Strategies and Methodologies for Developing Microbial Detoxification Systems to Mitigate Mycotoxins
Abstract
:1. Introduction
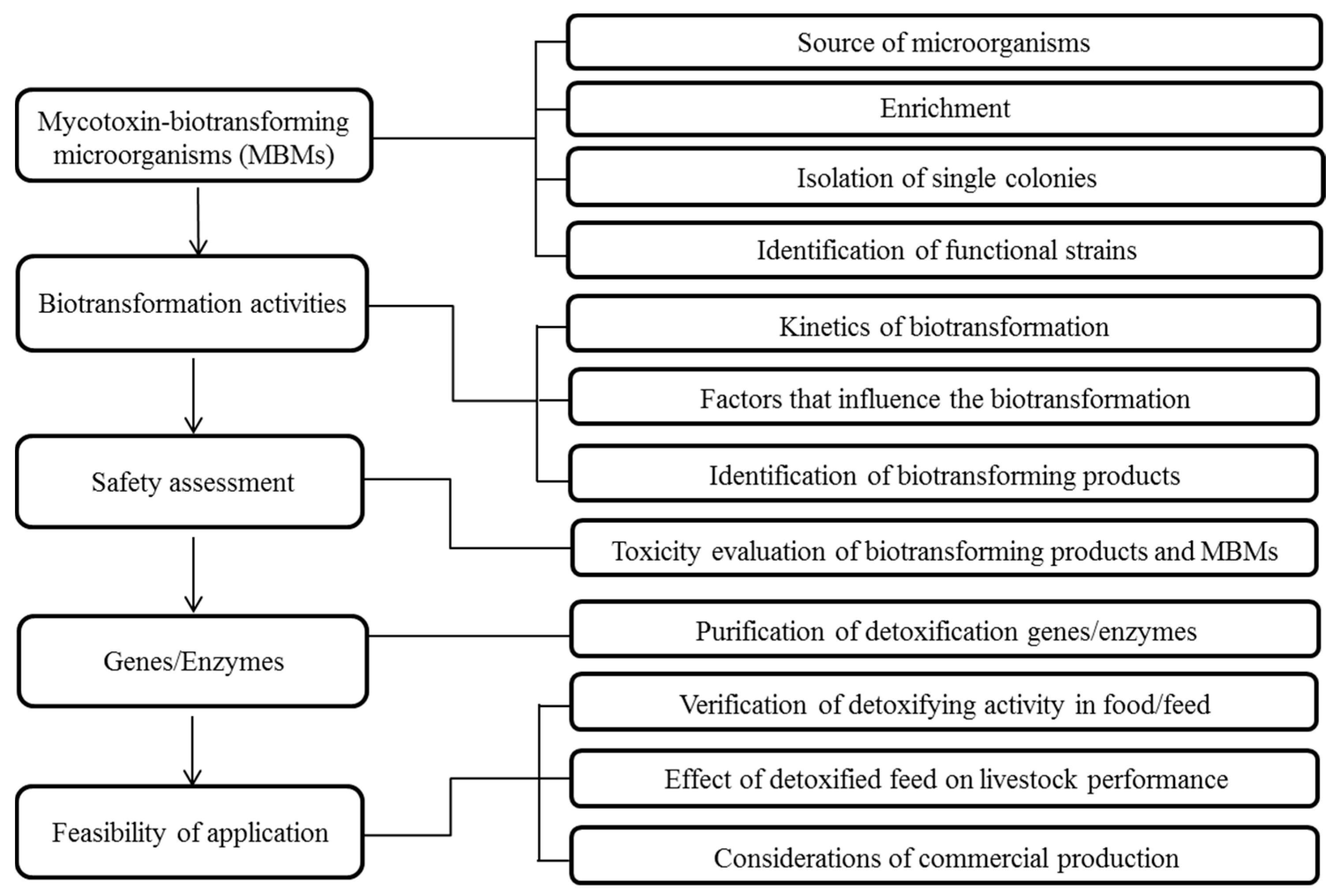
2. Workflow
3. Strategies and Methodologies
3.1. Sources of Microorganisms
3.2. Enrichment
3.3. Isolation of Single Colonies
3.4. Identification of Functional Microorganisms
3.5. Physiological Characterization of Mycotoxin Biotransformation Activity
3.6. Identification of Mycotoxins and Their Biotransformation Products
3.7. Evaluation of Toxicity of Biotransforming Product
3.8. Genes and Enzymes
3.9. Feasibility of Commercial Application
4. Conclusions and Research Trends
Acknowledgments
Conflicts of Interest
Abbreviations
| AFB1 | aflatoxin B1 |
| AFB2 | aflatoxin B2 |
| AFG1 | aflatoxin G1 |
| AFM1 | aflatoxin M1 |
| CIT | citrinin |
| DON | deoxynivalenol |
| DOM-1 | deepoxy-deoxynivalenol |
| DPA | desoxypatulinic acid |
| FUM | fumonisin |
| FUB1 | fumonisin B1 |
| NIV | nivalenol |
| OTA | ochratoxin A |
| OTα | ochratoxin α |
| PAT | patulin |
| ZEA | zearalenone |
| ZAL | zearalanol |
| ZOL | zearalenol |
| ZAN | zearalenone |
| ZOM-1 | ((5S)-5-({2,4-dihydroxy-6-[(1E)-5-hydroxypent-1-en-1-yl]benzoyl}oxy)hexanoic acid) |
References
- Bryden, W.L. Mycotoxin contamination of the feed supply chain: Implications for animal productivity and feed security. Anim. Feed Sci. Technol. 2012, 173, 134–158. [Google Scholar] [CrossRef]
- Vardon, P.J.; McLaughlin, C.; Nardinelli, C. Potential economic costs of mycotoxins in the United States. In Mycotoxins: Risks in Plant, Animal, and Human Systems; Task Force Report No. 139; Council for Agricultural Science and Technology (CAST): Ames, IA, USA, 2003. [Google Scholar]
- Faneli, F.; Logrieko, F. Main mycotoxin concerns in Europe: MYCORED and ISM efforts to harmonize strategies for their reduction in food and feed chain. Proc. Nat. Sci. Matica Srpska Novi Sad 2012, 122, 7–16. [Google Scholar] [CrossRef]
- Peraica, M.; Radić, B.; Lucić, A.; Pavlović, M. Toxic effects of mycotoxins in humans. Bull. World Health Organ. 1999, 77, 754–766. [Google Scholar] [PubMed]
- European Commission. RASFF—The Rapid Alert System for Food and Feed; 2015 Annual Report; European Commission: Brussels, Belgium, 2016. [Google Scholar]
- European Commission (EC). Commission Regulation (EC) No. 583/2006 of 17 August 2006. Commission recommendation on the prevention and reduction of Fusarium toxins in cereals and cereal products. Off. J. Eur. Union 2006, L234, 35–40. [Google Scholar]
- He, J.; Zhou, T.; Young, J.C.; Boland, G.J.; Scott, P.M. Chemical and biological transformations for detoxification of trichothecene mycotoxins in human and animal food chains: A review. Trends Food Sci. Technol. 2010, 21, 67–76. [Google Scholar] [CrossRef]
- He, J.; Zhou, T. Patented techniques for detoxification of mycotoxins in feeds and food matrices. Recent Pat. Food Nutr. Agric. 2010, 2, 96–104. [Google Scholar] [CrossRef] [PubMed]
- Zhu, Y.; Hassan, Y.I.; Watts, C.; Zhou, T. Innovative technologies for the mitigation of mycotoxins in animal feed and ingredients—A review of recent patents. Anim. Feed Sci. Technol. 2016, 216, 19–29. [Google Scholar] [CrossRef]
- European Commission (EC). Commission regulation (EC) No 386/2009 of 12 May 2009. Amending regulation (EC) No. 1831/2003 of the European Parliament and of the Council as regards the establishment of a new functional group of feed additives. Off. J. Eur. Union 2009, L118, 166. [Google Scholar]
- Hahn, I.; Kunz-Vekiru, E.; Twaruzek, M.; Grajewski, J.; Krska, R.; Berthiller, F. Aerobic and anaerobic in vitro testing of feed additives claiming to detoxify deoxynivalenol and zearalenone. Food Addit. Contam. Part A 2015, 32, 922–933. [Google Scholar] [CrossRef] [PubMed]
- He, J.; Hassan, Y.I.; Perilla, N.; Li, X.; Boland, G.J.; Zhou, T. Bacterial epimerization as a route for deoxynivalenol detoxification: The influence of growth and environmental conditions. Front. Microbiol. 2016, 7, 572. [Google Scholar] [CrossRef] [PubMed]
- Ikunaga, Y.; Sato, I.; Grond, S.; Numaziri, N.; Yoshida, S.; Yamaya, H.; Hiradate, S.; Hasegawa, M.; Toshima, H.; Koitabashi, M.; et al. Nocardioides sp. strain WSN05-2, isolated from a wheat field, degrades deoxynivalenol, producing the novel intermediate 3-epi-deoxynivalenol. Appl. Microbiol. Biotechnol. 2011, 89, 419–427. [Google Scholar] [CrossRef] [PubMed]
- Islam, R.; Zhou, T.; Young, J.C.; Goodwin, P.H.; Pauls, K.P. Aerobic and anaerobic de-epoxydation of mycotoxin deoxynivalenol by bacteria originating from agricultural soil. World J. Microbiol. Biotechnol. 2012, 28, 7–13. [Google Scholar] [CrossRef] [PubMed]
- Sato, I.; Ito, M.; Ishizaka, M.; Ikunaga, Y.; Sato, Y.; Yoshida, S.; Koitabashi, M.; Tsushima, S. Thirteen novel deoxynivalenol-degrading bacteria are classified within two genera with distinct degradation mechanisms. FEMS Microbiol. Lett. 2012, 327, 110–117. [Google Scholar] [CrossRef] [PubMed]
- Shima, J.; Takase, S.; Takahashi, Y.; Iwai, Y.; Fujimoto, H.; Yamazaki, M.; Ochi, K. Novel detoxification of the trichothecene mycotoxin deoxynivalenol by a soil bacterium isolated by enrichment culture. Appl. Environ. Microbiol. 1997, 63, 3825–3830. [Google Scholar] [PubMed]
- He, W.; Yuan, Q.; Zhang, Y.; Guo, M.; Gong, A.; Zhang, J.; Wu, A.; Huang, T.; Qu, B.; Li, H.; et al. Aerobic de-epoxydation of trichothecene mycotoxins by a soil bacterial consortium isolated using in situ soil enrichment. Toxins 2016, 8, 277. [Google Scholar] [CrossRef] [PubMed]
- Tan, H.; Hu, Y.; He, J.; Wu, L.; Liao, F.; Luo, B.; He, Y.; Zuo, Z.; Ren, Z.; Zhong, Z.; et al. Zearalenone degradation by two Pseudomonas strains from soil. Mycotoxin Res. 2014, 30, 191–196. [Google Scholar] [CrossRef] [PubMed]
- Lei, Y.; Zhao, L.; Ma, Q.; Zhang, J.; Zhou, T.; Gao, C.; Ji, C. Degradation of zearalenone in swine feed and feed ingredients by Bacillus subtilis ANSB01G. World Mycotoxin J. 2014, 7, 143–151. [Google Scholar] [CrossRef]
- Cho, K.J.; Kang, J.S.; Cho, W.T.; Lee, C.H.; Ha, J.K.; Song, K.B. In vitro degradation of zearalenone by Bacillus subtilis. Biotechnol. Lett. 2010, 32, 1921–1924. [Google Scholar] [CrossRef] [PubMed]
- Yi, P.; Pai, C.; Liu, J. Isolation and characterization of a Bacillus licheniformis strain capable of degrading zearalenone. World J. Microbiol. Biotechnol. 2011, 27, 1035–1043. [Google Scholar] [CrossRef]
- Kakeya, H.; Takahashi-Ando, N.; Kimura, M.; Onose, R.; Yamaguchi, I.; Osada, H. Biotransformation of the mycotoxin, zearalenone, to a non-estrogenic compound by a fungal strain of Clonostachys sp. Biosci. Biotechnol. Biochem. 2002, 66, 2723–2726. [Google Scholar] [CrossRef] [PubMed]
- Yu, Y.; Qiu, L.; Wu, H.; Tang, Y.; Yu, Y.; Li, X.; Liu, D. Degradation of zearalenone by the extracellular extracts of Acinetobacter sp. SM04 liquid cultures. Biodegradation 2011, 22, 613–622. [Google Scholar] [CrossRef] [PubMed]
