Electrochemical Voltammogram Recording for Identifying Varieties of Ornamental Plants
Abstract
:1. Introduction
2. Materials and Methods
3. Results
4. Conclusions
Author Contributions
Funding
Conflicts of Interest
References
- Anandan, S.; Rudolph, A.; Speck, T.; Speck, O. Comparative morphological and anatomical study of self-repair in succulent cylindrical plant organs. Flora 2018, 241, 1–7. [Google Scholar] [CrossRef]
- Sil, S.; Mallick, T.; De, K.K.; Pramanik, A.; Ghosh, A. Comparative morphological study of three species of Saraca L. (Fabaceae) by the statistical approach to find out the logic of potent morphological markers. Beni-Suef Univ. J. Basic Appl. Sci. 2018, 7, 612–619. [Google Scholar] [CrossRef]
- Yao, R.; Heinrich, M.; Zou, Y.; Reich, E.; Zhang, X.; Chen, Y.; Weckerle, C.S. Quality variation of Goji (fruits of Lycium spp.) in China: A comparative morphological and metabolomic analysis. Front. Pharmacol. 2018, 9, 151. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Pimenov, M.; Degtjareva, G.; Ostroumova, T.; Samigullin, T.; Zakharova, E. What is Seseli diffusum? A comparative morphological and molecular appraisal of a critical species of the Umbelliferae. Plant Syst. Evol. 2019, 305, 49–59. [Google Scholar] [CrossRef]
- Meng, Z.; Zhang, Z.; Yan, T.; Lin, Q.; Wang, Y.; Huang, W.; Huang, Y.; Li, Z.; Yu, Q.; Wang, J. Comprehensively characterizing the cytological features of Saccharum spontaneum by the development of a complete set of chromosome-specific oligo probes. Front. Plant Sci. 2018, 9, 1624. [Google Scholar] [CrossRef] [PubMed]
- Suzuki, A.; Hirokawa, M.; Ito, A.; Takada, N.; Higuchi, M.; Hayashi, T.; Kuma, S.; Miyauchi, A. Identification of cytological features distinguishing mucosa-associated lymphoid tissue lymphoma from reactive lymphoid proliferation using thyroid liquid-based cytology. Acta Cytol. 2018, 62, 93–98. [Google Scholar] [CrossRef]
- Song, F.; Meng, Z.; Luo, T.; Xin, J.; Xian, M.; Rao, N.; Chen, Q.; Wang, Y.; Khan, M.N.; Hu, L. Cytological identification of new-type Brassica napus materials and their physiological response to drought. Crop Pasture Sci. 2019, 70, 876–889. [Google Scholar] [CrossRef]
- Lan, Y.; Lianwei, Q.; Xin, H.; Gong, H.; Lei, J.; Xi, M. Physical mapping of rDNA and karyotype analysis in Tulipa sinkiangensis and T. schrenkii. Sci. Hortic. 2018, 240, 638–644. [Google Scholar] [CrossRef]
