Cone Snails: A Big Store of Conotoxins for Novel Drug Discovery
Abstract
:1. Introduction
2. Diversity of Cone Snails
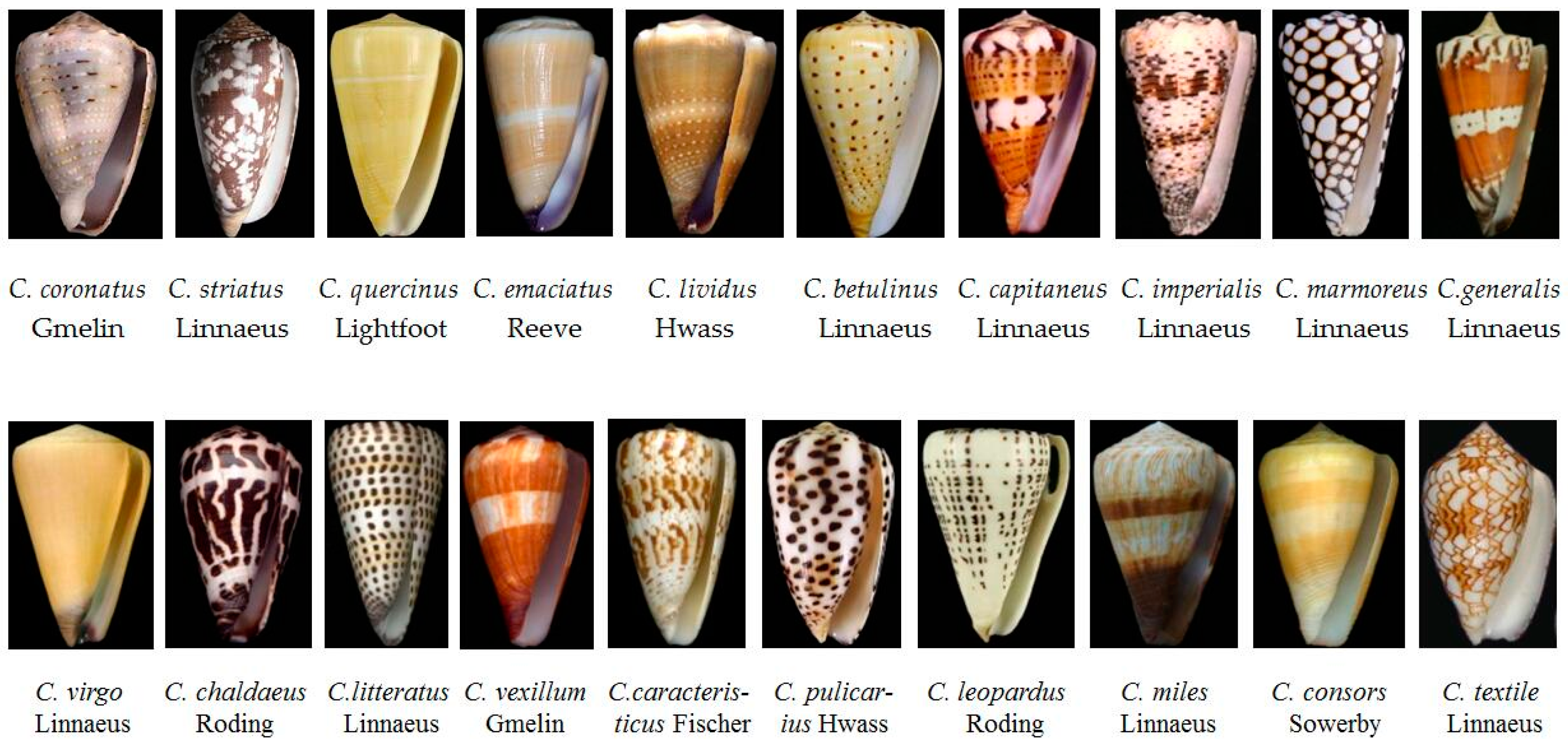
2.1. Various Phenotypes
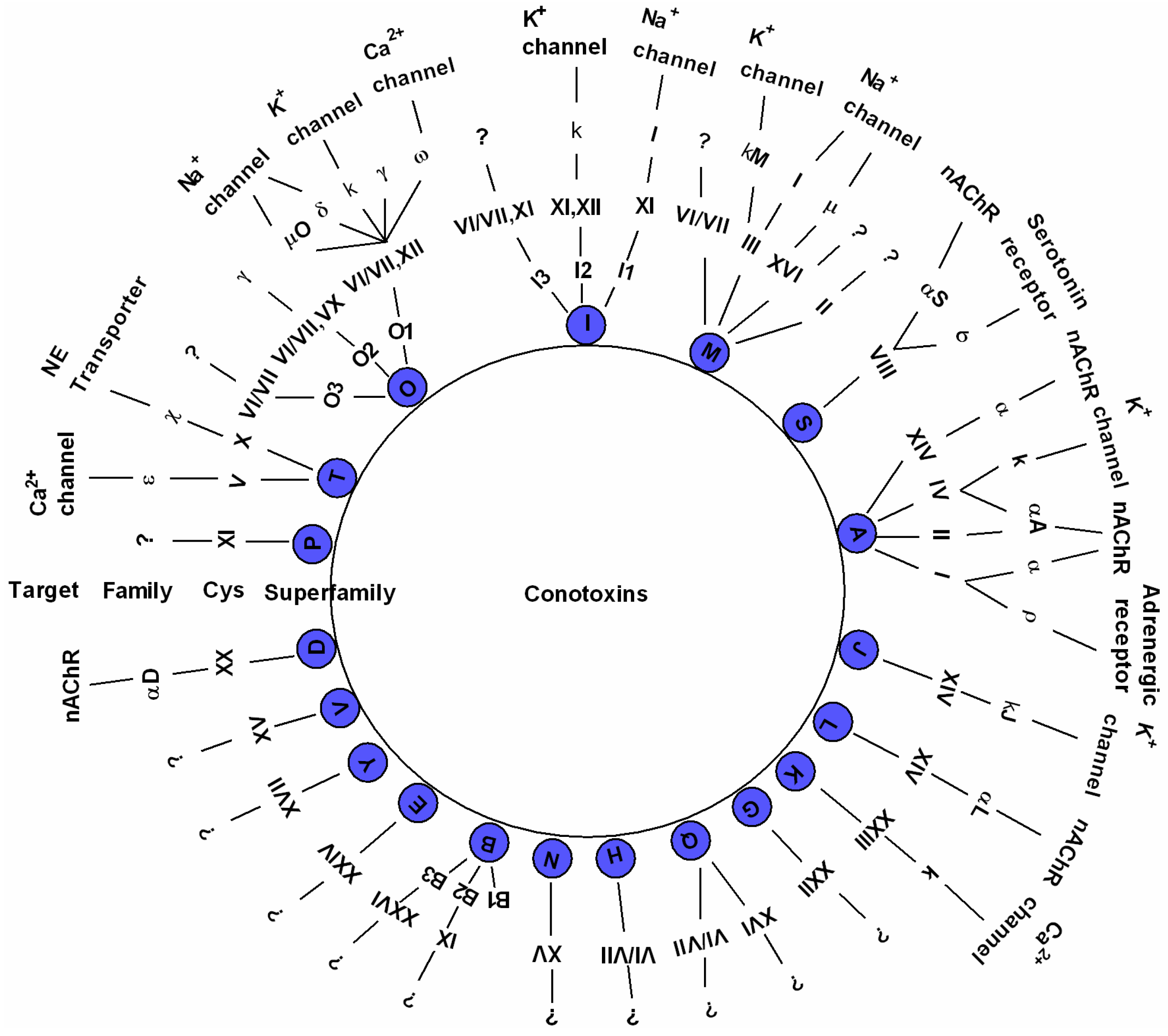
2.2. Diverse Conotoxins and Targets
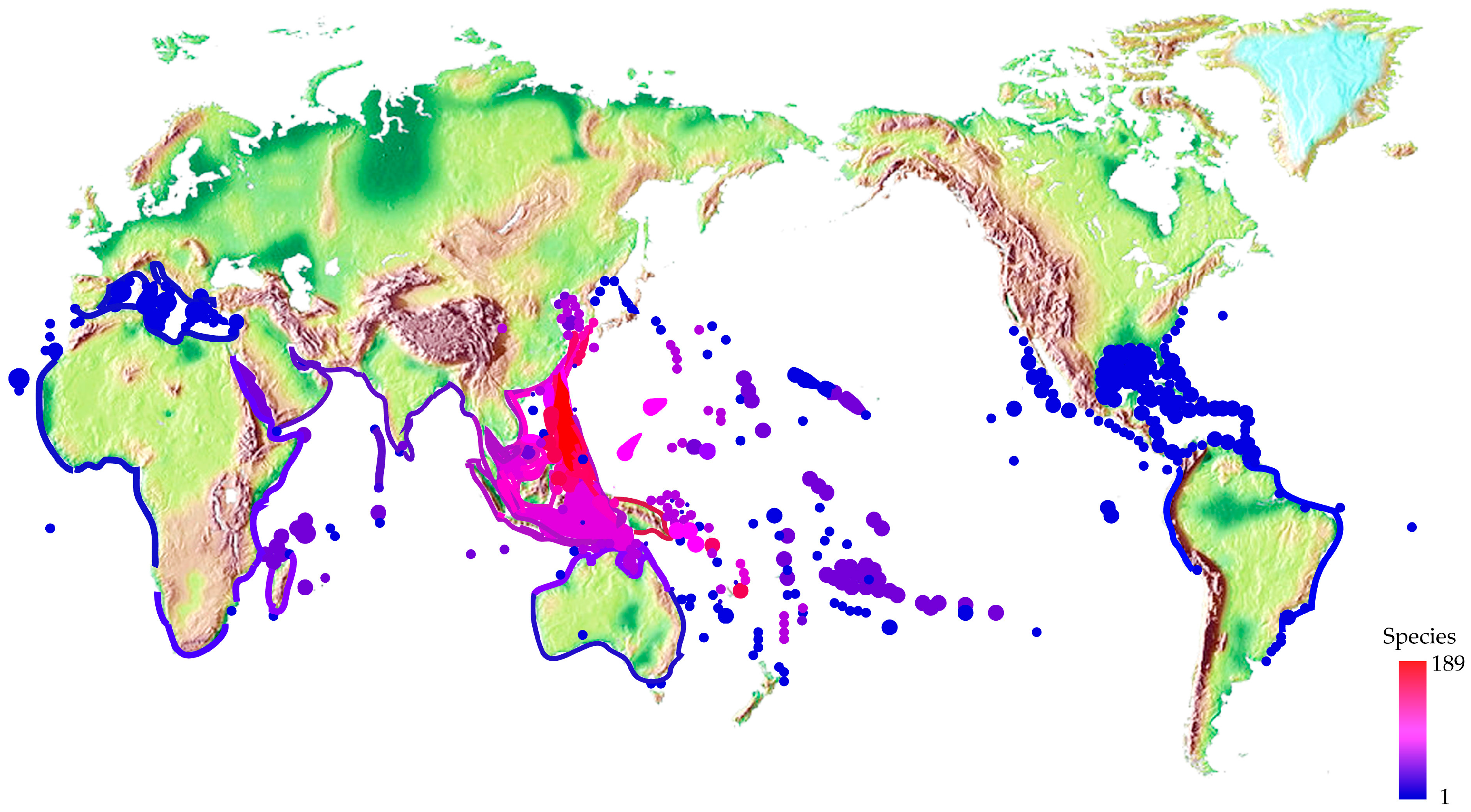
2.3. Different Distribution and Ecology
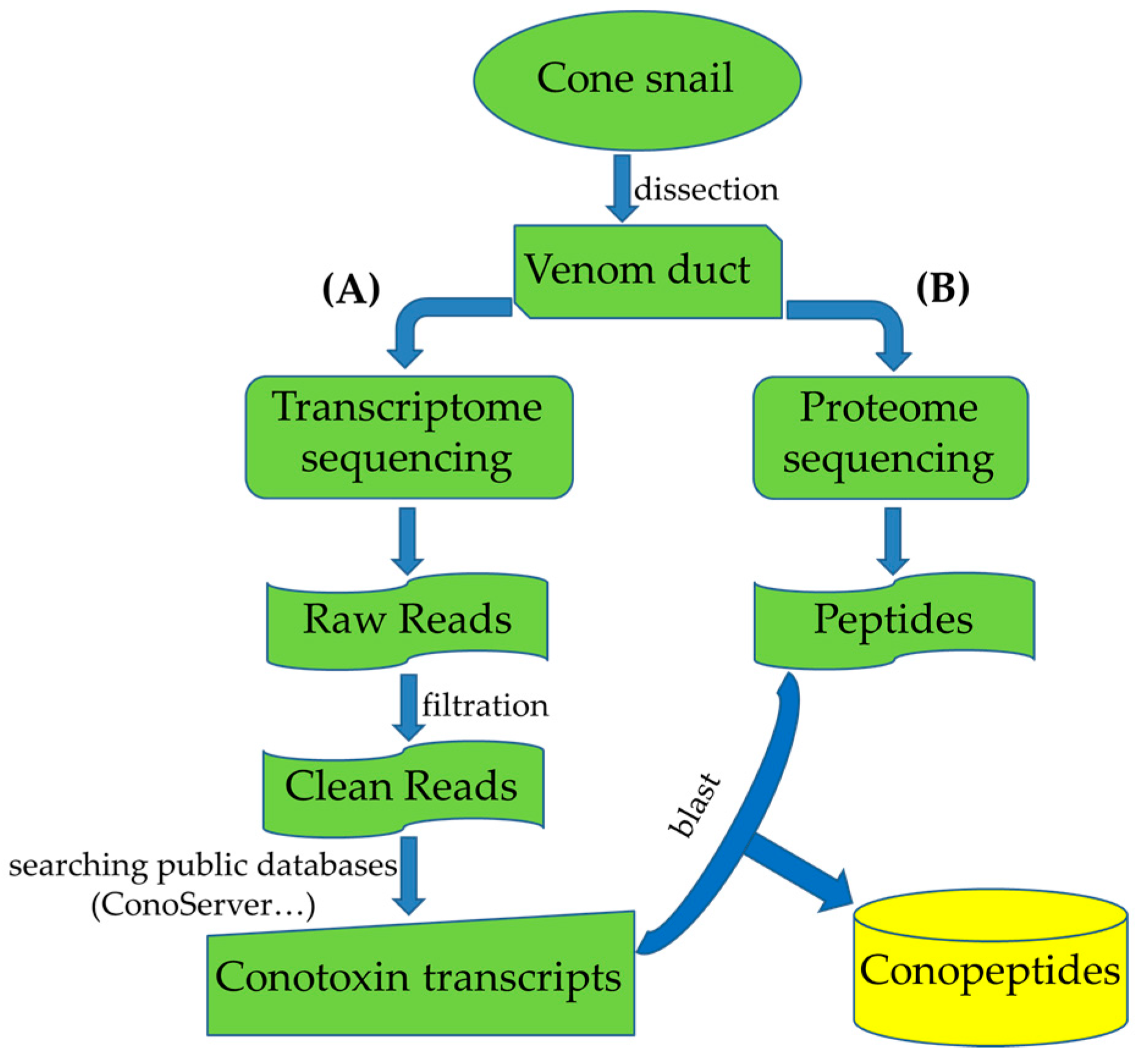
3. Multi-Omics Sequencing for High-Throughput Identification of New Conotoxins
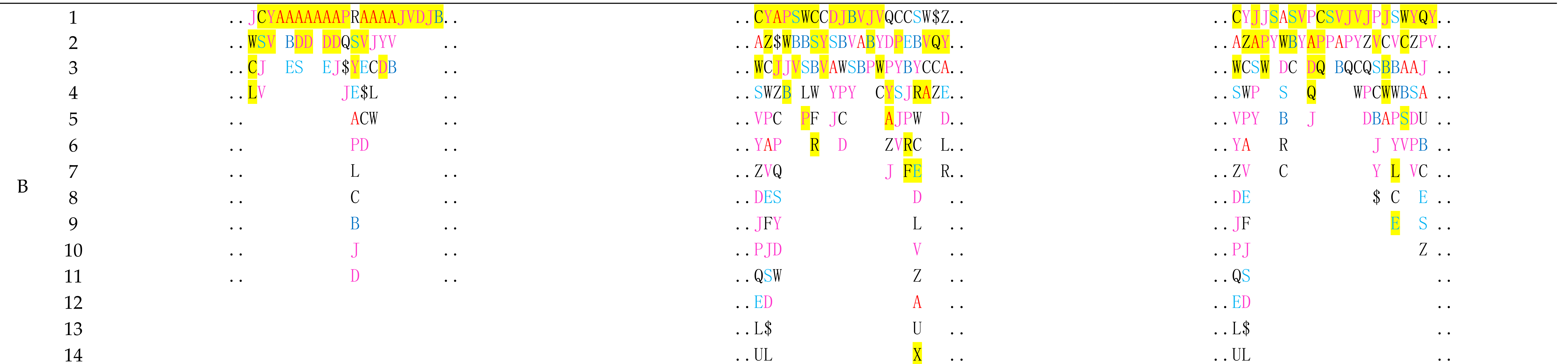
4. Structural Prediction with Protein Structure Fingerprinting for Novel Conformations
5. Recent Advances in Conotoxins for Drug Development
6. Conclusive Remarks
Acknowledgments
Author Contributions
Conflicts of Interest
References
- Prashanth, J.R.; Dutertre, S.; Jin, A.H.; Lavergne, V.; Hamilton, B.; Cardoso, F.C.; Griffin, J.; Venter, D.J.; Alewood, P.F.; Lewis, R.J. The role of defensive ecological interactions in the evolution of conotoxins. Mol. Ecol. 2016, 25, 598–615. [Google Scholar] [CrossRef] [PubMed]
- Himaya, S.W.; Jin, A.H.; Dutertre, S.; Giacomotto, J.; Mohialdeen, H.; Vetter, I.; Alewood, P.F.; Lewis, R.J. Comparative Venomics Reveals the Complex Prey Capture Strategy of the Piscivorous Cone Snail Conus catus. J. Proteom. Res. 2015, 14, 4372–4381. [Google Scholar] [CrossRef] [PubMed]
- Lewis, R.J.; Dutertre, S.; Vetter, I.; Christie, M.J. Conus venom peptide pharmacology. Pharmacol. Rev. 2012, 64, 259–298. [Google Scholar] [CrossRef] [PubMed]
- Kumar, P.S.; Kumar, D.S.; Umamaheswari, S. A perspective on toxicology of Conus venom peptides. Asian Pac. J. Trop. Med. 2015, 8, 337–351. [Google Scholar] [CrossRef]
- Prashanth, J.R.; Brust, A.; Jin, A.H.; Alewood, P.F.; Dutertre, S.; Lewis, R.J. Cone snail venomics: From novel biology to novel therapeutics. Future Med. Chem. 2014, 6, 1659–1675. [Google Scholar] [CrossRef] [PubMed]
- Vetter, I.; Lewis, R.J. Therapeutic potential of cone snail venom peptides (conopeptides). Curr. Top. Med. Chem. 2012, 12, 1546–1552. [Google Scholar] [CrossRef] [PubMed]
- Peng, C.; Yao, G.; Gao, B.M.; Fan, C.X.; Bian, C.; Wang, J.; Cao, Y.; Wen, B.; Zhu, Y.; Ruan, Z.; et al. High-throughput identification of novel conotoxins from the Chinese tubular cone snail (Conus betulinus) by multi-transcriptome sequencing. Gigascience 2016, 5, 17. [Google Scholar] [CrossRef] [PubMed]
- Barghi, N.; Concepcion, G.P.; Olivera, B.M.; Lluisma, A.O. High conopeptide diversity in Conus tribblei revealed through analysis of venom duct transcriptome using two high-throughput sequencing platforms. Mar. Biotechnol. 2015, 17, 81–98. [Google Scholar] [CrossRef] [PubMed]
