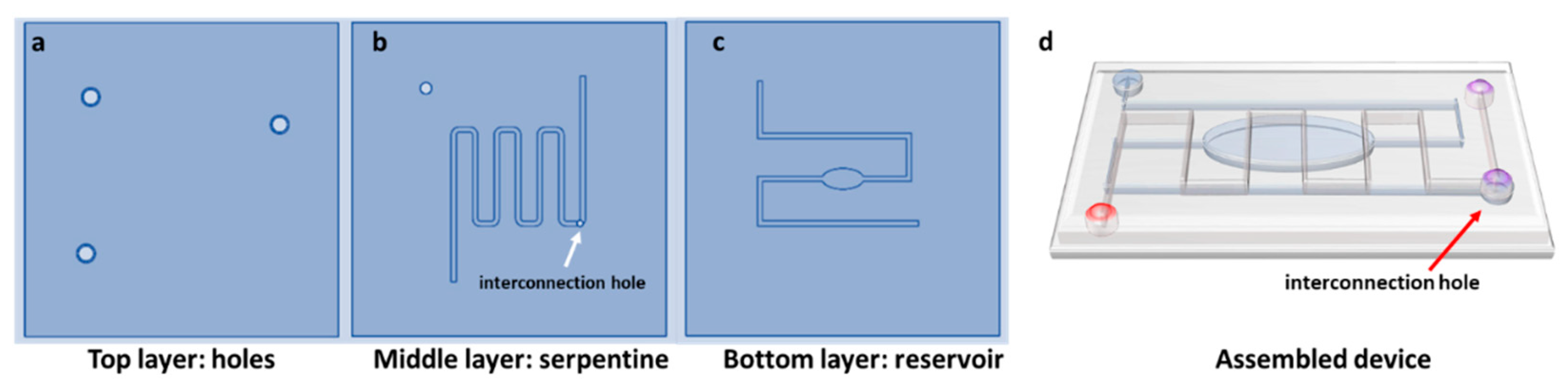
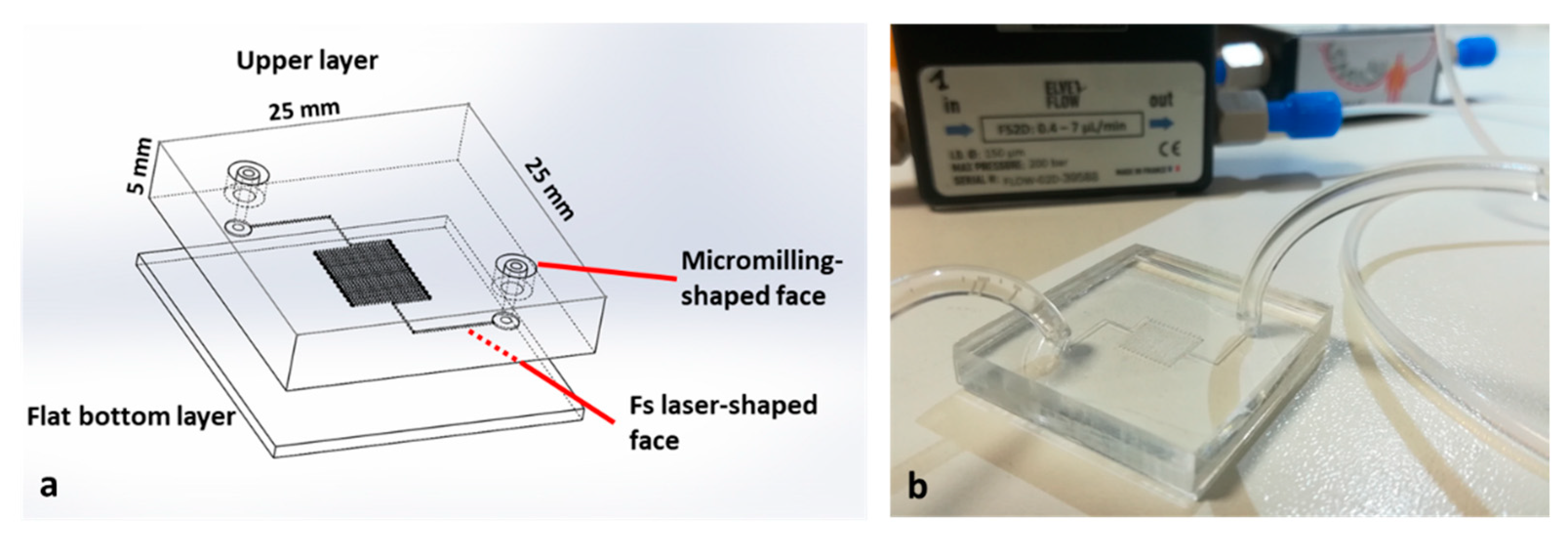
3.2. Design and Fabrication of the LOC Devices
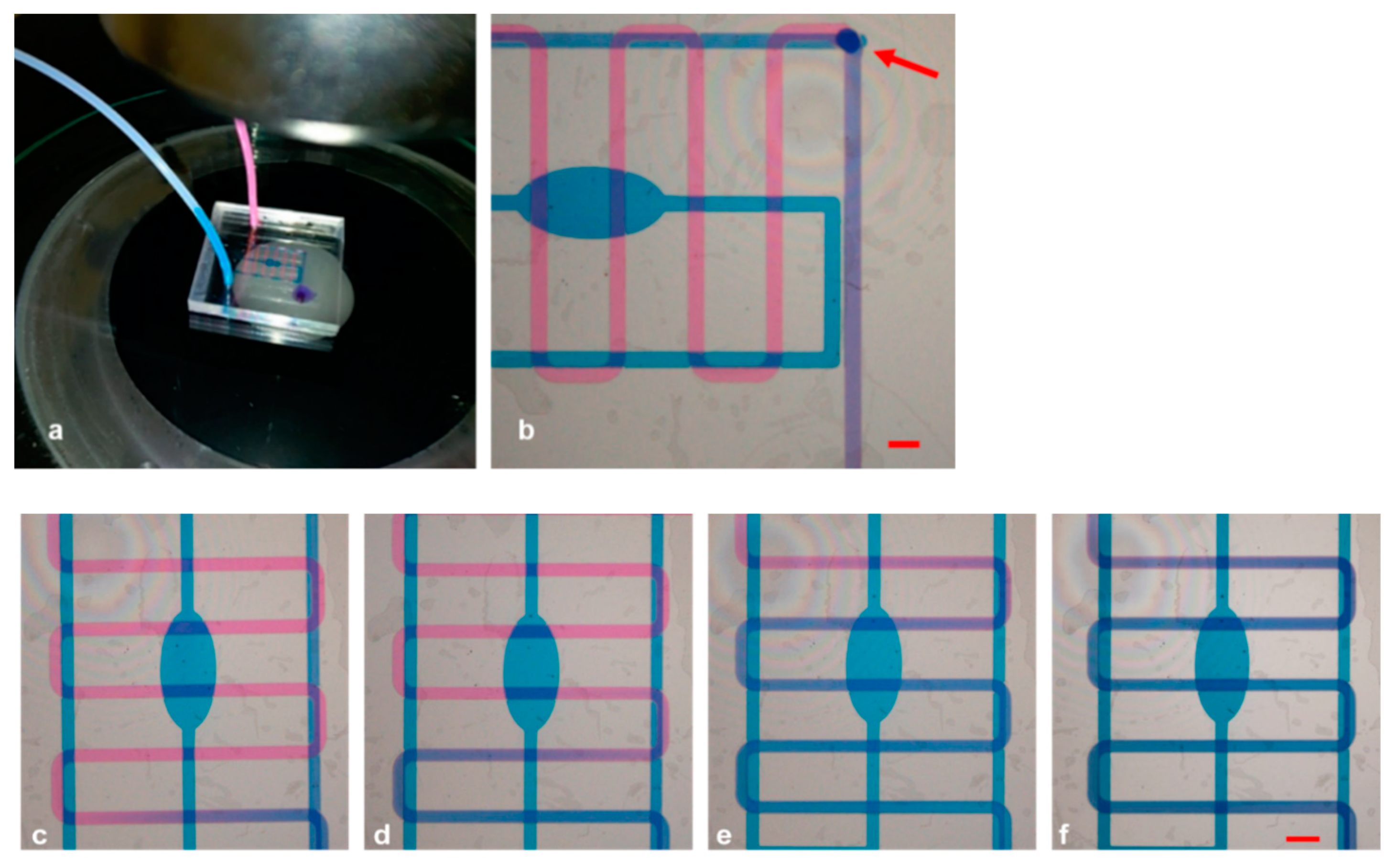
3.3. Mixing and Gradient Generation Experiments
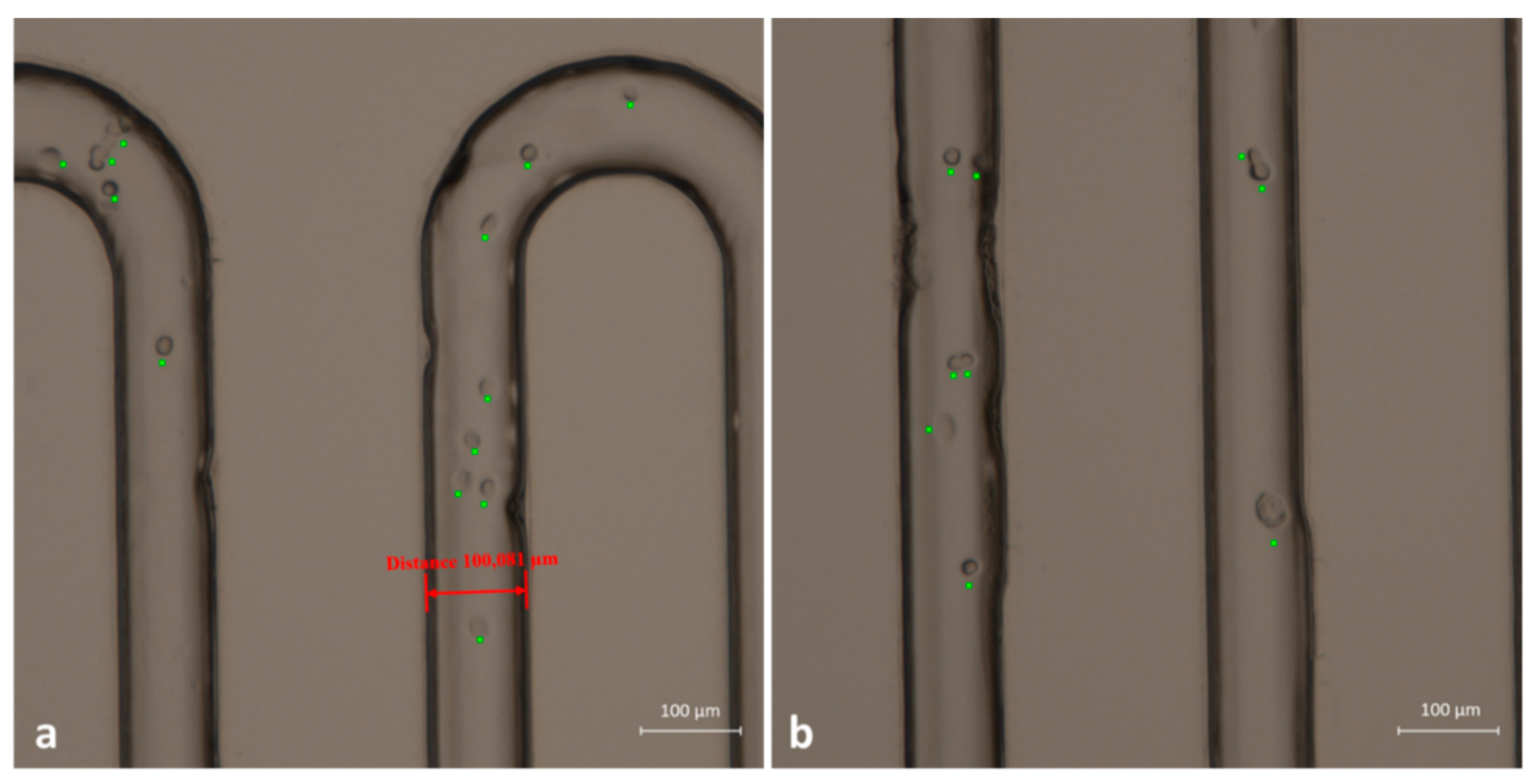
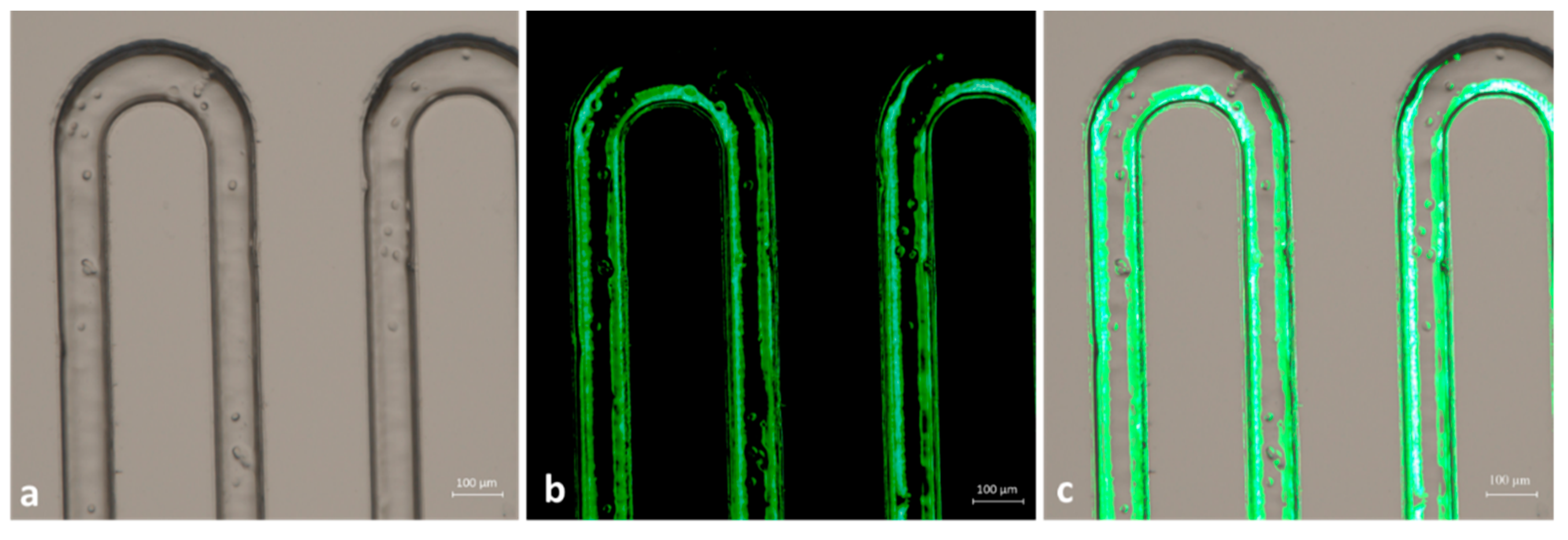
3.4. CTC Capture Experiments
4. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Siegel, R.L.; Miller, K.D.; Jemal, A. Cancer statistics, 2020. CA A Cancer J. Clin. 2020, 70, 7–30. [Google Scholar] [CrossRef] [PubMed]
- Vergaro, V.; Baldassarre, F.; de Santisa, F.; Ciccarella, G.; Giannelli, G.; Leporatti, S. TGF-beta inihibitor-loaded polyelectrolyte multilayers capsules for sustained targeting of hepatocarcinoma cells. Curr. Pharm. Des. 2012, 18, 4155–4164. [Google Scholar] [CrossRef]
- Gupta, B.; Johnson, N.W.; Kumar, N. Global Epidemiology of Head and Neck Cancers: A Continuing Challenge. Oncology 2016, 91, 13–23. [Google Scholar] [CrossRef]
- Hashibe, M.; Brennan, P.; Benhamou, S.; Castellsague, X.; Chu, C.; Curado, M.P.; Dal Maso, L.; Dauct, A.W.; Fabianova, E.; Wunsch, V.; et al. Alcohol drinking in never users of tobacco, cigarette smoking in never drinkers, and the risk of head and neck cancer: Pooled analysis in the international head and neck cancer epidemiology consortium. J. Natl. Cancer Inst. 2007, 99, 777–789. [Google Scholar] [CrossRef]