- Megharaj, M.; Garthwaite, I.; Thiele, J.H. Total biodegradation of the oestrogenic mycotoxin zearalenone by a bacterial culture. Lett. Appl. Microbiol. 1997, 24, 329–333. [Google Scholar] [CrossRef] [PubMed]
- Sangare, L.; Zhao, Y.; Folly, Y.M.E.; Chang, J.; Li, J.; Selvaraj, J.N.; Xing, F.; Zhou, L.; Wang, Y.; Liu, Y. Aflatoxin B1 degradation by a Pseudomonas strain. Toxins 2014, 6, 3028–3040. [Google Scholar] [CrossRef] [PubMed]
- Chen, Y.; Sheu, S.; Mau, J.; Hsieh, P. Isolation and characterization of a strain of Klebsiella pneumoniae with citrinin-degrading activity. World J. Microbiol. Biotechnol. 2011, 27, 487–493. [Google Scholar] [CrossRef]
- Kanpiengjai, A.; Mahawan, R.; Lumyong, S.; Khanongnuch, C. A soil bacterium Rhizobium borbori and its potential for citrinin-degrading application. Ann. Microbiol. 2016, 66, 807–816. [Google Scholar] [CrossRef]
- Benedetti, R.; Nazzi, F.; Locci, R.; Firrao, G. Degradation of fumonisin B1 by a bacterial strain isolated from soil. Biodegradation 2006, 17, 31–38. [Google Scholar] [CrossRef] [PubMed]
- Ito, M.; Sato, I.; Ishizaka, M.; Yoshida, S.; Koitabash, M.; Yoshida, S.; Tsushima, S. Bacterial cytochrome p450 system catabolizing the Fusarium toxin deoxynivalenol. Appl. Environ. Microbiol. 2013, 79, 1619–1628. [Google Scholar] [CrossRef] [PubMed]
- Völkl, A.; Vogle, B.; Schollenberger, M.; Karlovsky, P. Microbial detoxification of mycotoxin deoxynivalenol. J. Basic Microbiol. 2004, 44, 147–156. [Google Scholar] [CrossRef] [PubMed]
- El-Deeb, B.; Altalhi, A.; Khiralla, G.; Hassan, S.; Gherbawy, Y. Isolation and characterization of endophytic Bacilli bacterium from maize grains able to detoxify aflatoxin B1. Food Biotechnol. 2013, 27, 199–212. [Google Scholar] [CrossRef]
- Ito, M.; Sato, I.; Koitabashi, M.; Yoshida, S.; Imai, M.; Tsushima, S. A novel actinomycete derived from wheat heads degrades deoxynivalenol in the grain of wheat and barley affected by Fusarium head blight. Appl. Microbiol. Biotechnol. 2012, 96, 1059–1070. [Google Scholar] [CrossRef] [PubMed]
- Chang, X.; Wu, Z.; Wu, S.; Dai, Y.; Sun, C. Degradation of ochratoxin A by Bacillus amyloliquefaciens ASAG1. Food Addit. Contam. Part A 2015, 32, 564–571. [Google Scholar] [CrossRef] [PubMed]
- Zhang, W.; Xue, B.; Li, M.; Mu, Y.; Chen, Z.; Li, J.; Shan, A. Screening a strain of Aspergillus niger and optimization of fermentation conditions for degradation of aflatoxin B1. Toxins 2014, 6, 3157–3172. [Google Scholar] [CrossRef] [PubMed]
- Petchkongkaew, A.; Taillandier, P.; Gasaluck, P.; Lebrihi, A. Isolation of Bacillus spp. from Thai fermented soybean (Thua-nao): Screening for aflatoxin B1 and ochratoxin A detoxification. J. Appl. Microbiol. 2008, 104, 1495–1502. [Google Scholar] [CrossRef] [PubMed]
- Jard, G.; Liboz, T.; Mathieu, F.; Guyonvarc’h, A.; André, F.; Delaforge, M.; Lebrihi, A. Transformation of zearalenone to zearalenone-sulfate by Aspergillus spp. World Mycotoxin J. 2010, 3, 183–191. [Google Scholar] [CrossRef]
- Ricelli, A.; Baruzzi, F.; Solfrizzo, M.; Morea, M.; Fanizzi, F. Biotransformation of patulin by Gluconobacter oxydans. Appl. Environ. Microbiol. 2007, 73, 785–792. [Google Scholar] [CrossRef] [PubMed]
- Farzaneh, M.; Shi, Z.; Ghassempour, A.; Sedaghat, N.; Ahmadzadeh, M.; Mirabolfathy, M.; Javan-Nikkhah, M. Aflatoxin B1 degradation by Bacillus subtilis UTBSP1 isolated from pistachio nuts of Iran. Food Control 2012, 23, 100–106. [Google Scholar] [CrossRef]
- He, P.; Young, L.G.; Forsberg, C. Microbial transformation of deoxynivalenol (vomitoxin). Appl. Environ. Microbiol. 1992, 58, 3857–3863. [Google Scholar] [PubMed]
- Swanson, S.P.; Nicoletti, J.; Rood, H.D.; Buck, W.B.; Cote, L.M.; Yoshizawa, T. Metabolism of three trichothecene mycotoxins, T-2 toxin, diacetoxyscirpenol and deoxynivalenol, by bovine rumen microorganisms. J. Chromatogr. 1987, 414, 335–342. [Google Scholar] [CrossRef]
- Tan, H.; Zhang, Z.; Hu, Y.; Wu, L.; Liao, F.; He, J.; Luo, B.; He, Y.; Zuo, Z.; Ren, Z.; et al. Isolation and characterization of Pseudomonas otitidis TH-N1 capable of degrading zearalenone. Food Control 2015, 47, 285–290. [Google Scholar] [CrossRef]
- Kiessling, K.H.; Pettersson, H.; Sandholm, K.; Olsen, M. Metabolism of aflatoxin, ochratoxin, zearalenone, and three trichothecenes by intact rumen fluid, rumen protozoa, and rumen bacteria. Appl. Environ. Microbiol. 1984, 47, 1070–1073. [Google Scholar] [PubMed]
- Yu, H.; Zhou, T.; Gong, J.; Young, C.; Su, X.; Li, X.; Zhu, H.; Tsao, R.; Yang, R. Isolation of deoxynivalenol-transforming bacteria from the chicken intestines using the approach of PCR-DGGE guided microbial selection. BMC Microbiol. 2010, 10, 182. [Google Scholar] [CrossRef] [PubMed]
- Kollarczik, B.; Gareis, M.; Hanelt, M. In vitro transformation of the Fusarium mycotoxins deoxynivalenol and zearalenone by the normal gut microflora of pigs. Nat. Toxins 1994, 2, 105–110. [Google Scholar] [CrossRef] [PubMed]
- Gao, X.; Ma, Q.; Zhao, L.; Lei, Y.; Shan, Y.; Ji, C. Isolation of bacillus subtilis: Screening for aflatoxins B1, M1, and G1 detoxification. Eur. Food Res. Technol. 2011, 232, 957–962. [Google Scholar] [CrossRef]
- Guan, S.; Ji, C.; Zhou, T.; Li, J.; Ma, Q.; Niu, T. Aflatoxin B1 degradation by Stenotrophomonas maltophilia and other microbes selected using coumarin medium. Int. J. Mol. Sci. 2008, 9, 1489–1503. [Google Scholar] [CrossRef] [PubMed]
- Shi, L.; Liang, Z.; Li, J.; Hao, J.; Xu, Y.; Huang, K.; Tian, J.; He, X.; Xu, W. Ochratoxin A biocontrol and biodegradation by Bacillus subtilis CW 14. J. Sci. Food Agric. 2014, 94, 1879–1885. [Google Scholar] [CrossRef] [PubMed]
- Guan, S.; He, J.; Young, J.C.; Zhu, H.; Li, X.; Ji, C.; Zhou, T. Transformation of trichothecene mycotoxins by microorganisms from fish digesta. Aquaculture 2009, 290, 290–295. [Google Scholar] [CrossRef]
- Abrunhosa, L.; Serra, R.; Venâncio, A. Biodegradation of ochratoxin A by fungi isolated from grapes. J. Agric. Food Chem. 2002, 50, 7493–7496. [Google Scholar] [CrossRef] [PubMed]
- Devi, P.; Naik, C.G.; Rodrigues, C. Biotransformation of citrinin to decarboxycitrinin using an organic solvent-tolerant marine bacterium, Moraxella sp. MB1. Mar. Biotechnol. 2006, 8, 129–138. [Google Scholar] [CrossRef] [PubMed]
- Rodriguez, H.; Reveron, I.; Doria, F.; Costantini, A.; de Las Rivas, B.; Muňoz, R.; Garcia-Moruno, E. Degradation of ochratoxin A by Brevibacterium species. J. Agric. Food Chem. 2011, 59, 10755–10760. [Google Scholar] [CrossRef] [PubMed]
- Péteri, Z.; Téren, J.; Vágvölgyi, C.; Varga, J. Ochratoxin degradation and adsorption caused by astaxanthin-producing yeasts. Food Microbiol. 2007, 24, 205–210. [Google Scholar] [CrossRef] [PubMed]
- Abrunhosa, L.; Inês, A.; Rodrigues, A.I.; Guimarães, A.; Pereira, V.L.; Parpot, P.; Mendes-Faia, A.; Venâncio, A. Biodegradation of ochratoxin A by Pediococcus parvulus isolated from Douro wines. Int. J. Food Microbiol. 2014, 188, 45–52. [Google Scholar] [CrossRef] [PubMed]
- Zhang, X.; Guo, Y.; Ma, Y.; Chai, Y.; Li, Y. Biodegradation of patulin by a Byssochlamys nivea strain. Food Control 2016, 64, 142–150. [Google Scholar] [CrossRef]
