Digital Microfluidics for Single Bacteria Capture and Selective Retrieval Using Optical Tweezers
Abstract
1. Introduction
2. Materials and Methods
2.1. Materials
2.2. Digital Microfluidic Platform
2.3. Cell Culture
2.4. MB Functionalization
2.5. Working Buffer Optimization
2.6. Salmonella Capture on Microarray
2.7. Selective Retrieval of Single Salmonella Bacterium
3. Results and Discussion
3.1. Buffer Selection for OT-Mediated Retrieval of MBs
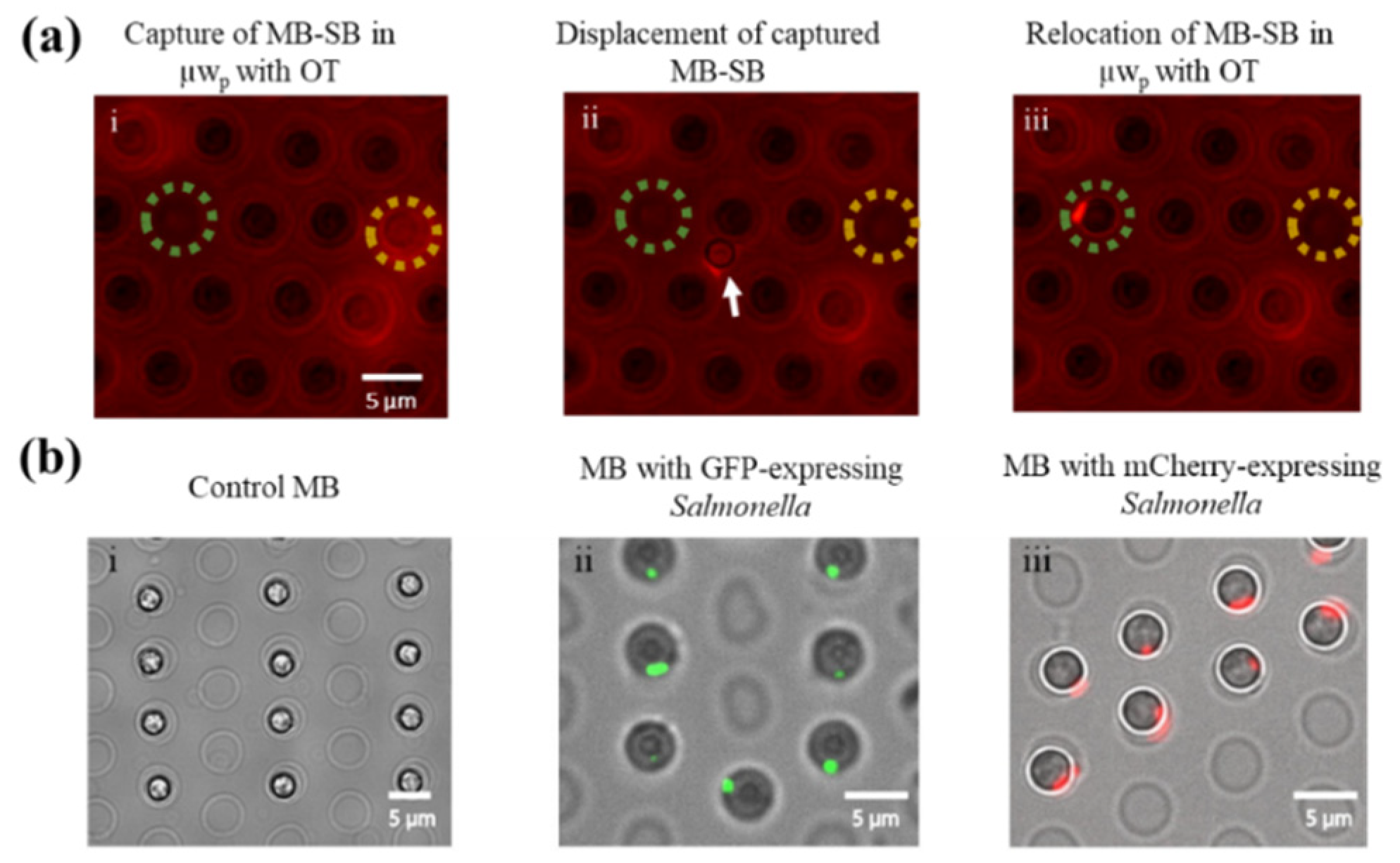
3.2. Capturing Single Salmonella Cells on MBs
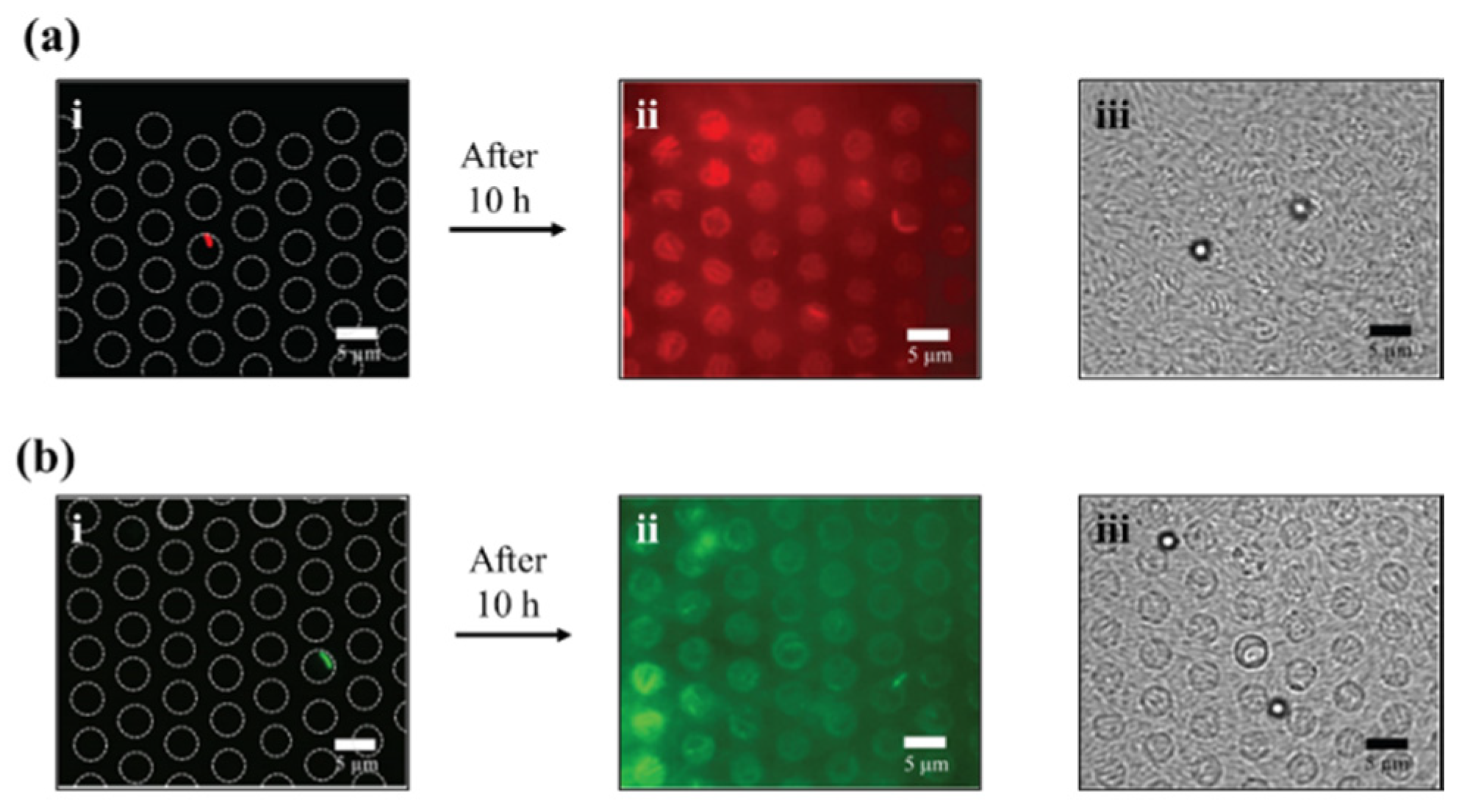
3.3. Growth in Agar Patch
4. Conclusions
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Elowitz, M.B.; Levine, A.J.; Siggia, E.D.; Swain, P.S. Stochastic gene expression in a single cell. Science 2002, 297, 1183–1186. [Google Scholar] [CrossRef]
- Glauche, I.; Thielecke, L.; Roeder, I. Cellular aging leads to functional heterogeneity of hematopoietic stem cells: A modeling perspective. Aging Cell 2011, 10, 457–465. [Google Scholar] [CrossRef]
- Huh, D.; Paulsson, J. Random partitioning of molecules at cell division. Proc. Natl. Acad. Sci. USA 2011, 108, 15004–15009. [Google Scholar] [CrossRef]
- Levchenko, A.; Nemenman, I. Cellular noise and information transmission. Curr. Opin. Biotechnol. 2014, 28, 156–164. [Google Scholar] [CrossRef]
- Govers, S.K.; Mortier, J.; Adam, A.; Aertsen, A. Protein aggregates encode epigenetic memory of stressful encounters in individual Escherichia coli cells. PLoS Biol. 2018, 16, e2003853. [Google Scholar] [CrossRef]
- Wu, H.; Zhang, X.-Y.; Hu, Z.; Hou, Q.; Zhang, H.; Li, Y.; Li, S.; Yue, J.; Jiang, Z.; Weissman, S.M.; et al. Evolution and heterogeneity of non-hereditary colorectal cancer revealed by single-cell exome sequencing. Oncogene 2017, 36, 2857–2867. [Google Scholar] [CrossRef] [PubMed]
- Kuipers, J.; Jahn, K.; Beerenwinkel, N. Advances in understanding tumour evolution through single-cell sequencing. Biochim. Biophys. Acta - Rev. Cancer 2017, 1867, 127–138. [Google Scholar] [CrossRef] [PubMed]
- Helaine, S.; Holden, D.W.; Mouton, J.M.; Sampson, S.L. Elucidating population-wide mycobacterial replication dynamics at the single-cell level. Microbiology 2016, 162, 966–978. [Google Scholar]
- Davis, K.M.; Isberg, R.R. Defining heterogeneity within bacterial populations via single cell approaches. BioEssays 2016, 38, 782–790. [Google Scholar] [CrossRef]