- Perot, P.; Falguieres, M.; Arowas, L.; Laude, H.; Foy, J.P.; Goudot, P.; Corre-Catelin, N.; Ungeheuer, M.N.; Caro, V.; Heard, I.; et al. Investigation of viral etiology in potentially malignant disorders and oral squamous cell carcinomas in non-smoking, non-drinking patients. PLoS ONE 2020, 15. [Google Scholar] [CrossRef]
- Petersen, P.E.; Bourgeois, D.; Ogawa, H.; Estupinan-Day, S.; Ndiaye, C. The global burden of oral diseases and risks to oral health. Bull. World Health Organ. 2005, 83, 661–669. [Google Scholar] [PubMed]
- Li, L.; Zhang, Z.T. Genetic Association between NFKBIA and NFKB1 Gene Polymorphisms and the Susceptibility to Head and Neck Cancer: A Meta-Analysis. Dis. Markers 2019, 2019. [Google Scholar] [CrossRef] [Green Version]
- Tsompana, M.; Gluck, C.; Sethi, I.; Joshi, I.; Bard, J.; Nowak, N.J.; Sinha, S.; Buck, M.J. Reactivation of super-enhancers by KLF4 in human Head and Neck Squamous Cell Carcinoma. Oncogene 2020, 39, 262–277. [Google Scholar] [CrossRef]
- Ogino, S.; Nowak, J.A.; Hamada, T.; Milner, D.A., Jr.; Nishihara, R. Insights into Pathogenic Interactions among Environment, Host, and Tumor at the Crossroads of Molecular Pathology and Epidemiology. Annu. Rev. Pathol. 2019, 14, 83–103. [Google Scholar] [CrossRef] [PubMed]
