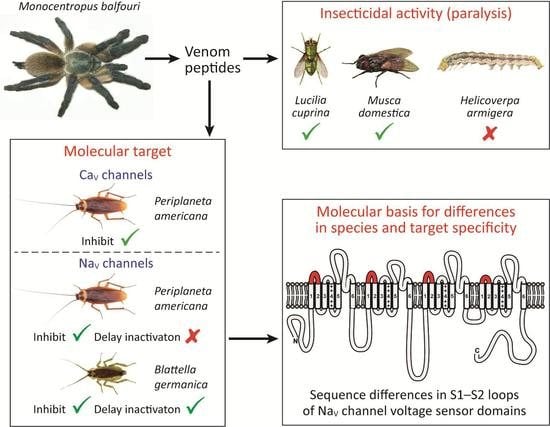
Insect-Active Toxins with Promiscuous Pharmacology from the African Theraphosid Spider Monocentropus balfouri
Abstract
:1. Introduction
2. Results
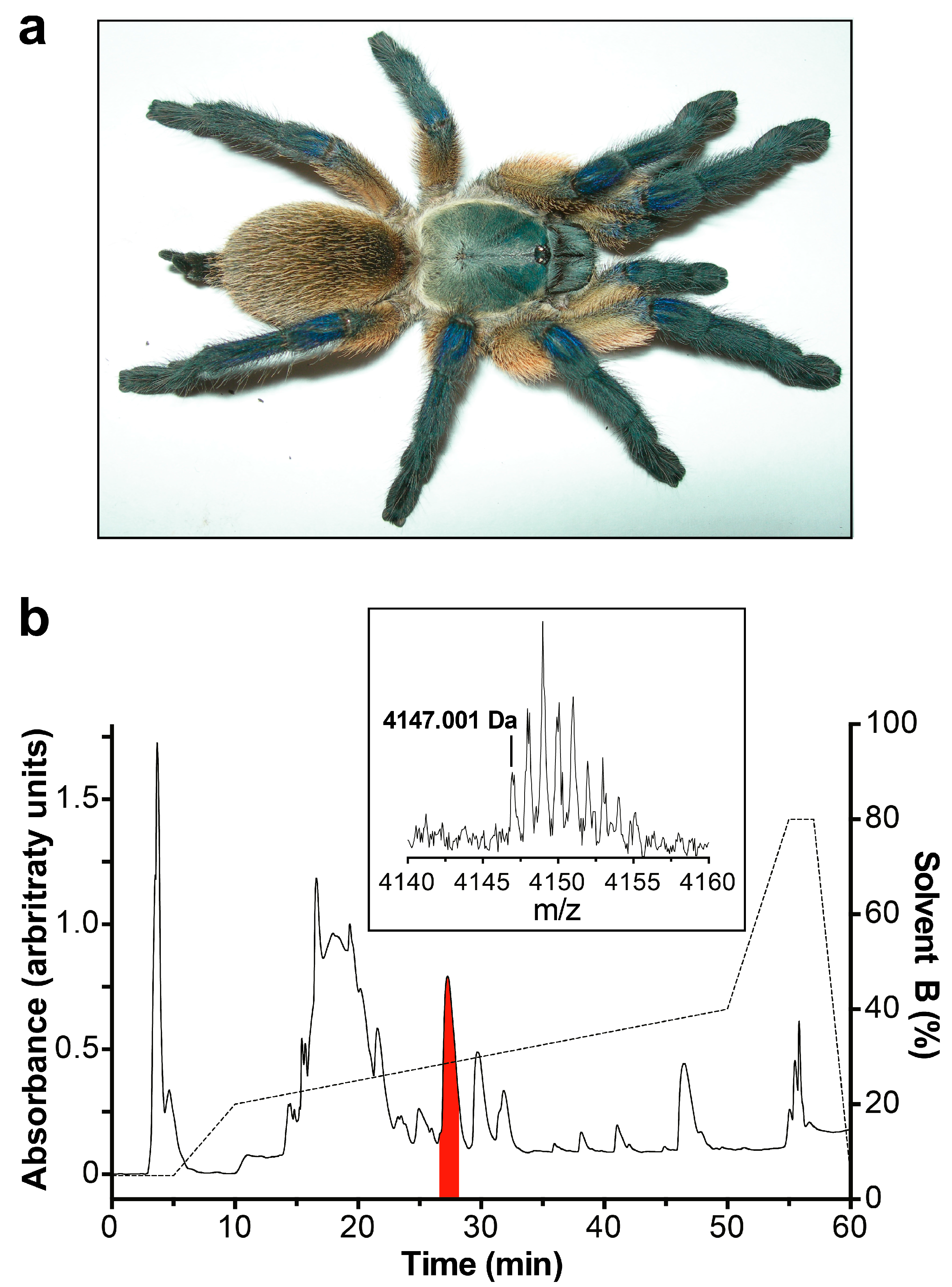
2.1. Isolation and Sequencing of µ/ω-TRTX-Mb1a and -Mb1b
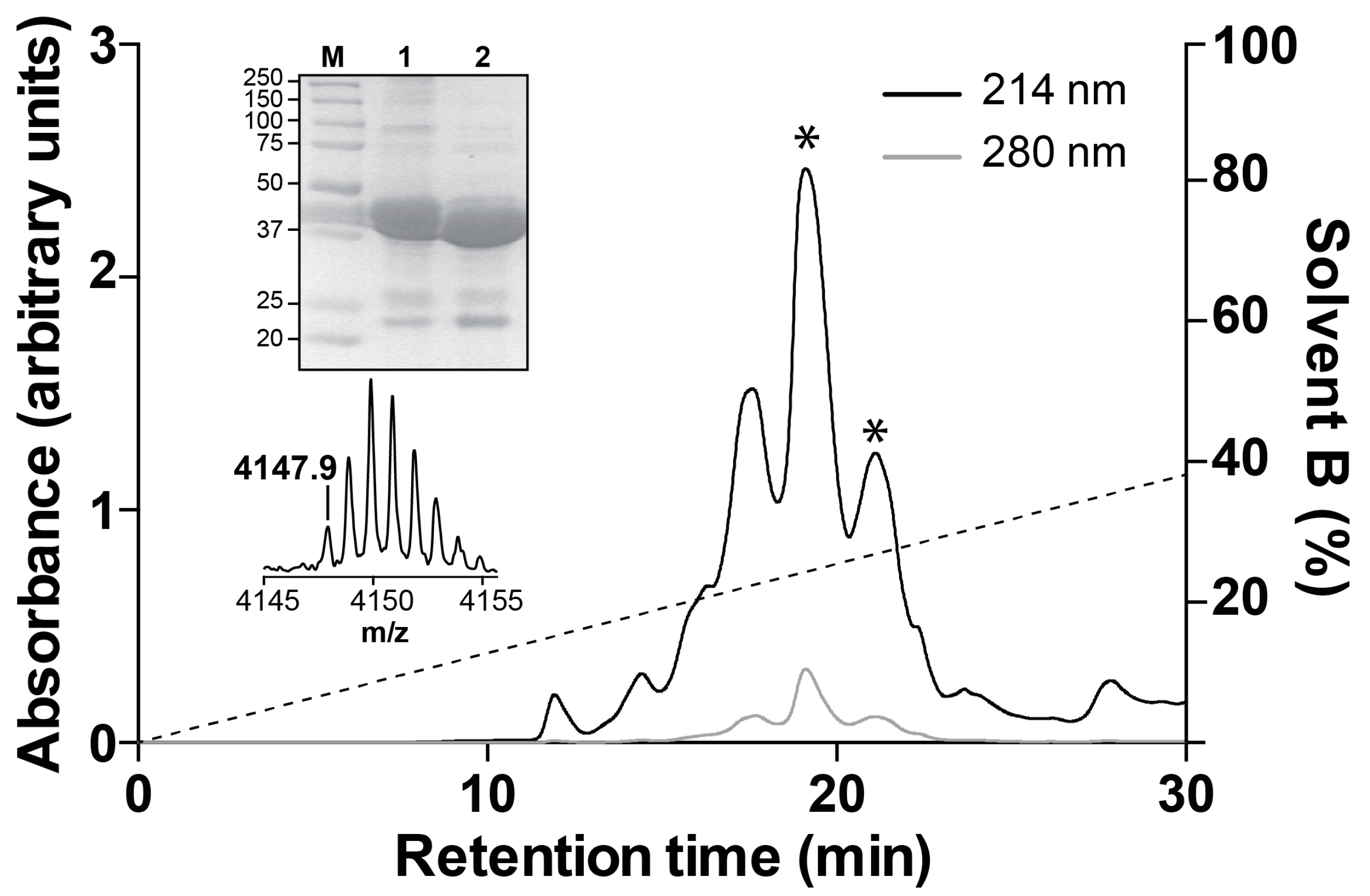
2.2. Recombinant Production of Mb1a and Mb1b
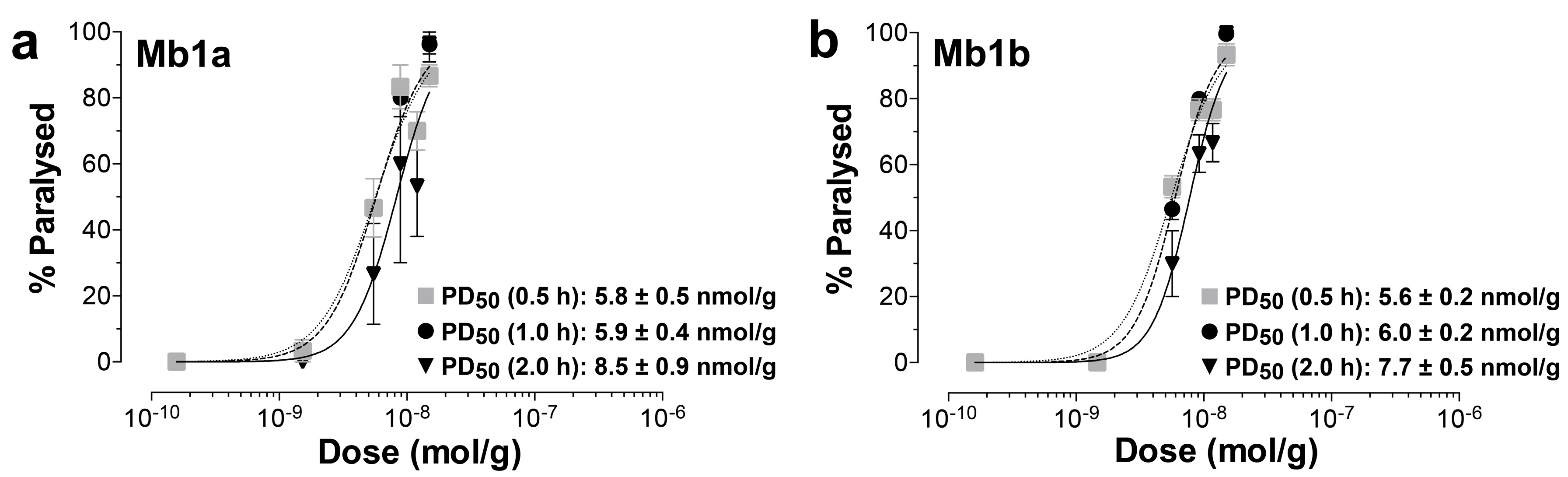
2.3. Insecticidal Activity of µ/ω-TRTX-Mb1a and -Mb1b
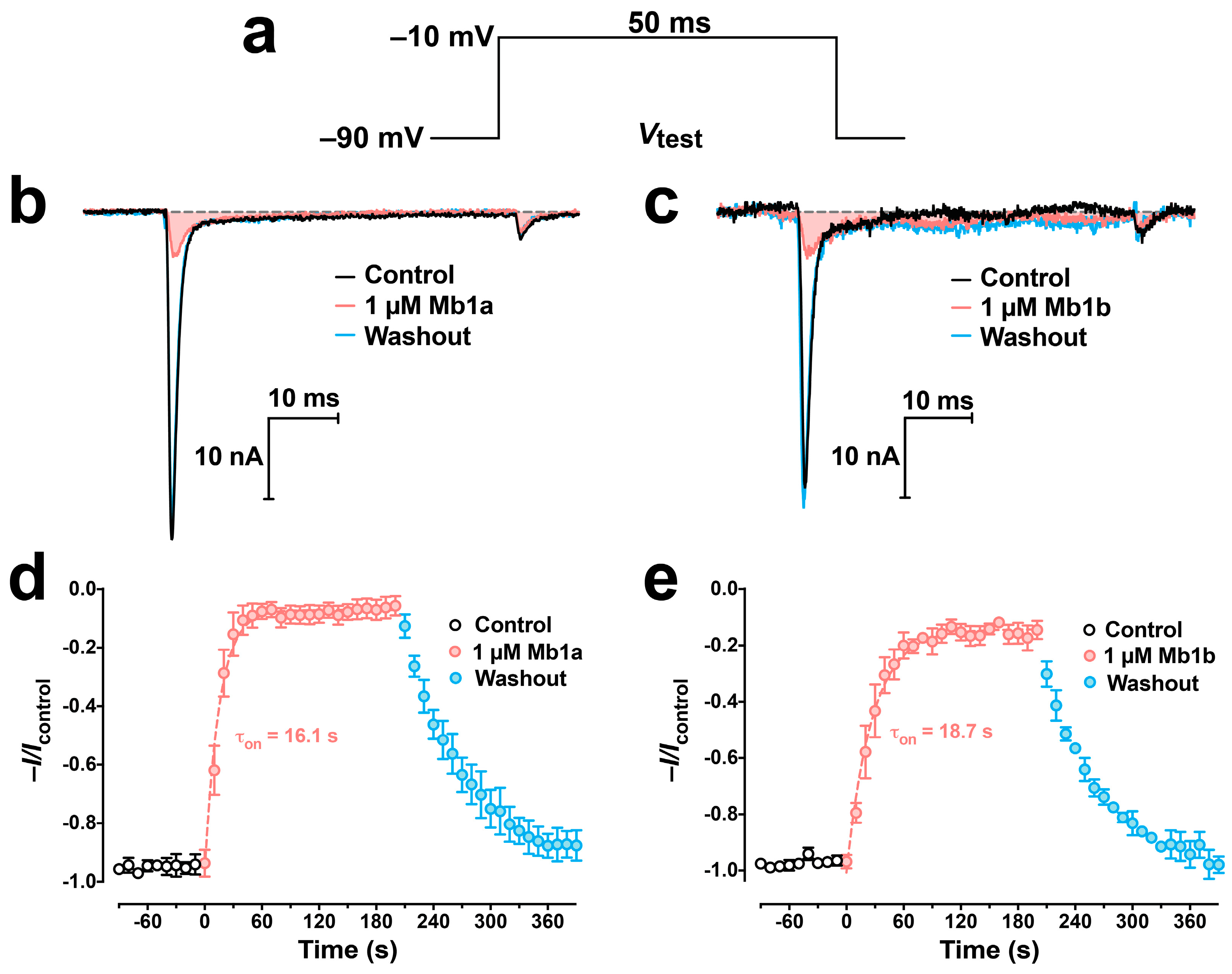
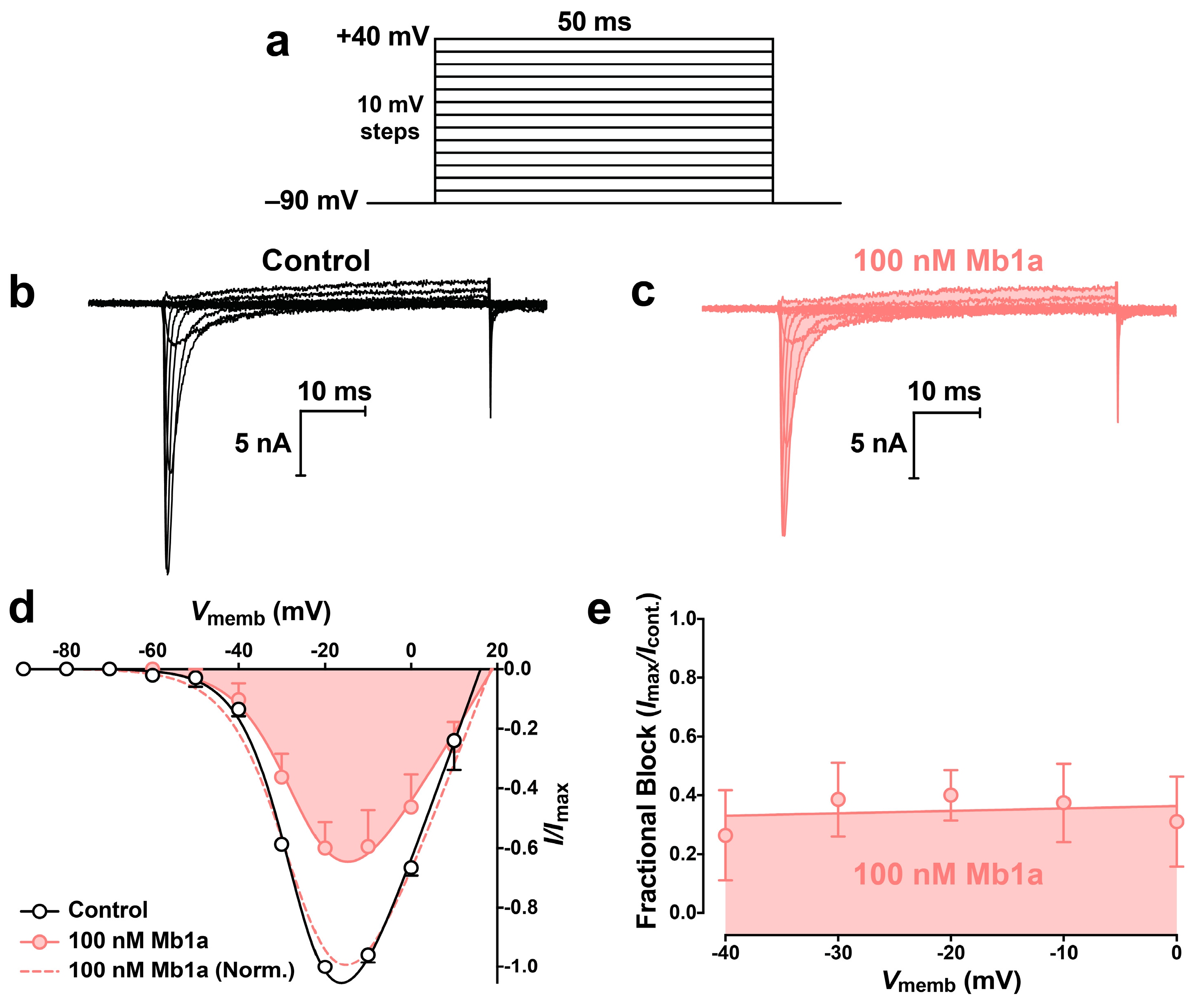
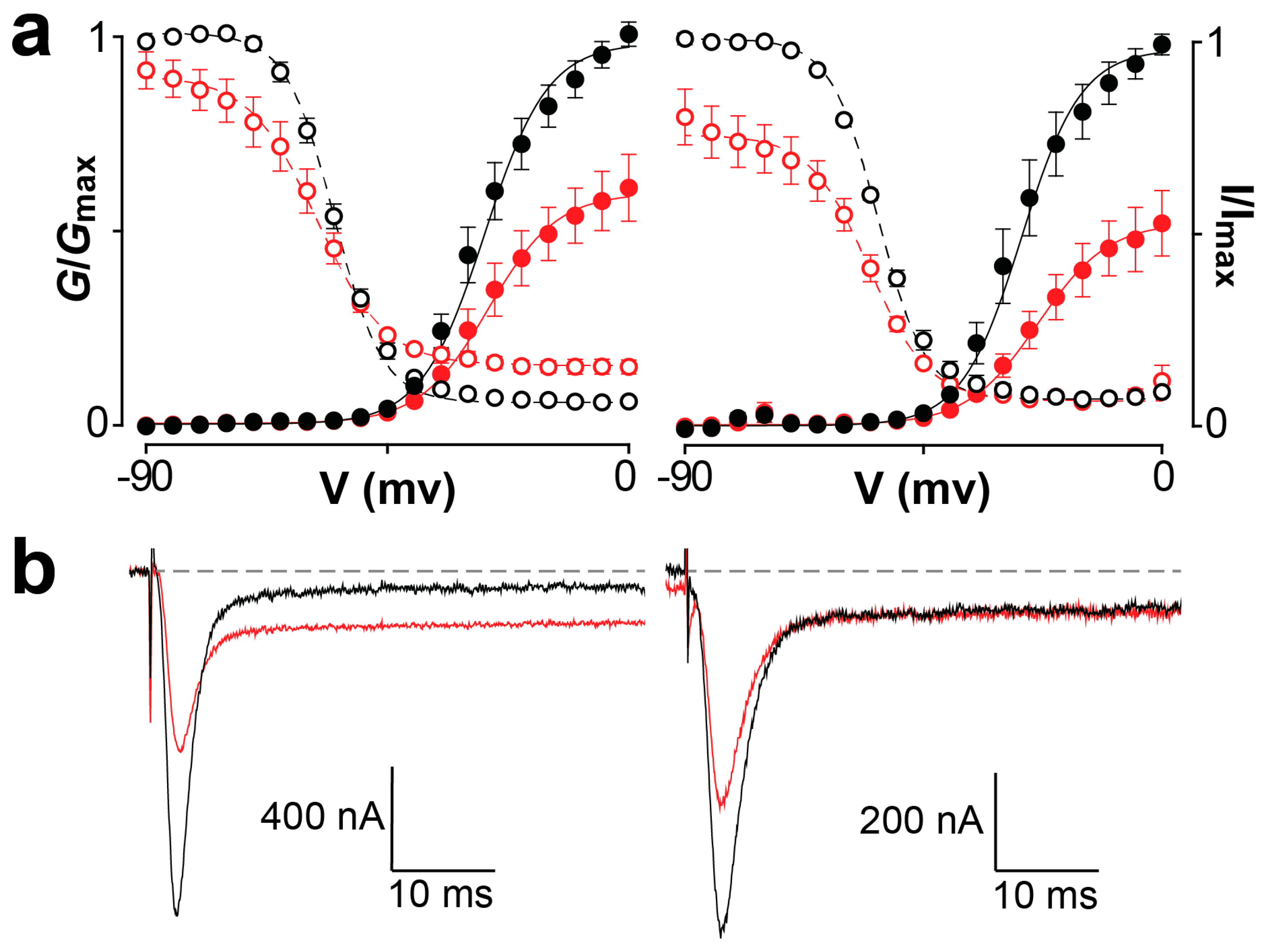
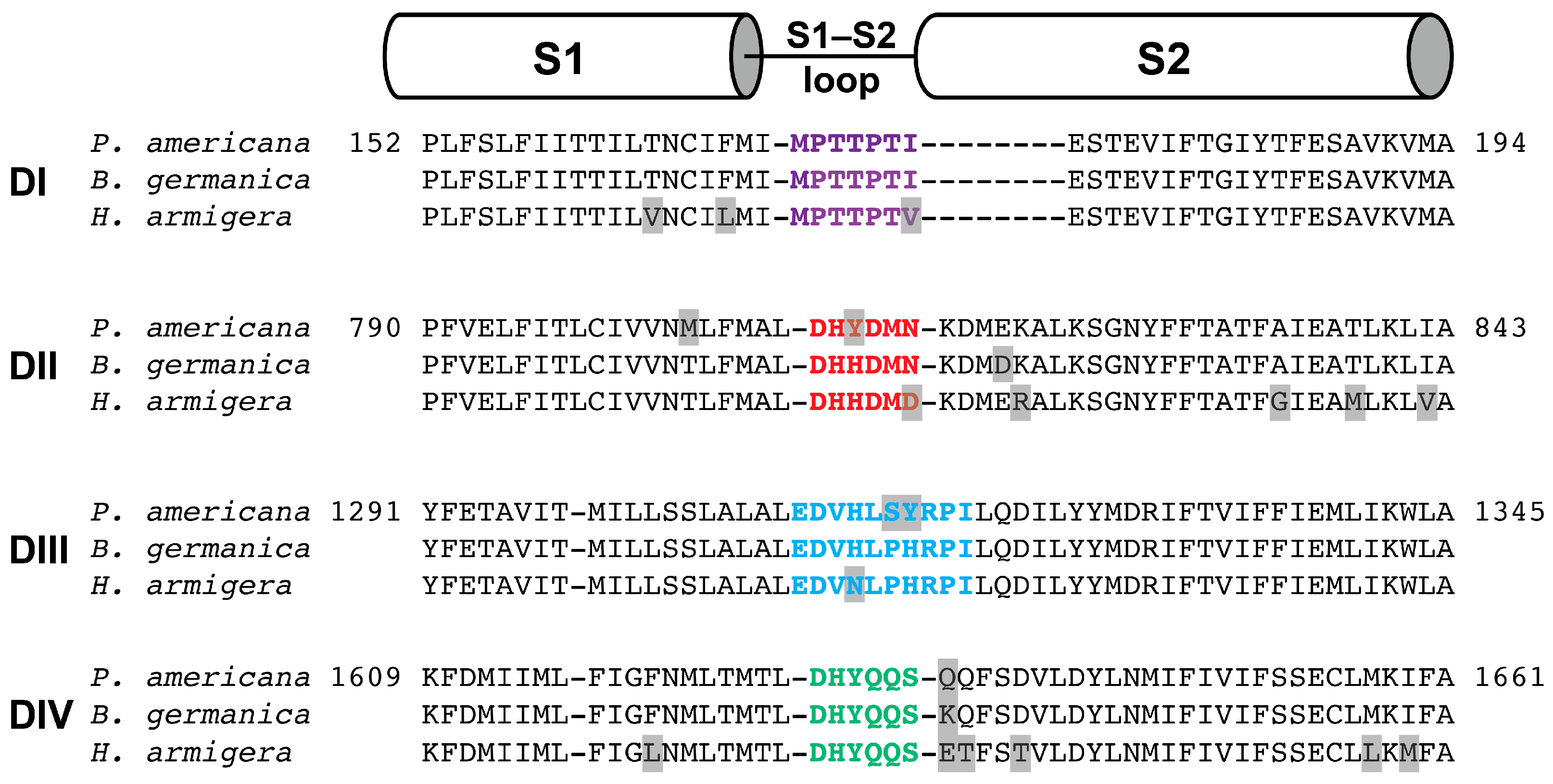
2.4. Activity of Mb1a and Mb1b on Insect NaV Channel Currents
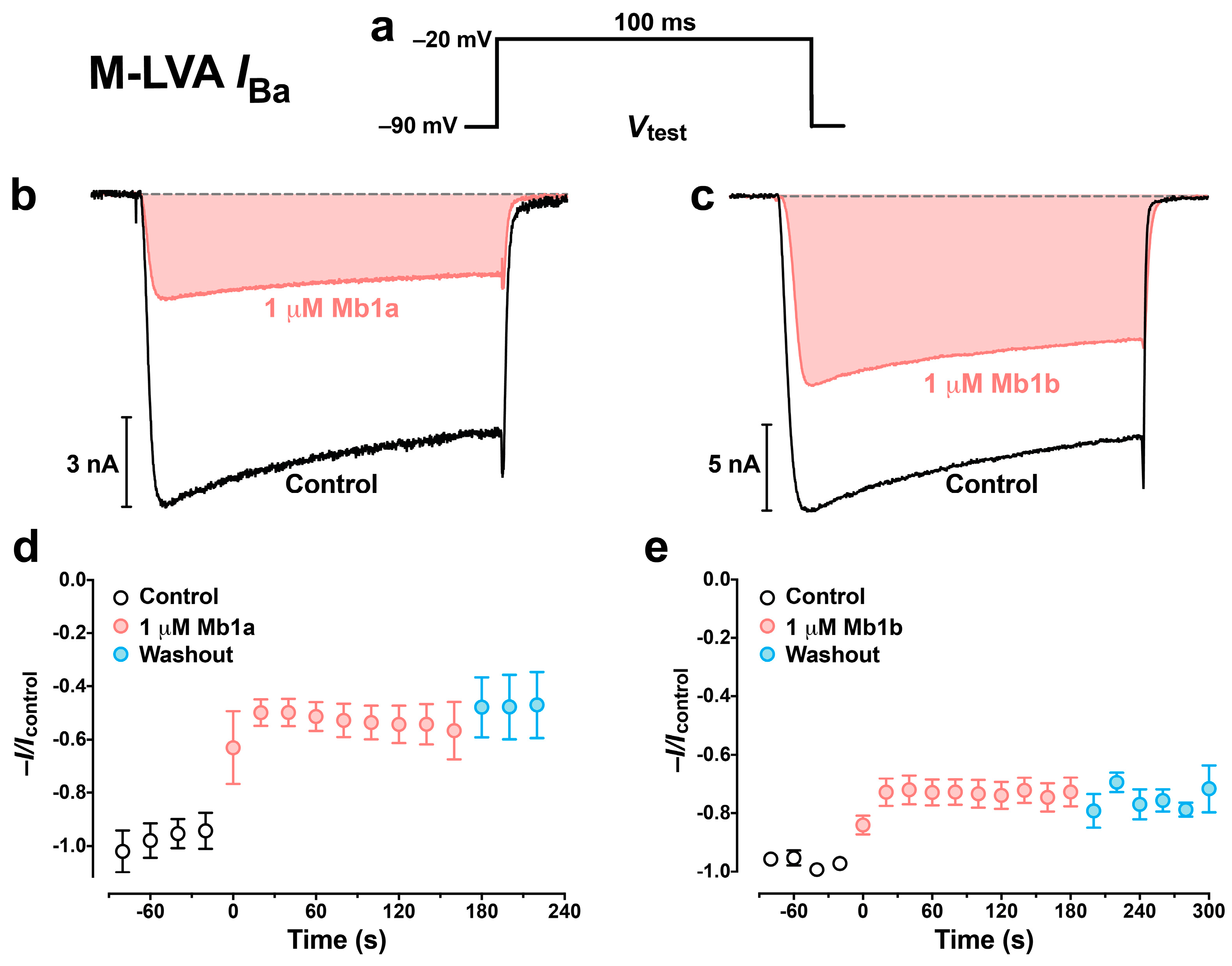
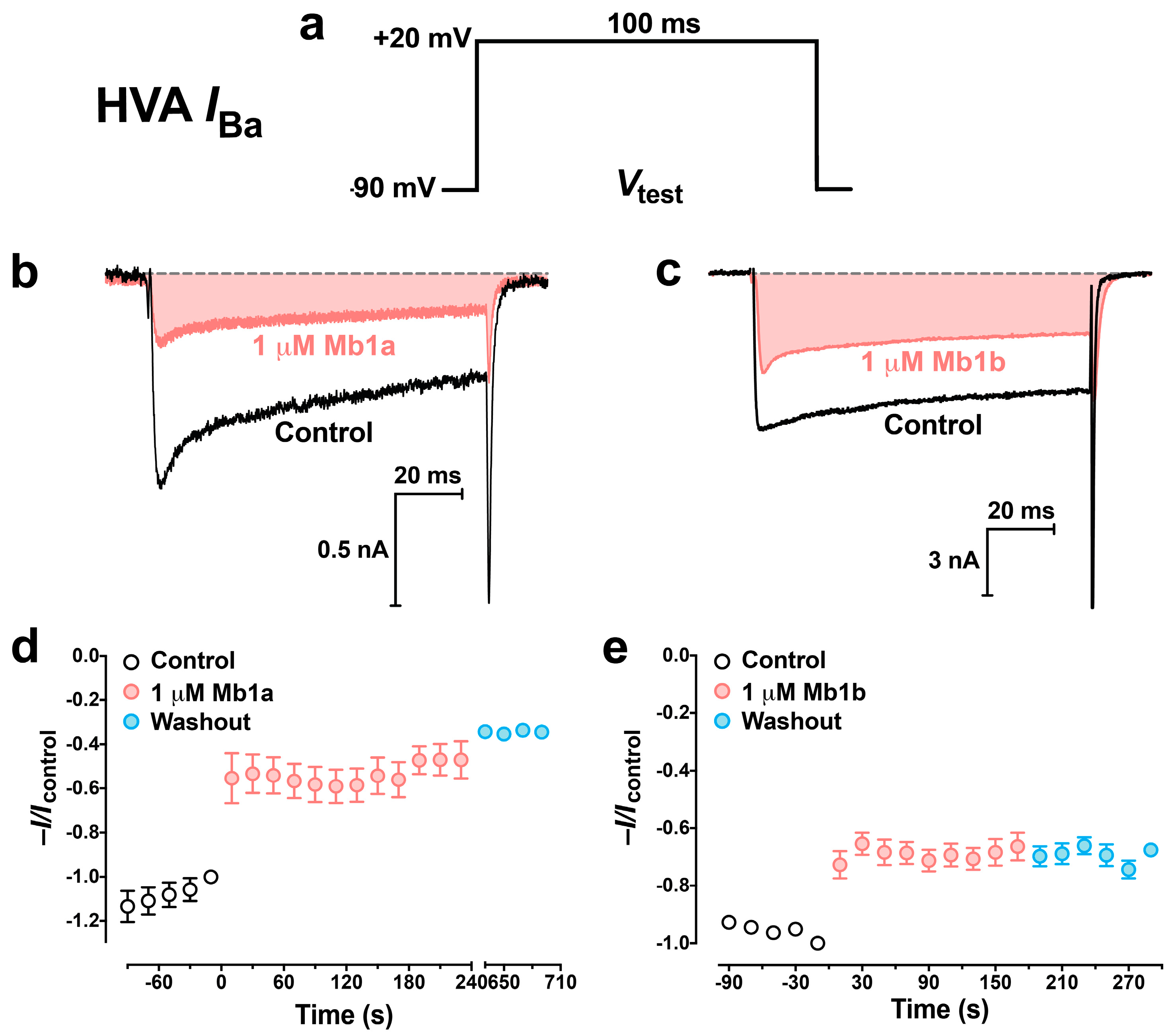
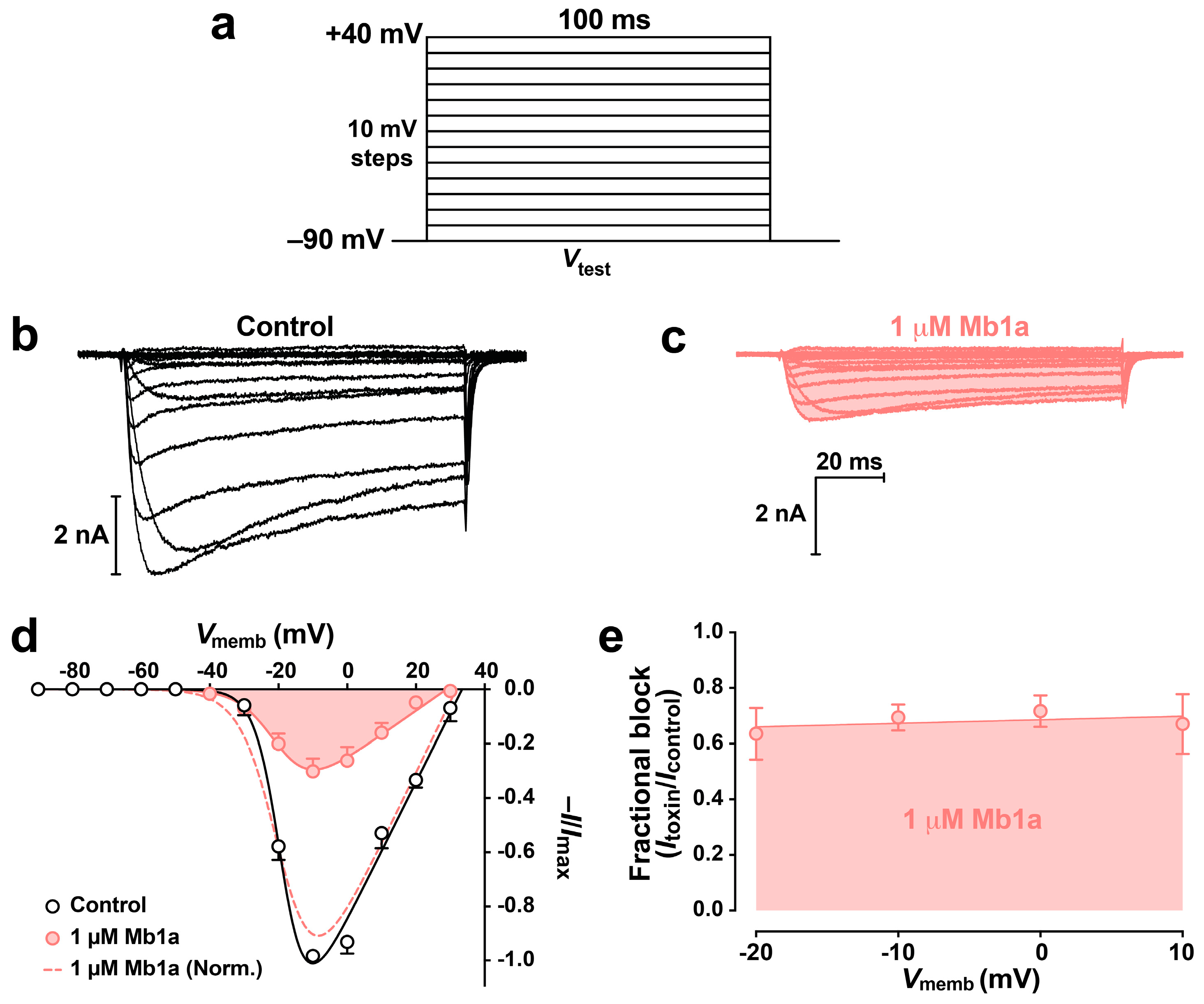
2.5. Effect of Mb1a and Mb1b on Insect CaV Channel Currents
3. Discussion
3.1. Promiscuous Pharmacology of Mb1a/1b
3.2. Phyletic Selectivity of Mb1a/1b
4. Conclusions
5. Materials and Methods
5.1 Venom Collection
5.2. Fractionation of Crude Venom
5.3. Sequencing of Active Peptide
5.4. Recombinant Production of Mb1a and Mb1b
5.5. Insecticidal Assays
5.5.1. Sheep Blowflies and House Flies
5.5.2. Cotton Bollworms
5.6. Patch Clamp Electrophysiology Using P. americana Neurons
5.7. Two–Electrode Voltage-Clamp Electrophysiology
Acknowledgments
Author Contributions
Conflicts of Interest
References
- Saez, N.J.; Senff, S.; Jensen, J.E.; Er, S.Y.; Herzig, V.; Rash, L.D.; King, G.F. Spider-venom peptides as therapeutics. Toxins 2010, 2, 2851–2871. [Google Scholar] [CrossRef] [PubMed]
- Pineda, S.S.; Undheim, E.A.; Rupasinghe, D.B.; Ikonomopoulou, M.P.; King, G.F. Spider venomics: Implications for drug discovery. Future Med. Chem. 2014, 6, 1699–1714. [Google Scholar] [CrossRef] [PubMed]
- Bohlen, C.J.; Julius, D. Receptor-targeting mechanisms of pain-causing toxins: How ow? Toxicon 2012, 60, 254–264. [Google Scholar] [CrossRef] [PubMed]