- Bracewell, R.; Chatla, K.; Nalley, M.J.; Bachtrog, D. Dynamic turnover of centromeres drives karyotype evolution in Drosophila. eLife 2019, 8, e49002. [Google Scholar] [CrossRef]
- Deng, H.; Cai, Z.; Xiang, S.; Guo, Q.; Huang, W.; Liang, G. Karyotype Analysis of Diploid and Spontaneously Occurring Tetraploid Blood Orange [Citrus sinensis (L.) Osbeck] Using Multicolor FISH With Repetitive DNA Sequences as Probes. Front. Plant Sci. 2019, 10, 331. [Google Scholar] [CrossRef]
- Xin, H.; Zhang, T.; Wu, Y.; Zhang, W.; Zhang, P.; Xi, M.; Jiang, J. An extraordinarily stable karyotype of the woody Populus species revealed by chromosome painting. Plant J. 2020, 101, 253–264. [Google Scholar] [CrossRef] [PubMed]
- Leontidou, K.; Vernesi, C.; De Groeve, J.; Cristofolini, F.; Vokou, D.; Cristofori, A. DNA metabarcoding of airborne pollen: New protocols for improved taxonomic identification of environmental samples. Aerobiologia 2018, 34, 63–74. [Google Scholar] [CrossRef]
- Zhang, X.; Huang, K.-Y.; Zheng, Z.; Zhang, Y.-Z.; Wan, Q.-C.; Tian, L.-P. Pollen morphology of Quercus sect. Ilex and its relevance for fossil pollen identification in southwest China. Grana 2018, 57, 401–414. [Google Scholar] [CrossRef]
- Zimmermann, B. Chemical characterization and identification of Pinaceae pollen by infrared microspectroscopy. Planta 2018, 247, 171–180. [Google Scholar] [CrossRef]
- Geisen, S.; Snoek, L.B.; ten Hooven, F.C.; Duyts, H.; Kostenko, O.; Bloem, J.; Martens, H.; Quist, C.W.; Helder, J.A.; van der Putten, W.H. Integrating quantitative morphological and qualitative molecular methods to analyse soil nematode community responses to plant range expansion. Methods Ecol. Evol. 2018, 9, 1366–1378. [Google Scholar] [CrossRef] [Green Version]
- Mi, J.; Jia, K.-P.; Wang, J.Y.; Al-Babili, S. A rapid LC-MS method for qualitative and quantitative profiling of plant apocarotenoids. Anal. Chim. Acta 2018, 1035, 87–95. [Google Scholar] [CrossRef]
- He, K.; Chang, L.; Dong, Y.; Cui, T.; Qu, J.; Liu, X.; Xu, S.; Xue, J.; Liu, J. Identification of quantitative trait loci for agronomic and physiological traits in maize (Zea mays L.) under high-nitrogen and low-nitrogen conditions. Euphytica 2018, 214, 15. [Google Scholar] [CrossRef]
- Nomura, T.; Kuchida, R.; Kitaoka, N.; Kato, Y. Molecular diversity of tuliposide B-converting enzyme in tulip (Tulipa gesneriana): Identification of the third isozyme with a distinct expression profile. Biosci. Biotechnol. Biochem. 2018, 82, 810–820. [Google Scholar] [CrossRef]