- Mima, K.; Kosumi, K.; Baba, Y.; Hamada, T.; Baba, H.; Ogino, S. The microbiome, genetics, and gastrointestinal neoplasms: The evolving field of molecular pathological epidemiology to analyze the tumor-immune-microbiome interaction. Hum. Genet. 2021, 140, 725–746. [Google Scholar] [CrossRef] [PubMed]
- Shiga, K.; Tateda, M.; Saijo, S.; Hori, T.; Sato, I.; Tateno, H.; Matsuura, K.; Takasaka, T.; Miyagi, T. Presence of Streptococcus infection in extra-oropharyngeal head and neck squamous cell carcinoma and its implication in carcinogenesis. Oncol. Rep. 2001, 8, 245–248. [Google Scholar] [CrossRef]
- Zhang, L.; Liu, Y.; Zheng, H.J.; Zhang, C.P. The Oral Microbiota May Have Influence on Oral Cancer. Front. Cell. Infect. Microbiol. 2020, 9, 11. [Google Scholar] [CrossRef] [PubMed]
- Sinevici, N.; Mittermayr, S.; Davey, G.P.; Bones, J.; O’Sullivan, J. Salivary N-glycosylation as a biomarker of oral cancer: A pilot study. Glycobiology 2019, 29, 726–734. [Google Scholar] [CrossRef] [PubMed]
- Zhang, C.Z.; Cheng, X.Q.; Li, J.Y.; Zhang, P.; Yi, P.; Xu, X.; Zhou, X.D. Saliva in the diagnosis of diseases. Int. J. Oral Sci. 2016, 8, 133–137. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Vesty, A.; Gear, K.; Biswas, K.; Radcliff, F.J.; Taylor, M.W.; Douglas, R.G. Microbial and inflammatory-based salivary biomarkers of head and neck squamous cell carcinoma. Clin. Exp. Dent. Res. 2018, 4, 255–262. [Google Scholar] [CrossRef] [Green Version]