- Escoubas, P.; Bosmans, F. Spider peptide toxins as leads for drug development. Expert Opin. Drug Discov. 2007, 2, 823–835. [Google Scholar] [CrossRef] [PubMed]
- Escoubas, P.; Rash, L. Tarantulas: Eight-legged pharmacists and combinatorial chemists. Toxicon 2004, 43, 555–574. [Google Scholar] [CrossRef] [PubMed]
- King, G.F.; Hardy, M.C. Spider-venom peptides: Structure, pharmacology, and potential for control of insect pests. Annu. Rev. Entomol. 2013, 58, 475–496. [Google Scholar] [CrossRef] [PubMed]
- Bern, N.H.M. World Spider Catalog. Available online: http://wsc.nmbe.ch (accessed on 3 May 2017).
- Escoubas, P.; Sollod, B.; King, G.F. Venom landscapes: Mining the complexity of spider venoms via a combined cDNA and mass spectrometric approach. Toxicon 2006, 47, 650–663. [Google Scholar] [CrossRef] [PubMed]
- Smith, J.J.; Herzig, V.; King, G.F.; Alewood, P.F. The insecticidal potential of venom peptides. Cell. Mol. Life Sci. 2013, 70, 3665–3693. [Google Scholar] [CrossRef] [PubMed]
- Smith, J.J.; Lau, C.H.Y.; Herzig, V.; Ikonomopoulou, M.P.; Rash, L.D.; King, G.F. Therapeutic applications of spider-venom peptides. In Venoms to Drugs: Venom as a Source for the Development of Human Therapeutics; King, G.F., Ed.; Royal Society of Chemistry: Cambridge, UK, 2015; pp. 221–244. [Google Scholar]
- Klint, J.K.; Senff, S.; Rupasinghe, D.B.; Er, S.Y.; Herzig, V.; Nicholson, G.M.; King, G.F. Spider-venom peptides that target voltage-gated sodium channels: Pharmacological tools and potential therapeutic leads. Toxicon 2012, 60, 478–491. [Google Scholar] [CrossRef] [PubMed]
- Ahern, C.A.; Payandeh, J.; Bosmans, F.; Chanda, B. The hitchhiker’s guide to the voltage-gated sodium channel galaxy. J. Gen. Physiol. 2016, 147, 1–24. [Google Scholar] [CrossRef] [PubMed]
- Davies, T.G.; Field, L.M.; Usherwood, P.N.; Williamson, M.S. DDT, pyrethrins, pyrethroids and insect sodium channels. IUBMB Life 2007, 59, 151–162. [Google Scholar] [CrossRef] [PubMed]