- Passaris, I.; Cambré, A.; Govers, S.K.; Aertsen, A. Bimodal expression of the salmonella typhimurium spv operon. Genetics 2018. [Google Scholar] [CrossRef]
- Clark, S.J.; Lee, H.J.; Smallwood, S.A.; Kelsey, G.; Reik, W. Single-cell epigenomics: powerful new methods for understanding gene regulation and cell identity. Genome Biol. 2016, 17, 72. [Google Scholar] [CrossRef] [PubMed]
- Cheow, L.F.; Courtois, E.T.; Tan, Y.; Viswanathan, R.; Xing, Q.; Tan, R.Z.; Tan, D.S.W.; Robson, P.; Loh, Y.-H.; Quake, S.R.; et al. Single-cell multimodal profiling reveals cellular epigenetic heterogeneity. Nat. Methods 2016, 13, 833–836. [Google Scholar] [CrossRef] [PubMed]
- Patel, A.P.; Tirosh, I.; Trombetta, J.J.; Shalek, A.K.; Gillespie, S.M.; Wakimoto, H.; Cahill, D.P.; Nahed, B.V.; Curry, W.T.; Martuza, R.L.; et al. Single-cell RNA-seq highlights intratumoral heterogeneity in primary glioblastoma. Science 2014, 344, 1396–1401. [Google Scholar] [CrossRef] [PubMed]
- Chen, C.-L.; Mahalingam, D.; Osmulski, P.; Jadhav, R.R.; Wang, C.-M.; Leach, R.J.; Chang, T.-C.; Weitman, S.D.; Kumar, A.P.; Sun, L.; et al. Single-cell analysis of circulating tumor cells identifies cumulative expression patterns of EMT-related genes in metastatic prostate cancer. Prostate 2013, 73, 813–826. [Google Scholar] [CrossRef]
- Eisenstein, M. Cell sorting: Divide and conquer. Nature 2006, 441, 1179–1185. [Google Scholar] [CrossRef]
- Yanez, L.Z.; Camarillo, D.B. Microfluidic analysis of oocyte and embryo biomechanical properties to improve outcomes in assisted reproductive technologies. MHR Basic Sci. Reprod. Med. 2017, 23, 235–247. [Google Scholar] [CrossRef]
- Mellors, J.S.; Jorabchi, K.; Smith, L.M.; Ramsey, J.M. Integrated Microfluidic Device for Automated Single Cell Analysis Using Electrophoretic Separation and Electrospray Ionization Mass Spectrometry. Anal. Chem. 2010, 82, 967–973. [Google Scholar] [CrossRef]
- Attayek, P.J.; Waugh, J.P.; Hunsucker, S.A.; Grayeski, P.J.; Sims, C.E.; Armistead, P.M.; Allbritton, N.L. Automated microraft platform to identify and collect non-adherent cells successfully gene-edited with CRISPR-Cas9. Biosens. Bioelectron. 2017, 91, 175–182. [Google Scholar] [CrossRef][Green Version]
- Kokkinis, G.; Cardoso, S.; Keplinger, F.; Giouroudi, I. Microfluidic platform with integrated GMR sensors for quantification of cancer cells. Sensors Actuators B Chem. 2017, 241, 438–445. [Google Scholar] [CrossRef]