- Diesch, T.; Filippi, C.; Fritschi, N.; Filippi, A.; Ritz, N. Cytokines in saliva as biomarkers of oral and systemic oncological or infectious diseases: A systematic review. Cytokine 2021, 143. [Google Scholar] [CrossRef]
- Song, X.; Yang, X.; Narayanan, R.; Shankar, V.; Ethiraj, S.; Wang, X.; Duan, N.; Ni, Y.-H.; Hu, Q.; Zare, R.N. Oral squamous cell carcinoma diagnosed from saliva metabolic profiling. Proc. Natl. Acad. Sci. USA 2020, 117, 16167–16173. [Google Scholar] [CrossRef]
- Day, A.T.; Fakhry, C.; Tiro, J.A.; Dahlstrom, K.R.; Sturgis, E.M. Considerations in Human Papillomavirus-Associated Oropharyngeal Cancer Screening A Review. JAMA Otolaryngol-Head Neck Surg. 2020, 146, 656–664. [Google Scholar] [CrossRef]
- Tenore, G.; Nuvoli, A.; Mohsen, A.; Cassoni, A.; Battisti, A.; Terenzi, V.; Della Monaca, M.; Raponi, I.; Brauner, E.; De Felice, F.; et al. Tobacco, Alcohol and Family History of Cancer as Risk Factors of Oral Squamous Cell Carcinoma: Case-Control Retrospective Study. Appl. Sci. Basel 2020, 10, 11. [Google Scholar] [CrossRef]