- Binder, J.; Horvath, E.M.; Schatzmayr, G.; Ellend, N.; Danner, H.; Krska, R.; Braun, R. Screening for deoxynivalenol-detoxifying anaerobic rumen microorganisms. Cereal Res. Commun. 1997, 25, 343–346. [Google Scholar]
- Schatzmayr, G.; Zehner, F.; Täubel, M.; Schatzmayr, D.; Klimitsch, A.; Loibner, A.P.; Binder, E.M. Microbiologicals for deactivating mycotoxins. Mol. Nutr. Food Res. 2006, 50, 543–551. [Google Scholar] [CrossRef] [PubMed]
- He, C.; Fan, Y.; Liu, G.; Zhang, H. Isolation and identification of a strain of Aspergillus tubingensis with deoxynivalenol biotransformation capability. Int. J. Mol. Sci. 2008, 9, 2366–2375. [Google Scholar] [CrossRef] [PubMed]
- Liang, Z.; Li, J.; He, Y.; Guan, S.; Wang, N.; Ji, C.; Niu, T. AFB1 bio-degradation by a new strain—Stenotrophomonas. sp. Agric. Sci. China 2008, 7, 1433–1437. [Google Scholar] [CrossRef]
- Hormisch, D.; Brost, I.; Kohring, G.W.; Giffhorn, F.; Kroppenstedt, R.M.; Stackebrandt, E.; Färber, P.; Holzapfel, W.H. Mycobacterium fluoranthenivorans sp. nov., a fluoranthene and aflatoxin B1 degrading bacterium from contaminated soil of a former coal gas plant. Syst. Appl. Microbiol. 2004, 27, 653–660. [Google Scholar] [CrossRef] [PubMed]
- Dong, X.; Jiang, W.; Li, C.; Ma, N.; Xu, Y.; Meng, X. Patulin biodegradation by marine yeast Kodameae ohmeri. Food Addit. Contam. Part A 2015, 32, 352–360. [Google Scholar]
- Varga, J.; Rigó, K.; Téren, J. Degradation of ochratoxin A by Aspergillus species. Int. J. Food Microbiol. 2000, 59, 1–7. [Google Scholar] [CrossRef]
- Hwanga, C.; Draughon, F.A. Degradation of ochratoxin A by Acinetobacter calcoaceticus. J. Food Prot. 1994, 57, 410–414. [Google Scholar] [CrossRef]
- Barer, M.R.; Harwood, C.R. Bacterial viability and culturability. Adv. Microb. Physiol. 1999, 41, 93–137. [Google Scholar] [PubMed]
- Janssen, P.H.; Yates, P.S.; Grinton, B.E.; Taylor, P.M.; Sait, M. Improved culturability of soil bacteria and isolation in pure culture of novel members of the divisions Acidobacteria, Actinobacteria, Proteobacteria, and Verrucomicrobia. Appl. Environ. Microbiol. 2002, 68, 2391–2396. [Google Scholar] [CrossRef] [PubMed]
- Lane, D.J.; Pace, B.; Olsen, G.J.; Stahl, D.A.; Sogin, M.L.; Pace, N.R. Rapid determination of 16S ribosomal RNA sequences for phylogenetic analyses. Proc. Natl. Acad. Sci. USA 1985, 82, 6955–6959. [Google Scholar] [CrossRef] [PubMed]
- Vos, M.; Quince, C.; Pijl, A.S.; Hollander, M.D.; Kowalchuk, G.A. A comparison of rpoB and 16S rRNA as markers in pyrosequencing studies of bacterial diversity. PLoS ONE 2012, 7, e30600. [Google Scholar] [CrossRef] [PubMed]
- Conlan, S.; Kong, H.H.; Segre, J.A. Species-level analysis of DNA sequence data from the NIH Human Microbiome Project. PLoS ONE 2012, 7, e47075. [Google Scholar] [CrossRef] [PubMed]
- Fettweis, J.M.; Serrano, M.G.; Sheth, N.U.; Mayer, C.M.; Glascock, A.L.; Brooks, J.P.; Jefferson, K.K.; Buck, G.A. Species-level classification of the vaginal microbiome. BMC Genom. 2012, 13 (Suppl. S8), S17. [Google Scholar]
- Srinivasan, R.; Karaoz, U.; Volegova, M.; MacKichan, J.; Kato-Maeda, M.; Miller, S.; Nadarajan, R.; Brodie, E.L.; Lynch, S.V. Use of 16S rRNA gene for identification of a broad range of clinically relevant bacterial pathogens. PLoS ONE 2015, 10, e0117617. [Google Scholar] [CrossRef] [PubMed]
- Quast, C.; Pruesse, E.; Yilmaz, P.; Gerken, J.; Schweer, T.; Yarza, P.; Peplies, J.; Glöckner, F.O. The SILVA ribosomal RNA gene database project: Improved data processing and web-based tools. Nucleic Acids Res. 2013, 41, D590–D596. [Google Scholar] [CrossRef] [PubMed]
- Cole, J.R.; Wang, Q.; Fish, J.A.; Chai, B.; McGarrell, D.M.; Sun, Y.; Brown, C.T.; Porras-Alfaro, A.; Kuske, C.R.; Tiedje, J.M. Ribosomal Database Project: Data and tools for high throughput rRNA analysis. Nucleic Acids Res. 2014, 42, D633–D642. [Google Scholar] [CrossRef] [PubMed]
- DeSantis, T.Z.; Hugenholtz, P.; Larsen, N.; Rojas, M.; Brodie, E.L.; Keller, K.; Huber, T.; Dalevi, D.; Hu, P.; Andersen, G.L. Greengenes, a chimera-checked 16S rRNA gene database and workbench compatible with ARB. Appl. Environ. Microbiol. 2006, 72, 5069–5072. [Google Scholar] [CrossRef] [PubMed]
- Klindworth, A.; Pruesse, E.; Schweer, T.; Peplies, J.; Quast, C.; Horn, M.; Glöckner, F.O. Evaluation of general 16S ribosomal RNA gene PCR primers for classical and next-generation sequencing-based diversity studies. Nucleic Acids Res. 2013, 41, e1. [Google Scholar] [CrossRef] [PubMed]
- Wang, Y.; Qian, P.-Y. Conservative fragments in bacterial 16S rRNA genes and primer design for 16S ribosomal DNA amplicons in metagenomic studies. PLoS ONE 2009, 4, e7401. [Google Scholar] [CrossRef] [PubMed]
- Liu, Z.; DeSantis, T.Z.; Andersen, G.L.; Knight, R. Accurate taxonomy assignments from 16S rRNA sequences produced by highly parallel pyrosequencers. Nucleic Acids Res. 2008, 36, e120. [Google Scholar] [CrossRef] [PubMed]
- Navas-Molina, J.A.; Peralta-Sánchez, J.M.; González, A.; McMurdie, P.J.; Vázquez-Baeza, Y.; Xu, Z.; Ursell, L.K.; Lauber, C.; Zhou, H.; Song, S.J.; et al. Advancing our understanding of the human microbiome using QIIME. Methods Enzymol. 2013, 531, 371–444. [Google Scholar] [PubMed]
- Schloss, P.D.; Westcott, S.L.; Ryabin, T.; Hall, J.R.; Hartmann, M.; Hollister, E.B.; Lesniewski, R.A.; Oakley, B.B.; Parks, D.H.; Robinson, C.J.; et al. Introducing mothur: Open-source, platform-independent, community-supported software for describing and comparing microbial communities. Appl. Environ. Microbiol. 2009, 75, 7537–7541. [Google Scholar] [CrossRef] [PubMed]
- D’Amore, R.; Ijaz, U.Z.; Schirmer, M.; Kenny, J.G.; Gregory, R.; Darby, A.C.; Shakya, M.; Podar, M.; Quince, C.; Hall, N. A comprehensive benchmarking study of protocols and sequencing platforms for 16S rRNA community profiling. BMC Genom. 2016, 17, 55. [Google Scholar] [CrossRef] [PubMed]
- Fouhy, F.; Clooney, A.G.; Stanton, C.; Claesson, M.J.; Cotter, P.D. 16S rRNA gene sequencing of mock microbial populations—Impact of DNA extraction method, primer choice and sequencing platform. BMC Microbiol. 2016, 16, 123. [Google Scholar] [CrossRef] [PubMed]
- Li, W.; Raoult, D.; Fournier, P.E. Bacterial strain typing in the genomic era. FEMS Microbiol. Rev. 2009, 33, 892–916. [Google Scholar] [CrossRef] [PubMed]
- Molnar, O.; Schatzmayr, G.; Fuchs, E.; Prillinger, H. Trichosporon mycotoxinivorans sp. nov., a new yeast species useful in biological detoxification of various mycotoxins. Syst. Appl. Microbiol. 2004, 27, 661–671. [Google Scholar] [CrossRef] [PubMed]
- From, C.; Pukall, R.; Schumann, P.; Hormazábal, V.; Granum, P.E. Toxin-producing ability among Bacillus spp. outside the Bacillus cereus group. Appl. Environ. Microbiol. 2005, 71, 1178–1183. [Google Scholar] [CrossRef] [PubMed]
- Generally Recognized As Safe (GRAS). Available online: https://www.Fda.Gov/food/ingredientspackaginglabeling/gras/ (accessed on 10 March 2017).