- Dutertre, S.; Jin, A.H.; Kaas, Q.; Jones, A.; Alewood, P.F.; Lewis, R.J. Deep venomics reveals the mechanism for expanded peptide diversity in cone snail venom. Mol. Cell. Proteom. 2013, 12, 312–329. [Google Scholar] [CrossRef] [PubMed]
- Robinson, S.D.; Safavi-Hemami, H.; McIntosh, L.D.; Purcell, A.W.; Norton, R.S.; Papenfuss, A.T. Diversity of conotoxin gene superfamilies in the venomous snail, Conus victoriae. PLoS ONE 2014, 9, e87648. [Google Scholar] [CrossRef] [PubMed]
- Lewis, R.J.; Garcia, M.L. Therapeutic potential of venom peptides. Nat. Rev. Drug Discov. 2003, 2, 790–802. [Google Scholar] [CrossRef] [PubMed]
- Rigo, F.K.; Dalmolin, G.D.; Trevisan, G.; Tonello, R.; Silva, M.A.; Rossato, M.F.; Klafke, J.Z.; Cordeiro Mdo, N.; Castro Junior, C.J.; Montijo, D.; et al. Effect of omega-conotoxin MVIIA and Phalpha1beta on paclitaxel-induced acute and chronic pain. Pharmacol. Biochem. Behav. 2013, 114, 16–22. [Google Scholar] [CrossRef] [PubMed]
- Eisapoor, S.S.; Jamili, S.; Shahbazzadeh, D.; Ghavam Mostafavi, P.; Pooshang Bagheri, K. A New, High Yield, Rapid, and Cost-Effective Protocol to Deprotection of Cysteine-Rich Conopeptide, Omega-Conotoxin MVIIA. Chem. Biol. Drug Des. 2016, 87, 687–693. [Google Scholar] [CrossRef] [PubMed]
- Han, T.S.; Teichert, R.W.; Olivera, B.M.; Bulaj, G. Conus venoms-a rich source of peptide-based therapeutics. Curr. Pharm. Des. 2008, 14, 2462–2479. [Google Scholar] [CrossRef] [PubMed]
- Olivera, B.M.; Teichert, R.W. Diversity of the neurotoxic Conus peptides: A model for concerted pharmacological discovery. Mol. Interv. 2007, 7, 251–260. [Google Scholar] [CrossRef] [PubMed]
- Fedosov, A.E.; Moshkovskii, S.A.; Kuznetsova, K.G.; Olivera, B.M. Conotoxins: From the biodiversity of gastropods to new drugs. Biomed. Khim. 2013, 59, 267–294. [Google Scholar] [CrossRef] [PubMed]
- Fallon, P.J. Taxonomic review of tropical western Atlantic shallow water Drilliidae (Mollusca: Gastropoda: Conoidea) including descriptions of 100 new species. Zootaxa 2016, 4090, 1–363. [Google Scholar] [CrossRef] [PubMed]
- Marshall, J.; Kelley, W.P.; Rubakhin, S.S.; Bingham, J.P.; Sweedler, J.V.; Gilly, W.F. Anatomical correlates of venom production in Conus californicus. Biol. Bull. 2002, 203, 27–41. [Google Scholar] [CrossRef] [PubMed]
- Chen, P.W.; Hsiao, S.T.; Huang, C.W.; Chen, K.S.; Tseng, C.T.; Wu, W.L.; Hwang, D.F. The complete mitochondrial genome of Conus tulipa (Neogastropoda: Conidae). Mitochondrial DNA Part A 2016, 27, 2738–2739. [Google Scholar]