- Du, Y.; Nomura, Y.; Satar, G.; Hu, Z.; Nauen, R.; He, S.Y.; Zhorov, B.S.; Dong, K. Molecular evidence for dual pyrethroid-receptor sites on a mosquito sodium channel. Proc. Natl. Acad. Sci. USA 2013, 110, 11785–11790. [Google Scholar] [CrossRef] [PubMed]
- Du, Y.; Nomura, Y.; Zhorov, B.S.; Dong, K. Sodium channel mutations and pyrethroid resistance in Aedes aegypti. Insects 2016, 7, 60. [Google Scholar] [CrossRef] [PubMed]
- Zhang, Y.; Du, Y.; Jiang, D.; Behnke, C.; Nomura, Y.; Zhorov, B.S.; Dong, K. The receptor site and mechanism of action of sodium channel blocker insecticides. J. Biol. Chem. 2016, 291, 20113–20124. [Google Scholar] [CrossRef] [PubMed]
- Gilchrist, J.; Olivera, B.M.; Bosmans, F. Animal toxins influence voltage-gated sodium channel function. Handb. Exp. Pharmacol. 2014, 221, 203–229. [Google Scholar] [PubMed]
- King, G.F. Modulation of insect CaV channels by peptidic spider toxins. Toxicon 2007, 49, 513–530. [Google Scholar] [CrossRef] [PubMed]
- Bomgardner, M. Spider venom: An insecticide whose time has come? Chem. Eng. News 2017, 95, 30–31. [Google Scholar]
- King, G.F.; Gentz, M.C.; Escoubas, P.; Nicholson, G.M. A rational nomenclature for naming peptide toxins from spiders and other venomous animals. Toxicon 2008, 52, 264–276. [Google Scholar] [CrossRef] [PubMed]
- Klint, J.K.; Senff, S.; Saez, N.J.; Seshadri, R.; Lau, H.Y.; Bende, N.S.; Undheim, E.A.; Rash, L.D.; Mobli, M.; King, G.F. Production of recombinant disulfide-rich venom peptides for structural and functional analysis via expression in the periplasm of E. coli. PLoS ONE 2013, 8. [Google Scholar] [CrossRef] [PubMed]
- Liu, Z.; Song, W.; Dong, K. Persistent tetrodotoxin-sensitive sodium current resulting from U-to-C RNA editing of an insect sodium channel. Proc. Natl. Acad. Sci. USA 2004, 101, 11862–11867. [Google Scholar] [CrossRef] [PubMed]