- Mano, J.; Kanameda, S.; Kuramitsu, R.; Matsuura, N.; Yamauchi, Y. Detoxification of reactive carbonyl species by glutathione transferase Tau isozymes. Front. Plant Sci. 2019, 10, 487. [Google Scholar] [CrossRef] [Green Version]
- Chaichompoo, W.; Chokchaisiri, R.; Sangkaew, A.; Pabuprapap, W.; Yompakdee, C.; Suksamrarn, A. Alkaloids with anti-human carbonic anhydrase isozyme II activity from the bulbs of Crinum asiaticum L. var. asiaticum. Phytochem. Lett. 2020, 37, 101–105. [Google Scholar] [CrossRef]
- Hussein, E.; Aly, A.; El-Awamri, A.; Habeb, M. Use of Electrophoretic Patterns of Proteins and Isozymes to Characterize Trichoderma Isolates from Cotton Roots. J. Agric. Chem. Biotechnol. 2018, 9, 219–222. [Google Scholar] [CrossRef] [Green Version]
- Caburatan, L.; Kim, J.; Park, J. Expression Profiles and Post-Translational Modifications of Phosphoenolpyruvate Carboxylase Isozymes of Bienertia sinuspersici during Leaf Development. Russ. J. Plant Physiol. 2019, 66, 738–747. [Google Scholar] [CrossRef]
- Garrido-Cardenas, J.A.; Mesa-Valle, C.; Manzano-Agugliaro, F. Trends in plant research using molecular markers. Planta 2018, 247, 543–557. [Google Scholar] [CrossRef]
- Nadeem, M.A.; Nawaz, M.A.; Shahid, M.Q.; Doğan, Y.; Comertpay, G.; Yıldız, M.; Hatipoğlu, R.; Ahmad, F.; Alsaleh, A.; Labhane, N.; et al. DNA molecular markers in plant breeding: Current status and recent advancements in genomic selection and genome editing. Biotechnol. Biotechnol. Equip. 2018, 32, 261–285. [Google Scholar] [CrossRef] [Green Version]
- Afshar, S.; Zamani, H.A.; Karimi-Maleh, H. NiO/SWCNTs coupled with an ionic liquid composite for amplified carbon paste electrode; A feasible approach for improving sensing ability of adrenalone and folic acid in dosage form. J. Pharm. Biomed. Anal. 2020, 188, 113393. [Google Scholar] [CrossRef]
- Alamgholiloo, H.; Rostamnia, S.; Hassankhani, A.; Liu, X.; Eftekhari, A.; Hasanzadeh, A.; Zhang, K.; Karimi-Maleh, H.; Khaksar, S.; Varma, R.S.; et al. Formation and stabilization of colloidal ultra-small palladium nanoparticles on diamine-modified Cr-MIL-101: Synergic boost to hydrogen production from formic acid. J. Colloid Interface Sci. 2020, 567, 126–135. [Google Scholar] [CrossRef]