- Zoupanou, S.; Chiriacò, M.S.; Tarantini, I.; Ferrara, F. Innovative 3D Microfluidic Tools for On-Chip Fluids and Particles Manipulation: From Design to Experimental Validation. Micromachines 2021, 12, 104. [Google Scholar] [CrossRef] [PubMed]
- Volpe, A.; Krishnan, U.; Chiriacò, M.S.; Primiceri, E.; Ancona, A.; Ferrara, F. A smart procedure for the femtosecond laser-based fabrication of a polymeric lab-on-a-chip for capturing tumor cell. Engineering 2020, 12. [Google Scholar] [CrossRef]
- Trotta, G.; Volpe, A.; Ancona, A.; Fassi, I. Flexible micro manufacturing platform for the fabrication of PMMA microfluidic devices. J. Manuf. Process. 2018, 35, 107–117. [Google Scholar] [CrossRef]
- Volpe, A.; Paie, P.; Ancona, A.; Osellame, R. Polymeric fully inertial lab-on-a-chip with enhanced-throughput sorting capabilities. Microfluid. Nanofluidics 2019, 23. [Google Scholar] [CrossRef]
- Niculescu, A.G.; Chircov, C.; Birca, A.C.; Grumezescu, A.M. Fabrication and Applications of Microfluidic Devices: A Review. Int. J. Mol. Sci. 2021, 22, 2011. [Google Scholar] [CrossRef]
- Abgrall, P.; Gue, A.M. Lab-on-chip technologies: Making a microfluidic network and coupling it into a complete microsystem-A review. J. Micromech. Microeng. 2007, 17, R15–R49. [Google Scholar] [CrossRef]
- Abdelgawad, M.; Watson, M.W.L.; Young, E.W.K.; Mudrik, J.M.; Ungrin, M.D.; Wheeler, A.R. Soft lithography: Masters on demand. Lab Chip 2008, 8, 1379–1385. [Google Scholar] [CrossRef] [PubMed]