- Bende, N.S.; Kang, E.; Herzig, V.; Bosmans, F.; Nicholson, G.M.; Mobli, M.; King, G.F. The insecticidal neurotoxin Aps III is an atypical knottin peptide that potently blocks insect voltage-gated sodium channels. Biochem. Pharmacol. 2013, 85, 1542–1554. [Google Scholar] [CrossRef] [PubMed]
- Bende, N.S.; Dziemborowicz, S.; Mobli, M.; Herzig, V.; Gilchrist, J.; Wagner, J.; Nicholson, G.M.; King, G.F.; Bosmans, F. A distinct sodium channel voltage-sensor locus determines insect selectivity of the spider toxin Dc1a. Nat. Commun. 2014, 5, 4350. [Google Scholar] [CrossRef] [PubMed]
- Newcomb, R.; Szoke, B.; Palma, A.; Wang, G.; Chen, X.; Hopkins, W.; Cong, R.; Miller, J.; Urge, L.; Tarczy-Hornoch, K.; et al. Selective peptide antagonist of the class E calcium channel from the venom of the tarantula Hysterocrates gigas. Biochemistry 1998, 37, 15353–15362. [Google Scholar] [CrossRef] [PubMed]
- Bourinet, E.; Stotz, S.C.; Spaetgens, R.L.; Dayanithi, G.; Lemos, J.; Nargeot, J.; Zamponi, G.W. Interaction of SNX482 with domains III and IV inhibits activation gating of α1E (CaV2.3) calcium channels. Biophys. J. 2001, 81, 79–88. [Google Scholar] [CrossRef]
- Bosmans, F.; Rash, L.; Zhu, S.; Diochot, S.; Lazdunski, M.; Escoubas, P.; Tytgat, J. Four novel tarantula toxins as selective modulators of voltage-gated sodium channel subtypes. Mol. Pharmacol. 2006, 69, 419–429. [Google Scholar] [CrossRef] [PubMed]
- Klint, J.K.; Berecki, G.; Durek, T.; Mobli, M.; Knapp, O.; King, G.F.; Adams, D.J.; Alewood, P.F.; Rash, L.D. Isolation, synthesis and characterization of ω-TRTX-Cc1a, a novel tarantula venom peptide that selectively targets L-type Cav channels. Biochem. Pharmacol. 2014, 89, 276–286. [Google Scholar] [CrossRef] [PubMed]
- Arroyo, G.; Aldea, M.; Fuentealba, J.; Albillos, A.; Garcia, A.G. SNX482 selectively blocks P/Q Ca2+ channels and delays the inactivation of Na+ channels of chromaffin cells. Eur. J. Pharmacol. 2003, 475, 11–18. [Google Scholar] [CrossRef]