- Fouladgar, M.; Karimi-Maleh, H.; Opoku, F.; Govender, P.P. Electrochemical anticancer drug sensor for determination of raloxifene in the presence of tamoxifen using graphene-CuO-polypyrrole nanocomposite structure modified pencil graphite electrode: Theoretical and experimental investigation. J. Mol. Liq. 2020, 311, 113314. [Google Scholar] [CrossRef]
- Fu, L.; Zheng, Y.; Zhang, P.; Zhang, H.; Xu, Y.; Zhou, J.; Zhang, H.; Karimi-Maleh, H.; Lai, G.; Zhao, S.; et al. Development of an electrochemical biosensor for phylogenetic analysis of Amaryllidaceae based on the enhanced electrochemical fingerprint recorded from plant tissue. Biosens. Bioelectron. 2020, 159, 112212. [Google Scholar] [CrossRef]
- Karimi-Maleh, H.; Karimi, F.; Malekmohammadi, S.; Zakariae, N.; Esmaeili, R.; Rostamnia, S.; Yola, M.L.; Atar, N.; Movaghgharnezhad, S.; Rajendran, S.; et al. An amplified voltametric sensor based on platinum nanoparticle/polyoxometalate/two-dimensional hexagonal boron nitride nanosheets composite and ionic liquid for determination of N-hydroxysuccinimide in water samples. J. Mol. Liq. 2020, 310, 113185. [Google Scholar] [CrossRef]
- Karimi-Maleh, H.; Cellat, K.; Arıkan, K.; Savk, A.; Karimi, F.; Şen, F. Palladium–Nickel nanoparticles decorated on Functionalized-MWCNT for high precision non-enzymatic glucose sensing. Mater. Chem. Phys. 2020, 250, 123042. [Google Scholar] [CrossRef]
- Karimi-Maleh, H.; Kumar, B.G.; Rajendran, S.; Qin, J.; Vadivel, S.; Durgalakshmi, D.; Gracia, F.; Soto-Moscoso, M.; Orooji, Y.; Karimi, F. Tuning of metal oxides photocatalytic performance using Ag nanoparticles integration. J. Mol. Liq. 2020, 314, 113588. [Google Scholar] [CrossRef]
- Mohanraj, J.; Durgalakshmi, D.; Rakkesh, R.A.; Balakumar, S.; Rajendran, S.; Karimi-Maleh, H. Facile synthesis of paper based graphene electrodes for point of care devices: A double stranded DNA (dsDNA) biosensor. J. Colloid Interface Sci. 2020, 566, 463–472. [Google Scholar] [CrossRef] [PubMed]
- Tavana, T.; Rezvani, A.R.; Karimi-Maleh, H. Pt-Pd-doped NiO nanoparticle decorated at single-wall carbon nanotubes: An excellent, powerful electrocatalyst for the fabrication of An electrochemical sensor to determine nalbuphine in the presence of tramadol as two opioid analgesic drugs. J. Pharm. Biomed. Anal. 2020, 189, 113397. [Google Scholar] [CrossRef] [PubMed]
- Zabihpour, T.; Shahidi, S.-A.; Karimi-Maleh, H.; Ghorbani-HasanSaraei, A. An ultrasensitive electroanalytical sensor based on MgO/SWCNTs- 1-Butyl-3-methylimidazolium bis(trifluoromethylsulfonyl)imide paste electrode for the determination of ferulic acid in the presence sulfite in food samples. Microchem. J. 2020, 154, 104572. [Google Scholar] [CrossRef]