- Bandyopadhyay, P.K.; Stevenson, B.J.; Ownby, J.P.; Cady, M.T.; Watkins, M.; Olivera, B.M. The mitochondrial genome of Conus textile, coxI-coxII intergenic sequences and Conoidean evolution. Mol. Phylogenet. Evol. 2008, 46, 215–223. [Google Scholar] [CrossRef] [PubMed]
- Barghi, N.; Concepcion, G.P.; Olivera, B.M.; Lluisma, A.O. Characterization of the complete mitochondrial genome of Conus tribblei Walls, 1977. Mitochondrial DNA Part A 2016, 27, 4451–4452. [Google Scholar] [CrossRef] [PubMed]
- Cunha, R.L.; Grande, C.; Zardoya, R. Neogastropod phylogenetic relationships based on entire mitochondrial genomes. BMC Evol. Biol. 2009, 9, 210. [Google Scholar] [CrossRef] [PubMed]
- Chen, P.W.; Hsiao, S.T.; Chen, K.S.; Tseng, C.T.; Wu, W.L.; Hwang, D.F. The complete mitochondrial genome of Conus capitaneus (Neogastropoda: Conidae). Mitochondrial DNA Part B 2016, 1, 520–521. [Google Scholar] [CrossRef]
- Boore, J.L. Animal mitochondrial genomes. Nucleic Acids Res. 1999, 27, 1767–1780. [Google Scholar] [CrossRef] [PubMed]
- Pastukh, V.M.; Gorodnya, O.M.; Gillespie, M.N.; Ruchko, M.V. Regulation of mitochondrial genome replication by hypoxia: The role of DNA oxidation in D-loop region. Free Radic. Biol. Med. 2016, 96, 78–88. [Google Scholar] [CrossRef] [PubMed]
- Remigio, E.A.; Duda, T.F., Jr. Evolution of ecological specialization and venom of a predatory marine gastropod. Mol. Ecol. 2008, 17, 1156–1162. [Google Scholar] [CrossRef] [PubMed]
- Kohn, A.J. Maximal species richness in Conus: Diversity, diet and habitat on reefs of northeast Papua New Guinea. Coral Reefs 2001, 20, 25–38. [Google Scholar]
- Duda, T. Origins of diverse feeding ecologies within Conus, a genus of venomous marine gastropods. Biol. J. Linn. Soc. 2001, 73, 391–409. [Google Scholar] [CrossRef]
- Puillandre, N.; Bouchet, P.; Duda, T.F., Jr.; Kauferstein, S.; Kohn, A.J.; Olivera, B.M.; Watkins, M.; Meyer, C. Molecular phylogeny and evolution of the cone snails (Gastropoda, Conoidea). Mol. Phylogenet. Evol. 2014, 78, 290–303. [Google Scholar] [CrossRef] [PubMed]
- Guindon, S.; Gascuel, O. A simple, fast, and accurate algorithm to estimate large phylogenies by maximum likelihood. Syst. Boil. 2003, 52, 696–704. [Google Scholar] [CrossRef]
- Olivera, B.M.; Seger, J.; Horvath, M.P.; Fedosov, A.E. Prey-Capture Strategies of Fish-Hunting Cone Snails: Behavior, Neurobiology and Evolution. Brain Behav. Evol. 2015, 86, 58–74. [Google Scholar] [CrossRef] [PubMed]