- Qualified Presumption of Safety (QPS). Available online: https://www.Efsa.Europa.Eu/en/topics/topic/qualified-presumption-safety-qps (accessed on 10 March 2017).
- Moss, M.O.; Long, M.T. Fate of patulin in the presence of the yeast Saccharomyces cerevisiae. Food Addit. Contam. 2002, 19, 387–399. [Google Scholar] [CrossRef] [PubMed]
- Abrunhosa, L.; Santos, L.; Venâncio, A. Degradation of ochratoxin A by proteases and by a crude enzyme of Aspergillus niger. Food Biotechnol. 2006, 20, 231–242. [Google Scholar] [CrossRef]
- Bejaoui, H.; Mathieu, F.; Taillandier, P.; Lebrihi, A. Biodegradation of ochratoxin A by Aspergillus section nigri species isolated from French grapes: A potential means of ochratoxin A decontamination in grape juices and musts. FEMS Microbiol. Lett. 2006, 255, 203–208. [Google Scholar] [CrossRef] [PubMed]
- Guan, S.; Zhao, L.; Ma, Q.; Zhou, T.; Wang, N.; Hu, X.; Ji, C. In vitro efficacy of Myxococcus fulvus ANSM068 to biotransform aflatoxin B1. Int. J. Mol. Sci. 2010, 11, 4063–4079. [Google Scholar] [CrossRef] [PubMed]
- Tinyiro, S.E.; Wokadala, C.; Xu, D.; Yao, W. Adsorption and degradation of zearalenone by Bacillus strains. Folia Microbiol. 2011, 56, 321–327. [Google Scholar] [CrossRef] [PubMed]
- D’Souza, D.H.; Brackeet, R.E. The role of trace metal ions in aflatoxin B1 degradation by Flavobacterium aurantiacum. J. Food Prot. 1998, 61, 1666–1669. [Google Scholar] [CrossRef] [PubMed]
- D’Souza, D.H.; Brackett, R.E. The influence of divalent cations and chelators on aflatoxin B1 degradation by Flavobacterium aurantiacum. J. Food Prot. 2000, 63, 102–105. [Google Scholar] [CrossRef] [PubMed]
- D’Souza, D.H.; Brackett, R.E. Aflatoxin B1 degradation by Flavobacterium aurantiacum in the presence of reducing conditions and seryl and sulfhydryl group inhibitors. J. Food Prot. 2001, 64, 268–271. [Google Scholar] [CrossRef] [PubMed]
- Das, A.; Bhattacharya, S.; Palaniswamy, M.; Angayarkanni, J. Biodegradation of aflatoxin B1 in contaminated rice straw by Pleurotus ostreatus MTCC 142 and Pleurotus ostreatus GHBBF10 in the presence of metal salts and surfactants. World J. Microbiol. Biotechnol. 2014, 30, 2315–2324. [Google Scholar] [CrossRef] [PubMed]
- Hamid, A.B.; Smith, J.E. Degradation of aflatoxin by Aspergillus flavus. J. Gen. Microbiol. 1987, 133, 2023–2029. [Google Scholar] [CrossRef] [PubMed]
- Yang, Q.; Wang, J.; Zhang, H.; Li, C.; Zhang, X. Ochratoxin A is degraded by Yarrowia lipolytica and generates non-toxic degradation products. World Mycotoxin J. 2016, 9, 269–278. [Google Scholar] [CrossRef]
- Eshelli, M.; Harvey, L.; Edrada-Ebel, R.; McNeil, B. Metabolomics of the bio-degradation process of aflatoxin B1 by actinomycetes at an initial pH of 6.0. Toxins 2015, 7, 439–456. [Google Scholar] [CrossRef] [PubMed]
- Kong, Q.; Zhai, C.; Guan, B.; Li, C.; Shan, S.; Yu, J. Mathematic modeling for optimum conditions on aflatoxin B1 degradation by the aerobic bacterium Rhodococcus erythropolis. Toxins 2012, 4, 1181–1195. [Google Scholar] [CrossRef] [PubMed]
- Sun, X.; He, X.; Xue, K.; Li, Y.; Xu, D.; Qian, H. Biological detoxification of zearalenone by Aspergillus niger strain FS10. Food Chem. Toxicol. 2014, 72, 76–82. [Google Scholar] [CrossRef] [PubMed]
- Zhu, R.; Feussner, K.; Wu, T.; Yan, F.; Karlovsky, P.; Zheng, X. Detoxification of mycotoxin patulin by the yeast Rhodosporidium paludigenum. Food Chem. 2015, 179, 1–5. [Google Scholar] [CrossRef] [PubMed]
- Li, X.; Zhu, C.; Lange, C.F.M.d.; Zhou, T.; He, J.; Yu, H.; Gong, J.; Young, J.C. Efficacy of detoxification of deoxynivalenol-contaminated corn by Bacillus sp. LS100 in reducing the adverse effects of the mycotoxin on swine growth performance. Food Addit. Contam. Part A 2011, 28, 894–901. [Google Scholar] [CrossRef] [PubMed]
- He, J.; Yang, R.; Zhou, T.; Tsao, R.; Young, J.C.; Zhu, H.; Li, X.-Z.; Boland, G.J. Purification of deoxynivalenol from Fusarium graminearum rice culture and mouldy corn by high-speed counter-current chromatography. J. Chromatogr. A 2007, 1151, 187–192. [Google Scholar] [CrossRef] [PubMed]
- He, J.; Yang, R.; Zhou, T.; Boland, G.J.; Scott, P.M.; Bondy, G.S. An epimer of deoxynivalenol: Purification and structure identification of 3-epi-deoxynivalenol. Food Addit. Contam. Part A 2015, 32, 1523–1530. [Google Scholar] [CrossRef] [PubMed]
- El-Sharkawy, S.; Abul-Hajj, Y. Microbial transformation of zearalenone, I. Formation of zearalenone-4-O-β-glucoside. J. Nat. Prod. 1987, 50, 520–521. [Google Scholar] [CrossRef]
- El-Sharkawy, S.H.; Abul-Hajj, Y.J. Microbial transformation of zearalenone. 2. Reduction, hydroxylation, and methylation products. J. Org. Chem. 1988, 53, 515–519. [Google Scholar] [CrossRef]
- El-Sharkawy, S.H.; Selim, M.I.; Afifi, M.S.; Halaweish, F.T. Microbial transformation of zearalenone to a zearalenone sulfate. Appl. Environ. Microbiol. 1991, 57, 549–552. [Google Scholar]
- Vekiru, E.; Hametner, C.; Mitterbauer, R.; Rechthaler, J.; Adam, G.; Schatzmayr, G.; Krska, R.; Schuhmacher, R. Cleavage of zearalenone by Trichosporon mycotoxinivorans to a novel nonestrogenic metabolite. Appl. Environ. Microbiol. 2010, 76, 2353–2359. [Google Scholar] [CrossRef] [PubMed]
- Brodehl, A.; Möller, A.; Kunte, H.J.; Koch, M.; Maul, R. Biotransformation of the mycotoxin zearalenone by fungi of the genera Rhizopus and Aspergillus. FEMS Microbiol. Lett. 2014, 359, 124–130. [Google Scholar] [CrossRef] [PubMed]
- Chen, Y.; Kong, Q.; Chi, C.; Shan, S.; Guan, B. Biotransformation of aflatoxin B1 and aflatoxin G1 in peanut meal by anaerobic solid fermentation of Streptococcus thermophilus and Lactobacillus delbrueckii subsp. bulgaricus. Int. J. Food Microbiol. 2015, 211, 1–5. [Google Scholar] [CrossRef] [PubMed]
- Alberts, J.F.; Engelbrecht, Y.; Steyn, P.S.; Holzapfel, W.H.; van Zyl, W.H. Biological degradation of aflatoxin B1 by Rhodococcus erythropolis cultures. Int. J. Food Microbiol. 2006, 109, 121–126. [Google Scholar] [CrossRef] [PubMed]
- Das, A.; Bhattacharya, S.; Palaniswamy, M.; Angayarkanni, J. Aflatoxin B1 degradation during co-cultivation of Aspergillus flavus and Pleurotus ostreatus strains on rice straw. 3 Biotech 2015, 5, 279–284. [Google Scholar] [CrossRef] [PubMed]
- Samuel, M.S.; Sivaramakrishna, A.; Mehta, A. Degradation and detoxification of aflatoxin B1 by Pseudomonas putida. Int. Biodeterior. Biodegrad. 2014, 86, 202–209. [Google Scholar] [CrossRef]
- Castoria, R.; Mannina, L.; Durán-Patrón, R.; Maffei, F.; Sobolev, A.P.; de Felice, D.V.; Pinedo-Rivilla, C.; Ritieni, A.; Ferracane, R.; Wright, S.A. Conversion of the mycotoxin patulin to the less toxic desoxypatulinic acid by the biocontrol yeast Rhodosporidium kratochvilovae strain LS11. J. Agric. Food Chem. 2011, 59, 11571–11578. [Google Scholar] [CrossRef] [PubMed]