- Jiang, M.; Wang, J.; Zhang, H. The complete plastome sequence of Clematis guniuensis (Ranunculaceae), a new plant species endemic to China. Mitochondrial DNA Part B 2020, 5, 408–409. [Google Scholar] [CrossRef] [Green Version]
- Mao, C.; Zhang, X.; Shi, J.; Chen, S. The complete chloroplast genome sequence of Clematis Montana Buch.-Ham. (Ranunculaceae) and its phylogenetic analysis. Mitochondrial DNA Part B 2020, 5, 2246–2247. [Google Scholar] [CrossRef]
- Yang, Y.-C.; Wang, N.; Zhang, W.; Zhou, T. The complete chloroplast genome of Clematis fruticosa Turcz. (Ranunculaceae). Mitochondrial DNA Part B 2020, 5, 1908–1909. [Google Scholar] [CrossRef]
- Zhou, J.; Wu, M.; Xu, Y.; Li, Z.; Yao, Y.; Fu, L. 2D Pattern Recognition of White Spirit Based on the Electrochemical Profile Recorded by Screen-Printed Electrode. Int. J. Electrochem. Sci. 2020, 15, 5793–5802. [Google Scholar] [CrossRef]
- Fu, L.; Zhang, H.; Zheng, Y.; Zhang, H.; Liu, Q. An electroanalytical method for brewing vinegar authentic identification. Rev. Mex. Ing. Química 2020, 19, 803–812. [Google Scholar]
- Fu, L.; Zheng, Y.; Zhang, P.; Zhang, H.; Wu, M.; Zhang, H.; Wang, A.; Su, W.; Chen, F.; Yu, J.; et al. An electrochemical method for plant species determination and classification based on fingerprinting petal tissue. Bioelectrochemistry 2019, 129, 199–205. [Google Scholar] [CrossRef]
- Fu, L.; Wang, Q.; Zhang, M.; Zheng, Y.; Wu, M.; Lan, Z.; Pu, J.; Zhang, H.; Chen, F.; Su, W. Electrochemical sex determination of dioecious plants using polydopamine-functionalized graphene sheets. Front. Chem. 2020, 8, 92. [Google Scholar] [CrossRef]
- Fu, L.; Wang, A.; Xie, K.; Zhu, J.; Chen, F.; Wang, H.; Zhang, H.; Su, W.; Wang, Z.; Zhou, C.; et al. Electrochemical detection of silver ions by using sulfur quantum dots modified gold electrode. Sens. Actuators B Chem. 2020, 304, 127390. [Google Scholar] [CrossRef]
- Xu, Y.; Lu, Y.; Zhang, P.; Wang, Y.; Zheng, Y.; Fu, L.; Zhang, H.; Lin, C.-T.; Yu, A. Infrageneric phylogenetics investigation of Chimonanthus based on electroactive compound profiles. Bioelectrochemistry 2020, 133, 107455. [Google Scholar] [CrossRef] [PubMed]
- Fu, L.; Wu, M.; Zheng, Y.; Zhang, P.; Ye, C.; Zhang, H.; Wang, K.; Su, W.; Chen, F.; Yu, J.; et al. Lycoris species identification and infrageneric relationship investigation via graphene enhanced electrochemical fingerprinting of pollen. Sens. Actuators B Chem. 2019, 298, 126836. [Google Scholar] [CrossRef]