- Jin, A.H.; Dutertre, S.; Kaas, Q.; Lavergne, V.; Kubala, P.; Lewis, R.J.; Alewood, P.F. Transcriptomic messiness in the venom duct of Conus miles contributes to conotoxin diversity. Mol. Cell. Proteom. 2013, 12, 3824–3833. [Google Scholar] [CrossRef] [PubMed]
- Robinson, S.D.; Norton, R.S. Conotoxin gene superfamilies. Mar. Drugs 2014, 12, 6058–6101. [Google Scholar] [CrossRef] [PubMed]
- Halai, R.; Craik, D.J. Conotoxins: Natural product drug leads. Nat. Prod. Rep. 2009, 26, 526–536. [Google Scholar] [CrossRef] [PubMed]
- Akondi, K.B.; Muttenthaler, M.; Dutertre, S.; Kaas, Q.; Craik, D.J.; Lewis, R.J.; Alewood, P.F. Discovery, synthesis, and structure-activity relationships of conotoxins. Chem. Rev. 2014, 114, 5815–5847. [Google Scholar] [CrossRef] [PubMed]
- Kohn, A.J. Superfamily conoidea. In Mollusca: The Southern Synthesis. Fauna of Australia; Beesley, P.L., Ross, G.J.B., Wells, A., Eds.; CSIRO Publishing: Melbourne, Australia, 1998; Volume 5, pp. 846–854. [Google Scholar]
- Briggs, J.C. Marine Zoogeography; McGraw-Hill: New York, NY, USA, 1974. [Google Scholar]
- Kohn, A.J.; Nybakken, J.W. Ecology of Conus on eastern Indian Ocean fringing reefs: Diversity of species and resource utilization. Mar. Biol. 1975, 29, 211–234. [Google Scholar] [CrossRef]
- Kohn, A.J. Tempo and mode of evolution in Conidae. Malacologia 1990, 32, 55–67. [Google Scholar]
- Kohn, A.J. The feeding process in Conus victoriae. In The Marine Flora and Fauna of Dampier, Western Australia; Wells, F.E., Walker, D.I., Jones, D.S., Eds.; Western Australian Museum: Perth, Australia, 2003. [Google Scholar]
- Kohn, A.J. The conidae (Mollusca: Gastropoda) of India. J. Nat. Hist. 1978, 12, 295–335. [Google Scholar] [CrossRef]
- Clench, W.J.; Kondo, Y. The poison cone shell. Am. J. Trop. Med. Hyg. 1943, 23, 105–121. [Google Scholar] [CrossRef]
- Kohn, A.J. Cone shell stings; recent cases of human injury due to venomous marine snails of the genus Conus. Hawaii Med. J. 1958, 17, 528–532. [Google Scholar] [PubMed]
- Endean, R.; Rudkin, C. Studies of the venoms of some Conidae. Toxicon 1963, 1, 49–64. [Google Scholar] [CrossRef]
- Cruz, L.J.; Gray, W.R.; Olivera, B.M. Purification and properties of a myotoxin from Conus geographus venom. Arch. Biochem. Biophys. 1978, 190, 539–548. [Google Scholar] [CrossRef]
- Mullis, K.B.; Faloona, F. Specific synthesis of DNA in vitro via a polymerase catalyzed chain reaction. Meth. Enzymol. 1987, 155, 335–350. [Google Scholar] [PubMed]
- Hillyard, D.R.; Monje, V.D.; Mintz, I.M.; Bean, B.P.; Nadasdi, L.; Ramachandran, J.; Miljanich, G.; Azimi-Zoonooz, A.; McIntosh, J.M.; Cruz, L.J.; et al. A new Conus peptide ligand for mammalian presynaptic Ca2+ channels. Neuron 1992, 9, 69–77. [Google Scholar] [CrossRef]