- Benítez, J.J.; Topolancik, J.; Tian, H.C.; Wallin, C.B.; Latulippe, D.R.; Szeto, K.; Murphy, P.J.; Cipriany, B.R.; Levy, S.L.; Soloway, P.D.; et al. Microfluidic extraction, stretching and analysis of human chromosomal DNA from single cells. Lab Chip 2012, 12, 4848. [Google Scholar] [CrossRef]
- Iino, R.; Hayama, K.; Amezawa, H.; Sakakihara, S.; Kim, S.H.; Matsumono, Y.; Nishino, K.; Yamaguchi, A.; Noji, H. A single-cell drug efflux assay in bacteria by using a directly accessible femtoliter droplet array. Lab Chip 2012, 12, 3923. [Google Scholar] [CrossRef] [PubMed]
- Snider, A.; Pirozzi, I.; Tripathi, A. Centrifugal Microfluidics Traps for Parallel Isolation and Imaging of Single Cells. Micromachines 2020, 11, 149. [Google Scholar] [CrossRef] [PubMed]
- Zhu, C.; Luo, X.; Espulgar, W.V.; Koyama, S.; Kumanogoh, A.; Saito, M.; Takamatsu, H.; Tamiya, E. Real-Time Monitoring and Detection of Single-Cell Level Cytokine Secretion Using LSPR Technology. Micromachines 2020, 11, 107. [Google Scholar] [CrossRef] [PubMed]
- Steubing, R.W.; Cheng, S.; Wright, W.H.; Numajiri, Y.; Berns, M.W. Laser induced cell fusion in combination with optical tweezers: The laser cell fusion trap. Cytometry 1991, 12, 505–510. [Google Scholar] [CrossRef] [PubMed]
- Zhu, R.; Avsievich, T.; Bykov, A.; Popov, A.; Meglinski, I. Influence of Pulsed He–Ne Laser Irradiation on the Red Blood Cell Interaction Studied by Optical Tweezers. Micromachines 2019, 10, 853. [Google Scholar] [CrossRef] [PubMed]
- Liang, Y.; Liang, G.; Xiang, Y.; Lamstein, J.; Gautam, R.; Bezryadina, A.; Chen, Z. Manipulation and Assessment of Human Red Blood Cells with Tunable “tug-of-War” Optical Tweezers. Phys. Rev. Appl. 2019, 12, 064060. [Google Scholar] [CrossRef]
- Berns, M.W.; Wright, W.H.; Tromberg, B.J.; Profeta, G.A.; Andrews, J.J.; Walter, R.J. Use of a laser-induced optical force trap to study chromosome movement on the mitotic spindle. Proc. Natl. Acad. Sci. USA 1989, 86, 4539–4543. [Google Scholar] [CrossRef]
- Hou, J.; Luo, T.; Ng, K.L.; Leung, A.Y.H.; Liang, R.; Sun, D. Characterization of Drug Effect on Leukemia Cells Through Single Cell Assay With Optical Tweezers and Dielectrophoresis. IEEE Trans. Nanobiosci. 2016, 15, 820–827. [Google Scholar] [CrossRef]