- Zhang, M.; Pan, B.; Wang, Y.; Du, X.; Fu, L.; Zheng, Y.; Chen, F.; Wu, W.; Zhou, Q.; Ding, S. Recording the Electrochemical Profile of Pueraria Leaves for Polyphyly Analysis. ChemistrySelect 2020, 5, 5035–5040. [Google Scholar] [CrossRef]
- Liu, Z.; Bruins, M.E.; de Bruijn, W.J.C.; Vincken, J.-P. A comparison of the phenolic composition of old and young tea leaves reveals a decrease in flavanols and phenolic acids and an increase in flavonols upon tea leaf maturation. J. Food Compos. Anal. 2020, 86, 103385. [Google Scholar] [CrossRef]
- Arvand, M.; Farahpour, M.; Ardaki, M.S. Electrochemical characterization of in situ functionalized gold organosulfur self-assembled monolayer with conducting polymer and carbon nanotubes for determination of rutin. Talanta 2018, 176, 92–101. [Google Scholar] [CrossRef]
- Lu, Y.; Xu, Y.; Shi, H.; Zhang, P.Z.H.; Fu, L. Feasibility of electrochemical fingerprinting for plant phylogeography study: A case of Chimonanthus praecox. Int. J. Electrochem. Sci. 2020, 15, 758–764. [Google Scholar] [CrossRef]
- Xing, R.; Tong, L.; Zhao, X.; Liu, H.; Ma, P.; Zhao, J.; Liu, X.; Liu, S. Rapid and sensitive electrochemical detection of myricetin based on polyoxometalates/SnO2/gold nanoparticles ternary nanocomposite film electrode. Sens. Actuators B Chem. 2019, 283, 35–41. [Google Scholar] [CrossRef]
- Irakli, M.; Chatzopoulou, P.; Ekateriniadou, L. Optimization of ultrasound-assisted extraction of phenolic compounds: Oleuropein, phenolic acids, phenolic alcohols and flavonoids from olive leaves and evaluation of its antioxidant activities. Ind. Crop. Prod. 2018, 124, 382–388. [Google Scholar] [CrossRef]
- Di Marino, D.; Jestel, T.; Marks, C.; Viell, J.; Blindert, M.; Kriescher, S.M.; Spiess, A.C.; Wessling, M. Carboxylic acids production via electrochemical depolymerization of lignin. ChemElectroChem 2019, 6, 1434–1442. [Google Scholar] [CrossRef]
- García-Carmona, L.; González, M.C.; Escarpa, A. Nanomaterial-based electrochemical (bio)-sensing: One step ahead in diagnostic and monitoring of metabolic rare diseases. TrAC Trends Anal. Chem. 2019, 118, 29–42. [Google Scholar] [CrossRef]
- Suprun, E.V. Protein post-translational modifications–A challenge for bioelectrochemistry. TrAC Trends Anal. Chem. 2019, 116, 44–60. [Google Scholar] [CrossRef]