- Wilson, M.E.; Kota, N.; Kim, Y.; Wang, Y.; Stolz, D.B.; LeDuc, P.R.; Ozdoganlar, O.B. Fabrication of circular microfluidic channels by combining mechanical micromilling and soft lithography. Lab Chip 2011, 11, 1550–1555. [Google Scholar] [CrossRef] [PubMed]
- Chiriaco, M.S.; Bianco, M.; Amato, F.; Primiceri, E.; Ferrara, F.; Arima, V.; Maruccio, G. Fabrication of interconnected multilevel channels in a monolithic SU-8 structure using a LOR sacrificial layer. Microelectron. Eng. 2016, 164, 30–35. [Google Scholar] [CrossRef]
- Hardy, B.S.; Uechi, K.; Zhen, J.; Pirouz Kavehpour, H. The deformation of flexible PDMS microchannels under a pressure driven flow. Lab Chip 2009, 9, 935–938. [Google Scholar] [CrossRef] [PubMed]
- Bhattacharjee, N.; Urrios, A.; Kanga, S.; Folch, A. The upcoming 3D-printing revolution in microfluidics. Lab Chip 2016, 16, 1720–1742. [Google Scholar] [CrossRef] [Green Version]
- Yazdi, A.A.; Popma, A.; Wong, W.; Nguyen, T.; Pan, Y.Y.; Xu, J. 3D printing: An emerging tool for novel microfluidics and lab-on-a-chip applications. Microfluid. Nanofluidics 2016, 20. [Google Scholar] [CrossRef]
- Available online: https://ttconsultants.com/3d-printing-limitations/ (accessed on 23 July 2021).