- Duda, T.F., Jr.; Chang, D.; Lewis, B.D.; Lee, T. Geographic variation in venom allelic composition and diets of the widespread predatory marine gastropod Conus ebraeus. PLoS ONE 2009, 4, e6245. [Google Scholar] [CrossRef] [PubMed]
- Liu, Z.; Xu, N.; Hu, J.; Zhao, C.; Yu, Z.; Dai, Q. Identification of novel I-superfamily conopeptides from several clades of Conus species found in the South China Sea. Peptides 2009, 30, 1782–1787. [Google Scholar] [CrossRef] [PubMed]
- Conticello, S.G.; Gilad, Y.; Avidan, N.; Ben-Asher, E.; Levy, Z.; Fainzilber, M. Mechanisms for evolving hypervariability: The case of conopeptides. Mol. Biol. Evol. 2001, 18, 120–131. [Google Scholar] [CrossRef] [PubMed]
- Ansorge, W.J. Next-generation DNA sequencing techniques. Nat. Biotechnol. 2009, 25, 195–203. [Google Scholar] [CrossRef] [PubMed]
- Hu, H.; Bandyopadhyay, P.K.; Olivera, B.M.; Yandell, M. Characterization of the Conus bullatus genome and its venom-duct transcriptome. BMC Genom. 2011, 12, 60. [Google Scholar] [CrossRef] [PubMed]
- Lavergne, V.; Dutertre, S.; Jin, A.H.; Lewis, R.J.; Taft, R.J.; Alewood, P.F. Systematic interrogation of the Conus marmoreus venom duct transcriptome with ConoSorter reveals 158 novel conotoxins and 13 new gene superfamilies. BMC Genom. 2013, 14, 708. [Google Scholar] [CrossRef] [PubMed]
- Lavergne, V.; Harliwong, I.; Jones, A.; Miller, D.; Taft, R.J.; Alewood, P.F. Optimized deep-targeted proteotranscriptomic profiling reveals unexplored Conus toxin diversity and novel cysteine frameworks. Proc. Natl. Acad. Sci. USA 2015, 112, E3782–E3791. [Google Scholar] [CrossRef] [PubMed]
- Jin, A.H.; Vetter, I.; Himaya, S.W.; Alewood, P.F.; Lewis, R.J.; Dutertre, S. Transcriptome and proteome of Conus planorbis identify the nicotinic receptors as primary target for the defensive venom. Proteomics 2015, 15, 4030–4040. [Google Scholar] [CrossRef] [PubMed]
- Huang, Y.; Peng, C.; Yi, Y.; Gao, B.; Shi, Q. A Transcriptomic Survey of Ion Channel-Based Conotoxins in the Chinese Tubular Cone Snail (Conus betulinus). Mar. Drugs 2017, 15, 228. [Google Scholar] [CrossRef] [PubMed]
- Gao, B.; Peng, C.; Lin, B.; Chen, Q.; Zhang, J.; Shi, Q. Screening and Validation of Highly-Efficient Insecticidal Conotoxins from a Transcriptome-Based Dataset of Chinese Tubular Cone Snail. Toxins 2017, 9, 214. [Google Scholar] [CrossRef] [PubMed]