- Švorc, Ľ.; Haššo, M.; Sarakhman, O.; Kianičková, K.; Stanković, D.M.; Otřísal, P. A progressive electrochemical sensor for food quality control: Reliable determination of theobromine in chocolate products using a miniaturized boron-doped diamond electrode. Microchem. J. 2018, 142, 297–304. [Google Scholar] [CrossRef]
- Serrano, N.; Cetó, X.; Núñez, O.; Aragó, M.; Gámez, A.; Ariño, C.; Díaz-Cruz, J.M. Characterization and classification of Spanish paprika (Capsicum annuum L.) by liquid chromatography coupled to electrochemical detection with screen-printed carbon-based nanomaterials electrodes. Talanta 2018, 189, 296–301. [Google Scholar] [CrossRef]
- Kramer, K.; Yeboah-Awudzi, M.; Magazine, N.; King, J.M.; Xu, Z.; Losso, J.N. Procyanidin B2 rich cocoa extracts inhibit inflammation in Caco-2 cell model of in vitro celiac disease by down-regulating interferon-gamma- or gliadin peptide 31-43-induced transglutaminase-2 and interleukin-15. J. Funct. Foods 2019, 57, 112–120. [Google Scholar] [CrossRef]
- Marín-Sáez, J.; Romero-González, R.; Garrido Frenich, A. Effect of tea making and boiling processes on the degradation of tropane alkaloids in tea and pasta samples contaminated with Solanaceae seeds and coca leaf. Food Chem. 2019, 287, 265–272. [Google Scholar] [CrossRef]
- Volochanskyi, O.; Švecová, M.; Prokopec, V. Detection and identification of medically important alkaloids using the surface-enhanced Raman scattering spectroscopy. Spectrochim. Acta Part A Mol. Biomol. Spectrosc. 2019, 207, 143–149. [Google Scholar] [CrossRef]
- Baran, E.; Cakir, A.; Yazici, B. Inhibitory effect of Gentiana olivieri extracts on the corrosion of mild steel in 0.5M HCl: Electrochemical and phytochemical evaluation. Arab. J. Chem. 2019, 12, 4303–4319. [Google Scholar] [CrossRef] [Green Version]
- Chou, S.; Chen, B.; Chen, J.; Wang, M.; Wang, S.; Croft, H.; Shi, Q. Estimation of leaf photosynthetic capacity from the photochemical reflectance index and leaf pigments. Ecol. Indic. 2020, 110, 105867. [Google Scholar] [CrossRef]
- Çakal, D.; Ertan, S.; Cihaner, A.; Önal, A.M. Electrochemical and optical properties of substituted phthalimide based monomers and electrochemical polymerization of 3, 4-ethylenedioxythiophene-polyhedral oligomeric silsesquioxane (POSS) analogue. Dye. Pigment. 2019, 161, 411–418. [Google Scholar] [CrossRef]