- Farson, D.F.; Choi, H.W.; Zimmerman, B.; Steach, J.K.; Chalmers, J.J.; Olesik, S.V.; Lee, L.J. Femtosecond laser micromachining of dielectric materials for biomedical applications. J. Micromech. Microeng. 2008, 18, 035020. [Google Scholar] [CrossRef]
- Volpe, A.; Gaudiuso, C.; Ancona, A. Sorting of Particles Using Inertial Focusing and Laminar Vortex Technology: A Review. Micromachines 2019, 10, 594. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Kononenko, T.V.; Konov, V.I.; Garnov, S.V.; Danielius, R.; Piskarskas, A.; Tamosauskas, G.; Dausinger, F. Comparative study of the ablation of materials by femtosecond and pico- or nanosecond laser pulses. Quantum Electron. 1999, 29, 724–728. [Google Scholar] [CrossRef]
- Volpe, A.; Covella, S.; Gaudiuso, C.; Ancona, A. Improving the Laser Texture Strategy to Get Superhydrophobic Aluminum Alloy Surfaces. Coatings 2021, 11, 369. [Google Scholar] [CrossRef]
- Ashkenasi, D.; Müller, G.; Rosenfeld, A.; Stoian, R.; Hertel, I.V.; Bulgakova, N.M.; Campbell, E.E.B. Fundamentals and advantages of ultrafast micro-structuring of transparent materials. Appl. Phys. A 2003, 77, 223–228. [Google Scholar] [CrossRef]
- Faustino, V.; Catarino, S.O.; Lima, R.; Minas, G. Biomedical microfluidic devices by using low-cost fabrication techniques: A review. J. Biomech. 2016, 49, 2280–2292. [Google Scholar] [CrossRef] [Green Version]
- Guckenberger, D.J.; de Groot, T.E.; Wan, A.M.D.; Beebe, D.J.; Young, E.W.K. Micromilling: A method for ultra-rapid prototyping of plastic microfluidic devices. Lab Chip 2015, 15, 2364–2378. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Song, I.-H.; Park, T. Connector-Free World-to-Chip Interconnection for Microfluidic Devices. Micromachines 2019, 10, 166. [Google Scholar] [CrossRef] [Green Version]
- Yuen, P.K. A reconfigurable stick-n-play modular microfluidic system using magnetic interconnects. Lab Chip 2016, 16, 3700–3707. [Google Scholar] [CrossRef]
- Berthier, E.; Young, E.W.K.; Beebe, D. Engineers are from PDMS-land, Biologists are from Polystyrenia. Lab Chip 2012, 12, 1224–1237. [Google Scholar] [CrossRef]
- Rotem, A.; Abate, A.R.; Utada, A.S.; Van Steijn, V.; Weitz, D.A. Drop formation in non-planar microfluidic devices. Lab Chip 2012, 12, 4263–4268. [Google Scholar] [CrossRef] [Green Version]
- Nguyen, T.; Chidambara, V.A.; Andreasen, S.Z.; Golabi, M.; Huynh, V.N.; Linh, Q.T.; Bang, D.D.; Wolff, A. Point-of-care devices for pathogen detections: The three most important factors to realise towards commercialization. TrAC Trends Anal. Chem. 2020, 131. [Google Scholar] [CrossRef]
- Temiz, Y.; Lovchik, R.D.; Kaigala, G.V.; Delamarche, E. Lab-on-a-chip devices: How to close and plug the lab? Microelectron. Eng. 2015, 132, 156–175. [Google Scholar] [CrossRef]
- Vazquez, R.M.; Trotta, G.; Volpe, A.; Bernava, G.; Basile, V.; Paturzo, M.; Ferraro, P.; Ancona, A.; Fassi, I.; Osellame, R. Rapid Prototyping of Plastic Lab-on-a-Chip by Femtosecond Laser Micromachining and Removable Insert Microinjection Molding. Micromachines 2017, 8, 328. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Chaudhuri, P.K.; Ebrahimi Warkiani, M.; Jing, T.; Kenry; Lim, C.T. Microfluidics for research and applications in oncology. Analyst 2016, 141, 504–524. [Google Scholar] [CrossRef] [PubMed]
- Song, W.; Li, X.; Zhao, Y.; Liu, C.; Xu, J.; Wang, H.; Zhang, T. Functional, UV-curable coating for the capture of circulating tumor cells. Biomater. Sci. 2019, 7, 2383–2393. [Google Scholar] [CrossRef] [PubMed]