- Paiva, A.L.; Matavel, A.; Peigneur, S.; Cordeiro, M.N.; Tytgat, J.; Diniz, M.R.; de Lima, M.E. Differential effects of the recombinant toxin PnTx4(5–5) from the spider Phoneutria nigriventer on mammalian and insect sodium channels. Biochimie 2016, 121, 326–335. [Google Scholar] [CrossRef] [PubMed]
- Bosmans, F.; Martin-Eauclaire, M.F.; Swartz, K.J. Deconstructing voltage sensor function and pharmacology in sodium channels. Nature 2008, 456, 202–208. [Google Scholar] [CrossRef] [PubMed]
- Capes, D.L.; Goldschen-Ohm, M.P.; Arcisio-Miranda, M.; Bezanilla, F.; Chanda, B. Domain IV voltage-sensor movement is both sufficient and rate limiting for fast inactivation in sodium channels. J. Gen. Physiol. 2013, 142, 101–112. [Google Scholar] [CrossRef] [PubMed]
- Zhang, J.Z.; Yarov-Yarovoy, V.; Scheuer, T.; Karbat, I.; Cohen, L.; Gordon, D.; Gurevitz, M.; Catterall, W.A. Structure-function map of the receptor site for β-scorpion toxins in domain II of voltage-gated sodium channels. J. Biol. Chem. 2011, 286, 33641–33651. [Google Scholar] [CrossRef] [PubMed]
- Xiao, Y.; Blumenthal, K.; Jackson, J.O.; Liang, S.; Cummins, T.R. The tarantula toxins ProTx-II and huwentoxin-IV differentially interact with human Nav1.7 voltage sensors to inhibit channel activation and inactivation. Mol. Pharmacol. 2010, 78, 1124–1134. [Google Scholar] [CrossRef] [PubMed]
- Xiao, Y.; Jackson, J.O.; Liang, S.; Cummins, T.R. Common molecular determinants of tarantula huwentoxin-IV inhibition of Na+ channel voltage sensors in domains II and IV. J. Biol. Chem. 2011, 286, 27301–27310. [Google Scholar] [CrossRef] [PubMed]
- Osteen, J.D.; Herzig, V.; Gilchrist, J.; Emrick, J.J.; Zhang, C.; Wang, X.; Castro, J.; Garcia-Caraballo, S.; Grundy, L.; Rychkov, G.Y.; et al. Selective spider toxins reveal a role for the NaV1.1 channel in mechanical pain. Nature 2016, 534, 494–499. [Google Scholar] [CrossRef] [PubMed]