- Xue, Y.-N.; Xue, X.-Z.; Miao, M.; Liu, J.-K. Mass preparation and anticorrosion mechanism of highly triple-effective corrosion inhibition performance for co-modified zinc phosphate-based pigments. Dye. Pigment. 2019, 161, 489–499. [Google Scholar] [CrossRef]
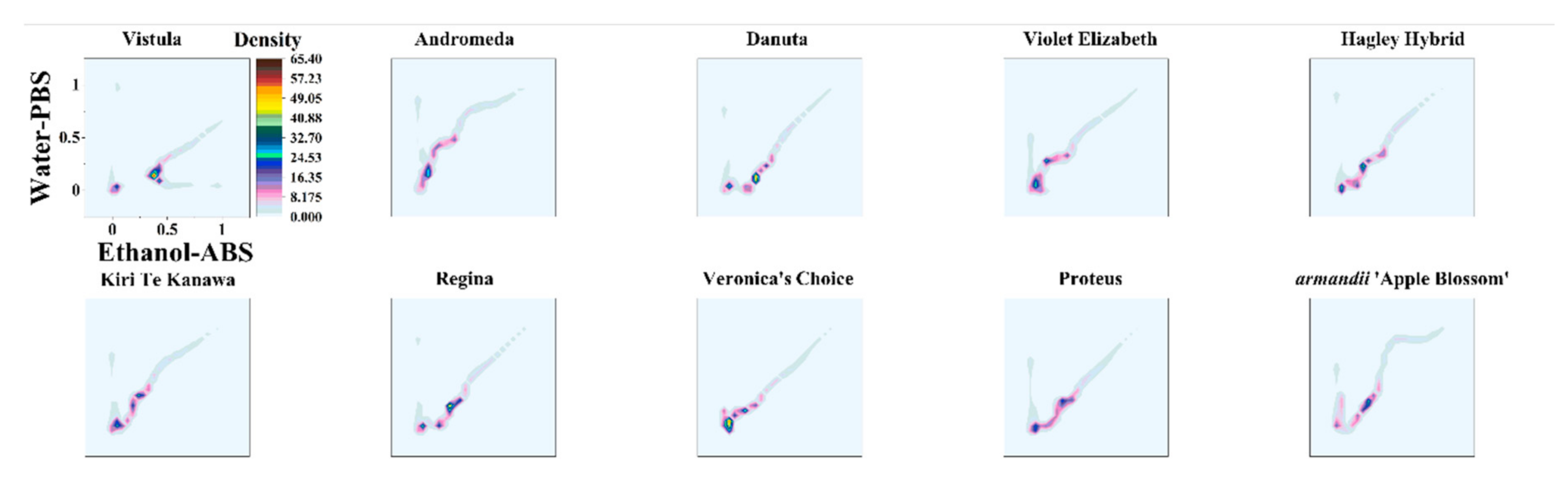
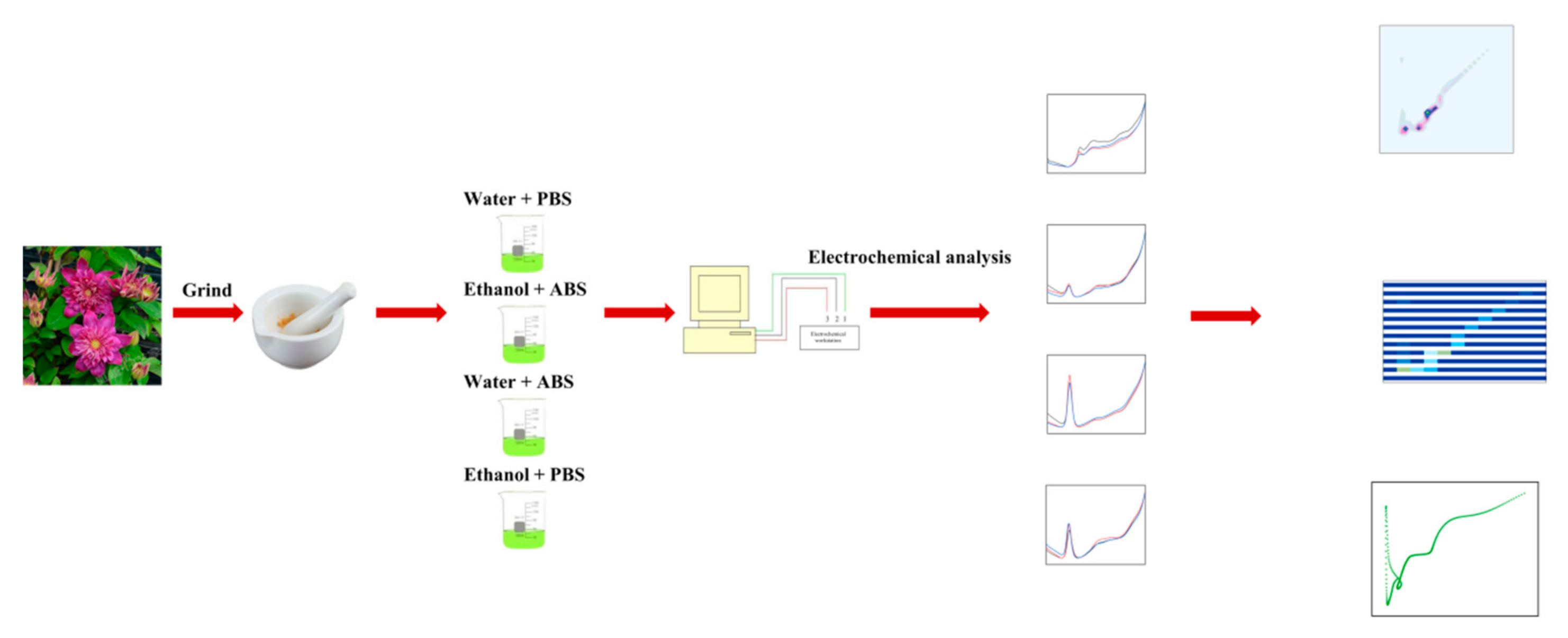
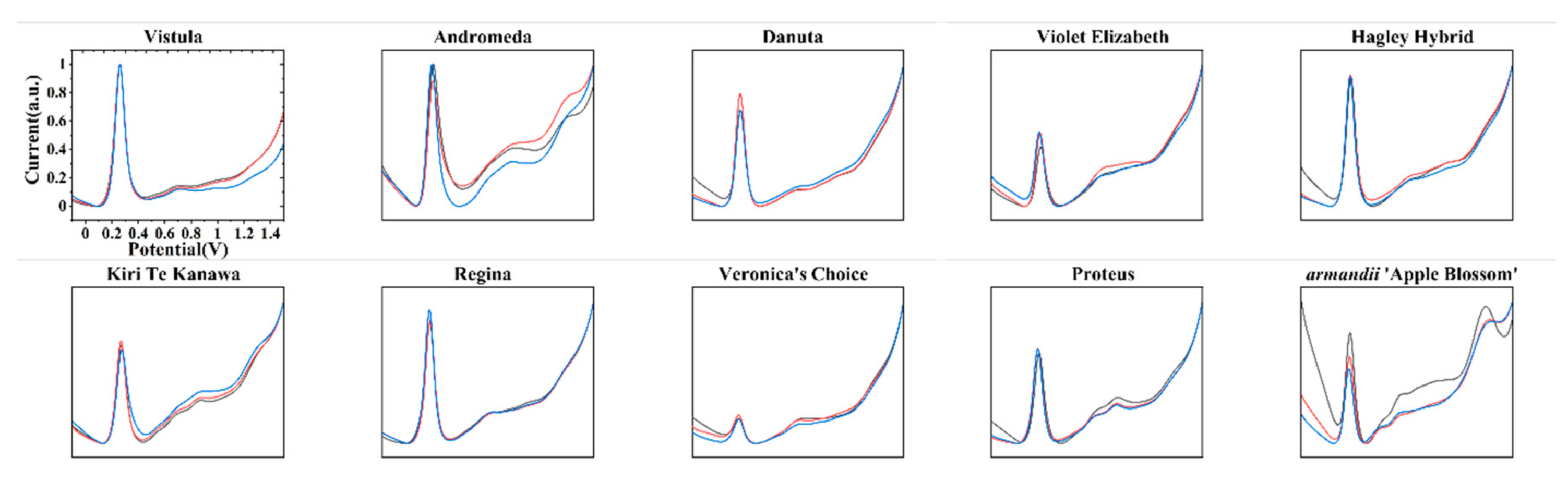
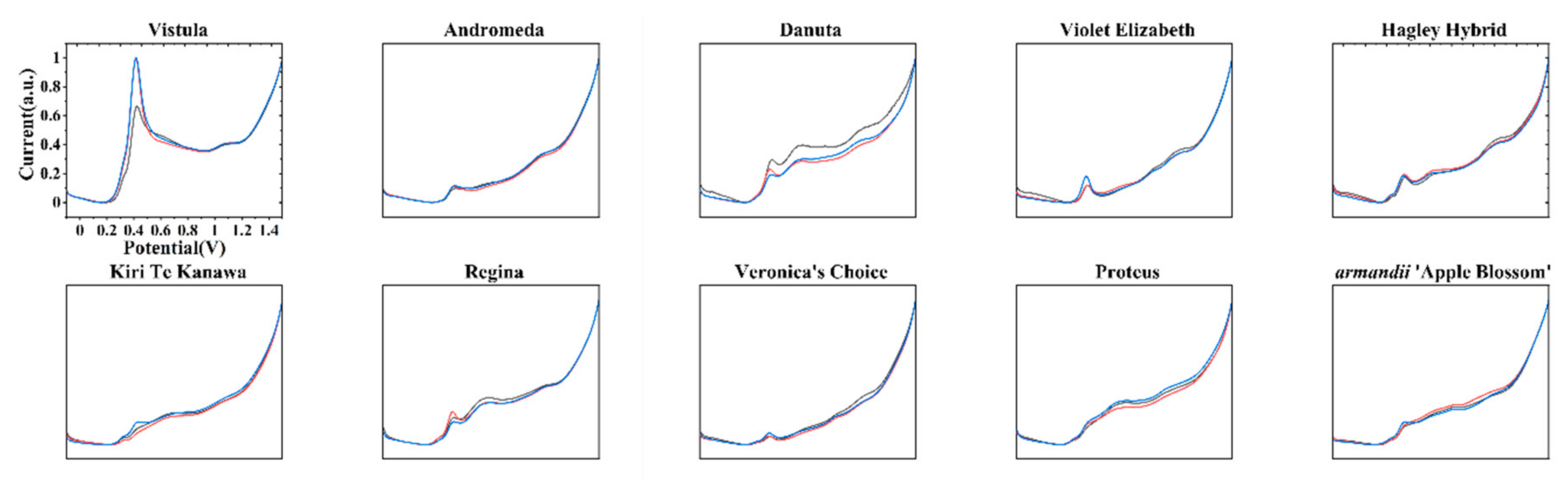


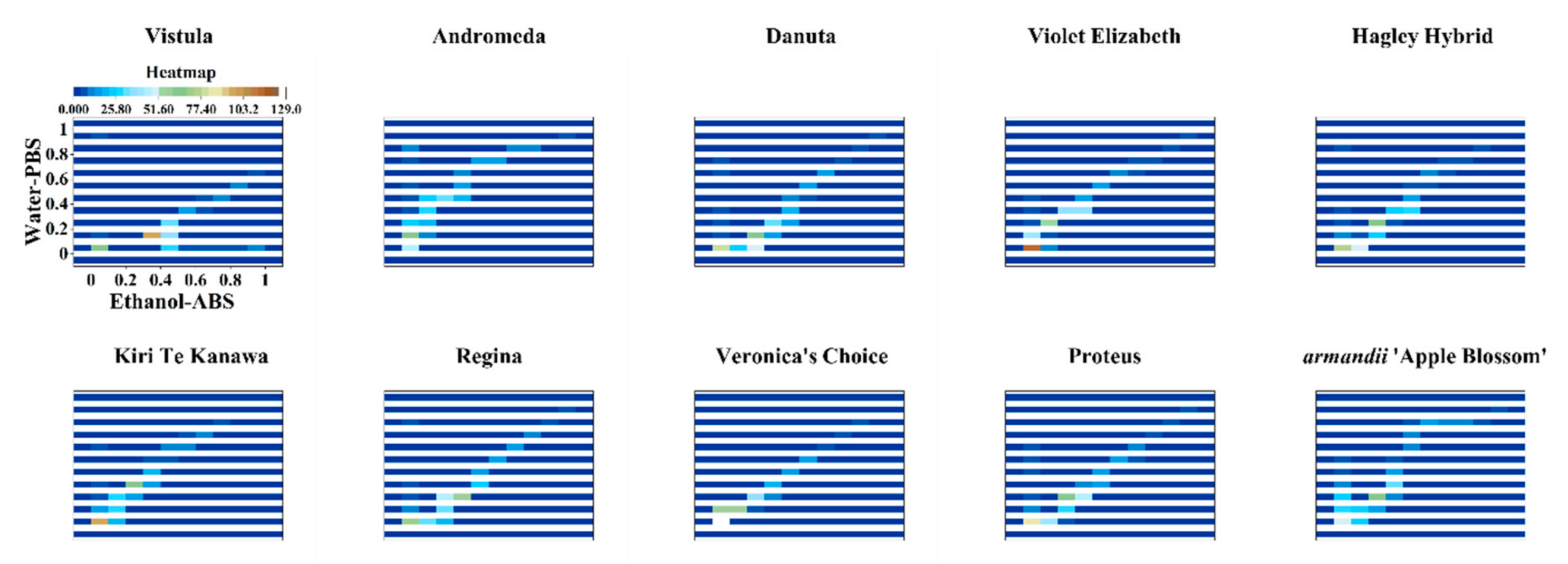
| Variety Name | Group | Approximate Height | Country of Origin | Parentage |
|---|---|---|---|---|
| Vistula | Early large-flowered group | 2.0–3.0 m | Poland | Unknown |
| Andromeda | Early large-flowered group | 2.0–4.0 m | United Kingdom | Seedling of ‘General Sikorski’ |
| Danuta | - | 2.0–2.5 m | Poland | Unknown |
| Armandii ‘Apple Blossom’ | Armandii group | 5.0–7.0 m | China | Unknown |
| Proteus | Early large-flowered group | 2.5–3.0 m | United Kingdom | C. viticella ‘Grandiflora’ x ‘Fortunei |
| Hagley Hybrid | Late large-flowered group | 2.0–3.0 m | United Kingdom | Unknown |
| Violet Elizabeth | Early large-flowered group | 2.0–3.5 m | United Kingdom | ‘Vyvyan Pennell’ x ‘Mrs. Spencer |
| Kiri Te Kanawa | Early large-flowered group | 2.0–3.0 m | United Kingdom | ‘Beauty of Worcester’ x ‘Chalcedony’ |
| Regina | Early large-flowered group | 1.5–2.0 m | Poland | Unknown |
| Veronica’s Choice | Early large-flowered group | 2.5–3.0 m | United Kingdom | ‘Vyvyan Pennell’ x ‘Percy Lake’ |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Yang, R.; Fan, B.; Wang, S.; Li, L.; Li, Y.; Li, S.; Zheng, Y.; Fu, L.; Lin, C.-T. Electrochemical Voltammogram Recording for Identifying Varieties of Ornamental Plants. Micromachines 2020, 11, 967. https://doi.org/10.3390/mi11110967
Yang R, Fan B, Wang S, Li L, Li Y, Li S, Zheng Y, Fu L, Lin C-T. Electrochemical Voltammogram Recording for Identifying Varieties of Ornamental Plants. Micromachines. 2020; 11(11):967. https://doi.org/10.3390/mi11110967
Chicago/Turabian StyleYang, Rutong, Boyuan Fan, Shu’an Wang, Linfang Li, Ya Li, Sumei Li, Yuhong Zheng, Li Fu, and Cheng-Te Lin. 2020. "Electrochemical Voltammogram Recording for Identifying Varieties of Ornamental Plants" Micromachines 11, no. 11: 967. https://doi.org/10.3390/mi11110967
APA StyleYang, R., Fan, B., Wang, S., Li, L., Li, Y., Li, S., Zheng, Y., Fu, L., & Lin, C.-T. (2020). Electrochemical Voltammogram Recording for Identifying Varieties of Ornamental Plants. Micromachines, 11(11), 967. https://doi.org/10.3390/mi11110967