- Sathish, S.; Ishizu, N.; Shen, A.Q. Air Plasma-Enhanced Covalent Functionalization of Poly(methyl methacrylate): High-Throughput Protein Immobilization for Miniaturized Bioassays. Acs Appl. Mater. Interfaces 2019, 11, 46350–46360. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Wang, X.; Deng, L.H.; Gjertsen, B.T. A microfluidic device for differential capture of heterogeneous rare tumor cells with epithelial and mesenchymal phenotypes. Anal. Chim. Acta 2020, 1129, 1–11. [Google Scholar] [CrossRef]
- Wu, X.; Bai, Z.; Wang, L.; Cui, G.; Yang, M.; Yang, Q.; Ma, B.; Song, Q.; Tian, D.; Ceyssens, F.; et al. Magnetic Cell Centrifuge Platform Performance Study with Different Microsieve Pore Geometries. Sensors 2020, 20, 48. [Google Scholar] [CrossRef] [Green Version]
- Lim, L.S.; Hu, M.; Huang, M.C.; Cheong, W.C.; Gan, A.T.L.; Looi, X.L.; Leong, S.M.; Koay, E.S.-C.; Li, M.-H. Microsieve lab-chip device for rapid enumeration and fluorescence in situ hybridization of circulating tumor cells. Lab Chip 2012, 12, 4388–4396. [Google Scholar] [CrossRef]
- To, T.D.; Truong, A.T.T.; Nguyen, A.T.; Doan, T.C.D.; Dang, C.M. Filtration of circulating tumour cells MCF-7 in whole blood using non-modified and modified silicon nitride microsieves. Int. J. Nanotechnol. 2018, 15, 39–52. [Google Scholar] [CrossRef]
- Volpe, A.; Paie, P.; Ancona, A.; Osellame, R.; Lugara, P.M.; Pascazio, G. A computational approach to the characterization of a microfluidic device for continuous size-based inertial sorting. J. Phys. D Appl. Phys. 2017, 50. [Google Scholar] [CrossRef]
- Pereiro, I.; Fomitcheva Khartchenko, A.; Petrini, L.; Kaigala, G.V. Nip the bubble in the bud: A guide to avoid gas nucleation in microfluidics. Lab Chip 2019, 19, 2296–2314. [Google Scholar] [CrossRef] [PubMed]
- Blonski, S.; Zaremba, D.; Jachimek, M.; Jakiela, S.; Wacławczyk, T.; Korczyk, P.M. Impact of inertia and channel angles on flow distribution in microfluidic junctions. Microfluid. Nanofluidics 2020, 24, 14. [Google Scholar] [CrossRef] [Green Version]


| Repetition Rate (RR) | Pulse Energy | Scan Speed | Hatch Distance |
|---|---|---|---|
| 50 kHz | 12 μJ | 40 mm·s–1 | 5 μm |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Zoupanou, S.; Volpe, A.; Primiceri, E.; Gaudiuso, C.; Ancona, A.; Ferrara, F.; Chiriacò, M.S. SMILE Platform: An Innovative Microfluidic Approach for On-Chip Sample Manipulation and Analysis in Oral Cancer Diagnosis. Micromachines 2021, 12, 885. https://doi.org/10.3390/mi12080885
Zoupanou S, Volpe A, Primiceri E, Gaudiuso C, Ancona A, Ferrara F, Chiriacò MS. SMILE Platform: An Innovative Microfluidic Approach for On-Chip Sample Manipulation and Analysis in Oral Cancer Diagnosis. Micromachines. 2021; 12(8):885. https://doi.org/10.3390/mi12080885
Chicago/Turabian StyleZoupanou, Sofia, Annalisa Volpe, Elisabetta Primiceri, Caterina Gaudiuso, Antonio Ancona, Francesco Ferrara, and Maria Serena Chiriacò. 2021. "SMILE Platform: An Innovative Microfluidic Approach for On-Chip Sample Manipulation and Analysis in Oral Cancer Diagnosis" Micromachines 12, no. 8: 885. https://doi.org/10.3390/mi12080885
APA StyleZoupanou, S., Volpe, A., Primiceri, E., Gaudiuso, C., Ancona, A., Ferrara, F., & Chiriacò, M. S. (2021). SMILE Platform: An Innovative Microfluidic Approach for On-Chip Sample Manipulation and Analysis in Oral Cancer Diagnosis. Micromachines, 12(8), 885. https://doi.org/10.3390/mi12080885