- Heinl, S.; Hartinger, D.; Thamhesl, M.; Vekiru, E.; Krska, R.; Schatzmayr, G.; Moll, W.D.; Grabherr, R. Degradation of fumonisin B1 by the consecutive action of two bacterial enzymes. J. Biotechnol. 2010, 145, 120–129. [Google Scholar] [CrossRef] [PubMed]
- Heinl, S.; Hartinger, D.; Thamhesl, M.; Schatzmayr, G.; Moll, W.D.; Grabherr, R. An aminotransferase from bacterium ATCC 55552 deaminates hydrolyzed fumonisin B1. Biodegradation 2011, 22, 25–30. [Google Scholar] [CrossRef] [PubMed]
- Hartinger, D.; Moll, W.D. Fumonisin elimination and prospects for detoxification by enzymatic transformation. World Mycotoxin J. 2011, 4, 271–283. [Google Scholar] [CrossRef]
- De Zutter, N.; Audenaert, K.; Arroyo-Manzanares, N.; de Boevre, M.; van Poucke, C.; de Saeger, S.; Haesaert, G.; Smagghe, G. Aphids transform and detoxify the mycotoxin deoxynivalenol via a type II biotransformation mechanism yet unknown in animals. Sci. Rep. 2016, 6, 38640. [Google Scholar] [CrossRef] [PubMed]
- Mannaa, M.; Kim, K.D. Microbe-mediated control of mycotoxigenic grain fungi in stored rice with focus on aflatoxin biodegradation and biosynthesis inhibition. Mycobiology 2016, 44, 67–78. [Google Scholar] [CrossRef] [PubMed]
- Vanhoutte, I.; Audenaert, K.; de Gelder, L. Biodegradation of mycotoxins: Tales from known and unexplored worlds. Front. Microbiol. 2016, 7, 561. [Google Scholar] [CrossRef] [PubMed]
- Tian, Y.; Tan, Y.; Liu, N.; Liao, Y.; Sun, C.; Wang, S.; Wu, A. Functional agents to biologically control deoxynivalenol contamination in cereal grains. Front. Microbiol. 2016, 7, 395. [Google Scholar] [CrossRef] [PubMed]
- Fernandez-Blanco, C.; Frizzell, C.; Shannon, M.; Ruiz, M.J.; Connolly, L. An in vitro investigation on the cytotoxic and nuclear receptor transcriptional activity of the mycotoxins fumonisin B1 and beauvericin. Toxicol. Lett. 2016, 257, 1–10. [Google Scholar] [CrossRef] [PubMed]
- Reisinger, N.; Dohnal, I.; Nagl, V.; Schaumberger, S.; Schatzmayr, G.; Mayer, E. Fumonisin B1 (FB1) induces lamellar separation and alters sphingolipid metabolism of in vitro cultured hoof explants. Toxins 2016, 8, 89. [Google Scholar] [CrossRef] [PubMed]
- Wang, X.; Wu, Q.; Wan, D.; Liu, Q.; Chen, D.; Liu, Z.; Martinez-Larranaga, M.R.; Martinez, M.A.; Anadon, A.; Yuan, Z. Fumonisins: Oxidative stress-mediated toxicity and metabolism in vivo and in vitro. Arch. Toxicol. 2016, 90, 81–101. [Google Scholar] [CrossRef] [PubMed]
- Peng, Z.; Chen, L.; Nussler, A.K.; Liu, L.; Yang, W. Current sights for mechanisms of deoxynivalenol-induced hepatotoxicity and prospective views for future scientific research: A mini review. J. Appl. Toxicol. 2016. [Google Scholar] [CrossRef] [PubMed]
- Boevre, M.D.; Graniczkowska, K.; Saeger, S.D. Metabolism of modified mycotoxins studied through in vitro and in vivo models: An overview. Toxicol. Lett. 2015, 233, 24–28. [Google Scholar] [CrossRef] [PubMed]
- Chen, X.; Murdoch, R.; Shafer, D.J.; Ajuwon, K.M.; Applegate, T.J. Cytotoxicity of various chemicals and mycotoxins in fresh primary duck embryonic fibroblasts: A comparison to HepG2 cells. J. Appl. Toxicol. 2016, 36, 1437–1445. [Google Scholar] [CrossRef] [PubMed]
- Jakimiuk, E.; Gajecka, M.; Jana, B.; Brzuzan, P.; Zielonka, L.; Skorska-Wyszynska, E.; Gajecki, M. Factors determining sensitivity of prepubertal gilts to hormonal influence of zearalenone. Pol. J. Vet. Sci. 2009, 12, 149–158. [Google Scholar] [PubMed]
- He, J.; Bondy, G.S.; Zhou, T.; Caldwell, D.; Boland, G.J.; Scott, P.M. Toxicology of 3-epi-deoxynivalenol, a deoxynivalenol-transformation product by Devosia mutans 17-2-E-8. Food Chem. Toxicol. 2015, 84, 250–259. [Google Scholar] [CrossRef] [PubMed]
- Minervini, F.; Giannoccaro, A.; Cavallini, A.; Visconti, A. Investigations on cellular proliferation induced by zearalenone and its derivatives in relation to the estrogenic parameters. Toxicol. Lett. 2005, 159, 272–283. [Google Scholar] [CrossRef] [PubMed]
- Zhao, L.; Lei, Y.; Bao, Y.; Jia, R.; Ma, Q.; Zhang, J.; Chen, J.; Ji, C. Ameliorative effects of Bacillus subtilis ANSB01G on zearalenone toxicosis in pre-pubertal female gilts. Food Addit. Contam. Part A 2015, 32, 617–625. [Google Scholar] [CrossRef] [PubMed]
- Kriszt, R.; Krifaton, C.; Szoboszlay, S.; Cserháti, M.; Kriszt, B.; Kukolya, J.; Czéh, A.; Fehér-Tóth, S.; Török, L.; Szőke, Z.; et al. A new zearalenone biodegradation strategy using non-pathogenic Rhodococcus pyridinivorans K408 strain. PLoS ONE 2012, 7, e43608. [Google Scholar] [CrossRef] [PubMed]
- Haq, M.; Gonzalez, N.; Mintz, K.; Jaja-Chimedza, A.; Jesus, C.L.D.; Lydon, C.; Welch, A.Z.; Berry, J.P. Teratogenicity of ochratoxin A and the degradation product, ochratoxin α, in the zebrafish (Danio rerio) embryo model of vertebrate development. Toxins 2016, 8, 40. [Google Scholar] [CrossRef] [PubMed]
- Dearfield, K.L.; Gollapudi, B.B.; Bemis, J.C.; Benz, R.D.; Douglas, G.R.; Elespuru, R.K.; Johnson, G.E.; Kirkland, D.J.; LeBaron, M.J.; Li, A.P.; et al. Next generation testing strategy for assessment of genomic damage: A conceptual framework and considerations. Environ. Mol. Mutagen. 2016. [Google Scholar] [CrossRef] [PubMed]
- Lean, I.J.; Lucy, M.C.; McNamara, J.P.; Bradford, B.J.; Block, E.; Thomson, J.M.; Morton, J.M.; Celi, P.; Rabiee, A.R.; Santos, J.E.; et al. Invited review: Recommendations for reporting intervention studies on reproductive performance in dairy cattle: Improving design, analysis, and interpretation of research on reproduction. J. Dairy Sci. 2016, 99, 1–17. [Google Scholar] [CrossRef] [PubMed]
- Park, S.H.; Kim, J.; Kim, D.; Moon, Y. Mycotoxin detoxifiers attenuate deoxynivalenol-induced pro-inflammatory barrier insult in porcine enterocytes as an in vitro evaluation model of feed mycotoxin reduction. Toxicol. In Vitro 2017, 38, 108–116. [Google Scholar] [CrossRef] [PubMed]
- Juan-Garcia, A.; Manyes, L.; Ruiz, M.J.; Font, G. Applications of flow cytometry to toxicological mycotoxin effects in cultured mammalian cells: A review. Food Chem. Toxicol. 2013, 56, 40–59. [Google Scholar] [CrossRef] [PubMed]
- Guillouzo, A.; Corlu, A.; Aninat, C.; Glaise, D.; Morel, F.; Guguen-Guillouzo, C. The human hepatoma HepaRG cells: A highly differentiated model for studies of liver metabolism and toxicity of xenobiotics. Chem. Biol. Interact. 2007, 168, 66–73. [Google Scholar] [CrossRef] [PubMed]
- Berger, V.; Gabriel, A.F.; Sergent, T.; Trouet, A.; Larondelle, Y.; Schneider, Y.J. Interaction of ochratoxin A with human intestinal Caco-2 cells: Possible implication of a multidrug resistance-associated protein (MRP2). Toxicol. Lett. 2003, 140–141, 465–476. [Google Scholar] [CrossRef]
- Mace, K.; Offord, E.A.; Harris, C.C.; Pfeifer, A.M. Development of in vitro models for cellular and molecular studies in toxicology and chemoprevention. Arch. Toxicol. Suppl. 1998, 20, 227–236. [Google Scholar] [PubMed]