- Terrat, Y.; Biass, D.; Dutertre, S.; Favreau, P.; Remm, M.; Stöcklin, R.; Piquemal, D.; Ducancel, F. High-resolution picture of a venom gland transcriptome: Case study with the marine snail Conus consors. Toxicon 2012, 59, 34–46. [Google Scholar] [CrossRef] [PubMed]
- Lluisma, A.O.; Milash, B.A.; Moore, B.; Olivera, B.M.; Bandyopadhyay, P.K. Novel venom peptides from the cone snail Conus pulicarius discovered through next-generation sequencing of its venom duct transcriptome. Mar. Genom. 2012, 5, 43–51. [Google Scholar] [CrossRef] [PubMed]
- Hu, H.; Bandyopadhyay, P.K.; Olivera, B.M.; Yandell, M. Elucidation of the molecular envenomation strategy of the cone snail Conus geographus through transcriptome sequencing of its venom duct. BMC Genom. 2012, 13, 284. [Google Scholar] [CrossRef] [PubMed]
- Barghi, N.; Concepcion, G.P.; Olivera, B.M.; Lluisma, A.O. Comparison of the Venom Peptides and Their Expression in Closely Related Conus Species: Insights into Adaptive Post-Speciation Evolution of Conus Exogenomes. Genome Biol. Evol. 2015, 7, 1797–1814. [Google Scholar] [CrossRef] [PubMed]
- Phuong, M.A.; Mahardika, G.N.; Alfaro, M.E. Dietary breadth is positively correlated with venom complexity in cone snails. BMC Genom. 2016, 17, 401. [Google Scholar] [CrossRef] [PubMed]
- Robinson, S.D.; Li, Q.; Lu, A.; Bandyopadhyay, P.K.; Yandell, M.; Olivera, B.M.; Safavi-Hemami, H. The Venom Repertoire of Conus gloriamaris (Chemnitz, 1777), the Glory of the Sea. Mar. Drugs 2017, 15, 145. [Google Scholar] [CrossRef] [PubMed]
- Kaas, Q.; Yu, R.; Jin, A.H.; Dutertre, S.; Craik, D.J. ConoServer: Updated content, knowledge, and discovery tools in the conopeptide database. Nucleic Acids Res. 2012, 40, D325–D330. [Google Scholar] [CrossRef] [PubMed]
- Kaas, Q.; Westermann, J.C.; Halai, R.; Wang, C.K.; Craik, D.J. ConoServer, a database for conopeptide sequences and structures. Bioinformatics 2008, 24, 445–446. [Google Scholar] [CrossRef] [PubMed]
- Yang, J. Comprehensive description of protein structures using protein folding shape code. Proteins 2008, 71, 1497–1518. [Google Scholar] [CrossRef] [PubMed]
- Imperial, J.S.; Bansal, P.S.; Alewood, P.F.; Daly, N.L.; Craik, D.J.; Sporning, A.; Terlau, H.; López-Vera, E.; Bandyopadhyay, P.K.; Olivera, B.M. A novel conotoxin inhibitor of Kv1.6 channel and nAChR subtypes defines a new superfamily of conotoxins. Biochemistry 2006, 45, 8331–8340. [Google Scholar] [CrossRef] [PubMed]
- Park, K.H.; Suk, J.E.; Jacobsen, R.; Gray, W.R.; McIntosh, J.M.; Han, K.H. Solution conformation of alpha-conotoxin EI, a neuromuscular toxin specific for the alpha 1/delta subunit interface of torpedo nicotinic acetylcholine receptor. J. Biol. Chem. 2001, 276, 49028–49033. [Google Scholar] [CrossRef] [PubMed]