- Ericsson, M.; Hanstorp, D.; Hagberg, P.; Enger, J.; Nyström, T. Sorting out bacterial viability with optical tweezers. J. Bacteriol. 2000, 182, 5551–5555. [Google Scholar] [CrossRef]
- Zhao, H.; Chin, L.K.; Shi, Y.; Nguyen, K.T.; Liu, P.Y.; Zhang, Y.; Zhang, M.; Zhang, J.; Cai, H.; Yap, E.P.H.; et al. Massive nanophotonic trapping and alignment of rod-shaped bacteria for parallel single-cell studies. Sens. Actuators B Chem. 2020, 306. [Google Scholar] [CrossRef]
- Probst, C.; Grünberger, A.; Wiechert, W.; Kohlheyer, D. Microfluidic growth chambers with optical tweezers for full spatial single-cell control and analysis of evolving microbes. J. Microbiol. Methods 2013, 95, 470–476. [Google Scholar] [CrossRef] [PubMed]
- Wessel, A.K.; Hmelo, L.; Parsek, M.R.; Whiteley, M. Going local: technologies for exploring bacterial microenvironments. Nat. Rev. Microbiol. 2013, 11, 337–348. [Google Scholar] [CrossRef] [PubMed]
- Quake, S.R.; Fu, A.Y.; Spence, C.; Scherer, A.; Arnold, F.H. A microfabricated fluorescence-activated cell sorter. Nat. Biotechnol. 1999, 17, 1109–1111. [Google Scholar]
- Jahan-Tigh, R.R.; Ryan, C.; Obermoser, G.; Schwarzenberger, K. Flow Cytometry. J. Invest. Dermatol. 2012, 132, 1–6. [Google Scholar] [CrossRef] [PubMed]
- Keloth, A.; Anderson, O.; Risbridger, D.; Paterson, L. Single cell isolation using optical tweezers. Micromachines 2018, 9, 434. [Google Scholar]
- Ng, A.H.C.; Dean Chamberlain, M.; Situ, H.; Lee, V.; Wheeler, A.R. Digital microfluidic immunocytochemistry in single cells. Nat. Commun. 2015, 6, 7513. [Google Scholar] [CrossRef] [PubMed]
- Kumar, P.T.; Vriens, K.; Cornaglia, M.; Gijs, M.; Kokalj, T.; Thevissen, K.; Geeraerd, A.; Cammue, B.P.A.; Puers, R.; Lammertyn, J. Digital microfluidics for time-resolved cytotoxicity studies on single non-adherent yeast cells. Lab Chip 2015, 15, 1852–1860. [Google Scholar] [CrossRef]
- Decrop, D.; Brans, T.; Gijsenbergh, P.; Lu, J.; Spasic, D.; Kokalj, T.; Beunis, F.; Goos, P.; Puers, R.; Lammertyn, J. Optical Manipulation of Single Magnetic Beads in a Microwell Array on a Digital Microfluidic Chip. Anal. Chem. 2016, 88, 8596–8603. [Google Scholar] [CrossRef]
- Decrop, D.; Brans, T.; Kokalj, T.; Beunis, F.; Lammertyn, J. Magnetic particle retrieval and positioning in a microwell array by integrating optical tweezers in a digital microfluidic platform. In Proceedings of the 18th International Conference on Miniaturized Systems for Chemistry and Life Sciences, MicroTAS 2014, San Antonio, TX, USA, 26–30 October 2014; pp. 1455–1457. [Google Scholar]