- Herzig, V.; Ikonomopoulou, M.; Smith, J.J.; Dziemborowicz, S.; Gilchrist, J.; Kuhn-Nentwig, L.; Rezende, F.O.; Moreira, L.A.; Nicholson, G.M.; Bosmans, F.; et al. Molecular basis of the remarkable species selectivity of an insecticidal sodium channel toxin from the African spider Augacephalus ezendami. Sci. Rep. 2016, 6. [Google Scholar] [CrossRef] [PubMed]
- Lau, C.H.; King, G.F.; Mobli, M. Molecular basis of the interaction between gating modifier spider toxins and the voltage sensor of voltage-gated ion channels. Sci. Rep. 2016, 6. [Google Scholar] [CrossRef] [PubMed]
- Shen, H.; Zhou, Q.; Pan, X.; Li, Z.; Wu, J.; Yan, N. Structure of a eukaryotic voltage-gated sodium channel at near-atomic resolution. Science 2017, 355. [Google Scholar] [CrossRef] [PubMed]
- Shon, K.J.; Olivera, B.M.; Watkins, M.; Jacobsen, R.B.; Gray, W.R.; Floresca, C.Z.; Cruz, L.J.; Hillyard, D.R.; Brink, A.; Terlau, H.; et al. μ-Conotoxin PIIIA, a new peptide for discriminating among tetrodotoxin-sensitive Na channel subtypes. J. Neurosci. 1998, 18, 4473–4481. [Google Scholar] [PubMed]
- Xiao, Y.; Bingham, J.P.; Zhu, W.; Moczydlowski, E.; Liang, S.; Cummins, T.R. Tarantula huwentoxin-IV inhibits neuronal sodium channels by binding to receptor site 4 and trapping the domain ii voltage sensor in the closed configuration. J. Biol. Chem. 2008, 283, 27300–27313. [Google Scholar] [CrossRef] [PubMed]
- Klint, J.K.; Smith, J.J.; Vetter, I.; Rupasinghe, D.B.; Er, S.Y.; Senff, S.; Herzig, V.; Mobli, M.; Lewis, R.J.; Bosmans, F.; et al. Seven novel modulators of the analgesic target NaV1.7 uncovered using a high-throughput venom-based discovery approach. Br. J. Pharmacol. 2015, 172, 2445–2458. [Google Scholar] [CrossRef] [PubMed]
- Salari, A.; Vega, B.S.; Milescu, L.S.; Milescu, M. Molecular interactions between tarantula toxins and low-voltage-activated calcium channels. Sci. Rep. 2016, 6, 23894. [Google Scholar] [CrossRef] [PubMed]
- Bourinet, E.; Zamponi, G.W. Block of voltage-gated calcium channels by peptide toxins. Neuropharmacology 2016, in press. [Google Scholar] [CrossRef] [PubMed]