- Nielsen, K.J.; Adams, D.; Thomas, L.; Bond, T.; Alewood, P.F.; Craik, D.J.; Lewis, R.J. Structure-activity relationships of omega-conotoxins MVIIA, MVIIC and 14 loop splice hybrids at N and P/Q-type calcium channels. J. Mol. Biol. 1999, 289, 1405–1421. [Google Scholar] [CrossRef] [PubMed]
- Abraham, N.; Healy, M.; Ragnarsson, L.; Brust, A.; Alewood, P.F.; Lewis, R.J. Structural mechanisms for alpha-conotoxin activity at the human alpha 3 beta 4 nicotinic acetylcholine receptor. Sci. Rep. 2017, 7, 45466. [Google Scholar] [CrossRef] [PubMed]
- Nicke, A.; Wonnacott, S.; Lewis, R.J. Alpha-conotoxins as tools for the elucidation of structure and function of neuronal nicotinic acetylcholine receptor subtypes. Eur. J. Biochem. 2004, 271, 2305–2319. [Google Scholar] [CrossRef] [PubMed]
- Leipold, E.; Hansel, A.; Olivera, B.M.; Terlau, H.; Heinemann, S.H. Molecular interaction of delta-conotoxins with voltage-gated sodium channels. FEBS Lett. 2005, 579, 3881–3884. [Google Scholar] [CrossRef] [PubMed]
- Shon, K.J.; Stocker, M.; Terlau, H.; Stühmer, W.; Jacobsen, R.; Walker, C.; Grilley, M.; Watkins, M.; Hillyard, D.R.; Gray, W.R.; et al. kappa-Conotoxin PVIIA is a peptide inhibiting the shaker K+ channel. J. Biol. Chem. 1998, 273, 33–38. [Google Scholar] [CrossRef] [PubMed]
- Li, R.A.; Tomaselli, G.F. Using the deadly mu-conotoxins as probes of voltage-gated sodium channels. Toxicon 2004, 44, 117–122. [Google Scholar] [CrossRef] [PubMed]
- Nielsen, K.J.; Schroeder, T.; Lewis, R. Structure-activity relationships of omega-conotoxins at N-type voltage-sensitive calcium channels (abstract). J. Mol. Recognit. 2000, 13, 55–70. [Google Scholar] [CrossRef]
- Mir, R.; Karim, S.; Kamal, M.A.; Wilson, C.M.; Mirza, Z. Conotoxins: Structure, Therapeutic Potential and Pharmacological Applications. Curr. Pharm. Des. 2016, 22, 582–589. [Google Scholar] [CrossRef] [PubMed]
- Tosti, E.; Boni, R.; Gallo, A. µ-Conotoxins Modulating Sodium Currents in Pain Perception and Transmission: A Therapeutic Potential. Mar. Drugs 2017, 15, 295. [Google Scholar] [CrossRef] [PubMed]
- SIPO. Available online: http://www.sipo.gov.cn/ (accessed on 15 November 2017).
- USPTO. Available online: https://www.uspto.gov/patent (accessed on 15 November 2017).
- EPO. Available online: http://www.epo.org/ (accessed on 15 November 2017).
- WIPO. Available online: http://www.wipo.int/ (accessed on 15 November 2017).