- Decrop, D.; Ruiz, E.P.; Kumar, P.T.; Tripodi, L.; Kokalj, T.; Lammertyn, J. Digital microfluidics assisted sealing of individual magnetic particles in femtoliter-sized reaction wells for single-molecule detection. In Methods in Molecular Biology; Humana Press: New York, NY, USA, 2017; Volume 1547, pp. 85–101. [Google Scholar]
- Witters, D.; Knez, K.; Ceyssens, F.; Puers, R.; Lammertyn, J. Digital microfluidics-enabled single-molecule detection by printing and sealing single magnetic beads in femtoliter droplets. Lab Chip 2013, 13, 2047. [Google Scholar] [CrossRef]
- Passaris, I.; Ghosh, A.; Cenens, W.; Michiels, C.W.; Lammertyn, J.; Aertsen, A. Isolation and Validation of an Endogenous Fluorescent Nucleoid Reporter in Salmonella Typhimurium. PLoS ONE 2014, 9, e93785. [Google Scholar] [CrossRef]
- Yonekura, K.; Maki-Yonekura, S.; Namba, K. Complete atomic model of the bacterial flagellar filament by electron cryomicroscopy. Nature 2003, 424, 643–650. [Google Scholar] [CrossRef] [PubMed]
- Aertsen, A.; Tesfazgi Mebrhatu, M.; Michiels, C.W. Activation of the Salmonella Typhimurium Mrr protein. Biochem. Biophys. Res. Commun. 2008, 367, 435–439. [Google Scholar] [CrossRef] [PubMed]
- Bertani, G. Lysogeny at mid-twentieth century: P1, P2, and other experimental systems. J. Bacteriol. 2004, 186, 595–600. [Google Scholar] [CrossRef] [PubMed]
- Park, G.J. Analytic Methods for Design Practice; Springer: London, UK, 2007; ISBN 9781846284724. [Google Scholar]
- Hajmeer, M.; Ceylan, E.; Marsden, J.L.; Fung, D.Y.C. Impact of sodium chloride on Escherichia coli O157:H7 and Staphylococcus aureus analysed using transmission electron microscopy. Food Microbiol. 2006, 23, 446–452. [Google Scholar] [CrossRef] [PubMed]
- Thayer, D.W.; Muller, W.S.; Buchanan, R.L.; Phillips, J.G. Effect of NaCl, pH, temperature, and atmosphere on growth of Salmonella typhimurium in glucose-mineral salts medium. Appl. Environ. Microbiol. 1987, 53, 1311–1315. [Google Scholar] [CrossRef] [PubMed]
- Hamzeloo-Moghadam, M.; Taiebi, N.; Mosaddegh, M.; Tehrani, B.E.; Esmaeili, S. The effect of some cosolvents and surfactants on viability of cancerous cell lines. Res. J. pharmacogn. 2014, 1, 41–45. [Google Scholar]