- Mintz, I.M. Block of Ca channels in rat central neurons by the spider toxin ω-Aga-IIIA. J. Neurosci. 1994, 14, 2844–2853. [Google Scholar] [PubMed]
- Newcomb, R.; Palma, A.; Fox, J.; Gaur, S.; Lau, K.; Chung, D.; Cong, R.; Bell, J.R.; Horne, B.; Nadasdi, L.; et al. SNX-325, a novel calcium antagonist from the spider Segestria florentina. Biochemistry 1995, 34, 8341–8347. [Google Scholar] [CrossRef] [PubMed]
- Adams, M.E.; Herold, E.E.; Venema, V.J. Two classes of channel-specific toxins from funnel web spider venom. J. Comp. Physiol. A 1989, 164, 333–342. [Google Scholar] [CrossRef] [PubMed]
- Bende, N.S.; Dziemborowicz, S.; Herzig, V.; Ramanujam, V.; Brown, G.W.; Bosmans, F.; Nicholson, G.M.; King, G.F.; Mobli, M. The insecticidal spider toxin SFI1 is a knottin peptide that blocks the pore of insect voltage-gated sodium channels via a large β-hairpin loop. FEBS J. 2015, 282, 904–920. [Google Scholar] [CrossRef] [PubMed]
- Mukherjee, A.K.; Sollod, B.L.; Wikel, S.K.; King, G.F. Orally active acaricidal peptide toxins from spider venom. Toxicon 2006, 47, 182–187. [Google Scholar] [CrossRef] [PubMed]
- Undheim, E.A.; Grimm, L.L.; Low, C.F.; Morgenstern, D.; Herzig, V.; Zobel-Thropp, P.; Pineda, S.S.; Habib, R.; Dziemborowicz, S.; Fry, B.G.; et al. Weaponization of a hormone: Convergent recruitment of hyperglycemic hormone into the venom of arthropod predators. Structure 2015, 23, 1283–1292. [Google Scholar] [CrossRef] [PubMed]
- Sahara, K.; Yoshido, A.; Shibata, F.; Fujikawa-Kojima, N.; Okabe, T.; Tanaka-Okuyama, M.; Yasukochi, Y. FISH identification of Helicoverpa armigera and Mamestra brassicae chromosomes by BAC and fosmid probes. Insect Biochem. Mol. Biol. 2013, 43, 644–653. [Google Scholar] [CrossRef] [PubMed]
- BioInformatics Platform for Agro-Ecosystems Arthropods. Available online: http://bipaa.genouest.org/is/lepidodb/ (accessed on 1 March 2017).