- Pierron, A.; Mimoun, S.; Murate, L.S.; Loiseau, N.; Lippi, Y.; Bracarense, A.P.; Schatzmayr, G.; He, J.; Zhou, T.; Moll, W.D.; et al. Microbial biotransformation of DON: Molecular basis for reduced toxicity. Sci. Rep. 2016, 6, 29105. [Google Scholar] [CrossRef] [PubMed]
- Payros, D.; Alassane-Kpembi, I.; Pierron, A.; Loiseau, N.; Pinton, P.; Oswald, I.P. Toxicology of deoxynivalenol and its acetylated and modified forms. Arch. Toxicol. 2016, 90, 2931–2957. [Google Scholar] [CrossRef] [PubMed]
- Harbeson, S.L.; Abelleira, S.M.; Akiyama, A.; Barrett, R., 3rd; Carroll, R.M.; Straub, J.A.; Tkacz, J.N.; Wu, C.; Musso, G.F. Stereospecific synthesis of peptidyl alpha-keto amides as inhibitors of calpain. J. Med. Chem. 1994, 37, 2918–2929. [Google Scholar] [CrossRef] [PubMed]
- Wu, Y.; Levons, J.; Narang, A.S.; Raghavan, K.; Rao, V.M. Reactive impurities in excipients: Profiling, identification and mitigation of drug-excipient incompatibility. AAPS PharmSciTech 2011, 12, 1248–1263. [Google Scholar] [CrossRef] [PubMed]
- Burgess, K.M.; Renaud, J.B.; McDowell, T.; Sumarah, M.W. Mechanistic insight into the biosynthesis and detoxification of fumonisin mycotoxins. ACS Chem. Biol. 2016, 11, 2618–2625. [Google Scholar] [CrossRef] [PubMed]
- Wetterhorn, K.M.; Newmister, S.A.; Caniza, R.K.; Busman, M.; McCormick, S.P.; Berthiller, F.; Adam, G.; Rayment, I. Crystal structure of Os79 (Os04g0206600) from Oryza sativa: A UDP-glucosyltransferase involved in the detoxification of deoxynivalenol. Biochemistry 2016, 55, 6175–6186. [Google Scholar] [CrossRef] [PubMed]
- Hassan, Y.I.; Zhu, H.L.; Zhu, Y.; Zhou, T. Beyond ribosomal binding: The increased polarity and aberrant molecular interactions of 3-epi-deoxynivalenol. Toxins 2016, 8, E261. [Google Scholar] [CrossRef] [PubMed]
- Kovalsky, P.; Kos, G.; Nahrer, K.; Schwab, C.; Jenkins, T.; Schatzmayr, G.; Sulyok, M.; Krska, R. Co-occurrence of regulated, masked and emerging mycotoxins and secondary metabolites in finished feed and maize–An extensive survey. Toxins 2016, 8, E363. [Google Scholar] [CrossRef] [PubMed]
- Berthiller, F.; Crews, C.; Dall’Asta, C.; Saeger, S.D.; Haesaert, G.; Karlovsky, P.; Oswald, I.P.; Seefelder, W.; Speijers, G.; Stroka, J. Masked mycotoxins: A review. Mol. Nutr. Food Res. 2013, 57, 165–186. [Google Scholar] [CrossRef] [PubMed]
- Gratz, S.W.; Dinesh, R.; Yoshinari, T.; Holtrop, G.; Richardson, A.J.; Duncan, G.; MacDonald, S.; Lloyd, A.; Tarbin, J. Masked trichothecene and zearalenone mycotoxins withstand digestion and absorption in the upper GI tract but are efficiently hydrolyzed by human gut microbiota in vitro. Mol. Nutr. Food Res. 2016. [Google Scholar] [CrossRef] [PubMed]
- Cirlini, M.; Barilli, A.; Galaverna, G.; Michlmayr, H.; Adam, G.; Berthiller, F.; Dall’Asta, C. Study on the uptake and deglycosylation of the masked forms of zearalenone in human intestinal Caco-2 cells. Food Chem. Toxicol. 2016, 98, 232–239. [Google Scholar] [CrossRef] [PubMed]
- Dall’Erta, A.; Cirlini, M.; Dall’Asta, M.; Del Rio, D.; Galaverna, G.; Dall’Asta, C. Masked mycotoxins are efficiently hydrolyzed by human colonic microbiota releasing their aglycones. Chem. Res. Toxicol. 2013, 26, 305–312. [Google Scholar] [CrossRef] [PubMed]
- Dellafiora, L.; Galaverna, G.; Righi, F.; Cozzini, P.; Dall’Asta, C. Assessing the hydrolytic fate of the masked mycotoxin zearalenone-14-glucoside—A warning light for the need to look at the “maskedome”. Food Chem. Toxicol. 2017, 99, 9–16. [Google Scholar] [CrossRef] [PubMed]
- Stoev, S.D. Foodborne mycotoxicoses, risk assessment and underestimated hazard of masked mycotoxins and joint mycotoxin effects or interaction. Environ. Toxicol. Pharmacol. 2015, 39, 794–809. [Google Scholar] [CrossRef] [PubMed]
- Bullerman, L.B.; Giesova, M.; Hassan, Y.; Deibert, D.; Ryu, D. Antifungal activity of sourdough bread cultures. Adv. Exp. Med. Biol. 2006, 571, 307–316. [Google Scholar] [PubMed]
- Hassan, Y.I.; Bullerman, L.B. Cell-surface binding of deoxynivalenol to lactobacillus paracasei subsp. tolerans isolated from sourdough starter culture. J. Microbiol. Biotechnol. Food Sci. 2013, 2, 2323–2325. [Google Scholar]
- Abe, K.; Ichikawa, H. Gene overexpression resources in cereals for functional genomics and discovery of useful genes. Front. Plant Sci. 2016, 7, 1359. [Google Scholar] [CrossRef] [PubMed]
- Adesioye, F.A.; Makhalanyane, T.P.; Biely, P.; Cowan, D.A. Phylogeny, classification and metagenomic bioprospecting of microbial acetyl xylan esterases. Enzym. Microb. Technol. 2016, 93, 79–91. [Google Scholar] [CrossRef] [PubMed]
- Chu, Q.; Ma, J.; Saghatelian, A. Identification and characterization of sORF-encoded polypeptides. Crit. Rev. Biochem. Mol. Biol. 2015, 50, 134–141. [Google Scholar] [CrossRef] [PubMed]
- Wrenbeck, E.E.; Faber, M.S.; Whitehead, T.A. Deep sequencing methods for protein engineering and design. Curr. Opin. Struct. Biol. 2016, 45, 36–44. [Google Scholar] [CrossRef] [PubMed]
- Mehta, D.; Satyanarayana, T. Bacterial and archaeal alpha-amylases: Diversity and amelioration of the desirable characteristics for industrial applications. Front. Microbiol. 2016, 7, 1129. [Google Scholar] [CrossRef] [PubMed]
- Heine, T.; Tucker, K.; Okonkwo, N.; Assefa, B.; Conrad, C.; Scholtissek, A.; Schlomann, M.; Gassner, G.; Tischler, D. Engineering styrene monooxygenase for biocatalysis: Reductase-epoxidase fusion proteins. Appl. Biochem. Biotechnol. 2016, 1–21. [Google Scholar] [CrossRef] [PubMed]
- Chen, H.; Zhu, Z.; Huang, R.; Zhang, Y.P. Coenzyme engineering of a hyperthermophilic 6-phosphogluconate dehydrogenase from NADP+ to NAD+ with its application to biobatteries. Sci. Rep. 2016, 6, 36311. [Google Scholar] [CrossRef] [PubMed]
- Wambacq, E.; Vanhoutte, I.; Audenaert, K.; Gelder, L.D.; Haesaert, G. Occurrence, prevention and remediation of toxigenic fungi and mycotoxins in silage: A review. J. Sci. Food Agric. 2016, 96, 2284–2302. [Google Scholar] [CrossRef] [PubMed]
- European Food Safety Authority (EFSA). Guidance for the preparation of dossiers for technological additives—EFSA panel on additives and products or substances used in animal feed (FEEDAP). EFSA J. 2012, 10, 2528. [Google Scholar]
- EU Authorizations for Mycofix® Secure and Biomin® BBSH 797. First-Ever Products with Official Anti-Mycotoxin Claim. Available online: http://www.Biomin.Net/en/press-releases/eu-authorizations-for-mycofixr-secure-and-biominr-bbsh-797-first-ever-products-with-official-anti-mycotoxin-claim/ (accessed on 10 March 2017).
- Positive EFSA Opinion for Mycofix® Ingredient Biomin® BBSH 797 for All Avian Species. Available online: http://www.Biomin.Net/en/press-releases/positive-efsa-opinion-for-mycofixr-ingredient-biominr-bbsh-797-for-all-avian-species/ (accessed on 10 March 2017).
- EU Authorization for FUMzyme® Proves Fumonisin Biotransformation. Available online: http://www.Biomin.Net/en/press-releases/eu-authorization-for-fumzymer-proves-fumonisin-biotransformation/ (accessed on 10 March 2017).