- Durek, T.; Craik, D.J. Therapeutic conotoxins: A US patent literature survey. Expert Opin. Ther. Pat. 2015, 25, 1159–1173. [Google Scholar] [CrossRef] [PubMed]
- Miljanich, G.P. Ziconotide: Neuronal calcium channel blocker for treating severe chronic pain. Curr. Med. Chem. 2004, 11, 3029–3040. [Google Scholar] [CrossRef] [PubMed]
- Nielsen, C.K.; Lewis, R.J.; Alewood, D.; Drinkwater, R.; Palant, E.; Patterson, M.; Yaksh, T.L.; McCumber, D.; Smith, M.T. Anti-allodynic efficacy of the chi-conopeptide, Xen2174, in rats with neuropathic pain. Pain 2005, 118, 112–124. [Google Scholar] [CrossRef] [PubMed]
- Adams, D.J.; Smith, A.B.; Schroeder, C.I.; Yasuda, T.; Lewis, R.J. Omega-conotoxin CVID inhibits a pharmacologically distinct voltage-sensitive calcium channel associated with transmitter release from preganglionic nerve terminals. J. Biol. Chem. 2003, 278, 4057–4062. [Google Scholar] [CrossRef] [PubMed]
- Craig, A.G.; Norberg, T.; Griffin, D.; Hoeger, C.; Akhtar, M.; Schmidt, K.; Low, W.; Dykert, J.; Richelson, E.; Navarro, V.; et al. Contulakin-G, an O-glycosylated invertebrate neurotensin. J. Biol. Chem. 1999, 274, 13752–13759. [Google Scholar] [CrossRef] [PubMed]
- Malmberg, A.B.; Gilbert, H.; McCabe, R.T.; Basbaum, A.I. Powerful antinociceptive effects of the cone snail venom-derived subtype-selective NMDA receptor antagonists conantokins G and T. Pain 2003, 101, 109–116. [Google Scholar] [CrossRef]
- Satkunanathan, N.; Livett, B.; Gayler, K.; Sandall, D.; Down, J.; Khalil, Z. Alpha-conotoxin Vc1.1 alleviates neuropathic pain and accelerates functional recovery of injured neurones. Brain Res. 2005, 1059, 149–158. [Google Scholar] [CrossRef] [PubMed]
- Lubbers, N.L.; Campbell, T.J.; Polakowski, J.S.; Bulaj, G.; Layer, R.T.; Moore, J.; Gross, G.J.; Cox, B.F. Postischemic administration of CGX-1051, a peptide from cone snail venom, reduces infarct size in both rat and dog models of myocardial ischemia and reperfusion. J. Cardiovasc. Pharmacol. 2005, 46, 141–146. [Google Scholar] [CrossRef] [PubMed]
- Ekberg, J.; Jayamanne, A.; Vaughan, C.W.; Aslan, S.; Thomas, L.; Mould, J.; Drinkwater, R.; Baker, M.D.; Abrahamsen, B.; Wood, J.N.; et al. muO-conotoxin MrVIB selectively blocks Nav1.8 sensory neuron specific sodium channels and chronic pain behavior without motor deficits. Proc. Natl. Acad. Sci. USA 2006, 103, 17030–17035. [Google Scholar] [CrossRef] [PubMed]

  |
| Alpha-Conotoxin in Structure (PDB ID: 5T90 for LsIA) | ||||
| Chain | Start | End | Sequence | PFSC |
| F | 1 | 17 | SGCCSNPACRVNNPNIC | ..AAAJVAAAAAJVA.. |
| Image of LsIA (yellow; PDB ID: 5T90) binding with human α3β4 nicotinic acetylcholine receptor | 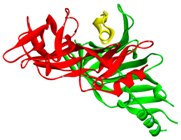 | 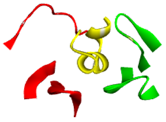 | ||
| Chain A (red); Chain C (green); Conotoxin as Chain F (yellow) | Conotoxin binding site with five fragments | |||
| Binding Fragments of nAChR for Alpha-Conotoxin (PDB ID = 5T90) | ||||
| Chain | Start | End | Sequence | PFSC |
| A | 53 | 57 | WQQTT | RBBEE |
| A | 102 | 116 | LARVVSDGEVLYMPS | BLREWYQSBBEBEWR |
| A | 155 | 165 | TTENSDDSEYF | CYJVAJVAAAP |
| C | 141 | 149 | GSWTHHSRE | WSVAJWZAD |
| C | 183 | 193 | VTYSCCPEAYE | BEWYAJVPREE |
| Clinical Application | Conopeptide | Molecular Target | Clinical Status | Reference |
|---|---|---|---|---|
| Pain | ω-MVIIA (Ziconitide) | Ca2+ channel (CaV2.2) N-type calcium channels/blocker | FDA-approved | [83] |
| Pain | χ-MrIA (Xen2174) | Norepinephrine transporter | Phase IIa * | [84] |
| Pain | ω-CVID (AM336) | Ca2+ channel (CaV2.2) N-type calcium channels/blocker | Phase IIa * | [85] |
| Pain | Contulakin-G (CGX-1160) | Neurotensin receptor | Phase Ib * | [86] |
| Pain/Neuro protection | Conantokin-G (CGX-1007) | NMDA receptor (NR2B) | Preclinical * | [87] |
| Pain | α-Vc1.1 (ACV1) | nAChR (α9α10) | Phase II * | [88] |
| Myocardial infarction | κ-PVIIA (CGX-1051) | K+ channel (KV1) | Preclinical | [89] |
| Neuropathic pain | μO-MrVIB (CGX-1002) | Sodium channels/subtype selective blocker | Preclinical * | [90] |
© 2017 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Gao, B.; Peng, C.; Yang, J.; Yi, Y.; Zhang, J.; Shi, Q. Cone Snails: A Big Store of Conotoxins for Novel Drug Discovery. Toxins 2017, 9, 397. https://doi.org/10.3390/toxins9120397
Gao B, Peng C, Yang J, Yi Y, Zhang J, Shi Q. Cone Snails: A Big Store of Conotoxins for Novel Drug Discovery. Toxins. 2017; 9(12):397. https://doi.org/10.3390/toxins9120397
Chicago/Turabian StyleGao, Bingmiao, Chao Peng, Jiaan Yang, Yunhai Yi, Junqing Zhang, and Qiong Shi. 2017. "Cone Snails: A Big Store of Conotoxins for Novel Drug Discovery" Toxins 9, no. 12: 397. https://doi.org/10.3390/toxins9120397
APA StyleGao, B., Peng, C., Yang, J., Yi, Y., Zhang, J., & Shi, Q. (2017). Cone Snails: A Big Store of Conotoxins for Novel Drug Discovery. Toxins, 9(12), 397. https://doi.org/10.3390/toxins9120397