- Herzig, V.; Hodgson, W.C. Intersexual variations in the pharmacological properties of Coremiocnemis tropix (Araneae, Theraphosidae) spider venom. Toxicon 2009, 53, 196–205. [Google Scholar] [CrossRef] [PubMed]
- Eitan, M.; Fowler, E.; Herrmann, R.; Duval, A.; Pelhate, M.; Zlotkin, E. A scorpion venom neurotoxin paralytic to insects that affects sodium current inactivation: Purification, primary structure, and mode of action. Biochemistry 1990, 29, 5941–5947. [Google Scholar] [CrossRef] [PubMed]
- Herzig, V.; Hodgson, W.C. Neurotoxic and insecticidal properties of venom from the Australian theraphosid spider Selenotholus foelschei. Neurotoxicology 2008, 29, 471–475. [Google Scholar] [CrossRef] [PubMed]
- Gunning, S.J.; Maggio, F.; Windley, M.J.; Valenzuela, S.M.; King, G.F.; Nicholson, G.M. The Janus-faced atracotoxins are specific blockers of invertebrate KCa channels. FEBS J. 2008, 275, 4045–4059. [Google Scholar] [CrossRef] [PubMed]
- Chong, Y.; Hayes, J.L.; Sollod, B.; Wen, S.; Wilson, D.T.; Hains, P.G.; Hodgson, W.C.; Broady, K.W.; King, G.F.; Nicholson, G.M. The ω-atracotoxins: Selective blockers of insect M-LVA and HVA calcium channels. Biochem. Pharmacol. 2007, 74, 623–638. [Google Scholar] [CrossRef] [PubMed]
- Wicher, D.; Penzlin, H. Ca2+ currents in central insect neurons: Electrophysiological and pharmacological properties. J. Neurophysiol. 1997, 77, 186–199. [Google Scholar] [PubMed]
- Dong, K. A single amino acid change in the para sodium channel protein is associated with knockdown-resistance (kdr) to pyrethroid insecticides in German cockroach. Insect Biochem. Mol. Biol. 1997, 27, 93–100. [Google Scholar] [CrossRef]
- Feng, G.; Deak, P.; Chopra, M.; Hall, L.M. Cloning and functional analysis of TipE, a novel membrane protein that enhances Drosophila para sodium channel function. Cell 1995, 82, 1001–1011. [Google Scholar] [CrossRef]
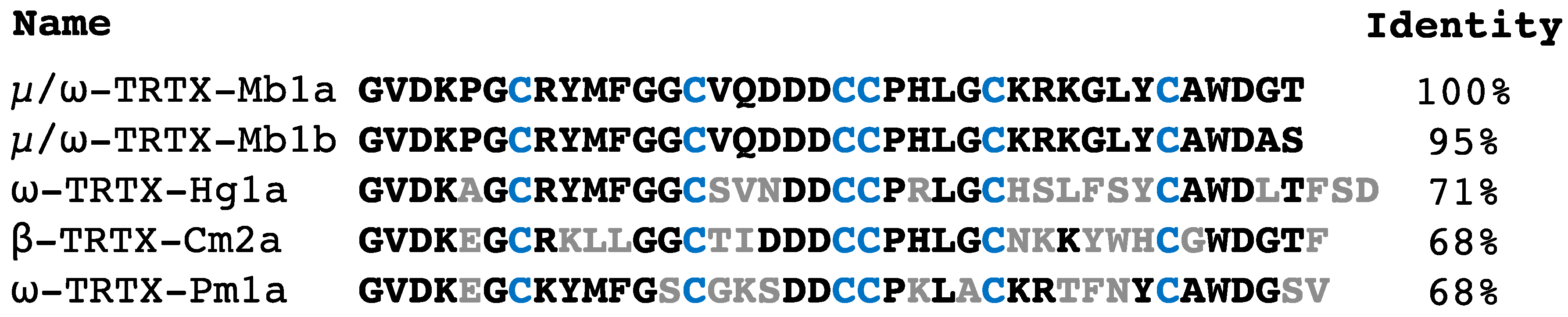
© 2017 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Smith, J.J.; Herzig, V.; Ikonomopoulou, M.P.; Dziemborowicz, S.; Bosmans, F.; Nicholson, G.M.; King, G.F. Insect-Active Toxins with Promiscuous Pharmacology from the African Theraphosid Spider Monocentropus balfouri. Toxins 2017, 9, 155. https://doi.org/10.3390/toxins9050155
Smith JJ, Herzig V, Ikonomopoulou MP, Dziemborowicz S, Bosmans F, Nicholson GM, King GF. Insect-Active Toxins with Promiscuous Pharmacology from the African Theraphosid Spider Monocentropus balfouri. Toxins. 2017; 9(5):155. https://doi.org/10.3390/toxins9050155
Chicago/Turabian StyleSmith, Jennifer J., Volker Herzig, Maria P. Ikonomopoulou, Sławomir Dziemborowicz, Frank Bosmans, Graham M. Nicholson, and Glenn F. King. 2017. "Insect-Active Toxins with Promiscuous Pharmacology from the African Theraphosid Spider Monocentropus balfouri" Toxins 9, no. 5: 155. https://doi.org/10.3390/toxins9050155
APA StyleSmith, J. J., Herzig, V., Ikonomopoulou, M. P., Dziemborowicz, S., Bosmans, F., Nicholson, G. M., & King, G. F. (2017). Insect-Active Toxins with Promiscuous Pharmacology from the African Theraphosid Spider Monocentropus balfouri. Toxins, 9(5), 155. https://doi.org/10.3390/toxins9050155