- Varga, J.; Péteri, Z.; Tábori, K.; Téren, J.; Vágvölgyi, C. Degradation of ochratoxin A and other mycotoxins by rhizopus isolates. Int. J. Food Microbiol. 2005, 99, 321–328. [Google Scholar] [CrossRef] [PubMed]
- Döll, S.; Dänicke, S. In vivo detoxification of Fusarium toxins. Arch. Anim. Nutr. 2004, 58, 419–441. [Google Scholar] [CrossRef] [PubMed]
- Kong, C.; Shin, S.Y.; Kim, B.G. Evaluation of mycotoxin sequestering agents for aflatoxin and deoxynivalenol: An in vitro approach. SpringerPlus 2014, 3, 346. [Google Scholar] [CrossRef] [PubMed]
- Masching, S.; Naehrer, K.; Schwartz-Zimmermann, H.E.; Sarandan, M.; Schaumberger, S.; Dohnal, I.; Nagl, V.; Schatzmayr, D. Gastrointestinal degradation of fumonisin B1 by carboxylesterase fumD prevents fumonisin induced alteration of sphingolipid metabolism in turkey and swine. Toxins 2016, 8, 84. [Google Scholar] [CrossRef] [PubMed]
- Lee, J.T.; Jessen, K.A.; Beltran, R.; Starkl, V.; Schatzmayr, G.; Borutova, R.; Caldwell, D.J. Effects of mycotoxin-contaminated diets and deactivating compound in laying hens: 2. Effects on white shell egg quality and characteristics. Poult. Sci. 2012, 91, 2096–2104. [Google Scholar] [CrossRef] [PubMed]
- Pinton, P.; Tsybulskyy, D.; Lucioli, J.; Laffitte, J.; Callu, P.; Lyazhri, F.; Grosjean, F.; Bracarense, A.P.; Kolf-Clauw, M.; Oswald, I.P. Toxicity of deoxynivalenol and its acetylated derivatives on the intestine: Differential effects on morphology, barrier function, tight junction proteins, and mitogen-activated protein kinases. Toxicol. Sci. 2012, 130, 180–190. [Google Scholar] [CrossRef] [PubMed]
| Mycotoxin | Enrichment | Isolation | ||||
|---|---|---|---|---|---|---|
| Medium | Strategy/Methodology | Medium | Strategy/Methodology | Biotransforming Strains | Reference | |
| DON | Corn meal broth | Soil samples enriched from the corn contaminated by the DON-producing fungi | Corn meal agar | Single colonies screening; extended incubation time for slow-growth strains | Devosia mutans 17-2-E-8 | [12] |
| Anaerobic incubation medium with 10% chicken cecal digesta extract | In vivo enrichment with moldy wheat; antibiotics treatment; guiding the enrichment by PCR-DGGE | L10 agar | Single colonies screening | Bacillus sp. LS-100 | [43] | |
| M10 medium + DON (100 µg/mL) | Treatment by antibiotics and hemin | M1 medium | Single colonies screening | Eubacterium sp. BBSH 797 | [55,56] | |
| Mineral salts with peptone medium + DON (50 µg/mL) | Antibiotics and heat treatment; guiding the enrichment by T-RFLP | Mineral salts with peptone agar | Single colonies screening; extended incubation time for slow-growth strains | Microbial consortium with at least 6 bacterial genera | [14] | |
| Mineral medium + DON (100 µg/mL) | DON as a sole carbon source | 1/100 nutrient agar | Single colonies screening | Nocardioides sp. WSN05-2 | [13] | |
| Mineral salt medium + DON (100 µg/mL) | In situ plant enrichment in contaminated wheat head by spraying DON | MRDG medium | Single colonies screening; using gellan gum rather than agar | Marmoricola sp. MIM116 | [32] | |
| Mineral medium + DON (100 µg/mL) | DON as a sole carbon source | Reasoner’s 2A (R2A) agar, 1/100 nutrient agar | Single colonies screening | 9 Nocardioides spp. and 4 Devosia spp. | [15] | |
| BYE medium + DON (200 µg/mL) | Repeated sub-culturing in fresh medium with high level of DON (200 µg/mL) | 1/10 nutrient agar | Single colonies screening | E3-39 (belonging to Agrobacterium or Rhizobium) | [16] | |
| Inorganic salt culture medium + DON (4 µg/mL) | Enrichment with minimal nutrients | Czapek’s agar, LB agar | Single colonies screening | Aspergillus tubingensis NJA-1 | [57] | |
| Mineral salts with peptone medium + DON (50 µg/mL) | In situ soil enrichment by spraying DON; guiding the enrichment by PCR-DGGE | - | - | Microbial consortium | [17] | |
| ZEA | Minimal salt medium + ZEA (2 µg/mL) | ZEA as a sole carbon source | LB agar | Single colonies screening | Pseudomonas alcaliphila TH-C1, Pseudomonas plecoglossicida TH-L1 | [18] |
| LB broth | Selective screening Bacillus strains by heat treatment | LB agar | Single colonies screening | Bacillus subtilis ANSB01G | [19] | |
| Minimal salt medium + ZEA (2 µg/mL) | ZEA as a sole carbon source | LB agar | Single colonies screening | Pseudomonas otitidis TH-N1 | [41] | |
| M1 + ZEA (25 µg/mL) + nystatin (15 µg/mL), M2 + ZEA (500 µg/mL) | ZEA as a sole carbon source | Nutrient agar | Single colonies screening | Acinetobacter sp. SM04 | [23] | |
| M9 medium + ZEA (50 µg/mL) | ZEA as a sole carbon source | LB agar | Single colonies screening | Microbial consortium | [24] | |
| AFB1 | Coumarin medium (with 1% coumarin) | Coumarin, a basic molecular structure of aflatoxins, as a sole carbon source | Coumarin medium | Single colonies screening; coumarin as a sole carbon source | Stenotrophomonas maltophilia 35-3 | [46] |
| Coumarin medium (with 1% coumarin) | Coumarin, a basic molecular structure of aflatoxins, as a sole carbon source | Coumarin medium | Single colonies screening; coumarin as a sole carbon source | Pseudomonas aeruginosa N17-1 | [25] | |
| - | - | Modified Hormisch medium (with 0.1% coumarin) | Coumarin as a sole carbon source | Stenotrophomonas sp. NMO-3 | [58] | |
| Nutrient broth | Non-selective enrichment | Coumarin medium (with 0.1% coumarin) | Single colonies screening; coumarin as a sole carbon source; K-B disk diffusion | Aspergillus niger ND-1 | [34] | |
| Minimal salt/vitamin medium + fluoranthene (10 mg/mL) | Fluoranthene as a sole carbon source | R2A agar | Single colonies screening | Mycobacterium fluoranthenivorans sp. nov. | [59] | |
| Minimal salt medium + AFB1 (10 µg/mL) | AFB1 as a sole carbon source | Minimal salt agar + AFB1 (10 µg/mL) | Single colonies screening | Bacillus sp. TUBF1 | [31] | |
| - | - | Nutrient agar | Single colonies screening | Bacillus licheniformis CM21, Bacillus subtilis MHS 13 | [35] | |
| AFB1, AFM1, AFG1 | LB broth | Selective screening Bacillus strains by heat treatment | LB agar | Single colonies screening | Bacillus subtilis ANSB060 | [45] |
| PAT | Mineral salt medium + increased concentration (300–600 µg/mL) of PAT | PAT as a sole carbon source | Mineral salt agar + PAT (600 µg/mL) | Single colonies screening | Byssochlamys nivea FF1-2 | [54] |
| - | - | YEPD medium + PAT (10 µg/mL) | Screening in liquid medium | Kodameae ohmeri HYJM34 | [60] | |
| CIT | Mineral broth + (1–4 µg/mL) of CIT | CIT as a sole carbon source | Mineral salt agar + CIT (10 µg/mL) | Single colonies screening | Klebsiella pneumoniae NPUST-B11 | [26] |
| Mineral broth + CIT (1 µg/mL) | CIT as a sole carbon source | Mineral salt agar + CIT (1–5 µg/mL) | Single colonies screening | Rhizobium borbori PS45 | [27] | |
| - | - | Nutrient agar | Screening strains by disc plate diffusion assay (50 µg/disk of CIT) | Moraxella sp. MB1 | [50] | |
| FUB1 | BYE medium + FUB1 (500 µg/mL) | Increasing population of FUB1-transforming microbes; antibiotics treatment | Nutrient agar (NA), NA + sucrose, NA + skim milk, PYEI agar, BYE agar | Single colonies screening | NCB 1492 (belonging to Delftia or Comamonas) | [28] |
| OTA | - | - | YES medium + OTA (2 µg/mL) | Screening in liquid medium | Aspergillus niger CBS 120.49 | [61] |
| - | - | Czapek-Dox medium + OTA (40 µg/plate) | Screening point-pated colonies by observing the loss of fluorescence | Acinetobacter calcoaceticus NRRL B-551 | [62] | |
| - | - | LB agar + OTA (3 µg/mL); medium with isocoumarin as the sole carbon source | Screening microbes using isocoumarin as a sole carbon source | Bacillus subtilis CW 14 | [47] | |
| Factor | Mycotoxin | Biotransforming Product(s) | Optimal Condition/Reverse Effect |
|---|---|---|---|
| Carbon source | AFB1 | U.I. a | Starch (4.0%) [34] |
| CIT | U.I. | Glucose (1.2%) [26] | |
| Nitrogen source | AFB1 | U.I. | Yeast extract (0.5%) [88]; tryptone (0.5%) [34] |
| CIT | U.I. | Peptone (0.3%) [26] | |
| Vitamins | CIT | U.I. | Vitamin C (100 µg/mL) [26] |
| Metals ions | DON | 3-epi-DON | Minerals added in the corn steep liquor and peptone [12] |
| ZEA | U.I. | Zn2+, Mn2+, Ca2+, Mg2+ (10 mmol/L) [89] | |
| AFB1 | U.I. | Mg2+, Cu2+ (10 mmol/L) [46]; Ca2+, Mg2+ (10 mmol/L) [90,91,92]; Mn2+, Cu2+ (10 mmol/L) [25]; Mg2+, Zn2+, Cu2+, Mn2+ (10 mmol/L) [93] | |
| Enzyme inhibitor/enhancer | ZEA | U.I. | Reverse effect: chelating agents of EDTA, OPT (10 mmol/L) [89] |