- D’Aoust, J.-Y.; Maishment, C.; Stotland, P.; Boville, A. Surfactants for the Effective Recovery of Salmonella in Fatty Foods. J. Food Prot. 1982, 45, 249–252. [Google Scholar] [CrossRef]
- Breukers, J.; Struyfs, C.; Decrop, D.; Kil, D.; Thevissen, K.; Cammue, B.; Puers, R.; Lammertyn, J. Tuning the surface interactions between single cells and an OSTE+ microwell array for enhanced optical tweezers manipulation. In Proceedings of the Miniaturized Systems for Chemistry and Life Sciences (MicroTAS) Conference, Kaohsiung, Taiwan, 11–15 November 2018. [Google Scholar]
- Molloy, J.E.; Padgett, M.J. Lights, action: Optical tweezers. Contemp. Phys. 2002, 43, 241–258. [Google Scholar] [CrossRef]
- Zhang, H.; Liu, K.-K. Optical tweezers for single cells. J. R. Soc. Interface 2008, 5, 671–690. [Google Scholar] [CrossRef]
- Zhang, Z.; Kimkes, T.E.P.; Heinemann, M. Manipulating rod-shaped bacteria with optical tweezers. Sci. Rep. 2019, 9, 19086. [Google Scholar] [CrossRef]
- Neuman, K.C.; Nagy, A. Single-molecule force spectroscopy: Optical tweezers, magnetic tweezers and atomic force microscopy. Nat. Methods 2008, 5, 491–505. [Google Scholar] [CrossRef] [PubMed]



© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Tewari Kumar, P.; Decrop, D.; Safdar, S.; Passaris, I.; Kokalj, T.; Puers, R.; Aertsen, A.; Spasic, D.; Lammertyn, J. Digital Microfluidics for Single Bacteria Capture and Selective Retrieval Using Optical Tweezers. Micromachines 2020, 11, 308. https://doi.org/10.3390/mi11030308
Tewari Kumar P, Decrop D, Safdar S, Passaris I, Kokalj T, Puers R, Aertsen A, Spasic D, Lammertyn J. Digital Microfluidics for Single Bacteria Capture and Selective Retrieval Using Optical Tweezers. Micromachines. 2020; 11(3):308. https://doi.org/10.3390/mi11030308
Chicago/Turabian StyleTewari Kumar, Phalguni, Deborah Decrop, Saba Safdar, Ioannis Passaris, Tadej Kokalj, Robert Puers, Abram Aertsen, Dragana Spasic, and Jeroen Lammertyn. 2020. "Digital Microfluidics for Single Bacteria Capture and Selective Retrieval Using Optical Tweezers" Micromachines 11, no. 3: 308. https://doi.org/10.3390/mi11030308
APA StyleTewari Kumar, P., Decrop, D., Safdar, S., Passaris, I., Kokalj, T., Puers, R., Aertsen, A., Spasic, D., & Lammertyn, J. (2020). Digital Microfluidics for Single Bacteria Capture and Selective Retrieval Using Optical Tweezers. Micromachines, 11(3), 308. https://doi.org/10.3390/mi11030308