| AFB1 | U.I. | Reverse effect: chelating agents of EDTA, OPT (10 mmol/L) [90,91,92] | |
| OTA | OTα | Reverse effect: chelating agents of EDTA (10 mmol/L), OPT (1 mmol/L) [52] | |
| AFB1 | U.I. | Tween 80, Triton X-100 (0.05%) [93] | |
| AFB1 | U.I. | NADPH (0.2 mmol/L), NaIO4 (3 mmol/L) [94] | |
| Concentration of mycotoxins | AFB1 | U.I. | 0.5 µg/mL [93] |
| OTA | U.I. a | 0.1 µg/mL [95] | |
| Concentration of cells | OTA | U.I. | 108 CFU/mL [95] |
| OTA | OTα | 109 CFU/mL [53] | |
| Initial pH | DON | 3-epi-DON | pH = 7 [12] |
| DON | DOM-1 | pH = 6.5–7 [14]; pH = 5–10 [17] | |
| ZEA | U.I. | pH = 7–8 [89]; pH = 4.5 [41] | |
| AFB1 | U.I. | pH = 5–6 [96]; pH = 6–7 [31,34,88]; pH = 8 [46] | |
| PAT | E- and Z-ascladiol | pH = 3–6 [60] | |
| PAT | U.I. | pH = 3–5 [54] | |
| CIT | U.I. | pH = 7 [26] | |
| OTA | U.I. | pH = 4 [95] | |
| Temperature | DON | 3-epi-DON | 20–35 °C [12] |
| DON | DOM-1 | 20–35 °C [14]; 20–37 °C [17] | |
| ZEA | U.I. | 30–37 °C [41]; 42 °C [89] | |
| AFB1 | U.I. | 30–37 °C [31,34,46,88,96] | |
| PAT | E- and Z-ascladiol | 35 °C [60] | |
| PAT | U.I. | 37 °C [54] | |
| CIT | U.I. | 37 °C [26] | |
| OTA | OTα | 25–35 °C [52] | |
| OTA | U.I. | 28 °C [95] | |
| Shaking rate | CIT | U.I. | 200 RPM [26] |
| Oxygen preference | DON | DOM-1 | Aerobic condition [14] |
| Concentration of mycotoxins | OTA | U.I. | 0.1 µg/mL [95] |
| Pre-incubation time | CIT | U.I. | 36–48 h [26] |
| Mycotoxins/Biotransforming Products | Extraction Solvents | Analytical Method |
|---|---|---|
| DON | 50% methanol [12,43,48]; 84% acetonitrile [100,101]; ethyl acetate [30] | HPLC [12,13,15,100,101]; LC-MS [48,101,102]; ELISA [32]; HSCCC [101]; NMR [102] |
| 3-epi-DON | 50% methanol [12]; ethyl acetate [13] | HPLC [12,13,15]; LC-MS [102]; NMR [13,102] |
| DOM-1 (deepoxy DON) | 84% acetonitrile [100]; 50% methanol [48] | HPLC [100]; LC-MS [14,48]; GC-MS [17]; MS [43] |
| 3-keto-DON | Ethyl acetate [16,30] | MS [16,30]; NMR [16,30] |
| ZEA | 50% methanol [18,23,41]; 90% acetonitrile [19]; 84% acetonitrile [21]; 50% acetonitrile [98] | HPLC [18,19,21,24,41]; LC-MS [23,89,98]; ELISA [24] |
| 1-(3,5-dihydroxy-phenyl)-10′-hydroxy-1′E-undecene-6′-one | Chloroform [22] | TLC, MS, NMR [22] |
| ZEA-sulfate | 60% methanol [36] | LC-MS [36] |
| ZEA-4-O-β-glucoside | TLC, MS, NMR, IR [103] | |
| α-ZAL, β-ZAL, α-ZOL, β-ZOL, ZAN, 8′(S)-hydroxyzearalenone, ZEA-2,4-bis(methyl ether), ZEA-2-methyl ether | 50% chloroform [104] | TLC, MS, NMR, IR [104] |
| ZEN-4-O-sulfate | 33% chloroform:methanol (9:1) [105] | MS, NMR, infrared [105] |
| ZOM-1 ((5S)-5-({2,4-dihydroxy-6-[(1E)-5-hydroxypent-1-en-1-yl]benzoyl}oxy)hexanoic acid) | Ethyl acetate [106] | HPLC, LC-MS, NMR [106] |
| α-ZOL, α-ZOL-S, ZEA-14-sulfate, ZEA-16-sulfate, ZEA-14-Glc, ZEA-16-Glc | 50% acetonitrile [107] | LC-MS [107] |
| AFB1 | 60% methanol [108]; 50% methanol [45]; chloroform [38,46,88,109]; dichloromethane [110] | HPLC [25,38,45,46,88,96,108,109,110]; TLC [38,96,109]; ESMS [109]; LC-MS [38,96,109]; HR-FTMS [96]; ELISA [34] |
| AFB2 | Chloroform [25] | HPLC [25] |
| AFG1 | 60% methanol [108]; 50% methanol [45] | HPLC [45,108] |
| AFM1 | Chloroform [25]; 50% methanol [45] | HPLC [25,45] |
| AFD1, AFD2, AFD3 | Chloroform [111] | TLC, HPLC, GC-MS, FT-IR [111] |
| PAT | Ethyl acetate [54,60,99] | TLC [85]; HPLC([37,54,60,85]; LC-MS [99]; NMR [85] |
| DPA (desoxypatulinic acid) | Ethyl acetate [99] | HPLC [112]; LC-MS [99]; NMR [112] |
| E- and Z-ascladiol | Ethyl acetate [60,85] | TLC [85]; HPLC [37,60,85]; LC-MS [37,60]; NMR [37,85] |
| CIT | Acetone:ethyl acetate (1:1) [27]; ethyl acetate [50] | TLC(C04); HPLC [26,27] |
| Decarboxycitrinin | Ethyl acetate [50] | MS, NMR [50] |
| FUB1 | TLC [28]; HPLC [28]; GC-MS[28]; LC-MS [113,114] | |
| Heptadecanone, isononadecene, octadecenal, eicosane | GC-MS [28] | |
| Hydrolyzed FUB1 | LC-MS [113,114] | |
| 2-keto-hydrolyzed FUB1 | LC-MS, NMR [115] | |
| OTA | Methanol [51]; dichloromethane [33,52,61]; ethyl acetate [53] | TLC [52,61]; HPLC [33,51,52,53,61] |
| OTα | Methanol [51]; dichloromethane [52]; | HPLC[51,52]; LC-MS [51] |
| l-β-phenylalanine | Methanol [51] | HPLC [51] |
| Mycotoxins | Biotransforming Products | Model | Methodologies | Reference |
|---|---|---|---|---|
| DON | 3-epi-DON | Caco-2 cells | Evaluation of metabolic activity by an MTT cell proliferation assay | [127] |
| 3-epi-DON | 3T3 cells | Evaluation of DNA synthesis activity by a cell proliferation ELISA employing BrdU incorporation | [127] | |
| 3-epi-DON | Female B6C3F1 mice | Evaluation of effects on body weight gain, relative organ weights, food consumption, hematology and clinical chemistry | [127] | |
| DOM-1 | Swine kidney cells | Evaluation of metabolic activity by an MTT cell proliferation assay | [44] | |
| DOM-1 | Chicken lymphocytes | Evaluation of DNA synthesis activity by a cell proliferation ELISA employing BrdU incorporation | [56] | |
| DOM-1 | Starter pigs | Evaluation of effects on growth performance and serum metabolites | [100] | |
| 3-keto-DON | Mouse spleen lymphocytes | Evaluation of immunosuppressive activity by a cell proliferation assay | [16] | |
| ZEA | 1-(3,5-dihydroxy-phenyl)-10′-hydroxy-1′E-undecene-6′-one | MCF-7 cells | Evaluation of estrogenic activity by a WST cell proliferation assay | [22] |
| ZEA-sulfate | MCF-7 cells | Evaluation of estrogenic activity by an MTS cell proliferation assay | [36] | |
| α-ZAL, β-ZAL, α-ZOL, β-ZOL | MCF-7 and MDA-MB-231 cells | Evaluation of estrogenic activity by an MTT cell proliferation assay | [128] | |
| α-ZAL, β-ZAL, α-ZOL, β-ZOL, ZAN, 8′(S)-Hydroxyzearalenone, ZEA-2,4-bis(methyl ether), ZEA-2-methyl ether | Rat uteri | Evaluation of relative binding affinity by an estrogen receptor binding assay | [104] | |
| ZOM-1 | Yeast YZRM7 | Evaluation of estrogenic activity by a sensitive yeast assay | [106] | |
| ZOM-1 | Human estrogen receptor-α | Evaluation of estrogenic activity by a HitHunter EFC estrogen chemiluminescence assay | [106] | |
| U.I. a | Pre-pubertal female gilts | Evaluation of effects on growth performance, genital organs, serum hormones and histopathological changes | [129] | |
| U.I. | Yeast BLYES | Evaluation of estrogenic activity by a sensitive yeast assay | [130] | |
| U.I. | Pre-pubertal female rats | Evaluation of estrogenic activity by an immature uterotrophic assay | [130] | |
| AFB1 | AFD1, AFD2, AFD3 | Hela cells | Evaluation of cytotoxicity by an MTT cell proliferation assay | [111] |
| U.I. | L929 cells | Evaluation of cytotoxicity by an MTT cell proliferation assay | [108] | |
| U.I. | Salmonella typhimurium TA100 | Evaluation of mutagenicity by an Ames assay | [109] | |
| U.I. | Artemia salina | Evaluation of toxicity by an insect larvae survival assay | [31] | |
| PAT | DPA | Escherichia coli | Evaluation of microbial toxicity | [99] |
| DPA | Seeds of Arabidopsis thaliana | Evaluation of phytotoxicity | [99] | |
| DPA | Human hepatocytes LO2 | Evaluation of cytotoxicity by an MTT cell proliferation assay | [99] | |
| DPA | Human lymphocytes | Evaluation of cytotoxicity by a trypan blue cell proliferation assay | [112] | |
| CIT | U.I. | Bacillus subtilis TISTR | Evaluation of microbial toxicity | [27] |
| OTA | U.I. | HepG2 cells | Evaluation of cytotoxicity by an MTT cell proliferation assay | [95] |
| OTα | Zebrafish (Danio rerio) embryo | Evaluation of teratogenicity | [131] |
© 2017 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Zhu, Y.; Hassan, Y.I.; Lepp, D.; Shao, S.; Zhou, T. Strategies and Methodologies for Developing Microbial Detoxification Systems to Mitigate Mycotoxins. Toxins 2017, 9, 130. https://doi.org/10.3390/toxins9040130
Zhu Y, Hassan YI, Lepp D, Shao S, Zhou T. Strategies and Methodologies for Developing Microbial Detoxification Systems to Mitigate Mycotoxins. Toxins. 2017; 9(4):130. https://doi.org/10.3390/toxins9040130
Chicago/Turabian StyleZhu, Yan, Yousef I. Hassan, Dion Lepp, Suqin Shao, and Ting Zhou. 2017. "Strategies and Methodologies for Developing Microbial Detoxification Systems to Mitigate Mycotoxins" Toxins 9, no. 4: 130. https://doi.org/10.3390/toxins9040130
APA StyleZhu, Y., Hassan, Y. I., Lepp, D., Shao, S., & Zhou, T. (2017). Strategies and Methodologies for Developing Microbial Detoxification Systems to Mitigate Mycotoxins. Toxins, 9(4), 130. https://doi.org/10.3390/toxins9040130







